Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
|
|
|
- Alvin Gilmore
- 5 years ago
- Views:
Transcription
1 Ligand Type Name 6 Crystallization-grade After D After V3 cocktail Receptor CD4 Resonance Units Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units VRC1 Resonance Units Supplementary Data Set 1 Characterization of purified BG55 SOSIP.664 after negative selection with V3 antibodies. BG55 SOSIP.664 antigenicity before (left panel) and after (middle and right panel) negative selection with V3 antibodies by SPR. CD4- Ig or antibodies were captured on an anti-fc surface, and a nm solution of BG55 SOSIP.664 was used to measure binding. Nature Structural & Molecular Biology: doi:1.138/nsmb.351
2 a ECL units Antigenicity of purified BG55 SOSIP.664 and mutants by MSD-ECLIA 1. X X X X X X X X X X X X 1 5 BG55 SOSIP Y191W Q432P A433P 1C 433C [Trimer], µg/ml $ b VRC1 b12 F15 1.5e 17b 17b+sCD4 PGT121 PGT128 2G12 PGT145 VRC D D+sCD4 PGT ANC195 Temporal Stability of SOSIP, I1C A433C and A433P Retained quaternary structure integrity (fraction) SOSIP 1C 433C A433P c Antigenicity of purified BG55 SOSIP.664 and I1C A433C in absence and presence of CD4 by MSD-ECLIA d Antigenicity of purified BG55 SOSIP.664 and I1C A433C by ELISA ECL units 1. X X X X X X X X 1 5 BG55 SOSIP.664 1C 433C -scd4 +scd4 -scd4 +scd4 OD (45 nm) BG55 SOSIP.664 1C-433C $ VRC1 b12 F15 1.5e 17b 17b+sCD4 PGT121 PGT128 2G12 PGT145 VRC D e 1. X X X X [Trimer], µg/ml Affinity and kinetics of interaction of BG55 SOSIP.664 and 1C 433C by Surface Plasmon Resonance [Antibodies], µg/ml PGT145 PGT151 VRC1 35O22 PGT128 VRC26 8ANC D+sCD4 BG55 SOSIP.664 1C 433C f Affinity and kinetics of interaction of BG55 SOSIP.664 and 1C 433C by Biolayer interferometry VRC26 PGT145 VRC1 35O22 PGT151 b12 F15 17b D BG55 SOSIP.664 1C 433C Supplementary Data Set 2 Antigenicity of purified BG55 SOSIP.664 and selected variants. (a) Antigenicity of BG55 SOSIP.664 and mutants by MSD-ECLIA. Plots show binding of neutralizing (green), non- or weakly neutralizing antibodies (magenta) and antibodies in presence of CD4 (yellow) to BG55 SOSIP.664 and mutants. (b) Temporal stability of BG55 SOSIP.664, A433P and I1C A433C. (c) Comparison of antigenicity of BG55 SOSIP.664 and 1C 433C in absence and presence of soluble, 2-domain CD4 by MSD-ECLIA. Antibodies are color coded as in (a). (d) Antigenicity of BG55 SOSIP.664 and 1C 433C assessed by ELISA. Antibodies are color coded as in (a). (e) Binding of BG55 SOSIP.664 and 1C 433C measured by Surface Plasmon Resonance (SPR) and (f) by Biolayer Interferometry (BLI). Supplementary Table 5 lists kinetics and affinity parameters obtained from SPR and BLI measurements. Supplementary Table 6 lists pairwise comparison statistics between the four methods (MSD-ECLIA, ELISA, SPR and BLI) described above. Nature Structural & Molecular Biology: doi:1.138/nsmb.351
3 a 8 6 Conformational triggering of co-receptor site: 17b detection (-/+ CD4) BG55 SOSIP.664 BG55 SOSIP.664 Q432P BG55 SOSIP.664 A433P BG55 SOSIP.664 I1C A433C +scd4 -scd b Enhanced CD4-induction of P313W mutant:17b detection (+scd4) BG55 SOSIP Conformational triggering of HR2-site: C34 detection (-/+ CD4) BG55 SOSIP.664 BG55 SOSIP.664 Q432P BG55 SOSIP.664 A433P BG55 SOSIP.664 I1C A433C +scd4 +scd4 -scd4 -scd BG55 SOSIP.664 P313W 1 3 c Affinity and Kinetics of 17b binding to activated trimer BG55 SOSIP.664 BG55.SOSIP.664 Q432P BG55.SOSIP.664 A433P BG55.SOSIP.664 I1C A433C Response Units 5 nm to nm 5 nm to nm d Time course of CD4 activation of BG55 SOSIP detection 17b detection P313W BG55 SOSIP.664 I1C A433C P313W BG55 SOSIP.664 I1C A433C Time (8 min -1 hr) Supplementary Data Set 3 CD4-induced activation of HIV-1 Env. (a) SPR analysis of (top panel) co-receptor site exposure and (bottom panel) HR2-site exposure upon CD4 activation. (b) CD4-induced binding of SOSIP and P313W mutant to 17b. Trimer samples at concentrations from nm to 2.5 nm in 2-fold dilutions were combined with nm scd4 and injected on a RU 17b IgG surface. (c) SPR single-cycle kinetics analysis of 17b Fab binding to soluble trimers activated with scd4. Under conditions of constant 5 nm scd4 flow, 17b Fab was injected at incrementally in 2-fold dilutions from 25 nm to 1.5 nm on trimer captured on a 2G12 chip. A433P and I1C A433C were further subjected to 17b Fab concentrations ranging from 5 nm to nm (insets). (d) Time-dependent increase in exposure of V3 epitope (374 detection) and bridging sheet (17b detection) in SOSIP trimers on incubation with scd4. Nature Structural & Molecular Biology: doi:1.138/nsmb.351
4 a b c Molecule BG55 SOSIP.664 N322 BG55 SOSIP.664 1C 433C CD4 d1d2 CD4 d1d4 Mass (19,994 ± 225 Da) x 3 = 329,982 ± 675 Da (19,6 ± 884 Da) x 3 = 327,18 ± 2652 Da,329 ± 131 Da 44,629 ± 167 Da sum of residuals (1-12 ) Fitted stoichiometry CD4:SOSIP d CD4 D1-D2 e CD4 D1-D4 Fit A1 BG55SOSIP.664: Fit A2 BG55SOSIP.664: 1 SOSIP with1 bound CD4 Fit A3 BG55SOSIP.664: 1 SOSIP with 2 bound CD4 Fit A4 BG55SOSIP.664: 1 SOSIP with 3 bound CD4 Fit A1 BG55SOSIP.664: Fit A2 BG55SOSIP.664: 1 bound SOSIP Fit A3 BG55SOSIP.664: 2 bound SOSIP Fit A4 BG55SOSIP.664: 3 bound SOSIP Fit B1 SOSIP 1C 433C : Fit B2 SOSIP 1C 433C : 1 SOSIP 1C 433C Fit B3 SOSIP 1C 433C : 1 SOSIP 1C 433C with 2 bound CD4 Fit B4 SOSIP 1C 433C : 1 SOSIP 1C 433C with 3 bound CD4 Fit B1 SOSIP 1C 433C : Fit B2 SOSIP 1C 433C : 1 SOSIP 1C 433C Fit B3 SOSIP 1C 433C : 1 SOSIP 1C 433C with 2 bound CD4 Fit B4 SOSIP 1C 433C : 1 SOSIP 1C 433C with 3 bound CD4 Supplementary Data Set 4 Analysis of stoichiometry of CD4-binding to BG55 SOSIP.664 or the DS variant, BG55 SOSIP.664 1C 433C. (a) Masses determined for each molecule by MALDI-TOF mass spectrometry. For the BG55 SOSIP molecules, monomer masses were determined, but trimeric masses were used in the analytical ultracentrifugation fitting. (b) Dependence of fit quality as a function of stoichiometry is shown for ultracentrifugation runs performed with four CD4 D1D2 molecules for each trimer. The residual errors are plotted for each. For wild-type trimer, as expected the best fit is at stoichiometry of three CD4 D1D2 molecules per trimer. For the DS mutant, BG55 SOSIP, the residual errors are lowest at 1:1 stoichiometry and fail to fit higher numbers of bound CD4 molecules, indicating 1 CD4 molecule bound per DS trimer. (c) Same as (b), but using CD4 D1- D4, which has a larger mass. Identical results were obtained with best fit at three CD4 D1D4 per trimer for wild-type, and one CD4 D1D4 for the DS mutant. Fitting of the raw sedimentation equilibrium data and residual errors of the fits are shown in panels (d) and (e) for CD4 D1D2 and CD4 D1D4, respectively. Calculations shown were made using the program HeteroAnalysis ( Similar trends were observed in fits with the program SEDPHAT (Vistica et al.). Nature Structural & Molecular Biology: doi:1.138/nsmb.351
5 Relative Light Units Time BG55 (s) BG55 Q432P BG55 A433P BG55 I1C A433C Dilution Dilution Dilution Dilution Supplementary Data Set 5 Virus entry. Ability of HIV-1 BG55 and mutants to enter CD4+CCR5+ cells. TZM-bl cells were infected with pseudoviruses bearing the indicated Env construct. The cells expressed luciferase upon infection; luciferase activity measured in Relative Light Units. Nature Structural & Molecular Biology: doi:1.138/nsmb.351
6 Supplementary Data Set 6 Uptake plots for all observable peptides for ligand-free BG55 SOSIP.664 (WT) (blue), WT+sCD4 (red), ligand-free DS-SOSIP (1C 433C) (cyan), and DS-SOSIP+sCD4 (purple). Error bars show the standard deviation from duplicate measurements. Nature Structural & Molecular Biology: doi:1.138/nsmb.351
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary information. Early development of broad neutralizing antibodies in HIV-1 infected infants
 Supplementary information Early development of broad neutralizing antibodies in HIV-1 infected infants Leslie Goo, Vrasha Chohan, Ruth Nduati, Julie Overbaugh Supplementary Figure 1. Neutralization profile
Supplementary information Early development of broad neutralizing antibodies in HIV-1 infected infants Leslie Goo, Vrasha Chohan, Ruth Nduati, Julie Overbaugh Supplementary Figure 1. Neutralization profile
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) (b) (c) (d) (e) (f) (g) .
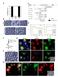 Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Yasmeen et al. Retrovirology 2014, 11:41
 Yasmeen et al. Retrovirology 214, 11:41 RESEARCH Open Access Differential binding of neutralizing and non-neutralizing antibodies to native-like soluble HIV-1 Env trimers, uncleaved Env proteins, and monomeric
Yasmeen et al. Retrovirology 214, 11:41 RESEARCH Open Access Differential binding of neutralizing and non-neutralizing antibodies to native-like soluble HIV-1 Env trimers, uncleaved Env proteins, and monomeric
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Bispecific Fusion Antibodies. Exhibit 100% Breadth and Picomolar Potency. Craig Pace, PhD
 The Aaron Diamond AIDS Research Center Affiliate of The Rockefeller University Bispecific Fusion Antibodies PG9 Ibalizumab & VRC Ibalizumab Exhibit % Breadth and Picomolar Potency Craig Pace, PhD AIDS
The Aaron Diamond AIDS Research Center Affiliate of The Rockefeller University Bispecific Fusion Antibodies PG9 Ibalizumab & VRC Ibalizumab Exhibit % Breadth and Picomolar Potency Craig Pace, PhD AIDS
Surface plasmon resonance (SPR) analysis
 Surface plasmon resonance (SPR) analysis Soluble CD8αα and was manufactured as described previously. 1,2 The C12C heterodimeric TCR was produced using an engineered disulfide bridge between the constant
Surface plasmon resonance (SPR) analysis Soluble CD8αα and was manufactured as described previously. 1,2 The C12C heterodimeric TCR was produced using an engineered disulfide bridge between the constant
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
EXOTESTTM. ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids
 DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Nair S, Branagan AR, Liu J, Boddupalli CS, Mistry PK, Dhodapkar
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Nair S, Branagan AR, Liu J, Boddupalli CS, Mistry PK, Dhodapkar
Identification of Mutation(s) in. Associated with Neutralization Resistance. Miah Blomquist
 Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Supplementary Material
 Supplementary Material HLA-DM Captures Partially Empty HLA-DR Molecules for Catalyzed Peptide Removal Anne-Kathrin Anders, Melissa J. Call, Monika-Sarah E. D. Schulze, Kevin D. Fowler, David A. Schuert,
Supplementary Material HLA-DM Captures Partially Empty HLA-DR Molecules for Catalyzed Peptide Removal Anne-Kathrin Anders, Melissa J. Call, Monika-Sarah E. D. Schulze, Kevin D. Fowler, David A. Schuert,
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen
 Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV. (Summary of the recommendations from an Enterprise Working Group)
 AIDS Vaccine 07, Seattle, August 20-23, 2007 EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV (Summary of the recommendations from an Enterprise Working Group) The Working Group Reston, Virginia,
AIDS Vaccine 07, Seattle, August 20-23, 2007 EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV (Summary of the recommendations from an Enterprise Working Group) The Working Group Reston, Virginia,
Spike Trimer RNA. dsdna
 Spike Trimer RNA dsdna Spike Trimer RNA Spike trimer subunits xxx gp120: receptor and coreceptor binding xxxxxxx gp41: membrane anchoring and target cell fusion dsdna Spike Trimer HIV gp120 binds to host
Spike Trimer RNA dsdna Spike Trimer RNA Spike trimer subunits xxx gp120: receptor and coreceptor binding xxxxxxx gp41: membrane anchoring and target cell fusion dsdna Spike Trimer HIV gp120 binds to host
Measuring interaction complexity and time to equilibrium for biotherapeutics in whole cell assays. Karl Andersson Uppsala University Sweden
 Measuring interaction complexity and time to equilibrium for biotherapeutics in whole cell assays Karl Andersson Uppsala University Sweden Who am I? Affiliated with Uppsala University and Ridgeview Instruments
Measuring interaction complexity and time to equilibrium for biotherapeutics in whole cell assays Karl Andersson Uppsala University Sweden Who am I? Affiliated with Uppsala University and Ridgeview Instruments
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles
 Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1
 Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1 CHRISTOPH RADER, 2 MIKHAIL POPKOV, JOHN A. NEVES, AND CARLOS F. BARBAS III 2 Department of Molecular Biology and The
Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1 CHRISTOPH RADER, 2 MIKHAIL POPKOV, JOHN A. NEVES, AND CARLOS F. BARBAS III 2 Department of Molecular Biology and The
Nature Immunology: doi: /ni.3631
 Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Differential glycosylation of envelope gp120 is associated with differential recognition of HIV-1 by virus-specific antibodies and cell infection
 Raska et al. AIDS Research and Therapy 2014, 11:23 RESEARCH Open Access Differential glycosylation of envelope gp120 is associated with differential recognition of HIV-1 by virus-specific antibodies and
Raska et al. AIDS Research and Therapy 2014, 11:23 RESEARCH Open Access Differential glycosylation of envelope gp120 is associated with differential recognition of HIV-1 by virus-specific antibodies and
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
Supplementary Figure 1: Equilibrium binding analyses of the interaction of efgartigimod with human FcRn
 SULEMENTARY DATA Supplementary Figure 1: Equilibrium binding analyses of the interaction of with human FcRn using SR. The coupling density of was 275 RU. Human FcRn was injected over the flow cells at
SULEMENTARY DATA Supplementary Figure 1: Equilibrium binding analyses of the interaction of with human FcRn using SR. The coupling density of was 275 RU. Human FcRn was injected over the flow cells at
of Nebraska - Lincoln
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 2007 Characterization of Human Immunodeficiency Virus Type 1 Monomeric and
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 2007 Characterization of Human Immunodeficiency Virus Type 1 Monomeric and
Stalking influenza by vaccination with pre-fusion headless HA mini-stem
 Stalking influenza by vaccination with pre-fusion headless HA mini-stem Sophie A Valkenburg a,b, V Vamsee Aditya Mallajosyula c, Olive TW Li b, Alex WH hin b, George arnell d, Nigel Temperton d, Raghavan
Stalking influenza by vaccination with pre-fusion headless HA mini-stem Sophie A Valkenburg a,b, V Vamsee Aditya Mallajosyula c, Olive TW Li b, Alex WH hin b, George arnell d, Nigel Temperton d, Raghavan
IMMUNOBIOLOGY, BIOL 537 Exam # 2 Spring 1997
 Name I. TRUE-FALSE (1 point each) IMMUNOBIOLOGY, BIOL 537 Exam # 2 Spring 1997 Which of the following is TRUE or FALSE relating to immunogenicity of an antigen and T and B cell responsiveness to antigen?
Name I. TRUE-FALSE (1 point each) IMMUNOBIOLOGY, BIOL 537 Exam # 2 Spring 1997 Which of the following is TRUE or FALSE relating to immunogenicity of an antigen and T and B cell responsiveness to antigen?
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
B. SDS-PAGE. Western blot MW (kda) Supplementary Information. Supplementary Figure 1. MW (kda) Glutelin. Prolamin. 3 E.coli. 3 E.
 Supplementary Information Supplementary Figure 1. A. MW (kda) B. SDS-PAGE Western blot MW (kda) 50 40 100 75 50 37 30 Glutelin 25 20 15 20 Prolamin 10 ARP1 1 2 3 E.coli MucoRice ARP1 (RNAi +) 4 5 ARP1
Supplementary Information Supplementary Figure 1. A. MW (kda) B. SDS-PAGE Western blot MW (kda) 50 40 100 75 50 37 30 Glutelin 25 20 15 20 Prolamin 10 ARP1 1 2 3 E.coli MucoRice ARP1 (RNAi +) 4 5 ARP1
Influences on the design and purification of soluble, recombinant native-like HIV-1 envelope glycoprotein trimers
 JVI Accepted Manuscript Posted Online 26 August 2015 J. Virol. doi:10.1128/jvi.01768-15 Copyright 2015, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
JVI Accepted Manuscript Posted Online 26 August 2015 J. Virol. doi:10.1128/jvi.01768-15 Copyright 2015, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
Increased Functional Stability and Homogeneity of Viral Envelope Spikes through Directed Evolution
 Increased Functional Stability and Homogeneity of Viral Envelope Spikes through Directed Evolution Daniel P. Leaman, Michael B. Zwick* Department of Immunology and Microbial Science, The Scripps Research
Increased Functional Stability and Homogeneity of Viral Envelope Spikes through Directed Evolution Daniel P. Leaman, Michael B. Zwick* Department of Immunology and Microbial Science, The Scripps Research
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supporting Information
 Supporting Information Walker et al. 1.173/pnas.111753118 - JR-CSF 1 3 1 4 1 5 1 6 1 7 1 8 blank beads protein A beads JR-FL - 1 3 1 4 1 5 1 6 1 7 1 8 - MGRM-C26 1 3 1 4 1 5 1 6 1 7 1 8 reciprocal serum
Supporting Information Walker et al. 1.173/pnas.111753118 - JR-CSF 1 3 1 4 1 5 1 6 1 7 1 8 blank beads protein A beads JR-FL - 1 3 1 4 1 5 1 6 1 7 1 8 - MGRM-C26 1 3 1 4 1 5 1 6 1 7 1 8 reciprocal serum
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
Antigen Recognition by T cells
 Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Broad and Potent Neutralizing Antibodies from an African Donor Reveal a New HIV-1 Vaccine Target
 Broad and Potent Neutralizing Antibodies from an African Donor Reveal a New HIV-1 Vaccine Target Laura M. Walker, 1 * Sanjay K. Phogat, 2 * Po-Ying Chan-Hui, 3 Denise Wagner, 2 Pham Phung, 4 Julie L. Goss,
Broad and Potent Neutralizing Antibodies from an African Donor Reveal a New HIV-1 Vaccine Target Laura M. Walker, 1 * Sanjay K. Phogat, 2 * Po-Ying Chan-Hui, 3 Denise Wagner, 2 Pham Phung, 4 Julie L. Goss,
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Thesis by. Joshua Simon Klein. In Partial Fulfillment of the Requirements. for the Degree of. Doctor of Philosophy
 INVESTIGATIONS IN THE DESIGN AND CHARACTERIZATION OF HIV-1 NEUTRALIZING MOLECULES Thesis by Joshua Simon Klein In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy California
INVESTIGATIONS IN THE DESIGN AND CHARACTERIZATION OF HIV-1 NEUTRALIZING MOLECULES Thesis by Joshua Simon Klein In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy California
Figure S1. Western blot analysis of clathrin RNA interference in human DCs Human immature DCs were transfected with 100 nm Clathrin SMARTpool or
 Figure S1. Western blot analysis of clathrin RNA interference in human DCs Human immature DCs were transfected with 100 nm Clathrin SMARTpool or control nontargeting sirnas. At 90 hr after transfection,
Figure S1. Western blot analysis of clathrin RNA interference in human DCs Human immature DCs were transfected with 100 nm Clathrin SMARTpool or control nontargeting sirnas. At 90 hr after transfection,
Supplementary Figure 1 Protease allergens induce IgE and IgG1 production. (a-c)
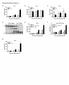 1 Supplementary Figure 1 Protease allergens induce IgE and IgG1 production. (a-c) Serum IgG1 (a), IgM (b) and IgG2 (c) concentrations in response to papain immediately before primary immunization (day
1 Supplementary Figure 1 Protease allergens induce IgE and IgG1 production. (a-c) Serum IgG1 (a), IgM (b) and IgG2 (c) concentrations in response to papain immediately before primary immunization (day
Antigen Receptor Structures October 14, Ram Savan
 Antigen Receptor Structures October 14, 2016 Ram Savan savanram@uw.edu 441 Lecture #8 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Today): Structural features of the BCR and TCR Janeway Chapter
Antigen Receptor Structures October 14, 2016 Ram Savan savanram@uw.edu 441 Lecture #8 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Today): Structural features of the BCR and TCR Janeway Chapter
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Interactions of peptide triazole thiols with Env gp120 induce irreversible breakdown and inactivation of HIV-1 virions
 Interactions of peptide triazole thiols with Env gp120 induce irreversible breakdown and inactivation of HIV-1 virions Bastian et al. Bastian et al. Retrovirology 2013, 10:153 Bastian et al. Retrovirology
Interactions of peptide triazole thiols with Env gp120 induce irreversible breakdown and inactivation of HIV-1 virions Bastian et al. Bastian et al. Retrovirology 2013, 10:153 Bastian et al. Retrovirology
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supporting Information
 Supporting Information Horwitz et al. 73/pnas.35295 A Copies ml - C 3NC7 7 697 698 7 7 73 76-2 2 Days Gp2 residue G458D G459D T278A 7/36 N28 K D 28 459 A28T ID# 697 ID# 698 ID# 7 ID# 7 ID# 73 ID# 76 ID#
Supporting Information Horwitz et al. 73/pnas.35295 A Copies ml - C 3NC7 7 697 698 7 7 73 76-2 2 Days Gp2 residue G458D G459D T278A 7/36 N28 K D 28 459 A28T ID# 697 ID# 698 ID# 7 ID# 7 ID# 73 ID# 76 ID#
Isolation of a Broadly Neutralizing Antibody with Low Somatic Mutation from a Chronically Infected HIV-1 Patient
 Isolation of a Broadly Neutralizing Antibody with Low Somatic Mutation from a Chronically Infected HIV-1 Patient Amanda Fabra García, Carolina Beltrán Pavez, Alberto Merino Mansilla, Cristina Xufré, Isabel
Isolation of a Broadly Neutralizing Antibody with Low Somatic Mutation from a Chronically Infected HIV-1 Patient Amanda Fabra García, Carolina Beltrán Pavez, Alberto Merino Mansilla, Cristina Xufré, Isabel
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69
 Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
Supplementary Information Appendix
 1 Supplementary Information Appendix Supplementary Materials and Methods Construct design Fifteen clade C env genes were assessed by various parameters that appear to correlate with native-like trimer
1 Supplementary Information Appendix Supplementary Materials and Methods Construct design Fifteen clade C env genes were assessed by various parameters that appear to correlate with native-like trimer
SUPPLEMENTARY FIG. S1. MVC inhibition curves in NP2-CD4/CCR5 cells. Luciferase reporter viruses pseudotyped with baseline (black solid lines) and MVC
 Supplementary Data SUPPLEMENTARY FIG. S1. MVC inhibition curves in NP2-CD4/CCR5 cells. Luciferase reporter viruses pseudotyped with baseline (black solid lines) and MVC failure Envs (black dotted lines)
Supplementary Data SUPPLEMENTARY FIG. S1. MVC inhibition curves in NP2-CD4/CCR5 cells. Luciferase reporter viruses pseudotyped with baseline (black solid lines) and MVC failure Envs (black dotted lines)
A V3 Loop-Dependent gp120 Element Disrupted by CD4 Binding Stabilizes the Human Immunodeficiency Virus Envelope Glycoprotein Trimer
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 4-2010 A V3 Loop-Dependent gp120 Element Disrupted by CD4 Binding Stabilizes
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 4-2010 A V3 Loop-Dependent gp120 Element Disrupted by CD4 Binding Stabilizes
Binding Interactions between Soluble HIV Envelope Glycoproteins and Quaternary-Structure-Specific Monoclonal Antibodies PG9 and PG16
 JOURNAL OF VIROLOGY, July 2011, p. 7095 7107 Vol. 85, No. 14 0022-538X/11/$12.00 doi:10.1128/jvi.00411-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Binding Interactions between
JOURNAL OF VIROLOGY, July 2011, p. 7095 7107 Vol. 85, No. 14 0022-538X/11/$12.00 doi:10.1128/jvi.00411-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Binding Interactions between
Biology Open (2014) 000, 1 10 doi: /bio
 (2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
(2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit
 PROTOCOL Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit DESCRIPTION Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit Sufficient materials
PROTOCOL Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit DESCRIPTION Mitochondrial Trifunctional Protein (TFP) Protein Quantity Microplate Assay Kit Sufficient materials
Revised JVI Binding interactions between soluble HIV envelope glycoproteins and quaternarystructure-specific
 JVI Accepts, published online ahead of print on 4 May 2011 J. Virol. doi:10.1128/jvi.00411-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 4 May 2011 J. Virol. doi:10.1128/jvi.00411-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
HIV Anti-HIV Neutralizing Antibodies
 ,**/ The Japanese Society for AIDS Research The Journal of AIDS Research : HIV HIV Anti-HIV Neutralizing Antibodies * Junji SHIBATA and Shuzo MATSUSHITA * Division of Clinical Retrovirology and Infectious
,**/ The Japanese Society for AIDS Research The Journal of AIDS Research : HIV HIV Anti-HIV Neutralizing Antibodies * Junji SHIBATA and Shuzo MATSUSHITA * Division of Clinical Retrovirology and Infectious
Retrovirology. Open Access RESEARCH
 DOI 10.1186/s12977-016-0312-7 Retrovirology RESEARCH Open Access Membrane bound modified form of clade B Env, JRCSF is suitable for immunogen design as it is efficiently cleaved and displays all the broadly
DOI 10.1186/s12977-016-0312-7 Retrovirology RESEARCH Open Access Membrane bound modified form of clade B Env, JRCSF is suitable for immunogen design as it is efficiently cleaved and displays all the broadly
a surface permeabilized
 a surface permeabilized RAW 64.7 P388D1 J774 b CD11b + Ly-6G - Blood Monocytes WT Supplementary Figure 1. Cell surface expression on macrophages and DCs. (a) RAW64.7, P388D1, and J774 cells were subjected
a surface permeabilized RAW 64.7 P388D1 J774 b CD11b + Ly-6G - Blood Monocytes WT Supplementary Figure 1. Cell surface expression on macrophages and DCs. (a) RAW64.7, P388D1, and J774 cells were subjected
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting
 T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
Structural Insights into HIV-1 Neutralization by Broadly Neutralizing Antibodies PG9 and PG16
 Structural Insights into HIV-1 Neutralization by Broadly Neutralizing Antibodies PG9 and PG16 Robert Pejchal, Laura M. Walker, Robyn L. Stanfield, Wayne C. Koff, Sanjay K. Phogat, Pascal Poignard, Dennis
Structural Insights into HIV-1 Neutralization by Broadly Neutralizing Antibodies PG9 and PG16 Robert Pejchal, Laura M. Walker, Robyn L. Stanfield, Wayne C. Koff, Sanjay K. Phogat, Pascal Poignard, Dennis
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/305/ra106/dc1 Supplementary Materials for Controlling Long-Term Signaling: Receptor Dynamics Determine Attenuation and Refractory Behavior of the TGF-β Pathway
www.sciencesignaling.org/cgi/content/full/6/305/ra106/dc1 Supplementary Materials for Controlling Long-Term Signaling: Receptor Dynamics Determine Attenuation and Refractory Behavior of the TGF-β Pathway
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Computational Prediction of Broadly Neutralizing HIV-1 Antibody Epitopes from Neutralization Activity Data
 Computational Prediction of Broadly Neutralizing HIV-1 Antibody Epitopes from Neutralization Activity Data The MIT Faculty has made this article openly available. Please share how this access benefits
Computational Prediction of Broadly Neutralizing HIV-1 Antibody Epitopes from Neutralization Activity Data The MIT Faculty has made this article openly available. Please share how this access benefits
Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes
 Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes Siew Leong Chan Protein Analytics, AbbVie Bioresearch Center Worcester MA Acknowledgement
Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes Siew Leong Chan Protein Analytics, AbbVie Bioresearch Center Worcester MA Acknowledgement
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
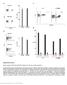 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Modeling Virus- and Antibody-Specific Factors to Predict Human Immunodeficiency Virus Neutralization Efficiency
 Article Modeling Virus- and Antibody-Specific Factors to Predict Human Immunodeficiency Virus Neutralization Efficiency Hillel Haim, 1,2 Ignacio Salas, 1 Kathleen McGee, 1 Noah Eichelberger, 2 Elizabeth
Article Modeling Virus- and Antibody-Specific Factors to Predict Human Immunodeficiency Virus Neutralization Efficiency Hillel Haim, 1,2 Ignacio Salas, 1 Kathleen McGee, 1 Noah Eichelberger, 2 Elizabeth
Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP. ratio
 Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio Mathew Tantama, Juan Ramón Martínez-François, Rebecca Mongeon, Gary Yellen* Department of Neurobiology,
Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio Mathew Tantama, Juan Ramón Martínez-François, Rebecca Mongeon, Gary Yellen* Department of Neurobiology,
SUPPLEMENTARY INFORMATION
 Supplementary Notes 1: accuracy of prediction algorithms for peptide binding affinities to HLA and Mamu alleles For each HLA and Mamu allele we have analyzed the accuracy of four predictive algorithms
Supplementary Notes 1: accuracy of prediction algorithms for peptide binding affinities to HLA and Mamu alleles For each HLA and Mamu allele we have analyzed the accuracy of four predictive algorithms
Nature Methods: doi: /nmeth Supplementary Figure 1. Salipro lipid particles.
 Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
In situ drug-receptor binding kinetics in single cells: a quantitative label-free study of anti-tumor drug resistance
 Supplementary Information for In situ drug-receptor binding kinetics in single cells: a quantitative label-free study of anti-tumor drug resistance Wei Wang 1, Linliang Yin 2,3, Laura Gonzalez-Malerva
Supplementary Information for In situ drug-receptor binding kinetics in single cells: a quantitative label-free study of anti-tumor drug resistance Wei Wang 1, Linliang Yin 2,3, Laura Gonzalez-Malerva
Bead Based Assays for Cytokine Detection
 Bead Based Assays for Cytokine Detection September 27, 2014 6 th EFIS-EJI South East European Immunology School SEEIS 2014 Timisoara, Romania The Cells of the Immune System The Immune Reaction (Th2) (Th1)
Bead Based Assays for Cytokine Detection September 27, 2014 6 th EFIS-EJI South East European Immunology School SEEIS 2014 Timisoara, Romania The Cells of the Immune System The Immune Reaction (Th2) (Th1)
BMDCs were generated in vitro from bone marrow cells cultured in 10 % RPMI supplemented
 Supplemental Materials Figure S1. Cultured BMDCs express CD11c BMDCs were generated in vitro from bone marrow cells cultured in 10 % RPMI supplemented with 15 ng/ml GM-CSF. Media was changed and fresh
Supplemental Materials Figure S1. Cultured BMDCs express CD11c BMDCs were generated in vitro from bone marrow cells cultured in 10 % RPMI supplemented with 15 ng/ml GM-CSF. Media was changed and fresh
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11095 Supplementary Table 1. Summary of the binding between Angptls and various Igdomain containing receptors as determined by flow cytometry analysis. The results were summarized from
doi:10.1038/nature11095 Supplementary Table 1. Summary of the binding between Angptls and various Igdomain containing receptors as determined by flow cytometry analysis. The results were summarized from
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
Supplementary Figures
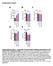 Supplementary Figures a b c d PDI activity in % ERp72 activity in % 4 3 2 1 1 1 ERp activity in % e ΔRFU min -1 1 1 ERp7 activity in % 1 1 Supplementary Figure 1. Selectivity of the bepristat-mediated
Supplementary Figures a b c d PDI activity in % ERp72 activity in % 4 3 2 1 1 1 ERp activity in % e ΔRFU min -1 1 1 ERp7 activity in % 1 1 Supplementary Figure 1. Selectivity of the bepristat-mediated
HIV-1 p24 ANTIGEN CAPTURE ASSAY
 HIV-1 p24 ANTIGEN CAPTURE ASSAY Enzyme Immunoassay for the detection of Human Immunodeficiency Virus Type 1 (HIV-1) p24 in tissue culture media. Catalog # 5421 株式会社東京未来スタイル Tokyo Future Style, Inc 305-0047
HIV-1 p24 ANTIGEN CAPTURE ASSAY Enzyme Immunoassay for the detection of Human Immunodeficiency Virus Type 1 (HIV-1) p24 in tissue culture media. Catalog # 5421 株式会社東京未来スタイル Tokyo Future Style, Inc 305-0047
PFK Activity Assay Kit (Colorimetric)
 PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
Maturation and Diversity of the VRC01-Antibody Lineage over 15 Years of Chronic HIV-1 Infection
 Article Maturation and Diversity of the VRC01-Antibody Lineage over 15 Years of Chronic HIV-1 Infection Graphical Abstract Authors Xueling Wu, Zhenhai Zhang,..., John R. Mascola, Lawrence Shapiro Correspondence
Article Maturation and Diversity of the VRC01-Antibody Lineage over 15 Years of Chronic HIV-1 Infection Graphical Abstract Authors Xueling Wu, Zhenhai Zhang,..., John R. Mascola, Lawrence Shapiro Correspondence
Computational Prediction of Broadly Neutralizing HIV-1 Antibody Epitopes from Neutralization Activity Data
 Computational Prediction of Broadly Neutralizing HIV-1 Antibody Epitopes from Neutralization Activity Data Andrew L. Ferguson 1, Emilia Falkowska 2,3, Laura M. Walker 3, Michael S. Seaman 4, Dennis R.
Computational Prediction of Broadly Neutralizing HIV-1 Antibody Epitopes from Neutralization Activity Data Andrew L. Ferguson 1, Emilia Falkowska 2,3, Laura M. Walker 3, Michael S. Seaman 4, Dennis R.
Supplementary Information for. Inhibitors of Hedgehog Acyltransferase Block Sonic Hedgehog Signaling. Marilyn D. Resh 1,2,6*
 Supplementary Information for Inhibitors of Hedgehog Acyltransferase Block Sonic Hedgehog Signaling Elissaveta Petrova 1,5, Jessica Rios-Esteves 1,2, Ouathek Ouerfelli 3, J. Fraser Glickman 4, and Marilyn
Supplementary Information for Inhibitors of Hedgehog Acyltransferase Block Sonic Hedgehog Signaling Elissaveta Petrova 1,5, Jessica Rios-Esteves 1,2, Ouathek Ouerfelli 3, J. Fraser Glickman 4, and Marilyn
RAISON D ETRE OF THE IMMUNE SYSTEM:
 RAISON D ETRE OF THE IMMUNE SYSTEM: To Distinguish Self from Non-Self Thereby Protecting Us From Our Hostile Environment. Innate Immunity Adaptive Immunity Innate immunity: (Antigen - nonspecific) defense
RAISON D ETRE OF THE IMMUNE SYSTEM: To Distinguish Self from Non-Self Thereby Protecting Us From Our Hostile Environment. Innate Immunity Adaptive Immunity Innate immunity: (Antigen - nonspecific) defense
N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis
 JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
SUPPLEMENTARY INFORMATION. Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was
 SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
BIOASSAYS IN PRODUCT DEVELOPMENT: An Immuno-Oncology Perspective
 BIOASSAYS IN PRODUCT DEVELOPMENT: An Immuno-Oncology Perspective CASSS Bioassays May 09, 2017 Tara Stauffer 1 Overview Introduction: Cell-based bioassays in IO product development Immuno-Oncology (IO)
BIOASSAYS IN PRODUCT DEVELOPMENT: An Immuno-Oncology Perspective CASSS Bioassays May 09, 2017 Tara Stauffer 1 Overview Introduction: Cell-based bioassays in IO product development Immuno-Oncology (IO)
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Nature Biotechnology: doi: /nbt.2354
 a b c Summed up peptide ion signals (arb. u.) 1E+03 INSR 1E+02 1E+01 1E+00-10 - 5 0 5 10 Summed up peptide ion signals (arb. u.) 1E+03 INSR 1E+02 1E+01 1E+00-10 -5 0 5 10 1E+03 1E+02 1E+01 INSR 1E+00-10
a b c Summed up peptide ion signals (arb. u.) 1E+03 INSR 1E+02 1E+01 1E+00-10 - 5 0 5 10 Summed up peptide ion signals (arb. u.) 1E+03 INSR 1E+02 1E+01 1E+00-10 -5 0 5 10 1E+03 1E+02 1E+01 INSR 1E+00-10
Aminoglycoside activity observed on single pre-translocation ribosome complexes
 correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
Insulin (Porcine/Canine) ELISA
 Insulin (Porcine/Canine) ELISA For the quantitative measurement of insulin in Porcine/Canine serum and plasma (EDTA) For Research Use Only. Not For Use In Diagnostic Procedures. Catalog Number: 80-INSPO-E01
Insulin (Porcine/Canine) ELISA For the quantitative measurement of insulin in Porcine/Canine serum and plasma (EDTA) For Research Use Only. Not For Use In Diagnostic Procedures. Catalog Number: 80-INSPO-E01
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Direct Interaction of Ligand-Receptor Pairs Specifying Stomatal Patterning
 Lee_Jin Suk 1 Direct Interaction of Ligand-Receptor Pairs Specifying Stomatal Patterning Jin Suk Lee, Takeshi Kuroha, Marketa Hnilova, Dmitriy Khatayevich, Masahiro M. Kanaoka, Jessica M. McAbee, Mehmet
Lee_Jin Suk 1 Direct Interaction of Ligand-Receptor Pairs Specifying Stomatal Patterning Jin Suk Lee, Takeshi Kuroha, Marketa Hnilova, Dmitriy Khatayevich, Masahiro M. Kanaoka, Jessica M. McAbee, Mehmet
Generation of Robust Antibody Responses to HIV-1 MPER Antigens in Mice Reconstituted with Cultured B cells
 Generation of Robust Antibody Responses to HIV-1 MPER Antigens in Mice Reconstituted with Cultured B cells T. Matt Holl 1, Masayuki Kuraoka 1, Dongmei Liao 1, Laurent Verkoczy 2, M. Anthony Moody 2, Munir
Generation of Robust Antibody Responses to HIV-1 MPER Antigens in Mice Reconstituted with Cultured B cells T. Matt Holl 1, Masayuki Kuraoka 1, Dongmei Liao 1, Laurent Verkoczy 2, M. Anthony Moody 2, Munir
SUPPLEMENTARY INFORMATION
 Supplementary discussion of the role of C05 HCDR1 in antigen binding. Deletion of the 5 amino-acid insertion in HCDR1 del(fgest) and scanning mutagenesis of the tip of the loop had differing effects on
Supplementary discussion of the role of C05 HCDR1 in antigen binding. Deletion of the 5 amino-acid insertion in HCDR1 del(fgest) and scanning mutagenesis of the tip of the loop had differing effects on
