Constructs expressing recombinant proteins were generated by PCR; mutant variants were
|
|
|
- Abel Sanders
- 6 years ago
- Views:
Transcription
1 Cell, Volume 136 Supplemental Data The Ste5 Scaffold Directs Mating Signaling by Catalytically Unlocking the Fus3 MAP Kinase for Activation Matthew Good, Grace Tang, Julie Singleton, Attila Remenyi, and Wendell A. Lim SUPPLEMENTAL EXPERIMENTAL PROCEDURES Protein Expression and Purification (expanded) Constructs expressing recombinant proteins were generated by PCR; mutant variants were generated by two-step PCR, using internal mutagenic primers. Fus3 and Fus3-K42R were expressed at 18 o C in Rosetta(DE3)pLysS E.coli cells (Novagen) using the pbh4 vector to generate an N-terminal His6-tagged, TEV-cleavable protein. Purification was carried out as described previously (Remenyi et al., 2005). Kss1 and Ste7 variants were expressed in Spodoptera frugiperda (SF9) cells, using the Bac-to-Bac Baculoviral Expression System (Invitrogen), at 27 o C. Kss1 was expressed with an N-terminal TEV-cleavable His6 tag, while Ste7 was expressed with N-terminal TEV cleavable GST, and C-terminal His6 tag. Kss1 purification was similar to Fus3, while GST-Ste7 was purified by successive affinity columns, Ni-NTA (Qiagen) and Glutathione Agarose (Sigma). Ste5 variant proteins were expressed at 18 o C in Rosetta(DE3)pLysS E.coli cells as either GST-fusions (petara vector) or MBP-
2 fusions (pmbp-mg vector, modified NEB pmal vector). Proteins were purified using Ni- NTA, Glutathione Agarose beads or Amylose resin (NEB), and dialyzed into Standard Buffer (150mM NaCl, 25mM Tris ph8, 10% Glycerol, 2mM DTT). The Ste5-ms protein used for crystallography was expressed as an N-terminal TEV-cleavable His6 fusion (pbh4 vector) in Rosetta cells, and purified in manner similar to Fus3 (Ni-Nta, TEV cleavage, ResourceQ ion exchange) Structure Determination (expanded) All diffraction data was processed using HKL2000 (Otwinowski and Minor, 1997). Heavy atoms positions were found by SOLVE (Terwilliger and Berendzen, 1999) and initial solvent flattened maps were generated RESOLVE (Terwilliger, 2003). An preliminary automated model was built using ArpWarp6.0 (Morris et al., 2003). Additional model building was done manually with Coot (Emsley and Cowtan, 2004), and TLS refinement was done using Refmac5 (Murshudov et al., 1997) and Phenix (Adams et al., 2002). The cryopreservant for x-ray data collection (not mentioned in the main methods) was made by mixing 12% PEG 3350 and 34% Glycerol 1:1 (vol:vol) with hanging drop solution).
3
4
5
6
7
8
9
10
11
12 Table S1. Constructs used in this study Plasmid Parent Vector Gene N-term. fusion C-term. fusion Expression System vmg500 pbh4 Fus3wt TEV cleavable His6 E. coli vmg510 pbh4 Fus3-K42R TEV cleavable His6 E. coli vmg515 pbh4 Fus3-K42R-(I161L, ) TEV cleavable His6 E. coli vmg530 vmg950 Fus3wt TEV cleavable His6 E. coli vmg531 vmg950 Fus3-I161C TEV cleavable His6 E. coli vmg532 vmg950 Fus3-I161L TEV cleavable His6 E. coli vmg533 vmg950 Fus3-( ) TEV cleavable His6 E. coli vmg534 vmg950 Fus3-( ) TEV cleavable His6 E. coli vmg540 vmg950 Fus3-(I161L, ) TEV cleavable His6 E. coli vmg541 vmg950 Fus3-(I161L, ) TEV cleavable His6 E. coli vmg542 vmg950 Fus3-(I161L, , ) TEV cleavable His6 E. coli vmg543 vmg950 Fus3-( , ) TEV cleavable His6 E. coli vmg544 vmg950 Fus3-( , ) TEV cleavable His6 E. coli vmg550 vmg950 Fus3/Kss1 TEV cleavable His6 E. coli vmg551 vmg950 Kss1/Fus3 TEV cleavable His6 E. coli vmg800 pfbht-b Kss1 TEV cleavable His6 SF9 vmg600 pfb1mg Ste7wt TEV cleavable GST His6 SF9 vmg610 pfb1mg Ste7EE TEV cleavable GST His6 SF9 vmg612 pfb1mg Ste7EE-ND2 TEV cleavable GST His6 SF9 vmg613 pfb1mg Ste7EE-ND1/2 TEV cleavable GST His6 SF9 vmg700 pbh4 Ste5-ms TEV cleavable His6 E. coli vmg705 pbh4 KCK-Ste5ms (C724A) TEV cleavable His6 E. coli vmg720 petara ΔN-Ste5 TEV cleavable GST His6 E. coli vmg711 petara ΔN-Ste5-ND TEV cleavable GST His6 E. coli vmg721 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg723 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg724 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg726 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg733 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg734 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg735 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg738 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg739 petara Ste5 ( ) TEV cleavable GST His6 E. coli vmg750- petara Ste5-ms (mutant variants) TEV cleavable GST His6 E. coli vmg770 [21 variants] vmg775 pmbp Ste5-ms (mutant variants) TEV cleavable MBP His6 E. coli vmg795 [21 variants] pbh4 is a pet derived bacterial expression vector that makes your protein with a TEV-cleavable N-terminal His6 fusion VMG950 is a vector for bicistronic expression of gene of interest with GST-lambda phosphatase (gene is expressed with a TEV-cleavable N-terminal His6 tag) petara is a pet derived bacterial expression that makes your protein with a TEV-cleavable N-terminal GST fusion and C-terminal His6 fusion pmbp is a pet derived bacterial expression that makes your protein with a TEV-cleavable N-terminal MBP fusion and C-terminal His6 fusion pfbht-b is a shuttle vector (pfastbac-ht-b, Invitrogen) for SF9 expression that makes your protein with a TEV-cleavable N-terminal His6 fusion pfb1mg is modified pfastbac1 vector (Invitrogen) for SF9 expression that makes your protein with a TEV-cleavable N-terminal GST fusion and C-terminal His6 * numbers for Fus3 genes denote regions swapped with Kss1 sequence
13 Table S2. Crystallographic Statistics HgCl 2 Data Statistics native λ1 (peak) λ2 (inflection) λ3 (remote) Space Group P P Cell 46.8, 63.6, 69.1, 90, 90, , 63.2, 68.9, 90, 90, 90 Wavelength Å Å Å Å Resolution (last shell) Å ( Å) Å ( Å) Å ( Å) Å ( Å) Unique Reflections Redundancy 4.3 (3.9) 7.4 (7.4) 5.2 (5.1) 7.5 (7.4) Completeness 92.4% (93.9%) 99.8% (99.8%) 99.7% (100%) 99.8% (99.8%) I /σ 19.3 (4.0) 20.8 (5.0) 18.1 (4.4) 20.8 (5.0) R sym (0.311) 0.07 (0.326) (0.334) (0.357) Mean Figure of Merit 0.563; 0.75 after solvent flattening Refinement Statistics Resolution Range Reflections Used Work (Test) (1224) R cryst /R free / Overall Figure of Merit r.m.s.d. bonds/angles 0.005Å / o Average B factor 29.91Å2 Overall B wilson 20.23Å 2 Ramachandran Analysis 94.4.% / 4.2% (Preferred / Allowed) r.m.s.d. is the root-mean squared deviation from ideal geometry R sym = Σ hkl Σ i ΙI hkl,i <I hkl,i >Ι/Σ hkl Σ i ΙI hkl,i Ι R cyst and R free = ΣΙF obs F calc Ι/ΣΙF obs Ι. F obs and F calc are observed and calculated structure factors. R free is calculated from a set of randomly chosen 5% of reflections, and R cyst is calculated with the remaining 95% of reflections.
14 SUPPLEMENTAL REFERENCES Adams, P. D., Grosse-Kunstleve, R. W., Hung, L. W., Ioerger, T. R., McCoy, A. J., Moriarty, N. W., Read, R. J., Sacchettini, J. C., Sauter, N. K., and Terwilliger, T. C. (2002). PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr D Biol Crystallogr 58, Emsley, P., and Cowtan, K. (2004). Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60, Kurtzman, C. P. (2003). Phylogenetic circumscription of Saccharomyces, Kluyveromyces and other members of the Saccharomycetaceae, and the proposal of the new genera Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora. FEMS Yeast Res 4, Morris, R. J., Perrakis, A., and Lamzin, V. S. (2003). ARP/wARP and automatic interpretation of protein electron density maps. Methods Enzymol 374, Murshudov, G. N., Vagin, A. A., and Dodson, E. J. (1997). Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr 53, Otwinowski, Z., and Minor, W. (1997). Processing of X-ray Diffraction Data Collected in Oscillation Mode. In Macromolecular Crystallography, Part A, J. N. Abelson, M. I. Simon, C. W. Carter Jr., and R. M. Sweet, eds. (Academic Press), pp Remenyi, A., Good, M. C., Bhattacharyya, R. P., and Lim, W. A. (2005). The role of docking interactions in mediating signaling input, output, and discrimination in the yeast MAPK network. Mol Cell 20, Scannell, D. R., Frank, A. C., Conant, G. C., Byrne, K. P., Woolfit, M., and Wolfe, K. H. (2007). Independent sorting-out of thousands of duplicated gene pairs in two yeast species descended from a whole-genome duplication. Proc Natl Acad Sci U S A 104, Terwilliger, T. C. (2003). Automated main-chain model building by template matching and iterative fragment extension. Acta Crystallogr D Biol Crystallogr 59, Terwilliger, T. C., and Berendzen, J. (1999). Automated MAD and MIR structure solution. Acta Crystallogr D Biol Crystallogr 55, Tsui, C. K., Daniel, H. M., Robert, V., and Meyer, W. (2008). Re-examining the phylogeny of clinically relevant Candida species and allied genera based on multigene analyses. FEMS Yeast Res 8, Wapinski, I., Pfeffer, A., Friedman, N., and Regev, A. (2007). Natural history and evolutionary principles of gene duplication in fungi. Nature 449,
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.
![172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2. 172R 172K TAM-2/172R TAM-2/172K. AZT concentration [nm] AZT concentration [nm] MgCl 2 2.5K 2.5K 5K 2.5K 5K 2.5K K 5K 2.5K 5K 2.5K 50 2.](/thumbs/82/85138523.jpg) 5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
5 5 5 5 A MgCl 2 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] B 172R 172K TAM-2/172R TAM-2/172K AZT concentration [nm] ATP + ATP - Supplemental Figure 1. Primer extension of HIV-1 RT polymorphisms
Structure and functional analysis of the IGF-II/IGF2R interaction
 Structure and functional analysis of the IGF-II/IGF2R interaction James Brown 1, Carlie Delaine 2, Oliver J. Zaccheo 3, Christian Siebold 1, Robert J. Gilbert 1, Gijs van Boxel 1, Adam Denley 2, John C.
Structure and functional analysis of the IGF-II/IGF2R interaction James Brown 1, Carlie Delaine 2, Oliver J. Zaccheo 3, Christian Siebold 1, Robert J. Gilbert 1, Gijs van Boxel 1, Adam Denley 2, John C.
In situ proteolysis for protein crystallization and structure determination
 In situ proteolysis for protein crystallization and structure determination Aiping Dong, Xiaohui Xu, Aled M Edwards, Midwest Center for Structural Genomics Members & Structural Genomics Consortium Supplementary
In situ proteolysis for protein crystallization and structure determination Aiping Dong, Xiaohui Xu, Aled M Edwards, Midwest Center for Structural Genomics Members & Structural Genomics Consortium Supplementary
Supporting Information
 Supporting Information Lin et al. 10.1073/pnas.1010907107 SI Methods Dehydrosqualene Synthase (CrtM) Expression, Purification, Inhibition and Site-Directed Mutagenesis. CrtM with a histidine tag was overexpressed
Supporting Information Lin et al. 10.1073/pnas.1010907107 SI Methods Dehydrosqualene Synthase (CrtM) Expression, Purification, Inhibition and Site-Directed Mutagenesis. CrtM with a histidine tag was overexpressed
research papers Automated side-chain model building and sequence assignment by template matching 1. Introduction Thomas C.
 Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Automated side-chain model building and sequence assignment by template matching Thomas C. Terwilliger Mail Stop M888, Los Alamos
Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Automated side-chain model building and sequence assignment by template matching Thomas C. Terwilliger Mail Stop M888, Los Alamos
Improving amino-acid identification, fit and C ff prediction using the Simplex method in automated model building
 Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Editors: E. N. Baker and Z. Dauter Improving amino-acid identification, fit and C ff prediction using the Simplex method in automated
Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Editors: E. N. Baker and Z. Dauter Improving amino-acid identification, fit and C ff prediction using the Simplex method in automated
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
KDM2A. Reactions. containing. Reactions
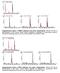 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Characterizing the mesophase behavior of hydrated 9.9 MAG at 20 C and at increasing concentrations of DDM by SAXS.
 Supplementary Figure 1 Characterizing the mesophase behavior of hydrated 9.9 MAG at 20 C and at increasing concentrations of DDM by SAXS. Individual panels show I/2 SAXS profiles at the specified concentration
Supplementary Figure 1 Characterizing the mesophase behavior of hydrated 9.9 MAG at 20 C and at increasing concentrations of DDM by SAXS. Individual panels show I/2 SAXS profiles at the specified concentration
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Cheiron School BL Practice of BL38B1 (Protein Crystallography)
 Cheiron School BL Practice of BL38B1 (Protein Crystallography) Title: Data Collection and S-SAD Phasing of Protein Crystals Staff: Kazuya Hasegawa, Nobuhiro Mizuno (Structural Biology Group, SPring-8/JASRI)
Cheiron School BL Practice of BL38B1 (Protein Crystallography) Title: Data Collection and S-SAD Phasing of Protein Crystals Staff: Kazuya Hasegawa, Nobuhiro Mizuno (Structural Biology Group, SPring-8/JASRI)
Supplementary information. macrophages in foam cells
 Supplementary information The Helicobacter cinaedi antigen CAIP participates in atherosclerotic inflammation by promoting the differentiation of macrophages in foam cells Mario Milco D Elios, Francesca
Supplementary information The Helicobacter cinaedi antigen CAIP participates in atherosclerotic inflammation by promoting the differentiation of macrophages in foam cells Mario Milco D Elios, Francesca
Supplementary Information. A Single Amino Acid Switch Alters the Isoprene Donor Specificity in RiPP Prenyltransferases
 Supplementary Information A Single Amino Acid Switch Alters the Isoprene Donor Specificity in RiPP Prenyltransferases Paola Estrada 1,2,#, Maho Morita 3,#, Yue Hao 1,#,+, Eric W. Schmidt 3,*, and Satish
Supplementary Information A Single Amino Acid Switch Alters the Isoprene Donor Specificity in RiPP Prenyltransferases Paola Estrada 1,2,#, Maho Morita 3,#, Yue Hao 1,#,+, Eric W. Schmidt 3,*, and Satish
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Received 7 May 2009/Accepted 1 July 2009
 JOURNAL OF VIROLOGY, Sept. 2009, p. 9024 9030 Vol. 83, No. 18 0022-538X/09/$08.00 0 doi:10.1128/jvi.00911-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Nucleoside Monophosphate
JOURNAL OF VIROLOGY, Sept. 2009, p. 9024 9030 Vol. 83, No. 18 0022-538X/09/$08.00 0 doi:10.1128/jvi.00911-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Nucleoside Monophosphate
MEK1 Assay Kit 1 Catalog # Lot # 16875
 MEK1 Assay Kit 1 Kit Components Assay Dilution Buffer (ADB), Catalog # 20-108. Three vials, each containing 1.0ml of assay dilution buffer (20mM MOPS, ph 7.2, 25mM ß-glycerol phosphate, 5mM EGTA, 1mM sodium
MEK1 Assay Kit 1 Kit Components Assay Dilution Buffer (ADB), Catalog # 20-108. Three vials, each containing 1.0ml of assay dilution buffer (20mM MOPS, ph 7.2, 25mM ß-glycerol phosphate, 5mM EGTA, 1mM sodium
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Overview of the Expressway Cell-Free Expression Systems. Expressway Mini Cell-Free Expression System
 Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Structural Biology of Membrane Proteins: Are There Any Rules? D.C. Rees Caltech/HHMI NIH Roadmap Meeting
 Structural Biology of Membrane Proteins: Are There Any Rules? D.C. Rees Caltech/HHMI NIH Roadmap Meeting Are there any rules? no magic bullets (shots on goal) proteins that are polydisperse crystallize
Structural Biology of Membrane Proteins: Are There Any Rules? D.C. Rees Caltech/HHMI NIH Roadmap Meeting Are there any rules? no magic bullets (shots on goal) proteins that are polydisperse crystallize
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
Computational Design of an Unnatural Amino Acid Dependent Metalloprotein with Atomic Level Accuracy
 pubs.acs.org/jacs Computational Design of an Unnatural Amino Acid Dependent Metalloprotein with Atomic Level Accuracy Jeremy H. Mills, Sagar D. Khare,,# Jill M. Bolduc, Farhad Forouhar, Vikram Khipple
pubs.acs.org/jacs Computational Design of an Unnatural Amino Acid Dependent Metalloprotein with Atomic Level Accuracy Jeremy H. Mills, Sagar D. Khare,,# Jill M. Bolduc, Farhad Forouhar, Vikram Khipple
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1: Features shared between IL-1β decoy and signaling complex. (a) Superposition of the IL-1β signaling complex with the IL- 1β decoy receptor complex (IL-1β ligands
Supplementary Figures Supplementary Figure 1: Features shared between IL-1β decoy and signaling complex. (a) Superposition of the IL-1β signaling complex with the IL- 1β decoy receptor complex (IL-1β ligands
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Correction Notice: Structural insight into mammalian sialyltransferases
 CORRECTION NOTICE Correction Notice: Structural insight into mammalian sialyltransferases Francesco V Rao, Jamie R Rich, Bojana Rakić, Sai Buddai, Marc F Schwartz, Karl Johnson, Caryn Bowe, Warren W Wakarchuk,
CORRECTION NOTICE Correction Notice: Structural insight into mammalian sialyltransferases Francesco V Rao, Jamie R Rich, Bojana Rakić, Sai Buddai, Marc F Schwartz, Karl Johnson, Caryn Bowe, Warren W Wakarchuk,
Selective protection of an ARF1-GTP signaling axis by a bacterial scaffold induces bidirectional trafficking arrest.
 Selective protection of an ARF1-GTP signaling axis by a bacterial scaffold induces bidirectional trafficking arrest. Andrey S. Selyunin, L. Evan Reddick, Bethany A. Weigele, and Neal M. Alto Supplemental
Selective protection of an ARF1-GTP signaling axis by a bacterial scaffold induces bidirectional trafficking arrest. Andrey S. Selyunin, L. Evan Reddick, Bethany A. Weigele, and Neal M. Alto Supplemental
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
Table S1. Sequence of human and mouse primers used for RT-qPCR measurements.
 Table S1. Sequence of human and mouse primers used for RT-qPCR measurements. Ca9, carbonic anhydrase IX; Ndrg1, N-myc downstream regulated gene 1; L28, ribosomal protein L28; Hif1a, hypoxia inducible factor
Table S1. Sequence of human and mouse primers used for RT-qPCR measurements. Ca9, carbonic anhydrase IX; Ndrg1, N-myc downstream regulated gene 1; L28, ribosomal protein L28; Hif1a, hypoxia inducible factor
BabyBio IMAC columns DATA SHEET DS
 BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Production and crystallization of membrane proteins. So Iwata Imperial College London / Diamond Light Source
 Production and crystallization of membrane proteins So Iwata Imperial College London / Diamond Light Source Membrane Proteins: the Last Frontier in Structural Biology Only ~190 unique membrane protein
Production and crystallization of membrane proteins So Iwata Imperial College London / Diamond Light Source Membrane Proteins: the Last Frontier in Structural Biology Only ~190 unique membrane protein
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Moore s law in information technology. exponential growth!
 Moore s law in information technology exponential growth! ... and how it compares to developments in bio sciences Biology is driven by an avalanche of new information... DNA sequencing methods and throughput
Moore s law in information technology exponential growth! ... and how it compares to developments in bio sciences Biology is driven by an avalanche of new information... DNA sequencing methods and throughput
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Analysis of Cancer Proteins: Pattern Identification on Escherichia coli Species
 Research Article imedpub Journals http://www.imedpub.com Colorectal Cancer: Open Access DOI: 10.21767/2471-9943.100005 Abstract Analysis of Cancer Proteins: Pattern Identification on Escherichia coli Species
Research Article imedpub Journals http://www.imedpub.com Colorectal Cancer: Open Access DOI: 10.21767/2471-9943.100005 Abstract Analysis of Cancer Proteins: Pattern Identification on Escherichia coli Species
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Supplemental Data Functional characterization of core components of the Bacillus subtilis c- di-
 Supplemental Data Functional characterization of core components of the Bacillus subtilis c- di- GMP signaling pathway Xiaohui Gao 1,3, Sampriti Mukherjee 2, Paige M. Matthews 1, Loubna A. Hammad 1, Daniel
Supplemental Data Functional characterization of core components of the Bacillus subtilis c- di- GMP signaling pathway Xiaohui Gao 1,3, Sampriti Mukherjee 2, Paige M. Matthews 1, Loubna A. Hammad 1, Daniel
Center for High-Throughput Structural Biology
 Center for High-Throughput Structural Biology Michael Malkowski, Ph.D. Protein Structure Initiative 5th Bottlenecks Meeting April 13, 2006 Mission To overcome the most significant obstacles to structure
Center for High-Throughput Structural Biology Michael Malkowski, Ph.D. Protein Structure Initiative 5th Bottlenecks Meeting April 13, 2006 Mission To overcome the most significant obstacles to structure
Supplementary Information
 Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
Supplementary Information Using the pimeloyl-coa synthetase adenylation fold to synthesise fatty acid thioesters Menglu Wang 1a ; Lucile Moynié 2a, Peter J. Harrison 1, Van Kelly 1, Andrew Piper 1, James
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
research papers A Rac1 GDP trimer complex binds zinc with tetrahedral and octahedral coordination, displacing magnesium
 Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 A Rac1 GDP trimer complex binds zinc with tetrahedral and octahedral coordination, displacing magnesium Gerd Prehna and C. Erec
Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 A Rac1 GDP trimer complex binds zinc with tetrahedral and octahedral coordination, displacing magnesium Gerd Prehna and C. Erec
Supplementary Figure S1. Supplementary Figure S1. Chemical structure of CH
 Supplementary Figure S1 Supplementary Figure S1. Chemical structure of CH7057288. Supplementary Figure S2 Kinase % competition TRKA 99 TRKC 95 TRKB 92 MEK5 87 AXL 84 CLK1 82 MERTK 82 GCN2 (Kin.Dom.2,S808G)
Supplementary Figure S1 Supplementary Figure S1. Chemical structure of CH7057288. Supplementary Figure S2 Kinase % competition TRKA 99 TRKC 95 TRKB 92 MEK5 87 AXL 84 CLK1 82 MERTK 82 GCN2 (Kin.Dom.2,S808G)
X-ray crystallographic analysis of 72 Data collection A colorless prism crystal of C Br 4 having approximate dimensions of mm was
 Total synthesis of cortistatins A and J Shuji Yamashita, *,a Kentaro Iso, a Kazuki Kitajima, a Masafumi imuro, a and Masahiro irama *,a, b a Department of Chemistry, Graduate School of Science, Tohoku
Total synthesis of cortistatins A and J Shuji Yamashita, *,a Kentaro Iso, a Kazuki Kitajima, a Masafumi imuro, a and Masahiro irama *,a, b a Department of Chemistry, Graduate School of Science, Tohoku
I nfluenza A and B viruses cause the infectious disease influenza, commonly referred to as flu, in the respiratory
 OPEN SUBJECT AREAS: X-RAY CRYSTALLOGRAPHY MOLECULAR BIOPHYSICS INFECTIOUS DISEASES Received 2 June 2014 Accepted 18 July 2014 Published 4 August 2014 Structural analysis of H1N1 and H7N9 influenza A virus
OPEN SUBJECT AREAS: X-RAY CRYSTALLOGRAPHY MOLECULAR BIOPHYSICS INFECTIOUS DISEASES Received 2 June 2014 Accepted 18 July 2014 Published 4 August 2014 Structural analysis of H1N1 and H7N9 influenza A virus
The proteins encoded by the classical HLA class I loci in
 The Journal of Immunology Structural Studies of Allelic Diversity of the MHC Class I Homolog MIC-B, a Stress-Inducible Ligand for the Activating Immunoreceptor NKG2D 1 Margaret A. Holmes, 2 * Pingwei Li,
The Journal of Immunology Structural Studies of Allelic Diversity of the MHC Class I Homolog MIC-B, a Stress-Inducible Ligand for the Activating Immunoreceptor NKG2D 1 Margaret A. Holmes, 2 * Pingwei Li,
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene
 Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for crystallization trials
 Supporting information 1 2 3 Volume 71 (2015) Supporting information for article: 4 5 6 7 8 Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for
Supporting information 1 2 3 Volume 71 (2015) Supporting information for article: 4 5 6 7 8 Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for
SUPPLEMENTAL DATA RESULTS
 SUPPLEMENTAL DATA RESULTS Ded1-mediated ribosomal scanning is less leaky than scanning promoted by eifs 4A/4B/4F. The efficiency of leaky scanning in the presence of Ded1 or of eifs 4A/4B/4F was investigated
SUPPLEMENTAL DATA RESULTS Ded1-mediated ribosomal scanning is less leaky than scanning promoted by eifs 4A/4B/4F. The efficiency of leaky scanning in the presence of Ded1 or of eifs 4A/4B/4F was investigated
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplements. Figure S1. B Phalloidin Alexa488
 Supplements A, DMSO, PP2, PP3 Crk-myc Figure S1. (A) Src kinase activity is necessary for recruitment of Crk to Nephrin cytoplasmic domain. Human podocytes expressing /7-NephrinCD () were treated with
Supplements A, DMSO, PP2, PP3 Crk-myc Figure S1. (A) Src kinase activity is necessary for recruitment of Crk to Nephrin cytoplasmic domain. Human podocytes expressing /7-NephrinCD () were treated with
Supplementary Materials for
 www.sciencemag.org/content/349/6244/187/suppl/dc1 Supplementary Materials for Crystal structure of a mycobacterial Insig homolog provides insight into how these sensors monitor sterol levels Ruobing Ren,
www.sciencemag.org/content/349/6244/187/suppl/dc1 Supplementary Materials for Crystal structure of a mycobacterial Insig homolog provides insight into how these sensors monitor sterol levels Ruobing Ren,
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
Protein Stability Measurements Using Hydrogen/Deuterium Exchange, Mass Spectrometry and NMR Spectroscopy. By Tatiana Borissova
 Protein Stability Measurements Using Hydrogen/Deuterium Exchange, Mass Spectrometry and NMR Spectroscopy By Tatiana Borissova ACKNOWLEDGEMENTS Thank you to: CECILIA DELLA VALLE Dr. Gaetano Montelione Dr.
Protein Stability Measurements Using Hydrogen/Deuterium Exchange, Mass Spectrometry and NMR Spectroscopy By Tatiana Borissova ACKNOWLEDGEMENTS Thank you to: CECILIA DELLA VALLE Dr. Gaetano Montelione Dr.
Workshop on Analysis and prediction of contacts in proteins
 Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
supplementary information
 DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
Probing protein interactions in living cells of Pseudomonas aeruginosa by chemical cross-linking
 Probing protein interactions in living cells of Pseudomonas aeruginosa by chemical cross-linking Arti Navare, Richard Siehnel, Kirsten Beck, Alejandro Wolf-Yadlin, Pradeep Singh, James E. Bruce University
Probing protein interactions in living cells of Pseudomonas aeruginosa by chemical cross-linking Arti Navare, Richard Siehnel, Kirsten Beck, Alejandro Wolf-Yadlin, Pradeep Singh, James E. Bruce University
Biochemical and Biophysical Research Communications 346 (2006) Crystal structure of rat carnitine palmitoyltransferase II (CPT-II)
 Biochemical and Biophysical Research Communications 346 (2006) 974 980 BBRC www.elsevier.com/locate/ybbrc Crystal structure of rat carnitine palmitoyltransferase II (CPT-II) Yu-Shan Hsiao a, Gerwald Jogl
Biochemical and Biophysical Research Communications 346 (2006) 974 980 BBRC www.elsevier.com/locate/ybbrc Crystal structure of rat carnitine palmitoyltransferase II (CPT-II) Yu-Shan Hsiao a, Gerwald Jogl
CHTC Case Studies. Impossible is (almost) nothing Challenging projects in CHTC
 CHTC Case Studies Impossible is (almost) nothing Challenging projects in CHTC Cathrin Weiss UW Madison cweiss@cs.wisc.edu 1 1 Most projects 2 2 Most projects Scientist 2 2 Most projects Scientist compute
CHTC Case Studies Impossible is (almost) nothing Challenging projects in CHTC Cathrin Weiss UW Madison cweiss@cs.wisc.edu 1 1 Most projects 2 2 Most projects Scientist 2 2 Most projects Scientist compute
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
by Micah Johnson, Michael Turner, Elena Poiata, Stephanie Vadasz, Nathan Yardley, and Maria Moreno
 MCDB 201L: Molecular Biology Laboratory Professor: Maria Moreno By submitting this essay, I attest that it is my own work, completed in accordance with University regulations. Micah Johnson Abstract Cloning
MCDB 201L: Molecular Biology Laboratory Professor: Maria Moreno By submitting this essay, I attest that it is my own work, completed in accordance with University regulations. Micah Johnson Abstract Cloning
Advances in membrane-protein crystallization: From detergent-free crystallization to in situ approaches. Dr. Jana Broecker
 Advances in membrane-protein crystallization: From detergent-free crystallization to in situ approaches Dr. Jana Broecker jana.broecker@utoronto.ca 6 th International Symposium on HOS of Protein Therapeutics
Advances in membrane-protein crystallization: From detergent-free crystallization to in situ approaches Dr. Jana Broecker jana.broecker@utoronto.ca 6 th International Symposium on HOS of Protein Therapeutics
Strongly Fluorescent Quaternary Cu-In-Zn-S Nanocrystals Prepared from Cu 1-x InS 2 Nanocrystals by Partial Cation Exchange
 SUPPORTING INFORMATION FOR: Strongly Fluorescent Quaternary Cu-In-Zn-S Nanocrystals Prepared from Cu 1-x InS 2 Nanocrystals by Partial Cation Exchange Inductively Coupled Plasma and emission analysis In:Cu
SUPPORTING INFORMATION FOR: Strongly Fluorescent Quaternary Cu-In-Zn-S Nanocrystals Prepared from Cu 1-x InS 2 Nanocrystals by Partial Cation Exchange Inductively Coupled Plasma and emission analysis In:Cu
Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa
 SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
SUPPLEMENTARY FIGURES Figure S1. HP1α localizes to centromeres in mitosis and interacts with INCENP. (A&B) HeLa tet-on cells that stably express HP1α-CFP, HP1β-CFP, or HP1γ-CFP were monitored with livecell
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Quality control of Saccharomyces yeasts: differentiation of species level and strain grouping using COX 2 gene analysis and MALDI-TOF-MS analysis
 Quality control of Saccharomyces yeasts: differentiation of species level and strain grouping using COX 2 gene analysis and MALDI-TOF-MS analysis Hiroko Kawasaki 1, Shinya Ninomiya 1, Kanae Teramoto 2,
Quality control of Saccharomyces yeasts: differentiation of species level and strain grouping using COX 2 gene analysis and MALDI-TOF-MS analysis Hiroko Kawasaki 1, Shinya Ninomiya 1, Kanae Teramoto 2,
Experimental. Crystal data
 organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 N,N 0 -(Biphenyl-2,2 0 -diyl)bis(furan-2- carboxamide) Chin Hui Kee, Noel F. Thomas, Azhar Ariffin, Khalijah Awang
organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 N,N 0 -(Biphenyl-2,2 0 -diyl)bis(furan-2- carboxamide) Chin Hui Kee, Noel F. Thomas, Azhar Ariffin, Khalijah Awang
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Insulin SEC analysis. Insulin Samples:
 EE1 Insulin SEC analysis Insulin Samples: Lispro, identical in structure to Insulin Human, except that it has lysine and proline at positions 8 and 9, respectively, of chain B, whereas this sequence is
EE1 Insulin SEC analysis Insulin Samples: Lispro, identical in structure to Insulin Human, except that it has lysine and proline at positions 8 and 9, respectively, of chain B, whereas this sequence is
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supplementary Material
 Supplementary Material HLA-DM Captures Partially Empty HLA-DR Molecules for Catalyzed Peptide Removal Anne-Kathrin Anders, Melissa J. Call, Monika-Sarah E. D. Schulze, Kevin D. Fowler, David A. Schuert,
Supplementary Material HLA-DM Captures Partially Empty HLA-DR Molecules for Catalyzed Peptide Removal Anne-Kathrin Anders, Melissa J. Call, Monika-Sarah E. D. Schulze, Kevin D. Fowler, David A. Schuert,
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/20326 holds various files of this Leiden University dissertation. Author: Somireddy Venkata, Bharat Kumar Reddy Title: Dynamics of protein-protein interactions
Cover Page The handle http://hdl.handle.net/1887/20326 holds various files of this Leiden University dissertation. Author: Somireddy Venkata, Bharat Kumar Reddy Title: Dynamics of protein-protein interactions
Supplementary Figure 1
 Supplementary Figure 1 Isolation of mt-trnas and RNA-MS analysis of mt-trna Asn from M. nudus (a)m. nudus mt-trnas were isolated by RCC and resolved by 10% denaturing PAGE. The gel was stained with SYBR
Supplementary Figure 1 Isolation of mt-trnas and RNA-MS analysis of mt-trna Asn from M. nudus (a)m. nudus mt-trnas were isolated by RCC and resolved by 10% denaturing PAGE. The gel was stained with SYBR
Glycosylation analyses of recombinant proteins by LC-ESI mass spectrometry
 Glycosylation analyses of recombinant proteins by LC-ESI mass spectrometry Dr Malin Bäckström Mammalian Protein Expression Core Facility P4EU meeting Porto Nov 11-12, 2013 MPE - A tissue culture facility
Glycosylation analyses of recombinant proteins by LC-ESI mass spectrometry Dr Malin Bäckström Mammalian Protein Expression Core Facility P4EU meeting Porto Nov 11-12, 2013 MPE - A tissue culture facility
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
Serial Femtosecond Crystallography
 Serial Femtosecond Crystallography technical journal club 02.06.2015 Manuela Pfammatter outline principle of x-ray free electron laser (XFEL) serial femtosecond crystallography (SFX) Chapman et al., Nature,
Serial Femtosecond Crystallography technical journal club 02.06.2015 Manuela Pfammatter outline principle of x-ray free electron laser (XFEL) serial femtosecond crystallography (SFX) Chapman et al., Nature,
Data collection. Refinement. R[F 2 >2(F 2 )] = wr(f 2 ) = S = reflections 326 parameters 2 restraints
![Data collection. Refinement. R[F 2 >2(F 2 )] = wr(f 2 ) = S = reflections 326 parameters 2 restraints Data collection. Refinement. R[F 2 >2(F 2 )] = wr(f 2 ) = S = reflections 326 parameters 2 restraints](/thumbs/93/111285625.jpg) organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 2-Chloro-N-(2,3-dichlorophenyl)- benzamide B. Thimme Gowda, a * Sabine Foro, b B. P. Sowmya a and Hartmut Fuess
organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 2-Chloro-N-(2,3-dichlorophenyl)- benzamide B. Thimme Gowda, a * Sabine Foro, b B. P. Sowmya a and Hartmut Fuess
Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag
 Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag Jonathan A. Brain Galina Gulis, Ph.D. 1 Kevin E. Redding, Ph.D. 2 Associate Professor of Chemistry Adjunct
Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag Jonathan A. Brain Galina Gulis, Ph.D. 1 Kevin E. Redding, Ph.D. 2 Associate Professor of Chemistry Adjunct
Purification of Glucagon3 Interleukin-2 Fusion Protein Derived from E. coli
 Purification of Glucagon3 Interleukin-2 Fusion Protein Derived from E. coli Hye Soon Won Dept. of Chem. Eng. Chungnam National University INTRODUCTION Human interleukin-2(hil-2) - known as T Cell Growth
Purification of Glucagon3 Interleukin-2 Fusion Protein Derived from E. coli Hye Soon Won Dept. of Chem. Eng. Chungnam National University INTRODUCTION Human interleukin-2(hil-2) - known as T Cell Growth
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Designing successful membrane protein crystallisation screens: how we designed MemGold. Simon Newstead
 Designing successful membrane protein crystallisation screens: how we designed MemGold Simon Newstead Barriers to structure X-ray diffraction is currently the most successful method Requires good crystals
Designing successful membrane protein crystallisation screens: how we designed MemGold Simon Newstead Barriers to structure X-ray diffraction is currently the most successful method Requires good crystals
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
SUPPLEMENTAL INFORMATION. Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays
 SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
Structural Impact of the Leukemia Drug 1- -D-Arabinofuranosylcytosine (Ara-C) on the Covalent Human Topoisomerase I-DNA Complex*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 14, Issue of April 4, pp. 12461 12466, 2003 Printed in U.S.A. Structural Impact of the Leukemia Drug 1- -D-Arabinofuranosylcytosine (Ara-C) on the Covalent
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 14, Issue of April 4, pp. 12461 12466, 2003 Printed in U.S.A. Structural Impact of the Leukemia Drug 1- -D-Arabinofuranosylcytosine (Ara-C) on the Covalent
Structural Determinants for Improved Stability of Designed Ankyrin Repeat Proteins with a Redesigned C-Capping Module
 doi:10.1016/j.jmb.2010.09.023 J. Mol. Biol. (2010) 404, 381 391 Contents lists available at www.sciencedirect.com Journal of Molecular Biology journal homepage: http://ees.elsevier.com.jmb Structural Determinants
doi:10.1016/j.jmb.2010.09.023 J. Mol. Biol. (2010) 404, 381 391 Contents lists available at www.sciencedirect.com Journal of Molecular Biology journal homepage: http://ees.elsevier.com.jmb Structural Determinants
2-Methyl-4-nitro-1-(4-nitrophenyl)-1H-imidazole
 University of Wollongong Research Online Australian Institute for Innovative Materials - Papers Australian Institute for Innovative Materials 2007 2-Methyl-4-nitro-1-(4-nitrophenyl)-1H-imidazole Pawel
University of Wollongong Research Online Australian Institute for Innovative Materials - Papers Australian Institute for Innovative Materials 2007 2-Methyl-4-nitro-1-(4-nitrophenyl)-1H-imidazole Pawel
X-ray Structures of Isopentenyl Phosphate Kinase
 ARTICLE X-ray Structures of Isopentenyl Phosphate Kinase Mark F. Mabanglo, Heidi L. Schubert, Mo Chen,, Christopher P. Hill, and C. Dale Poulter, * Department of Chemistry, University of Utah, 315 South
ARTICLE X-ray Structures of Isopentenyl Phosphate Kinase Mark F. Mabanglo, Heidi L. Schubert, Mo Chen,, Christopher P. Hill, and C. Dale Poulter, * Department of Chemistry, University of Utah, 315 South
Clostridium thermocellum JW-20 Proteins Expressed into Esherichia coli
 Improving Solubility of Shewanella oneidensis MR-1 and Clostridium thermocellum JW-20 Proteins Expressed into Esherichia coli Irina Kataeva,*, Jessie Chang, Hao Xu, Chi-Hao Luan, Jizhong Zhou, Vladimir
Improving Solubility of Shewanella oneidensis MR-1 and Clostridium thermocellum JW-20 Proteins Expressed into Esherichia coli Irina Kataeva,*, Jessie Chang, Hao Xu, Chi-Hao Luan, Jizhong Zhou, Vladimir
How mutations in trna distant from the anticodon affect the fidelity of decoding
 Europe PMC Funders Group Author Manuscript Published in final edited form as: Nat Struct Mol Biol. 2011 April ; 18(4): 432 436. doi:10.1038/nsmb.2003. How mutations in trna distant from the anticodon affect
Europe PMC Funders Group Author Manuscript Published in final edited form as: Nat Struct Mol Biol. 2011 April ; 18(4): 432 436. doi:10.1038/nsmb.2003. How mutations in trna distant from the anticodon affect
ASSESMENT OF CRYOPRESERVATION SYSTEMS INFLUENCE ON THE SURVAVIAL OF E. COLI RECOMBINANT STRAINS
 Lucrări ştiinńifice Zootehnie şi Biotehnologii, vol. 41(1) (2008), Timişoara ASSESMENT OF CRYOPRESERVATION SYSTEMS INFLUENCE ON THE SURVAVIAL OF E. COLI RECOMBINANT STRAINS TESTAREA INFLUENłEI SISTEMELOR
Lucrări ştiinńifice Zootehnie şi Biotehnologii, vol. 41(1) (2008), Timişoara ASSESMENT OF CRYOPRESERVATION SYSTEMS INFLUENCE ON THE SURVAVIAL OF E. COLI RECOMBINANT STRAINS TESTAREA INFLUENłEI SISTEMELOR
Supporting Information. Copyright Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2009
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2009 DOI: 10.1002/cbic.200900584 Supporting Information for Exploring the Substrate Promiscuity of Drug-modifying Enzymes
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2009 DOI: 10.1002/cbic.200900584 Supporting Information for Exploring the Substrate Promiscuity of Drug-modifying Enzymes
Antibodies: LB1 buffer For 50 ml For 10ml For 30 ml Final 1 M HEPES, ph 2.5 ml 0.5 ml 1.5 ml 50mM. 5 M NaCl 1.4 ml 280 µl 0.
 Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
The most common type of oxidative damage identified in
 8-Oxoguanine rearranges the active site of human topoisomerase I Diem-Thu Thieu Lesher*, Yves Pommier, Lance Stewart, and Matthew R. Redinbo* Departments of *Chemistry and Biochemistry and Biophysics,
8-Oxoguanine rearranges the active site of human topoisomerase I Diem-Thu Thieu Lesher*, Yves Pommier, Lance Stewart, and Matthew R. Redinbo* Departments of *Chemistry and Biochemistry and Biophysics,
metal-organic compounds
 metal-organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 Chlorido(2-chloronicotinato)triphenylantimony(V) Li Quan, Handong Yin* and Daqi Wang College of Chemistry
metal-organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 Chlorido(2-chloronicotinato)triphenylantimony(V) Li Quan, Handong Yin* and Daqi Wang College of Chemistry
