Supplementary Table 1. T-cell receptor sequences of HERV-K(HML-2)-specific CD8 + T cell clone.
|
|
|
- Rachel Ray
- 6 years ago
- Views:
Transcription
1 Supplementary Table 1. T-cell receptor sequences of HERV-K(HML-2)-specific CD8 + T cell clone. alpha beta ATGCTCCTGCTGCTCGTCCCAGTGCTCGAGGTGATTTTTACTCTGGGAGGAACCAGAGCC CAGTCGGTGACCCAGCTTGACAGCCACGTCTCTGTCTCTGAAGGAACCCCGGTGCTGCTG AGGTGCAACTACTCATCTTCTTATTCACCATCTCTCTTCTGGTATGTGCAACACCCCAAC AAAGGACTCCAGCTTCTCCTGAAGTACACATCAGCGGCCACCCTGGTTAAAGGCATCAAC GGTTTTGAGGCTGAATTTAAGAAGAGTGAAACCTCCTTCCACCTGACGAAACCCTCAGCC CATATGAGCGACGCGGCTGAGTACTTCTGTGTTGTGAGTACTCTCAAGATCATCTTTGGA AAAGGGACACGACTTCATATTCTCCCCAATATCCAGAACCCTGACCCTGCCGTGTACCAG CTGAGAGACTCTAAATCCAGTGACAAGTCTGTCTGCCTATTCACCGATTTTGATTCTCAA ACAAATGTGTCACAAAGTAAGGATTCTGATGTGTATATCACAGACAAAACTGTGCTAGAC ATGAGGTCTATGGACTTCAAGAGCAACAGTGCTGTGGCCTGGAGCAACAAATCTGACTTT GCATGTGCAAACGCCTTCAACAACAGCATTATTCCAGAAGACACCTTCTTCCCCAGCCCA GAAAGTTCCTGTGATGTCAAGCTGGTCGAGAAAAGCTTTGAAACAGATACGAACCTAAA CTTTCAAAACCTGTCAGTGATTGGGTTCCGAATCCTCCTCCTGAAAGTGGCCGGGTTTAAT CTGCTCATGACGCTGCGGCTGTGGTCCAGCTGA ATGGGCACCAGCCTCCTCTGCTGGATGGCCCTGTGTCTCCTGGGGGCAGATCACGCAGAT ACTGGAGTCTCCCAGGACCCCAGACACAAGATCACAAAGAGGGGACAGAATGTAACTTT CAGGTGTGATCCAATTTCTGAACACAACCGCCTTTATTGGTACCGACAGACCCTGGGGCA GGGCCCAGAGTTTCTGACTTACTTCCAGAATGAAGCTCAACTAGAAAAATCAAGGCTGCT CAGTGATCGGTTCTCTGCAGAGAGGCCTAAGGGATCTTTCTCCACCTTGGAGATCCAGCG CACAGAGCAGGGGGACTCGGCCATGTATCTCTGTGCCAGCAGCATAGGCCCGTCTGAAG CTTTCTTTGGACAAGGCACCAGACTCACAGTTGTAGAGGACCTGAACAAGGTGTTCCCAC CCGAGGTCGCTGTGTTTGAGCCATCAGAAGCAGAGATCTCCCACACCCAAAAGGCCACA CTGGTGTGCCTGGCCACAGGCTTCTTCCCTGACCACGTGGAGCTGAGCTGGTGGGTGAAT GGGAAGGAGGTGCACAGTGGGGTCAGCACGGACCCGCAGCCCCTCAAGGAGCAGCCCGC CCTCAATGACTCCAGATACTGCCTGAGCAGCCGCCTGAGGGTCTCGGCCACCTTCTGGCA GAACCCCCGCAACCACTTCCGCTGTCAAGTCCAGTTCTACGGGCTCTCGGAGAATGACGA GTGGACCCAGGATAGGGCCAAACCCGTCACCCAGATCGTCAGCGCCGAGGCCTGGGGTA GAGCAGACTGTGGCTTTACCTCGGTGTCCTACCAGCAAGGGGTCCTGTCTGCCACCATCC TCTATGAGATCCTGCTAGGGAAGGCCACCCTGTATGCTGTGCTGGTCAGCGCCCTTGTGTT GATG GCCATGGTCAAGAGAAAGGATTTCTGA 1
2 Supplementary Table 2. Results summary from IMGT/V-QUEST tool alpha chain of HERV-K(HML-2)-specific CD8 + T cell clone TCR. Result summary: Productive TRA rearranged sequence (no stop codon and in-frame junction) V-GENE and allele Homsap TRAV8-2*01 F score = 1360 identity = 100,00% (273/273 nt) J-GENE and allele Homsap TRAJ30*01 F score = 209 identity = 91,84% (45/49 nt) FR-IMGT lengths, CDR-IMGT lengths and AA JUNCTION [ ] [6.8.8] CVVSTLKIIF Supplementary Table 3. Results summary from IMGT/V-QUEST tool beta chain of HERV-K(HML-2)-specific CD8 + T cell clone TCR. Result summary: Productive TRB rearranged sequence (no stop codon and in-frame junction) V-GENE and allele Homsap TRBV7-9*03 F score = 1375 identity = 100,00% (276/276 nt) J-GENE and allele Homsap TRBJ1-1*01 F score = 186 identity = 87,50% (42/48 nt) D-GENE and allele by IMGT/JunctionAnalysis Homsap TRBD1*01 F D-REGION is in reading frame 1 FR-IMGT lengths, CDR-IMGT lengths and AA JUNCTION [ ] [5.6.10] CASSIGPSEAFF 2
3 Supplementary Table 4. HIV-1 viruses. Clade/CRF Catalog # Reference ID Full Length Accession Coreceptor A 2521 ELI K03454 X4 A KE_KNH1135 AF47605 R5 A KE_KNH1209 AF R5 A KE_KNH1088 AF R5 A KE_KNH1207 AF47068 R5 B US_33931N AY R5 B _US1 AY R5 B TH_BK132 AY X4 B US_Ba_L AY R5 C ET_14 AY R5 C SM_145 AY R5 C _ET288 AY R5 C US_MSC5016 AY R5 C IN_101 N/A R5 D UG_57128 AF R5 D UG_J32228M4 AF R5 D MB N/A Not tested G 3187 G3 NA R5 CRF02AG CM_1475MV AY R5 CRF02AG DJ_263 AF R5 3
4 A 3512 HIV A N/A Not tested A 3513 HIV K N/A Not tested N/A 3060 SIVhu N/A Not tested N/A 253 SIVmac251 N/A Not tested Supplementary Table 5. Quantitative PCR primers and probes. Forward Primer (5'-3') Reverse Primer (5'-3') Probe (5'-3') HERV-K-HML-2-gag TCGGGAAACGAGCAAAGG GAATTGGGAATGCCCCAGTT 6FAM-CTCAGGCCCCACAAC-MGBNFQ HERV-K-HML-2-pol GGGAATGCTTAATAGTCCAACTATTTG TGAAAACTTGTCTCTAACTGGTTGAAG 6FAM-CAGACTTTTGTAGCTCAAG-MGBNFQ HERV-K-HML-2-env GGGTACCTGGCCCCATAGA CATCATCCCTTCTTCCTCAGGTT 6FAM-ATCGCTGCCCTGCC-MGBNFQ HERV-K-HML-2-rec GTGACACAAACCCCAGAGAGTATG CTGCAGACACCATTGATACAATCA 6FAM-TGCTTGCAGCCTTG-MGBNFQ TBP AAGTTGGGTTTTCCAGCTAAGTTC CATCACAGCTCCCCACCATAT 6FAM-TGGACTTCAAGATTCAG-MGBNFQ PP1A TCTGCCCACCTTACAGACC GATCAAATCCGCCACCTCTA N/A GAPDH TTGACCTCAGCTGCACATTC AGGATGGTCTCGAGTGCTTG N/A β-actin CACAGGGGAGGTGATAGCAT CACGAAGGCTCATCATTCAA N/A Supplementary Results. Additional Characterization of HERV-K(HML-2)-Env-specific T cell clone. Additional Characterization of T-cell Determinant. The EC 50 of the HERV-K(HML-2)-Env-specific T cell clone to the optimal epitope, at approximately 4µM, is considerably higher than that typically observed with virus-specific T cell responses. One possible explanation for this is that this T cell clone is of relatively low avidity. However it is also possible that this response was primarily raised against a distinct HERV-K-Env sequence containing mismatches with the consensus HERV-K(HML-2)-Env CIDSTFNWHQRI sequence. We assembled an alignment of HERV-K-Env sequences and selected the following variants of this sequence: CIDLTFNWQHRI, CIDSTFDWQHHI, CIDSTFDWQHYI, CIDSTSDWQHHI. Peptides corresponding to these sequences were manufactured, but 4
5 each of these completely failed to stimulate the clone (data not shown). This does not represent an exhaustive list of HERV-K variant sequences, however, and we feel that it may be important for future studies to identify precisely which subset of HERV-K insertions are induced upon HIV-1 infection. A third possible explanation for the high EC 50 of the clone recognizes is that it may recognize only a modified version of the determinant that is present at low abundance in our synthetic peptide. It is important to note that we can strictly rule out the possibility that the clone could be recognizing a minor foreign contaminating peptide by virtue of the consistent recognition of 6 different batches of the CIDSTFNWQHRI peptide (from two different manfacturers), by the logical fine-mapping of the epitope where any peptide containing the minimal epitope is recognized while shorter peptides are not, and by the very similar titration curves observed with different peptide preparations (ex. CIDSTFNWQHRILLV vs CIDSTFNWQHRI, Figure 3D). One possibility is that the clone recognizes the native peptide sequence but that this sequence is present at low abundance in the synthetic peptide. It has previously been demonstrated that sulfhydryl modification of cysteine is responsible for reducing the antigenicity of the influenza virus nuclear protein determinants NP and NP and that replacing cysteine residues in these determinants with either serine or alanine resulted in substantial increases in antigenicity (1). Similarly, substitution of the cysteine residue in the lympocytic choriomeningitis virus T cell determinant KAVYNFATC with methionine, to give KAVYNFATM, results in a decrease in the EC 50 from >5µM to 21nM by preventing the formation of cysteine dimers (2). We tested the effect of replacing the N-terminal cysteine in CIDSTFNWQHRI with either alanine, serine methionine or tyrosine 5
6 (AIDSTFNWQHRI, SIDSTFNWQHRI, MIDSTFNWQHRI and YIDSTFNWQHRI) on the antigenicity of the peptide. Substitution with either alanine, serine, or threonine completely abolished recognition by the HERV-K(HML-2)-Env-specific T cell clone, while the methionine-substituted peptide was recognized at a reduced level (data not shown). We further tested the response of the T cell clone to reduced (TCEP treated) CIDSTFNWQHRI peptide as well as oxidized dimeric peptide and observed lesser antigenicity of both of these modified peptides as compared to the untreated peptide (data not shown). Taken together, these data suggest that a modification of the cysteine residue in this peptide may be required for optimal immunogenicity but that this is not simply related to redox state. Next, we tested the possibility that a minor species of the CIDSTFNWQHRILLV peptide bearing an unknown modification is the primary T cell determinant recognized by the HERV-K(HML-2)-Env specific T cell clone, 50mg of crude peptide was divided into 60 fractions by C18 reversed-phase HPLC (JPT Peptide Technologies) and each fraction was tested for its ability to stimulate the T cell clone in IFN-γ ELISPOT assays. The T cell clone responded maximally to fraction 8 with lesser responses to fractions 9-11 and no response observed to later fraction. This overlapped only partially with the elution profile of the native CIDSTFNWQHRILLV peptide (identified by a 923 mw/z peak) that was the primary product in fractions 8 18 (data not shown). The elution of CIDSTFNWQHRILLV peaked at fractions 12 and 13, while the clone did not respond to these fractions (data not shown). These data indicate that the T cell determinant of the HERV-K(HML-2)-Env specific T cell clone is a modified version of CIDSTFNWQHRILLV, which is slightly less hydrophobic than the native peptide (elutes 6
7 earlier). Peptide titration experiments performed on fraction 8 demonstrated an EC 50 of 2µM, only slightly lower than the 4µM of the previously tested >98% pure CIDSTFNWQHRILLV. This corresponds with the observation that fraction 8 does not represent an isolated modified species but rather is still comprised primarily of native peptide. In fact, no unique peak could be observed in fractions 8-11 on the corresponding mass spectra, suggesting that the T-cell agonist is still present as only a minor component in these fractions. This has prevented us from using these data to identify the peptide modification needed for antigenicity. Deamidation of asparagine or glutamine residues represents one common peptide modification that has been shown to alter antigenicity(3). We therefore tested whether demamidation of the asparagine (N) residue in CIDSTFNWQHRI to asparatate or iso-aspartate (CIDSTFDWHQRI or CIDSTF-iso-Asp- WQHRI) resulted in a more antigenic peptide. Both of these modified peptides completely failed to stimulate the HERV-K(HML-2)-Env-specific T-cell clone (data not shown). Taken together, these data indicate that the HERV-K(HML-2)-Env-specific T-cell clone is remarkably specific for its cognate determinant, as a number of very subtle changes in the peptide sequence abolished recognition. The peptide fractionation experiment clearly demonstrates that the unmodified CIDSTFNWQHRILLV peptide is not the determinant recognized by the clone however, we have been unable to identify the precise identity of the modification required to render this peptide antigenic. The fact that the exact chemical composition of determinants recognized by T-cells is generally unknown has been highlighted by others (4) 7
8 Recognition of low-level HIV-1 infections at 16 hours post-infection by HERV- K(HML-2)-Env-specific T-cells. The majority of recognition assays presented in this manuscript tested the ability of HERV-K(HML-2)-Env-specific T-cells to recognize high-levels of HIV-1 infection. One of the side observations from the ARV suppression recognition assay presented in Figure 6 was that even the small amount of HIV-1 infection that occurred in nevirapinetreated cells was sufficient to induce measurable recognition by the T-cell clone, although this was reduced as compared to cells infected to high-levels in the absence of ARVs. We repeated this experiment and again observed that even a low-level HIV-1 infection defined by both a low frequency of Gag + cells and low-levels of Gag expression on a percell basis was sufficient to trigger robust recognition by the HERV-K(HML-2)-Envspecific T-cell clone. Also of note, the co-culture between clone and target was initiated at 16 hours post-infection. The addition of brefledin A at the time of co-culture would have blocked additional MHC-I antigen presentation over the course of this co-culture. Thus, low levels of HIV-1 infection are detectable by HERV-K(HML-2)-Env-specific T- cell clones within at least 16 hours of infection. 8
9 Supplementary Figure 1 CD4+ Targets ARV Mix Nevirapine No ARVs CD HIV-1-Gag HERV-K(HML-2)-Env-specific clone ARV Mix Nevirapine No ARVs CD107a IFN-! Supplementary Figure 1. HERV-K(HML-2)-Env-specific T-cells recognize lowlevels of HIV-1 infection within 16 hours of infection. Primary CD4 + T-cells autologous to the HERV-K(HML-2)-Env-specific T-cell clone were treated with either nevirapine, a mix of ARVs (see Figure 6 legend), or maintained as no ARV controls. These cells were subsequently infected with HIV-1. At 16 hours post-infection, these infected targets were co-cultured with the HERV-K(HML-2)-Env-specific T-cell clone in the presence of brefeldin A for 12 hours. Shown are flow cytometry plots gated on CD8 + cells and depicting CD107a by IFN-γ. 9
10 Elimination of SIV-infected cells by HERV-K(HML-2)-Env-specific T-cells. Here we present additional data supporting that the HERV-K(HML-2)-Envspecific T-cell clone specifically eliminates SIV-infected cells. The percentage of SIVinfected primary CD4 + T-cells was substantially reduced upon co-culture with HERV- K(HML-2)-specific, but not with CMV-pp65- or HIV-1-Gag-specific CD8 + T-cell clones (Supplementary Figure 2A, B). Pretreatment of these target cells with the anti-mhc-i antibody DX17 reduced this effect. We also observed dose-dependent elimination of SIVmac251-infected targets upon the co-culture with different clone (effector) to target ratios of HERV-K(HML-2)-Env-specific T-cell clone (Supplementary Figure 2C). Supplementary Figure 2 10
11 Supplementary Figure 2. HERV-K(HML-2)-Env-specific T-cells eliminate SIVinfected cells. Primary CD4+ T-cells from subject OM9 were infected with the indicated SIV viruses. Infections were allowed to proceed for 72 hours and then either CMV-pp65-, HIV-1-Gag, or HERV-K(HML-2)-Env-specific T-cell clones were added at a ratio of 1 clone:10 targets (A, B) or at the indicated effector (clone) to target ratios and cultured for 24 hours. Where indicated, these target cells were pre-incubated with the anti-mhc-i antibody DX17 at 10µg/ml prior to the addition of T-cell clone. (C) SIVmac239-infected target cells were co-cultured with the HERV-K(HML-2)-Env-specific T-cell clone at the indicated effector (clone) to target ratios. Levels of infection were measured by flow cytometry staining for CD4 and intracellular HIV-1-Gag. Supplementary References. 1. Chen, W., Yewdell, J.W., Levine, R.L., and Bennink, J.R Modification of cysteine residues in vitro and in vivo affects the immunogenicity and antigenicity of major histocompatibility complex class I- restricted viral determinants. J Exp Med 189: Puglielli, M.T., Zajac, A.J., van der Most, R.G., Dzuris, J.L., Sette, A., Altman, J.D., and Ahmed, R In vivo selection of a lymphocytic choriomeningitis virus variant that affects recognition of the GP33-43 epitope by H- 2Db but not H- 2Kb. J Virol 75: Arentz- Hansen, H., Korner, R., Molberg, O., Quarsten, H., Vader, W., Kooy, Y.M., Lundin, K.E., Koning, F., Roepstorff, P., Sollid, L.M., et al The intestinal T cell response to alpha- gliadin in adult celiac disease is focused on a single deamidated glutamine targeted by tissue transglutaminase. J Exp Med 191: Yewdell, J.W The seven dirty little secrets of major histocompatibility complex class I antigen processing. Immunol Rev 207:
Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide
 Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide pulsed T2 cells. clone avidity by 4-hour 51 Cr-release assay 50% lysis at E:T 10:1 [LML peptide, M] #24
Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide pulsed T2 cells. clone avidity by 4-hour 51 Cr-release assay 50% lysis at E:T 10:1 [LML peptide, M] #24
NK mediated Antibody Dependent Cellular Cytotoxicity in HIV infections
 NK mediated Antibody Dependent Cellular Cytotoxicity in HIV infections Amy Chung Dr. Ivan Stratov Prof. Stephen Kent ADCC process consists of Target cell QuickTime and a TIFF (Uncompressed) FcγR decompressor
NK mediated Antibody Dependent Cellular Cytotoxicity in HIV infections Amy Chung Dr. Ivan Stratov Prof. Stephen Kent ADCC process consists of Target cell QuickTime and a TIFF (Uncompressed) FcγR decompressor
HERV-K specific T cells eliminate diverse HIV-1/2 and SIV primary isolates
 Research article HERV-K specific T cells eliminate diverse HIV-1/2 and SIV primary isolates R. Brad Jones, 1 Keith E. Garrison, 2 Shariq Mujib, 1 Vesna Mihajlovic, 1 Nasra Aidarus, 1 Diana V. Hunter, 1
Research article HERV-K specific T cells eliminate diverse HIV-1/2 and SIV primary isolates R. Brad Jones, 1 Keith E. Garrison, 2 Shariq Mujib, 1 Vesna Mihajlovic, 1 Nasra Aidarus, 1 Diana V. Hunter, 1
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
DEBATE ON HIV ENVELOPE AS A T CELL IMMUNOGEN HAS BEEN GAG-GED
 DEBATE ON HIV ENVELOPE AS A T CELL IMMUNOGEN HAS BEEN GAG-GED Viv Peut Kent Laboratory, University of Melbourne, Australia WHY ENVELOPE? Env subject to both humoral and cellular immune responses Perhaps
DEBATE ON HIV ENVELOPE AS A T CELL IMMUNOGEN HAS BEEN GAG-GED Viv Peut Kent Laboratory, University of Melbourne, Australia WHY ENVELOPE? Env subject to both humoral and cellular immune responses Perhaps
Supporting Information
 Supporting Information Sui et al..7/pnas.997 Pre-CLP CM9 LA9 SL Tat# Pol Vif % Tetramer + CD + CD + Vac+IL- +IL- Vac Fig. S. Frequencies of six different CD + CD + Mamu-A*-tetramer + cells were measured
Supporting Information Sui et al..7/pnas.997 Pre-CLP CM9 LA9 SL Tat# Pol Vif % Tetramer + CD + CD + Vac+IL- +IL- Vac Fig. S. Frequencies of six different CD + CD + Mamu-A*-tetramer + cells were measured
Mechanisms of antagonism of HIVspecific CD4+ T cell responses BSRI
 Mechanisms of antagonism of HIVspecific CD4+ T cell responses BSRI Problems Virus escape from immune recognition Antagonism of T cell responses Peptide-MHC-TCR interaction T cell antagonism Variants of
Mechanisms of antagonism of HIVspecific CD4+ T cell responses BSRI Problems Virus escape from immune recognition Antagonism of T cell responses Peptide-MHC-TCR interaction T cell antagonism Variants of
Surface plasmon resonance (SPR) analysis
 Surface plasmon resonance (SPR) analysis Soluble CD8αα and was manufactured as described previously. 1,2 The C12C heterodimeric TCR was produced using an engineered disulfide bridge between the constant
Surface plasmon resonance (SPR) analysis Soluble CD8αα and was manufactured as described previously. 1,2 The C12C heterodimeric TCR was produced using an engineered disulfide bridge between the constant
Cytotoxicity assays. Rory D. de Vries, PhD 1. Viroscience lab, Erasmus MC, Rotterdam, the Netherlands
 Cytotoxicity assays Rory D. de Vries, PhD 1 1 Viroscience lab, Erasmus MC, Rotterdam, the Netherlands Anti-influenza immunity Humoral / CD4+ / CD8+ / NK? Function of CTL Elimination of virus-infected cells?
Cytotoxicity assays Rory D. de Vries, PhD 1 1 Viroscience lab, Erasmus MC, Rotterdam, the Netherlands Anti-influenza immunity Humoral / CD4+ / CD8+ / NK? Function of CTL Elimination of virus-infected cells?
Antigen Recognition by T cells
 Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific
 SUPPLEMENTARY FIGURE LEGEND Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific CD4 + T cells correlates with intracellular CTLA-4 levels. (a) Comparative CTLA-4 levels
SUPPLEMENTARY FIGURE LEGEND Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific CD4 + T cells correlates with intracellular CTLA-4 levels. (a) Comparative CTLA-4 levels
Potential cross reactions between HIV 1 specific T cells and the microbiome. Andrew McMichael Suzanne Campion
 Potential cross reactions between HIV 1 specific T cells and the microbiome Andrew McMichael Suzanne Campion Role of the Microbiome? T cell (and B cell) immune responses to HIV and Vaccines are influenced
Potential cross reactions between HIV 1 specific T cells and the microbiome Andrew McMichael Suzanne Campion Role of the Microbiome? T cell (and B cell) immune responses to HIV and Vaccines are influenced
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Figure 1. Example of gating strategy
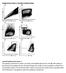 Supplementary Figure 1. Example of gating strategy Legend Supplementary Figure 1: First, gating is performed to include only single cells (singlets) (A) and CD3+ cells (B). After gating on the lymphocyte
Supplementary Figure 1. Example of gating strategy Legend Supplementary Figure 1: First, gating is performed to include only single cells (singlets) (A) and CD3+ cells (B). After gating on the lymphocyte
CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads (Miltenyi Biotec) for depletion of CD25 +
 Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
T Cell Development. Xuefang Cao, MD, PhD. November 3, 2015
 T Cell Development Xuefang Cao, MD, PhD November 3, 2015 Thymocytes in the cortex of the thymus Early thymocytes development Positive and negative selection Lineage commitment Exit from the thymus and
T Cell Development Xuefang Cao, MD, PhD November 3, 2015 Thymocytes in the cortex of the thymus Early thymocytes development Positive and negative selection Lineage commitment Exit from the thymus and
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Significance of the MHC
 CHAPTER 8 Major Histocompatibility Complex (MHC) What is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) - MHC molecules were initially discovered during studies aimed
CHAPTER 8 Major Histocompatibility Complex (MHC) What is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) - MHC molecules were initially discovered during studies aimed
staining and flow cytometry
 Detection of influenza virus-specific T cell responses by intracellular by cytokine intracellular staining cytokine staining and flow cytometry Detection of influenza virus-specific T cell responses and
Detection of influenza virus-specific T cell responses by intracellular by cytokine intracellular staining cytokine staining and flow cytometry Detection of influenza virus-specific T cell responses and
Rajesh Kannangai Phone: ; Fax: ; *Corresponding author
 Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Recombinant Vaccinia Virus-Induced T-Cell Immunity: Quantitation of the Response to the Virus Vector and the Foreign Epitope
 JOURNAL OF VIROLOGY, Apr. 2002, p. 3329 3337 Vol. 76, No. 7 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.7.3329 3337.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Recombinant
JOURNAL OF VIROLOGY, Apr. 2002, p. 3329 3337 Vol. 76, No. 7 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.7.3329 3337.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Recombinant
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
RAISON D ETRE OF THE IMMUNE SYSTEM:
 RAISON D ETRE OF THE IMMUNE SYSTEM: To Distinguish Self from Non-Self Thereby Protecting Us From Our Hostile Environment. Innate Immunity Acquired Immunity Innate immunity: (Antigen nonspecific) defense
RAISON D ETRE OF THE IMMUNE SYSTEM: To Distinguish Self from Non-Self Thereby Protecting Us From Our Hostile Environment. Innate Immunity Acquired Immunity Innate immunity: (Antigen nonspecific) defense
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD-
 Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE
 DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE Eustache Paramithiotis PhD Vice President, Biomarker Discovery & Diagnostics 17 March 2016 PEPTIDE PRESENTATION BY MHC MHC I Antigen presentation by
DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE Eustache Paramithiotis PhD Vice President, Biomarker Discovery & Diagnostics 17 March 2016 PEPTIDE PRESENTATION BY MHC MHC I Antigen presentation by
Rapid perforin upregulation directly ex vivo by CD8 + T cells is a defining characteristic of HIV elite controllers
 Rapid perforin upregulation directly ex vivo by CD8 + T cells is a defining characteristic of HIV elite controllers Adam R. Hersperger Department of Microbiology University of Pennsylvania Evidence for
Rapid perforin upregulation directly ex vivo by CD8 + T cells is a defining characteristic of HIV elite controllers Adam R. Hersperger Department of Microbiology University of Pennsylvania Evidence for
TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer
 AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
Mutations and Disease Mutations in the Myosin Gene
 Biological Sciences Initiative HHMI Mutations and Disease Mutations in the Myosin Gene Goals Explore how mutations can lead to disease using the myosin gene as a model system. Explore how changes in the
Biological Sciences Initiative HHMI Mutations and Disease Mutations in the Myosin Gene Goals Explore how mutations can lead to disease using the myosin gene as a model system. Explore how changes in the
Structural Analysis of TCRpMHC Complexes Using Computational Tools. Feroze Mohideen Briarcliff High School
 Structural Analysis of TCRpMHC Complexes Using Computational Tools Feroze Mohideen Briarcliff High School TCR-pMHC Complexes Peptide Structure of TCR-pMHC complex PDB AO7 Crossreactivity is the ability
Structural Analysis of TCRpMHC Complexes Using Computational Tools Feroze Mohideen Briarcliff High School TCR-pMHC Complexes Peptide Structure of TCR-pMHC complex PDB AO7 Crossreactivity is the ability
Evidence for antigen-driven TCRβ chain convergence in the tumor infiltrating T cell repertoire
 Evidence for antigen-driven TCRβ chain convergence in the tumor infiltrating T cell repertoire Geoffrey M. Lowman, PhD Senior Staff Scientist EACR - 02 July 2018 For Research Use Only. Not for use in diagnostic
Evidence for antigen-driven TCRβ chain convergence in the tumor infiltrating T cell repertoire Geoffrey M. Lowman, PhD Senior Staff Scientist EACR - 02 July 2018 For Research Use Only. Not for use in diagnostic
CD44
 MR1-5-OP-RU CD24 CD24 CD44 MAIT cells 2.78 11.2 WT RORγt- GFP reporter 1 5 1 4 1 3 2.28 1 5 1 4 1 3 4.8 1.6 8.1 1 5 1 4 1 3 1 5 1 4 1 3 3.7 3.21 8.5 61.7 1 2 1 3 1 4 1 5 TCRβ 2 1 1 3 1 4 1 5 CD44 1 2 GFP
MR1-5-OP-RU CD24 CD24 CD44 MAIT cells 2.78 11.2 WT RORγt- GFP reporter 1 5 1 4 1 3 2.28 1 5 1 4 1 3 4.8 1.6 8.1 1 5 1 4 1 3 1 5 1 4 1 3 3.7 3.21 8.5 61.7 1 2 1 3 1 4 1 5 TCRβ 2 1 1 3 1 4 1 5 CD44 1 2 GFP
Introduction. and Department of Medical Oncology, Dana-Farber Cancer Institute, 77 Avenue Louis Pasteur, Boston, MA 02115, USA
 doi:10.1016/j.jmb.2007.06.057 J. Mol. Biol. (2007) 372, 535 548 CTL Recognition of a Protective Immunodominant Influenza A Virus Nucleoprotein Epitope Utilizes a Highly Restricted Vβ but Diverse Vα Repertoire:
doi:10.1016/j.jmb.2007.06.057 J. Mol. Biol. (2007) 372, 535 548 CTL Recognition of a Protective Immunodominant Influenza A Virus Nucleoprotein Epitope Utilizes a Highly Restricted Vβ but Diverse Vα Repertoire:
Principles of Adaptive Immunity
 Principles of Adaptive Immunity Chapter 3 Parham Hans de Haard 17 th of May 2010 Agenda Recognition molecules of adaptive immune system Features adaptive immune system Immunoglobulins and T-cell receptors
Principles of Adaptive Immunity Chapter 3 Parham Hans de Haard 17 th of May 2010 Agenda Recognition molecules of adaptive immune system Features adaptive immune system Immunoglobulins and T-cell receptors
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
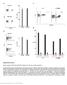 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created.
 938 939 940 941 942 Figure S1 Schematic of the in silico TCRminer and MiXCR validation. 1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created. Then, 100,000 simulated 80 bp
938 939 940 941 942 Figure S1 Schematic of the in silico TCRminer and MiXCR validation. 1,000 in silico simulated alpha, beta, gamma and delta TCR repertoires were created. Then, 100,000 simulated 80 bp
Defining kinetic properties of HIV-specific CD8 + T-cell responses in acute infection
 Defining kinetic properties of HIV-specific CD8 + T-cell responses in acute infection Yiding Yang 1 and Vitaly V. Ganusov 1,2,3 1 Department of Microbiology, University of Tennessee, Knoxville, TN 37996,
Defining kinetic properties of HIV-specific CD8 + T-cell responses in acute infection Yiding Yang 1 and Vitaly V. Ganusov 1,2,3 1 Department of Microbiology, University of Tennessee, Knoxville, TN 37996,
How T cells recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do? Monoclonal antibody approach
 How T cells recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about T cell function was known, but the receptor genes had not been identified
How T cells recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about T cell function was known, but the receptor genes had not been identified
A second type of TCR TCR: An αβ heterodimer
 How s recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about function was known, but the receptor genes had not been identified Recall
How s recognize antigen: The T Cell Receptor (TCR) Identifying the TCR: Why was it so hard to do By the early 1980s, much about function was known, but the receptor genes had not been identified Recall
SUPPLEMENTARY INFORMATION
 Supplementary Notes 1: accuracy of prediction algorithms for peptide binding affinities to HLA and Mamu alleles For each HLA and Mamu allele we have analyzed the accuracy of four predictive algorithms
Supplementary Notes 1: accuracy of prediction algorithms for peptide binding affinities to HLA and Mamu alleles For each HLA and Mamu allele we have analyzed the accuracy of four predictive algorithms
HD1 (FLU) HD2 (EBV) HD2 (FLU)
 ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
Therapeutic effect of baicalin on experimental autoimmune encephalomyelitis. is mediated by SOCS3 regulatory pathway
 Therapeutic effect of baicalin on experimental autoimmune encephalomyelitis is mediated by SOCS3 regulatory pathway Yuan Zhang 1,2, Xing Li 1,2, Bogoljub Ciric 1, Cun-gen Ma 3, Bruno Gran 4, Abdolmohamad
Therapeutic effect of baicalin on experimental autoimmune encephalomyelitis is mediated by SOCS3 regulatory pathway Yuan Zhang 1,2, Xing Li 1,2, Bogoljub Ciric 1, Cun-gen Ma 3, Bruno Gran 4, Abdolmohamad
Why are validated immunogenicity assays important for HIV vaccine development?
 Why are validated immunogenicity assays important for HIV vaccine development? There is a need to compare immunogenicity of products in the pipeline, when similar or different in class when developed by
Why are validated immunogenicity assays important for HIV vaccine development? There is a need to compare immunogenicity of products in the pipeline, when similar or different in class when developed by
Copyright 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings
 Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
MHC class I MHC class II Structure of MHC antigens:
 MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
T cell Receptor. Chapter 9. Comparison of TCR αβ T cells
 Chapter 9 The αβ TCR is similar in size and structure to an antibody Fab fragment T cell Receptor Kuby Figure 9-3 The αβ T cell receptor - Two chains - α and β - Two domains per chain - constant (C) domain
Chapter 9 The αβ TCR is similar in size and structure to an antibody Fab fragment T cell Receptor Kuby Figure 9-3 The αβ T cell receptor - Two chains - α and β - Two domains per chain - constant (C) domain
Narrowed TCR repertoire and viral escape as a consequence of heterologous immunity
 Research article Narrowed TCR repertoire and viral escape as a consequence of heterologous immunity Markus Cornberg, 1,2 Alex T. Chen, 1 Lee A. Wilkinson, 1 Michael A. Brehm, 1 Sung-Kwon Kim, 1 Claudia
Research article Narrowed TCR repertoire and viral escape as a consequence of heterologous immunity Markus Cornberg, 1,2 Alex T. Chen, 1 Lee A. Wilkinson, 1 Michael A. Brehm, 1 Sung-Kwon Kim, 1 Claudia
Treatment with IL-7 Prevents the Decline of Circulating CD4 + T Cells during the Acute Phase of SIV Infection in Rhesus Macaques
 SUPPORTING INFORMATION FOR: Treatment with IL-7 Prevents the Decline of Circulating CD4 + T Cells during the Acute Phase of SIV Infection in Rhesus Macaques Lia Vassena, 1,2 Huiyi Miao, 1 Raffaello Cimbro,
SUPPORTING INFORMATION FOR: Treatment with IL-7 Prevents the Decline of Circulating CD4 + T Cells during the Acute Phase of SIV Infection in Rhesus Macaques Lia Vassena, 1,2 Huiyi Miao, 1 Raffaello Cimbro,
Key Concept B F. How do peptides get loaded onto the proper kind of MHC molecule?
 Location of MHC class I pockets termed B and F that bind P and P9 amino acid side chains of the peptide Different MHC alleles confer different functional properties on the adaptive immune system by specifying
Location of MHC class I pockets termed B and F that bind P and P9 amino acid side chains of the peptide Different MHC alleles confer different functional properties on the adaptive immune system by specifying
B F. Location of MHC class I pockets termed B and F that bind P2 and P9 amino acid side chains of the peptide
 Different MHC alleles confer different functional properties on the adaptive immune system by specifying molecules that have different peptide binding abilities Location of MHC class I pockets termed B
Different MHC alleles confer different functional properties on the adaptive immune system by specifying molecules that have different peptide binding abilities Location of MHC class I pockets termed B
Significance of the MHC
 CHAPTER 8 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
CHAPTER 8 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
Supplementary Figure 1.
 Supplementary Figure 1. Female Pro-ins2 -/- mice at 5-6 weeks of age were either inoculated i.p. with a single dose of CVB4 (1x10 5 PFU/mouse) or PBS and treated with αgalcer or control vehicle. On day
Supplementary Figure 1. Female Pro-ins2 -/- mice at 5-6 weeks of age were either inoculated i.p. with a single dose of CVB4 (1x10 5 PFU/mouse) or PBS and treated with αgalcer or control vehicle. On day
Antigen Presentation to T lymphocytes
 Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antigen processing: How are
Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antigen processing: How are
Supporting Information
 Supporting Information Horwitz et al. 73/pnas.35295 A Copies ml - C 3NC7 7 697 698 7 7 73 76-2 2 Days Gp2 residue G458D G459D T278A 7/36 N28 K D 28 459 A28T ID# 697 ID# 698 ID# 7 ID# 7 ID# 73 ID# 76 ID#
Supporting Information Horwitz et al. 73/pnas.35295 A Copies ml - C 3NC7 7 697 698 7 7 73 76-2 2 Days Gp2 residue G458D G459D T278A 7/36 N28 K D 28 459 A28T ID# 697 ID# 698 ID# 7 ID# 7 ID# 73 ID# 76 ID#
Determinants of Immunogenicity and Tolerance. Abul K. Abbas, MD Department of Pathology University of California San Francisco
 Determinants of Immunogenicity and Tolerance Abul K. Abbas, MD Department of Pathology University of California San Francisco EIP Symposium Feb 2016 Why do some people respond to therapeutic proteins?
Determinants of Immunogenicity and Tolerance Abul K. Abbas, MD Department of Pathology University of California San Francisco EIP Symposium Feb 2016 Why do some people respond to therapeutic proteins?
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
T Cell Development II: Positive and Negative Selection
 T Cell Development II: Positive and Negative Selection 8 88 The two phases of thymic development: - production of T cell receptors for antigen, by rearrangement of the TCR genes CD4 - selection of T cells
T Cell Development II: Positive and Negative Selection 8 88 The two phases of thymic development: - production of T cell receptors for antigen, by rearrangement of the TCR genes CD4 - selection of T cells
Profiling HLA motifs by large scale peptide sequencing Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009
 Profiling HLA motifs by large scale peptide sequencing 2009 Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009 HLA Background The human leukocyte antigen system (HLA) is the
Profiling HLA motifs by large scale peptide sequencing 2009 Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009 HLA Background The human leukocyte antigen system (HLA) is the
Lecture 11. Immunology and disease: parasite antigenic diversity
 Lecture 11 Immunology and disease: parasite antigenic diversity RNAi interference video and tutorial (you are responsible for this material, so check it out.) http://www.pbs.org/wgbh/nova/sciencenow/3210/02.html
Lecture 11 Immunology and disease: parasite antigenic diversity RNAi interference video and tutorial (you are responsible for this material, so check it out.) http://www.pbs.org/wgbh/nova/sciencenow/3210/02.html
READ THIS FIRST. Your Name
 Introduction to Biochemistry Final Examination - Individual (Part I) Monday, 24 May 2010 7:00 8:45 PM H. B. White Instructor 120 Points Your Name "Ability is what you're capable of doing. Motivation determines
Introduction to Biochemistry Final Examination - Individual (Part I) Monday, 24 May 2010 7:00 8:45 PM H. B. White Instructor 120 Points Your Name "Ability is what you're capable of doing. Motivation determines
Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12
 1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
Chapter 22: The Lymphatic System and Immunity
 Bio40C schedule Lecture Immune system Lab Quiz 2 this week; bring a scantron! Study guide on my website (see lab assignments) Extra credit Critical thinking questions at end of chapters 5 pts/chapter Due
Bio40C schedule Lecture Immune system Lab Quiz 2 this week; bring a scantron! Study guide on my website (see lab assignments) Extra credit Critical thinking questions at end of chapters 5 pts/chapter Due
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1175194/dc1 Supporting Online Material for A Vital Role for Interleukin-21 in the Control of a Chronic Viral Infection John S. Yi, Ming Du, Allan J. Zajac* *To whom
www.sciencemag.org/cgi/content/full/1175194/dc1 Supporting Online Material for A Vital Role for Interleukin-21 in the Control of a Chronic Viral Infection John S. Yi, Ming Du, Allan J. Zajac* *To whom
Third line of Defense
 Chapter 15 Specific Immunity and Immunization Topics -3 rd of Defense - B cells - T cells - Specific Immunities Third line of Defense Specific immunity is a complex interaction of immune cells (leukocytes)
Chapter 15 Specific Immunity and Immunization Topics -3 rd of Defense - B cells - T cells - Specific Immunities Third line of Defense Specific immunity is a complex interaction of immune cells (leukocytes)
Received 19 September 2007/Accepted 21 November 2007
 JOURNAL OF VIROLOGY, Feb. 2008, p. 1723 1738 Vol. 82, No. 4 0022-538X/08/$08.00 0 doi:10.1128/jvi.02084-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Patterns of CD8 Immunodominance
JOURNAL OF VIROLOGY, Feb. 2008, p. 1723 1738 Vol. 82, No. 4 0022-538X/08/$08.00 0 doi:10.1128/jvi.02084-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Patterns of CD8 Immunodominance
Chapter 6. Antigen Presentation to T lymphocytes
 Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
Cellular Immunity in Aging and HIV: Correlates of Protection. Immune Senescence
 Cellular Immunity in Aging and HIV: Correlates of Protection Janet E. McElhaney, MD Professor of Medicine Allan M. McGavin Chair in Research Geriatrics University of British Columbia Vancouver, BC and
Cellular Immunity in Aging and HIV: Correlates of Protection Janet E. McElhaney, MD Professor of Medicine Allan M. McGavin Chair in Research Geriatrics University of British Columbia Vancouver, BC and
Significance of the MHC
 CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
Cell Mediated Immunity (I) Dr. Aws Alshamsan Department of Pharmaceu5cs Office: AA87 Tel:
 Cell Mediated Immunity (I) Dr. Aws Alshamsan Department of Pharmaceu5cs Office: AA87 Tel: 4677363 aalshamsan@ksu.edu.sa Learning Objectives By the end of this lecture you will be able to: 1 Understand
Cell Mediated Immunity (I) Dr. Aws Alshamsan Department of Pharmaceu5cs Office: AA87 Tel: 4677363 aalshamsan@ksu.edu.sa Learning Objectives By the end of this lecture you will be able to: 1 Understand
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
There are 2 major lines of defense: Non-specific (Innate Immunity) and. Specific. (Adaptive Immunity) Photo of macrophage cell
 There are 2 major lines of defense: Non-specific (Innate Immunity) and Specific (Adaptive Immunity) Photo of macrophage cell Development of the Immune System ery pl neu mφ nk CD8 + CTL CD4 + thy TH1 mye
There are 2 major lines of defense: Non-specific (Innate Immunity) and Specific (Adaptive Immunity) Photo of macrophage cell Development of the Immune System ery pl neu mφ nk CD8 + CTL CD4 + thy TH1 mye
Antigen presenting cells
 Antigen recognition by T and B cells - T and B cells exhibit fundamental differences in antigen recognition - B cells recognize antigen free in solution (native antigen). - T cells recognize antigen after
Antigen recognition by T and B cells - T and B cells exhibit fundamental differences in antigen recognition - B cells recognize antigen free in solution (native antigen). - T cells recognize antigen after
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Innate and Cellular Immunology Control of Infection by Cell-mediated Immunity
 Innate & adaptive Immunity Innate and Cellular Immunology Control of Infection by Cell-mediated Immunity Helen Horton PhD Seattle Biomedical Research Institute Depts of Global Health & Medicine, UW Cellular
Innate & adaptive Immunity Innate and Cellular Immunology Control of Infection by Cell-mediated Immunity Helen Horton PhD Seattle Biomedical Research Institute Depts of Global Health & Medicine, UW Cellular
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
Medical Virology Immunology. Dr. Sameer Naji, MB, BCh, PhD (UK) Head of Basic Medical Sciences Dept. Faculty of Medicine The Hashemite University
 Medical Virology Immunology Dr. Sameer Naji, MB, BCh, PhD (UK) Head of Basic Medical Sciences Dept. Faculty of Medicine The Hashemite University Human blood cells Phases of immune responses Microbe Naïve
Medical Virology Immunology Dr. Sameer Naji, MB, BCh, PhD (UK) Head of Basic Medical Sciences Dept. Faculty of Medicine The Hashemite University Human blood cells Phases of immune responses Microbe Naïve
Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs.
 Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs. C.J. SAVOIE, N. KAMIKAWAJI, T. SASAZUKI Dept. of Genetics, Medical Institute of Bioregulation, Kyushu
Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs. C.J. SAVOIE, N. KAMIKAWAJI, T. SASAZUKI Dept. of Genetics, Medical Institute of Bioregulation, Kyushu
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Degenerate T-cell Recognition of Peptides on MHC Molecules Creates Large Holes in the T-cell Repertoire
 Degenerate T-cell Recognition of Peptides on MHC Molecules Creates Large Holes in the T-cell Repertoire Jorg J. A. Calis*, Rob J. de Boer, Can Keşmir Theoretical Biology & Bioinformatics, Utrecht University,
Degenerate T-cell Recognition of Peptides on MHC Molecules Creates Large Holes in the T-cell Repertoire Jorg J. A. Calis*, Rob J. de Boer, Can Keşmir Theoretical Biology & Bioinformatics, Utrecht University,
Supplemental Materials for. Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to. FTY720 during neuroinflammation
 Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Lecture 7: Signaling Through Lymphocyte Receptors
 Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
RAISON D ETRE OF THE IMMUNE SYSTEM:
 RAISON D ETRE OF THE IMMUNE SYSTEM: To Distinguish Self from Non-Self Thereby Protecting Us From Our Hostile Environment. Innate Immunity Adaptive Immunity Innate immunity: (Antigen - nonspecific) defense
RAISON D ETRE OF THE IMMUNE SYSTEM: To Distinguish Self from Non-Self Thereby Protecting Us From Our Hostile Environment. Innate Immunity Adaptive Immunity Innate immunity: (Antigen - nonspecific) defense
Supporting Information
 Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Developing Understanding of CMI. Dr Tom Wilkinson Associate Professor of Respiratory Medicine Faculty of Medicine University of Southampton UK
 Pre-existing influenza-specific CD4+ T cells correlate with homologous and heterotypic response and disease protection against influenza challenge in humans Do At Risk Groups Rely more on CMI to Viruses?
Pre-existing influenza-specific CD4+ T cells correlate with homologous and heterotypic response and disease protection against influenza challenge in humans Do At Risk Groups Rely more on CMI to Viruses?
Epitope discovery. Using predicted MHC binding CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS
 Epitope discovery Using predicted MHC binding SARS corona virus The 2003 outbreak The disease Lung Pathology Inflammatory exudation in alveoli and interstitial spaces Monocytic and lymphocytic infiltration
Epitope discovery Using predicted MHC binding SARS corona virus The 2003 outbreak The disease Lung Pathology Inflammatory exudation in alveoli and interstitial spaces Monocytic and lymphocytic infiltration
Antiviral CD8 + T Cells Restricted by Human Leukocyte Antigen Class II Exist during Natural HIV Infection and Exhibit Clonal Expansion
 Article Antiviral CD8 + T Cells Restricted by Human Leukocyte Antigen Class II Exist during Natural HIV Infection and Exhibit Clonal Expansion Highlights d CD8 + T cells restricted by HLA-DRB1 exist in
Article Antiviral CD8 + T Cells Restricted by Human Leukocyte Antigen Class II Exist during Natural HIV Infection and Exhibit Clonal Expansion Highlights d CD8 + T cells restricted by HLA-DRB1 exist in
PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human
 Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
On an individual level. Time since infection. NEJM, April HIV-1 evolution in response to immune selection pressures
 HIV-1 evolution in response to immune selection pressures BISC 441 guest lecture Zabrina Brumme, Ph.D. Assistant Professor, Faculty of Health Sciences Simon Fraser University http://www3.niaid.nih.gov/topics/hivaids/understanding/biology/structure.htm
HIV-1 evolution in response to immune selection pressures BISC 441 guest lecture Zabrina Brumme, Ph.D. Assistant Professor, Faculty of Health Sciences Simon Fraser University http://www3.niaid.nih.gov/topics/hivaids/understanding/biology/structure.htm
Received 25 April 2002/Accepted 21 May 2002
 JOURNAL OF VIROLOGY, Sept. 2002, p. 8757 8768 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8757 8768.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Clustering
JOURNAL OF VIROLOGY, Sept. 2002, p. 8757 8768 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8757 8768.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Clustering
MATERIALS AND METHODS. Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All
 MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
Variable HIV peptide stability in human cytosol is critical to epitope presentation and immune escape
 Research article Variable HIV peptide stability in human cytosol is critical to epitope presentation and immune escape Estibaliz Lazaro, 1 Carl Kadie, 2 Pamela Stamegna, 1 Shao Chong Zhang, 1 Pauline Gourdain,
Research article Variable HIV peptide stability in human cytosol is critical to epitope presentation and immune escape Estibaliz Lazaro, 1 Carl Kadie, 2 Pamela Stamegna, 1 Shao Chong Zhang, 1 Pauline Gourdain,
Bioinformation Volume 5
 Do N-glycoproteins have preference for specific sequons? R Shyama Prasad Rao 1, 2, *, Bernd Wollenweber 1 1 Aarhus University, Department of Genetics and Biotechnology, Forsøgsvej 1, Slagelse 4200, Denmark;
Do N-glycoproteins have preference for specific sequons? R Shyama Prasad Rao 1, 2, *, Bernd Wollenweber 1 1 Aarhus University, Department of Genetics and Biotechnology, Forsøgsvej 1, Slagelse 4200, Denmark;
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV. (Summary of the recommendations from an Enterprise Working Group)
 AIDS Vaccine 07, Seattle, August 20-23, 2007 EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV (Summary of the recommendations from an Enterprise Working Group) The Working Group Reston, Virginia,
AIDS Vaccine 07, Seattle, August 20-23, 2007 EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV (Summary of the recommendations from an Enterprise Working Group) The Working Group Reston, Virginia,
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Defensive mechanisms include :
 Acquired Immunity Defensive mechanisms include : 1) Innate immunity (Natural or Non specific) 2) Acquired immunity (Adaptive or Specific) Cell-mediated immunity Humoral immunity Two mechanisms 1) Humoral
Acquired Immunity Defensive mechanisms include : 1) Innate immunity (Natural or Non specific) 2) Acquired immunity (Adaptive or Specific) Cell-mediated immunity Humoral immunity Two mechanisms 1) Humoral
