Figure S1! CrFK! DNA Synthesis! Copies viral DNA/ 100 ng DNA (Log 10 ) % Infected (GFP+) cells! 100! 80! 60! 40! 20! 0! 1! 10! 100! 1000!
|
|
|
- Ellen Watson
- 6 years ago
- Views:
Transcription
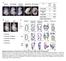
1 Figure S1! A! B! % Infected (GFP+) cells! 100! 80! 60! 40! 20! 0! 1! 10! 100! 1000! Dose μl! CrFK! B-MLV! N-MLV! Template:! Water! Ube2V2! Ube2V1! Primers:! V2! V1! V2! V1! V2! V1! bp! 500! 300! 200! 150! 100! 75! 50! 25! C! D! Copies viral DNA/ 100 ng DNA (Log 10 ) shrna:! Relative mrna copies shrna:! DNA Synthesis! B-MLV N-MLV None Scr Ube2V1 Scr Ube2V1
2 Figure S2A!
3 Figure S2B!
4 Figure S3! A! TRIM5α rh,-n-met +N-alpha acetyl group! (MW calc = 57,943 Da)! B! TRIM5α rh, -N-Met! (MW calc = 57,901 Da, species absent)! TRIM5α rh,-n-met +N-alpha acetyl group! (MW calc = 57,943 Da)! TRIM5α rh, +N-Met! (MW calc = 58,032 Da, species absent)!
5 Figure S4! A! MW! βme:! ! 97! 66! 45! 31! 21.5! 14.4! 6.5! (Marker)! E2:! Ube2B! Ube2C! Ube2H! *!! *! 1! 2! 3! 4! 5! 6!! *!! UBA1~Ub! UBA1! E2~Ub! E2*! Ub! B! MW! 116! 97! 66! C! (Marker)! (Marker)! βme:! ! 31! 21.5! 14.4! 6.5! E2:! Ube2W! Ube2N! Ube2L3! E2:! MW! 116! 97! 66!!! *! *! 1! 2! 3! 4! 5! 6! Ube2D3! βme:! + - UBA1~Ub! UBA1! *!! UBA1~Ub! UBA1! E2~Ub! E2*! Ub! 45! 31! 21.5! 14.4! 6.5! *! 1! 2!! E2~Ub! E2*! Ub!
6 Figure S5! A! K0! K63R! K48R! K33R! K29R! K27R! K11R! K6R! Ube2D3!! Ubiquitin:! -!! E2:! MW! IB: TRIM5αrh! 250! 150! (Ub)n-TRIM5αrh! 100! Ub-TRIM5αrh! TRIM5αrh! 50! 5! 6! 7! 8! 9! C! D! 4 3 shrna:! Boil! 2 Ube2D3! IB: Actin 5 Scr! IB: HA B-MLV N-MLV None! Ube2V Copies viral DNA/! 100 ng DNA (Log10)! 42 - Ube2D3 shrna: HA-Ube2D3: MW 15 - None DNA Synthesis! Infection! shrna:! Boil! B! 10! Ube2D3! 4! Scr! 3! None! 2! % Infected (GFP+) cells! 1!
7 Figure S6! A! E2s:! MW! -! -! N / V2! -! -! W! -! * W! D3! IB: Ub! 250! 150! 100! *! Unanchored ubiquitin chains! 50! 37! 25! 20! 15! 1! 2! 3! 4! 5! Relative lane intensity! B! MW! 250! 150! 100! IB: TRIM5α rh! (Lys63-Ub) n -TRIM5α rh! 50! 1! 2! 3! 4! 5! Ub-TRIM5α rh! TRIM5α rh!
8 Figure S7! A! E2/E3 ratio:! 1! 0.5! 1! 0.5! 1! ATP:! -! +! +! +! +! E2:! TRIM5α rh :! Ube2W! *! *! *! +! +! (Coomassie Stain)! Ub (3) -TRIM5α rh! B! 1! 2! 3! 4! 5! Ub (2) -TRIM5α rh! Ub-TRIM5α rh! TRIM5α rh! (MH 2 ) Selected for CID Theoretical mass = Full Scan MS S M L Y K E G E R G G Relative Abundance (MH 2 ) m/z m/z Full Scan MS / MS 2 + CID of m/z (MH 2 ) Relative Abundance y y b S M L Y K E G E R y y G G y b b y y a y6 (2+) b b b y m/z
9 Figure S8 A Copies viral DNA/ 100 ng DNA (Log 10 ) DNA Synthesis Vector T5 T5 3KR T5 7KR Boil B-MLV N-MLV B % Infected (GFP+) cells Infection Vector T5 T5 3KR T5 7KR Boil C TRIM5 -HA: 3KR 7KR MG132: E TRIM5 PRYSPRY v1 (PDB: 3UV9) MW TRIM5 -HA IB: HA K371 K v IB: Actin D Pulldown: His Input (1%) TRIM5 -HA: None 3KR 7KR None 3KR 7KR 6His-Ub: None Vector None Vector MW (Ub) n -TRIM5 - HA Ub-TRIM5 -HA TRIM5 -HA MW 40 - IB: HA IB: Actin
10 Table S1. Expression Plasmids, Antibodies, sirnas, Oligonucleotides, Real-Time Probes Used in This Study 1A. Bacterial Expression Vectors Plasmid Name Source 1 Internal ID Cloning Site Epitope Tags pet11hispp-ube2b NP_ WISP NheI-XhoI His-PP pet11hispp-ube2c NP_ WISP NheI-XhoI His-PP pet11hispp-ube2d3 NP_ WISP14-34 NheI-XhoI His-PP pet11hispp-ube2h NP_ WISP NheI-XhoI His-PP pet11hispp-ube2l3 NP_ WISP14-35 NheI-XhoI His-PP pet11hispp-ube2n NP_ WISP NheI-XhoI His-PP pet11hispp-ube2v1 NP_ WISP14-36 NheI-XhoI His-PP pet11hispp-ube2v2 NP_ WISP NheI-XhoI His-PP pet11hispp-ube2w Q96B02-2 WISP14-37 NheI-XhoI His-PP pet15b-ubiquitin Rachel Klevit WISP BamHI-KpnI none pet15b-ubiquitin K6R WISP WISP BamHI-KpnI none pet15b-ubiquitin K11R WISP WISP14-38 BamHI-KpnI none pet15b-ubiquitin K27R WISP WISP14-39 BamHI-KpnI none pet15b-ubiquitin K29R WISP WISP14-40 BamHI-KpnI none pet15b-ubiquitin K33R WISP WISP14-41 BamHI-KpnI none pet15b-ubiquitin K48R WISP WISP BamHI-KpnI none pet15b-ubiquitin K63R WISP WISP BamHI-KpnI none pet15b-ubiquitin K0 WISP WISP14-42 BamHI-KpnI none pet21d-uba1 Cynthia Wolberger WISP14-43 His 1B. Insect Expression Vectors Plasmid Name Source 1 Internal ID Cloning Site Epitope Tags pfastbac1 OSF-PP-TRIM5a (rhesus) DQ WISP13-56 SLIC OSF-PP 1C. Mammalian Expression Vectors Plasmid Name Source 1 Internal ID Cloning Site Epitope Tags phr-sin-6xhis-ubgfp- Paul Lehner BamHI-NotI His phr-sin-6xhis-ubgfp-k48r Paul Lehner BamHI-NotI His
11 phr-sin-6xhis-ubgfp-k63r Paul Lehner BamHI-NotI His EXN Paul Bieniasz EcoRI-NotI HA pcdna3.1(+) Invitrogen EcoRI-NotI pcdna4-his Invitrogen EcoRI-NotI His psiren-retroq Clontech SRQ BamHI-EcoRI None pmt123 (Treier et al, 1994) HA-Ub pmt107 (Treier et al, 1994) His-Ub 1D. Virus Production Vectors Plasmid Name Source 1 Internal ID Cloning Site Francoise Loic pcmvi Cosset pcig3-n (Bock et al, 2000) pcig3-b (Bock et al, 2000) pcncg p8.91ex Yasu Takeuchi Yasu Ikeda pmd2.g (Naldini et al, 1996) 1E. Antibodies Antigen Species Blocking Dilution Source TRIM5 Rhesus 2% milk 1/1000 Lampire Biological Laboratories, NIH Reagent Program (5D5-1-1) HA-tag Rat 1% milk 1/1000 Roche, 3F10 Ube2N Rabbit 1% milk 1/1000 Millipore, AB10025 β-actin Mouse None 1/40000 Abcam, AC-15 His-tag Mouse 3% milk 1/1000 Millipore, H8 Ubiquitin Mouse 2% milk 1/2000 Santa Cruz Biotechnology, P4D1 Ub Lys63 Rabbit 5% BSA 1/2000 Millipore, Apu3 1F. shrna 19mers Protein First nucleotide Sense sequence Reference Ube2W 920 (3 UTR) GCATGATAGGGCCTATGAA This study Ube2V (3 UTR) CCCTGGTTTCTTTAAGTCTTAA (Pertel et al, 2011) Ube2V2 324 (ORF) GAGCATACCAGTGTTAGCA This study Ube2D3 237 (ORF) CAGTAATGGCAGCATTTGT (Saville et al, 2004) Ube2N 788 (ORF) GAGCATGGACTAGGCTATA (Duncan et al, 2006) Scrambled GTTATAGGCTCGCAAAAGG This study 1G. Cloning Oligonucleotides
12 Target Sense sequence Anti-sense sequence TRIM5α hu ATGCCAATTGATGGCTTCTGGAATCCTGGTTAATGTAAAGG ATCGGCGGCCGCTCAAGAGCTTGGTGAGCACAGAG CGATGCGGCCGCTTAATTGTTGTATGTTTGTCCTTCTGGTG Ube2V2 CAGTGAATTCCACCATGGCGGTCTCCACAGGAGTTAAAG GCTG Ube2V1 ATGCGGATCCATGCCAGGAGAGGTTCAAGCGTC ATGCGCGGCCGCTTAATTGCTGTAACACTGTCCTTCG Ube2W ATGCGAATTCATGGCGTCAATGCAGACCACAG ATGCGCGGCCGCTCAACAAGTATCATCATG 1H. Real-Time Oligonucleotides and Probe Target Sense sequence Anti-sense sequence GFP CAACAGCCACAACGTCTATATCAT ATGTTGTGGCGGATCTTGAAG GFP probe 5'[6FAM]CCGACAAGCAGAAGAACGGCATCAA[TAM]3' Ube2V2 CCACCAAGGACAAATTATG TGCTAACACTGGTATGCTCC Ube2V1 GGGCCTCCAAGAACAATTTATG CTGATATGGCTCTTGGGTCC GAPDH GGCTGAGAACGGGAAGCTT AGGGATCTCGCTCCTGGAA Ube2W CTCTCCTCAGGTCATGTTTAC GAATTATCCGGTGGTCGTC 1 Source refers to vendors for commercial vectors, papers describing these constructs or the Uniprot accession numbers used for cloning. Abbreviations SLIC, Sequence and ligation independent cloning OSF, One STrEP and FLAG tag PP, PreScission Protease Cleavage site
TRIM5 requires Ube2W to anchor Lys63-linked ubiquitin chains and restrict reverse transcription
 Manuscript EMBO-2014-90361 TRIM5 requires Ube2W to anchor Lys63-linked ubiquitin chains and restrict reverse transcription Adam J Fletcher, Devin E Christensen, Chad Nelson, Choon Ping Tan, Torsten Schaller,
Manuscript EMBO-2014-90361 TRIM5 requires Ube2W to anchor Lys63-linked ubiquitin chains and restrict reverse transcription Adam J Fletcher, Devin E Christensen, Chad Nelson, Choon Ping Tan, Torsten Schaller,
Supplementary Information
 Supplementary Information Supplementary Figure 1. EBV-gB 23-431 mainly exists as trimer in HEK 293FT cells. (a) Western blotting analysis for DSS crosslinked FLAG-gB 23-431. HEK 293FT cells transfected
Supplementary Information Supplementary Figure 1. EBV-gB 23-431 mainly exists as trimer in HEK 293FT cells. (a) Western blotting analysis for DSS crosslinked FLAG-gB 23-431. HEK 293FT cells transfected
Supplemental Information
 Cell Host & Microbe, Volume 14 Supplemental Information HIV-1 Induces the Formation of Stable Microtubules to Enhance Early Infection Yosef Sabo, Derek Walsh, Denis S. Barry, Sedef Tinaztepe, Kenia de
Cell Host & Microbe, Volume 14 Supplemental Information HIV-1 Induces the Formation of Stable Microtubules to Enhance Early Infection Yosef Sabo, Derek Walsh, Denis S. Barry, Sedef Tinaztepe, Kenia de
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supporting Information
 Supporting Information Zhu et al. 1.173/pnas.11167618 SI Materials and Methods DNA Construction. The plasmid pcdna4to/myc-rzap, which expresses myc-tagged full-length rat ZAP, has been described previously
Supporting Information Zhu et al. 1.173/pnas.11167618 SI Materials and Methods DNA Construction. The plasmid pcdna4to/myc-rzap, which expresses myc-tagged full-length rat ZAP, has been described previously
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3
 Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature05732 SUPPLEMENTARY INFORMATION Supplemental Data Supplement Figure Legends Figure S1. RIG-I 2CARD undergo robust ubiquitination a, (top) At 48 h posttransfection with a GST, GST-RIG-I-2CARD
doi: 10.1038/nature05732 SUPPLEMENTARY INFORMATION Supplemental Data Supplement Figure Legends Figure S1. RIG-I 2CARD undergo robust ubiquitination a, (top) At 48 h posttransfection with a GST, GST-RIG-I-2CARD
Supplemental Figure 1
 Supplemental Figure 1 A S100A4: SFLGKRTDEAAFQKLMSNLDSNRDNEVDFQEYCVFLSCIAMMCNEFFEGFPDK Overlap: SF G DE KLM LD N D VDFQEY VFL I M N FF G PD S100A2: SFVGEKVDEEGLKKLMGSLDENSDQQVDFQEYAVFLALITVMCNDFFQGCPDR
Supplemental Figure 1 A S100A4: SFLGKRTDEAAFQKLMSNLDSNRDNEVDFQEYCVFLSCIAMMCNEFFEGFPDK Overlap: SF G DE KLM LD N D VDFQEY VFL I M N FF G PD S100A2: SFVGEKVDEEGLKKLMGSLDENSDQQVDFQEYAVFLALITVMCNDFFQGCPDR
condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1%
 FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
Supplemental Information
 Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
Supplemental Information Tobacco-specific Carcinogen Induces DNA Methyltransferases 1 Accumulation through AKT/GSK3β/βTrCP/hnRNP-U in Mice and Lung Cancer patients Ruo-Kai Lin, 1 Yi-Shuan Hsieh, 2 Pinpin
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Supplementary Figure S1 Supplementary Figure S2
 Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Supplementary methods:
 Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Neither the activation nor suppression of the MAPK pathway affects the ASK1/Vif interaction. (a, b) HEK293 cells were cotransfected with plasmids encoding
Supplementary Figure 1 Supplementary Figure 1. Neither the activation nor suppression of the MAPK pathway affects the ASK1/Vif interaction. (a, b) HEK293 cells were cotransfected with plasmids encoding
TRIM25 Is Required for the Antiviral Activity of Zinc-finger Antiviral Protein
 JVI Accepted Manuscript Posted Online 15 February 2017 J. Virol. doi:10.1128/jvi.00088-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 2 TRIM25 Is Required for the Antiviral
JVI Accepted Manuscript Posted Online 15 February 2017 J. Virol. doi:10.1128/jvi.00088-17 Copyright 2017 American Society for Microbiology. All Rights Reserved. 1 2 TRIM25 Is Required for the Antiviral
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supporting Information
 Supporting Information Gack et al. 1.173/pnas.8494715 SI Text Cell Culture, Transfection, and Reagents. HEK293T, MEF, Vero, EcoPack2-293 (D iosciences), HeLa, HCT116, Huh7, NHLF, 2fTGH wild-type, U3A,
Supporting Information Gack et al. 1.173/pnas.8494715 SI Text Cell Culture, Transfection, and Reagents. HEK293T, MEF, Vero, EcoPack2-293 (D iosciences), HeLa, HCT116, Huh7, NHLF, 2fTGH wild-type, U3A,
Scaffold function of long noncoding RNA HOTAIR in protein ubiquitination
 Yoon et al, page Scaffold function of long noncoding RNA HOTAIR in protein ubiquitination Je-Hyun Yoon,, Kotb Abdelmohsen, Jiyoung Kim, Xiaoling Yang, Jennifer L. Martindale, Kumiko Tominaga-Yamanaka,
Yoon et al, page Scaffold function of long noncoding RNA HOTAIR in protein ubiquitination Je-Hyun Yoon,, Kotb Abdelmohsen, Jiyoung Kim, Xiaoling Yang, Jennifer L. Martindale, Kumiko Tominaga-Yamanaka,
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Online Data Supplement. Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2
 Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Studying apoptosis in DT40 cells
 Studying apoptosis in DT40 cells Sandrine Ruchaud E12.5! Acridine Orange! Role of apoptosis in sculpting the mouse paw E13.5! E14.5! gift of William Wood & Paul Martin! University College, London! Normal
Studying apoptosis in DT40 cells Sandrine Ruchaud E12.5! Acridine Orange! Role of apoptosis in sculpting the mouse paw E13.5! E14.5! gift of William Wood & Paul Martin! University College, London! Normal
Supplementary information. The mitochondrial calcium uniporter is a multimer that can include a dominant-negative pore-forming subunit
 Supplementary information The mitochondrial calcium uniporter is a multimer that can include a dominant-negative pore-forming subunit Anna Raffaello 1,4, Diego De Stefani 1,4, Davide Sabbadin 2, Enrico
Supplementary information The mitochondrial calcium uniporter is a multimer that can include a dominant-negative pore-forming subunit Anna Raffaello 1,4, Diego De Stefani 1,4, Davide Sabbadin 2, Enrico
SUPPLEMENTARY MATERIAL
 SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
Supplementary Materials and Methods
 Supplementary Materials and Methods Reagents and antibodies was purchased from iaffin GmbH & Co KG. Cisplatin (ristol-myers Squibb Co.) and etoposide (Sandoz Pharma Ltd.) were used. Antibodies recognizing
Supplementary Materials and Methods Reagents and antibodies was purchased from iaffin GmbH & Co KG. Cisplatin (ristol-myers Squibb Co.) and etoposide (Sandoz Pharma Ltd.) were used. Antibodies recognizing
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth.
 Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
Overview of the Expressway Cell-Free Expression Systems. Expressway Mini Cell-Free Expression System
 Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Supplemental Information. Induction of Expansion and Folding. in Human Cerebral Organoids
 Cell Stem Cell, Volume 20 Supplemental Information Induction of Expansion and Folding in Human Cerebral Organoids Yun Li, Julien Muffat, Attya Omer, Irene Bosch, Madeline A. Lancaster, Mriganka Sur, Lee
Cell Stem Cell, Volume 20 Supplemental Information Induction of Expansion and Folding in Human Cerebral Organoids Yun Li, Julien Muffat, Attya Omer, Irene Bosch, Madeline A. Lancaster, Mriganka Sur, Lee
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/324/5924/261/dc1 Supporting Online Material for Glioma-Derived Mutations in IDH1 Dominantly Inhibit IDH1 Catalytic Activity and Induce HIF-1α Shimin Zhao, Yan Lin, Wei
www.sciencemag.org/cgi/content/full/324/5924/261/dc1 Supporting Online Material for Glioma-Derived Mutations in IDH1 Dominantly Inhibit IDH1 Catalytic Activity and Induce HIF-1α Shimin Zhao, Yan Lin, Wei
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Erzsebet Kokovay, Susan Goderie, Yue Wang, Steve Lotz, Gang Lin, Yu Sun, Badrinath Roysam, Qin Shen,
 Cell Stem Cell, Volume 7 Supplemental Information Adult SVZ Lineage Cells Home to and Leave the Vascular Niche via Differential Responses to SDF1/CXCR4 Signaling Erzsebet Kokovay, Susan Goderie, Yue Wang,
Cell Stem Cell, Volume 7 Supplemental Information Adult SVZ Lineage Cells Home to and Leave the Vascular Niche via Differential Responses to SDF1/CXCR4 Signaling Erzsebet Kokovay, Susan Goderie, Yue Wang,
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Plasmids Western blot analysis and immunostaining Flow Cytometry Cell surface biotinylation RNA isolation and cdna synthesis
 Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Marine Streptomyces sp. derived antimycin analogues. suppress HeLa cells via depletion HPV E6/E7 mediated by
 Marine Streptomyces sp. derived antimycin analogues suppress HeLa cells via depletion HPV E6/E7 mediated by ROS-dependent ubiquitin proteasome system Weiyi Zhang 1, +, Qian Che 1, 2, +, Hongsheng Tan 1,
Marine Streptomyces sp. derived antimycin analogues suppress HeLa cells via depletion HPV E6/E7 mediated by ROS-dependent ubiquitin proteasome system Weiyi Zhang 1, +, Qian Che 1, 2, +, Hongsheng Tan 1,
Supplementary Figure 1.
 Supplementary Figure 1. Increased β cell mass and islet diameter in βtsc2 -/- mice up to 35 weeks A: Reconstruction of multiple anti-insulin immunofluorescence images showing differences in β cell mass
Supplementary Figure 1. Increased β cell mass and islet diameter in βtsc2 -/- mice up to 35 weeks A: Reconstruction of multiple anti-insulin immunofluorescence images showing differences in β cell mass
Supporting Information
 Supporting Information Fujishita et al. 10.1073/pnas.0800041105 SI Text Polyp Scoring. Intestinal polyps were counted as described (1). Briefly, the small and large intestines were excised, washed with
Supporting Information Fujishita et al. 10.1073/pnas.0800041105 SI Text Polyp Scoring. Intestinal polyps were counted as described (1). Briefly, the small and large intestines were excised, washed with
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis. Midbrain Forebrain/ Forebrain/ Hindbrain Spinal cord Hindbrain Hindbrain
 A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supporting Information. Supporting Tables. S-Table 1 Primer pairs for RT-PCR. Product size. Gene Primer pairs
 Supporting Information Supporting Tables S-Table 1 Primer pairs for RT-PCR. Gene Primer pairs Product size (bp) FAS F: 5 TCTTGGAAGCGATGGGTA 3 429 R: 5 GGGATGTATCATTCTTGGAC 3 SREBP-1c F: 5 CGCTACCGTTCCTCTATCA
Supporting Information Supporting Tables S-Table 1 Primer pairs for RT-PCR. Gene Primer pairs Product size (bp) FAS F: 5 TCTTGGAAGCGATGGGTA 3 429 R: 5 GGGATGTATCATTCTTGGAC 3 SREBP-1c F: 5 CGCTACCGTTCCTCTATCA
E2Select Ubiquitin Conjugation Kit
 E2Select Ubiquitin Conjugation Kit Catalog Number K-982 This kit contains reagents sufficient to perform 2 Ubiquitination reactions for each E2 included in the kit. This package insert must be read in
E2Select Ubiquitin Conjugation Kit Catalog Number K-982 This kit contains reagents sufficient to perform 2 Ubiquitination reactions for each E2 included in the kit. This package insert must be read in
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study.
 Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study. In this study, one fluorophosphonate (FP, 1), three nitrophenol phosphonate probes
Supplementary Figure 1. Chemical structures of activity-based probes (ABPs) and of click reagents used in this study. In this study, one fluorophosphonate (FP, 1), three nitrophenol phosphonate probes
PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65
 SUPPLEMENTARY INFORMATION TITLE: PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65 RUNNING TITLE: PKCζ-NFκB Signaling in Breast Cancer
SUPPLEMENTARY INFORMATION TITLE: PKCζ Promotes Breast Cancer Invasion by Regulating Expression of E-cadherin and Zonula Occludens-1 (ZO-1) via NFκB-p65 RUNNING TITLE: PKCζ-NFκB Signaling in Breast Cancer
DATA SHEET. Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter calf thymus DNA.
 Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
a b G75 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server.
 a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Tumor stage : I II III IV. well differentiated. moderately differentiated. adenocarcinoma. normal colon (adjacent to cancer) Log (T/H) SLAP mrna level
 moderately differentiated well differentiated Log (T/H) mrna level a Tumor stage : I II III IV.4.4.8 1.2 1.6 2. 2.4 2.8 3.2 N 1 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 # patient b normal colon (adjacent
moderately differentiated well differentiated Log (T/H) mrna level a Tumor stage : I II III IV.4.4.8 1.2 1.6 2. 2.4 2.8 3.2 N 1 2 3 4 5 6 7 8 9 1 11 12 13 14 15 16 17 # patient b normal colon (adjacent
Electron micrograph of phosphotungstanic acid-stained exosomes derived from murine
 1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
Supplemental Table 1. Plasmid Construct backbone. Restriction sites. PCR template. HLA DRα pcdna3.1 Zeo+
 SupplementalTable1 Plasmid Construct backbone HLA DRα Zeo+ Restriction sites PCR template PCRCloningstrategy Primerpairs(F/R)andpairsofprimersforchimericconstructs(N terminalandc terminal codingregion)weregiven
SupplementalTable1 Plasmid Construct backbone HLA DRα Zeo+ Restriction sites PCR template PCRCloningstrategy Primerpairs(F/R)andpairsofprimersforchimericconstructs(N terminalandc terminal codingregion)weregiven
Supplementary Materials for
 immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery
 Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery Presenter: April 12, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. Outline Introduction to GeneCopoeia Lentiviral
Choosing Between Lentivirus and Adeno-associated Virus For DNA Delivery Presenter: April 12, 2017 Ed Davis, Ph.D. Senior Application Scientist GeneCopoeia, Inc. Outline Introduction to GeneCopoeia Lentiviral
Supplementary Figure 1 Cell line TRIB2 status. Supplementary Figure 2 TRIB2 status has no impact on the cell cycle after PI3K inhibition. a. b.
 Supplementary Figure 1 Cell line TRIB2 status. TRIB2 protein expression to determine endogenous expression and to determine the effectiveness of each of our TRIB2 knockdown constructs. Supplementary Figure
Supplementary Figure 1 Cell line TRIB2 status. TRIB2 protein expression to determine endogenous expression and to determine the effectiveness of each of our TRIB2 knockdown constructs. Supplementary Figure
Supporting Information
 Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
The Cellular RNA Helicase DDX1 Interacts with Coronavirus Nonstructural Protein 14 and Enhances Viral Replication
 JOURNAL OF VIROLOGY, Sept. 2010, p. 8571 8583 Vol. 84, No. 17 0022-538X/10/$12.00 doi:10.1128/jvi.00392-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. The Cellular RNA Helicase
JOURNAL OF VIROLOGY, Sept. 2010, p. 8571 8583 Vol. 84, No. 17 0022-538X/10/$12.00 doi:10.1128/jvi.00392-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. The Cellular RNA Helicase
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document
 Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
SUPPLEMENTARY INFORMATION
 Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
SUPPLEMENTARY DATA. Supplementary Table 2. Antibodies used for Immunofluoresence. Supplementary Table 3. Real-time PCR primer sequences.
 Supplementary Table 2. Antibodies used for Immunofluoresence. Antibody Dilution Source Goat anti-pdx1 1:100 R&D Systems Rabbit anti-hnf6 1:100 Santa Cruz Biotechnology Mouse anti-nkx6.1 1:200 Developmental
Supplementary Table 2. Antibodies used for Immunofluoresence. Antibody Dilution Source Goat anti-pdx1 1:100 R&D Systems Rabbit anti-hnf6 1:100 Santa Cruz Biotechnology Mouse anti-nkx6.1 1:200 Developmental
(Stratagene, La Jolla, CA) (Supplemental Fig. 1A). A 5.4-kb EcoRI fragment
 SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Song et al. identified FBXW7 as a new positive regulator for RIG-Itriggered type I IFN signaling pathway. The authors observed
Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Song et al. identified FBXW7 as a new positive regulator for RIG-Itriggered type I IFN signaling pathway. The authors observed
Supplement Figure S1. Real Time PCR analysis of mrna levels of C/EBPα and PU.1 in wild type (WT) and NQO1-null (NQO1-/-) mice.
 competes with 20S proteasome for binding with C/EBP leading to its stabilization and Relative mrna levels Supplement Figure S1. Real Time PCR analysis of mrna levels of C/EBPα and PU.1 in wild type (WT)
competes with 20S proteasome for binding with C/EBP leading to its stabilization and Relative mrna levels Supplement Figure S1. Real Time PCR analysis of mrna levels of C/EBPα and PU.1 in wild type (WT)
Tel: ; Fax: ;
 Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
Name Animal source Vendor Cat # Dilutions
 Supplementary data Table S1. Primary and Secondary antibody sources Devi et al, TXNIP in mitophagy A. Primary Antibodies Name Animal source Vendor Cat # Dilutions 1. TXNIP mouse MBL KO205-2 1:2000 (WB)
Supplementary data Table S1. Primary and Secondary antibody sources Devi et al, TXNIP in mitophagy A. Primary Antibodies Name Animal source Vendor Cat # Dilutions 1. TXNIP mouse MBL KO205-2 1:2000 (WB)
RESEARCH COMMUNICATION. sirna Mediated Silencing of NIN1/RPN12 Binding Protein 1 Homolog Inhibits Proliferation and Growth of Breast Cancer Cells
 RESEARCH COMMUNICATION sirna Mediated Silencing of NIN1/RPN12 Binding Protein 1 Homolog Inhibits Proliferation and Growth of Breast Cancer Cells Wei-Yi Huang, Dong-Hui Chen, Li Ning, Li-Wei Wang* Abstract
RESEARCH COMMUNICATION sirna Mediated Silencing of NIN1/RPN12 Binding Protein 1 Homolog Inhibits Proliferation and Growth of Breast Cancer Cells Wei-Yi Huang, Dong-Hui Chen, Li Ning, Li-Wei Wang* Abstract
Doctoral Degree Program in Marine Biotechnology, College of Marine Sciences, Doctoral Degree Program in Marine Biotechnology, Academia Sinica, Taipei,
 Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Cyclooxygenase 2 facilitates dengue virus replication and serves as a potential target for developing antiviral agents Chun-Kuang Lin 1,2, Chin-Kai Tseng 3,4, Yu-Hsuan Wu 3,4, Chih-Chuang Liaw 1,5, Chun-
Tumor suppressor Spred2 interaction with LC3 promotes autophagosome maturation and induces autophagy-dependent cell death
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Tumor suppressor Spred2 interaction with LC3 promotes autophagosome maturation and induces autophagy-dependent cell death Supplementary
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Tumor suppressor Spred2 interaction with LC3 promotes autophagosome maturation and induces autophagy-dependent cell death Supplementary
Nature Immunology doi: /ni Supplementary Figure 1. Raf-1 inhibition does not affect TLR4-induced type I IFN responses.
 Supplementary Figure 1 Raf-1 inhibition does not affect TLR4-induced type I IFN responses. Real-time PCR analyses of IFNB, ISG15, TRIM5, TRIM22 and APOBEC3G mrna in modcs 6 h after stimulation with TLR4
Supplementary Figure 1 Raf-1 inhibition does not affect TLR4-induced type I IFN responses. Real-time PCR analyses of IFNB, ISG15, TRIM5, TRIM22 and APOBEC3G mrna in modcs 6 h after stimulation with TLR4
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
The C-Terminal Region but Not the Arg-X-Pro Repeat of Epstein-Barr Virus Protein EB2 Is Required for Its Effect on RNA Splicing and Transport
 JOURNAL OF VIROLOGY, May 1999, p. 4090 4100 Vol. 73, No. 5 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The C-Terminal Region but Not the Arg-X-Pro Repeat
JOURNAL OF VIROLOGY, May 1999, p. 4090 4100 Vol. 73, No. 5 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. The C-Terminal Region but Not the Arg-X-Pro Repeat
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/406/ra126/dc1 Supplementary Materials for The microrna mir-485 targets host and influenza virus transcripts to regulate antiviral immunity and restrict viral
www.sciencesignaling.org/cgi/content/full/8/406/ra126/dc1 Supplementary Materials for The microrna mir-485 targets host and influenza virus transcripts to regulate antiviral immunity and restrict viral
Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or
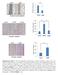 Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or 3xflag-CagA expression vector were wounded using a pipette
Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or 3xflag-CagA expression vector were wounded using a pipette
Supplementary Materials and Methods
 Supplementary Materials and Methods Hepatocyte toxicity assay. Freshly isolated hepatocytes were incubated for overnight with varying concentrations (-25 µm) of sodium glycochenodeoxycholate (GCDC) or
Supplementary Materials and Methods Hepatocyte toxicity assay. Freshly isolated hepatocytes were incubated for overnight with varying concentrations (-25 µm) of sodium glycochenodeoxycholate (GCDC) or
Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved
 1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/366/ra25/dc1 Supplementary Materials for Viral entry route determines how human plasmacytoid dendritic cells produce type I interferons Daniela Bruni, Maxime
www.sciencesignaling.org/cgi/content/full/8/366/ra25/dc1 Supplementary Materials for Viral entry route determines how human plasmacytoid dendritic cells produce type I interferons Daniela Bruni, Maxime
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical
 Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
SUPPLEMENTARY INFORMATION
 a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
