DESIGNING, DOCKING AND TOXICITY STUDIES OF NOVEL HIV-1 PROTEASE INHIBITORS
|
|
|
- Bridget Henry
- 5 years ago
- Views:
Transcription
1 IMPACT: International Journal of Research in Applied atural and Social Sciences (IMPACT: IJRASS) ISS(E): ; ISS(P): Vol. 3, Issue 9, Sep 2015, Impact Journals DESIGIG, DCKIG AD TXICITY STUDIES F VEL HIV-1 PRTEASE IHIBITRS PAVAI ELIPILLA Department of Biotechnology, Acharya agarjuna University, Guntur, Andhra Pradesh, India ABSTRACT HIV virus causes Acquired immune deficiency syndrome (AIDS). HIV virus type-1 protease plays crucial role in the life cycle of the HIV viral particles. So this protein has been targeted as one of the antiretroviral treatment of AIDS, and HIV-1 Protease inhibitors as anti-hiv drugs. Due to the frequent development of drug resistance there is always a need to develop new drugs which are non toxic and efficient inhibitors. The present study aims to focus on designing of 10 non toxic, novel lead molecules which targets HIV 1 protease. And performing docking and toxicity studies to calculate their binding energies, and to predict their toxicity properties. Protein- ligand interactions were studied using HIV type 1 Protease protein, PDB ID - 1HXW extracted from PDB to evaluate the binding efficiency of various molecules towards the active site. And these values were compared with commercially available FDA approved HIV drugs Ritonavir, Saquinavir, Amprenavir, Indinavir, Lopinavir, elfinavir. The final docking and toxicity prediction studies proves that these novel drug molecules were satisfied all drug likeness rules, which are violated by FDA approved drugs, and have the binding energies and predicted inhibitory constant Ki nearly similar to commercially available drugs. KEYWRDS: HIV, HIV-1 Protease, Anti Retroviral Drugs, Drug Designing, Docking, Predicted Toxicity Studies ITRDUCTI Most of the HIV epidemics are caused by HIV type 1, and type 2 variants (Alexander Wlodawer, 2002). HIV viral cells specifically chooses T cells which are having CD4 receptors. Soon after binding to them, the viral genome enters the host cytoplasm and with the help of viral reverse transcriptase enzyme it converts its viral RA into DA then integrates into the host genome which is directed by viral integrase. During translation this genome will start synthesizing HIV viral proteins since it has integrated viral genome within it. Then HIV protease comes into action to process these proteins and activates them which eventually involves in the construction of viral mature proteins. These proteins and viral RA packed together to form new virions and released to infect the new healthy host cells to spread the disease. The most common checkpoints for the development of HIV drugs are, blocking the viral adhesion to host cells, blocking viral fusion and uncoating, inhibit the activity of reverse transcriptase, regulate the gene expression and block the HIV protease function. HIV 1 protease is a dimeric protein with 2 identical monomers. 25th aspartic acid of HIV protease imparts the catalytic activity of this protein (Ashraf Brik,Chi-Huey Wong, 2003). In addition to it, this protein's active site possessa signature amino acid sequence Asp-Thr-Gly which interacts strongly with the substrates or inhibitors. By designing ligands which could be able to bind and inhibit the activity of these catalytic amino acids, we can block the function of HIV protease. So ultimately it stops the processing and maturation of viral proteins thereby inhibits the formation and release of new viral particles. There are 2 ways to block this protein, either the active site of this protein Impact Factor(JCC): This article can be downloaded from
2 140 Pavani Elipilla should be mutated or block it by using some inhibitors (Alexander Wlodawer, Jiri Vondrasek,1998). This present study focused on designing novel ligands which can efficiently block the HIV 1 protease by binding its activesite. These novel ligands binding energies were calculated by docking them into protein and these docking scores and Ki were compared with commercially available FDA approved HIV drugs Ritonavir, Saquinavir, Amprenavir, Indinavir, Lopinavir, elfinavir(joseph J. Eron, Jr.,2000). MATERIALS AD METHDS Ligands Preparation 10 novel non peptide ligands were designed by means of ligand based drug design method. Their structures were drawn using ACD labs chemsketch tool (Figure 1). 6 FDA approved HIV drugs were selected for comparative studies and their structures were also drawn using chemsketch. To remove the steric clashes these ligands were taken for energy minimization and optimization step, by using Prodrg online server (A. W. Schüttelkopf and D. M. F. van Aalten (2004).PRDRG: a tool for high throughput crystallography of protein ligand complexes, Acta Crystallogr D60, ). Then the corrected PDB files of all ligands were collected for further steps. IUPAC names of the 10 novel drugs were mentioned below. 2-amino-5-{[3-amino-4-(4-amino-3-hydroxybutan-2-yl)piperidin-1-yl]oxy}--tert-butyl-3-methyl benzamide [7-amino-6-methyl-2-(4-methylpentan-2-yl)-8-(trifluoromethyl)quinolin-4-yl](4-aminopiperidin-2-yl)methanol -tert-butyl-4-{[3-(4,5-dimethyl-3h-pyrrol-2-yl)phenyl]methyl}-1-(2-hydroxypropyl)piperazine-2-carboxamide -tert-butyl-1-(2-hydroxypropyl)-4-[3-(3h-pyrrol-2-yl)phenoxy]piperazine-2-carboxamide -tert-butyl-2-{4-hydroxy-3-[(4-hydroxyphenyl)methyl]butyl}-decahydroisoquinoline-3-carboxamide 3-[6-(tert-butylamino)-8-(1-methoxypropan-2-yl)-3-methyl-7-oxo-decahydroisoquinolin-2-yl]butanamide 4-[5-(2-amino-4-tert-butylphenyl)pentan-2-yl]quinoline-3,4,6-triamine -tert-butyl-6-(4-oxo-6-phenyl hexan-2-yl)-decahydro-2,6-naphthyridine-3-carboxamide {2-[(tert-butylamino) methyl] 8 (pyridin-2-yl) quinolin-4-yl}(piperidin-2-yl) methanol 1-[4-amino-1-(4-amino-2-cyclopentylbenzenesulfonyl)-5-methylpiperidin-3-yl]-2-(oxolan-3-yl) ethan-1-one The 6 FDA approved drug structures taken for comparative studies were Amprenavir, Indinavir, Lopinavir, elfinavir, Ritonavir, Saquinavir. Index Copernicus Value: Articles can be sent to editor@impactjournals.us
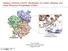
3 Designing, Docking and Toxicity Studies of ovel HIV-1 Protease Inhibitors 141 H 3 C H H H H 2 H H 2 H 3 C H H 3 C H 2 H 2 H 2 F F H 3 C H H 3 C F H H H 2 H C H 3 H 3 C H H 3 C H 3 C H H H H H H 2 H 2 H H H H 2 H H 2 S H 2 CH3 Protein Preparation Figure 1: 10-on Peptide ovel Ligands were Designed. Structures were Drawn in ACD Labs - Chem Sketch Tool (1-10, from Left to Right) In this present study a crystal structure of HIV-1 Protease (PDB: 1HXW) was extracted from RCSB Protein Data Bank. All HETATM were deleted. To repair distorted geometries and to fill the missing atoms in the crystallographic structure, the protein was subjected to protein optimization and energy minimization step by using SPDV tool, where it was processed using 200 steps of steepest descent method. Then this optimized structure was taken to docking step. Molecular Docking Molecular docking was carried out using Autodock4.0 tools. Above optimized protein (1 HXW) was taken and water molecules were removed then polar hydrogens and kollman charges were assigned. Ligand Torsions were allowed to rotate Auto grid generated grid parameter files (.gpf). The grid dimensions on X, Y, Z were set to A. It covers all the active site area of the protein which accommodates the ligands. Through literature it was known that the catalytic amino acid of HIV 1 protease is, 25th asparticacid. Autodock generated dock parameter files (.dpf). Then Lamarckian Genetic Algorithm (LGA) was run by using cygwin which has generated logarithmic files.glg,.dlg. Then autodock was run Impact Factor(JCC): This article can be downloaded from
4 142 Pavani Elipilla to dock the ligand into the protein active site and generates 10 best conformational poses, which can be visualized using UCSF-chimera and Ligplot to get 2d images of docked protein. We can visualize the hydrogen bonds, hydrophobic interactions and vanderwaal interactions between ligand and active site amino acids (Figures 2a -2g). Toxicity Prediction All of the10 novel ligands and FDA approved drugs were checked for the violation of Lipinsky filter, Viber filter, Ghose filter, lead likeness rules (Table 1 and 2). ther properties like LogP and LogS, Protease inhibitory property, Solvent Accessible Surface Area, Polar surface area, Estimated Binding Energy, Inhibition Constant Ki were predicted using online servers like chemicalize, Molinspiration, ALGPS. RESULTS AD DISCUSSIS A series of 10 non peptide novel ligands were designed (Figure 1) and they were along with 6 FDA approved commercially available HIV drugs subjected to docking with HIV-1 protease. All of the docking results and toxicity prediction studies values were incorporated in (Table 1 and 2). The docking studies reveals that all the novel ligands were having strong binding affinity towards the active site of HIV 1 protease. The predicted binding energy of these 10 ligands were in between to Kcal/mol and predicted inhibitory constant Ki values were in between 3.65 nm to nm. (Table 1). Protein-Ligand interactions were visualized using LigPlot software which generated 2d images of the formation of hydrogen bonding between active site amino acids of HIV protease and the ligand molecule (Figures 2a -2g). Ligands Table 1: Calculation of Binding Energy, Inhibitory Constant and ther Properties of ovel Ligands and FDA Approved HIV Drugs Estimated Binding Energy Kcal/mol Estimated Inhibition Constant Ki. H Bonds Formed Molecular weight LGP LGS Index Copernicus Value: Articles can be sent to editor@impactjournals.us Polar Surface Area: Ligand nm Ligand nm Ligand nm Ligand nM Ligand nm Ligand nm Ligand nm Ligand nm Ligand nm Ligand nm Amprenavir nm Indinavir pm Lopinavir nm elfinavir nm Ritonavir um Saquinavir nm And these 10 novel ligands and FDA approved drugs were checked for the violation of Lipinski filter, Viber filter, Ghose filter, like rules (Table 2). ther properties like LogP and LogS, Protease inhibitory property, Solvent Accessible Surface Area, Polar surface area values were predicted using online servers like chemicalize, Molinspiration, ALGPS. A ligand should satisfy all the ADME properties to be a promising drug molecule. These novel ligands were accepted by all the drug likeness filters like veber filter, lipinski rule, ghose filter, muegge filter. A drug molecule will have a poor
5 Designing, Docking and Toxicity Studies of ovel HIV-1 Protease Inhibitors 143 permeability if its polar surface area exceeds 140 Ų, The absorption of a drug will be more when it s LogP (lipophilicity) is < 5, and LogS (aqueous solubility) is 0 (Bob Gotwals, CSSM Chemistry 2009), and Molecular weight should be < 500 dalton. These 10 novel ligands were having polar surface area ranging from Ų to Ų, LogP values were ranging from 0.59 to 3.98, LogS values falls within the range of to and molecular weight ranged between to daltons, which were proven to be having good pharmacological properties. Ligands Table 2: Calculation of Predicted Pharmacological Properties of ovel and FDA Approved Drugs Solvent Accessible Surface Area: Protease Inhibitory Property Lipinski's Rule of five Bio Availability GHSE Lead Likeness MUEGGE VEBER Ligand Yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Ligand yes yes yes yes yes yes Amprenavir no no no no yes no Indinavir no no no no no no Lopinavir no no no no no no elfinavir no yes no no yes yes Ritonavir no no no no no no Saquinavir no no no no no no CCLUSIS It was concluded that, these newly designed 10 ligand molecules can be considered as a better HIV drugs, than the commercially available HIV drugs since they have shown nearly same binding efficiency and better pharmacological properties compared to FDA approved drugs. All the FDA approved 6 HIV protease inhibitors violated the Lipinski rule, veber rule and all other standard drug approval rules, whereas these novel ligands satisfied all the drug likeness rules which reduces the side effects and safe to clinical trials. If these designed novel analogues were synthesized and tested in animal models theywould gives us promising results for the discovery of better drugs. Figure 2a: 1HXW and Ligand 1 Figure 2a: Ligplot 2d Analysis of the Active Site of HIV-1 Protease Complexed with the ovel Ligand-1 and the Impact Factor(JCC): This article can be downloaded from
6 144 Pavani Elipilla Green Dots Represents the Hydrogen Bond Formation between Liand and Active Site Amino Acids of HIV Protease Figure 2b: 1hxw and Ligand 2 Figure 2b: Ligplot 2d Analysis of the Active Site of HIV-1 Protease Complexed with the ovel Ligand-2 and Ligand-2 and the Green Dots Represents the Hydrogen Bond Formation between Ligand 2 AD Active Site Amino Acids of HIV Protease. Figure 2c: 1HXW and Saquinavir Figure 2c: Ligplot 2D analysis of the Active Site of HIV-1 Protease Complexed with anti Retroviral Drug Saquinavir and the Green dots represents the Hydrogen bond Formation between Ligand and active site Amino Acids of HIV Protease. Index Copernicus Value: Articles can be sent to editor@impactjournals.us
7 Designing, Docking and Toxicity Studies of ovel HIV-1 Protease Inhibitors 145 Figure 2d: 1HXW and Ligand 9 Figure 2d: Ligplot 2D Analysis of the Active site of HIV-1 Protease Complexed with Anti Retroviral Drug Ligand 9 and the Green Dots Represents the Hydrogen Bond Formation between Ligand and Active Site Amino acids of HIV protease. Figure 2e: 1HXW and Indinavir Figure 2e: Ligplot 2D Analysis of the Active Site of HIV-1 Protease Complexed with Anti Retroviral Drug Indinavir and the Green Dots Represents the Hydrogen bond Formation between Ligand and active site amino acids of HIV Protease. Impact Factor(JCC): This article can be downloaded from
8 146 Pavani Elipilla Figure 2f: 1HXW and Ligand 10 Figure 2f: Schematic 2D Representation of the Interation between HIV Protease Active Site Amino Acids with ovel Ligand-10, Image was Generated Using Accelrys Discovery Studio. Figure 2g: 1HXW and Ligand 6 Figure 2g: Schematic 2D Representation of the Interation between HIV Protease Active Site Amino Acids with ovel Ligand-10, Image was Generated Using Accelrys Discovery Studio REFERECES 1. Ashraf Brik and Chi-Huey Wong, HIV-1 protease: mechanism and drug discovery, rg, Bio mol Chem.,2003,1, 5-14, DI: / b208248a 2. Alexander Wlodawer and Jiri Vondrasek, Inhibitors of HIV-1 protease: A Major Success of Structure-Assisted Drug Design, Annu. Rev. Biophys.Biomol. Struct : Alexander Wlodawer, Rational approach to aids drug design through structural biology, Annu. Rev. Med : Bob gotwals, Case Study: Drug Solubility, CSSM Chemistry, Feb,12, Joseph J. Eron, Jr., HIV-1 Protease Inhibitors, Clinical Infectious Diseases 2000; 30 (Suppl 2):S Index Copernicus Value: Articles can be sent to editor@impactjournals.us
INTERACTIONS OF GANODERIOL-F WITH ASPARTIC PROTEASES OF HIV AND PLASMEPSIN FOR ANTI-HIV AND ANTI-MALARIA DISCOVERY
 Innovare Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 6, Issue 5, 2014 Original Article INTERACTIONS OF GANODERIOL-F WITH ASPARTIC PROTEASES OF HIV
Innovare Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 6, Issue 5, 2014 Original Article INTERACTIONS OF GANODERIOL-F WITH ASPARTIC PROTEASES OF HIV
Docking Analysis of Darunavir as HIV Protease Inhibitors
 Journal of Computational Methods in Molecular Design, 2012, 2 (1):39-43 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Docking Analysis
Journal of Computational Methods in Molecular Design, 2012, 2 (1):39-43 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Docking Analysis
Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids
 www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Pelagia Research Library. Docking studies on PPARγ of novel α-phenoxy phenyl propionic acid derivatives as antidiabetic agents
 Available online at www.pelagiaresearchlibrary.com Der Pharmacia Sinica, 2011, 2 (2): 327-332 ISSN: 0976-8688 CDEN (USA): PSHIBD Docking studies on PPARγ of novel α-phenoxy phenyl propionic acid derivatives
Available online at www.pelagiaresearchlibrary.com Der Pharmacia Sinica, 2011, 2 (2): 327-332 ISSN: 0976-8688 CDEN (USA): PSHIBD Docking studies on PPARγ of novel α-phenoxy phenyl propionic acid derivatives
Available Online through
 ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors
 www.bioinformation.net Hypothesis Volume 9(11) Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors Monika 1 *, Janmeet Kour 1 & Kulwinder Singh 2 1Department of Biotechnology,
www.bioinformation.net Hypothesis Volume 9(11) Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors Monika 1 *, Janmeet Kour 1 & Kulwinder Singh 2 1Department of Biotechnology,
A Combinatorial approach: To design inhibitory molecules on Hemagglutinin protein of H1N1 virus (Swine Flu)
 www.bioinformation.net Hypothesis Volume 9(11) A Combinatorial approach: To design inhibitory molecules on Hemagglutinin protein of H1N1 virus (Swine Flu) Chekkara Venkata Satya Siva Prasad*, Kamal Kumar
www.bioinformation.net Hypothesis Volume 9(11) A Combinatorial approach: To design inhibitory molecules on Hemagglutinin protein of H1N1 virus (Swine Flu) Chekkara Venkata Satya Siva Prasad*, Kamal Kumar
Final Report. Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD. Wendy Fong ( 方美珍 ) CNIC; Beijing, China
 Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma (PPARγ)
 3rd International Conference on Computation for Science and Technology (ICCST-3) Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma
3rd International Conference on Computation for Science and Technology (ICCST-3) Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma
NIH Public Access Author Manuscript J Am Chem Soc. Author manuscript; available in PMC 2008 September 29.
 NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
QSAR Analysis of Chromone Derivatives as HIV-1 Protease Inhibitors
 Original Article Mahidol University Journal of Pharmaceutical Sciences 2013; 40 (3), 8-15 QSAR Analysis of Chromone Derivatives as HIV-1 Protease Inhibitors W. Samee a and J. Ungwitayatorn b * a Faculty
Original Article Mahidol University Journal of Pharmaceutical Sciences 2013; 40 (3), 8-15 QSAR Analysis of Chromone Derivatives as HIV-1 Protease Inhibitors W. Samee a and J. Ungwitayatorn b * a Faculty
Margaret A. Daugherty. Announcements! Fall Michaelis Menton Kinetics and Inhibition. Lecture 14: Enzymes & Kinetics III
 Lecture 14: Enzymes & Kinetics III Michaelis Menton Kinetics and Inhibition Margaret A. Daugherty Fall 2004 Announcements! Monday 10/11 lecture: starts at 10:15; Taught by Dr. Stephen Everse o ffice our/review
Lecture 14: Enzymes & Kinetics III Michaelis Menton Kinetics and Inhibition Margaret A. Daugherty Fall 2004 Announcements! Monday 10/11 lecture: starts at 10:15; Taught by Dr. Stephen Everse o ffice our/review
Workshop on Analysis and prediction of contacts in proteins
 Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents
 Research Article In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents C. N. Hemalatha 1, Elancheziyan K 1, D. Pavithra 2, M. Vijey Aanandhi 3 * ABSTRACT
Research Article In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents C. N. Hemalatha 1, Elancheziyan K 1, D. Pavithra 2, M. Vijey Aanandhi 3 * ABSTRACT
Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II inhibitors
 International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol. 3, No.3,pp 1622-1627, July-Sept 2011 Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II
International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol. 3, No.3,pp 1622-1627, July-Sept 2011 Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II
Analysis of Resistance to Human Immunodeficiency Virus Protease Inhibitors Using Molecular Mechanics and Machine Learning Strategies
 American Medical Journal 1 (2): 126-132, 2010 ISSN 1949-0070 2010 Science Publications Analysis of Resistance to Human Immunodeficiency Virus Protease Inhibitors Using Molecular Mechanics and Machine Learning
American Medical Journal 1 (2): 126-132, 2010 ISSN 1949-0070 2010 Science Publications Analysis of Resistance to Human Immunodeficiency Virus Protease Inhibitors Using Molecular Mechanics and Machine Learning
News. Volume 5: June 12th, The Research Team We re all members of Prof. Olson s Molecular Graphics Lab
 FightAIDS@Home News Volume 5: June 12th, 2008 Alex Perry man Ga rret t Mo r ri s Stefano Art Forl i Ol son Alex Gi l let The FightAIDS@Home Research Team We re all members of Prof. Olson s Molecular Graphics
FightAIDS@Home News Volume 5: June 12th, 2008 Alex Perry man Ga rret t Mo r ri s Stefano Art Forl i Ol son Alex Gi l let The FightAIDS@Home Research Team We re all members of Prof. Olson s Molecular Graphics
Exploring Energy and Fitness Landscapes of Proteins
 Exploring Energy and Fitness Landscapes of Proteins! Computer Simulations in Chemistry and Biophysics! Protein-Ligand Binding: structure based drug design! HIV Family and Kinase Family Protein Targets!
Exploring Energy and Fitness Landscapes of Proteins! Computer Simulations in Chemistry and Biophysics! Protein-Ligand Binding: structure based drug design! HIV Family and Kinase Family Protein Targets!
Term Definition Example Amino Acids
 Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
Molecular Docking Studies on Pyrazolopyrimidine and their Derivatives as Human Phosphoinositide 3-Kinase Inhibitors
 Cloud Publications International Journal of Advanced Bioinformatics and Computational Biology 2012, Volume 1, Issue 1, pp. 1-11, Article ID Med-30 Research Article Open Access Molecular Docking Studies
Cloud Publications International Journal of Advanced Bioinformatics and Computational Biology 2012, Volume 1, Issue 1, pp. 1-11, Article ID Med-30 Research Article Open Access Molecular Docking Studies
News. Volume 8: November 2, 2009
 FightAIDS@Home News Volume 8: November 2, 2009 The FightAIDS@Home Project uses the volunteered computing power of the World Community Grid to test candidate compounds against the variations (or mutants
FightAIDS@Home News Volume 8: November 2, 2009 The FightAIDS@Home Project uses the volunteered computing power of the World Community Grid to test candidate compounds against the variations (or mutants
ACQUIRED IMMUNODEFICIENCY SYNDROME AND ITS OCULAR COMPLICATIONS
 ACQUIRED IMMUNODEFICIENCY SYNDROME AND ITS OCULAR COMPLICATIONS Acquired immunodeficiency syndrome (AIDS ) is an infectious disease caused by a retrovirus, the human immunodeficiency virus(hiv). AIDS is
ACQUIRED IMMUNODEFICIENCY SYNDROME AND ITS OCULAR COMPLICATIONS Acquired immunodeficiency syndrome (AIDS ) is an infectious disease caused by a retrovirus, the human immunodeficiency virus(hiv). AIDS is
Saquinavir is a medicament which is
 Saquinavir Tobias Mongelli, Niklas Oefner, Jonathan Ritter, Melina Roemer Technische Universität Darmstadt tobias.mongelli@gmx.de Abstract Human immunodeficiency virus (HIV) is a subgroup of a retrovirus
Saquinavir Tobias Mongelli, Niklas Oefner, Jonathan Ritter, Melina Roemer Technische Universität Darmstadt tobias.mongelli@gmx.de Abstract Human immunodeficiency virus (HIV) is a subgroup of a retrovirus
PREDICTION AND COMPARISON OF HIV-1 PROTEASE INHIBITOR BINDING ENERGIES BY VARIOUS MOLECULAR DOCKING METHODS
 PREDICTION AND COMPARISON OF HIV-1 PROTEASE INHIBITOR BINDING ENERGIES BY VARIOUS MOLECULAR DOCKING METHODS THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF Master of Technology
PREDICTION AND COMPARISON OF HIV-1 PROTEASE INHIBITOR BINDING ENERGIES BY VARIOUS MOLECULAR DOCKING METHODS THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF Master of Technology
In silico identification of Novel HIV- Protease inhibitors (PIs) using ZINC drug Database
 In silico identification of Novel HIV- Protease inhibitors (PIs) using ZINC drug Database K.K. Srivastava 1, Shubha Srivastava 2, Tanweer Alam 1, Rituraj 1* 1 Department of Chemistry, Vinoba Bhave University,
In silico identification of Novel HIV- Protease inhibitors (PIs) using ZINC drug Database K.K. Srivastava 1, Shubha Srivastava 2, Tanweer Alam 1, Rituraj 1* 1 Department of Chemistry, Vinoba Bhave University,
Controlling ADME through Chemical Design. Marty Mulvihill Chris Vulpe
 Controlling ADME through Chemical Design Marty Mulvihill Chris Vulpe ADME Chemical Processes in ADME Wang and Skolnik, Chemistry and Biodiversity, 2009, 1887. Controlling toxicity through ADME Toward molecular
Controlling ADME through Chemical Design Marty Mulvihill Chris Vulpe ADME Chemical Processes in ADME Wang and Skolnik, Chemistry and Biodiversity, 2009, 1887. Controlling toxicity through ADME Toward molecular
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Bioinformation Volume 5
 Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER
 ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase in the treatment of dermatophytoses by molecular docking
 International Journal of Advanced Research in Medical Sciences 2017 Volume 1 Issue 1 Pages: 1-5 Original Research Paper Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase
International Journal of Advanced Research in Medical Sciences 2017 Volume 1 Issue 1 Pages: 1-5 Original Research Paper Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase
Int. J. Pharm. Sci. Rev. Res., 40(2), September October 2016; Article No. 34, Pages:
 Research Article In silico Screening of Various Natural Compounds to Predict the Potential Inhibitors that Target the HIV-1 Protease Sub Type A Jitender Singh, Ashvinder Raina* Dept of Experimental Medicine
Research Article In silico Screening of Various Natural Compounds to Predict the Potential Inhibitors that Target the HIV-1 Protease Sub Type A Jitender Singh, Ashvinder Raina* Dept of Experimental Medicine
PHARMACOPHORE MODELING
 CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
Creating a high-throughput screening database to propose ligands capable of modulating the HIV-1 protease receptor
 Vol. 5(7), pp. 224-234, July, 2013 DOI 10.5897/JAHR12.056 ISSN 2141-2359 2013 Academic Journals http://www.academicjournals.org/jahr Journal of AIDS and HIV Research Full Length Research Paper Creating
Vol. 5(7), pp. 224-234, July, 2013 DOI 10.5897/JAHR12.056 ISSN 2141-2359 2013 Academic Journals http://www.academicjournals.org/jahr Journal of AIDS and HIV Research Full Length Research Paper Creating
MedChem 401~ Retroviridae. Retroviridae
 MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
Molecular Dynamics of HIV-1 Reverse Transcriptase
 Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Asian Journal of Biomedical and Pharmaceutical Sciences 1 (6) 2012, 01-05
 Asian Journal of Biomedical and Pharmaceutical Sciences 1 (6) 2012, 01-05 RESEARCH ARTICLE ISSN 2249-622X Virtual Screening of Xanthones in Combating Malaria Targeting Plasmodium Falciparum Erythrocyte
Asian Journal of Biomedical and Pharmaceutical Sciences 1 (6) 2012, 01-05 RESEARCH ARTICLE ISSN 2249-622X Virtual Screening of Xanthones in Combating Malaria Targeting Plasmodium Falciparum Erythrocyte
Computational Drug Discovery of Potential Aldose Reductase Inhibitors Using in silico Studies
 Computational Drug Discovery of Potential Aldose Reductase Inhibitors Using in silico Studies Arumugam Madeswaran*, Muthuswamy Umamaheswari, Kuppusamy Asokkumar, Thirumalaisamy Sivashanmugam, Varadharajan
Computational Drug Discovery of Potential Aldose Reductase Inhibitors Using in silico Studies Arumugam Madeswaran*, Muthuswamy Umamaheswari, Kuppusamy Asokkumar, Thirumalaisamy Sivashanmugam, Varadharajan
LESSON 4.6 WORKBOOK. Designing an antiviral drug The challenge of HIV
 LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
Computer Aided Docking Studies on Anticancer Drugs for Breast Cancer
 International Journal of Information & Computation Technology. ISSN 0974-2239 Volume 2, Number 1 (2012), pp. 49-55 International Research Publications House http://www. ripublication.com Computer Aided
International Journal of Information & Computation Technology. ISSN 0974-2239 Volume 2, Number 1 (2012), pp. 49-55 International Research Publications House http://www. ripublication.com Computer Aided
Binding property of HIV p24 and Reverse transcriptase by chalcones from Pongamia pinnata seeds
 www.bioinformation.net Volume 14(6) Hypothesis Binding property of HIV p24 and Reverse transcriptase by chalcones from Pongamia pinnata seeds Manikannan Mathaiyan 1, Arumugam Suresh 1*, Rangasamy Balamurugan
www.bioinformation.net Volume 14(6) Hypothesis Binding property of HIV p24 and Reverse transcriptase by chalcones from Pongamia pinnata seeds Manikannan Mathaiyan 1, Arumugam Suresh 1*, Rangasamy Balamurugan
Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters
 Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters Chen-Hsiang Shen 1, Yuan-Fang Wang 1, Andrey Y. Kovalevsky 1, *, Robert W. Harrison 1,2 and Irene T.
Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters Chen-Hsiang Shen 1, Yuan-Fang Wang 1, Andrey Y. Kovalevsky 1, *, Robert W. Harrison 1,2 and Irene T.
Simulated Docking of Darunavir with the HIV-1 L76V Mutant Protease Active Site
 Simulated Docking of Darunavir with the HIV-1 L76V Mutant Protease Active Site Jack K. Horner P.O. Box 266 Los Alamos NM 87544 USA Abstract Darunavir is a second-generation HIV-1 protease inhibitor. Here
Simulated Docking of Darunavir with the HIV-1 L76V Mutant Protease Active Site Jack K. Horner P.O. Box 266 Los Alamos NM 87544 USA Abstract Darunavir is a second-generation HIV-1 protease inhibitor. Here
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
In Silico docking studies of lipoxygenase inhibitory activity of commercially available flavonoids
 Available online at www.scholarsresearchlibrary.com J. Comput. Method. Mol. Design, 2011, 1 (4):65-72 (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA In Silico docking
Available online at www.scholarsresearchlibrary.com J. Comput. Method. Mol. Design, 2011, 1 (4):65-72 (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA In Silico docking
Molecular Dynamics of Sialic Acid Analogues and their Interaction with Influenza Hemagglutinin
 Research Papers www.ijpsonline.com Molecular Dynamics of Sialic Acid Analogues and their Interaction with Influenza Hemagglutinin T. FEMLIN BLESSIA, V. S. RAPHEAL 1 AND D. J. S. SHARMILA* Department of
Research Papers www.ijpsonline.com Molecular Dynamics of Sialic Acid Analogues and their Interaction with Influenza Hemagglutinin T. FEMLIN BLESSIA, V. S. RAPHEAL 1 AND D. J. S. SHARMILA* Department of
Available online Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2016, 8(4):912-918 Research Article ISS : 0975-7384 CODE(USA) : JCPRC5 Synthesis of -(2-nitrophenyl) pyrazine-2-carboxamide
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2016, 8(4):912-918 Research Article ISS : 0975-7384 CODE(USA) : JCPRC5 Synthesis of -(2-nitrophenyl) pyrazine-2-carboxamide
We are IntechOpen, the world s leading publisher of Open Access books Built by scientists, for scientists. International authors and editors
 We are IntechOpen, the world s leading publisher of Open Access books Built by scientists, for scientists 4,100 116,000 120M Open access books available International authors and editors Downloads Our
We are IntechOpen, the world s leading publisher of Open Access books Built by scientists, for scientists 4,100 116,000 120M Open access books available International authors and editors Downloads Our
fibromyalgia and early genetic studies revealed that the gene SLC6A4 showed an increased frequency among patients
 ISSN: 0975-766X CODEN: IJPTFI Available Online through Research Article www.ijptonline.com IN-SILICO SCREENING FOR THE IDENTIFICATION OF SLC6A4 DRUG TARGETS AND ANTI- RHEUMATIC ACTIVITY OF PHYTOLIGANDS
ISSN: 0975-766X CODEN: IJPTFI Available Online through Research Article www.ijptonline.com IN-SILICO SCREENING FOR THE IDENTIFICATION OF SLC6A4 DRUG TARGETS AND ANTI- RHEUMATIC ACTIVITY OF PHYTOLIGANDS
Lab 5: Proteins and the small molecules that love them (AKA Computer Modeling with PyMol #2)
 Lab 5: Proteins and the small molecules that love them (AKA Computer Modeling with PyMol #2) Goals: The objective of this lab is to provide you with an understanding of: 1. Catalysis 2. Small molecule
Lab 5: Proteins and the small molecules that love them (AKA Computer Modeling with PyMol #2) Goals: The objective of this lab is to provide you with an understanding of: 1. Catalysis 2. Small molecule
Design and Molecular Docking Studies of luteolin derivatives, from Biebersteinia multifida DC., as novel HMG-CoA reductase inhibitors
 International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol.4, No.2, pp 733-738, April-June 2012 Design and Molecular Docking Studies of luteolin derivatives, from Biebersteinia
International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol.4, No.2, pp 733-738, April-June 2012 Design and Molecular Docking Studies of luteolin derivatives, from Biebersteinia
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies
 Research Article Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies Pawan Gupta 1,2 *, Prabha Garg 2 ABSTRACT Background and Aim: The
Research Article Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies Pawan Gupta 1,2 *, Prabha Garg 2 ABSTRACT Background and Aim: The
Analysis of protein modeling for envelope glycoprotein GP120 for HIV via bioinformatics approaches
 International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
Research Article. Insilico predictions of inhibitors of novel statin structural analogues with HMG-CoA reductase
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(3):942-950 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico predictions of inhibitors of novel statin
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(3):942-950 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico predictions of inhibitors of novel statin
STRUCTURE BASED DESIGN OF FEW SUBSTITUTED PIPERIDONES AS DIPEPTIDYL PEPTIDASE INHIBITORS
 Academic ciences International Journal of Pharmacy and Pharmaceutical ciences I- 0975-1491 Vol 4, uppl 5, 2012 Research Article TRUCTURE BAED DEIG EW UBTITUTED PIPERIDE A DIPEPTIDYL PEPTIDAE IIBITR URAV
Academic ciences International Journal of Pharmacy and Pharmaceutical ciences I- 0975-1491 Vol 4, uppl 5, 2012 Research Article TRUCTURE BAED DEIG EW UBTITUTED PIPERIDE A DIPEPTIDYL PEPTIDAE IIBITR URAV
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Time Minimizing in anticancer drug discovery. Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia
 Time Minimizing in anticancer drug discovery Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia Abstract Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia
Time Minimizing in anticancer drug discovery Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia Abstract Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia
Microbiology Review Division of Antiviral Drug Products (HFD-530)
 Microbiology Review Division of Antiviral Drug Products (HFD-530) NDA#: 21-785 Serial #: 000 Reviewer: N. Battula Date submitted: June 17,2004 Date received: June 18,2004 Date assigned: July 01,2004 Date
Microbiology Review Division of Antiviral Drug Products (HFD-530) NDA#: 21-785 Serial #: 000 Reviewer: N. Battula Date submitted: June 17,2004 Date received: June 18,2004 Date assigned: July 01,2004 Date
BMC Bioinformatics. Open Access. Abstract. BioMed Central
 BMC Bioinformatics BioMed Central Research Molecular docking studies of dithionitrobenzoic and its related compounds to protein disulfide isomerase: computational screening of inhibitors to HIV-1 entry
BMC Bioinformatics BioMed Central Research Molecular docking studies of dithionitrobenzoic and its related compounds to protein disulfide isomerase: computational screening of inhibitors to HIV-1 entry
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Structure-based analysis of curcumin and conventional drugs targeting tumor-inducing protein PHF20
 www.bioinformation.net Volume 14(9) Hypothesis Structure-based analysis of curcumin and conventional drugs targeting tumor-inducing protein PHF20 Vibha Agrawal 1, Aradhya Mishra 1, Shivani Tiwari 1, Akhileshwar
www.bioinformation.net Volume 14(9) Hypothesis Structure-based analysis of curcumin and conventional drugs targeting tumor-inducing protein PHF20 Vibha Agrawal 1, Aradhya Mishra 1, Shivani Tiwari 1, Akhileshwar
DEVELOPMENT AND VALIDATION OF RP-HPLC METHOD FOR THE ESTIMATION OF ANTIRETROVIRAL DRUGS IN TABLET DOSAGE FORMS
 Int. J. Chem. Sci.: 14(4), 2016, 2461-2466 ISSN 0972-768X www.sadgurupublications.com DEVELOPMENT AND VALIDATION OF RP-HPLC METHOD FOR THE ESTIMATION OF ANTIRETROVIRAL DRUGS IN TABLET DOSAGE FORMS K. PARAMESWARA
Int. J. Chem. Sci.: 14(4), 2016, 2461-2466 ISSN 0972-768X www.sadgurupublications.com DEVELOPMENT AND VALIDATION OF RP-HPLC METHOD FOR THE ESTIMATION OF ANTIRETROVIRAL DRUGS IN TABLET DOSAGE FORMS K. PARAMESWARA
HIV - Life cycle. HIV Life Cyle
 Human Immunodeficiency Virus Retrovirus - integrated into host genome ne single-strand RA 7,000 bases HIV1 > HIV2 > HIV0 Pathology Destruction of CD4+ T lymphocytes Loss of immune function pportunistic
Human Immunodeficiency Virus Retrovirus - integrated into host genome ne single-strand RA 7,000 bases HIV1 > HIV2 > HIV0 Pathology Destruction of CD4+ T lymphocytes Loss of immune function pportunistic
Identification of Structural Mechanisms of HIV-1 Protease Specificity Using Computational Peptide Docking: Implications for Drug Resistance
 Article Identification of Structural Mechanisms of HIV-1 Protease Specificity Using Computational Peptide Docking: Implications for Drug Resistance Sidhartha Chaudhury 1 and Jeffrey J. Gray 1,2, * 1 Program
Article Identification of Structural Mechanisms of HIV-1 Protease Specificity Using Computational Peptide Docking: Implications for Drug Resistance Sidhartha Chaudhury 1 and Jeffrey J. Gray 1,2, * 1 Program
CHAPTER 9: CATALYTIC STRATEGIES. Chess vs Enzymes King vs Substrate
 CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
Austroinulin from Stevia rebaudiana a Potential Agonist of the Peroxisome Proliferator-activated Receptor (PPARγ): an in-silico Study.
 Research Article http://www.alliedacademies.org/systems-biology-proteome-research/ Austroinulin from Stevia rebaudiana a Potential Agonist of the Peroxisome Proliferator-activated Receptor (PPARγ): an
Research Article http://www.alliedacademies.org/systems-biology-proteome-research/ Austroinulin from Stevia rebaudiana a Potential Agonist of the Peroxisome Proliferator-activated Receptor (PPARγ): an
The Binding Mode of by Electron Crystallography
 The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
Molecular Docking Studies of Shc1 Protein: A Drug Target to Treat Obesity
 IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 9, Issue 6 Ver. III (Nov -Dec. 2014), PP 63-68 Molecular Docking Studies of Shc1 Protein: A Drug
IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 9, Issue 6 Ver. III (Nov -Dec. 2014), PP 63-68 Molecular Docking Studies of Shc1 Protein: A Drug
REPORT DOCUMENTATION PAGE (SF298) (Continuation Sheet)
 REPRT DCUMENTATIN PAGE (SF298) (Continuation Sheet) Key Features of the Report: (i) (ii) (iii) (iv) We have synthesized a polymer that could form both unimolecular micelles and inverse micelles, depending
REPRT DCUMENTATIN PAGE (SF298) (Continuation Sheet) Key Features of the Report: (i) (ii) (iii) (iv) We have synthesized a polymer that could form both unimolecular micelles and inverse micelles, depending
Trilateral Project WM4
 ANNEX 2: Comments of the JPO Trilateral Project WM4 Comparative studies in new technologies Theme: Comparative study on protein 3-dimensional (3-D) structure related claims 1. Introduction As more 3-D
ANNEX 2: Comments of the JPO Trilateral Project WM4 Comparative studies in new technologies Theme: Comparative study on protein 3-dimensional (3-D) structure related claims 1. Introduction As more 3-D
A Random Forest Model for the Analysis of Chemical Descriptors for the Elucidation of HIV 1 Protease Protein Ligand Interactions
 A Random Forest Model for the Analysis of Chemical Descriptors for the Elucidation of HIV 1 Protease Protein Ligand Interactions Gene M. Ko, A. Srinivas Reddy, Sunil Kumar, Barbara A. Bailey, and Rajni
A Random Forest Model for the Analysis of Chemical Descriptors for the Elucidation of HIV 1 Protease Protein Ligand Interactions Gene M. Ko, A. Srinivas Reddy, Sunil Kumar, Barbara A. Bailey, and Rajni
Week 5 Section. Junaid Malek, M.D.
 Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Proteomics and Rational Drug Design. Dr Gilda Tachedjian Molecular Interactions Group Dec 2006
 Proteomics and Rational Drug Design Dr Gilda Tachedjian Molecular Interactions Group Dec 2006 Topics to be Covered in this Presentation What is Proteomics? Tools used to study the Proteome. Application
Proteomics and Rational Drug Design Dr Gilda Tachedjian Molecular Interactions Group Dec 2006 Topics to be Covered in this Presentation What is Proteomics? Tools used to study the Proteome. Application
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
7.014 Problem Set 2 Solutions
 7.014 Problem Set 2 Solutions Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in at the box outside 68-120 by 11:45 Friday, February
7.014 Problem Set 2 Solutions Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in at the box outside 68-120 by 11:45 Friday, February
Chapter 3: Amino Acids and Peptides
 Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Separation of Macrocyclic Lactones (Avermectins) on FLARE C18 MM & FLARE C18+ Columns
 Separation of Macrocyclic Lactones (Avermectins) on FLARE C8 MM & FLARE C8+ Columns Introduction Diamond Analytics Technical Note: T05- Avermectins are a series of 6-membered macrocyclic lactone derivatives
Separation of Macrocyclic Lactones (Avermectins) on FLARE C8 MM & FLARE C8+ Columns Introduction Diamond Analytics Technical Note: T05- Avermectins are a series of 6-membered macrocyclic lactone derivatives
HIV-1 REVERSE TRANSCRIPTASE INHIBIT THE NON-NUCLEOSIDE REVERSE TRANSCRIPTASE BY INSILICO METHOD
 ISS: 2321-676X Kanaga Durga, et al. / star Bio. Info. 06 (2013) IV-1 REVERSE TRASCRIPTASE IIBIT TE -UCLESIDE REVERSE TRASCRIPTASE BY ISILIC METD V. Kanaga Durga 1, T. Dhinesh Kumar 2, Dr. D. Velmurugan
ISS: 2321-676X Kanaga Durga, et al. / star Bio. Info. 06 (2013) IV-1 REVERSE TRASCRIPTASE IIBIT TE -UCLESIDE REVERSE TRASCRIPTASE BY ISILIC METD V. Kanaga Durga 1, T. Dhinesh Kumar 2, Dr. D. Velmurugan
Journal of Chemical and Pharmaceutical Research, 2012, 4(4): Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2012, 4(4):2037-2042 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico designing and development of potent drug
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2012, 4(4):2037-2042 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico designing and development of potent drug
Biochemical and Biophysical Research Communications 305 (2003)
 Biochemical and Biophysical Research Communications 305 (2003) 322 326 BBRC www.elsevier.com/locate/ybbrc Comprehensive mutagenesis of HIV-1 protease: a computational geometry approach q Majid Masso and
Biochemical and Biophysical Research Communications 305 (2003) 322 326 BBRC www.elsevier.com/locate/ybbrc Comprehensive mutagenesis of HIV-1 protease: a computational geometry approach q Majid Masso and
Structural Analysis of TCRpMHC Complexes Using Computational Tools. Feroze Mohideen Briarcliff High School
 Structural Analysis of TCRpMHC Complexes Using Computational Tools Feroze Mohideen Briarcliff High School TCR-pMHC Complexes Peptide Structure of TCR-pMHC complex PDB AO7 Crossreactivity is the ability
Structural Analysis of TCRpMHC Complexes Using Computational Tools Feroze Mohideen Briarcliff High School TCR-pMHC Complexes Peptide Structure of TCR-pMHC complex PDB AO7 Crossreactivity is the ability
Natural Flavonoids as CDK5 inhibitors: A Homology modeling and Molecular Docking Study
 International Journal of ChemTech Research CDEN (USA): IJCRGG, ISSN: 0974-4290, ISSN(nline):2455-9555 Vol.9, No.09 pp 210-215, 2016 Natural Flavonoids as CDK5 inhibitors: A Homology modeling and Molecular
International Journal of ChemTech Research CDEN (USA): IJCRGG, ISSN: 0974-4290, ISSN(nline):2455-9555 Vol.9, No.09 pp 210-215, 2016 Natural Flavonoids as CDK5 inhibitors: A Homology modeling and Molecular
Computational molecular docking studies on anticancer drugs
 S734 Asian Pacific Journal of Tropical Disease (2012)S734-S738 Contents lists available at ScienceDirect Asian Pacific Journal of Tropical Disease journal homepage: www.elsevier.com/locate/apjtd Document
S734 Asian Pacific Journal of Tropical Disease (2012)S734-S738 Contents lists available at ScienceDirect Asian Pacific Journal of Tropical Disease journal homepage: www.elsevier.com/locate/apjtd Document
Article Teaching Foundational Topics and Scientific Skills in Biochemistry Within the Conceptual Framework of HIV Protease hs
 Article Teaching Foundational Topics and Scientific Skills in Biochemistry Within the Conceptual Framework of HIV Protease hs R. Jeremy Johnson* From the Department of Chemistry, Butler University, Indianapolis,
Article Teaching Foundational Topics and Scientific Skills in Biochemistry Within the Conceptual Framework of HIV Protease hs R. Jeremy Johnson* From the Department of Chemistry, Butler University, Indianapolis,
EGFRIndb: Epidermal Growth Factor Receptor Inhibitor database
 EGFRIndb: Epidermal Growth Factor Receptor Inhibitor database Inderjit S Yadav 1,3, Harinder Singh 2, Mohd. Imran Khan 1, Ashok Chaudhury 3, G.P.S. Raghava 2 and Subhash M. Agarwal 1 * 1 Bioinformatics
EGFRIndb: Epidermal Growth Factor Receptor Inhibitor database Inderjit S Yadav 1,3, Harinder Singh 2, Mohd. Imran Khan 1, Ashok Chaudhury 3, G.P.S. Raghava 2 and Subhash M. Agarwal 1 * 1 Bioinformatics
DOCKING STUDY OF METHYL HESPERIDIN AS NUCLEOSIDE REVERSE TRANSCRIPTASE INHIBITOR
 International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 10, Issue 3, 2018 Original Article DOCKING STUDY OF METHYL HESPERIDIN AS NUCLEOSIDE REVERSE TRANSCRIPTASE INHIBITOR HAFRIZAL
International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 10, Issue 3, 2018 Original Article DOCKING STUDY OF METHYL HESPERIDIN AS NUCLEOSIDE REVERSE TRANSCRIPTASE INHIBITOR HAFRIZAL
APORPHINE ALKALOIDS AS INHIBITORS FOR MALONYL-COA: ACYL CARRIER PROTEIN TRANSACYLASE (MCAT) FROM HELICOBACTER PYLORI
 Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 4, Issue 3, 2012 Research Article APORPHINE ALKALOIDS AS INHIBITORS FOR MALONYL-COA: ACYL CARRIER PROTEIN
Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 4, Issue 3, 2012 Research Article APORPHINE ALKALOIDS AS INHIBITORS FOR MALONYL-COA: ACYL CARRIER PROTEIN
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Drug resistance mechanisms and drug design strategies for human immunodeficiency virus and hepatitis c virus proteases
 Wayne State University Wayne State University Dissertations 1-1-2012 Drug resistance mechanisms and drug design strategies for human immunodeficiency virus and hepatitis c virus proteases Yong Wang Wayne
Wayne State University Wayne State University Dissertations 1-1-2012 Drug resistance mechanisms and drug design strategies for human immunodeficiency virus and hepatitis c virus proteases Yong Wang Wayne
IN-SILICO APPROACH FOR THE ASSESSMENT OF ORAL CANCER PROPERTY ON LIMONIA ACIDISSIMA
 IJPSR (2016), Vol. 7, Issue 3 (Research Article) Received on 16 September, 2015; received in revised form, 29 October, 2015; accepted, 12 December, 2015; published 01 March, 2016 IN-SILICO APPROACH FOR
IJPSR (2016), Vol. 7, Issue 3 (Research Article) Received on 16 September, 2015; received in revised form, 29 October, 2015; accepted, 12 December, 2015; published 01 March, 2016 IN-SILICO APPROACH FOR
M Lakshmi Vasavi Devi et al /J. Pharm. Sci. & Res. Vol.3(4), 2011,
 ANALOGS OF CARBIDOPA: IN SILICO DESIGN & DEVELOPMENT OF NOVEL DOPA DECARBOXYLASE INHIBITORS IN THE TREATMENT OF PARKINSON S DISEASE M Lakshmi Vasavi Devi 1 *, Deepak Reddy Gade 2, T Gopi Raju 3, Pavan
ANALOGS OF CARBIDOPA: IN SILICO DESIGN & DEVELOPMENT OF NOVEL DOPA DECARBOXYLASE INHIBITORS IN THE TREATMENT OF PARKINSON S DISEASE M Lakshmi Vasavi Devi 1 *, Deepak Reddy Gade 2, T Gopi Raju 3, Pavan
Journal of Chemical and Pharmaceutical Research, 2016, 8(9): Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2016, 8(9):73-80 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 QSAR and Docking Studies of Pubchem Derivatives as
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2016, 8(9):73-80 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 QSAR and Docking Studies of Pubchem Derivatives as
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor!
 Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
CHAPTER 1 1. Molecular recognition 1.1 Lock-and-key versus induced-fit mechanism of ligand binding
 CHAPTER 1 1. Molecular recognition In each cell of an organism, a myriad of fundamental molecular reactions, covalent and non-covalent, co-ordinate and regulate the way in which biological molecules interact
CHAPTER 1 1. Molecular recognition In each cell of an organism, a myriad of fundamental molecular reactions, covalent and non-covalent, co-ordinate and regulate the way in which biological molecules interact
Predicting Inactive Conformations of Protein Kinases Using Active Structures: Conformational Selection of Type-II Inhibitors
 Predicting Inactive Conformations of Protein Kinases Using Active Structures: Conformational Selection of Type-II Inhibitors Min Xu, Lu Yu, Bo Wan, Long Yu, Qiang Huang* State Key Laboratory of Genetic
Predicting Inactive Conformations of Protein Kinases Using Active Structures: Conformational Selection of Type-II Inhibitors Min Xu, Lu Yu, Bo Wan, Long Yu, Qiang Huang* State Key Laboratory of Genetic
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
