SUPPLEMENTARY MATERIAL
|
|
|
- Bennett McKinney
- 6 years ago
- Views:
Transcription
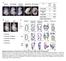
1 SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates. Numbers under the x-axis represent relative locations on the chromosome in base pairs. The y-axis shows different species. (+) means transcription from centromere to telomere; (-) means transcription from telomere to centromere. (B) 3 -UTRs of Casp2 are highly conserved in mammals. Letters in purple represent the seed regions for mir-183; letters in red indicate the seed regions for mir-96 and mir-182. Letters in the red box denote the conserved region for the mir-183/96/182 cluster. Figure S2. Creation of the mir-183/96/182 cluster sponge construct. (A) a. Schematic representation of the sponge construct: Ten copies of binding sites for each mir-183/96/182 cluster mir were inserted into the 3 -UTR of EGFP. The construct under control of a mouse opsin promoter ended with a SV40 polyadenylation signal. The whole fragment containing 6.8 kb was cut out by NotI/MluI. The SpongeFor/SpongeRev primer pair was designed for genotyping analysis and the BindingFor/BindingRev primer pair was included to check expression of the sponge element. Binding sequences of mir-183, mir-96 and mir-182 are shown in b. (B) Testing for the mouse Opsin promoter. Only Y79 cells transfected with the sponge construct showed a bright EGFP signal. Phase contrast images are shown on the right. Scale bars: 20 μm. Figure S3. Test of cluster probe specificities. There were no cross-reactions between different probes. Figure S4. Creation of mir-183/96/182 stable cell lines. (A) Inserted partial pri-mir sequences of mir-183/96/182 cluster mirs. Letters in purple indicate mature mir sequences; letters in red represent the remaining regions of pre-mir sequences; letters in blue denote cloning and detecting primer sequences. (B) Confirmation of mir expression levels in stable cell lines. Note that a demonstrates the specific pri-mir expression of target mirs detected by RT-PCR in different stable cell lines, whereas b shows specific mature mir expression in different cell lines by splinted ligation. Figure S5. Creation of target luciferase reporters. (A) Schematic representation of luciferase reporter constructs with four copies of the targeted regions from each target gene s 3 -UTR. The targeted regions in target genes 3-UTRs are listed; letters in green represent the seed regions, letters in red denote the complementary nucleotides. (B) Testing the linear range of genetically engineered luciferase reporter constructs. From ng was found to be in the linear range for all plasmid constructs and the empty pgl3p plasmid whereas ng was in the linear range for the Renilla plasmid. Figure S6. Supplementary data for the establishment of mir-183/96/182 sponge transgenic mouse cell lines. (A) A standard curve of sponge element DNA is shown with Ct-values derived from quantitative real-time PCR. The slope (efficiency), R 2 (coefficients) and y-intercept (sensitivity) are listed. A transgene size of 1 kb was estimated as 0.17 picograms of transgene per microgram of mouse genomic DNA. (B) Tissue specificity of sponge component expression. The opsin promoter mrna of sponge component constructs was detected only in retina and not in brain, heart, liver or intestine by semiquantitative RT-PCR. RT-minus [RT (-)] and Gapdh reactions were used as negative and positive controls. Amplification cycle numbers are indicated on the right. (C) Statistical analysis of mature mirs in different transgenic mouse lines. Error bars represent the standard errors of the means (n=3), * p<0.05. (D) Immunoblotting results show that Opsin 1
2 expression levels were not affected by the exogenous mouse Opsin promoter in all three transgenic mouse lines. β-tubulin was used as the internal control. Figure S7. Casp2 expression levels in 4-month-old wt mouse retina under different light intensity exposure. Immunoblotting data show that both pro-casp2 and cleaved-casp2 were increased after 5,000 and 10,000 lux light intensity exposure for 30 min. β-tubulin was used as the internal control. 2
3 SUPPLEMENTARY TABLES Table S1. Primer Sets for Real Time PCR. 1 Name mrna Forward Reverse Neurod4 NM_ AGACCGGGCTCTTATGGAAT GAATCTTTCAAGGCGAGCTT Hes1 NM_ GGCCTCTGAGCACAGAAAGT GCCGGGAGCTATCTTTCTTA Dcx NM_ CAGAAGCGATCAAACTGGAA GGACCACAAGCAATGAACAC Ncald NM_ TGGGACTGCACACATTGAA ACTCCTGGATCTCGTGCTCT Rarg NM_ TCATCTGTGGAGACCGAATG CCTTGGGAACATGTAGGGTT Gnai3 NM_ TTGATGTAGGTGGCCAAAGA ATGCATTCGGTTCATTTCCT Clock NM_ GGCACCACCAATAATAGGCT TGCATTAAGTGCTCGTGACA Bhlhe22 NM_ GGTTTAGTGCGGATGAAGTG TTTCGCCTCCAAAGAAGACT Casp2 NM_ GCACAGGAAATGCAAGAGAA CTTGGAGCTGAAGCAGTTTG Bcl2 NM_ GAGTACCTGAACCGGCATCT GAAATCAAACAGAGGTCGCA Arrdc3 NM_ TCTGGGAAGACAGAGACGTG ATCCACGTACACCATTAGCG Binding TCCTGCTGGAGTTCGTGAC CGTCCCCGGGTTTAAACTA Pri-miR96 GGGCCATAAACAGAGCAGAG GGCAGTGAAAGGTGATCTGG Pri-miR182 TACAGGCCGAAGGACCATAG GGCGCAGGGAAACATTAAG Pri-miR183 CTGGTGAGGAGGGTTGCTAA GTACTGGAACAGGCCCTCTG Gapdh NM_ GTGTTCCTACCCCCAATGTG GGAGACAACCTGGTCCTCAG 1 Primer sets used in this research. The first column shows the identities of the primer sets; the second column indicates the corresponding accession number; and the third and fourth columns present the sequences of the forward and reverse primers, respectively. 3
4 Table S2. Oligonucleotides used for Splinted Ligation. 1 Name Bridge-mmu-miR-96 Bridge-mmu-miR-182 Bridge-mmu-miR-183 Bridge-mmu-miR-125a Bridge-mmu-Let-7 Bridge-mmu-miR-107 Mimic-mmu-miR-96 Mimic-mmu-miR-182 Mimic-mmu-miR-183 Sequence GAATGTCATAAGCGAGCAAAAATGTGCTAGTGCCAAA GAATGTCATAAGCGCGGTGTGAGTTCTACCATTGCCAAA GAATGTCATAAGCGAGTGAATTCTACCAGTGCCATA GAATGTCATAAGCGTCACAGGTTAAAGGGTCTCAGGGA GAATGTCATAAGCGAACTATACAACCTACTACCTCA GAATGTCATAAGCGTGATAGCCCTGTACAATGCTGCT TTTGGCACTAGCACATTTTTGCT TTTGGCAATGGTAGAACTCACACCG TATGGCACTGGTAGAATTCACT 1 Oligonucleotides used for splinted ligation. Bridge oligonucleotides were used to hybridize the detection probe and target mirs; mimic oligonucleotides simulating the endogenous mir sequences were used as positive controls. 4
5 Table S3. Statistical analysis of Figure 3Cb. 1 wt transgene 0 min 10 min 30 min 0 min 10 min 30 min rod Opsin 1.0± ± ± ± ±0.1 * 0.4±0.0 * Mw-opsin 1.0± ± ± ± ± ±0.1 * 1 Immunoblotting data were quantified by Image J. Both Opsin and Mw-opsin levels were normalized by Tubulin. Data represent means±s.d. (n=3), * p<
6 Table S4. Statistical analysis of Figure 5Cb. 1 NIH3T3 mir-96 mir-182 mir-183 pro-casp2 1.0± ±0.1 * 0.5±0.1 * 1.1±0.1 cleaved-casp2 1.0± ±0.2 * 0.5±0.2 * 1.0±0.2 1 Immunoblotting data were quantified by Image J. Both pro-casp2 and cleaved-casp2 levels were normalized by Tubulin. Data represent mean±s.d. (n=3), * p<
7 Table S5. Statistical analysis of Figure 6Ab. 1 0 h 1 h 3 h 6 h 12 h 24 h 72 h mir ± ± ± ± ±0.2* 2.5±0.4* 1.1±0.2 mir ± ± ± ± ± ± ±0.1 mir ± ± ± ± ±0.1* 2.1±0.1* 1.0±0.0 1 Splinted ligation data were quantified by Image J. mir-96, mir-182 and mir-183 levels were normalized by Let-7. Data represent means±s.d. (n=3), * p<
8 Table S6. Statistical analysis of Figure 6Bb. 1 wt transgene not treated light-induced not treated light-induced pro-casp2 1.0± ± ± ±0.4 cleaved-casp2 1.0± ± ± ±0.3 * Bid 1.0± ± ± ±0.3 tbid 1.0± ±0.5 * 1.6±0.2 * 3.3±0.2 * 1 Immunoblotting data were quantified by Image J. All protein levels were normalized by Tubulin. Data represent means±s.d. (n=3), * p<
9 Table S7. Statistical analysis of Figure S lux 500 lux 1,000 lux 2,500 lux 5,000 lux 10,000 lux pro-casp2 1.0± ± ± ± ±0.3 * 1.6±0.4 cleaved-casp2 1.0± ± ± ± ±0.3 * 2.0±0.3 * 1 Immunoblotting data were quantified by Image J. Both pro-casp2 and cleaved-casp2 levels were normalized by Tubulin. Data represent means±s.d. (n=3), * p<
10 B A Danio rerio (+) Tetraodon nigroviridis (-) Fugu rubripes (-) Xenopus tropicalis (-) Monodelphis domestica (-) Mus musculus (-) Canis familiaris (+) Bos taurus (-) Macaca mulatta (-) Pan troglodytes (-) Homo sapiens (-) mir-183 space1 mir-96 space2 mir (kb) Homo_sapiens AGAGAACCTTTCCACT-CCCACTGCCAGGATTTTGTATTGCCATCGGGTGCCAAATAAAT Pan_troglodytes AGAGAACCTTTCCACT-CCCACTGCCAGGATTTTGTATTGCCATCGGGTGCCAAATAAAT Pongo_pygmaeus CGAGAATCTTTCCACT-CCCACTGCCAGGATTTTGTATTGCCATCAGGTGCCAAATAAAT Rattus_norvegicu GTAGACTCTTCACACTTCCCACTGCCAAGATTTTGTATTGCCATCAGGTGCCAAATAAAT Mus_musculus GTAGAATCTTCACACTTCCCACTGCCAAGATTTTGTATTGCCATCAGGTGCCAAATAAAT Monodelphis_dome TGAGCCTCTCCCCACTCCCCACTGCCAGGGTTTTGTATTGTTTTCAGGTGCCAAATAAAT Bos_taurus ATAGAACCTTTCCAGTTCCCACTGCCAGGATTTTGTATTGCCATCTGGTGCCAAATAAAT Canis_familiaris ATAGAACCTTTCCACTTCCCACTGCCAGGATTTTACATTGCCTTCAGGTGCCAAATAAAT Equus_caballus ATAGAACCTTTCCACTTCCCACTGCCAGGATTTTGTATTGCCATCAGGTGCCAAATAAAT ** ** ** * ********** * **** **** ** ************** Figure S1
11 A a 6.8 kb 0.1 kb NotI XhoI AclI PmeI MluI 5 - mouse opsin promoter EGFP SV40-3 b SpongeFor SpongeRev mir-183 binding sequence: TGG UTR 5 -AGTGAATTCTTGG AGTGCCATA mir UCACUUAAGAUG UCACGGUAU-5 UGG B BindingFor Y79 cells transfected GFP BindingRev phase contrast mir-96 binding sequence: ACG UTR 5 -AGCAAAAATGACG AGTGCCAAA mir UCGUUUUUACACGUCACGGUUU-5 ACGA Y79 cells not transfected mir-182 binding sequence: GAGG UTR 5 -CGGTGTGAGTTGAGG ATTGCCAAA mir GCCACACUCAAGAGGUAACGGUUU-5 GAUGG HEK293 cells transfected Figure S2
12 mir-96 mimic mir-182 mimic mir-183 mimic mir-96 probe mir-182 probe mir-183 probe Figure S3
13 A Mmu-miR-96 pri-mir GGGCCATAAACAGAGCAGAGACAGATCCGCGAGCACCTTGGAGCTCCTCACCCCTTTCTGCCTAGACCTCTGTTTCCAGGGGTGCCAGGGTACAAAGACCTCCTCTGCT CCTTCCCCAGAGGGCCTGTTCCAGTACCATCTGCTTGGCCGATTTTGGCACTAGCACATTTTTGCTTGTGTCTCTCCGCTGTGAGCAATCATGTGTAGTGCCAATATGGGA AAAGCGGGCTGCTGCGGCCACGTTCACCTCCCCCGGCATCCCAGGGTCTGTGTGTCTCACTGGCTCCCTGGCCCATCTGGCTTACTGCTGGGTGAGGAGGGTACAGCCCT ACCCTGGTGAACAGCCAGATCACCTTTCACTGCC Mmu-miR-182 pri-mir TACAGGCCGAAGGACCATAGTCTGGACCTTGTGTTAACTGTGGGAAGAGCGCCCTCCTAAAACCACCCTAACTGCTTCTTCTTCAGCATAGGCTTACTGGTCTGGCTGCT GGAGGCCTCCCACCATTTTTGGCAATGGTAGAACTCACACCGGTAAGGTAATGGGACCCGGTGGTTCTAGACTTGCCAACTATGGTGTAAGTGCTGAGCTGCTGAAGGT CTGCACCGTGCCGGAACCTGCCGATCACCAGGAAGGAGAGGGGACTCCTGTCTCCAGACCACCAGGCAGTGGCAGAGGGTGGGCGCAGCTGGAAGTGACCCTTAATGT TTCCCTGCGCC Mmu-miR-183 pri-mir CTGGTGAGGAGGGTTGCTAAACTGCGGATGACAGCAGTGGGGGTGGGGTGGGGGGAGTAGGTTGTAGGACCTCCAGGAGAAGGCAGCTGACCCCTCTGCAGGGTCTGC AGGCTGGAGAGTGTGACTCCTGTCCTGTGTATGGCACTGGTAGAATTCACTGTGAACAGTCTCAGTCAGTGAATTACCGAAGGGCCATAAACAGAGCAGAGACAGATCC GCGAGCACCTTGGAGCTCCTCACCCCTTTCTGCCTAGACCTCTGTTTCCAGGGGTGCCAGGGTACAAAGACCTCCTCTGCTCCTTCCCCAGAGGGCCTGTTCCAGTAC B a NIH3T3 mir-96 mir-182 mir-183 b mir-96 NIH3T3 mir-96 mir-182 mir-183 pri-mir-96 mir-182 pri-mir-182 pri-mir-183 mir-183 GAPDH Let-7 Figure S4
14 A Xbal FseI Luciferase SV40 Arrdc3-3-UTR TCAG-GTGAATGCATCACTTGCCAAA mir-96 UCGUUUUUACACGAUC-ACGGUUU mir-182 GCCACACUCAAGAUGGUAACGGUUU mir-183 UCACUUAAGAUGGUCACGGUAU Neurod4-3-UTR CAGGCCCTGTTCCAATGGTGCCAAA mir-96 UCGUUUUUACACGAUCACGGUUU mir-182 GCCACACUCAAGAUGGUAACGGUUU mir-183 UCACUUAAGAUGGUCACGGUAU Casp2-3-UTR TTTTGTATTGCCATCAGGTGCCAAA mir-96 UCGUUUUUACACGAUCACGGUUU mir-182 GCCACACUCAAGAUGGUAACGGUUU mir-183 UCACUUAAGAUGGUCACGGUAU B Luciferase activity (AU) pgl3p-4xneurod4-3 UTR pgl3p-4xarrdc3-3 UTR pgl3p-4xcasp2-3 UTR pgl3p Rellina Plasmid amount (ng) Figure S5
15 A Standard Curve Ct (cycles) Slope: Y-intercept: R 2 : Efficiency: 0.88 B mir-96 * * mir-182 mir-183 #1128 wt #1128 trans #1135 wt #1135 trans #1231 wt #1231 trans C retina brain heart liver intestine D #1128 wt #1128 trans #1135 wt #1135 trans #1231 wt #1231 trans Binding Primer RT (+) 33X rhodopsin Binding Primer RT (-) GAPDH RT (+) 33X 29X β-tubulin GAPDH RT (-) 29X Figure S6
16 0 lux 500 lux 1,000 lux 2,500 lux 5,000 lux 10,000 lux 49 kda pro-casp2 12 kda cleaved-casp2 β-tubulin Figure S7
Supplementary Figure 1. CFTR protein structure and domain architecture.
 A Plasma Membrane NH ₂ COOH Supplementary Figure. CFT protein structure and domain architecture. (A) Open state CFT homology model, ribbon representation from Serohijos et al. 8 PNAS 5:356. CFT domains
A Plasma Membrane NH ₂ COOH Supplementary Figure. CFT protein structure and domain architecture. (A) Open state CFT homology model, ribbon representation from Serohijos et al. 8 PNAS 5:356. CFT domains
Supplementary figure legends
 Supplementary figure legends SUPPLEMENTRY FIGURE S1. Lentiviral construct. Schematic representation of the PCR fragment encompassing the genomic locus of mir-33a that was introduced in the lentiviral construct.
Supplementary figure legends SUPPLEMENTRY FIGURE S1. Lentiviral construct. Schematic representation of the PCR fragment encompassing the genomic locus of mir-33a that was introduced in the lentiviral construct.
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent Protein.
 prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons
 Supplemental Material mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons Antoine de Chevigny, Nathalie Coré, Philipp Follert, Marion Gaudin, Pascal
Supplemental Material mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons Antoine de Chevigny, Nathalie Coré, Philipp Follert, Marion Gaudin, Pascal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis. Midbrain Forebrain/ Forebrain/ Hindbrain Spinal cord Hindbrain Hindbrain
 A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
A Normal Exencephaly Craniora- Spina bifida Microcephaly chischisis NTD Number of embryos % among NTD Embryos Exencephaly 52 74.3% Craniorachischisis 6 8.6% Spina bifida 5 7.1% Microcephaly 7 1% B Normal
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Uninformative BRCA Tests. Rebecca Sutphen, M.D. Professor, USF College of Medicine Chief Medical Officer, InformedDNA
 Uninformative RCA Tests Rebecca Sutphen, M.D. Professor, USF College of Medicine Chief Medical Officer, InformedDNA Uninformative RCA tests 1. R/OV cancer patient with normal results 2. Cancer-free person
Uninformative RCA Tests Rebecca Sutphen, M.D. Professor, USF College of Medicine Chief Medical Officer, InformedDNA Uninformative RCA tests 1. R/OV cancer patient with normal results 2. Cancer-free person
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Data to: Marco Mariotti and Roderic Guigó
 Supplementary Data to: Selenoprofiles: profile-based scanning of eukaryotic genome sequences for selenoprotein genes Marco Mariotti and Roderic Guigó Section S1: patterns used with SECISearch We report
Supplementary Data to: Selenoprofiles: profile-based scanning of eukaryotic genome sequences for selenoprotein genes Marco Mariotti and Roderic Guigó Section S1: patterns used with SECISearch We report
MII. Supplement Figure 1. CapZ β2. Merge. 250ng. 500ng DIC. Merge. Journal of Cell Science Supplementary Material. GFP-CapZ β2 DNA
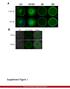 A GV GVBD MI DNA CapZ β2 CapZ β2 Merge B DIC GFP-CapZ β2 Merge CapZ β2-gfp 250ng 500ng Supplement Figure 1. MII A early MI late MI Control RNAi CapZαβ DNA Actin Tubulin B Phalloidin Intensity(A.U.) n=10
A GV GVBD MI DNA CapZ β2 CapZ β2 Merge B DIC GFP-CapZ β2 Merge CapZ β2-gfp 250ng 500ng Supplement Figure 1. MII A early MI late MI Control RNAi CapZαβ DNA Actin Tubulin B Phalloidin Intensity(A.U.) n=10
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Differential expression of mirnas from the pri-mir-17-92a locus.
 Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein
 Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Figure 1. AdipoR1 silencing and overexpression controls. (a) Representative blots (upper and lower panels) showing the AdipoR1 protein content relative to GAPDH in two independent experiments.
Supplementary Information
 Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Information mediates STAT3 activation at retromer-positive structures to promote colitis and colitis-associated carcinogenesis Zhang et al. a b d e g h Rel. Luc. Act. Rel. mrna Rel. mrna
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical
 Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Figure SⅠ: Expression of mir-155, mir-122 and mir-196a in allografts compared with
 Figure SⅠ: Expression of mir-155, mir-122 and mir-196a in allografts compared with isografts (control) at the 2nd week, 4th and 8th week by RT-PCR. At the advanced stage, the expression of these three
Figure SⅠ: Expression of mir-155, mir-122 and mir-196a in allografts compared with isografts (control) at the 2nd week, 4th and 8th week by RT-PCR. At the advanced stage, the expression of these three
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
mirna-guided regulation at the molecular level
 molecular level Hervé Seitz IGH du CNRS, Montpellier, France March 3, 2016 microrna target prediction . microrna target prediction mirna: target: 2 7 5 N NNNNNNNNNNNNNN 3 NNNNNN the seed microrna target
molecular level Hervé Seitz IGH du CNRS, Montpellier, France March 3, 2016 microrna target prediction . microrna target prediction mirna: target: 2 7 5 N NNNNNNNNNNNNNN 3 NNNNNN the seed microrna target
SUPPLEMENTARY INFORMATION
 -. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
-. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna
 Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
P. Mathijs Voorhoeve, Carlos le Sage, Mariette Schrier, Ad J.M. Gillis, Hans Stoop,
 Supplemental Data A Genetic Screen Implicates mirna-372 and mirna-373 As Oncogenes in Testicular Germ Cell Tumors P. Mathijs Voorhoeve, Carlos le Sage, Mariette Schrier, Ad J.M. Gillis, Hans Stoop, Remco
Supplemental Data A Genetic Screen Implicates mirna-372 and mirna-373 As Oncogenes in Testicular Germ Cell Tumors P. Mathijs Voorhoeve, Carlos le Sage, Mariette Schrier, Ad J.M. Gillis, Hans Stoop, Remco
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1
 S U P P L E M E N TA R Y I N F O R M AT I O N DOI: 10.1038/ncb2896 Supplementary Figure 1 Supplementary Figure 1. Sequence alignment of TERB1 homologs in vertebrates. M. musculus TERB1 was derived from
S U P P L E M E N TA R Y I N F O R M AT I O N DOI: 10.1038/ncb2896 Supplementary Figure 1 Supplementary Figure 1. Sequence alignment of TERB1 homologs in vertebrates. M. musculus TERB1 was derived from
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained
 Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
Supplementary Table 1. List of primers used in this study
 Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Supplementary Information
 Supplementary Information Supplementary Figure 1. EBV-gB 23-431 mainly exists as trimer in HEK 293FT cells. (a) Western blotting analysis for DSS crosslinked FLAG-gB 23-431. HEK 293FT cells transfected
Supplementary Information Supplementary Figure 1. EBV-gB 23-431 mainly exists as trimer in HEK 293FT cells. (a) Western blotting analysis for DSS crosslinked FLAG-gB 23-431. HEK 293FT cells transfected
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
MOLECULE PHYLOGENETIC ANALYSIS OF GENE ENCODING ASPARTATE AMINOTRANSFERASE
 Analele Ştiinţifice ale Universităţii Alexandru Ioan Cuza, Secţiunea Genetică şi Biologie Moleculară, TOM IX, 2008 MOLECULE PHYLOGENETIC ANALYSIS OF GENE ENCODING ASPARTATE AMINOTRANSFERASE XIAOSHENG DING
Analele Ştiinţifice ale Universităţii Alexandru Ioan Cuza, Secţiunea Genetică şi Biologie Moleculară, TOM IX, 2008 MOLECULE PHYLOGENETIC ANALYSIS OF GENE ENCODING ASPARTATE AMINOTRANSFERASE XIAOSHENG DING
Tyrosine phosphorylation and protein degradation control the transcriptional activity of WRKY involved in benzylisoquinoline alkaloid biosynthesis
 Supplementary information Tyrosine phosphorylation and protein degradation control the transcriptional activity of WRKY involved in benzylisoquinoline alkaloid biosynthesis Yasuyuki Yamada, Fumihiko Sato
Supplementary information Tyrosine phosphorylation and protein degradation control the transcriptional activity of WRKY involved in benzylisoquinoline alkaloid biosynthesis Yasuyuki Yamada, Fumihiko Sato
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Table. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Table File Name: Peer Review File Description: Supplementary Table 1 Primers and taqman probes used were the following:
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Table File Name: Peer Review File Description: Supplementary Table 1 Primers and taqman probes used were the following:
Supporting Information
 Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
Supporting Information Palmisano et al. 10.1073/pnas.1202174109 Fig. S1. Expression of different transgenes, driven by either viral or human promoters, is up-regulated by amino acid starvation. (A) Quantification
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
An Epstein-Barr virus-encoded microrna targets PUMA to promote host cell survival
 An Epstein-Barr virus-encoded microrna targets to promote host cell survival The Journal of Experimental Medicine 205(11): 2551-2560, 2008. 1 Elizabeth Yee-Wai Choy, Kam-Leung Siu, Kin-Hang Kok, Raymond
An Epstein-Barr virus-encoded microrna targets to promote host cell survival The Journal of Experimental Medicine 205(11): 2551-2560, 2008. 1 Elizabeth Yee-Wai Choy, Kam-Leung Siu, Kin-Hang Kok, Raymond
Supplementary Figure 1. Expression of phospho-sik3 in normal and osteoarthritic articular cartilage in the knee. (a) Semiserial histological sections
 Supplementary Figure 1. Expression of phospho-sik3 in normal and osteoarthritic articular cartilage in the knee. (a) Semiserial histological sections of normal cartilage were stained with safranin O-fast
Supplementary Figure 1. Expression of phospho-sik3 in normal and osteoarthritic articular cartilage in the knee. (a) Semiserial histological sections of normal cartilage were stained with safranin O-fast
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Amelio et al., http://www.jcb.org/cgi/content/full/jcb.201203134/dc1 Figure S1. mir-24 regulates proliferation and by itself induces
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Amelio et al., http://www.jcb.org/cgi/content/full/jcb.201203134/dc1 Figure S1. mir-24 regulates proliferation and by itself induces
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
Supplementary Information
 Supplementary Information Overexpression of Fto leads to increased food intake and results in obesity Chris Church, Lee Moir, Fiona McMurray, Christophe Girard, Gareth T Banks, Lydia Teboul, Sara Wells,
Supplementary Information Overexpression of Fto leads to increased food intake and results in obesity Chris Church, Lee Moir, Fiona McMurray, Christophe Girard, Gareth T Banks, Lydia Teboul, Sara Wells,
Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples.
 Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists
 Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists of: (i) the YPet domain (an enhanced YFP); (ii) the
Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists of: (i) the YPet domain (an enhanced YFP); (ii) the
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
Ccdc124: A novel protein involved in cytokinesis.
 Ccdc124: A novel protein involved in cytokinesis. Uygar H. Tazebay Gebze Technical University Department of Molecular Biology and Genetics Gebze, Turkey International Conference on Science and Technology
Ccdc124: A novel protein involved in cytokinesis. Uygar H. Tazebay Gebze Technical University Department of Molecular Biology and Genetics Gebze, Turkey International Conference on Science and Technology
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Nature Genetics: doi: /ng.3731
 Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supporting Information Table of Contents
 Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
Supporting Information Table of Contents Supporting Information Figure 1 Page 2 Supporting Information Figure 2 Page 4 Supporting Information Figure 3 Page 5 Supporting Information Figure 4 Page 6 Supporting
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
Supplemental Data. TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim.
 Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
Supplementary Information. Induction of p53-independent apoptosis by ectopic expression of HOXA5
 Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Supplementary Information Induction of p53-independent apoptosis by ectopic expression of in human liposarcomas Dhong Hyun Lee 1, *, Charles Forscher 1, Dolores Di Vizio 2, 3, and H. Phillip Koeffler 1,
Protein SD Units (P-value) Cluster order
 SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1. Formation of the AA5x. a, Camera lucida drawing of embryo at 48 hours post fertilization (hpf, modified from Kimmel et al. Dev Dyn. 1995 203:253-310). b, Confocal microangiogram
Supplementary Figure 1. Formation of the AA5x. a, Camera lucida drawing of embryo at 48 hours post fertilization (hpf, modified from Kimmel et al. Dev Dyn. 1995 203:253-310). b, Confocal microangiogram
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Research Article Base Composition Characteristics of Mammalian mirnas
 Journal of Nucleic Acids Volume 2013, Article ID 951570, 6 pages http://dx.doi.org/10.1155/2013/951570 Research Article Base Composition Characteristics of Mammalian mirnas Bin Wang Department of Chemistry,
Journal of Nucleic Acids Volume 2013, Article ID 951570, 6 pages http://dx.doi.org/10.1155/2013/951570 Research Article Base Composition Characteristics of Mammalian mirnas Bin Wang Department of Chemistry,
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
SUPPLEMENTARY LEGENDS...
 TABLE OF CONTENTS SUPPLEMENTARY LEGENDS... 2 11 MOVIE S1... 2 FIGURE S1 LEGEND... 3 FIGURE S2 LEGEND... 4 FIGURE S3 LEGEND... 5 FIGURE S4 LEGEND... 6 FIGURE S5 LEGEND... 7 FIGURE S6 LEGEND... 8 FIGURE
TABLE OF CONTENTS SUPPLEMENTARY LEGENDS... 2 11 MOVIE S1... 2 FIGURE S1 LEGEND... 3 FIGURE S2 LEGEND... 4 FIGURE S3 LEGEND... 5 FIGURE S4 LEGEND... 6 FIGURE S5 LEGEND... 7 FIGURE S6 LEGEND... 8 FIGURE
SUPPLEMENTARY FIGURES AND TABLES
 SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
Supplemental Table 1. Primers used for RT-PCR analysis of inflammatory cytokines Gene Primer Sequence
 Supplemental Table 1. Primers used for RT-PCR analysis of inflammatory cytokines Gene Primer Sequence IL-1α Forward primer 5 -CAAGATGGCCAAAGTTCGTGAC-3' Reverse primer 5 -GTCTCATGAAGTGAGCCATAGC-3 IL-1β
Supplemental Table 1. Primers used for RT-PCR analysis of inflammatory cytokines Gene Primer Sequence IL-1α Forward primer 5 -CAAGATGGCCAAAGTTCGTGAC-3' Reverse primer 5 -GTCTCATGAAGTGAGCCATAGC-3 IL-1β
Supplementary Materials
 Supplementary Materials Figure S1. MTT Cell viability assay. To measure the cytotoxic potential of the oxidative treatment, the MTT [3-(4,5-dimethylthiazol- 2-yl)-2,5-diphenyl tetrazolium bromide] assay
Supplementary Materials Figure S1. MTT Cell viability assay. To measure the cytotoxic potential of the oxidative treatment, the MTT [3-(4,5-dimethylthiazol- 2-yl)-2,5-diphenyl tetrazolium bromide] assay
Supplementary Figure 1
 Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
doi: 1.138/nature89 IFN- (ng ml ) 5 4 3 1 Splenocytes NS IFN- (ng ml ) 6 4 Lymph node cells NS Nfkbiz / Nfkbiz / Nfkbiz / Nfkbiz / IL- (ng ml ) 3 1 Splenocytes IL- (ng ml ) 1 8 6 4 *** ** Lymph node cells
<10. IL-1β IL-6 TNF + _ TGF-β + IL-23
 3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice
 SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
Studying apoptosis in DT40 cells
 Studying apoptosis in DT40 cells Sandrine Ruchaud E12.5! Acridine Orange! Role of apoptosis in sculpting the mouse paw E13.5! E14.5! gift of William Wood & Paul Martin! University College, London! Normal
Studying apoptosis in DT40 cells Sandrine Ruchaud E12.5! Acridine Orange! Role of apoptosis in sculpting the mouse paw E13.5! E14.5! gift of William Wood & Paul Martin! University College, London! Normal
Rescue of mutant rhodopsin traffic by metformin-induced AMPK activation accelerates photoreceptor degeneration Athanasiou et al
 Supplementary Material Rescue of mutant rhodopsin traffic by metformin-induced AMPK activation accelerates photoreceptor degeneration Athanasiou et al Supplementary Figure 1. AICAR improves P23H rod opsin
Supplementary Material Rescue of mutant rhodopsin traffic by metformin-induced AMPK activation accelerates photoreceptor degeneration Athanasiou et al Supplementary Figure 1. AICAR improves P23H rod opsin
Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or
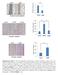 Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or 3xflag-CagA expression vector were wounded using a pipette
Supplementary Figure 1. The CagA-dependent wound healing or transwell migration of gastric cancer cell. AGS cells transfected with vector control or 3xflag-CagA expression vector were wounded using a pipette
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3461 In the format provided by the authors and unedited. Supplementary Figure 1 (associated to Figure 1). Cpeb4 gene-targeted mice develop liver steatosis. a, Immunoblot displaying CPEB4
DOI: 10.1038/ncb3461 In the format provided by the authors and unedited. Supplementary Figure 1 (associated to Figure 1). Cpeb4 gene-targeted mice develop liver steatosis. a, Immunoblot displaying CPEB4
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Table.
 Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tale. Type of file: VI Title of file for HTML: Supplementary Movie 1 Description:
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures and Supplementary Tale. Type of file: VI Title of file for HTML: Supplementary Movie 1 Description:
Supporting Information
 Supporting Information ou et al..73/pnas.08791112 dd Thymidine Release & transfection dd Thymidine Release dd MG132 Fix and IF -14 h 0 h 8 h 24 h 34 h 36 h siontrol simps1-1 simps1-1 simps1-1 simps1-2
Supporting Information ou et al..73/pnas.08791112 dd Thymidine Release & transfection dd Thymidine Release dd MG132 Fix and IF -14 h 0 h 8 h 24 h 34 h 36 h siontrol simps1-1 simps1-1 simps1-1 simps1-2
A. List of selected proteins with high SILAC (H/L) ratios identified in mass
 Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables. File Name: Peer Review File Description:
 File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables File Name: Peer Review File Description: Primer Name Sequence (5'-3') AT ( C) RT-PCR USP21 F 5'-TTCCCATGGCTCCTTCCACATGAT-3'
Supplementary Figure 1. HGL-DTG levels in uninduced and herbivory induced leaves. Total HGL-
 Supplementary Figure 1. HGL-DTG levels in uninduced and herbivory induced leaves. Total HGL- DTG concentrations [F 1,4 = 36.88, P 0.0037; significant differences (threshold: P 0.05) between means (± SE)
Supplementary Figure 1. HGL-DTG levels in uninduced and herbivory induced leaves. Total HGL- DTG concentrations [F 1,4 = 36.88, P 0.0037; significant differences (threshold: P 0.05) between means (± SE)
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplementary Information and Figure legends
 Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
