microrna analysis Merete Molton Worren Ståle Nygård
|
|
|
- Joy Rogers
- 5 years ago
- Views:
Transcription
1 microrna analysis Merete Molton Worren Ståle Nygård Help personnel: Daniel Vodak
2 Background Dysregulation of mirna expression has been connected to progression and development of atherosclerosis The hypothesis: mirna expression profiles differ between baboons with high and low serum LDL-C A subset of these mirnas regulate genes are relevant to dyslipidemia and risk of atherosclerosis
3 Experiment Baboons divided into two groups based on their LDL-C response levels Two diets Chow (baseline diet) HCHF (High Cholesterol High Fat diet) Compared mirna expression profiles from liver biopsies collected before and after the HCHF challenge diet
4 Paper Karere GM, Glenn JP, Vandeberg JL, Cox LA Differential microrna response to a high-cholesterol, high-fat diet in livers of low and high LDL-C baboons. BMC Genomics 13(1):32
5 Experiment setup 12 samples available from baboon liver biopsies 3 replicates from high-ldl, baseline (chow) diet 3 replicates from high-ldl, HCHF diet 3 replicates from low-ldl, baseline (chow) diet 3 replicates from low-ldl, HCHF diet We will work with the high-ldl samples only
6 QUESTIONS?
7 mirna analysis workflow Preparations Alignment Annotation Read count per mirna Normalization Differential expression analysis
8 Preparations, read files Starting point: sequence read files Practical info File format (.fastq,.fasta,.sff) Base quality scale used (Phred+33, Phred+64) Quality control Adapters
9 Preparations, reference Find sequences you will align to Resources: UCSC: NCBI: ENSEMBL: We will use sequences from mirbase that we will customize to our needs
10 Preparations, mirna Collapse files for increased speed Check that reads/reference are both RNA or both DNA
11 QUESTIONS?
12 Next: Alignment
13 Alignment The sequence read is not very informative in itself. Alignment to a reference -> location of origin TAGCTTATCAGACTGATGTTGA
14 Alignment The sequence read is not very informative in itself. Alignment to a reference -> location of origin TAGCTTATCAGACTGATGTTGA TACACTGAGTCGACTGAGTGGCCTAGCTTATCAGACTGATGTTGAATTTATATAGGCGTGACGTA
15 Alignment The sequence read is not very informative in itself. Alignment to a reference -> location of origin TAGCTTATCAGACTGATGTTGA TACACTGAGTCGACTGAGTGGCCTAGCTTATCAGACTGATGTTGAATTTATATAGGCGTGACGTA mir-21-5p
16 What to align to Often several possible references The genome of the organism or a closely related organism Specific sequences like mrna or mirnas Databases such as Rfam filter out reads check what the unaligned reads are
17 Aligners Many different aligners Novoalign Bowtie/Bowtie2 BWA (Burrows-Wheeler Aligner) SOAPaligner/soap2 (Short Oligonucleotide Analysis Package)
18 Alignment challenges Sequencing errors True differences from reference Repeat regions Similar coding sequences
19 Alignment process Indexing An index is made of the genome Mapping based on index Finds several possible alignment location Final alignment Thorough search of possible mapping locations
20 Simplified example, k-mer=2 Genome : TTATATTTTATTAATTTTA Possible 2-mers Location positions TT 1,6,7,8,11,15,16,17 TA 2,4,9,12,18 AT 3,5,10,14 AA 13 Our read: ATT Start 2-mer AT
21 Options Different options for different aligners Reading reference manuals is a big part of doing bioinformatics
22 Some common options Level of similarity, genome/reads Output formats Options to control multiple mappers
23 QUESTIONS?
24 Next: Annotation
25 Annotation Aligned to sequence of interest The annotation is already in place Aligned to genome Either: coordinates for the features of interest are already known Or: part of project is to annotate the genome
26 Annotation Some genomes are better annotated than others Coordinates for specific genomic features can be found through several sites ENSEMBL UCSC Table browser mirbase
27 QUESTIONS?
28 Next: Read counts
29 Read counts How much do we have? Read counts is the starting point for finding expression and differential expression When counting, multiple mappers must be considered
30 Read count mirna Multiple mappers are common Size of reads Similarity of different mirnas This leads to some additional challenges when counting the reads of mirnas
31 Aligning mirnas to mirnas Reads\Reference hsa mirs all mirs hsa mirs (2578 reads) 2578/ mirs lost 2578/ mirs lost all mirs (30424 reads) 10111/ mirs lost 30424/ mirs lost
32 Counting multiple mappers Disregard all the reads that maps to multiple locations Keeping one or more randomly Keeping them all
33 QUESTIONS?
34 Next: Normalization
35 Normalization The purpose of normalization is to remove technical bias while maintaining true biological signal.
36 Normalization Normalization is necessary because the true expression is not the only factor influencing the read counts Other factors may be Sample size Differences in extraction Differences during the sequencing procedure
37 Normalization methods for mirna No method for best practice yet Normalization by read counts Several other methods tried with better results TMM
38 QUESTIONS?
39 Next: Differential expression analysis
40 Differential expression analysis Differential expression analysis aims at finding the mirnas that are truly differentially expressed under the relevant conditions The challenge is to be able to find as many as possible of the true ones, while avoiding to call a mirna differentially expressed when the changes are random or due to other factors
41 Differential expression analysis FLU (subject1) HEALTHY (subject2) Is this a good way of checking what mirnas are differentially expressed when having the flu?
42 Differential expression analysis FLU (subject1) Male Arthritis Eats vegetables HEALTHY (subject2) Female Allergy Eats burgers and fries Is this a good way of checking what mirnas are differentially expressed when having the flu?
43 Biological replicates There will be a lot of mirnas that have different expression in two samples Some due to the condition you are investigating Some due to other factors The solution: biological replicates Average out unknown/unwanted differences Call differential expression only where it is due to the factor you are investigating
44 QUESTIONS?
45 Next: A little statistics with Ståle Nygård (Eivind Hovig s group)
46 QUESTIONS?
47 Next: Discussion of the results
48 Comparison Lifeportal analysis mirna FDR mir mir-133b mir-1-3p mir-133a-3p mir-95-3p mir-499a-5p mir-1.5p ---- Command line mirna FDR mir mir-133b mir-1-3p mir-133a-3p mir-95-3p mir-499a-5p mir-1-5p 0.024
49 Comparison Our analysis mirna FDR mir mir mir-133a-3p mir-133b mir-95-3p mir-499a-5p mir mir-452-5p Paper mirna FDR mir-29b 0.01 mir mir mir
50 Possible sources of difference From fastq files to read counts Differences in aligner/reference Different way of counting Statistics Different normalization Different statistical tests used
51 Learning points Different pipelines will give different results Very important to give good descriptions of exactly what you have done PCR validation is important
Micro-RNA web tools. Introduction. UBio Training Courses. mirnas, target prediction, biology. Gonzalo
 Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
Small RNA Sequencing. Project Workflow. Service Description. Sequencing Service Specification BGISEQ-500 SERVICE OVERVIEW SAMPLE PREPARATION
 BGISEQ-500 SERVICE OVERVIEW Small RNA Sequencing Service Description Small RNAs are a type of non-coding RNA (ncrna) molecules that are less than 200nt in length. They are often involved in gene silencing
BGISEQ-500 SERVICE OVERVIEW Small RNA Sequencing Service Description Small RNAs are a type of non-coding RNA (ncrna) molecules that are less than 200nt in length. They are often involved in gene silencing
Obstacles and challenges in the analysis of microrna sequencing data
 Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
A Quick-Start Guide for rseqdiff
 A Quick-Start Guide for rseqdiff Yang Shi (email: shyboy@umich.edu) and Hui Jiang (email: jianghui@umich.edu) 09/05/2013 Introduction rseqdiff is an R package that can detect differential gene and isoform
A Quick-Start Guide for rseqdiff Yang Shi (email: shyboy@umich.edu) and Hui Jiang (email: jianghui@umich.edu) 09/05/2013 Introduction rseqdiff is an R package that can detect differential gene and isoform
Eukaryotic small RNA Small RNAseq data analysis for mirna identification
 Eukaryotic small RNA Small RNAseq data analysis for mirna identification P. Bardou, C. Gaspin, S. Maman, J. Mariette, O. Rué, M. Zytnicki INRA Sigenae Toulouse INRA MIA Toulouse GenoToul Bioinfo INRA MaIAGE
Eukaryotic small RNA Small RNAseq data analysis for mirna identification P. Bardou, C. Gaspin, S. Maman, J. Mariette, O. Rué, M. Zytnicki INRA Sigenae Toulouse INRA MIA Toulouse GenoToul Bioinfo INRA MaIAGE
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS
 Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Supplementary Figure 1
 Supplementary Figure 1 a Location of GUUUCA motif relative to RNA position 4e+05 3e+05 Reads 2e+05 1e+05 0e+00 35 40 45 50 55 60 65 Distance upstream b 1: Egg 2: L3 3: Adult 4: HES 4e+05 3e+05 2e+05 start
Supplementary Figure 1 a Location of GUUUCA motif relative to RNA position 4e+05 3e+05 Reads 2e+05 1e+05 0e+00 35 40 45 50 55 60 65 Distance upstream b 1: Egg 2: L3 3: Adult 4: HES 4e+05 3e+05 2e+05 start
DNA Sequence Bioinformatics Analysis with the Galaxy Platform
 DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
Human breast milk mirna, maternal probiotic supplementation and atopic dermatitis in offsrping
 Human breast milk mirna, maternal probiotic supplementation and atopic dermatitis in offsrping Melanie Rae Simpson PhD candidate Department of Public Health and General Practice Norwegian University of
Human breast milk mirna, maternal probiotic supplementation and atopic dermatitis in offsrping Melanie Rae Simpson PhD candidate Department of Public Health and General Practice Norwegian University of
RmiR package vignette
 RmiR package vignette Francesco Favero * October 17, 2016 Contents 1 Introduction 1 2 Coupling mirna and gene expression data 2 2.1 Default parameters.............................. 3 2.2 Change the at.least
RmiR package vignette Francesco Favero * October 17, 2016 Contents 1 Introduction 1 2 Coupling mirna and gene expression data 2 2.1 Default parameters.............................. 3 2.2 Change the at.least
VirusDetect pipeline - virus detection with small RNA sequencing
 VirusDetect pipeline - virus detection with small RNA sequencing CSC webinar 16.1.2018 Eija Korpelainen, Kimmo Mattila, Maria Lehtivaara Big thanks to Jan Kreuze and Jari Valkonen! Outline Small interfering
VirusDetect pipeline - virus detection with small RNA sequencing CSC webinar 16.1.2018 Eija Korpelainen, Kimmo Mattila, Maria Lehtivaara Big thanks to Jan Kreuze and Jari Valkonen! Outline Small interfering
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis
 A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
Golden Helix s End-to-End Solution for Clinical Labs
 Golden Helix s End-to-End Solution for Clinical Labs Steven Hystad - Field Application Scientist Nathan Fortier Senior Software Engineer 20 most promising Biotech Technology Providers Top 10 Analytics
Golden Helix s End-to-End Solution for Clinical Labs Steven Hystad - Field Application Scientist Nathan Fortier Senior Software Engineer 20 most promising Biotech Technology Providers Top 10 Analytics
Package TargetScoreData
 Title TargetScoreData Version 1.14.0 Author Yue Li Package TargetScoreData Maintainer Yue Li April 12, 2018 Precompiled and processed mirna-overexpression fold-changes from 84 Gene
Title TargetScoreData Version 1.14.0 Author Yue Li Package TargetScoreData Maintainer Yue Li April 12, 2018 Precompiled and processed mirna-overexpression fold-changes from 84 Gene
Arabidopsis thaliana small RNA Sequencing. Report
 Arabidopsis thaliana small RNA Sequencing Report September 2015 Project Information Client Name Client Company / Institution Macrogen Order Number Order ID Species Arabidopsis thaliana Reference UCSC hg19
Arabidopsis thaliana small RNA Sequencing Report September 2015 Project Information Client Name Client Company / Institution Macrogen Order Number Order ID Species Arabidopsis thaliana Reference UCSC hg19
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Analysis with SureCall 2.1
 Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
High AU content: a signature of upregulated mirna in cardiac diseases
 https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
https://helda.helsinki.fi High AU content: a signature of upregulated mirna in cardiac diseases Gupta, Richa 2010-09-20 Gupta, R, Soni, N, Patnaik, P, Sood, I, Singh, R, Rawal, K & Rani, V 2010, ' High
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities
 Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Metabolomic and Proteomics Solutions for Integrated Biology. Christine Miller Omics Market Manager ASMS 2015
 Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
RNA Secondary Structures: A Case Study on Viruses Bioinformatics Senior Project John Acampado Under the guidance of Dr. Jason Wang
 RNA Secondary Structures: A Case Study on Viruses Bioinformatics Senior Project John Acampado Under the guidance of Dr. Jason Wang Table of Contents Overview RSpredict JAVA RSpredict WebServer RNAstructure
RNA Secondary Structures: A Case Study on Viruses Bioinformatics Senior Project John Acampado Under the guidance of Dr. Jason Wang Table of Contents Overview RSpredict JAVA RSpredict WebServer RNAstructure
P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( 鄧致剛 ); g ( 黄栢榕 ) Bioinformatics Center, Chang Gung University.
 Small RNA High Throughput Sequencing Analysis I P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( 鄧致剛 ); g ( 黄栢榕 ) Bioinformatics Center, Chang Gung University. Prominent members of the RNA family Classic RNAs mediating
Small RNA High Throughput Sequencing Analysis I P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( 鄧致剛 ); g ( 黄栢榕 ) Bioinformatics Center, Chang Gung University. Prominent members of the RNA family Classic RNAs mediating
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies
 Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
ChIP-seq hands-on. Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs
 ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
A Statistical Framework for Classification of Tumor Type from microrna Data
 DEGREE PROJECT IN MATHEMATICS, SECOND CYCLE, 30 CREDITS STOCKHOLM, SWEDEN 2016 A Statistical Framework for Classification of Tumor Type from microrna Data JOSEFINE RÖHSS KTH ROYAL INSTITUTE OF TECHNOLOGY
DEGREE PROJECT IN MATHEMATICS, SECOND CYCLE, 30 CREDITS STOCKHOLM, SWEDEN 2016 A Statistical Framework for Classification of Tumor Type from microrna Data JOSEFINE RÖHSS KTH ROYAL INSTITUTE OF TECHNOLOGY
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University.
 Databases and Tools for High Throughput Sequencing Analysis P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University. HTseq Platforms Applications on Biomedical Sciences
Databases and Tools for High Throughput Sequencing Analysis P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University. HTseq Platforms Applications on Biomedical Sciences
Transcriptome Analysis
 Transcriptome Analysis Data Preprocessing Sample Preparation Illumina Sequencing Demultiplexing Raw FastQ Reference Genome (fasta) Reference Annotation (GTF) Reference Genome Analysis Tophat Accepted hits
Transcriptome Analysis Data Preprocessing Sample Preparation Illumina Sequencing Demultiplexing Raw FastQ Reference Genome (fasta) Reference Annotation (GTF) Reference Genome Analysis Tophat Accepted hits
Methods 59 (2013) Contents lists available at SciVerse ScienceDirect. Methods. journal homepage:
 Methods 59 (2013) 154 163 Contents lists available at SciVerse ScienceDirect Methods journal homepage: www.elsevier.com/locate/ymeth Identifying transcriptional mirna biomarkers by integrating high-throughput
Methods 59 (2013) 154 163 Contents lists available at SciVerse ScienceDirect Methods journal homepage: www.elsevier.com/locate/ymeth Identifying transcriptional mirna biomarkers by integrating high-throughput
omiras: MicroRNA regulation of gene expression
 omiras: MicroRNA regulation of gene expression Sören Müller, Goethe University of Frankfurt am Main Molecular Bioinformatics Group, Institute of Computer Science Plant Molecular Biology Group, Institute
omiras: MicroRNA regulation of gene expression Sören Müller, Goethe University of Frankfurt am Main Molecular Bioinformatics Group, Institute of Computer Science Plant Molecular Biology Group, Institute
HBV. Next Generation Sequencing, data analysis and reporting. Presenter Leen-Jan van Doorn
 HBV Next Generation Sequencing, data analysis and reporting Presenter Leen-Jan van Doorn HBV Forum 3 October 24 th, 2017 Marriott Marquis, Washington DC www.forumresearch.org HBV Biomarkers HBV biomarkers:
HBV Next Generation Sequencing, data analysis and reporting Presenter Leen-Jan van Doorn HBV Forum 3 October 24 th, 2017 Marriott Marquis, Washington DC www.forumresearch.org HBV Biomarkers HBV biomarkers:
STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells
 CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
Synthetic microrna Reference Standards Genomics Research Group ABRF 2015
 Synthetic microrna Reference Standards Genomics Research Group ABRF 2015 Don A. Baldwin, Ph.D. support@signalbiology.com Reference samples for Platform evaluation Protocol development Assay service improvement
Synthetic microrna Reference Standards Genomics Research Group ABRF 2015 Don A. Baldwin, Ph.D. support@signalbiology.com Reference samples for Platform evaluation Protocol development Assay service improvement
Deciphering the Role of micrornas in BRD4-NUT Fusion Gene Induced NUT Midline Carcinoma
 www.bioinformation.net Volume 13(6) Hypothesis Deciphering the Role of micrornas in BRD4-NUT Fusion Gene Induced NUT Midline Carcinoma Ekta Pathak 1, Bhavya 1, Divya Mishra 1, Neelam Atri 1, 2, Rajeev
www.bioinformation.net Volume 13(6) Hypothesis Deciphering the Role of micrornas in BRD4-NUT Fusion Gene Induced NUT Midline Carcinoma Ekta Pathak 1, Bhavya 1, Divya Mishra 1, Neelam Atri 1, 2, Rajeev
Supplemental Figure 1. Small RNA size distribution from different soybean tissues.
 Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Lombardi Cancer Center Georgetown University Washington, DC
 Lombardi Cancer Center Georgetown University Washington, DC Lombardi Cancer Center Georgetown University Washington, DC microrna and other nucleic acids analysis in liquid biopsies Anton Wellstein World
Lombardi Cancer Center Georgetown University Washington, DC Lombardi Cancer Center Georgetown University Washington, DC microrna and other nucleic acids analysis in liquid biopsies Anton Wellstein World
Small RNA-Seq and profiling
 Small RNA-Seq and profiling Y. Hoogstrate 1,2 1 Department of Bioinformatics & Department of Urology ErasmusMC, Rotterdam 2 CTMM Translational Research IT (TraIT) BioSB: 5th RNA-seq data analysis course,
Small RNA-Seq and profiling Y. Hoogstrate 1,2 1 Department of Bioinformatics & Department of Urology ErasmusMC, Rotterdam 2 CTMM Translational Research IT (TraIT) BioSB: 5th RNA-seq data analysis course,
Multi-omics data integration colon cancer using proteogenomics approach
 Dept. of Medical Oncology Multi-omics data integration colon cancer using proteogenomics approach DTL Focus meeting, 29 August 2016 Thang Pham OncoProteomics Laboratory, Dept. of Medical Oncology VU University
Dept. of Medical Oncology Multi-omics data integration colon cancer using proteogenomics approach DTL Focus meeting, 29 August 2016 Thang Pham OncoProteomics Laboratory, Dept. of Medical Oncology VU University
CRS4 Seminar series. Inferring the functional role of micrornas from gene expression data CRS4. Biomedicine. Bioinformatics. Paolo Uva July 11, 2012
 CRS4 Seminar series Inferring the functional role of micrornas from gene expression data CRS4 Biomedicine Bioinformatics Paolo Uva July 11, 2012 Partners Pharmaceutical company Fondazione San Raffaele,
CRS4 Seminar series Inferring the functional role of micrornas from gene expression data CRS4 Biomedicine Bioinformatics Paolo Uva July 11, 2012 Partners Pharmaceutical company Fondazione San Raffaele,
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Zhao et al. BMC Bioinformatics (2017) 18:180 DOI /s
 Zhao et al. BMC Bioinformatics (2017) 18:180 DOI 10.1186/s12859-017-1601-4 SOFTWARE QuickMIRSeq: a pipeline for quick and accurate quantification of both known mirnas and isomirs by jointly processing
Zhao et al. BMC Bioinformatics (2017) 18:180 DOI 10.1186/s12859-017-1601-4 SOFTWARE QuickMIRSeq: a pipeline for quick and accurate quantification of both known mirnas and isomirs by jointly processing
Profiling micrornas in lung tissue from pigs infected with Actinobacillus pleuropneumoniae
 Podolska et al. BMC Genomics 2012, 13:459 RESEARCH ARTICLE Open Access Profiling micrornas in lung tissue from pigs infected with Actinobacillus pleuropneumoniae Agnieszka Podolska 1,2, Christian Anthon
Podolska et al. BMC Genomics 2012, 13:459 RESEARCH ARTICLE Open Access Profiling micrornas in lung tissue from pigs infected with Actinobacillus pleuropneumoniae Agnieszka Podolska 1,2, Christian Anthon
Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009
 Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009 1 Abstract A stretch of chimpanzee DNA was annotated using tools including BLAST, BLAT, and Genscan. Analysis of Genscan predicted genes revealed
Annotation of Chimp Chunk 2-10 Jerome M Molleston 5/4/2009 1 Abstract A stretch of chimpanzee DNA was annotated using tools including BLAST, BLAT, and Genscan. Analysis of Genscan predicted genes revealed
Long non-coding RNAs
 Long non-coding RNAs Dominic Rose Bioinformatics Group, University of Freiburg Bled, Feb. 2011 Outline De novo prediction of long non-coding RNAs (lncrnas) Genome-wide RNA gene-finding Intrinsic properties
Long non-coding RNAs Dominic Rose Bioinformatics Group, University of Freiburg Bled, Feb. 2011 Outline De novo prediction of long non-coding RNAs (lncrnas) Genome-wide RNA gene-finding Intrinsic properties
microrna PCR System (Exiqon), following the manufacturer s instructions. In brief, 10ng of
 SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
IPA Advanced Training Course
 IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
ChIP-seq data analysis
 ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
Simple, rapid, and reliable RNA sequencing
 Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Chip Seq Peak Calling in Galaxy
 Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
PSSV User Manual (V1.0)
 PSSV User Manual (V1.0) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
PSSV User Manual (V1.0) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
Deploying the full transcriptome using RNA sequencing. Jo Vandesompele, CSO and co-founder The Non-Coding Genome May 12, 2016, Leuven
 Deploying the full transcriptome using RNA sequencing Jo Vandesompele, CSO and co-founder The Non-Coding Genome May 12, 2016, Leuven Roadmap Biogazelle the power of RNA reasons to study non-coding RNA
Deploying the full transcriptome using RNA sequencing Jo Vandesompele, CSO and co-founder The Non-Coding Genome May 12, 2016, Leuven Roadmap Biogazelle the power of RNA reasons to study non-coding RNA
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success
 micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
How To Use MiRSEA. Junwei Han. July 1, Overview 1. 2 Get the pathway-mirna correlation profile(pmset) and a weighting matrix 2
 How To Use MiRSEA Junwei Han July 1, 2015 Contents 1 Overview 1 2 Get the pathway-mirna correlation profile(pmset) and a weighting matrix 2 3 Discovering the dysregulated pathways(or prior gene sets) based
How To Use MiRSEA Junwei Han July 1, 2015 Contents 1 Overview 1 2 Get the pathway-mirna correlation profile(pmset) and a weighting matrix 2 3 Discovering the dysregulated pathways(or prior gene sets) based
Profiles of gene expression & diagnosis/prognosis of cancer. MCs in Advanced Genetics Ainoa Planas Riverola
 Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles
 On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles Ying-Wooi Wan 1,2,4, Claire M. Mach 2,3, Genevera I. Allen 1,7,8, Matthew L. Anderson 2,4,5 *, Zhandong Liu 1,5,6,7 * 1 Departments of Pediatrics
On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles Ying-Wooi Wan 1,2,4, Claire M. Mach 2,3, Genevera I. Allen 1,7,8, Matthew L. Anderson 2,4,5 *, Zhandong Liu 1,5,6,7 * 1 Departments of Pediatrics
Prediction of micrornas and their targets
 Prediction of micrornas and their targets Introduction Brief history mirna Biogenesis Computational Methods Mature and precursor mirna prediction mirna target gene prediction Summary micrornas? RNA can
Prediction of micrornas and their targets Introduction Brief history mirna Biogenesis Computational Methods Mature and precursor mirna prediction mirna target gene prediction Summary micrornas? RNA can
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Accurate detection for a wide range of mutation and editing sites of micrornas from small RNA high-throughput sequencing profiles
 Published online 26 May 2016 Nucleic Acids Research, 2016, Vol. 44, No. 14 e123 doi: 10.1093/nar/gkw471 Accurate detection for a wide range of mutation and editing sites of micrornas from small RNA high-throughput
Published online 26 May 2016 Nucleic Acids Research, 2016, Vol. 44, No. 14 e123 doi: 10.1093/nar/gkw471 Accurate detection for a wide range of mutation and editing sites of micrornas from small RNA high-throughput
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP)
 Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Reliable Prediction of Viral RNA Structures
 Reliable Prediction of Viral RNA Structures Winterseminar 2016, Bled Roman Ochsenreiter TBI Wien Bled, 15.2.2016 Roman Ochsenreiter TBI Wien Reliable Prediction of Viral RNA Structures Bled, 15.2.2016
Reliable Prediction of Viral RNA Structures Winterseminar 2016, Bled Roman Ochsenreiter TBI Wien Bled, 15.2.2016 Roman Ochsenreiter TBI Wien Reliable Prediction of Viral RNA Structures Bled, 15.2.2016
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Supplementary materials and methods.
 Supplementary materials and methods. Microarray printing and data analysis. The LNA-modified oligonucleotide probe set for all annotated mirnas from mouse (Mus musculus) and human (Homo sapiens) in the
Supplementary materials and methods. Microarray printing and data analysis. The LNA-modified oligonucleotide probe set for all annotated mirnas from mouse (Mus musculus) and human (Homo sapiens) in the
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
The Value of Omics in Cardiovascular Research: Getting More Comfortable, More Frustrated or More Curious
 The Value of Omics in Cardiovascular Research: Getting More Comfortable, More Frustrated or More Curious Daniel Levy, MD Framingham Heart Study Population Sciences Branch National Heart, Lung, and Blood
The Value of Omics in Cardiovascular Research: Getting More Comfortable, More Frustrated or More Curious Daniel Levy, MD Framingham Heart Study Population Sciences Branch National Heart, Lung, and Blood
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
ChromHMM Tutorial. Jason Ernst Assistant Professor University of California, Los Angeles
 ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
mirviewer: A multispecies microrna homologous viewer
 BMC Research Notes This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. mirviewer: A multispecies
BMC Research Notes This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. mirviewer: A multispecies
I conducted an internship at the Texas Biomedical Research Institute in the
 Ashley D. Falk ashleydfalk@gmail.com My Internship at Texas Biomedical Research Institute I conducted an internship at the Texas Biomedical Research Institute in the spring of 2013. In this report I will
Ashley D. Falk ashleydfalk@gmail.com My Internship at Texas Biomedical Research Institute I conducted an internship at the Texas Biomedical Research Institute in the spring of 2013. In this report I will
Deep Sequencing the MicroRNA Transcriptome in Colorectal Cancer
 Deep Sequencing the MicroRNA Transcriptome in Colorectal Cancer Kristina Schee 1, Susanne Lorenz 1,2, Merete Molton Worren 3, Clara-Cecilie Günther 4, Marit Holden 4, Eivind Hovig 1,3,5, Øystein Fodstad
Deep Sequencing the MicroRNA Transcriptome in Colorectal Cancer Kristina Schee 1, Susanne Lorenz 1,2, Merete Molton Worren 3, Clara-Cecilie Günther 4, Marit Holden 4, Eivind Hovig 1,3,5, Øystein Fodstad
Bioinformatic analyses: methodology for allergen similarity search. Zoltán Divéki, Ana Gomes EFSA GMO Unit
 Bioinformatic analyses: methodology for allergen similarity search Zoltán Divéki, Ana Gomes EFSA GMO Unit EFSA info session on applications - GMO Parma, Italy 28 October 2014 BIOINFORMATIC ANALYSES Analysis
Bioinformatic analyses: methodology for allergen similarity search Zoltán Divéki, Ana Gomes EFSA GMO Unit EFSA info session on applications - GMO Parma, Italy 28 October 2014 BIOINFORMATIC ANALYSES Analysis
High Throughput Sequence (HTS) data analysis. Lei Zhou
 High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
The Focused Exome service at Bristol Genetics Laboratory
 The Focused Exome service at Bristol Genetics Laboratory Chris Buxton Maggie Williams July 2016 Bristol Clinical exome Service to mid July Validation: Agilent FE kit, NextSeq 500 and new pipeline 1st reports
The Focused Exome service at Bristol Genetics Laboratory Chris Buxton Maggie Williams July 2016 Bristol Clinical exome Service to mid July Validation: Agilent FE kit, NextSeq 500 and new pipeline 1st reports
sirna count per 50 kb small RNAs matching the direct strand Repeat length (bp) per 50 kb repeats in the chromosome
 Qi et al. 26-3-2564C Qi et al., Figure S1 sirna count per 5 kb small RNAs matching the direct strand sirna count per 5 kb small RNAs matching the complementary strand Repeat length (bp) per 5 kb repeats
Qi et al. 26-3-2564C Qi et al., Figure S1 sirna count per 5 kb small RNAs matching the direct strand sirna count per 5 kb small RNAs matching the complementary strand Repeat length (bp) per 5 kb repeats
VARIANT PRIORIZATION AND ANALYSIS INCORPORATING PROBLEMATIC REGIONS OF THE GENOME ANIL PATWARDHAN
 VARIANT PRIORIZATION AND ANALYSIS INCORPORATING PROBLEMATIC REGIONS OF THE GENOME ANIL PATWARDHAN Email: apatwardhan@personalis.com MICHAEL CLARK Email: michael.clark@personalis.com ALEX MORGAN Email:
VARIANT PRIORIZATION AND ANALYSIS INCORPORATING PROBLEMATIC REGIONS OF THE GENOME ANIL PATWARDHAN Email: apatwardhan@personalis.com MICHAEL CLARK Email: michael.clark@personalis.com ALEX MORGAN Email:
ncounter Assay Automated Process Immobilize and align reporter for image collecting and barcode counting ncounter Prep Station
 ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
MicroRNAs: novel regulators in skin research
 MicroRNAs: novel regulators in skin research Eniko Sonkoly, Andor Pivarcsi KI, Department of Medicine, Unit of Dermatology and Venerology What are micrornas? Small, ~21-mer RNAs 1993: The first mirna discovered,
MicroRNAs: novel regulators in skin research Eniko Sonkoly, Andor Pivarcsi KI, Department of Medicine, Unit of Dermatology and Venerology What are micrornas? Small, ~21-mer RNAs 1993: The first mirna discovered,
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Medtech Training Guide
 Medtech Training Guide Clinical Audit Tool Copyright Medtech Limited Page 1 of 80 Clinical Audit Tool v4.0 Contents Chapter 1: Getting Started... 3 Logging In... 3 Logging Out... 3 Setting the Preliminary
Medtech Training Guide Clinical Audit Tool Copyright Medtech Limited Page 1 of 80 Clinical Audit Tool v4.0 Contents Chapter 1: Getting Started... 3 Logging In... 3 Logging Out... 3 Setting the Preliminary
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
ncounter Assay Automated Process Capture & Reporter Probes Bind reporter to surface Remove excess reporters Hybridize CodeSet to RNA
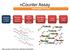 ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
Identification of mirnas in Eucalyptus globulus Plant by Computational Methods
 International Journal of Pharmaceutical Science Invention ISSN (Online): 2319 6718, ISSN (Print): 2319 670X Volume 2 Issue 5 May 2013 PP.70-74 Identification of mirnas in Eucalyptus globulus Plant by Computational
International Journal of Pharmaceutical Science Invention ISSN (Online): 2319 6718, ISSN (Print): 2319 670X Volume 2 Issue 5 May 2013 PP.70-74 Identification of mirnas in Eucalyptus globulus Plant by Computational
Below, we included the point-to-point response to the comments of both reviewers.
 To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
Transcript reconstruction
 Transcript reconstruction Summary I Data types, file formats and utilities Annotation: Genomic regions Genes Peaks bedtools Alignment: Map reads BAM/SAM Samtools Aggregation: Summary files Wig (UCSC) TDF
Transcript reconstruction Summary I Data types, file formats and utilities Annotation: Genomic regions Genes Peaks bedtools Alignment: Map reads BAM/SAM Samtools Aggregation: Summary files Wig (UCSC) TDF
Viral genome sequencing: applications to clinical management and public health. Professor Judy Breuer
 Viral genome sequencing: applications to clinical management and public health Professor Judy Breuer Why do whole viral genome sequencing Genome sequencing allows detection of multigenic resistance in
Viral genome sequencing: applications to clinical management and public health Professor Judy Breuer Why do whole viral genome sequencing Genome sequencing allows detection of multigenic resistance in
Long non coding RNA in the pea aphid; iden3fica3on and compara3ve expression in sexual and asexual embryos
 Long non coding RNA in the pea aphid; iden3fica3on and compara3ve expression in sexual and asexual embryos Fabrice Legeai, Thomas Derrien, Valen3n Wucher, Audrey David, Gael Le Trionnaire and Denis Tagu
Long non coding RNA in the pea aphid; iden3fica3on and compara3ve expression in sexual and asexual embryos Fabrice Legeai, Thomas Derrien, Valen3n Wucher, Audrey David, Gael Le Trionnaire and Denis Tagu
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
PSSV User Manual (V2.1)
 PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
Gene-microRNA network module analysis for ovarian cancer
 Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice
 SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
SUPPLEMENTARY INFORMATION RNA interference induced hepatotoxicity results from loss of the first synthesized isoform of microrna-122 in mice Paul N Valdmanis, Shuo Gu, Kirk Chu, Lan Jin, Feijie Zhang,
Package leukemiaseset
 Package leukemiaseset August 14, 2018 Type Package Title Leukemia's microarray gene expression data (expressionset). Version 1.16.0 Date 2013-03-20 Author Sara Aibar, Celia Fontanillo and Javier De Las
Package leukemiaseset August 14, 2018 Type Package Title Leukemia's microarray gene expression data (expressionset). Version 1.16.0 Date 2013-03-20 Author Sara Aibar, Celia Fontanillo and Javier De Las
SUPPLEMENTAL DATA AGING, July 2014, Vol. 6 No. 7
 SUPPLEMENTAL DATA Figure S1. Muscle mass changes in different anatomical regions with age. (A) The TA and gastrocnemius muscle showed a significant loss of weight in aged mice (24 month old) compared to
SUPPLEMENTAL DATA Figure S1. Muscle mass changes in different anatomical regions with age. (A) The TA and gastrocnemius muscle showed a significant loss of weight in aged mice (24 month old) compared to
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
The Role of micrornas in Pain. Phd Work of: Rehman Qureshi Advisor: Ahmet Sacan
 The Role of micrornas in Pain Phd Work of: Rehman Qureshi Advisor: Ahmet Sacan 1 Motivation for molecular analysis of Pain Chronic pain one of the most common causes for seeking medical treatment Approximate
The Role of micrornas in Pain Phd Work of: Rehman Qureshi Advisor: Ahmet Sacan 1 Motivation for molecular analysis of Pain Chronic pain one of the most common causes for seeking medical treatment Approximate
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
Screening for novel oncology biomarker panels using both DNA and protein microarrays. John Anson, PhD VP Biomarker Discovery
 Screening for novel oncology biomarker panels using both DNA and protein microarrays John Anson, PhD VP Biomarker Discovery Outline of presentation Introduction to OGT and our approach to biomarker studies
Screening for novel oncology biomarker panels using both DNA and protein microarrays John Anson, PhD VP Biomarker Discovery Outline of presentation Introduction to OGT and our approach to biomarker studies
