Supplemental Information. A Highly Sensitive and Robust Method. for Genome-wide 5hmC Profiling. of Rare Cell Populations
|
|
|
- Tyler Parsons
- 5 years ago
- Views:
Transcription
1 Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle Xu, ri M. Melnick, Ross L. Levine, and huan He
2 ng ng ng ng cell cell hm-seal hm-seal D B E F hm-seal hm-seal cell cell ng ng ng ng Pearson's r % 8% 6% % % % hm-sealng ng cell Portion of tags in hm clusters Distinct reads (million) number of called hm peaks otal reads (million) hm-seal ng ng cell % 8% 6% % % % hm-seal ng ng cell 8 6 Portion of B-seq - validated peaks Figure S hm-seal nano-hm-seal μg rep μg rep ng rep ng rep ng rep ng rep cell rep cell rep Normalized hm reads B.. -k SS S k -k SS S k. H - - Distance to B-seq hm sites 3 Normalized hm reads hm low hm medium hm high Input - - Distance to oxrrbs hm Is nano-hm-seal rep nano-hm-seal rep Input Normalized hm reads
3 Figure S B %m+hm MP MP MEP MEP hyper DMR 6 8 %m+hm hypo DMR hm MP hm MEP - - Distance to DMR Normalized hm reads - - Distance to DMR MP MEP Normalized reads Normalized hm reads Normalized reads hm (log RPKM) -- Distance to DMR 7 -- Distance to DMR MP MP MEP MEP D hm LSK cells me3 H3K7 ac hm MP cells me3 H3K7 ac hm MP cells MEP cells me3 H3K7 ac hm me3 H3K7 ac ene expression E LSK hm (log RPKM) hm hm (log RPKM) (log RPKM) MP MP MEP (log RPKM) (log RPKM) me3 (log RPKM) H3K7c (log RPKM) -k SS S k -k SS S k -k SS S k -k SS S k
4 Figure S3 hm LSK MP MP MEP H3K7 me3 ac hm H3K7 me3 ac hm H3K7 me3 ac hm H3K7 me3 ac -.k.k -k k B hm -.k.k -k k LSK MP MP MEP MPP MP MP MEP -.k.k -k k me3 MPP MP MP MEP MPP MP MP MEP -.k.k -k k H3K7c MPP MP MP MEP I II III IV -k k -k k -k k cluster I cluster II cluster III cluster IV immune system development hemopoiesis leukocyte activation cell activation apoptotic mitochondrial changes nucleoside triphosphate metabolic histone modification peptidyl-lysine modification erythrocyte differentiation tetrapyrrole biosynthetic process leukocyte differentiation cell chemotaxis lymphocyte differentiation cell differentiation leukocyte migration carbohydrate derivative catabolic vacuole organization peptide hormone stimulus protein kinase negative regulation leukocyte activation 3 3 -lg(p-value) D -k k abnormal hematopoietic cell number abnormal immune system cell morphology abnormal leukocyte morphology abnormal adaptive immunity abnormal immune cell physiology decreased erythrocyte cell number reticulocytosis abnormal mean corpuscular volume hemolytic anemia abnormal erythroid progenitor cell morphology abnormal mononuclear cell differentiation abnormal myeloblast morphology/development abnormal lymphopoiesis abnormal B cell morphology abnormal cell differentiation -k k cluster I cluster II cluster III cluster IV abnormal mononuclear phagocyte morphology abnormal immune tolerance autoimmune response decreased cholesterol level abnormal acute inflammation 6 8 -lg(p-value)
5 Figure S L HS S HS MPP MP MP MF N Mono LP B D D8 NK MEP Ery EryB Log / NK cells Size: 7 () B cells Size: 87 8 Erythroid Progenitors Size: 66 6 Erythroid Size: 66 Lymphoid Progenitors Size: 66 ommon Size: 799 Myeloid Size: 673 Myeloid Progenitors Size: 866 Progenitors Size: 6337 Normalized hm reads 8 6 -% -% % % % % anyon F3 mpp W mpp Input B hm loss region hm gain region Motif Factor P-value Motif Factor P-value EV e-63 ER e-9 ata e-8 ata e-6 ES e- PU.-IRF e-7 SpiB e-6 PU. e-6 IRF e-96 ELF e-8 EWS:ER e- ata e-3 IRF e-6 IRF e-6 Fli e-3 BP e-8 EHF e- S e-7 S e- ES e-3
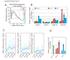
6 Supplementary Figure Legends Figure S. lobal comparison of conventional hm-seal and nano-hm-seal sequencing data Related to Figure ( and B) verage profiles () and heatmap (B) across gene regions ±, bp for nano-hm-seal libraries. Each row represents a gene, ordered by the mean value signals. Regular hm-seal libraries were generated from μg genomic DN and used as references. () enome-wide correlations (,bp tiling windows) of sequencing results obtained using conventional hm-seal and nano-hm-seal (with ng and ng DN as well as genomic DN isolated from, cells). (D) Fraction of reads located in hm high-density clusters for libraries constructed using different amounts of input DN. (E) Preseq library complexity curves for different libraries. (F) he number of high confident hm-enriched peaks (right axis) called from different libraries and the portion of peaks validated by B-seq hm sites (left axis). he hm-enriched peaks were considered as validated if B-seq detected hm sites reside in the area of bp surrounding the peak center. () he distribution of nano-hm-seal (ng) signals at hm sites detected by B-seq. hm sites were further divided into low ( - %), medium ( - %), high (% and above) subgroups (, sites were randomly selected for each subgroup) (H) he distribution of nano-hm-seal (ng) signals at 6 hm-containing Is detected by ox-rrbs method. Figure S. hm levels correlate with DN methylation and with histone marks in HS and progenitor cells Related to Figure () Boxplot to show the level of DN modification (m+hm) at hypo (upper) and hyper (lower) DMRs. (B-) he distribution of hm (B) and () signals at hypo (upper) and hyper (lower) DMRs. (D) Heatmap displaying the reads density distribution of hm and indicated histone modifications in all annotated genes ordered by decreasing expression in LSK, MP, MP cells and MEP cells. Each row represents a gene. Due to lack of LSK histone modification data in published datasets, LSK hm data was compared with histone modification data obtained from MPP cells. (E) Scatter plot displaying the correlation of hm with,, e3 and H3K7ac in LSK, MP, MP cells and MEP cells. Each dot represents a gene. Due to lack of LSK histone modification data in published dataset, LSK hm data was compared with histone modification data obtained from MPP cells. ll values are represented as log RPKM. he spearman rank correlation coefficient is shown (ρ) in each comparison.
7 Figure S3. he distribution of hm and histone modifications at selected genomic regions Related to Figure () Heatmap displaying read densities of -seq, hm and histone modifications around the -seq signal-enriched peaks. -seq peaks were divided into four clusters by k-means clustering and ranked according to decreased signal values. Due to lack of LSK histone modification data in the published dataset, LSK -seq and hm data were compared with histone modification data obtained from MPP cells. (B) Heatmap displaying read densities of hm,,, me3 and H3K7c around DhMRs across differentiation stages. lusters were generated by k-means clustering of hm signals. Histone modification datasets were arranged to match the order of hm heatmap. (-D) Functional annotation of DhMRs in each cluster was performed using RE. he top over-represented categories belonging to ene Ontology biological process () and Mouse enome Informatics phenotype ontology (D) are shown. he x axis values correspond to the log-transformed binomial P-values. Figure S. he relationship between hm and functional regulatory elements in W or ML model mice Related to Figure and Figure 3 () he activity of lineage specific enhancers during hematopoiesis. Heatmap showing lineage-specific hematopoiesis enhancers with k-means cluster analysis of signals (K=9). he genomic location of hematopoietic enhancers and hip-seq dataset were obtained from a previously published study (Lara-stiaso et al., ). K-mean cluster analysis is performed by R package pheatmap. (B) op enriched known transcription factor binding motifs detected at DhMRs (left: hm loss; right: hm gain) in MPP cells. Motif information was obtained from the Homer motif database. () Normalized distribution profiles of hm in MPP cells across DN methylation canyon regions detected in hematopoietic stem cells. he genomic location of DN canyon was obtained from a previously published study (Jeong et al., ).
8 ables: Summary statistics for the nano-hm-seal sequencing experiments. Related to Figure Sample rawreads mappedreads mapratio UniqueReads UniqueRatio mes_ng_ mes_ng_ mes_ng_ mes_ng_ mes_cell_ mes_cell_ mes_ng_input mes_ng_input mes_cell_input hematopoiesis_cmp_ hematopoiesis_cmp_ hematopoiesis_cmp_ hematopoiesis_gmp_ hematopoiesis_gmp_ hematopoiesis_gmp_ hematopoiesis_lsk_ hematopoiesis_lsk_ hematopoiesis_lsk_ hematopoiesis_mep_ hematopoiesis_mep_ hematopoiesis_mep_ hematopoiesis_input leukemia_f3 gmp leukemia_f3 mpp leukemia_f3 gmp leukemia_f3 mpp leukemia_f3_3_gmp leukemia_f3_3_mpp leukemia_w gmp leukemia_w mpp leukemia_w gmp leukemia_w mpp leukemia_w_3_gmp leukemia_w_3_mpp leukemia_w gmp leukemia_w mpp leukemia_w gmp leukemia_w mpp leukemia_f3_gmp_input leukemia_f3_mpp_input leukemia_w_mpp_input
9 References Jeong, M., Sun, D., Luo, M., Huang, Y., hallen,.., Rodriguez, B., Zhang, X., havez, L., Wang, H., Hannah, R., et al. (). Large conserved domains of low DN methylation maintained by Dnmt3a. Nature genetics 6, 7-3. Lara-stiaso, D., Weiner,., Lorenzo-Vivas, E., Zaretsky, I., Jaitin, D.., David, E., Keren-Shaul, H., Mildner,., Winter, D., Jung, S., et al. (). Immunogenetics. hromatin state dynamics during blood formation. Science 3,
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
Supplemental Information. Granulocyte-Monocyte Progenitors and. Monocyte-Dendritic Cell Progenitors Independently
 Immunity, Volume 47 Supplemental Information Granulocyte-Monocyte Progenitors and Monocyte-endritic ell Progenitors Independently Produce Functionally istinct Monocytes lberto Yáñez, Simon G. oetzee, ndre
Immunity, Volume 47 Supplemental Information Granulocyte-Monocyte Progenitors and Monocyte-endritic ell Progenitors Independently Produce Functionally istinct Monocytes lberto Yáñez, Simon G. oetzee, ndre
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Supplemental Information. Genomic Characterization of Murine. Monocytes Reveals C/EBPb Transcription. Factor Dependence of Ly6C Cells
 Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Large conserved domains of low DNA methylation maintained by Dnmt3a
 Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Figure 1. Dnmt3b expression in murine and human knee joint cartilage. (A) Representative images
 Figure Legends Figure. expression in murine and human knee joint cartilage. () Representative images showing that Dnmta is not expressed in chondrocytes from mo W articular cartilage [Dnmta expression
Figure Legends Figure. expression in murine and human knee joint cartilage. () Representative images showing that Dnmta is not expressed in chondrocytes from mo W articular cartilage [Dnmta expression
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Rice in vivo RNA structurome reveals RNA secondary structure conservation and divergence in plants
 Rice in vivo RN structurome reveals RN secondary structure conservation and divergence in plants Hongjing Deng 1,2,,5, Jitender heema 3, Hang Zhang 2, Hugh Woolfenden 2, Matthew Norris 2, Zhenshan Liu
Rice in vivo RN structurome reveals RN secondary structure conservation and divergence in plants Hongjing Deng 1,2,,5, Jitender heema 3, Hang Zhang 2, Hugh Woolfenden 2, Matthew Norris 2, Zhenshan Liu
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
Supplementary information
 Supplementary information High fat diet-induced changes of mouse hepatic transcription and enhancer activity can be reversed by subsequent weight loss Majken Siersbæk, Lyuba Varticovski, Shutong Yang,
Supplementary information High fat diet-induced changes of mouse hepatic transcription and enhancer activity can be reversed by subsequent weight loss Majken Siersbæk, Lyuba Varticovski, Shutong Yang,
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Results. Abstract. Introduc4on. Conclusions. Methods. Funding
 . expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
. expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature19360 Supplementary Tables Supplementary Table 1. Number of monoclonal reads in each sample Sample Number of cells Total reads Aligned reads Monoclonal reads
SUPPLEMENTARY INFORMATION doi:10.1038/nature19360 Supplementary Tables Supplementary Table 1. Number of monoclonal reads in each sample Sample Number of cells Total reads Aligned reads Monoclonal reads
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and
 Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
cis-regulatory enrichment analysis in human, mouse and fly
 cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types. Epigenome Informatics Workshop Bioinformatics Research Laboratory
 Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Gene-microRNA network module analysis for ovarian cancer
 Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Supplementary Fig.S1. MeDIP RPKM distribution of 5kb windows. Relative expression of DNMT. Percentage of genome. (fold change)
 Supplementary Fig.S1 Percentage of genome 10% 8% 6% 4% 2% 0 MeDIP RPKM distribution of 5kb windows 0 0. 0.5 0.75 >1 MeDIP RPKM value EC Relative expression of DNMT (fold change) 12 6 4 3 2 1 EC DNMT1 DNMT2
Supplementary Fig.S1 Percentage of genome 10% 8% 6% 4% 2% 0 MeDIP RPKM distribution of 5kb windows 0 0. 0.5 0.75 >1 MeDIP RPKM value EC Relative expression of DNMT (fold change) 12 6 4 3 2 1 EC DNMT1 DNMT2
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
Supplementary Information
 Supplementary Information 5-hydroxymethylcytosine-mediated epigenetic dynamics during postnatal neurodevelopment and aging By Keith E. Szulwach 1,8, Xuekun Li 1,8, Yujing Li 1, Chun-Xiao Song 2, Hao Wu
Supplementary Information 5-hydroxymethylcytosine-mediated epigenetic dynamics during postnatal neurodevelopment and aging By Keith E. Szulwach 1,8, Xuekun Li 1,8, Yujing Li 1, Chun-Xiao Song 2, Hao Wu
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Supplemental Information. Genetic Regulation of Plasma Lipid Species. and Their Association with Metabolic Phenotypes
 Cell Systems, Volume 6 Supplemental Information Genetic Regulation of Plasma Lipid Species and Their Association with Metabolic Phenotypes Pooja Jha, Molly T. McDevitt, Emina Halilbasic, Evan G. Williams,
Cell Systems, Volume 6 Supplemental Information Genetic Regulation of Plasma Lipid Species and Their Association with Metabolic Phenotypes Pooja Jha, Molly T. McDevitt, Emina Halilbasic, Evan G. Williams,
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
MIR retrotransposon sequences provide insulators to the human genome
 Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
DNA Methylation Dynamics of Human Hematopoietic Stem Cell Differentiation
 Resource DNA Methylation Dynamics of Human Hematopoietic Stem Cell Differentiation Graphical Abstract Authors Matthias Farlik, Florian Halbritter, Fabian M uller,..., Thomas Lengauer, Mattia Frontini,
Resource DNA Methylation Dynamics of Human Hematopoietic Stem Cell Differentiation Graphical Abstract Authors Matthias Farlik, Florian Halbritter, Fabian M uller,..., Thomas Lengauer, Mattia Frontini,
Assignment 5: Integrative epigenomics analysis
 Assignment 5: Integrative epigenomics analysis Due date: Friday, 2/24 10am. Note: no late assignments will be accepted. Introduction CpG islands (CGIs) are important regulatory regions in the genome. What
Assignment 5: Integrative epigenomics analysis Due date: Friday, 2/24 10am. Note: no late assignments will be accepted. Introduction CpG islands (CGIs) are important regulatory regions in the genome. What
Patient characteristics of training and validation set. Patient selection and inclusion overview can be found in Supp Data 9. Training set (103)
 Roepman P, et al. An immune response enriched 72-gene prognostic profile for early stage Non-Small- Supplementary Data 1. Patient characteristics of training and validation set. Patient selection and inclusion
Roepman P, et al. An immune response enriched 72-gene prognostic profile for early stage Non-Small- Supplementary Data 1. Patient characteristics of training and validation set. Patient selection and inclusion
ChIP-seq analysis. J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand
 ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5
 sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
Figure S1, Beyer et al.
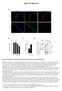 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Integrated analysis of sequencing data
 Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Measuring DNA Methylation with the MinION
 Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University Epigenetics: Modern Modern Definition of epigenetics involves heritable changes other
Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University Epigenetics: Modern Modern Definition of epigenetics involves heritable changes other
Nature Immunology: doi: /ni.3412
 Supplementary Figure 1 Gata1 expression in heamatopoietic stem and progenitor populations. (a) Unsupervised clustering according to 100 top variable genes across single pre-gm cells. The two main cell
Supplementary Figure 1 Gata1 expression in heamatopoietic stem and progenitor populations. (a) Unsupervised clustering according to 100 top variable genes across single pre-gm cells. The two main cell
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Supplementary Figures and Tables
 Supplementary Figures and Tables Supplementary Figure 1. Study design and sample collection. S.japonicum were harvested from C57 mice at 8 time points after infection. Total number of samples for RNA-Seq:
Supplementary Figures and Tables Supplementary Figure 1. Study design and sample collection. S.japonicum were harvested from C57 mice at 8 time points after infection. Total number of samples for RNA-Seq:
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB
 RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi
 Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi BioC 2017: Where Software and Biology Connect Profiling Tumor-Immune Ecosystem in Breast Cancer Immunotherapy treatments
Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi BioC 2017: Where Software and Biology Connect Profiling Tumor-Immune Ecosystem in Breast Cancer Immunotherapy treatments
Peak-calling for ChIP-seq and ATAC-seq
 Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Computational aspects of ChIP-seq. John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory
 Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
The Immune System. A macrophage. ! Functions of the Immune System. ! Types of Immune Responses. ! Organization of the Immune System
 The Immune System! Functions of the Immune System! Types of Immune Responses! Organization of the Immune System! Innate Defense Mechanisms! Acquired Defense Mechanisms! Applied Immunology A macrophage
The Immune System! Functions of the Immune System! Types of Immune Responses! Organization of the Immune System! Innate Defense Mechanisms! Acquired Defense Mechanisms! Applied Immunology A macrophage
Yingying Wei George Wu Hongkai Ji
 Stat Biosci (2013) 5:156 178 DOI 10.1007/s12561-012-9066-5 Global Mapping of Transcription Factor Binding Sites by Sequencing Chromatin Surrogates: a Perspective on Experimental Design, Data Analysis,
Stat Biosci (2013) 5:156 178 DOI 10.1007/s12561-012-9066-5 Global Mapping of Transcription Factor Binding Sites by Sequencing Chromatin Surrogates: a Perspective on Experimental Design, Data Analysis,
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Normal & Leukaemic haematopoiesis. Dr. Liu Te Chih Dept of Haematology / Oncology National University Health Services Singapore
 Normal & Leukaemic haematopoiesis 2010 Dr. Liu Te Chih Dept of Haematology / Oncology National University Health Services Singapore Use of Immunophenotyping today Lineage assignment Differentiation of
Normal & Leukaemic haematopoiesis 2010 Dr. Liu Te Chih Dept of Haematology / Oncology National University Health Services Singapore Use of Immunophenotyping today Lineage assignment Differentiation of
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
levels of genes were separated by their expression levels; 2,000 high, medium, and low
 Figure S1. Histone modification profiles near transcription start sites. The overall histone modification around transcription start sites (TSSs) was calculated. Histone modification levels of genes were
Figure S1. Histone modification profiles near transcription start sites. The overall histone modification around transcription start sites (TSSs) was calculated. Histone modification levels of genes were
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
T cell maturation. T-cell Maturation. What allows T cell maturation?
 T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation
 Published online: June 1, 15 Article Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation Evelyn N Kouwenhoven 1,, Martin Oti, Hanna Niehues 3, Simon J van
Published online: June 1, 15 Article Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation Evelyn N Kouwenhoven 1,, Martin Oti, Hanna Niehues 3, Simon J van
Long-term innate immune memory via effects on bone marrow progenitors
 Long-term innate immune memory via effects on bone marrow progenitors Helen S Goodridge, PhD helen.goodridge@csmc.edu Regenerative Medicine Institute, Cedars-Sinai Medical Center, Los Angeles, USA Fondation
Long-term innate immune memory via effects on bone marrow progenitors Helen S Goodridge, PhD helen.goodridge@csmc.edu Regenerative Medicine Institute, Cedars-Sinai Medical Center, Los Angeles, USA Fondation
CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies
 CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies Jennifer Gori American Society of Gene & Cell Therapy May 11, 2017 editasmedicine.com 1 Highlights Developed
CRISPR-mediated Editing of Hematopoietic Stem Cells for the Treatment of β-hemoglobinopathies Jennifer Gori American Society of Gene & Cell Therapy May 11, 2017 editasmedicine.com 1 Highlights Developed
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Cytokines, adhesion molecules and apoptosis markers. A comprehensive product line for human and veterinary ELISAs
 Cytokines, adhesion molecules and apoptosis markers A comprehensive product line for human and veterinary ELISAs IBL International s cytokine product line... is extremely comprehensive. The assays are
Cytokines, adhesion molecules and apoptosis markers A comprehensive product line for human and veterinary ELISAs IBL International s cytokine product line... is extremely comprehensive. The assays are
What can we contribute to cancer research and treatment from Computer Science or Mathematics? How do we adapt our expertise for them
 From Bioinformatics to Health Information Technology Outline What can we contribute to cancer research and treatment from Computer Science or Mathematics? How do we adapt our expertise for them Introduction
From Bioinformatics to Health Information Technology Outline What can we contribute to cancer research and treatment from Computer Science or Mathematics? How do we adapt our expertise for them Introduction
Supplement Material. Spleen weight (mg) LN cells (X106) Acat1-/- Acat1-/- Mouse weight (g)
 Supplement Material A Spleen weight (mg) C Mouse weight (g) 1 5 1 2 9 6 3 2 5 2 1 5 Male LN cells (X16) 4 ** ** Female B 3 2 1 Supplemental Figure I. Spleen weight (A), Inguinal lymph node (LN) cell number
Supplement Material A Spleen weight (mg) C Mouse weight (g) 1 5 1 2 9 6 3 2 5 2 1 5 Male LN cells (X16) 4 ** ** Female B 3 2 1 Supplemental Figure I. Spleen weight (A), Inguinal lymph node (LN) cell number
SUPPLEMENTARY INFORMATION doi: /nature12026
 doi:1.138/nature1226 a 4 35 3 MCSF level (pg/ml) 25 2 15 1 5 1h3 3h 5h 7h 15h 24h b MPP (CD135 KSL) HSC (CD34 CD15 KSLF) c % 4 ** LPS 3 GFP pos cells 2 PU.1 GFP LPS 1 FSCA Ctl NI 24h LPS Sup.Fig.1 Effect
doi:1.138/nature1226 a 4 35 3 MCSF level (pg/ml) 25 2 15 1 5 1h3 3h 5h 7h 15h 24h b MPP (CD135 KSL) HSC (CD34 CD15 KSLF) c % 4 ** LPS 3 GFP pos cells 2 PU.1 GFP LPS 1 FSCA Ctl NI 24h LPS Sup.Fig.1 Effect
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Nature Genetics: doi: /ng.3812
 Nature Genetics: doi:10.1038/ng.3812 Supplementary Figure 1 Smarcd2-knockout mice die perinatally with impaired energy homeostasis. (a) Generation of the Smarcd2 conditional knockout allele. Deletion of
Nature Genetics: doi:10.1038/ng.3812 Supplementary Figure 1 Smarcd2-knockout mice die perinatally with impaired energy homeostasis. (a) Generation of the Smarcd2 conditional knockout allele. Deletion of
Hematopoiesis. Hematopoiesis. Hematopoiesis
 Chapter. Cells and Organs of the Immune System Hematopoiesis Hematopoiesis- formation and development of WBC and RBC bone marrow. Hematopoietic stem cell- give rise to any blood cells (constant number,
Chapter. Cells and Organs of the Immune System Hematopoiesis Hematopoiesis- formation and development of WBC and RBC bone marrow. Hematopoietic stem cell- give rise to any blood cells (constant number,
Epigenetic programming in chronic lymphocytic leukemia
 Epigenetic programming in chronic lymphocytic leukemia Christopher Oakes 10 th Canadian CLL Research Meeting September 18-19 th, 2014 Epigenetics and DNA methylation programming in normal and tumor cells:
Epigenetic programming in chronic lymphocytic leukemia Christopher Oakes 10 th Canadian CLL Research Meeting September 18-19 th, 2014 Epigenetics and DNA methylation programming in normal and tumor cells:
Inferring Biological Meaning from Cap Analysis Gene Expression Data
 Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Cluster Dendrogram. dist(cor(na.omit(tss.exprs.chip[, c(1:10, 24, 27, 30, 48:50, dist(cor(na.omit(tss.exprs.chip[, c(1:99, 103, 104, 109, 110,
 A Transcriptome (RNA-seq) Transcriptome (RNA-seq) 3. 2.5 2..5..5...5..5 2. 2.5 3. 2.5 2..5..5...5..5 2. 2.5 Cluster Dendrogram RS_ES3.2 RS_ES3. RS_SHS5.2 RS_SHS5. PS_SHS5.2 PS_SHS5. RS_LJ3 PS_LJ3..4 _SHS5.2
A Transcriptome (RNA-seq) Transcriptome (RNA-seq) 3. 2.5 2..5..5...5..5 2. 2.5 3. 2.5 2..5..5...5..5 2. 2.5 Cluster Dendrogram RS_ES3.2 RS_ES3. RS_SHS5.2 RS_SHS5. PS_SHS5.2 PS_SHS5. RS_LJ3 PS_LJ3..4 _SHS5.2
STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells
 CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
Control shrna#9 shrna#12. shrna#12 CD14-PE CD14-PE
 a Control shrna#9 shrna#12 c Control shrna#9 shrna#12 e Control shrna#9 shrna#12 h 14 12 CFU-E BFU-E GEMM GM b Colony number 7 6 5 4 3 2 1 6 pm A pa pc CFU-E BFU-E GEMM GM pu pgm A p pg B d f CD11b-APC
a Control shrna#9 shrna#12 c Control shrna#9 shrna#12 e Control shrna#9 shrna#12 h 14 12 CFU-E BFU-E GEMM GM b Colony number 7 6 5 4 3 2 1 6 pm A pa pc CFU-E BFU-E GEMM GM pu pgm A p pg B d f CD11b-APC
Expanded View Figures
 Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells
 Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
Nature Immunology: doi: /ni Supplementary Figure 1. Examples of staining for each antibody used for the mass cytometry analysis.
 Supplementary Figure 1 Examples of staining for each antibody used for the mass cytometry analysis. To illustrate the functionality of each antibody probe, representative plots illustrating the expected
Supplementary Figure 1 Examples of staining for each antibody used for the mass cytometry analysis. To illustrate the functionality of each antibody probe, representative plots illustrating the expected
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Hematopoiesis. - Process of generation of mature blood cells. - Daily turnover of blood cells (70 kg human)
 Hematopoiesis - Process of generation of mature blood cells - Daily turnover of blood cells (70 kg human) 1,000,000,000,000 total cells 200,000,000,000 red blood cells 70,000,000,000 neutrophils Hematopoiesis
Hematopoiesis - Process of generation of mature blood cells - Daily turnover of blood cells (70 kg human) 1,000,000,000,000 total cells 200,000,000,000 red blood cells 70,000,000,000 neutrophils Hematopoiesis
Hao D. H., Ma W. G., Sheng Y. L., Zhang J. B., Jin Y. F., Yang H. Q., Li Z. G., Wang S. S., GONG Ming*
 Comparison of transcriptomes and gene expression profiles of two chilling- and drought-tolerant and intolerant Nicotiana tabacum varieties under low temperature and drought stress Hao D. H., Ma W. G.,
Comparison of transcriptomes and gene expression profiles of two chilling- and drought-tolerant and intolerant Nicotiana tabacum varieties under low temperature and drought stress Hao D. H., Ma W. G.,
Supplementary Materials for
 www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
15. Supplementary Figure 9. Predicted gene module expression changes at 24hpi during HIV
 Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
Supplementary Information Table of content 1. Supplementary Table 1. Summary of RNAseq data and mapping statistics 2. Supplementary Table 2. Biological functions enriched in 12 hpi DE genes, derived from
DISCOVERING ATCC IMMUNOLOGICAL CELLS - MODEL SYSTEMS TO STUDY THE IMMUNE AND CARDIOVASCULAR SYSTEMS
 DISCOVERING ATCC IMMUNOLOGICAL CELLS - MODEL SYSTEMS TO STUDY THE IMMUNE AND CARDIOVASCULAR SYSTEMS James Clinton, Ph.D. Scientist, ATCC February 19, 2015 About ATCC Founded in 1925, ATCC is a non-profit
DISCOVERING ATCC IMMUNOLOGICAL CELLS - MODEL SYSTEMS TO STUDY THE IMMUNE AND CARDIOVASCULAR SYSTEMS James Clinton, Ph.D. Scientist, ATCC February 19, 2015 About ATCC Founded in 1925, ATCC is a non-profit
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Chapter 1. Chapter 1 Concepts. MCMP422 Immunology and Biologics Immunology is important personally and professionally!
 MCMP422 Immunology and Biologics Immunology is important personally and professionally! Learn the language - use the glossary and index RNR - Reading, Note taking, Reviewing All materials in Chapters 1-3
MCMP422 Immunology and Biologics Immunology is important personally and professionally! Learn the language - use the glossary and index RNR - Reading, Note taking, Reviewing All materials in Chapters 1-3
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well
 Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Accelerate Your Research with Conversant Bio
 Accelerate Your Research with Conversant Bio 400+ Participating MDs 50+ Partner sites for tissue procurement Continuous expansion of sourcing capabilities Closely monitored chain of custody Full regulatory
Accelerate Your Research with Conversant Bio 400+ Participating MDs 50+ Partner sites for tissue procurement Continuous expansion of sourcing capabilities Closely monitored chain of custody Full regulatory
Epigenetic and genetic alterations and their influence on gene regulation in chronic lymphocytic leukemia
 Huang and Ovcharenko BMC Genomics (2017) 18:236 DOI 10.1186/s12864-017-3617-6 RESEARCH ARTICLE Open Access Epigenetic and genetic alterations and their influence on gene regulation in chronic lymphocytic
Huang and Ovcharenko BMC Genomics (2017) 18:236 DOI 10.1186/s12864-017-3617-6 RESEARCH ARTICLE Open Access Epigenetic and genetic alterations and their influence on gene regulation in chronic lymphocytic
Supplementary Methods
 Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
