SUPPLEMENTARY INFORMATION
|
|
|
- Melissa McCarthy
- 6 years ago
- Views:
Transcription
1 SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa 3,6, Xin Zhang 3,4, Manikandan Karuppasamy 1,2, Karine Huard 1,2, Guy Schoehn 1,2,5, Imre Berger 1,2, Shu-ou Shan 3, Christiane Schaffitzel 1,2 1 European Molecular Biology Laboratory (EMBL), Grenoble Outstation, Grenoble, France 2 Unit of Virus Host-Cell Interactions, Université Joseph Fourier (UJF)-EMBL-Centre National de la Recherche Scientifique (CNRS), UMI 3265, Grenoble, France 3 Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125, United States 4 Department of Molecular and Experimental Medicine, The Scripps Research Institute, La Jolla, CA 90037, United States 5 CNRS-CEA (Commissariat à l'énergie atomique et aux énergies alternatives)-ujf - Institut de Biologie Structurale-Jean-Pierre Ebel, UMR 5075, Grenoble, France 6 These authors contributed equally to this work. Correspondence should be addressed to: S.S., sshan@caltech.edu; C.S., schaffitzel@embl.fr
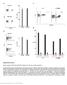
2 Supplementary Figure 1: In vitro translocation of EspP signal sequence variants and SRP binding to ribosomal complexes. (a) In vitro co-translational targeting and translocation through microsomal membranes. The EspP signal sequence variants were fused N-terminally to the signal peptidase cleavage site and to the mature region of pre-prolactin. Translocation of the preprotein leads to cleavage of the signal sequence. (b) Coomassie-stained SDS-PAGE gel showing binding of singlechain SRP (scsrp) to 70S, RNC FtsQ, and RNC EspP analyzed by ribosomal pelleting. The protein band corresponding to scsrp is highlighted by an arrow. In all experiments, except the 70S control, ribosomes were incubated with a five-fold molar excess of scsrp in the absence of nucleotides.
3 Supplementary Figure 2: Computational Sorting of the RNC EspP -SRP-FtsY data set. The dataset was sorted for compositional and conformational heterogeneity to obtain four subgroups: Particles were separated for the ratcheted (EMDB ID ) and for the not-ratcheted ribosome conformation (EMBD ID ) as well as for the presence and the absence of SRP and FtsY (scsrp) at the exit of the ribosomal tunnel. Upper row: resulting maps after computational sorting 44. Second row: Additional views of the non-ratcheted (left) and ratcheted (right) RNC EspP -SRP-FtsY complexes. The first view (1 st and 3 rd image) on the ribosomal tunnel exit shows a second connection of the SRP to the ribosome formed by the Ffh M-domain. The second view (2 nd and 4 th image) is on the exit of the ribosomal tunnel and the SRP-FtsY complex. Both SRP-FtsY containing maps have a second ribosomal connection formed by the M-domain (second row, 1 st and 3 rd image) and a non-symmetric NG-domain arrangement. The map of the not ratcheted RNC EspP -SRP-FtsY conformation corresponding to 36% of the dataset was refined to higher resolution.
4 Supplementary Figure 3: Resolution of the RNC EspP -SRP-FtsY reconstruction. The FSC function was calculated between two independent three-dimensional reconstructions of the RNC EspP -SRP-FtsY complex (continuous blue line). A set of 46,945 particles was split randomly into two equal halves to calculate the two reconstructions. FSC=0.5 indicates a resolution of 12.2 Å (dotted green line), and FSC= indicates a resolution of 8.3 Å (dotted purple line).
5 Supplementary Figure 4: Conformational rearrangements of the targeting complex assessed by placement of the the RNC FtsQ -SRP atomic model into the EM density of the false early complex. (a,b) The atomic model of the E. coli SRP bound to the translating ribosome (PDB ID: 2J28) 17 was placed into the EM density. Conformational changes of the Ffh NG-domain leading to a detachment from ribosomal protein L23 are indicated by black arrows. The detachment of the 4.5S RNA from the ribosomal protein L32 of the large ribosomal subunit is indicated by an orange arrow. The position in the EM density where the FtsY NG-domain is placed is highlighted by a purple arrow. The experimental density is depicted in light grey, 4.5S RNA in orange, the signal sequence in red, the Ffh M-domain in yellow, the Ffh NG-domain in greenyellow, the 50S rrna in dark gray, L29 in wheat and L23 in orange.
6 Supplementary Figure 5: Comparison of (a) the M-domain in the RNC EspP -SRP-FtsY false early complex and (b) the M-domain in the RNC FtsQ -SRP-FtsY early complex (EMDB ID: 1762) 20. Left: Close-up on the exit of the ribosomal tunnel and the M-domain of the RNC-SRP-FtsY complex in a surface representation. Right: Slice through the RNC-SRP-FtsY EM reconstruction. Ribosomal RNA helix h59 and the 4.5S SRP RNA are labelled for orientation. The arrow heads indicate the position of the signal-sequence binding part of the M-domain.
7 Supplementary Figure 6: Conformational heterogeneity in the RNC EspP -SRP-FtsY reconstruction. (a) Variance map of the RNC EspP -SRP-FtsY reconstruction calculated from hypergeometrically stratified resampled volumes in SPARX. After elimination of particles with low correlation coefficient, 45,200 particles were used for the hypergeometrically stratified resampling to calculate the 3D average- and variance maps 58. Left: view on the ribosomal tunnel exit site of the 50S and the SRP-FtsY complex; middle: crown view with the SRP-FtsY complex; right: view on the ribosomal tunnel exit showing the second connection of the SRP to the ribosome formed by the Ffh M- domain. Highly homogeneous regions (lower voxel variances) are coloured in blue, flexible or heterogeneous regions (higher voxel variances) are displayed in red. The surface representations were prepared in Chimera 59. (b-f) Two different conformations of the false early complex obtained by heterogeneity analysis using Xmipp. (b,c): A heterogeneity analysis using mlf_refine3d from Xmipp 57 resulted in two EM reconstructions showing distinct conformations of the SRP-FtsY complex. In both EM reconstructions, the M-domain contacts rrna helix 59. Arrows point to the connecting density between the Ffh G- and M- domains. (d,e) The EM density reveals two different Ffh/FtsY NG-domain conformations. (f) Overlay of the two maps shown in (d) and (e). In both conformations, the Ffh NGdomain is shifted towards the M-domain, and the NG-domains do not adopt a pseudo-symmetric V- shape.
8 SRP Binding K d (nm) Early Complex K d (nm) Early Closed Rearrangement k e-c (s -1 ) Closed Complex Assembly k on x 10 3 (M -1 s -1 ) RNC EspP [1] ± ± 1.1 RNC FtsQ [2] 1 80 ~ 0.3 ~ 1000 Supplementary Table 1: Comparison of kinetic and thermodynamic data of EspP as a non-srp substrate with FtsQ as a bona-fide SRP substrate. [1] Data from this study as reported in Figures 1d, 2b and 5c. [2] The data for the RNC FtsQ targeting complexes were originally reported in Zhang, X., Schaffitzel, C., Ban, N. & Shan, S.O. Multiple conformational switches in a GTPase complex control cotranslational protein targeting. Proc. Natl. Acad. Sci. USA 106, (2009).
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation
 HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Chapter 31. Completing the Protein Life Cycle: Folding, Processing and Degradation. Biochemistry by Reginald Garrett and Charles Grisham
 Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
I. Membrane Proteins II. Intracellular Compartments III. Protein Translocation
 Lecture 3 I. Membrane Proteins II. Intracellular Compartments III. Protein Translocation Ref: MBoC (5th Edition), Alberts Johnson Lewis Raff Roberts Walter Chapter 10 Membrane Structure Chapter 12 Intracellular
Lecture 3 I. Membrane Proteins II. Intracellular Compartments III. Protein Translocation Ref: MBoC (5th Edition), Alberts Johnson Lewis Raff Roberts Walter Chapter 10 Membrane Structure Chapter 12 Intracellular
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen
 Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Cryo EM structure of the ribosome SecYE complex in the
 Supplementary Information Cryo EM structure of the ribosome SecYE complex in the membrane environment Jens Frauenfeld,, James Gumbart, Eli O. van der Sluis,, Soledad Funes, Marco Gartmann,, Birgitta Beatrix,,
Supplementary Information Cryo EM structure of the ribosome SecYE complex in the membrane environment Jens Frauenfeld,, James Gumbart, Eli O. van der Sluis,, Soledad Funes, Marco Gartmann,, Birgitta Beatrix,,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1: Predicted structure of the most stable {110} antiphase boundary defect in magnetite (model APB-I). a) The same structure as that shown in Fig. 1b (main text)
Supplementary Figures Supplementary Figure 1: Predicted structure of the most stable {110} antiphase boundary defect in magnetite (model APB-I). a) The same structure as that shown in Fig. 1b (main text)
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplemental Information. Helicobacter pylori Employs a Unique Basolateral. Type IV Secretion Mechanism for CagA Delivery
 Cell Host & Microbe, Volume 22 Supplemental Information Helicobacter pylori Employs a Unique Basolateral Type IV Secretion Mechanism for CagA Delivery Nicole Tegtmeyer, Silja Wessler, Vittorio Necchi,
Cell Host & Microbe, Volume 22 Supplemental Information Helicobacter pylori Employs a Unique Basolateral Type IV Secretion Mechanism for CagA Delivery Nicole Tegtmeyer, Silja Wessler, Vittorio Necchi,
Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design
 Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design Crick's "Central Dogma"? DNA makes RNA makes protein Crick's "Central Dogma"? DNA makes
Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design Crick's "Central Dogma"? DNA makes RNA makes protein Crick's "Central Dogma"? DNA makes
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes
 Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes Siew Leong Chan Protein Analytics, AbbVie Bioresearch Center Worcester MA Acknowledgement
Electron Microscopy Imaging Reveals Unique Higher Order Structures of Adalimumab-TNFα and Infliximab-TNFα Complexes Siew Leong Chan Protein Analytics, AbbVie Bioresearch Center Worcester MA Acknowledgement
Rapid Characterization of Allosteric Networks. with Ensemble Normal Mode Analysis
 Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
TRANSLATION. Translation is a process where proteins are made by the ribosomes on the mrna strand.
 TRANSLATION Dr. Mahesha H B, Yuvaraja s College, University of Mysore, Mysuru. Translation is a process where proteins are made by the ribosomes on the mrna strand. Or The process in the ribosomes of a
TRANSLATION Dr. Mahesha H B, Yuvaraja s College, University of Mysore, Mysuru. Translation is a process where proteins are made by the ribosomes on the mrna strand. Or The process in the ribosomes of a
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Supplementary Information. Coupling ion channels to receptors for biomolecule sensing
 Supplementary Information Coupling ion channels to receptors for iomolecule sensing CHRISTOPHE J. MOREAU 1#, JULIEN P. DUPUIS 1, JEAN REVILLOUD 1, KARTHIK ARUMUGAM 2 AND MICHEL VIVAUDOU 1* 1 Institut de
Supplementary Information Coupling ion channels to receptors for iomolecule sensing CHRISTOPHE J. MOREAU 1#, JULIEN P. DUPUIS 1, JEAN REVILLOUD 1, KARTHIK ARUMUGAM 2 AND MICHEL VIVAUDOU 1* 1 Institut de
Translation Activity Guide
 Translation Activity Guide Student Handout β-globin Translation Translation occurs in the cytoplasm of the cell and is defined as the synthesis of a protein (polypeptide) using information encoded in an
Translation Activity Guide Student Handout β-globin Translation Translation occurs in the cytoplasm of the cell and is defined as the synthesis of a protein (polypeptide) using information encoded in an
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
Danish Research Institute of Translational Neuroscience DANDRITE, Nordic-EMBL Partnership
 Supplementary Information for Tuning of the Na,K-ATPase by the beta subunit Florian Hilbers 1,2,3, Wojciech Kopec 4, Toke Jost Isaksen 3,5, Thomas Hellesøe Holm 3,5, Karin Lykke- Hartmann 3,5,6, Poul Nissen
Supplementary Information for Tuning of the Na,K-ATPase by the beta subunit Florian Hilbers 1,2,3, Wojciech Kopec 4, Toke Jost Isaksen 3,5, Thomas Hellesøe Holm 3,5, Karin Lykke- Hartmann 3,5,6, Poul Nissen
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Use of double- stranded DNA mini- circles to characterize the covalent topoisomerase- DNA complex
 SUPPLEMENTARY DATA Use of double- stranded DNA mini- circles to characterize the covalent topoisomerase- DNA complex Armêl Millet 1, François Strauss 1 and Emmanuelle Delagoutte 1 1 Structure et Instabilité
SUPPLEMENTARY DATA Use of double- stranded DNA mini- circles to characterize the covalent topoisomerase- DNA complex Armêl Millet 1, François Strauss 1 and Emmanuelle Delagoutte 1 1 Structure et Instabilité
Molecular Dynamics of HIV-1 Reverse Transcriptase
 Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Reviewers' Comments: Reviewer #2: Remarks to the Author:
 Reviewers' Comments: Reviewer #1: Remarks to the Author: This manuscript presents cryo-em studies of a giant virus Paramecium bursaria chlorella virus 1 (PBCV-1), which is the first cryo-em structure of
Reviewers' Comments: Reviewer #1: Remarks to the Author: This manuscript presents cryo-em studies of a giant virus Paramecium bursaria chlorella virus 1 (PBCV-1), which is the first cryo-em structure of
Nature Neuroscience: doi: /nn Supplementary Figure 1. Confirmation that optogenetic inhibition of dopaminergic neurons affects choice
 Supplementary Figure 1 Confirmation that optogenetic inhibition of dopaminergic neurons affects choice (a) Sample behavioral trace as in Figure 1d, but with NpHR stimulation trials depicted as green blocks
Supplementary Figure 1 Confirmation that optogenetic inhibition of dopaminergic neurons affects choice (a) Sample behavioral trace as in Figure 1d, but with NpHR stimulation trials depicted as green blocks
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
TRANSPORT PROCESSES. 1b. moving proteins into membranes and organelles
 1b. moving proteins into membranes and organelles SLIDE 1 A typical mammalian cell contains up to 10,000 different kinds of proteins. The vast majority of these proteins are synthesized by cytosolic ribosomes,
1b. moving proteins into membranes and organelles SLIDE 1 A typical mammalian cell contains up to 10,000 different kinds of proteins. The vast majority of these proteins are synthesized by cytosolic ribosomes,
An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles
 REFERENCES CONTENT ALERTS An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles Jun Lin, Naiqian Cheng, Marie Chow, David J. Filman, Alasdair C. Steven,
REFERENCES CONTENT ALERTS An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles Jun Lin, Naiqian Cheng, Marie Chow, David J. Filman, Alasdair C. Steven,
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
<20 <20 <20 < <20 <20 <20 <20. Mock
 Cross-Lineage Neutralization PRNT 80 Titers Asian Asian West African Indian Ocean Group NHP Strain 181/25 Strain 99659 Strain 37997 Strain LR 142590 80 80 20 40 EILV/CHIKV 150844 640 640 160 320 Mock 150849
Cross-Lineage Neutralization PRNT 80 Titers Asian Asian West African Indian Ocean Group NHP Strain 181/25 Strain 99659 Strain 37997 Strain LR 142590 80 80 20 40 EILV/CHIKV 150844 640 640 160 320 Mock 150849
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR
 Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
Moving Proteins into Membranes and Organelles
 13 Moving Proteins into Membranes and Organelles Review the Concepts 1. In eukaryotes, protein translocation across the endoplasmic reticulum (ER) membrane is most commonly cotranslational; it can also
13 Moving Proteins into Membranes and Organelles Review the Concepts 1. In eukaryotes, protein translocation across the endoplasmic reticulum (ER) membrane is most commonly cotranslational; it can also
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting
 Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/11/e1701208/dc1 Supplementary Materials for Competitive chiral induction in a 2D molecular assembly: Intrinsic chirality versus coadsorber-induced chirality This
advances.sciencemag.org/cgi/content/full/3/11/e1701208/dc1 Supplementary Materials for Competitive chiral induction in a 2D molecular assembly: Intrinsic chirality versus coadsorber-induced chirality This
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Genetic information flows from mrna to protein through the process of translation
 Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Final Report. Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD. Wendy Fong ( 方美珍 ) CNIC; Beijing, China
 Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
High resolution structural evidence suggests the Sarcoplasmic Reticulum forms microdomains with Acidic Stores (lyososomes) in the heart.
 High resolution structural evidence suggests the Sarcoplasmic Reticulum forms microdomains with Acidic Stores (lyososomes) in the heart. Daniel Aston, Rebecca A. Capel, Kerrie L. Ford, Helen C. Christian,
High resolution structural evidence suggests the Sarcoplasmic Reticulum forms microdomains with Acidic Stores (lyososomes) in the heart. Daniel Aston, Rebecca A. Capel, Kerrie L. Ford, Helen C. Christian,
Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes
 Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
a b G75 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server.
 a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
BCH Graduate Survey of Biochemistry
 BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/9/eaao2182/dc1 Supplementary Materials for Potyvirus virion structure shows conserved protein fold and RNA binding site in ssrna viruses Miguel Zamora, Eduardo
advances.sciencemag.org/cgi/content/full/3/9/eaao2182/dc1 Supplementary Materials for Potyvirus virion structure shows conserved protein fold and RNA binding site in ssrna viruses Miguel Zamora, Eduardo
SAM Teacher s Guide Four Levels of Protein Structure
 SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
Supplementary Material
 Supplementary Material Supplementary Text Text S1. Time distributions of the high FRET efficiency at different concentrations of EF-G.GTP From Fig. 1, the model of ribosomal translocation at non-saturating
Supplementary Material Supplementary Text Text S1. Time distributions of the high FRET efficiency at different concentrations of EF-G.GTP From Fig. 1, the model of ribosomal translocation at non-saturating
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Newly synthesized proteins emerging from the peptide exit
 Trigger factor binds to ribosome signal-recognition particle (SRP) complexes and is excluded by binding of the SRP receptor Iwona Buskiewicz, Elke Deuerling, Shan-Qing Gu, Johannes Jöckel, Marina V. Rodnina,
Trigger factor binds to ribosome signal-recognition particle (SRP) complexes and is excluded by binding of the SRP receptor Iwona Buskiewicz, Elke Deuerling, Shan-Qing Gu, Johannes Jöckel, Marina V. Rodnina,
Ribosomes: Machines that Synthesize Proteins
 Ribosomes: Machines that Synthesize Proteins John Reader Department of Cell and Developmental Biology, University of North Carolina, at Chapel Hill 1 Protein Translation Enzymes that ligate amino acids
Ribosomes: Machines that Synthesize Proteins John Reader Department of Cell and Developmental Biology, University of North Carolina, at Chapel Hill 1 Protein Translation Enzymes that ligate amino acids
1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY).
 Problem Set 11 (Due Nov 25 th ) 1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY). a. Why don t trna molecules contain a 5 triphosphate like other RNA molecules
Problem Set 11 (Due Nov 25 th ) 1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY). a. Why don t trna molecules contain a 5 triphosphate like other RNA molecules
HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total. The ph at which a peptide has no net charge is its isoelectric point.
 BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
Virus Structure. Characteristics of capsids. Virus envelopes. Virion assembly John Wiley & Sons, Inc. All rights reserved.
 Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
RNA (Ribonucleic acid)
 RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
GTP-dependent ADP-ribosylation of a 22 kda protein in the endoplasmic reticulum membrane
 Volume 218, number, 63-67 FEB 04785 June 1987 GTP-dependent ADP-ribosylation of a 22 kda protein in the endoplasmic reticulum membrane Andrew Robinson and Brian Austen Department of Surgery, St. George's
Volume 218, number, 63-67 FEB 04785 June 1987 GTP-dependent ADP-ribosylation of a 22 kda protein in the endoplasmic reticulum membrane Andrew Robinson and Brian Austen Department of Surgery, St. George's
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Protein Trafficking in the Secretory and Endocytic Pathways
 Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
CHAPTER 1 Introduction to Heme Proteins
 CHAPTER 1 Introduction to Heme Proteins 1.1 Techniques Used to Study Protein Folding Pathways Proteins are synthesized by the ribosome in a stepwise fashion, emerging as long, linear polypeptides that
CHAPTER 1 Introduction to Heme Proteins 1.1 Techniques Used to Study Protein Folding Pathways Proteins are synthesized by the ribosome in a stepwise fashion, emerging as long, linear polypeptides that
Thursday, October 16 th
 Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
Triplex Structures in an RNA Pseudoknot Enhance Mechanical Stability and Increase Efficiency of 1 Ribosomal Frameshifting
 Triplex Structures in an RNA Pseudoknot Enhance Mechanical Stability and Increase Efficiency of 1 Ribosomal Frameshifting Gang CHEN Department of Molecular Biology The Scripps Research Institute 5 October,
Triplex Structures in an RNA Pseudoknot Enhance Mechanical Stability and Increase Efficiency of 1 Ribosomal Frameshifting Gang CHEN Department of Molecular Biology The Scripps Research Institute 5 October,
Supplemental Information
 Current Biology, Volume 22 Supplemental Information The Neural Correlates of Crowding-Induced Changes in Appearance Elaine J. Anderson, Steven C. Dakin, D. Samuel Schwarzkopf, Geraint Rees, and John Greenwood
Current Biology, Volume 22 Supplemental Information The Neural Correlates of Crowding-Induced Changes in Appearance Elaine J. Anderson, Steven C. Dakin, D. Samuel Schwarzkopf, Geraint Rees, and John Greenwood
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
BIOL 4374/BCHS 4313 Cell Biology Exam #2 March 22, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #2 March 22, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) In the
BIOL 4374/BCHS 4313 Cell Biology Exam #2 March 22, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) In the
Dynamic contact network between ribosomal subunits enables rapid large-scale rotation during spontaneous translocation
 Published online 24 June 2015 Nucleic Acids Research, 2015, Vol. 43, No. 14 6747 6760 doi: 10.1093/nar/gkv649 Dynamic contact network between ribosomal subunits enables rapid large-scale rotation during
Published online 24 June 2015 Nucleic Acids Research, 2015, Vol. 43, No. 14 6747 6760 doi: 10.1093/nar/gkv649 Dynamic contact network between ribosomal subunits enables rapid large-scale rotation during
Nature Methods: doi: /nmeth Supplementary Figure 1. Salipro lipid particles.
 Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
From Mosquitos to Humans: Genetic evolution of Zika Virus
 Article: From Mosquitos to Humans: Genetic evolution of Zika Virus Renata Pellegrino, PhD Director, Sequencing lab Center for Applied Genomics The Children s Hospital of Philadelphia Journal Club Clinical
Article: From Mosquitos to Humans: Genetic evolution of Zika Virus Renata Pellegrino, PhD Director, Sequencing lab Center for Applied Genomics The Children s Hospital of Philadelphia Journal Club Clinical
Nature Neuroscience: doi: /nn Supplementary Figure 1. MADM labeling of thalamic clones.
 Supplementary Figure 1 MADM labeling of thalamic clones. (a) Confocal images of an E12 Nestin-CreERT2;Ai9-tdTomato brain treated with TM at E10 and stained for BLBP (green), a radial glial progenitor-specific
Supplementary Figure 1 MADM labeling of thalamic clones. (a) Confocal images of an E12 Nestin-CreERT2;Ai9-tdTomato brain treated with TM at E10 and stained for BLBP (green), a radial glial progenitor-specific
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras
 Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
The protein stoichiometry of viral capsids via tiling theory
 The protein stoichiometry of viral capsids via tiling theory REIDUN TWAROCK Centre for Mathematical Science City University Northampton Square, London EC1V 0HB UNITED KINGDOM Abstract: - A vital part of
The protein stoichiometry of viral capsids via tiling theory REIDUN TWAROCK Centre for Mathematical Science City University Northampton Square, London EC1V 0HB UNITED KINGDOM Abstract: - A vital part of
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Summary of Endomembrane-system
 Summary of Endomembrane-system 1. Endomembrane System: The structural and functional relationship organelles including ER,Golgi complex, lysosome, endosomes, secretory vesicles. 2. Membrane-bound structures
Summary of Endomembrane-system 1. Endomembrane System: The structural and functional relationship organelles including ER,Golgi complex, lysosome, endosomes, secretory vesicles. 2. Membrane-bound structures
Supplementary Figures
 Supplementary Figures Spatial arrangement Variation in the morphology of central NCs (shape x size) x Variation in the morphology of satellite NCs (shape x size) x Variations in the spatial arrangement
Supplementary Figures Spatial arrangement Variation in the morphology of central NCs (shape x size) x Variation in the morphology of satellite NCs (shape x size) x Variations in the spatial arrangement
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/6/e1700338/dc1 Supplementary Materials for HIV virions sense plasma membrane heterogeneity for cell entry Sung-Tae Yang, Alex J. B. Kreutzberger, Volker Kiessling,
advances.sciencemag.org/cgi/content/full/3/6/e1700338/dc1 Supplementary Materials for HIV virions sense plasma membrane heterogeneity for cell entry Sung-Tae Yang, Alex J. B. Kreutzberger, Volker Kiessling,
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Aminoglycoside activity observed on single pre-translocation ribosome complexes
 correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation
 Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Dep. Control Time (min)
 aa Control Dep. RP 1s 1 mv 2s 1 mv b % potentiation of IPSP 2 15 1 5 Dep. * 1 2 3 4 Time (min) Supplementary Figure 1. Rebound potentiation of IPSPs in PCs. a, IPSPs recorded with a K + gluconate pipette
aa Control Dep. RP 1s 1 mv 2s 1 mv b % potentiation of IPSP 2 15 1 5 Dep. * 1 2 3 4 Time (min) Supplementary Figure 1. Rebound potentiation of IPSPs in PCs. a, IPSPs recorded with a K + gluconate pipette
Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex
 John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine
 Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
SUPPLEMENTARY INFORMATION. An orthogonal ribosome-trnas pair by the engineering of
 SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
Cbl ubiquitin ligase: Lord of the RINGs
 Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
Using OxRAM arrays to mimic bio-inspired short and long term synaptic plasticity Leti Memory Workshop Thilo Werner 27/06/2017
 Using OxRAM arrays to mimic bio-inspired short and long term synaptic plasticity Leti Memory Workshop Thilo Werner 27/06/2017 Collaboration by: T. Werner, E. Vianello, O. Bichler, A. Grossi, E. Nowak,
Using OxRAM arrays to mimic bio-inspired short and long term synaptic plasticity Leti Memory Workshop Thilo Werner 27/06/2017 Collaboration by: T. Werner, E. Vianello, O. Bichler, A. Grossi, E. Nowak,
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy
 7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
