SCIENCE CHINA Life Sciences
|
|
|
- Karin York
- 6 years ago
- Views:
Transcription
1 SCIENCE CHINA Life Sciences RESEARCH PAPER April 2013 Vol.56 No.4: 1 7 doi: /s x Table S1 Human paired-end RNA-Seq data sets from the Sequence Read Archive (SRA) database Experiment accession SRX SRX SRX SRX SRX SRX SRX SRX SRX Cell lines H1-hESC H1-hESC H1-hESC HepG2 HepG2 HUVEC NHEK HeLa-S3 GM12878 Table S2 Gene expression data sets from GEO database used to construct two-color co-expression network No. GEO accession Discription 1 GSE9988(62) Innate immune repsonses to TREM-1 activation 2 GSE11882 (173) Gene expression changes in the course of normal brain aging are sexually dimorphicgene expression changes in the course of normal brain aging are sexually dimorphic 3 GSE9196 (53) Comparison of ES, EB, and Blast cells to breast epithelial, leuckocytes, endothelial and stromal cells 4 GSE15431 (31) Global Gene Expression in the Human Fetal Testis and Ovary 5 GSE7214 (27) Comparison of gene expression data between wild-type and DM1-affected cells 6 GSE8823 (24) Overexpression of the Apoptotic Cell Removal Receptor, MERTK, in Alveolar Macrophages of Cigarette Smokers 7 GSE5372 (22) airway epithelium, large airways, pre and post-mechanical injury 8 GSE9254 (19) Normal human colorectal mucosa, cecum, ascending, transverse, sigmoid and rectum 9 GSE19599 (16) Expression data for normal flow sorted hematopietic cell subpopulations 10 GSE22167 (18) Reprogramming of T Cells from Human Peripheral Blood 11 GSE18265 (13) Transcriptomic analysis of pluripotent stem cells 12 GSE9709 (10) Human induced pluripotent stem (ips) cells from neonatal skin derived cells 13 GSE3526 (353) Comparison of gene expression profiles across the normal human body 14 GSE4888 (27) Molecular phenotyping of human endometrium 15 GSE15238 (10) Expression data from human embryonic (9-12w) and post-natal livers 16 GSE17251(24) Comparative analysis of gene regulation by the transcription factor PPARα_human 17 GSE14334 (29) Transcriptomic analysis of human lung development 18 GSE7821(40) Expression data from human intestinal biopsies 19 GSE11103(41) Study of human immune and memory T cells using microarray 20 GSE23968(14) Large intergenic non-coding RNAs as novel modulators of reprogramming: ESCs, fibroblast, and fibroblast-derived ipsc (gene expression) 21 GSE18897(80) Whole blood expression profiling of obese diet-sensitive, obese diet-resistant and lean human subjects 22 GSE9419(66) The skeletal muscle transcript profile reflects responses to inadequate protein intake in younger and older males 23 GSE9865(13) Expression profile of dermal fibroblasts reprogrammed to a pluripotent state 24 GSE7888(23) Expression data from human mesenchymal stem cells (six batches) 25 GSE13564(44) Gene expression in the human prefrontal cortex during postnatal development 26 GSE2125(45) isolated alveolar macrophages 27 GSE10041(72) Genomic Counter-Stress Changes Induced by Mind-Body Practice 28 GSE18791(57) Antiviral response dictated by choreographed cascade of transcription factors 29 GSE8658(63) PPARg regulated gene expression in human dendritic cells 30 GSE18637(20) Do Airway Epithelium Air-liquid Cultures Represent the In Vivo Airway Epithelium Transcriptome? 31 GSE16028(109) Longitudinal study of gene expression in healthy individuals The Author(s) This article is published with open access at Springerlink.com life.scichina.com
2 2 Sun L, et al. Sci China Life Sci April (2013) Vol.56 No.4 Table S3 Properties of networks using different number of datasets a) Number of connections (cc/nn/cn) (c/n) of degree coefficient topology criterion GO pairs (random) Number of genes Average number Network clustering Scale-free Proportion of same Number of dataset Gamma % (4%) % (5%) % (5%) % (8%) % (11%) % (14%) % (20%) a) c: coding genes; n: lincrnas; cc: coding-coding gene connections; nn: lincrnas-lincrnas connections; cn: coding gene-lincrnas connections. Table S4 Genetic mediators of protein coding genes Gene ID Gene symbol Validation CLEC7A no 4688 NCF2 no 7409 VAV1 yes 646 BNC1 no 6657 SOX2 yes 4815 NINJ2 no RASSF5 yes 4048 LTA4H yes 5321 PLA2G4A no 4922 NTS yes 1294 COL7A1 yes 5319 PLA2G1B yes HHIP no GKN2 no CIT yes 24 ABCA4 no P2RY10 yes 2304 FOXE1 no 722 C4BPA yes ZFP64 no ATP11A yes 3815 KIT yes RTKN2 no 1592 CYP26A1 no 2676 GFRA3 no 6098 ROS1 no RAB6B no 5744 PTHLH yes 190 NR0B1 yes 1399 CRKL no OPN3 no CABYR yes SUSD2 no 5045 FURIN no (To be continued on the next page)
3 (Continued) Gene ID Gene symbol Validation 961 CD47 yes ARNTL2 no LRRK2 yes PANX2 no BANK1 no CCL26 no 3641 INSL4 no TLR10 no SCGB3A1 yes CLDN18 yes 13 AADAC no 2719 GPC3 yes GPR87 no 8745 ADAM23 no 7090 TLE3 no RHOBTB2 yes GTSF1 no 5361 PLXNA1 no C1orf110 no 8999 CDKL2 no RGS17 yes 8547 FCN3 yes 5081 PAX7 yes OSGIN1 no 5275 SERPINB13 no 2196 FAT2 yes 2769 GNA15 no CHIA no HEY1 yes SOST no 6016 RIT1 no COX7B2 no FCRL2 no GPR110 no 9856 KIAA0319 no 7903 ST8SIA4 no 2702 GJA5 no WIF1 yes 9048 ARTN yes CAPN13 no ATP8A2 no 4973 OLR1 yes 5803 PTPRZ1 yes CDK5RAP2 yes 5590 PRKCZ no PIM2 yes 9182 RASSF9 no 3851 KRT4 yes 3853 KRT6A yes 6357 CCL13 no TREM1 no PITPNC1 no 7356 SCGB1A1 yes 6335 SCN9A no PCGF1 no 2729 GCLC yes 8626 TP63 yes PLUNC yes 6402 SELL yes 3694 ITGB6 no 1591 CYP24A1 yes IL20RB yes PHACTR3 yes Sun L, et al. Sci China Life Sci April (2013) Vol.56 No.4 3
4 4 Sun L, et al. Sci China Life Sci April (2013) Vol.56 No.4 Table S5 LincRNA-sets dysregulated in lung cancer. Star denotes the sets detected in both experiments Name of lincrnas state Functional annotation Est_480 Down Regulation of transcription(bp) and RNA metabolic process Est_884 Down chromatin modification Est_1088 Down Metabolism process(kegg) Est_1650 Down Metabolism process Est_669 Down MAPK signaling pathway(ke) Est_997 Down Regulation of metabolism Scrip_2185 Down* regulation of immune system process and leukocyte activation Both_202 Down embryonic cleavage Scrip_2379 Up* Regulation of cell cycle and cell death Est_835 Up* response to stimulus and regulation of cell cycle Est_810 Up* metabolic process, system development and embryonic development Scrip_3488 Up positive regulation of transcription and Wnt signaling pathway Figure S1 Computational pipelines for identification of human lincrnas.
5 Sun L, et al. Sci China Life Sci April (2013) Vol.56 No.4 5 Figure S2 Expression profiles of protein coding and lincrna genes. A, Expressional correlations of protein coding genes. Red line: distribution of pearson correlation coefficients for the expression rank of identical protein coding genes across six cell lines in RNA-Seq data and in the re-annotated Human Genome U133 Plus 2.0 Array. Light grey lines: distribution of pearson correlation coefficients for the expression rank of randomly selected protein coding genes pairs, repeated 1000 times. (Mean pearson correlation coefficient was 0.51, P-value of the KS test was less then 2.2e 10 ). B, Expressional correlations of lincrnas. Red line: distribution of pearson correlation coefficients for the expression rank of identical lincrnas across six cell lines in RNA-Seq data and in the re-annotated Human Genome U133 Plus 2.0 Array. Light grey lines: distribution of pearson correlation coefficients for the expression rank of randomly selected lincrna pairs, repeated 1000 times. (Mean pearson correlation coefficient was 0.41, P-value of the KS test was less than 2.2e 10 ). C, Expression profiles of lincrnas Both_148, Scrip_3188, Est_1593 and Both_2 in the re-annotated Human Genome U133 Plus 2.0 Array and RNA-Seq data.
6 6 Sun L, et al. Sci China Life Sci April (2013) Vol.56 No.4 Figure S3 Genes differently expressed response to CDK inhibitor treatment. A, Diagram showing number of protein coding genes differently expressed response to CDK inhibitor treatment in three cell lines. B, Diagram showing number of long ncrna genes differently expressed response to CDK inhibitor treatment in three cell lines. C, GO enrichment analysis results of 1254 coding genes which differently expressed response to CDK inhibitor treatment. Figure S4 Genomic contexts of three lincrna genes. LincRNAs as genetic mediators include Scrip_2341 (Up), Scrip_2379 (Middle) and Scrip_2616 (Down). Other information include H3K4me3 marker, RefSeq gene and conservation status.
7 Sun L, et al. Sci China Life Sci April (2013) Vol.56 No.4 7 Figure S5 The relationship between coding gene-centered gene sets and cancer. A, GO enrichment analysis result of central coding genes of 167 gene sets which significantly induced (or repressed) in lung cancer relative to normal lung. B, The left figure shows the genes in KIF11 related gene set induced (red) or repressed (green) in each array. The bottom bars denote the experiment sets which array belongs to. Cancer experiment sets were marked by yellow color. The right figure shows KIF11 related gene set. Genes colored in green are annotated as cell cycle genes by KEGG and colored in pink are annotated as cell cycle genes by GO. Yellow color denotes other functions. Edges colored in light grey (or green) are negative (or positive) correlations. Figure S6 An example of relationship between lincrna-set and cancer. A, Scrip_2185 related gene set. Genes colored in pink are immune related genes. Yellow color denotes other functions. Edges colored in light grey (or green) are negative (or positive) correlations. B, Genomic contexts of Scrip_2185 gene locus and UCSC genes.
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Chromatin marks identify critical cell-types for fine-mapping complex trait variants
 Chromatin marks identify critical cell-types for fine-mapping complex trait variants Gosia Trynka 1-4 *, Cynthia Sandor 1-4 *, Buhm Han 1-4, Han Xu 5, Barbara E Stranger 1,4#, X Shirley Liu 5, and Soumya
Chromatin marks identify critical cell-types for fine-mapping complex trait variants Gosia Trynka 1-4 *, Cynthia Sandor 1-4 *, Buhm Han 1-4, Han Xu 5, Barbara E Stranger 1,4#, X Shirley Liu 5, and Soumya
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Cluster Dendrogram. dist(cor(na.omit(tss.exprs.chip[, c(1:10, 24, 27, 30, 48:50, dist(cor(na.omit(tss.exprs.chip[, c(1:99, 103, 104, 109, 110,
 A Transcriptome (RNA-seq) Transcriptome (RNA-seq) 3. 2.5 2..5..5...5..5 2. 2.5 3. 2.5 2..5..5...5..5 2. 2.5 Cluster Dendrogram RS_ES3.2 RS_ES3. RS_SHS5.2 RS_SHS5. PS_SHS5.2 PS_SHS5. RS_LJ3 PS_LJ3..4 _SHS5.2
A Transcriptome (RNA-seq) Transcriptome (RNA-seq) 3. 2.5 2..5..5...5..5 2. 2.5 3. 2.5 2..5..5...5..5 2. 2.5 Cluster Dendrogram RS_ES3.2 RS_ES3. RS_SHS5.2 RS_SHS5. PS_SHS5.2 PS_SHS5. RS_LJ3 PS_LJ3..4 _SHS5.2
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
MIR retrotransposon sequences provide insulators to the human genome
 Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Profiling of the Exosomal Cargo of Bovine Milk Reveals the Presence of Immune- and Growthmodulatory Non-coding RNAs (ncrna)
 Animal Industry Report AS 664 ASL R3235 2018 Profiling of the Exosomal Cargo of Bovine Milk Reveals the Presence of Immune- and Growthmodulatory Non-coding RNAs (ncrna) Eric D. Testroet Washington State
Animal Industry Report AS 664 ASL R3235 2018 Profiling of the Exosomal Cargo of Bovine Milk Reveals the Presence of Immune- and Growthmodulatory Non-coding RNAs (ncrna) Eric D. Testroet Washington State
Supplemental Information. Aryl Hydrocarbon Receptor Controls. Monocyte Differentiation. into Dendritic Cells versus Macrophages
 Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
ChromHMM Tutorial. Jason Ernst Assistant Professor University of California, Los Angeles
 ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells
 Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
Single-cell RNA-Seq profiling of human pre-implantation embryos and embryonic stem cells Liying Yan,2,5, Mingyu Yang,5, Hongshan Guo, Lu Yang, Jun Wu, Rong Li,2, Ping Liu, Ying Lian, Xiaoying Zheng, Jie
Supplementary Data for:
 Supplementary Data for: Tumour sampling method can significantly influence gene expression profiles derived from neoadjuvant window studies Dominic A. Pearce 1, Laura M. Arthur 1, Arran K. Turnbull 1,
Supplementary Data for: Tumour sampling method can significantly influence gene expression profiles derived from neoadjuvant window studies Dominic A. Pearce 1, Laura M. Arthur 1, Arran K. Turnbull 1,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Supplementary figures legends
 Supplementary Information to Nijmegen Breakage Syndrome fibroblasts and ipscs: cellular models for uncovering diseaseassociated signaling pathways and establishing a screening platform for anti-oxidants
Supplementary Information to Nijmegen Breakage Syndrome fibroblasts and ipscs: cellular models for uncovering diseaseassociated signaling pathways and establishing a screening platform for anti-oxidants
STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells
 CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
cis-regulatory enrichment analysis in human, mouse and fly
 cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Lung Met 1 Lung Met 2 Lung Met Lung Met H3K4me1. Lung Met H3K27ac Primary H3K4me1
 a Gained Met-VELs 1.5 1.5 -.5 Lung Met 1 Lung Met Lung Met 3 1. Lung Met H3K4me1 Lung Met H3K4me1 1 Lung Met H3K4me1 Lung Met H3K7ac 1.5 Lung Met H3K7ac Lung Met H3K7ac.8 Primary H3K4me1 Primary H3K7ac
a Gained Met-VELs 1.5 1.5 -.5 Lung Met 1 Lung Met Lung Met 3 1. Lung Met H3K4me1 Lung Met H3K4me1 1 Lung Met H3K4me1 Lung Met H3K7ac 1.5 Lung Met H3K7ac Lung Met H3K7ac.8 Primary H3K4me1 Primary H3K7ac
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
Supporting Information
 Supporting Information Retinal expression of small non-coding RNAs in a murine model of proliferative retinopathy Chi-Hsiu Liu 1, Zhongxiao Wang 1, Ye Sun 1, John Paul SanGiovanni 2, Jing Chen 1, * 1 Department
Supporting Information Retinal expression of small non-coding RNAs in a murine model of proliferative retinopathy Chi-Hsiu Liu 1, Zhongxiao Wang 1, Ye Sun 1, John Paul SanGiovanni 2, Jing Chen 1, * 1 Department
Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles
 Author's response to reviews Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles Authors: Julia Tchou (julia.tchou@uphs.upenn.edu) Andrew V Kossenkov (akossenkov@wistar.org)
Author's response to reviews Title: Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles Authors: Julia Tchou (julia.tchou@uphs.upenn.edu) Andrew V Kossenkov (akossenkov@wistar.org)
Sudin Bhattacharya Institute for Integrative Toxicology
 Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells
 Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells Ondrej Podlaha 1, Subhajyoti De 2,3,4, Mithat Gonen 5, Franziska Michor 1 * 1 Department of Biostatistics
Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells Ondrej Podlaha 1, Subhajyoti De 2,3,4, Mithat Gonen 5, Franziska Michor 1 * 1 Department of Biostatistics
Supplementary Methods
 Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
Hao D. H., Ma W. G., Sheng Y. L., Zhang J. B., Jin Y. F., Yang H. Q., Li Z. G., Wang S. S., GONG Ming*
 Comparison of transcriptomes and gene expression profiles of two chilling- and drought-tolerant and intolerant Nicotiana tabacum varieties under low temperature and drought stress Hao D. H., Ma W. G.,
Comparison of transcriptomes and gene expression profiles of two chilling- and drought-tolerant and intolerant Nicotiana tabacum varieties under low temperature and drought stress Hao D. H., Ma W. G.,
Identification of candidate gonadal sex differentiation genes in the chicken embryo using RNA-seq
 Ayers et al. BMC Genomics (2015)16:704 DOI 10.1186/s12864-015-1886-5 RESEARCH ARTICLE Identification of candidate gonadal sex differentiation genes in the chicken embryo using RNA-seq Open Access Katie
Ayers et al. BMC Genomics (2015)16:704 DOI 10.1186/s12864-015-1886-5 RESEARCH ARTICLE Identification of candidate gonadal sex differentiation genes in the chicken embryo using RNA-seq Open Access Katie
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Paternal exposure and effects on microrna and mrna expression in developing embryo. Department of Chemical and Radiation Nur Duale
 Paternal exposure and effects on microrna and mrna expression in developing embryo Department of Chemical and Radiation Nur Duale Our research question Can paternal preconceptional exposure to environmental
Paternal exposure and effects on microrna and mrna expression in developing embryo Department of Chemical and Radiation Nur Duale Our research question Can paternal preconceptional exposure to environmental
4. Th1-related gene expression in infected versus mock-infected controls from Fig. 2 with gene annotation.
 List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and
 Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
TEB. Id4 p63 DAPI Merge. Id4 CK8 DAPI Merge
 a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
Transduction of lentivirus to human primary CD4+ T cells
 Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Transduction of lentivirus to human primary CD4 + T cells Human primary CD4 T cells were stimulated with anti-cd3/cd28 antibodies (10 µl/2 5 10^6 cells of Dynabeads CD3/CD28 T cell expander, Invitrogen)
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Gonadal Developmental Systems Biology
 Spring 2018 Systems Biology of Reproduction Discussion Outline Gonadal Developmental Systems Biology Michael K. Skinner Biol 475/575 CUE 418, 10:35-11:50 am, Tuesday & Thursday February 15, 2018 Week 6
Spring 2018 Systems Biology of Reproduction Discussion Outline Gonadal Developmental Systems Biology Michael K. Skinner Biol 475/575 CUE 418, 10:35-11:50 am, Tuesday & Thursday February 15, 2018 Week 6
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
TRANSLATIONAL AND CLINICAL RESEARCH
 TRANSLATIONAL AND CLINICAL RESEARCH Airway Basal Cells of Healthy Smokers Express an Embryonic Stem Cell Signature Relevant to Lung Cancer RENAT SHAYKHIEV, a RUI WANG, a RACHEL K. ZWICK, a NEIL R. HACKETT,
TRANSLATIONAL AND CLINICAL RESEARCH Airway Basal Cells of Healthy Smokers Express an Embryonic Stem Cell Signature Relevant to Lung Cancer RENAT SHAYKHIEV, a RUI WANG, a RACHEL K. ZWICK, a NEIL R. HACKETT,
The corrected Figure S1J is shown below. The text changes are as follows, with additions in bold and deletions in bracketed italics:
 Correction H3K4me3 readth Is Linked to Cell Identity and Transcriptional Consistency érénice A. enayoun, Elizabeth A. Pollina, Duygu Ucar, Salah Mahmoudi, Kalpana Karra, Edith D. Wong, Keerthana Devarajan,
Correction H3K4me3 readth Is Linked to Cell Identity and Transcriptional Consistency érénice A. enayoun, Elizabeth A. Pollina, Duygu Ucar, Salah Mahmoudi, Kalpana Karra, Edith D. Wong, Keerthana Devarajan,
Obstacles and challenges in the analysis of microrna sequencing data
 Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Tissue of origin determines cancer-associated CpG island promoter hypermethylation patterns
 RESEARCH Open Access Tissue of origin determines cancer-associated CpG island promoter hypermethylation patterns Duncan Sproul 1,2, Robert R Kitchen 1,3, Colm E Nestor 1,2, J Michael Dixon 1, Andrew H
RESEARCH Open Access Tissue of origin determines cancer-associated CpG island promoter hypermethylation patterns Duncan Sproul 1,2, Robert R Kitchen 1,3, Colm E Nestor 1,2, J Michael Dixon 1, Andrew H
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier
 Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
RNA sequencing of cancer reveals novel splicing alterations
 RNA sequencing of cancer reveals novel splicing alterations Jeyanthy Eswaran, Anelia Horvath, Sucheta Godbole, Sirigiri Divijendra Reddy, Prakriti Mudvari, Kazufumi Ohshiro, Dinesh Cyanam, Sujit Nair,
RNA sequencing of cancer reveals novel splicing alterations Jeyanthy Eswaran, Anelia Horvath, Sucheta Godbole, Sirigiri Divijendra Reddy, Prakriti Mudvari, Kazufumi Ohshiro, Dinesh Cyanam, Sujit Nair,
The Avatar System TM Yields Biologically Relevant Results
 Application Note The Avatar System TM Yields Biologically Relevant Results Liquid biopsies stand to revolutionize the cancer field, enabling early detection and noninvasive monitoring of tumors. In the
Application Note The Avatar System TM Yields Biologically Relevant Results Liquid biopsies stand to revolutionize the cancer field, enabling early detection and noninvasive monitoring of tumors. In the
R2 Training Courses. Release The R2 support team
 R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC)
 Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Molecular Genetics of Paediatric Tumours. Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA
 Molecular Genetics of Paediatric Tumours Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA Financial Disclosure NanoString - conference costs for
Molecular Genetics of Paediatric Tumours Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA Financial Disclosure NanoString - conference costs for
Reduced expression of SOX7 in ovarian cancer: a novel tumor suppressor through the Wnt/β-catenin signaling pathway
 Reduced expression of SOX7 in ovarian cancer: a novel tumor suppressor through the Wnt/β-catenin signaling pathway Huidi Liu, M.D. & Ph.D Genomics Research Centre Harbin Medical University, China 15804606535@163.com
Reduced expression of SOX7 in ovarian cancer: a novel tumor suppressor through the Wnt/β-catenin signaling pathway Huidi Liu, M.D. & Ph.D Genomics Research Centre Harbin Medical University, China 15804606535@163.com
Network-assisted data analysis
 Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Regulation of cell signaling cascades by influenza A virus
 The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
HepaRG LX2. HepaRG HepaRG LX2 LX2
 C Supporting Figure 1. Experimental design of s between and cells. (A) -hepatocytes were isolated from a 30 days of -progenitors. Differentiation into mature hepatocytes was achieved following a 2-weeks
C Supporting Figure 1. Experimental design of s between and cells. (A) -hepatocytes were isolated from a 30 days of -progenitors. Differentiation into mature hepatocytes was achieved following a 2-weeks
Simple, rapid, and reliable RNA sequencing
 Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Simple, rapid, and reliable RNA sequencing RNA sequencing applications RNA sequencing provides fundamental insights into how genomes are organized and regulated, giving us valuable information about the
Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos 3, Hui Yang 1, and Guillermo Garcia-Manero 1
 Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Supplementary Material
 Supplementary Material Fig. S1. Gpa33 -/- mice do not express Gpa33 mrna or GPA33 protein. (A) The Gpa33 null allele contains a premature translational stop codon (STOP) in exon 2, and almost all of the
Supplementary Material Fig. S1. Gpa33 -/- mice do not express Gpa33 mrna or GPA33 protein. (A) The Gpa33 null allele contains a premature translational stop codon (STOP) in exon 2, and almost all of the
Figure S1, Beyer et al.
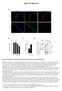 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Systematic exploration of autonomous modules in noisy microrna-target networks for testing the generality of the cerna hypothesis
 RESEARCH Systematic exploration of autonomous modules in noisy microrna-target networks for testing the generality of the cerna hypothesis Danny Kit-Sang Yip 1, Iris K. Pang 2 and Kevin Y. Yip 1,3,4* *
RESEARCH Systematic exploration of autonomous modules in noisy microrna-target networks for testing the generality of the cerna hypothesis Danny Kit-Sang Yip 1, Iris K. Pang 2 and Kevin Y. Yip 1,3,4* *
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
Large conserved domains of low DNA methylation maintained by Dnmt3a
 Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Nature Immunology: doi: /ni.3412
 Supplementary Figure 1 Gata1 expression in heamatopoietic stem and progenitor populations. (a) Unsupervised clustering according to 100 top variable genes across single pre-gm cells. The two main cell
Supplementary Figure 1 Gata1 expression in heamatopoietic stem and progenitor populations. (a) Unsupervised clustering according to 100 top variable genes across single pre-gm cells. The two main cell
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
A gene expression bar code for microarray data
 A gene expression bar code for microarray data Michael J Zilliox & Rafael A Irizarry Supplementary figures and text: Supplementary Figure 1: Bar code heatmaps, dendrograms, and summaries. Supplementary
A gene expression bar code for microarray data Michael J Zilliox & Rafael A Irizarry Supplementary figures and text: Supplementary Figure 1: Bar code heatmaps, dendrograms, and summaries. Supplementary
levels of genes were separated by their expression levels; 2,000 high, medium, and low
 Figure S1. Histone modification profiles near transcription start sites. The overall histone modification around transcription start sites (TSSs) was calculated. Histone modification levels of genes were
Figure S1. Histone modification profiles near transcription start sites. The overall histone modification around transcription start sites (TSSs) was calculated. Histone modification levels of genes were
Fibroblasts cell lines misclassified as cancer cell lines
 Fibroblasts cell lines misclassified as cancer cell lines Antoine de Weck 1, Hans Bitter 2, Audrey Kauffmann 1 1. Novartis Institutes for Biomedical Research, Basel CH-4002, Switzerland. 2. Novartis Institutes
Fibroblasts cell lines misclassified as cancer cell lines Antoine de Weck 1, Hans Bitter 2, Audrey Kauffmann 1 1. Novartis Institutes for Biomedical Research, Basel CH-4002, Switzerland. 2. Novartis Institutes
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells
 Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells Liying Yan 1,2,5, Mingyu Yang 1,5, Hongshan Guo 1, Lu Yang 1, Jun Wu 1, Rong Li 1,2, Ping Liu 1, Ying Lian 1, Xiaoying
Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells Liying Yan 1,2,5, Mingyu Yang 1,5, Hongshan Guo 1, Lu Yang 1, Jun Wu 1, Rong Li 1,2, Ping Liu 1, Ying Lian 1, Xiaoying
PCxN: The pathway co-activity map: a resource for the unification of functional biology
 PCxN: The pathway co-activity map: a resource for the unification of functional biology Sheffield Institute for Translational Neurosciences Center for Integrative Genome Translation GENOME INFORMATICS
PCxN: The pathway co-activity map: a resource for the unification of functional biology Sheffield Institute for Translational Neurosciences Center for Integrative Genome Translation GENOME INFORMATICS
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Reprogramming through micrornas Stefanie Dimmeler
 Klinikum der Johann Wolfgang Goethe Universität Frankfurt am Main Reprogramming through micrornas Stefanie Dimmeler Conflict of interest: T2cure GmbH, Miragen Non-coding DNA & RNA and micrornas Human Genome
Klinikum der Johann Wolfgang Goethe Universität Frankfurt am Main Reprogramming through micrornas Stefanie Dimmeler Conflict of interest: T2cure GmbH, Miragen Non-coding DNA & RNA and micrornas Human Genome
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
doi:10.1038/nature15260 Supplementary Data 1: Gene expression in individual basal/stem, luminal, and luminal progenitor cells. Box plots show expression levels for each gene from the 49-gene differentiation
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Gene Ontology 2 Function/Pathway Enrichment. Biol4559 Thurs, April 12, 2018 Bill Pearson Pinn 6-057
 Gene Ontology 2 Function/Pathway Enrichment Biol4559 Thurs, April 12, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Function/Pathway enrichment analysis do sets (subsets) of differentially expressed
Gene Ontology 2 Function/Pathway Enrichment Biol4559 Thurs, April 12, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Function/Pathway enrichment analysis do sets (subsets) of differentially expressed
Deconvolution of Heterogeneous Wound Tissue Samples into Relative Macrophage Phenotype Composition via Models based on Gene Expression
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2017 Supplementary Information for: Deconvolution of Heterogeneous Wound Tissue Samples into
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2017 Supplementary Information for: Deconvolution of Heterogeneous Wound Tissue Samples into
* Kyoto Encyclopedia of Genes and Genomes.
 Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
Supplemental Material Complete gene expression data using Affymetrix 3PRIME IVT ID Chip (54,614 genes) and human immature dendritic cells stimulated with rbmasnrs, IL-8 and control (media) has been deposited
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
SCIENCE CHINA Life Sciences
 SCIENCE CHINA Life Sciences RESEARCH PAPER June 2013 Vol.56 No.6: 503 512 doi: 10.1007/s11427-013-4485-1 Genome-wide identification of cancer-related polyadenylated and non-polyadenylated RNAs in human
SCIENCE CHINA Life Sciences RESEARCH PAPER June 2013 Vol.56 No.6: 503 512 doi: 10.1007/s11427-013-4485-1 Genome-wide identification of cancer-related polyadenylated and non-polyadenylated RNAs in human
Gene-microRNA network module analysis for ovarian cancer
 Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Gene-microRNA network module analysis for ovarian cancer Shuqin Zhang School of Mathematical Sciences Fudan University Oct. 4, 2016 Outline Introduction Materials and Methods Results Conclusions Introduction
Supplemental Information. Genomic Characterization of Murine. Monocytes Reveals C/EBPb Transcription. Factor Dependence of Ly6C Cells
 Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
Clasificación Molecular del Cáncer de Próstata. JM Piulats
 Clasificación Molecular del Cáncer de Próstata JM Piulats Introduction The Gleason score is the major method for prostate cancer tissue grading and the most important prognostic factor in this disease.
Clasificación Molecular del Cáncer de Próstata JM Piulats Introduction The Gleason score is the major method for prostate cancer tissue grading and the most important prognostic factor in this disease.
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells
 Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Cellular dissection of psoriasis for transcriptome analyses and the post-gwas era
 Swindell et al. BMC Medical Genomics 2014, 7:27 RESEARCH ARTICLE Open Access Cellular dissection of psoriasis for transcriptome analyses and the post-gwas era William R Swindell *, Philip E Stuart, Mrinal
Swindell et al. BMC Medical Genomics 2014, 7:27 RESEARCH ARTICLE Open Access Cellular dissection of psoriasis for transcriptome analyses and the post-gwas era William R Swindell *, Philip E Stuart, Mrinal
Supplementary information
 Supplementary information High fat diet-induced changes of mouse hepatic transcription and enhancer activity can be reversed by subsequent weight loss Majken Siersbæk, Lyuba Varticovski, Shutong Yang,
Supplementary information High fat diet-induced changes of mouse hepatic transcription and enhancer activity can be reversed by subsequent weight loss Majken Siersbæk, Lyuba Varticovski, Shutong Yang,
Broad Integration of Expression Maps and Co-Expression Networks Compassing Novel Gene Functions in the Brain
 Supplementary Information Broad Integration of Expression Maps and Co-Expression Networks Compassing Novel Gene Functions in the Brain Yuko Okamura-Oho a, b, *, Kazuro Shimokawa c, Masaomi Nishimura b,
Supplementary Information Broad Integration of Expression Maps and Co-Expression Networks Compassing Novel Gene Functions in the Brain Yuko Okamura-Oho a, b, *, Kazuro Shimokawa c, Masaomi Nishimura b,
Supplemental Information. A Highly Sensitive and Robust Method. for Genome-wide 5hmC Profiling. of Rare Cell Populations
 Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
