Journal: Nature Methods
|
|
|
- Hannah Phillips
- 6 years ago
- Views:
Transcription
1 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3 Supplementary Figure 4 Supplementary Figure 5 Supplementary Figure 6 Supplementary Figure 7 Supplementary Figure 8 Supplementary Figure 9 Supplementary Figure 10 Supplementary Figure 11 Supplementary Figure 12 Supplementary Figure 13 Supplementary Figure 14 Supplementary Figure 15 Supplementary Figure 16 Title or Caption An overview of the somatic mutation landscape of TCGA ovarian cancer cohort. Simulating across different networks Ovarian cancer association with overall survival Lung cancer association with overall survival Uterine cancer association with histological type Standard predictors of survival are independent of ovarian subtype A network view of genes with high network smoothed mutation scores in ovarian cancer, HumanNet, subtype 2 relative to other subtypes A network view of genes with high network smoothed mutation scores in ovarian cancer, HumanNet, subtype 3 relative to other subtypes A network view of genes with high smoothed mutation scores in ovarian cancer, HumanNet, subtype 4 relative to other subtypes. A network view of genes with high smoothed mutation scores in uterine cancer, STRING, subtype 1 relative to other subtypes. A network view of genes with high smoothed mutation scores in uterine cancer, STRING, subtype 2 relative to other subtypes. A network view of genes with high smoothed mutation scores in uterine cancer, STRING, subtype 3 relative to other subtypes. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 1 relative to other subtypes. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 2 relative to other subtypes. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 3 relative to other subtypes. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 5 relative to other subtypes. 1
2 Supplementary Figure 17 Supplementary Figure 18 Supplementary Figure 19 Supplementary Table 1 From mutation-derived subtypes to expression signatures Standard consensus clustering NMF used to recover subtypes in the Tothill et al. expression cohort of ovarian tumors Effects of progressively permuting proportions of the lung cancer dataset Summary of gene interaction networks Source data figure Figure 5, Supp Fig 7-9 Supp Fig Supp Fig File name OV_HM90_K4_SAM_diff.xlsx UCEC_ST90_K3_SAM_diff.xlsx LUAD_ST90_K3_SAM_diff.xlsx Supplementary Figure 1 Supplementary Figure 1. An overview of the somatic mutation landscape of TCGA ovarian cancer cohort. (a) Somatic mutations along the length of chromosome 17. (b) A histogram summing the frequency of mutations per gene for the entire exome. (c) A histogram summing the frequency of genes mutated per patient in the cohort. 2
3 Supplementary Figure 2 Supplementary Figure 2. Simulating across different networks. In this simulation network modules from the NCI- Nature cancer pathways network were used for the simulation and were recovered by NBS using the HumanNet network. Each subtype included between 2-6 driver modules totaling the specified size of genes and the driver gene frequency. Driver frequencies of 10%, 7.5%, 5% and driver modules comprising , 60-80, were used in panels (a),(b) and (c) respectively. Furthermore, a subset (0-4) of the modules was assigned to overlap across multiple subtypes. 3
4 Supplementary Figure 3 Supplementary Figure 3. Ovarian cancer association with overall survival. (a) Co-clustering matrices for ovarian cancer patients, comparing NBS (HumanNet) to standard consensus clustering. (b-c) Cox proportional hazards model logrank statistic for STRING and PathwayCommons. (d) Hazard ratio of each of the HumanNet subtypes compared to subtype 2 with confidence intervals (0.95, 0.8, 0.6 denoted in blue, yellow and orange respectively). (e) Mean and S.E survival in months for each of the subtypes. (f) A Kaplan-Meier plot of the probability of developing platinum drug resistance for HumanNet with four clusters, Logrank P= (Subtype 4 is dropped due to missing annotations for PFI for the majority of patients). 4
5 Supplementary Figure 4 Supplementary Figure 4. Lung cancer association with overall survival. (a) Co-clustering matrices for lung cancer patients, comparing NBS (HumanNet) to standard consensus clustering. (b) Lung cancer patient survival cox proportional hazard model logrank statistic for PathwayCommons. (c) A Kaplan-Meier survival plot with six subtypes. 5
6 Supplementary Figure 5 Supplementary Figure 5. Uterine cancer association with histological type. (a-c) Association with histological subtype vs. the number of clusters (K). (d-f) Association with tumor grade vs. the number of clusters (K) (g) Summary of histological types for each subtype. (h) Summary of tumor grade vs each subtype. 6
7 Supplementary Figure 6 Supplementary Figure 6. Standard predictors of survival are independent of ovarian subtype. (a) Percentage of patients receiving an optimal surgical resection (defined as less than 10mm of residual tumor) does not vary significantly between subtypes ( 2 P-value = 0.77). (b) Federation of Gynaecological Oncologists (FIGO) tumor stage does not show evidence for dependence on tumor subtype ( 2 P-value = 0.48). (c) Age at diagnosis does not show dependence on tumor subtype (One-way ANOVA P-value = 0.89). 7
8 Supplementary Figure 7 Supplementary Figure 7. A network view of genes with high network smoothed mutation scores in ovarian, HumanNet, subtype 2 relative to other subtypes. Node size corresponds to smoothed mutation score. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. 8
9 Supplementary Figure 8 Supplementary Figure 8. A network view of genes with high network smoothed mutation scores in ovarian, HumanNet, subtype 3 relative to other subtypes. Node size corresponds to smoothed mutation score. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. 9
10 Supplementary Figure 9 Supplementary Figure 9. A network view of genes with high smoothed mutation scores in ovarian, HumanNet, subtype 4 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. 10
11 Supplementary Figure 10 Supplementary Figure 10. A network view of genes with high smoothed mutation scores in uterine cancer, STRING, subtype 1 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. Edge thickness corresponds to relative edge confidence in the network, underlined gene names indicate the gene is mutated in this subtype. 11
12 Supplementary Figure 11 Supplementary Figure 11. A network view of genes with high smoothed mutation scores in uterine cancer, STRING, subtype 2 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. Edge thickness corresponds to relative edge confidence in the network, underlined gene names indicate the gene is mutated in this subtype. 12
13 Supplementary Figure 12 Supplementary Figure 12. A network view of genes with high smoothed mutation scores in uterine cancer, STRING, subtype 3 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. Edge thickness corresponds to relative edge confidence in the network, underlined gene names indicate the gene is mutated in this subtype. 13
14 Supplementary Figure 13 Supplementary Figure 13. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 1 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. Edge thickness corresponds to relative edge confidence in the network, underlined gene names indicate the gene is mutated in this subtype. 14
15 Supplementary Figure 14 Supplementary Figure 14. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 2 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. Edge thickness corresponds to relative edge confidence in the network, underlined gene names indicate the gene is mutated in this subtype. 15
16 Supplementary Figure 15 Supplementary Figure 15. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 3 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. Edge thickness corresponds to relative edge confidence in the network, underlined gene names indicate the gene is mutated in this subtype. 16
17 Supplementary Figure 16 Supplementary Figure 16. A network view of genes with high smoothed mutation scores in lung cancer, HumanNet, subtype 5 relative to other subtypes. Node size corresponds to smoothed mutation scores. Node color corresponds to a set of functional classes of interest recovered through manual examination of the resulting network with the aid of the GeneMania Cytoscape plugin. Thickened node outlines indicate genes which are known cancer genes from the Sanger list of cancer genes. Edge thickness corresponds to relative edge confidence in the network, underlined gene names indicate the gene is mutated in this subtype. 17
18 Supplementary Figure 17 Supplementary Figure 17. From mutation-derived subtypes to expression signatures. (a) A Kaplan-Meier analysis of the proportion of patients who acquire platinum resistance in the Tothill et al. expression cohort for subtypes defined in the TCGA dataset using somatic mutations and NBS. (b) Kaplan-Meier survival plots for the Bonome et al. ovarian cancer patients (c) Kaplan-Meier survival plots for a metastudy of ovarian cancer patients by Győrffy et al.. These subtypes were recovered using a shrunken centroid model trained on the TCGA expression data with somatic mutation NBS subtypes as labels. 18
19 Supplementary Figure 18 Supplementary Figure 18. Standard consensus clustering NMF used to recover subtypes in the Tothill et al. expression cohort of ovarian tumors. (a) Standard consensus clustering NMF was performed for 1000 rounds with random restarts on the top 4000 most variable genes in the cohort. Average linkage hierarchical clustering was performed on the co-occurrence matrix to recover the following subtypes. Kaplan-Meier plots are shown for three (b), four (c), and five subtypes (d). 19
20 Supplementary Figure 19 Supplementary Figure 19 Effects of progressively permuting proportions of the lung cancer dataset. Permuting a progressively larger number of mutation uniformly from the entire lung cohort. We report the median likelihood difference of a full model to a base model including just clinical covariates (age, grade, stage, mutation rate, residual tumor after surgery, as well as smoking). The colored regions represent the median absolute deviation (MAD). 20
21 Supplementary Table 1 Supplementary Table 1. Summary of gene interaction networks. The table shows the networks used as part of our analysis. The HumanNet and STRING networks where filtered to include the top 10% of interactions according to the interaction weights. After filtering all edges were treated as unweighted. Nodes Edges Links and description HumanNet v ,243 (7,949) STRING v ,560 (12,233) 476,399 (47,641) 1,638,830 (164,034) PathwayCommons 30 14, ,757 A database of gene interactions, derived using a naïve bayes approach by combining multiple lines of experimental evidence. Comprised of both protein-protein interactions (PPIs) and genetic interactions A database integrating a variety of evidence types including, experimental expression and literature mining approaches to derive a globally weighted network of gene interactions. Comprised of multiple types of gene interactions, including: PPIs, genetic and cocitation. An aggregated repository of gene interactions from several sources including BioGrid, HPRD, IntAct and the NCI set of cancer specific pathways. Comprised of mostly physical PPIs. 21
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
ARTICLE RESEARCH. Macmillan Publishers Limited. All rights reserved
 Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Patient characteristics of training and validation set. Patient selection and inclusion overview can be found in Supp Data 9. Training set (103)
 Roepman P, et al. An immune response enriched 72-gene prognostic profile for early stage Non-Small- Supplementary Data 1. Patient characteristics of training and validation set. Patient selection and inclusion
Roepman P, et al. An immune response enriched 72-gene prognostic profile for early stage Non-Small- Supplementary Data 1. Patient characteristics of training and validation set. Patient selection and inclusion
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant
 National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Surgical
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Surgical
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
List of Figures. List of Tables. Preface to the Second Edition. Preface to the First Edition
 List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
List of Figures List of Tables Preface to the Second Edition Preface to the First Edition xv xxv xxix xxxi 1 What Is R? 1 1.1 Introduction to R................................ 1 1.2 Downloading and Installing
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Evaluation of public cancer datasets and signatures identifies TP53 mutant signatures with robust prognostic and predictive value
 Lehmann et al. BMC Cancer (2015) 15:179 DOI 10.1186/s12885-015-1102-7 RESEARCH ARTICLE Open Access Evaluation of public cancer datasets and signatures identifies TP53 mutant signatures with robust prognostic
Lehmann et al. BMC Cancer (2015) 15:179 DOI 10.1186/s12885-015-1102-7 RESEARCH ARTICLE Open Access Evaluation of public cancer datasets and signatures identifies TP53 mutant signatures with robust prognostic
Expanded View Figures
 Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties
 Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties Bob Obenchain, Risk Benefit Statistics, August 2015 Our motivation for using a Cut-Point
Application of Local Control Strategy in analyses of the effects of Radon on Lung Cancer Mortality for 2,881 US Counties Bob Obenchain, Risk Benefit Statistics, August 2015 Our motivation for using a Cut-Point
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Integrative analysis of survival-associated gene sets in breast cancer
 Varn et al. BMC Medical Genomics (2015) 8:11 DOI 10.1186/s12920-015-0086-0 RESEARCH ARTICLE Open Access Integrative analysis of survival-associated gene sets in breast cancer Frederick S Varn 1, Matthew
Varn et al. BMC Medical Genomics (2015) 8:11 DOI 10.1186/s12920-015-0086-0 RESEARCH ARTICLE Open Access Integrative analysis of survival-associated gene sets in breast cancer Frederick S Varn 1, Matthew
Supplementary. properties of. network types. randomly sampled. subsets (75%
 Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD
 Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Supplementary Information
 Supplementary Information Guided Visual Exploration of Genomic Stratifications in Cancer Marc Streit 1,6, Alexander Lex 2,6, Samuel Gratzl¹, Christian Partl³, Dieter Schmalstieg³, Hanspeter Pfister², Peter
Supplementary Information Guided Visual Exploration of Genomic Stratifications in Cancer Marc Streit 1,6, Alexander Lex 2,6, Samuel Gratzl¹, Christian Partl³, Dieter Schmalstieg³, Hanspeter Pfister², Peter
Statistics 202: Data Mining. c Jonathan Taylor. Final review Based in part on slides from textbook, slides of Susan Holmes.
 Final review Based in part on slides from textbook, slides of Susan Holmes December 5, 2012 1 / 1 Final review Overview Before Midterm General goals of data mining. Datatypes. Preprocessing & dimension
Final review Based in part on slides from textbook, slides of Susan Holmes December 5, 2012 1 / 1 Final review Overview Before Midterm General goals of data mining. Datatypes. Preprocessing & dimension
From Biostatistics Using JMP: A Practical Guide. Full book available for purchase here. Chapter 1: Introduction... 1
 From Biostatistics Using JMP: A Practical Guide. Full book available for purchase here. Contents Dedication... iii Acknowledgments... xi About This Book... xiii About the Author... xvii Chapter 1: Introduction...
From Biostatistics Using JMP: A Practical Guide. Full book available for purchase here. Contents Dedication... iii Acknowledgments... xi About This Book... xiii About the Author... xvii Chapter 1: Introduction...
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplement for: CD4 cell dynamics in untreated HIV-1 infection: overall rates, and effects of age, viral load, gender and calendar time.
 Supplement for: CD4 cell dynamics in untreated HIV-1 infection: overall rates, and effects of age, viral load, gender and calendar time. Anne Cori* 1, Michael Pickles* 1, Ard van Sighem 2, Luuk Gras 2,
Supplement for: CD4 cell dynamics in untreated HIV-1 infection: overall rates, and effects of age, viral load, gender and calendar time. Anne Cori* 1, Michael Pickles* 1, Ard van Sighem 2, Luuk Gras 2,
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
An Example of Business Analytics in Healthcare
 An Example of Business Analytics in Healthcare Colleen McGahan Biostatistical Lead Cancer Surveillance & Outcomes BC Cancer Agency cmcgahan@bccancer.bc.ca Improve Ovarian Cancer Outcomes Business relevancy
An Example of Business Analytics in Healthcare Colleen McGahan Biostatistical Lead Cancer Surveillance & Outcomes BC Cancer Agency cmcgahan@bccancer.bc.ca Improve Ovarian Cancer Outcomes Business relevancy
Reveal Relationships in Categorical Data
 SPSS Categories 15.0 Specifications Reveal Relationships in Categorical Data Unleash the full potential of your data through perceptual mapping, optimal scaling, preference scaling, and dimension reduction
SPSS Categories 15.0 Specifications Reveal Relationships in Categorical Data Unleash the full potential of your data through perceptual mapping, optimal scaling, preference scaling, and dimension reduction
Nature Genetics: doi: /ng.2995
 Supplementary Figure 1 Kaplan-Meier survival curves of patients with brainstem tumors. (a) Comparison of patients with PPM1D mutation versus wild-type PPM1D. (b) Comparison of patients with PPM1D mutation
Supplementary Figure 1 Kaplan-Meier survival curves of patients with brainstem tumors. (a) Comparison of patients with PPM1D mutation versus wild-type PPM1D. (b) Comparison of patients with PPM1D mutation
PBZ FT01_PBZ FT01_TZ FT01_NZ. interface zone (I) tumor zone (TZ) necrotic zone (NZ)
 Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Colon cancer subtypes from gene expression data
 Colon cancer subtypes from gene expression data Nathan Cunningham Giuseppe Di Benedetto Sherman Ip Leon Law Module 6: Applied Statistics 26th February 2016 Aim Replicate findings of Felipe De Sousa et
Colon cancer subtypes from gene expression data Nathan Cunningham Giuseppe Di Benedetto Sherman Ip Leon Law Module 6: Applied Statistics 26th February 2016 Aim Replicate findings of Felipe De Sousa et
Surveillance report Published: 17 March 2016 nice.org.uk
 Surveillance report 2016 Ovarian Cancer (2011) NICE guideline CG122 Surveillance report Published: 17 March 2016 nice.org.uk NICE 2016. All rights reserved. Contents Surveillance decision... 3 Reason for
Surveillance report 2016 Ovarian Cancer (2011) NICE guideline CG122 Surveillance report Published: 17 March 2016 nice.org.uk NICE 2016. All rights reserved. Contents Surveillance decision... 3 Reason for
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
Carcinosarcoma Trial rial in s a in rare malign rare mali ancy
 Carcinosarcoma Trials in a rare malignancy BACKGROUND Rare and highly aggressive epithelial malignancies Biphasic tumors with epithelial and mesenchymal components Uterine carcinomas (UCS) uncommon with
Carcinosarcoma Trials in a rare malignancy BACKGROUND Rare and highly aggressive epithelial malignancies Biphasic tumors with epithelial and mesenchymal components Uterine carcinomas (UCS) uncommon with
Summary... 2 TRANSLATIONAL RESEARCH Tumour gene expression used to direct clinical decision-making for patients with advanced cancers...
 ESMO 2016 Congress 7-11 October, 2016 Copenhagen, Denmark Table of Contents Summary... 2 TRANSLATIONAL RESEARCH... 3 Tumour gene expression used to direct clinical decision-making for patients with advanced
ESMO 2016 Congress 7-11 October, 2016 Copenhagen, Denmark Table of Contents Summary... 2 TRANSLATIONAL RESEARCH... 3 Tumour gene expression used to direct clinical decision-making for patients with advanced
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Table of content. -Supplementary methods. -Figure S1. -Figure S2. -Figure S3. -Table legend
 Table of content -Supplementary methods -Figure S1 -Figure S2 -Figure S3 -Table legend Supplementary methods Yeast two-hybrid bait basal transactivation test Because bait constructs sometimes self-transactivate
Table of content -Supplementary methods -Figure S1 -Figure S2 -Figure S3 -Table legend Supplementary methods Yeast two-hybrid bait basal transactivation test Because bait constructs sometimes self-transactivate
Predicting Kidney Cancer Survival from Genomic Data
 Predicting Kidney Cancer Survival from Genomic Data Christopher Sauer, Rishi Bedi, Duc Nguyen, Benedikt Bünz Abstract Cancers are on par with heart disease as the leading cause for mortality in the United
Predicting Kidney Cancer Survival from Genomic Data Christopher Sauer, Rishi Bedi, Duc Nguyen, Benedikt Bünz Abstract Cancers are on par with heart disease as the leading cause for mortality in the United
VL Network Analysis ( ) SS2016 Week 3
 VL Network Analysis (19401701) SS2016 Week 3 Based on slides by J Ruan (U Texas) Tim Conrad AG Medical Bioinformatics Institut für Mathematik & Informatik, Freie Universität Berlin 1 Motivation 2 Lecture
VL Network Analysis (19401701) SS2016 Week 3 Based on slides by J Ruan (U Texas) Tim Conrad AG Medical Bioinformatics Institut für Mathematik & Informatik, Freie Universität Berlin 1 Motivation 2 Lecture
Nature Genetics: doi: /ng Supplementary Figure 1. Details of sequencing analysis.
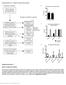 Supplementary Figure 1 Details of sequencing analysis. (a) Flow chart showing which patients fall into each category and were used for analysis. (b) Graph showing the average and median coverage for all
Supplementary Figure 1 Details of sequencing analysis. (a) Flow chart showing which patients fall into each category and were used for analysis. (b) Graph showing the average and median coverage for all
The Cancer Genome Atlas & International Cancer Genome Consortium
 The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
A Case-Study of Pancreatic Cancer in Michigan: Challenges in Reconstructing Residential Histories
 A Case-Study of Pancreatic Cancer in Michigan: Challenges in Reconstructing Residential Histories 1971 1987 2001 2002 Melissa Slotnick Research Associate BioMedware, Inc. slotnick@umich.edu Background
A Case-Study of Pancreatic Cancer in Michigan: Challenges in Reconstructing Residential Histories 1971 1987 2001 2002 Melissa Slotnick Research Associate BioMedware, Inc. slotnick@umich.edu Background
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
TEB. Id4 p63 DAPI Merge. Id4 CK8 DAPI Merge
 a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics
 SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics sapling.cshl.edu Wim Verleyen, Ph.D. Gillis Lab Outline Motivation Disease-gene analysis Enrichment analysis Gene network analysis:
SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics sapling.cshl.edu Wim Verleyen, Ph.D. Gillis Lab Outline Motivation Disease-gene analysis Enrichment analysis Gene network analysis:
Supplementary Figure 1: Classification scheme for non-synonymous and nonsense germline MC1R variants. The common variants with previously established
 Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
Transcriptional Profiles from Paired Normal Samples Offer Complementary Information on Cancer Patient Survival -- Evidence from TCGA Pan-Cancer Data
 Transcriptional Profiles from Paired Normal Samples Offer Complementary Information on Cancer Patient Survival -- Evidence from TCGA Pan-Cancer Data Supplementary Materials Xiu Huang, David Stern, and
Transcriptional Profiles from Paired Normal Samples Offer Complementary Information on Cancer Patient Survival -- Evidence from TCGA Pan-Cancer Data Supplementary Materials Xiu Huang, David Stern, and
Gene expression profiling predicts clinical outcome of prostate cancer. Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M.
 SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Theta sequences are essential for internally generated hippocampal firing fields.
 Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer
 Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Expanded View Figures
 EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
Figure S1: Effects on haptotaxis are independent of effects on cell velocity A)
 Supplemental Figures Figure S1: Effects on haptotaxis are independent of effects on cell velocity A) Velocity of MV D7 fibroblasts expressing different GFP-tagged Ena/VASP family proteins in the haptotaxis
Supplemental Figures Figure S1: Effects on haptotaxis are independent of effects on cell velocity A) Velocity of MV D7 fibroblasts expressing different GFP-tagged Ena/VASP family proteins in the haptotaxis
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
A 16-Gene Signature Distinguishes Anaplastic Astrocytoma from Glioblastoma
 A 16-Gene Signature Distinguishes Anaplastic Astrocytoma from Glioblastoma Soumya Alige Mahabala Rao 1, Sujaya Srinivasan 1, Irene Rosita Pia Patric 1, Alangar Sathyaranjandas Hegde 3, Bangalore Ashwathnarayanara
A 16-Gene Signature Distinguishes Anaplastic Astrocytoma from Glioblastoma Soumya Alige Mahabala Rao 1, Sujaya Srinivasan 1, Irene Rosita Pia Patric 1, Alangar Sathyaranjandas Hegde 3, Bangalore Ashwathnarayanara
Evaluating Classifiers for Disease Gene Discovery
 Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Statistics and Epidemiology Practice Questions
 1. Which of the following is not considered a measure of central tendency? a. Median b. Range c. Mode d. Average 2. Given the following set of values, what is the median? 4 5 9 3 8 3 7 1 5 3 a. 3 b. 5
1. Which of the following is not considered a measure of central tendency? a. Median b. Range c. Mode d. Average 2. Given the following set of values, what is the median? 4 5 9 3 8 3 7 1 5 3 a. 3 b. 5
Agilent GeneSpring/MPP Metadata Analysis Framework
 Agilent GeneSpring/MPP Metadata Analysis Framework Technical Overview Authors Srikanthi R., Pritha Aggarwal, Durairaj R., Maria Kammerer, and Pramila Tata Strand Life Sciences Bangalore, India Michael
Agilent GeneSpring/MPP Metadata Analysis Framework Technical Overview Authors Srikanthi R., Pritha Aggarwal, Durairaj R., Maria Kammerer, and Pramila Tata Strand Life Sciences Bangalore, India Michael
Reporting Checklist for Nature Neuroscience
 Corresponding Author: Manuscript Number: Manuscript Type: Alex Pouget NN-A46249B Article Reporting Checklist for Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 3 # Supplementary Tables:
Corresponding Author: Manuscript Number: Manuscript Type: Alex Pouget NN-A46249B Article Reporting Checklist for Nature Neuroscience # Main Figures: 7 # Supplementary Figures: 3 # Supplementary Tables:
Evidence Based Medicine
 Course Goals Goals 1. Understand basic concepts of evidence based medicine (EBM) and how EBM facilitates optimal patient care. 2. Develop a basic understanding of how clinical research studies are designed
Course Goals Goals 1. Understand basic concepts of evidence based medicine (EBM) and how EBM facilitates optimal patient care. 2. Develop a basic understanding of how clinical research studies are designed
Discontinuation and restarting in patients on statin treatment: prospective open cohort study using a primary care database
 open access Discontinuation and restarting in patients on statin treatment: prospective open cohort study using a primary care database Yana Vinogradova, 1 Carol Coupland, 1 Peter Brindle, 2,3 Julia Hippisley-Cox
open access Discontinuation and restarting in patients on statin treatment: prospective open cohort study using a primary care database Yana Vinogradova, 1 Carol Coupland, 1 Peter Brindle, 2,3 Julia Hippisley-Cox
Supplementary Online Content
 1 Supplementary Online Content Friedman DJ, Piccini JP, Wang T, et al. Association between left atrial appendage occlusion and readmission for thromboembolism among patients with atrial fibrillation undergoing
1 Supplementary Online Content Friedman DJ, Piccini JP, Wang T, et al. Association between left atrial appendage occlusion and readmission for thromboembolism among patients with atrial fibrillation undergoing
Supplemental Figure legends
 Supplemental Figure legends Supplemental Figure S1 Frequently mutated genes. Frequently mutated genes (mutated in at least four patients) with information about mutation frequency, RNA-expression and copy-number.
Supplemental Figure legends Supplemental Figure S1 Frequently mutated genes. Frequently mutated genes (mutated in at least four patients) with information about mutation frequency, RNA-expression and copy-number.
Mutational Impact on Diagnostic and Prognostic Evaluation of MDS
 Mutational Impact on Diagnostic and Prognostic Evaluation of MDS Elsa Bernard, PhD Papaemmanuil Lab, Computational Oncology, MSKCC MDS Foundation ASH 2018 Symposium Disclosure Research funds provided by
Mutational Impact on Diagnostic and Prognostic Evaluation of MDS Elsa Bernard, PhD Papaemmanuil Lab, Computational Oncology, MSKCC MDS Foundation ASH 2018 Symposium Disclosure Research funds provided by
A DNA Repair Pathway Focused Score for Prediction of Outcomes in Ovarian Cancer Treated With Platinum-Based Chemotherapy
 JNCI Journal of the National Cancer Institute Advance Access published April 13, 2012 DOI: 10.1093/jnci/djs177 ARTICLE The Author(s) 2012. Published by Oxford University Press. This is an Open Access article
JNCI Journal of the National Cancer Institute Advance Access published April 13, 2012 DOI: 10.1093/jnci/djs177 ARTICLE The Author(s) 2012. Published by Oxford University Press. This is an Open Access article
Quick start guide for using subscale reports produced by Integrity
 Quick start guide for using subscale reports produced by Integrity http://integrity.castlerockresearch.com Castle Rock Research Corp. November 17, 2005 Integrity refers to groups of items that measure
Quick start guide for using subscale reports produced by Integrity http://integrity.castlerockresearch.com Castle Rock Research Corp. November 17, 2005 Integrity refers to groups of items that measure
Supplemental Information. Molecular, Pathological, Radiological, and Immune. Profiling of Non-brainstem Pediatric High-Grade
 Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs.
 Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas
 Paul et al. Clinical Epigenetics (2017) 9:32 DOI 10.1186/s13148-017-0331-9 RESEARCH Open Access DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas Yashna Paul, Baisakhi
Paul et al. Clinical Epigenetics (2017) 9:32 DOI 10.1186/s13148-017-0331-9 RESEARCH Open Access DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas Yashna Paul, Baisakhi
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Name: BIOS 703 MIDTERM EXAMINATIONS (5 marks per question, total = 100 marks)
 Name: BIOS 703 MIDTERM EXAMINATIONS (5 marks per question, total = 100 marks) You will have 75 minutest to complete this examination. Some of the questions refer to Crizotinib in ROS1- Rearranged Non Small-
Name: BIOS 703 MIDTERM EXAMINATIONS (5 marks per question, total = 100 marks) You will have 75 minutest to complete this examination. Some of the questions refer to Crizotinib in ROS1- Rearranged Non Small-
Linear Regression in SAS
 1 Suppose we wish to examine factors that predict patient s hemoglobin levels. Simulated data for six patients is used throughout this tutorial. data hgb_data; input id age race $ bmi hgb; cards; 21 25
1 Suppose we wish to examine factors that predict patient s hemoglobin levels. Simulated data for six patients is used throughout this tutorial. data hgb_data; input id age race $ bmi hgb; cards; 21 25
Supplementary Figures
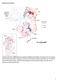 Supplementary Figures Supplementary Fig. 1: Map of the ten countries included in the analysis. Showing the first sub-national administrative boundaries in light grey, and DHS survey locations representing
Supplementary Figures Supplementary Fig. 1: Map of the ten countries included in the analysis. Showing the first sub-national administrative boundaries in light grey, and DHS survey locations representing
Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010
 Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010 C.J.Vaske et al. May 22, 2013 Presented by: Rami Eitan Complex Genomic
Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010 C.J.Vaske et al. May 22, 2013 Presented by: Rami Eitan Complex Genomic
Application of Artificial Neural Network-Based Survival Analysis on Two Breast Cancer Datasets
 Application of Artificial Neural Network-Based Survival Analysis on Two Breast Cancer Datasets Chih-Lin Chi a, W. Nick Street b, William H. Wolberg c a Health Informatics Program, University of Iowa b
Application of Artificial Neural Network-Based Survival Analysis on Two Breast Cancer Datasets Chih-Lin Chi a, W. Nick Street b, William H. Wolberg c a Health Informatics Program, University of Iowa b
Creating prognostic systems for cancer patients: A demonstration using breast cancer
 Received: 16 April 2018 Revised: 31 May 2018 DOI: 10.1002/cam4.1629 Accepted: 1 June 2018 ORIGINAL RESEARCH Creating prognostic systems for cancer patients: A demonstration using breast cancer Mathew T.
Received: 16 April 2018 Revised: 31 May 2018 DOI: 10.1002/cam4.1629 Accepted: 1 June 2018 ORIGINAL RESEARCH Creating prognostic systems for cancer patients: A demonstration using breast cancer Mathew T.
Test Category: Prognostic and Predictive. Clinical Scenario
 Use of Epidermal Growth Factor Receptor (EGFR) Mutation Analysis in Patients with Advanced Non-Small-Cell Lung Cancer (NSCLC) to Determine Erlotinib Use as First-line Therapy Test Category: Prognostic
Use of Epidermal Growth Factor Receptor (EGFR) Mutation Analysis in Patients with Advanced Non-Small-Cell Lung Cancer (NSCLC) to Determine Erlotinib Use as First-line Therapy Test Category: Prognostic
Classification of cancer profiles. ABDBM Ron Shamir
 Classification of cancer profiles 1 Background: Cancer Classification Cancer classification is central to cancer treatment; Traditional cancer classification methods: location; morphology, cytogenesis;
Classification of cancer profiles 1 Background: Cancer Classification Cancer classification is central to cancer treatment; Traditional cancer classification methods: location; morphology, cytogenesis;
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Experimental design and workflow utilized to generate the WMG Protein Atlas.
 Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION
 COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
Supplementary Methods
 Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Expert-guided Visual Exploration (EVE) for patient stratification. Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland
 Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
Expert-guided Visual Exploration (EVE) for patient stratification Hamid Bolouri, Lue-Ping Zhao, Eric C. Holland Oncoscape.sttrcancer.org Paul Lisa Ken Jenny Desert Eric The challenge Given - patient clinical
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant
 National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2012 June 30, 2013 Formula Grant Overview The National Surgical
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2012 June 30, 2013 Formula Grant Overview The National Surgical
Interactive analysis and quality assessment of single-cell copy-number variations
 Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Sum of Neurally Distinct Stimulus- and Task-Related Components.
 SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Inhibition of fatty acid oxidation as a therapy for MYC-overexpressing triplenegative
 SUPPLEMENTARY INFORMATION Inhibition of fatty acid oxidation as a therapy for MYC-overexpressing triplenegative breast cancer Roman Camarda, Alicia Y. Zhou, Rebecca A. Kohnz, Sanjeev Balakrishnan, Celine
SUPPLEMENTARY INFORMATION Inhibition of fatty acid oxidation as a therapy for MYC-overexpressing triplenegative breast cancer Roman Camarda, Alicia Y. Zhou, Rebecca A. Kohnz, Sanjeev Balakrishnan, Celine
