8 Suppression Analysis
|
|
|
- Griselda Fox
- 6 years ago
- Views:
Transcription
1 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: (Hardback); (Electronic) 8 Suppression Analysis OVERVIEW Suppression analysis is an elegant and highly favored molecular genetic tool used to identify genes that are functionally related to the gene of interest. It dates back to the very earliest days of genetics and the work of Sturtevant (1920) and Beadle & Ephrussi (1936) but it was not until the 1960s that the variety of suppression mechanisms and the capabilities of suppressor analysis were fully appreciated. Increased use of suppressor analysis, particularly in model genetic organisms like Saccharomyces, has refined the method making it one of the two accepted genetic methods for identification of functionally related genes or gene products. The second method, enhancer analysis, willbe described in Chapter 9. Functionally related genes encode products capable of carrying out the same or similar functions, are different components of the same metabolic or regulatory pathway, or function together in a structural or enzyme complex. Mutations in these genes might not have been revealed by the original mutant hunt for a variety of reasons. The mutant alleles might have a different and unpredicted phenotype or might not produce a detectable phenotype unless in conjunction with another mutation. Thus, a search for suppressor mutations has the potential forevealing an entirely new spectrum of genes. A suppressor mutation is a mutation that counteracts the effects of the original mutation such that the double mutant individual containing both the original mutation and the suppressor mutation has a phenotype similar to that of the wildtype. Suppressors are isolated when a mutant strain is reverted to restore the wildtype or wild-type-like phenotype. As with the isolation of the original mutation, one must devise a selection/screen that allows the identification of revertants having the wild-type phenotype. One determines whether a revertant is a true revertant, that is restores the original gene sequence, or if it carries a suppressor mutation by crossing the revertant to a wild-type strain and seeing whether the original mutation can be recovered in the progeny. Recovery of the original mutation can only occur if there has been a recombination event separating the original mutation from the suppressor mutation. The suppressor mutation alone will frequently have a mutant phenotype, but this phenotype could be novel or could be similar to the phenotype of the original mutation. INTRAGENIC SUPPRESSORS If the suppressor mutation and the original mutation are in the same gene, this is referred to as intragenic suppression. Recombination between the original mutation and the intragenic suppressor is expected to be rare because the two alterations are very tightly linked. Intragenic suppression can occur by a variety of mechanisms. A classic example is found in Crick et al. (1961). The original mutation used in this
2 . 92 GENETIC TECHNIQUES FOR BIOLOGICAL RESEARCH analysis was a +l frameshift mapping to the 5' end of the riib ORF of the T2 phage of E. coli. Intragenic suppressors of this mutation were produced by a nearby -1 frameshift thereby restoring the correct reading frame. One can also get intragenic suppression of a missense mutation. A missense mutation might functionally inactivate a protein by altering its shape or by increasing its rate of degradation to such an extent that inadequate amounts of the protein are produced. An intragenic suppressor could restore function if it enables the doubly mutant protein to fold into a more active shape or reduces the rate of degradation sufficiently to provide adequate levels of the protein. Changes in the promoter could increase the level of expression of a partially active mutant protein to a level adequate to restore a wildtype-like phenotype. INTERGENIC SUPPRESSION If the suppressor mutation and the original mutation are in different genes, this is referred to as intergenic suppression. Recombination between the original mutation and the suppressor mutation is likely to occur at a high rate since with rare exceptions the suppressor mutation will be unlinked to the original mutation. Intergenic suppressors fall into two broad categories: information suppressors and what we will call function suppressors. Information suppression Information suppressors are mutations in genes involved in the transmission of information from DNA to protein. As such, they act by improving the expression of the mutant gene. Moreover, an information suppressor will suppress any other gene, even those that are functionally unrelated to the original mutant, so long as the mutation in that gene has the same effect on the information transfer process as the mutation in the original gene. The reader is already familiar with nonsense, missense and frameshift suppressors that are mutations in trna genes. These were among the first types of information suppressor identified. Mutations affecting ribosome components also can be information suppressors. This type of information suppression acts at the level of the translation process and alters the reading of the encoded information. Information suppression can also act by increasing the amount of the transcript, such as by altering components of the transcription machinery or complexes involved in post-transcriptional processing. In summary, information suppressors suppress particular types of mutational alteration affecting the information transfer process and can do this in any gene that has that type of alteration. In genetic terms, information suppressors are allele-specific but not genespecific. Researchers interested in the processes of information transfer can use suppressor analysis to obtain mutations in genes involved in these processes but it is important to choose the original mutant strain carefully. For example, if one is interested in translation start-site selection one should work with a mutation in the leader sequence of a gene that exhibits reduced protein production but not reduced transcript levels. Intergenic suppressors that increase translation should be obtained, for example, in translation initiation factors or the small ribosomal subunit. Starting
3 SUPPRESSION ANALYSIS 93 with a promoter mutation or an alteration in the coding region of the gene would not give the desired types of suppressor. As part of the initial characterization of the suppressors, one should test the ability of the suppressor to suppress a comparable mutation in the leader sequence of another unrelated gene. This should sort the suppressor mutations into the desired class of information suppressor from uninteresting mutations affecting some other aspect of the expression of the original mutant gene. Function suppression Function suppressors act to restore or replace the altered gene function. This is accomplished by several means. The activity of a gene product can be appropriately modulated by mutations that affect its post-translational modification, subcellular localization, degradation, or interaction with activators, inhibitors, or other regulatory factors. Mutational changes can substitute another gene product for the mutant gene product either by changing the specificity or abundance of that alternate protein. Or, if the mutant gene encodes a component of a switch regulatory pathway, suppression can occur by the constitutive activation of a downstream component. The isolation of function suppressors provides a means of exploring the role of a gene product in a cellular process and identifies other functionally related gene products or other components of a pathway. Again, if one is interested in isolating function suppressors of a mutant gene, it is important to start the analysis with an appropriate mutation in that gene. The original mutation should affect the gene function, not expression, and thus should be confined to the coding region. Depending on the type of function suppressor desired, one could start with a null mutation, such as a deletion or an insertion, or with a missense mutation. Temperature-sensitive and cold-sensitive mutant alleles are frequently used for function suppression analysis because these types of mutation are almost always missense mutations. There are three mechanisms of function suppression: by-pass suppression, allelespecific suppression, and suppression by epistasis. BY-PASS SUPPRESSION A by-pass suppressor by-passes the need for the mutated gene product by providing the function of that gene product by alternate means. Two mechanisms of by-pass suppression of mutations in hypothetical GENI are shown in Figure 8.1. In Model 1, GENl and GEN2 carry out related but distinct processes that do not functionally overlap in the wild-type strain. The mutation of GENl blocks the reaction thereby creating a mutant. An alteration in GEN2 allows the GEN2 gene product to acquire a new function that enables it to substitute for the GENl product thereby bypassing the GENl mutation. The short arrow indicates that the by-pass may be less effective than the wild-type function. In Model 2, expressionof another geneis increased so that its product can substitute for the GENl product. Gen3 protein is capable of carrying out the same or a similar function as the Genl protein but not at rates adequate for a wild-type phenotype. Perhaps the specific activity of Gen3p
4 94 GENETIC TECHNIQUES FOR BIOLOGICAL RESEARCH Model 1 Wild-type Mutant Suppressed i I # I I genl GEN2-sup R GENl GENZ genl GENZ GENl 1 x z x z x z X 1 h Model 2 Wild-type Mutant Suppressed R R genii R GEN3 X h R GEN3 X genl f I GEN3-sup X X x x Figure 8.1 Models of by-pass suppression for the Genlp substrate R is very low. An increase in the expression levels of GEN3 by the suppressor mutation provides levels of Gen3 protein sufficient to allow it to substitute for the loss of GENI. It is important to note that by-pass suppression is not dependent on the original gene product and thus by-pass suppressors are able to suppress either missense mutations or null mutations of the original gene. In addition, the suppressor allele is dominant in both Model 1 and Model 2. ALLELE-SPECIFIC SUPPRESSION In allele-specific suppression, a suppressor mutation in GEN2 is able to suppress only a particular non-null allele or select a group of non-null alleles in GENl. Experience has demonstrated that allele specificity of function suppressors implies that the products of the two genes interact directly, i.e. they make physical contact with each other. Both by-pass suppression and suppression by epistasis (see below) can occur with null alleles, i.e. alleles that produce no protein product. Therefore, if a suppressor is not capable of suppressing a null mutant allele but is only capable of suppressing missense mutations of a gene, then the mechanism of suppression is allele specific. Allele-specific function suppression is considered strong evidence that the gene products of the original mutant gene and the suppressor gene physically interact. This interaction could indicate an enzyme-substrate type of interaction (as between a protein kinase and its target protein), an interaction between components of a heterodimer or heteromultimeric complex, or an interaction between the catalytic subunit of an enzyme complex and its regulatory subunit(s). To conceptualize allele-specific suppression, consider the fact that two proteins interact with each other only over a small portion of their surfaces, and that within
5 SUPPRESSION ANALYSIS Original mutation L Suppress r mutation 95 Figure 8.2 Model of allele-specific suppression these defined regions of interaction specific residues on the surfaces of the proteins make the most significant contributions to the binding strength of the interaction. This is illustrated in Figure 8.2. Mutations that alter these residues affect the binding strength of the interaction either by weakening or strengthening it. If the original mutation in GENl decreases the binding of the Genl protein to the Gen2 protein, a suppressor mutation in GEN2 might act to restore strong binding of Genlp to Gen2p. Suppression will occur if, and only if, the suppressor mutation in GENZ alters a residue mapping within the surface region of Gen2p that physically makes contact with Genlp. In the context of this explanation, the gen2 suppressor mutation can only be expected to suppress genl mutations that alter residuesin the region of Genlp involved in the Genlp-Gen2p interaction; that is, they would be specific to alleles altering residues in this region. Suppressor mutations in GEN2 would not be expected to suppress null mutations of GENZ (large deletions, 5 nonsense or frameshift mutations, insertions, or other rearrangements). Allele-specific suppressors also would not be expected to suppress mutations elsewhere in Genlp outside of the surface region of Genlp that interacts with Gen2p, for example in the Gen3p binding site. The suppressor mutation of gen2 alone, that is when not in combination with the original genl mutation, is likely to have a mutant phenotype similar to the genl mutant phenotype because both affecthe Genlp-Gen2p interaction. Allele-specific suppression is not limited to protein-protein interactions. It can be used to characterize DNA-protein or RNA-protein interactions. When two proteins (or a DNA sequence and a protein) are known to interact directly, allelespecific suppression can be used to explore the details of the binding, i.e.which specific residues and/or basepairs are involved. Alternately, one can identify novel proteins that interact with the product of a known gene by identifying allele-specific suppressors of a mutant allele of that gene. SUPPRESSION BY EPISTASIS Suppression by epistasis occurs between genes whose products are components of a switch regulatory pathway as described in Chapter 6 on epistasis. The component proteins of switch regulatory pathway alternate between the on state and the off
6 96 GENETIC TECHNIQUES FOR BIOLOGICAL RESEARCH state. Mutations in these proteins can permanently shift them to either state. Such mutations act to separate the switch regulatory pathway from the upstream signal. The pathway can be constitutive (that is, unregulated) by activating the pathway in the absence of the stimulatory signal or despite the presence of inhibitory signal. Alternately, mutations can block the pathway despite the presence of the stimulatory signal or in the absence of an inhibitory signal. JThe ultimate effectof a mutation in a component protein depends on whether the component is a positive regulator or a negative regulator. For example, in the switch regulatory pathway shown below, the signal activates protein l, protein 1 activates protein 2, activated protein 2 inhibits protein 3, and protein 3 - is an inhibitor of - GEN4 expression. (GEN4) Signal protein 1 protein 2 -+ protein 3 (GENl) (GEN2) (GEN3) Recessive mutations in GENl or GEN2 lead to a lack of GEN4 expression. Recessive mutations in GEN3 lead to the constitutive expression of GEN4. A strain doubly mutant for recessive mutations in GENl and GEN3 or GEN2 and GEN3 would be constitutive for GEN4 expression. Thus, a loss of function mutation in GEN3 will suppress lossof function mutations in GENI and GEN2, and restore GEN4 expression to genl and gen2 mutant strains. This suppression of genl and gen2 mutations by a mutation in gen3 is suppression by epistasis. Of course, in this case the suppression does not restore normal regulation but it does restore expression of GEN4, albeit constitutively. Suppression by epistasis can occur with null or non-null alleles of either the original mutant gene or the suppressor gene. For example, in the above pathway, a deletion of GEN3 will suppress either a missense mutation or a deletion mutation of GENl. A dominant constitutive mutation in GENl would activate this pathway in the absence of a signal. This constitutive activation would be suppressed by recessive mutations in GEN2. OVEREXPRESSION SUPPRESSION Because of the variety of plasmid vectors available for Saccharomyces it is possible to modulate the expression levels of a desired gene in a number of ways and not simplyby in situ alterations in the promoter region ofgenes. This type of suppression is called multicopy suppression or suppression by overexpression or overproduction. Multicopy plasmids are frequently used to obtain overproduction of a particular protein. A gene carried by a YEp vector will be present at up to 50 copieslcell and thus one might expect a 50-fold overproduction of the protein. Vectors are available that allow one to fuse the coding region of a gene to any one of several different Saccharomyces promoters that have different expression levels and some of these are easily regulated by environmental signals such as nutrient availability. These are both high and low copy vectors and, as a result, a wide range of expression levels of the inserted gene can be achieved (see Chapter l; Mumberg et al., 1995; Labbe & Thiele, 1999). Libraries can be made with these vectors and
7 SUPPRESSION 97 screened for suppression of mutant strains. The major strength of this type of suppressor hunt is that one can very simply identify the suppressing gene by recovering the suppression plasmid and sequencing its insert fragment. All three types of suppression can be obtained by multicopy or overexpression suppression. BY-PASS SUPPRESSION BY OVEREXPRESSION Overproduction of a gene product can lead to by-pass suppression. This is similar to Model 2 of by-pass suppression (above) except that the overproduction is from the plasmid-borne copy of the suppressing gene. ALLELE-SPECIFIC SUPPRESSION BY OVEREXPRESSION One might compensate for the reduced binding constant between mutant Protein 1 and wild-type Protein 2 by overproduction of wild-type Protein 2. Abundant amounts of Protein 2 could kinetically drive complex formation despite the lower binding constant by a mass action effect. This is a form of allele-specific suppression because it could not occur with null alleles of the gene encoding Protein 1. Mutant Protein 1 isnecessary for the suppression. Asisdiscussed above, overexpression allele-specific suppression of genl mutant alleles by GEN2 implies that the product of GEN2 binds to the product of GENl at the site of the mutant residue. Similar overproduction of Protein 3 should not suppress the same genl mutations as overproduction of Protein 2. OVEREXPRESSION SUPPRESSION BY EPISTASIS In a switch regulatory pathway, overproduction of an activator or repressor could, respectively, activate or block a pathway regardless of the upstream signal. For example, overproduction of an activator (positive regulator) might activate a blocked pathway by overcoming an inhibitory regulator (perhaps by binding to it and titrating out its effects). Or, if the activator has a very low constitutive activity, its overproduction might be sufficient to restore progress through the pathway. Comparable situations can be proposed for repressors (negative regulator). The suppressing gene product necessarily would be downstream of the original mutant gene product. Moreover, the effect of overexpression of a particular component will depend on the role of the component in the pathway. REFERENCES AND FURTHER READING Beadle, G.W. & B. Ephrussi (1936) Development of eye colors in Drosophila: transplantation experiments with suppressor of vermillion. Proc. Natl Acad. Sci. USA 22: 536. Crick, F., L. Barnett, S. Brenner, & R.J. Watts-Tobin (1961) General nature of the genetic code for proteins. Nature 192: Gorini, L. (1970) Informational suppression. Ann. Rev. Genet. 4: Gorini, L. & J. Beckwith (1966) Suppression. Ann. Rev. Microbiol. 20: Hartman, P.E. & J.R. Roth (1973) Mechanisms of suppression. Adv. Genet. 17:
8 98 GENETIC TECHNIQUES BIOLOGICAL FOR RESEARCH Labbe, S. & D.J. Thiele (1999) Copper ion inducible and repressible promoter systems in yeast. Meth. Enzymol. 306: Mount, S. & P. Anderson (2000) Expanding the definition of informational suppression. Trends Genet. 16: 157. Mumberg, D., R. Muller, & M. Funk (1995) Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene 156: Prelich, G. (1999) Suppression mechanisms: themes from variations. Trends Genet. 15: Rine, J. (1991) Gene overexpression in studies of Saccharomyces cerevisiue. Meth. Enzymol. 194: Sturtevant, A.H. (1920) The vermillion gene and gynandromorphism. Proc. Soc. Exp. Biol. Med. 17:
Genetic suppressors and enhancers provide clues to gene regulation and genetic pathways
 Genetic suppressors and enhancers provide clues to gene regulation and genetic pathways Suppressor mutation: a second mutation results in a less severe phenotype than the original mutation Suppressor mutations
Genetic suppressors and enhancers provide clues to gene regulation and genetic pathways Suppressor mutation: a second mutation results in a less severe phenotype than the original mutation Suppressor mutations
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Tumor suppressor genes D R. S H O S S E I N I - A S L
 Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
SUPPLEMENTARY INFORMATION
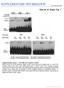 doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
doi:10.1038/nature10495 WWW.NATURE.COM/NATURE 1 2 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 3 4 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 5 6 WWW.NATURE.COM/NATURE WWW.NATURE.COM/NATURE 7 8 WWW.NATURE.COM/NATURE
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
HOW AND WHY GENES ARE REGULATED HOW AND WHY GENES ARE REGULATED. Patterns of Gene Expression in Differentiated Cells
 HOW AND WHY GENES ARE REGULATED 5 HOW AND WHY GENES ARE REGULATED 6 Every somatic cell in an organism contains identical genetic instructions. They all share the same genome. So what makes cells different
HOW AND WHY GENES ARE REGULATED 5 HOW AND WHY GENES ARE REGULATED 6 Every somatic cell in an organism contains identical genetic instructions. They all share the same genome. So what makes cells different
Early Embryonic Development
 Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Early cell death (FGF) B No RunX transcription factor produced Yes No differentiation
 Solution Key - Practice Questions Question 1 a) A recent publication has shown that the fat stem cells (FSC) can act as bone stem cells to repair cavities in the skull, when transplanted into immuno-compromised
Solution Key - Practice Questions Question 1 a) A recent publication has shown that the fat stem cells (FSC) can act as bone stem cells to repair cavities in the skull, when transplanted into immuno-compromised
Chapter 11 Gene Expression
 Chapter 11 Gene Expression 11-1 Control of Gene Expression Gene Expression- the activation of a gene to form a protein -a gene is on or expressed when it is transcribed. -cells do not always need to produce
Chapter 11 Gene Expression 11-1 Control of Gene Expression Gene Expression- the activation of a gene to form a protein -a gene is on or expressed when it is transcribed. -cells do not always need to produce
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Variations in Chromosome Structure & Function. Ch. 8
 Variations in Chromosome Structure & Function Ch. 8 1 INTRODUCTION! Genetic variation refers to differences between members of the same species or those of different species Allelic variations are due
Variations in Chromosome Structure & Function Ch. 8 1 INTRODUCTION! Genetic variation refers to differences between members of the same species or those of different species Allelic variations are due
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single
 8.3 A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single chromosome can alter their pattern of inheritance from those
8.3 A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single chromosome can alter their pattern of inheritance from those
Problem Set 5 KEY
 2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
Chapter 7 Conclusions
 VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
MCB140: Second Midterm Spring 2010
 MCB140: Second Midterm Spring 2010 Before you start, print your name and student identification number (S.I.D) at the top of each page. There are 11 pages including this page. You will have 150 minutes
MCB140: Second Midterm Spring 2010 Before you start, print your name and student identification number (S.I.D) at the top of each page. There are 11 pages including this page. You will have 150 minutes
Biology 105: Introduction to Genetics Midterm EXAM. Part1. Definitions. 1 Recessive allele. Name. Student ID. 2 Homologous chromosomes
 Biology 105: Introduction to Genetics Midterm EXAM Part1 Definitions 1 Recessive allele Name Student ID 2 Homologous chromosomes Before starting, write your name on the top of each page Make sure you have
Biology 105: Introduction to Genetics Midterm EXAM Part1 Definitions 1 Recessive allele Name Student ID 2 Homologous chromosomes Before starting, write your name on the top of each page Make sure you have
Laboratory. Mendelian Genetics
 Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Chapter 15: The Chromosomal Basis of Inheritance
 Name Period Chapter 15: The Chromosomal Basis of Inheritance Concept 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2.
Name Period Chapter 15: The Chromosomal Basis of Inheritance Concept 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2.
Section Chapter 14. Go to Section:
 Section 12-3 Chapter 14 Go to Section: Content Objectives Write these Down! I will be able to identify: The origin of genetic differences among organisms. The possible kinds of different mutations. The
Section 12-3 Chapter 14 Go to Section: Content Objectives Write these Down! I will be able to identify: The origin of genetic differences among organisms. The possible kinds of different mutations. The
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/22278 holds various files of this Leiden University dissertation. Author: Cunha Carvalho de Miranda, Noel Filipe da Title: Mismatch repair and MUTYH deficient
Cover Page The handle http://hdl.handle.net/1887/22278 holds various files of this Leiden University dissertation. Author: Cunha Carvalho de Miranda, Noel Filipe da Title: Mismatch repair and MUTYH deficient
RUNX1 and FPD/AML Translational Research. The Leukemia and Lymphoma Society / Babich Family Foundation Partnership. September 2016
 www.lls.org www.runx1.com RUNX1 and FPD/AML Translational Research The Leukemia and Lymphoma Society / Babich Family Foundation Partnership September 2016 Prepared by L. Greenberger, PhD Chief Scientific
www.lls.org www.runx1.com RUNX1 and FPD/AML Translational Research The Leukemia and Lymphoma Society / Babich Family Foundation Partnership September 2016 Prepared by L. Greenberger, PhD Chief Scientific
RNA and Protein Synthesis Guided Notes
 RNA and Protein Synthesis Guided Notes is responsible for controlling the production of in the cell, which is essential to life! o DNARNAProteins contain several thousand, each with directions to make
RNA and Protein Synthesis Guided Notes is responsible for controlling the production of in the cell, which is essential to life! o DNARNAProteins contain several thousand, each with directions to make
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
BIOLOGY - CLUTCH CH.15 - CHROMOSOMAL THEORY OF INHERITANCE
 !! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
!! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
C) The graph should look exactly like the graph on the left (Mut1 cells + Mating Pheromone for 3 hours at 25 degrees). The cells arrest in G1.
 706-2000-Exam 4 Answer Key 1) The question asks you to explain peaks A and B in the top graph. The other two graphs were there to give you hints. The question did not ask for these other two graphs to
706-2000-Exam 4 Answer Key 1) The question asks you to explain peaks A and B in the top graph. The other two graphs were there to give you hints. The question did not ask for these other two graphs to
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER
 REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
3. What law of heredity explains that traits, like texture and color, are inherited independently of each other?
 Section 2: Genetics Chapter 11 pg. 308-329 Part 1: Refer to the table of pea plant traits on the right. Then complete the table on the left by filling in the missing information for each cross. 6. What
Section 2: Genetics Chapter 11 pg. 308-329 Part 1: Refer to the table of pea plant traits on the right. Then complete the table on the left by filling in the missing information for each cross. 6. What
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Central Dogma. Central Dogma. Translation (mrna -> protein)
 Central Dogma Central Dogma Translation (mrna -> protein) mrna code for amino acids 1. Codons as Triplet code 2. Redundancy 3. Open reading frames 4. Start and stop codons 5. Mistakes in translation 6.
Central Dogma Central Dogma Translation (mrna -> protein) mrna code for amino acids 1. Codons as Triplet code 2. Redundancy 3. Open reading frames 4. Start and stop codons 5. Mistakes in translation 6.
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Function and Regulation of Yeast Hexose Transporters
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Sept. 1999, p. 554 569 Vol. 63, No. 3 1092-2172/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Function and Regulation of
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Sept. 1999, p. 554 569 Vol. 63, No. 3 1092-2172/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Function and Regulation of
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Activation of cellular proto-oncogenes to oncogenes. How was active Ras identified?
 Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Genetic information flows from mrna to protein through the process of translation
 Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
BIOL2005 WORKSHEET 2008
 BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
Chapter 15: The Chromosomal Basis of Inheritance
 Name Chapter 15: The Chromosomal Basis of Inheritance 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2. Explain the law
Name Chapter 15: The Chromosomal Basis of Inheritance 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2. Explain the law
TRANSLATION: 3 Stages to translation, can you guess what they are?
 TRANSLATION: Translation: is the process by which a ribosome interprets a genetic message on mrna to place amino acids in a specific sequence in order to synthesize polypeptide. 3 Stages to translation,
TRANSLATION: Translation: is the process by which a ribosome interprets a genetic message on mrna to place amino acids in a specific sequence in order to synthesize polypeptide. 3 Stages to translation,
Cancer. October is National Breast Cancer Awareness Month
 Cancer October is National Breast Cancer Awareness Month Objectives 1: Gene regulation Explain how cells in all the different parts of your body develop such different characteristics and functions. Contrast
Cancer October is National Breast Cancer Awareness Month Objectives 1: Gene regulation Explain how cells in all the different parts of your body develop such different characteristics and functions. Contrast
Chapter 16 Mutations. Practice Questions:
 Biology 234 J. G. Doheny Chapter 16 Mutations Practice Questions: Answer the following questions with one or two sentences. 1. List the name of one test that can be used to identify mutagens. 2. What is
Biology 234 J. G. Doheny Chapter 16 Mutations Practice Questions: Answer the following questions with one or two sentences. 1. List the name of one test that can be used to identify mutagens. 2. What is
Chapter 15 - The Chromosomal Basis of Inheritance. A. Bergeron +AP Biology PCHS
 Chapter 15 - The Chromosomal Basis of Inheritance A. Bergeron +AP Biology PCHS Do Now - Predicting Unpredictable Genotypes As an inexperienced (albeit precocious) gardener, I am always looking to maximize
Chapter 15 - The Chromosomal Basis of Inheritance A. Bergeron +AP Biology PCHS Do Now - Predicting Unpredictable Genotypes As an inexperienced (albeit precocious) gardener, I am always looking to maximize
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Mutations in the Caenorhabditis elegans U2AF Large Subunit UAF-1 Al= of a 3' Splice Site In Vivo
 Mutations in the Caenorhabditis elegans U2AF Large Subunit UAF-1 Al= of a 3' Splice Site In Vivo The MIT Faculty has made this article openly available. Please share how this access benefits you. Your
Mutations in the Caenorhabditis elegans U2AF Large Subunit UAF-1 Al= of a 3' Splice Site In Vivo The MIT Faculty has made this article openly available. Please share how this access benefits you. Your
Department of Chemistry, Université de Montréal, C.P. 6128, Succursale centre-ville, Montréal, Québec, H3C 3J7, Canada.
 Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Introduction 1/3. The protagonist of our story: the prokaryotic ribosome
 Introduction 1/3 The protagonist of our story: the prokaryotic ribosome Introduction 2/3 The core functional domains of the ribosome (the peptidyl trasferase center and the decoding center) are composed
Introduction 1/3 The protagonist of our story: the prokaryotic ribosome Introduction 2/3 The core functional domains of the ribosome (the peptidyl trasferase center and the decoding center) are composed
The Determination of the Genetic Order and Genetic Map for the Eye Color, Wing Size, and Bristle Morphology in Drosophila melanogaster
 Kudlac 1 Kaitie Kudlac March 24, 2015 Professor Ma Genetics 356 The Determination of the Genetic Order and Genetic Map for the Eye Color, Wing Size, and Bristle Morphology in Drosophila melanogaster Abstract:
Kudlac 1 Kaitie Kudlac March 24, 2015 Professor Ma Genetics 356 The Determination of the Genetic Order and Genetic Map for the Eye Color, Wing Size, and Bristle Morphology in Drosophila melanogaster Abstract:
Cellular Signaling Pathways. Signaling Overview
 Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Neoplasia 18 lecture 6. Dr Heyam Awad MD, FRCPath
 Neoplasia 18 lecture 6 Dr Heyam Awad MD, FRCPath ILOS 1. understand the role of TGF beta, contact inhibition and APC in tumorigenesis. 2. implement the above knowledge in understanding histopathology reports.
Neoplasia 18 lecture 6 Dr Heyam Awad MD, FRCPath ILOS 1. understand the role of TGF beta, contact inhibition and APC in tumorigenesis. 2. implement the above knowledge in understanding histopathology reports.
Introduction. Cancer Biology. Tumor-suppressor genes. Proto-oncogenes. DNA stability genes. Mechanisms of carcinogenesis.
 Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Introduction to Genetics
 Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Mutants and HBV vaccination. Dr. Ulus Salih Akarca Ege University, Izmir, Turkey
 Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
STRUCTURAL CHROMOSOMAL ABERRATIONS
 STRUCTURAL CHROMOSOMAL ABERRATIONS Structural chromosomal aberrations cause structural abnormalities in chromosome structure. They alter the sequence or the kind of genes present in chromosome. These are
STRUCTURAL CHROMOSOMAL ABERRATIONS Structural chromosomal aberrations cause structural abnormalities in chromosome structure. They alter the sequence or the kind of genes present in chromosome. These are
THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15
 THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15 What you must know: Inheritance in sex-linked genes. Inheritance of linked genes and chromosomal mapping. How alteration of chromosome number or structurally
THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15 What you must know: Inheritance in sex-linked genes. Inheritance of linked genes and chromosomal mapping. How alteration of chromosome number or structurally
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
6.5 Enzymes. Enzyme Active Site and Substrate Specificity
 180 Chapter 6 Metabolism 6.5 Enzymes By the end of this section, you will be able to: Describe the role of enzymes in metabolic pathways Explain how enzymes function as molecular catalysts Discuss enzyme
180 Chapter 6 Metabolism 6.5 Enzymes By the end of this section, you will be able to: Describe the role of enzymes in metabolic pathways Explain how enzymes function as molecular catalysts Discuss enzyme
Figure 1: Transmission of Wing Shape & Body Color Alleles: F0 Mating. Figure 1.1: Transmission of Wing Shape & Body Color Alleles: Expected F1 Outcome
 I. Chromosomal Theory of Inheritance As early cytologists worked out the mechanism of cell division in the late 1800 s, they began to notice similarities in the behavior of BOTH chromosomes & Mendel s
I. Chromosomal Theory of Inheritance As early cytologists worked out the mechanism of cell division in the late 1800 s, they began to notice similarities in the behavior of BOTH chromosomes & Mendel s
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Lecture 3-4. Identification of Positive Regulators downstream of SA
 Lecture 3-4 Identification of Positive Regulators downstream of SA Positive regulators between SA and resistance/pr Systemic resistance PR gene expression? SA Local infection Necrosis SA What could they
Lecture 3-4 Identification of Positive Regulators downstream of SA Positive regulators between SA and resistance/pr Systemic resistance PR gene expression? SA Local infection Necrosis SA What could they
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
p53 and Apoptosis: Master Guardian and Executioner Part 2
 p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer
 AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
AD Award Number: W81XWH-04-1-0325 TITLE: A Mouse Model to Investigate the Role of DBC2 in Breast Cancer PRINCIPAL INVESTIGATOR: Valerie Boka CONTRACTING ORGANIZATION: University of Texas Health Science
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
Chromosomes, Mapping, and the Meiosis-Inheritance Connection. Chapter 13
 Chromosomes, Mapping, and the Meiosis-Inheritance Connection Chapter 13 Chromosome Theory Chromosomal theory of inheritance - developed in 1902 by Walter Sutton - proposed that genes are present on chromosomes
Chromosomes, Mapping, and the Meiosis-Inheritance Connection Chapter 13 Chromosome Theory Chromosomal theory of inheritance - developed in 1902 by Walter Sutton - proposed that genes are present on chromosomes
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy
 Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
MECHANISM OF THE ORIGIN OF X-RAY INDUCED NOTCH. Summary.-Comparison has been made, using salivary gland chromosomes,
 24 GENETICS: DEMEREC AND FANO PROC. N. A. S. the male pronucleus, the breaks or potential breaks may remain capable of reunion for a limited time during which contacts with other chromosomes may be realized.
24 GENETICS: DEMEREC AND FANO PROC. N. A. S. the male pronucleus, the breaks or potential breaks may remain capable of reunion for a limited time during which contacts with other chromosomes may be realized.
Viral Genetics. BIT 220 Chapter 16
 Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
The Rts1 Regulatory Subunit of Protein Phosphatase 2A Is Required for Control of G1 Cyclin Transcription and Nutrient Modulation of Cell Size
 The Rts1 Regulatory Subunit of Protein Phosphatase 2A Is Required for Control of G1 Cyclin Transcription and Nutrient Modulation of Cell Size Karen Artiles, Stephanie Anastasia, Derek McCusker, Douglas
The Rts1 Regulatory Subunit of Protein Phosphatase 2A Is Required for Control of G1 Cyclin Transcription and Nutrient Modulation of Cell Size Karen Artiles, Stephanie Anastasia, Derek McCusker, Douglas
oncogenes-and- tumour-suppressor-genes)
 Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
GENE EXPRESSION. Amoeba Sisters video 3pk9YVo. Individuality & Mutations
 Amoeba Sisters video https://www.youtube.com/watch?v=giez 3pk9YVo GENE EXPRESSION Individuality & Mutations Complete video handout http://www.amoebasisters.com/uploads/ 2/1/9/0/21902384/video_recap_of_muta
Amoeba Sisters video https://www.youtube.com/watch?v=giez 3pk9YVo GENE EXPRESSION Individuality & Mutations Complete video handout http://www.amoebasisters.com/uploads/ 2/1/9/0/21902384/video_recap_of_muta
Non-Mendelian inheritance
 Non-Mendelian inheritance Focus on Human Disorders Peter K. Rogan, Ph.D. Laboratory of Human Molecular Genetics Children s Mercy Hospital Schools of Medicine & Computer Science and Engineering University
Non-Mendelian inheritance Focus on Human Disorders Peter K. Rogan, Ph.D. Laboratory of Human Molecular Genetics Children s Mercy Hospital Schools of Medicine & Computer Science and Engineering University
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
GONADAL MOSAICISM AS A FACTOR IN DETERMINING THE RATIO OF VISIBLE TO LETHAL, MUTATIONS IN DROSOPHILA'
 GONADAL MOSAICISM AS A FACTOR IN DETERMINING THE RATIO OF VISIBLE TO LETHAL, MUTATIONS IN DROSOPHILA' LUOLIN S. BROWNING AND EDGAR ALTENBURG Texas Medical Center, Inc., Houston, Texas, and Rice University,
GONADAL MOSAICISM AS A FACTOR IN DETERMINING THE RATIO OF VISIBLE TO LETHAL, MUTATIONS IN DROSOPHILA' LUOLIN S. BROWNING AND EDGAR ALTENBURG Texas Medical Center, Inc., Houston, Texas, and Rice University,
A Genetic Program for Embryonic Development
 Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
SSN SBPM Workshop Exam One. Short Answer Questions & Answers
 SSN SBPM Workshop Exam One Short Answer Questions & Answers 1. Describe the effects of DNA damage on the cell cycle. ANS : DNA damage causes cell cycle arrest at a G2 checkpoint. This arrest allows time
SSN SBPM Workshop Exam One Short Answer Questions & Answers 1. Describe the effects of DNA damage on the cell cycle. ANS : DNA damage causes cell cycle arrest at a G2 checkpoint. This arrest allows time
Biology Developmental Biology Spring Quarter Midterm 1 Version A
 Biology 411 - Developmental Biology Spring Quarter 2013 Midterm 1 Version A 75 Total Points Open Book Choose 15 out the 20 questions to answer (5 pts each). Only the first 15 questions that are answered
Biology 411 - Developmental Biology Spring Quarter 2013 Midterm 1 Version A 75 Total Points Open Book Choose 15 out the 20 questions to answer (5 pts each). Only the first 15 questions that are answered
Gene Regulation - 4. One view of the Lactose Operon
 Gene Regulation - 1 Regulating Genes We have been discussing the structure of DNA and that the information stored in DNA is used to direct protein synthesis. We've studied how RNA molecules are used to
Gene Regulation - 1 Regulating Genes We have been discussing the structure of DNA and that the information stored in DNA is used to direct protein synthesis. We've studied how RNA molecules are used to
F a coding sequence such that the frame of reference of translation is disturbed.
 Copyright 0 1983 by the Genetics Society of America NEW SUPPRESSORS OF FRAMESHIFT MUTATIONS IN SALMONELLA TYPHIMURIUM TADAHIKO KOHNO, LIONELLO BOSSI AND JOHN R. ROTH Department of Biology, University of
Copyright 0 1983 by the Genetics Society of America NEW SUPPRESSORS OF FRAMESHIFT MUTATIONS IN SALMONELLA TYPHIMURIUM TADAHIKO KOHNO, LIONELLO BOSSI AND JOHN R. ROTH Department of Biology, University of
Chapt 15: Molecular Genetics of Cell Cycle and Cancer
 Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
AP Biology Summer Assignment Chapter 3 Quiz
 AP Biology Summer Assignment Chapter 3 Quiz 2016-17 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. All of the following are found in a DNA nucleotide
AP Biology Summer Assignment Chapter 3 Quiz 2016-17 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. All of the following are found in a DNA nucleotide
Clinical Spectrum and Genetic Mechanism of GLUT1-DS. Yasushi ITO (Tokyo Women s Medical University, Japan)
 Clinical Spectrum and Genetic Mechanism of GLUT1-DS Yasushi ITO (Tokyo Women s Medical University, Japan) Glucose transporter type 1 (GLUT1) deficiency syndrome Mutation in the SLC2A1 / GLUT1 gene Deficiency
Clinical Spectrum and Genetic Mechanism of GLUT1-DS Yasushi ITO (Tokyo Women s Medical University, Japan) Glucose transporter type 1 (GLUT1) deficiency syndrome Mutation in the SLC2A1 / GLUT1 gene Deficiency
Viral vaccines. Lec. 3 أ.د.فائزة عبد هللا مخلص
 Lec. 3 أ.د.فائزة عبد هللا مخلص Viral vaccines 0bjectives 1-Define active immunity. 2-Describe the methods used for the preparation of attenuated live & killed virus vaccines. 3- Comparison of Characteristics
Lec. 3 أ.د.فائزة عبد هللا مخلص Viral vaccines 0bjectives 1-Define active immunity. 2-Describe the methods used for the preparation of attenuated live & killed virus vaccines. 3- Comparison of Characteristics
Problem Set 8 Key 1 of 8
 7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
Supplemental Material for. Figure S1. Identification of TetR responsive promoters in F. novicida and E. coli.
 Supplemental Material for Synthetic promoters functional in Francisella novicida and Escherichia coli Ralph L. McWhinnie and Francis E. Nano Department of Biochemistry and Microbiology, University of Victoria,
Supplemental Material for Synthetic promoters functional in Francisella novicida and Escherichia coli Ralph L. McWhinnie and Francis E. Nano Department of Biochemistry and Microbiology, University of Victoria,
7.012 Problem Set 6 Solutions
 Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
DNA codes for RNA, which guides protein synthesis.
 Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Single Gene (Monogenic) Disorders. Mendelian Inheritance: Definitions. Mendelian Inheritance: Definitions
 Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
MEDICAL GENOMICS LABORATORY. Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG)
 Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Section D: The Molecular Biology of Cancer
 CHAPTER 19 THE ORGANIZATION AND CONTROL OF EUKARYOTIC GENOMES Section D: The Molecular Biology of Cancer 1. Cancer results from genetic changes that affect the cell cycle 2. Oncogene proteins and faulty
CHAPTER 19 THE ORGANIZATION AND CONTROL OF EUKARYOTIC GENOMES Section D: The Molecular Biology of Cancer 1. Cancer results from genetic changes that affect the cell cycle 2. Oncogene proteins and faulty
