Investigating the Role of Inositol Polyphosphate 4-Phosphatase Type II (INPP4B) Overexpression on Autophagy in Acute Myeloid Leukemia (AML)
|
|
|
- Marylou Porter
- 6 years ago
- Views:
Transcription
1 Investigating the Role of Inositol Polyphosphate 4-Phosphatase Type II (INPP4B) Overexpression on Autophagy in Acute Myeloid Leukemia (AML) By Mark Hani Sharobim A thesis submitted in conformity with the requirements for the degree of Master of Science Graduate Department of Pharmacology and Toxicology University of Toronto Copyright by Mark Sharobim 2016
2 Investigating the Role of Inositol Polyphosphate 4-Phosphatase Type II (INPP4B) Overexpression on Autophagy in Acute Myeloid Leukemia (AML) ABSTRACT Mark H. Sharobim Master of Science Department of Pharmacology and Toxicology University of Toronto 2016 BACKGROUND: Inositol polyphosphate-4-phosphatase type-ii (INPP4B) is a lipid phosphatase that dephosphorylates PI(3,4)P2 into PI(3)P. The depletion of PI(3,4)P2 prevents aberrant AKT signalling. Therefore, INPP4B is classically known as a tumour suppressor. Recent studies however demonstrate INPP4B overexpression may promote cancer generation and progression. We investigated effects of INPP4B overexpression in acute myeloid leukemia (AML). METHODS: AML cell lines that overexpress wild-type (INPP4B wt ) and mutant (INPP4B mut ) INPP4B were used to determine what phenotypes are governed by its phosphatase activity. RESULTS: Overexpression of wild-type INPP4B conferred advantages in growth, chemoresistance and colony forming assays, suggesting a role for INPP4B opposite to that of a tumour suppressor. These phenotypes were either absent or greatly diminished in INPP4B mut cells. INPP4B wt cells also had increased accumulation of autophagosomes with or without autophagy inhibitors when compared to control. CONCLUSION: INPP4B-mediated phenotypes in AML are phosphatase-dependent and this is subsequently associated with increased potential to undergo autophagy. ii Word Count: 150/150
3 ACKNOWLEDGEMENTS It would be remiss of me to acknowledge anyone prior to my Lord Jesus Christ and His help throughout my entire graduate degree. I couldn t have done it without Him and His constant love. My family is a rock in my life that has kept me steady throughout the tough times and pushed me through obstacles I felt I could not overcome. It is with their support, guidance and love that I am where I am today. My dad is an example of integrity and virtue and not only do I learn from his actions everyday, the lessons I have learned from him I will keep close to my heart forever. My mother is the definition of a role-model and an exemplar for how I wish to carry myself in my personal and professional life throughout my years. My sisters are the greatest gift I could ever ask for and the relationship I have with them is something I will cherish forever. Their company keeps me cheerful throughout the tough times. My supervisor Dr. Lenny Salmena, PhD is by far and large the greatest inspiration I have in science and as a mentor. Though I was his first student and have technically graduated, I will consider him my mentor and teacher forever. His passion as well as dedication to his work and craft are attributes I aspire to attain one day. His love and enthusiasm for science is something I wish to pass on to those intending to pursue any sort of scientific career. Without him, without a doubt I would not be where I am today. His constant mentoring is something that cannot be understated. He has literally shaped the way I think and tackle science as a whole. Thank you so much Lenny. My lab mates, Martino Gabra, Emily Mangialardi, Anthony To, Meong-Hi Son, Lydia To, Thais Fontanezi-Maciel, Shayne Greenberg, Ayesha Rashid, Erik Dzneladze and John Woolley are a great family that has positively shaped my future forever. The mentoring I received from Dr. Michael Jain, Dr. Rob Laister, Dr. Vuk Stambolic and Ayesha Rashid are little gifts I cherish and I hope someone could be as lucky as me to have been able to pick their brains the past 3 years. I want to thank all those in the lab that taught me that science is an exercise in thinking. It is an open garden of creativity and is conducive to forming bright ideas, innovations and concepts. It is this I am most thankful that I learned while pursuing my degree, and a skill that could not have been cultivated elsewhere. iii
4 CONTRIBUTIONS Martino Gabra treated all the cells with daunorubicin and counted their viability (Figure 11). Emily Mangialardi created the immortalized Inpp4b knockout and wild type MEF cell lines (Figure 16). Anthony To helped with and performed western blots (Figures 15-17). Dr. Meong Hi Son did the colony forming cell experiment (Figure 10). Dr. John Woolley did the low serum viability counts (Figure 9B). iv
5 Table of Contents Abstract... ii Acknowledgements... iii Contributions... iv Table of Contents... v List of Abbreviations... vii List of Figures... x List of Tables... xi List of Appendices... xii 1. Introduction Inositol Phospholipids Phosphoinositide Signalling Overview Phosphoinositide Signalling in Cancer The PI3K Pathway Antagonizing PI3K signalling Inositol Polyphosphate 4-Phosphatase, Type II (INPP4B) Discovery Structure Functions INPP4B and Cancer INPP4B is a Tumour Suppressor INPP4B Overexpression in Cancer Acute Myeloid Leukemia Overview The PI3K pathway in AML Autophagy Overview Macroautophagy The PI3K-mTOR Pathway and Autophagy Autophagy and Cancer: a Double Edged Sword? Rationale, Aims and Hypothesis Rationale and Aims v
6 2.1.1 Investigating mechanisms of INPP4B-mediated phenotypes in AML cells INPP4B Overexpression and PI(3)P signalling: Implications for Autophagy Hypothesis Materials and Methods Cell Culture Lentivirus production DNA plasmids Methylcellulose Colony Formation Cell (CFC) Assay Phosphatase Assay Immunoblotting Cyto-ID assay Generation of Immortalized MEFs Genotyping Statistics Results Generation of a phosphatase-null INPP4B mutant (INPP4B mut ) protein INPP4B mut OCI-AML2 cells do not exhibit phenotypes observed in INPP4B wt overexpressing cells INPP4B wt Overexpressing Cells Have Increased Staining of Autophagosomes in an Untreated Condition INPP4B wt cells demonstrate increased autophagosome accumulation in the presence of inhibitors of autolysosomal acidification INPP4B wt NB4 cells demonstrate increased autophagosome accumulation when treated with chloroquine Inpp4b -/- MEFs have a reduced ability to undergo autophagy Discussion Summary of Results INPP4B Phosphatase Activity INPP4B and Autophagy Conclusions Future Directions References Appendices Publications and abstracts vi
7 LIST OF ABBREVIATIONS AKT V-akt murine thymoma viral oncogene homolog 1 or Protein Kinase B ALL Acute lymphoblastic leukemia or acute lymphocytic leukemia AML Acute myeloid leukemia or acute myelogenous leukemia AMPK AMP-activated protein kinase APL Acute promyelocytic leukemia Ara-C Cytarabine As2O3 Arsenic trioxide Atg7 Autophagy related protein 7 Atg8 Autophagy related protein 8 (yeast) or LC3A/B/C (human) ATRA All-trans retinoic acid BCR/ABL1 Philadelphia translocation fusion protein, Breakpoint cluster region (BCR)/ Abelson murine leukemia viral oncogene homolog 1 (ABL1) Beclin 1 or Atg6 Beclin 1 (human) or Autophagy related protein 6 (yeast) BRCA1 Breast cancer 1, early onset gene C. elegans Caenorhabditis elegans CDP-DAG Cytidine-diphosphate diacylglycerol CFC Colony forming cell c-kit Tyrosine-protein kinase Kit or CD117 Class III PI-3K VPS34 CMA Chaperone mediated autophagy CML Chronic myeloid leukemia or chronic myelogenous leukemia CR Complete remission DAG Diacyleglycerol DFCP1 Double FYVE-domain containing protein 1 EFS Event free survival EGF Epidermal growth factor ER Endoplasmic Reticulum ERec Estrogen receptor FLT3 Fms-like tyrosine kinase 3 FLT3-ITD Fms-like tyrosine kinase 3 internal tandem duplication FOXO3A Forkhead box O3 FYVE Fab1p, YOTB, Vac1p and Early Endosome Antigen 1 GPCRs G protein couple receptors HER2 Human epidermal growth factor receptor 2 HIF-1α and HIF-1β Hypoxia-inducible factor 1-alpha and beta HMECs Human mammary epithelial cells HSC Haematopoietic stem cell HSPCs Haematopoietic stem and progenitor cells INPP4A Inositol polyphosphate 4-phosphatase type I INPP4B Inositol polyphosphate 4-phosphatase type II INPP4B mut Denotes overexpression of mutant full length INPP4B protein INPP4B wt Denotes overexpression of wild-type full length INPP4B protein INPP5A-J Inositol polyphosphate 5-phosphatases Ins(1,3)P2 Inositol 1,3-bisphosphate vii
8 Ins(1,3,4)P3 Inositol 1,3,4-trisphosphate Ins(1,4,5)P3 or IP3 Inositol-1,4,5-trisphosphate Ins(3)P Inositol 3-phosphate IRS1 Insulin receptor substrate 1 kbp Kilo base pairs KFERQ Lys-Phe-Glu-Arg-Gln KNMT2A or MLL Histone-lysine N-methyltransfeRASe 2A KRAS Kirsten rat sarcoma virus oncogene LAMP2A Lysosome-associated membrane protein 2 LFS Leukemia-free survival LOH Loss of heterozygosity LSC Leukemic stem cell LSK cells Lin - Sca-1 + c-kit + progenitor cells MAP1LC3A/B/C or LC3A/B/C Microtubule-associated protein light chain 3 A/B/C (human) or Atg8 (yeast) MAPK Mitogen activated protein kinase MEFs Mouse embryonic fibroblasts mirna microrna mtor Mammalian target of rapamycin or mechanistic target of rapamycin mtorc1 mtor complex 1 mtorc2 mtor complex 2 NHR2 Nervy-homology 2 NPC Nasopharyngeal carcinoma NPM1 Nucleophosmin NRAS Neuroblastoma RAS Viral (V-RAS) Oncogene NSG NOD scid gamma OS Overrall Survival p53 Tumor protein p53 PA Phosphatidic acid p-akt or phospho-akt Phosphorylated-AKT, otherwise activated AKT PDPK1 or PDK1 3-phosphoinositide dependent protein kinase-1 PE Phosphatidylethanolamine PEST Proline, glutamate/aspartate, serine/threonine rich PH Pleckstrin homology phospho-sgk3 or p-sgk3 Phosphorylated-SGK3, otherwise activated SGK3 PHTS PTEN hamartoma tumor syndrome PI(3)P Phosphatidylinositol-3-phosphate PI(3,4)P2 Phosphatidylinositol (3,4)-bisphosphate PI(3,4,5)P3 or PIP3 Phosphatidylinositol (3,4,5)-trisphosphate PI(4)P Phosphatidylinositol-4-phosphate PI(4,5)P2 Phosphatidylinositol-4,5-bisphosphate PI(5)P Phosphatidylinositol-5-phosphate PI3K Phosphatidylinositol 3-kinase PI3K C2α Class II PI3K alpha PIKK PI3K-related protein kinase PIs Phosphatidylinositols viii
9 PIS Phosphatidylinositol synthase PKB Protein kinase B or AKT PKC Protein kinase C PLC Phospholipase C PML-RARα promyelocytic leukemia gene (PML) - retinoic acid receptor α (RARα) fusion protein PEG Polyethylene glycol PTEN Phosphatase and tensin homolog deleted on chromosome 10 PTP Protein tyrosine phosphatase PX Phox homology RAS Rat sarcoma virus oncogene RHEB RAS homolog enriched in brain RNase A Ribonuclease A RNase-S peptide Ribonuclease S peptide RTKs Receptor tyrosine kinases SGK3 Serine/threonine-protein kinase 3 SH2 Src homology 2 SHIP1 or INPP5D SH2-containing inositol 5-phosphatase 1 SHIP2 or INPPL1 SH2-containing inositol 5-phosphatase 2 shrna Small or short hairpin RNA SNPs Single nucleotide polymorphisms TAPP1 Tandem PH domain-containing protein 1 TAPP2 Tandem PH domain-containing protein 2 TSC 1/2 Tuberous sclerosis complex 1 and 2 ULK1 Unc-51 like autophagy activating kinase 1 ULK2 Unc-51 like autophagy activating kinase 2 VPS34 or hvps34 (human) Class III PI3K ix
10 LIST OF FIGURES Figure 1. Schematic of a Phosphatidylinositol Figure 2. Seven phosphoinositides found in cells, derived from phosphatidylinositol (PIs) Figure 3. Schematic depicting main INPP4B function Figure 4. Both the Substrate (PI(3,4)P2) and Product (PI(3)P) of INPP4B Function Activate Similar Kinases Figure 5. Overview of Autophagic Flux Figure 6. AKT activation results in indirect stimulation of mtor and inhibition of autophagy. 38 Figure 7. Sequencing Data Encoding INPP4B wt and INPP4B mut phosphatase domains Figure 8. Characterization of OCI-AML2 cells expressing vector control and INPP4B wt and INPP4B mut proteins Figure 9. INPP4B wt cells proliferate more rapidly than control or mutant cells in normal and low serum conditions Figure 10. INPP4B wt cells form colonies preferentially in vitro Figure 11. INPP4B wt cells are more resistant to daunorubicin in vitro Figure 12 INPP4B expression in OCI-AML3 cells leads to a higher level of autophagosomes in an unstimulated condition Figure 13. OCI-AML3 INPP4B wt have a greater propensity for autophagosome biogenesis Figure 14. Expression of INPP4B in OCI-AML2 increases amount of autophagosome accumulation Figure 15. Dose dependent effect of chloroquine in INPP4B wt NB4 cells Figure 16. Inpp4b -/- MEFs have a reduction in their ability to accumulate autophagosomes in a basal state x
11 LIST OF TABLES Table 1. Summary of INPP4B roles and functions in select cancers xi
12 LIST OF APPENDICES Appendix 1. INPP4B high AML patients have lower CR rates and shorter survival. (Taken from Dzneladze et al. 2015, Leukemia) Appendix 2. INPP4B high constitutes a significant hazard in total and CN-AML (Taken from Dzneladze et al. 2015, Leukemia) Appendix 3. Ectopic overexpression of INPP4B in AML cells leads to increased colony-forming potential and proliferation. (Taken from Dzneladze et al. 2015, Leukemia) Appendix 4. INPP4B overexpression is associated with resistance to chemotherapy and ionizing radiation. (Taken from Dzneladze et al. 2015, Leukemia) xii
13 1. INTRODUCTION 1.1 Inositol Phospholipids Phosphatidylinositols (PIs) are a family of membrane bound phospholipids found in all eukaryotic cells 1. Structurally, PIs consist of two fatty acids and a myo-inositol ring linked via phosphate group to a glycerol backbone (Figure 1). PIs are synthesized from cytidinediphosphate diacylglycerol (CDP-DAG) and myo-inositol by the action of phosphatidylinositol synthase (PIS) 2 and generally comprise 10-20% of all cellular phospholipids 3. Phosphatidylinositol kinases and phosphatidylinositol phosphatases tightly control the relative abundance of different PIs by the addition or removal of phosphate groups. PIs can be reversibly phosphorylated at the hydroxyl groups on positions 3, 4 and 5 of the inositol ring, thus generating seven distinct PI-derivatives in addition to PI itself (Figure 2). Such variable phosphorylation explains how PIs serve a wide range of cellular functions. Despite seemingly little difference between PI isoforms (i.e. only a slightly shifted phosphate between monophosphorylated PIs) PI-binding domains can distinguish PIs with varying degrees of specificity 4. Thus, proteins can be specifically "recruited" to intracellular membranes where PIs are located. For example, pleckstrin homology domains can bind the second messengers phosphatidylinositol (3,4,5)-trisphosphate (PI(3,4,5)P3 or PIP3) and phosphatidylinositol (3,4)- bisphosphate (PI(3,4)P2) with high specificity (discussed below) 5. Proteins such as protein kinase B (PKB or AKT) can be recruited to the plasma membrane where PIP3 and PI(3,4)P2 are located, they then can be activated and perform downstream functions. 1
14 1.2 Phosphoinositide Signalling Overview PIs are essential to cellular homeostasis, and it has been suggested that phosphoinositide signalling modulates virtually all biological processes 6. Their roles in signal transduction pathways controlling vesicle trafficking (reviewed in 7), DNA replication and repair 8 10, oncogenesis 11, apoptosis 12 and proliferation 13 are well documented. The most notable example is the phenomenon discovered in the 1980s by which phospholipase C (PLC) hydrolyzes membrane bound phosphatidylinositol-4,5-bisphosphate (PI(4,5)P2) to form diacyleglycerol (DAG) and the soluble inositol-1,4,5-trisphosphate (Ins(1,4,5)P3 or IP3) 14. Untethered, IP3 can then diffuse throughout the cytoplasm to bind and activate calcium channels on intracellular organelles, thereby increasing intracellular Ca 2+. DAG and IP3 can also lead to activation of protein kinase C (PKC), and thus subsequent phosphorylation of other cellular molecules. Therefore, although PIs do not constitute a majority of cellular lipids, their effectiveness as second messengers is imperative, as they are often one of the first events in signalling hierarchies. Overall, the importance of PI signalling is illustrated by the many human diseases that manifest because of dysregulation in the normal function of proteins and pathways controlling PI metabolism 15. 2
15 Figure 1. Schematic of a Phosphatidylinositol. Phosphatidylinositols (PIs) contain two fatty acids and a myo-inositol ring linked via phosphate group to a glycerol backbone. They can be phosphorylated at positions D3-D5 of the myo-inositol ring, generating PI derivatives with phosphate groups called phosphoinositides. PIs are generated through the enzymatic action of Phosphatidylinositol synthase (PIS); they are generated from cytidinediphosphate diacylglycerol (CDP-DAG) and myo-inositol 2. They are found on the cytosolic side of all eukaryotic membranes 16. 3
16 Figure 2. Seven phosphoinositides found in cells, derived from phosphatidylinositol (PIs). The bioavailability of each phosphoinositide is tightly controlled through the function of multiple phosphoinositide kinases and phosphatases that can add or remove phosphate groups on each PI. The most abundant phosphoinositide is PI(4,5)P2 6. In total, there are 7 derivatives excluding the non-phosphorylated PI. In terms of total cellular lipid content, PIs are not among the most abundant cellular lipids, but control various signalling pathways 3. This is due to their ability to be recognized by different protein structures and domains. Proteins such as AKT or PDK1 can recognize phosphoinositides with high specificity and accuracy, despite being similar in shape and structure. Thus, although there are for example 3 mono-phosphorylated PIs (PI(3)P, PI(4)P and PI(5)P), the proteins that can distinguish them are involved in very different pathways 4. Figure adapted from Le Roy and Wrana, Nature Reviews Molecular Cell Biology, 2005.(Image in Box 1) 17 4
17 1.3 Phosphoinositide Signalling in Cancer The PI3K Pathway The phosphatidylinositol 3-kinase (PI3K) pathway is a cellular signalling network centered around the activity of the PI3K, AKT and the mammalian target of rapamycin (mtor) proteins. PI3Ks are a group of evolutionarily conserved lipid kinases that phosphorylate the hydroxyl group at the D3 position of PIs 18. The preferred substrate for PI3K is PI(4,5)P2, but it is also able to phosphorylate phosphatidylinositol-5-phosphate (PI(5)P), phosphatidylinositol-4- phosphate (PI(4)P) and PI to some capacity 19. There are three main classes of PI3K: Class I, II and III, each largely differing based on primary structure and substrate specificity. Class I PI3Ks are further subdivided into types IA and IB, depending on whether they are activated by receptor tyrosine kinases (RTKs) or G protein couple receptors (GPCRs), respectively. Class I PI3Ks are heterodimers of two subunits, a functional p110 subunit and a p85 regulatory subunit, each of which have three isoforms 20. The main product of class I PI3K activity is PIP3 through the phosphorylation of PI(4,5)P2. PIP3 is the only triphosphorylated phosphoinositide and is found predominantly on the plasma membrane 21. PI3Ks are activated through stimulation of RTKs or GPCRs. To describe this briefly, when a ligand binds its matching receptor, the receptor dimerizes and an autophosphorylation event occurs on tyrosine residues on the cytosolic side of the receptor. Phosphorylated tyrosine residues create binding sites for adaptor proteins with Src homology 2 (SH2) protein domains such as insulin receptor substrate 1 (IRS1) or PI3K itself, allowing for increased PI3K activity 22. While basal levels of PIP3 and PI(3,4)P2 (generated from phosphatidylinositol-4-phosphate, PI(4)P) are low, receptor-ligand binding (e.g. epidermal growth factor (EGF) binding RTK) 5
18 results in elevated class I PI3K activation, thus causing both PIs to accumulate at the plasma membrane. Although all classes of PI3K are known to phosphorylate the D3 position of phosphoinositides, Class II PI3Ks have affinity for PI and PI(4)P, thus generating PI(3)P and PI(3,4)P2 respectively, while Class III PI3Ks (VPS34 or hvps34 in humans) generate PI(3)P from PI. It is possible that the products generated by each PI3K may differ based on classspecific localization in the cell (that is, some intracellular membranes may be enriched with a specific 3-phosphoinositide, such as endosomes and PI(3)P, which are accounted for by VPS34 on these structures) 23. PH domain containing proteins such as 3-phosphoinositide dependent protein kinase-1 (PDPK1 or PDK1) 24,25 and AKT 26 can bind to PIP3 and PI(3,4)P2, facilitating their recruitment to lipid membranes. Recruitment of AKT to the cell membrane is required for full activation, which is achieved by phosphorylation of two residues, Thr308 and Ser473 on its kinase and regulatory protein domains, respectively 27,28. PDK1 mediates T308 phosphorylation and membrane binding is also required for PDK1 activation of AKT 24. There have been many kinases proposed to mediate the phosphorylation of the S473 residue on AKT, but a recent consensus confirms that it is mediated by the mtor complex 2 (mtorc2) proteins 29. Once activated, the serine/threonine kinase, AKT controls many critical processes such as apoptosis, cell cycle and proliferation through phosphorylation of its numerous substrates 30,31. Recent estimates suggest that AKT has more than 100 substrates of which it can phosphorylate and modulate function 32,33. Hyperactive AKT is a common occurrence in many human cancers, and mouse models have demonstrated that on its own, constitutively active AKT is sufficient for carcinogenesis 34. One of the principal AKT effector proteins is the mtor kinase. mtor is a 289 kda protein that belongs to the PI3K-related protein kinase (PIKK) family 35. mtor is the catalytically active 6
19 member of two functionally unique protein kinase complexes called mtor complex 1 and 2 (mtorc1 and 2). The mtorc complexes differ in the effectiveness by which the macrolide fungicide rapamycin inhibits their activity, with mtorc2 being practically insensitive while mtorc1 exhibits sensitivity. Stimulation of AKT leads to the inhibition of tuberous sclerosis complex 1 and 2 (TSC 1/2) proteins which themselves normally prevent activation of mtorc1 by further inhibition of the GTPase RAS homolog enriched in brain (RHEB), a known activator of mtor (Figure 6). mtorc1 itself receives input from cellular energy sensors (Figure 6), and stimulation causes gene transcription, mrna translation as well as ribosome biogenesis which all together increase cell growth, proliferation and survival. Accordingly, it is desirable to control or prevent dysregulation of AKT in cancer Antagonizing PI3K signalling Phosphatase and tensin homolog deleted on chromosome 10 (PTEN) The primary biological mechanism by which PI3K signalling is negatively regulated is through the activity of phosphatase and tensin homolog deleted on chromosome 10 (PTEN). PTEN exhibits lipid phosphatase activity by dephosphorylating PIP3 at the D3 position of the inositol ring, generating PI(4,5)P2. By decreasing levels of PIP3, AKT activity is curtailed because it can no longer localize to the plasma membrane. PTEN also has a C2 lipid binding domain which localizes it to the plasma membrane, an association that is necessary for its lipid phosphatase activity 36. Germline losses or mutation of PTEN are associated with a myriad of syndromes and disorders including PTEN hamartoma tumor syndrome (PHTS, including but not limited to Cowden syndrome) 37,38 and autism spectrum disorder 39. PTEN is also a bona-fide 7
20 tumour suppressor; almost 50% of all human cancers contain inactive alleles of PTEN. Indeed, it is among the most frequently mutated or lost genes in cancer, and in some malignancies up to 80% of cases harbour some sort of PTEN inactivity SH2-containing inositol 5-phosphatase (SHIP) A large group of proteins that can also hydrolyze PIP3 are the inositol polyphosphate 5- phosphatases, of which there are 10 isoforms. Although a majority of isoforms exhibit D5 phosphatase activity on multiple PIs, the product generated when these phosphatases hydrolyze PIP3 is PI(3,4)P2. One such phosphatase, SH2-containing inositol 5-phosphatase 1 (SHIP1), is a 5-phosphatase encoded by the INPP5D gene. Its expression is mainly restricted to hematopoietic tissues while a closely related SHIP2 isoform is more widely expressed 41,42. It has been proposed that increased 5-phosphatase activity may lead to elevated or sustained AKT signalling, since like PIP3, PI(3,4)P2 can also bind the PH domain on AKT. Other studies demonstrate the opposite; that SHIP phosphatases suppress AKT activity This may be explained by the stability or differential binding of AKT to PIP3 or PI(3,4)P2. There are some studies showing SHIP proteins to be inactivated in cancer; for example, mice with B cell specific downregulation of SHIP1 develop acute lymphoblastic leukemia (ALL) 46 and ALL cell lines as well as primary tumour samples exhibit SHIP1 inactivation or loss 47. While its role in cancer may be attributed to the notion that many 5-phosphatases certainly terminate PI3K/AKT signalling, only one study has implicated a 5-phosphatase as a real tumour suppressor at the in vitro and in vivo level in cancer 48. In summary, through increased 5-phosphatase activity, there is accumulation of the preferred substrate for 4-phosphatases, the biphosphorylated PI(3,4)P2. 8
21 1.4 Inositol Polyphosphate 4-Phosphatase, Type II (INPP4B) Discovery The first evidence that there was enzymatic degradation of the D4 phosphate found on the inositol ring on radiolabelled inositol phospholipids was reported in 1987 by Inhorn et al. using crude tissue extracts of calf brain. In their study, they noticed that inositol 3,4-bisphosphate (Ins(3,4)P2) is converted to the soluble inositol 3-phosphate (Ins(3)P) 49. In the same year, it was discovered that inositol 1,3,4-trisphosphate (Ins(1,3,4)P3) can be dephosphorylated to inositol 1,3-bisphosphate (Ins(1,3)P2) using a similar method 50. The term inositol polyphosphate 4- phosphatase was the term used to describe the enzyme that performed this function, and it was isolated and characterized in Later purified from calf brain tissue extract, it was identified as a Mg 2+ independent 105 kda protein with affinity for both Ins(1,3,4)P3 and Ins(3,4)P2 51. Notably, both Ins(1,3,4)P3 and Ins(3,4)P2 are soluble inositol lipids, and that only in 1994 was it discovered that there was an enzyme capable of dephosphorylating the membrane bound PI(3,4)P2. In agreement with current knowledge, the purified inositol polyphosphate 4- phoshphatase had a greater affinity for PI(3,4)P2 than Ins(1,3,4)P3 or Ins(3,4)P2 52. cdna encoding the inositol polyphosphate 4-phosphatase enzymes revealed two isoforms, a type I and type II (also INPP4A and B, respectively) which share 37% amino acid sequence identity. With both having varying tissue specific expression levels, it is now understood the type I and type II isoforms are located at different genetic loci. INPP4A is highly expressed in brain tissue, and while INPP4B has a wider tissue distribution, expression is highest in skeletal muscle and heart 53 56, but is also present in epithelial tissues like the breast 57. There was also evidence that the type II isoenzyme was subject to alternative splicing, thus further 9
22 subdividing both types into α and β splice variants. The murine Inpp4bα isoform also has high expression in haematopoietic lineages, specifically in natural killer (NK) as well as mast cells and to a lesser degree in B-lymphocytes Structure The INPP4B gene is located on the long arm of chromosome 4, at 4q The coding exons span approximately 825 kilo base pairs (kbp) and the mrna sequence 4 kbp. The primary mrna transcript codes for a protein of 105 kda consisting of 924 amino acids with 3 conserved protein domains 55,60. INPP4B contains a N-terminal C2 lipid binding domain that is conserved among similar lipid binding proteins 61. The INPP4B C2-domain is said to have highest affinity for phosphatidic acid (PA, a major constituent of lipid membranes) and PIP3 58. INPP4B also has a central hydrophobic protein motif called the Nervy Homology 2 (NHR2) and that allows protein to protein interaction and oligomerization, such as is the case for fusion proteins such as AML1/ETO 62. The catalytic phosphatase domain of INPP4B is located in the extreme C-terminus. The INPP4 phosphatases have a conserved catalytically active phosphatase domain and in INPP4B, this is characterized by the amino acid motif CKSAKDR (starting at amino acid 842). This domain is conserved amongst similar acid, protein tyrosine and dual specificity (i.e. lipid and protein) phosphatases, identified by 5 amino acids flanked by cysteine and arginine residues (CX5R in similar proteins). This signature motif brings the oxygen atom on the phosphate into the catalytic pocket. The nucleophilic sulfhydryl group on the cysteine residue attacks the oxygen on the phosphorous atom resulting in an intermediate cysteine-po3 complex and eventually, an inorganic phosphate 63. Substitution of the Cys-842 residue to serine or 10
23 alanine will render the phosphatase activity null 63,64. Replacement of the cysteine residue to serine allows the phosphatase to access the substrate but stabilizes the intermediate complex formed with the phosphate group, preventing release of inorganic phosphate and this overall prevents dephosphorylation from occuring Functions The most well studied biological function of INPP4B is of its ability to mediate the hydrolysis of the D4 phosphate group (i.e. dephosphorylation) found on the inositol head ring of PI(3,4)P2, generating phosphatidylinositol-3-phosphate (PI(3)P), (shown in Figure 3). While PIP3 is most certainly a potent activator of AKT, PI(3,4)P2 has also been shown to regulate AKT activation in vitro 66. Both PI(3,4,5)P3 and PI(3,4)P2 can act as membrane anchors for proteins, however some studies suggests PI(3,4)P2 remains at the plasma membrane longer after receptormediated stimulation of PI3K, and is not subject to rapid degradation similar to PI(3,4,5)P3 67. Therefore prolonged inactivation of INPP4 phosphatases may result in continuous AKT activation. Accordingly, INPP4B has been shown to be lost or inactivated in cancers of the breast 57,68, ovarian 69, thyroid 70 and lung 71. While a role for INPP4B has been implicated in the progression and maintenance of cancer, the same has cannot be said for its paralog, INPP4A. This may be due to their tissue specific expression, as INPP4A expression is mainly restricted to neuronal tissues. Studies suggest it has a role in preventing excitotoxic cell death of neurons 72,73. INPP4A has also been shown to localize to endosomes and can modulate PI3K signalling there as well 74. Although this discussion is mainly focused on the proposed roles of INPP4B in cancer, it may have other biological functions. Inpp4b was identified as a novel regulator of osteoclastogenesis, where it 11
24 represses differentiation of osteoclasts and shows an ability to affect the transcription of osteoclast-specific target genes 75. Single nucleotide polymorphisms (SNPs) in the Inpp4b gene in mice changed nerve conduction velocity, and was associated with multiple sclerosis (MS) in patents 76. Inpp4b was also shown to positively control callosal axon formation in mice
25 Figure 3. Schematic depicting main INPP4B function. INPP4B normally dephosphorylates the D4 phosphate on the inositol ring found on the lipid membrane bound PI(3,4)P2 to generate the mono-phosphorylated PI(3)P. The phosphatase domain is catalyzed by a protein motif CKSAKDR. Substitution of the cysteine to a serine renders the phosphatase catalytically inactive
26 1.5 INPP4B and Cancer INPP4B is a Tumour Suppressor The first indication that INPP4B was implicated in cancer was published by Westbrook et al. in The authors used a shrna library to screen for suppressors of transformation in human mammary epithelial cells (HMECs). INPP4B was one of 8 genes that were lost in 90% of colonies formed as a result of small hairpin (shrna) knockdown 78. In other studies, INPP4B was lost during the transformation of malignant proerythroblast cell line models, and when overexpressed, it regulated phosphorylated-akt (p-akt) levels 79. A tumour suppressive role for INPP4B was later demonstrated by Gewinner et al. and Fedele et al. in breast cancer 57,68. The two groups showed that INPP4B knockdown increases anchorage independent growth, migratory as well as invasive potential and p-akt in multiple breast cancer cell lines. Xenograft models suggested INPP4B overexpression reduces tumour formation, whereas its knockdown mediated the opposite effect in mice. Also, loss of heterozygosity (LOH) at the INPP4B locus is a common feature in breast cancer patients from both studies 57,68. The genetic locus harbouring INPP4B was one of the 2 most frequently deleted regions in tumours of breast cancer patients 80. Along with PTEN, INPP4B is the most commonly lost or mutated gene associated within the PI3K pathway in the basal and human epidermal growth factor receptor 2 (HER2) subtypes of breast cancer 81. Consistent with these studies, INPP4B has also been shown to behave as a tumour suppressor in additional epithelial cancers. Loss of INPP4B is associated with poor survival and lymph node metastasis of ovarian cancer patients. Immunohistochemistry also revealed it is more frequently lost than PTEN or Tumor protein 53 (p53) in ovarian cancer tissue microarrays 68,69. 14
27 In prostate cancer, INPP4B expression is driven by androgen receptor and its loss increased activation of AKT. It is absent in primary prostate tumour samples and predicts disease recurrence in patients with rapidly proliferating tumour cells 82,83. Ectopic INPP4B expression in prostate cancer cell lines suppresses their invasive potential in vitro and in vivo. It is also lost in prostate carcinoma epithelium when compared to benign prostate epithelium 83,84. In melanoma, INPP4B is present in primary samples but is lost in tumour tissue, and INPP4B overexpression curtails receptor mediated AKT activation. As in previous studies, ectopic INPP4B expression suppressed tumour size and burden occurring in xenograft mouse models 85. In thyroid cancer, complete or partial Inpp4b knockout causes progression of benign thyroid adenomas found in Pten heterozygous mice into metastatic thyroid tumours. It is lost in thyroid cancer lines and its expression was able to specifically abrogate AKT signalling on endosomes, suggesting location-specific function for this phosphatase 70. In bladder cancer, INPP4B expression is strongly induced by estrogen receptor (ERec) expression and this is associated with inhibition of AKT activation. Knocking down INPP4B reverts this phenotype 86. Evidence of epigenetic inactivation of the INPP4B locus in nasopharyngeal carcinoma (NPC) tumours and cell lines 87 demonstrates there are multiple routes by which a tumour inactivates INPP4B. Taken together, there is a substantial body of work that demonstrates INPP4B is a bonafide tumour suppressor, and that when present, its main enzymatic function is to counteract PI3K/AKT signalling. 15
28 1.5.2 INPP4B Overexpression in Cancer The tumour suppressive characteristics of INPP4B have been well established in multiple cancer types. A number of studies now demonstrate that INPP4B is upregulated in cancer with pathological significance. For example, by analyzing gene expression from radioresistant laryngeal cancer cell lines, INPP4B mrna levels was proposed to be a marker of tumour resistance to radiotherapy. Radiation treatment of laryngeal cancer cell lines also induced expression of INPP4B. De novo INPP4B expression increased resistance to both radiation and chemotherapy drug by evasion of apoptosis 88. In a follow up study, the authors similarly noticed that INPP4B expression is induced by hypoxia, with predicted binding sites for the transcription factors hypoxia-inducible factor 1-alpha and beta (HIF-1α and HIF-1β) found on the INPP4B promoter sequence. Furthermore, the resistance phenotype in cells with high expression of INPP4B is associated with an increased production of glucose, a commonly observed characteristic of drug resistant tumor cells. Finally, up to 70% of their laryngeal cancer tissue samples stained positively for INPP4B 89. Prior to our work, there was little data regarding INPP4B function in leukemias or other blood cancers. One study showed that INPP4B is overexpressed in childhood acute lymphoblastic leukemia (ALL) that is Philadelphia translocation (BCR/ABL1) positive 90. Since INPP4B has never been investigated in acute myeloid leukemia (AML), we first interrogated gene expression data from AML patients to determine what role it may have. Unexpectedly, the transcript was upregulated in approximately 25% of patients across multiple patient gene expression databases. INPP4B high (in the top 25%) was associated with a variety of clinical factors that depict an overall worse disease for those with AML. This signature was accompanied by decreased overall survival (OS) and event free survival (EFS) in patients from all six 16
29 independent data sets. It was also identified that patients with INPP4B high are poor responders to standard chemotherapy used to treat AML (Appendices 1 and 2). In AML, prognostication of the survival risk associated with different subtypes is confirmed by the presence of biomarkers such as cytogenetic abnormalities and adverse molecular events. INPP4B was demonstrated to be an independent prognostic measure of poor disease; with predictability comparable or better than known risk factors such as the presence of Fms-like tyrosine kinase 3 internal tandem duplication (FLT3-ITD) or Nucleophosmin (NPM1) mutations (Appendix 2) 91. Taken together, AML patients with INPP4B transcript overexpression demonstrated a poorer outcome compared to those with low INPP4B transcript levels. We performed a series of in vitro experiments to validate these observations; cdna encoding the full-length wild-type INPP4B transcript was overexpressed in various AML cell lines. We observed that INPP4B overexpressing cell lines (INPP4B wt ) grew faster in full or low serum media, were more resistant to treatment with drug or ionizing radiation and formed colonies preferentially in colony forming cell (CFC) assays (Appendices 3 and 4). These in vitro results recapitulated our clinical findings for INPP4B and AML 91. In an independent study, data published by Rijal et al. reflected these findings where they show that INPP4B protein is upregulated in bone marrow from AML patients when compared to normal based on mass spectrometry. Moreover, INPP4B overexpressing leukemia cells injected into NOD scid gamma (NSG) mice showed greater infiltration into bone marrow compared to controls. Mice treated with the chemotherapeutic agent cytarabine showed chemoresistance under the same conditions 92. Altogether, these two independent studies demonstrate that INPP4B overexpression is associated with more aggressive AML, indicating that there may be a tumour-promoting role for INPP4B in some cancer types. 17
30 Indeed, context specific functions may partially explain differential INPP4B phenotypes (Figure 4). Gasser et al. showed in a study that upregulation of serine/threonine-protein kinase 3 (SGK3) in estrogen receptor (ERec) positive breast cancers may be in part due to INPP4B upregulation. The authors note that INPP4B expression is also induced by ERec 57, and that both have a functional connection. This is because the product of INPP4B catalysis is PI(3)P, and SGK3 (unlike its closely related isoforms, SGK1 and SGK2) contains a phox homology (PX) protein domain that binds PI(3)P which allows it to localize to membranes where it can be activated. Interestingly, SGK isoforms are 55% homologous to the AKT1-3 catalytic domains as well as phosphorylate the same target motif as AKT in effector proteins, and in some cases compensate for oncogenic AKT signalling even when AKT activation is absent 93,94. Gasser and colleagues performed starvation-stimulation experiments in vitro and showed that INPP4B is necessary for SGK3 activation (phospho-sgk3 or p-sgk3) after growth factor stimulation. INPP4B expression was also necessary for SGK3-dependent anchorage independent growth and xenograft tumour formation in nude mice 95. In colon cancer cell lines, Guo and colleagues demonstrated that INPP4B knockdown led to a decrease in SGK3 activation, presumably through a similar mechanism suggested by Gasser et al. in breast cancer 96. However, they also showed that INPP4B overexpression was accompanied by an increase in AKT activation in colon cancer cell lines. This finding appears paradoxical because the tumour suppressor function of INPP4B was reportedly dependent on its ability to decrease AKT activation. This was also the first reported instance that INPP4B upregulation leads to greater activation of AKT. Phenotypically, INPP4B overexpression in colon cancer cell lines led to increased cell proliferation as well as promoted colony forming 18
31 potential. Accordingly, knockdown of INPP4B decreased growth in colon cancer xenografts in nude mice. Although most reports to date describe INPP4B exclusively as a lipid phosphatase, Guo et al. suggest that INPP4B may also possess protein tyrosine phosphatase (PTP) activity 64,97. They report that this oncogenic regulator phenotype can be explained by PTP function of INPP4B on the tumour suppressor PTEN. This is demonstrated through experiments where overexpression of INPP4B downregulates PTEN expression levels, and simultaneously increases PI(3,4,5)P3 levels. This finding was corroborated in vitro where INPP4B was able to dephosphorylate total and immunoprecipitated PTEN 97. Therefore, the proposed mechanism for this phenomenon is that INPP4B dephosphorylates PTEN, thereby destabilizing it and making it unable to control AKT activation. It is generally understood however that phosphorylation of PTEN inhibits enzymatic activity 98 and dephosphorylation would activate it, notwithstanding in a majority of cases phosphorylation of PTEN has been shown to increase its half-life, albeit without preserving enzymatic acitivty 99. It is important to note that the PTEN protein sequence contains over 10 residues that are known to be phosphorylated 100, and in some cases phosphorylation could activate enzyme activity. The phosphorylation sites on PTEN proposed to be controlled by INPP4B in the study by Guo and colleagues were those of Serine 380 and 385, as well as Threonine 382 and 383 (Ser380-Ser385 cluster), all of which are found in the tail region of PTEN. Nonetheless, phosphorylation of this cluster is known to result in increased protein stability but not enzymatic activity 100. Taken altogether, this raises the possibility of another paradox, and the potential that INPP4B mediated regulation of PTEN is different than that by other proteins and post translational modifications. 19
32 It was previously mentioned that INPP4B functions as a tumour suppressor in melanoma. Recently, a study emerged that shows in a subset of melanomas it may function oppositely. The authors found that it was upregulated in primary and metastatic melanoma. In contrast to other findings, they observed that INPP4B expression did not correlate with any change in AKT activation. Instead, they showed that INPP4B upregulation is associated with SGK3 activation, and that this drove enhanced proliferation and clonogenic growth 101. In summary, there is growing evidence that demonstrates INPP4B overexpression promotes cellular phenotypes associated with more aggressive or worse cancer. This is in direct contrast to its previously cemented role as a tumour suppressor. It is especially interesting that in two cases, INPP4B was published to have both roles in the same cancer, though different subsets within the same cancer may explain this duality. A summary of proposed INPP4B roles in various cancers is found in Table 1. This warrants further investigation into what sways the role of INPP4B one way or another. More importantly, it demonstrates the importance of accounting for context in the analysis of signalling pathways implicated in different cancers. 20
33 A B C Figure 4. Both the Substrate (PI(3,4)P2) and Product (PI(3)P) of INPP4B Function Activate Similar Kinases. (A) Like the triphosphorylated PI(3,4,5)P3, the biphosphorylated PI(3,4)P2 can be bound by AKT, thus anchoring it to lipid membranes. This brings AKT into the proximity of the kinases that can phosphorylate it. Phosphorylation of AKT is essential to its kinase activity. When active, it can activate or deactivate effector proteins, thus accounting for several AKTmediated phenotypes. Aberrant activation of AKT is conducive to many of the characteristics cells acquire in becoming carcinogenic 34. INPP4B acts to prevent this by depleting the AKT activator, PI(3,4)P2. The figure in 5A is adapted from Bunney and Katan. Nature Reviews Cancer, (B) Like AKT, PDK1 can bind membrane bound phosphoinositides such as PI(3,4,5)P3 and PI(3,4)P2. PDK1 phosphorylates AKT at T308 and the mtorc2 complex phosphorylates AKT at S473. Also shown are some of the proteins whose activity is modulated by active AKT. The figure is 5B is adapted from Manning and Cantley, Cell (C) SGK3 is a kinase with ~55% sequence identity to AKT 94. It also is activated by PDK1. SGK3 is unique when compared to AKT and even its closely related SGK1 and SGK2 isoforms in that it has a Phox homology (PX) domain that can selectively bind PI(3)P at lipid membranes (shown here at the endosome). This binding is similar to AKT binding of PI(3,4,5)P3 and PI(3,4)P2. AKT and SGK3 share common target proteins and SGK3 can often compensate for the absence of AKT signalling 93. Therefore, in one instance, INPP4B may act to halt AKT activation (5A+B), in another, it may act to indirectly activate AKT-like phenotypes through the stimulation of the closely related SGK3. The figure in 4C is adapted from the record accessed from mutagenetix.utsouthwestern.edu
34 Table 1. Summary of INPP4B roles and functions in select cancers. Cancer Breast cancer, general 57,68 Breast cancer, ERec-positive 96 Ovarian Cancer 68,69 Colon Cancer 97 Prostate Cancer 82,103,104 Potential Role for INPP4B Tumour suppressor Cancer promoting, Driver of PI3K/AKT signalling Tumour suppressor Oncogenic Regulator, Tumour promoting Tumour suppressor Description of INPP4B role, phenotypes and mechanism - INPP4B loss increases AKT activation, INPP4B overexpression decreases AKT activation - INPP4B promotes anchorage independent growth - Overexpression of INPP4B decreases tumour formation in xenograft mouse models - LOH at INPP4B genetic locus found in breast cancer patients - ERec drives INPP4B expression - Accumulation of PI(3)P as a result of INPP4B upregulation leads to SGK3 activation - Loss of INPP4B is associated with poor survival and lymph node metastasis in patients. - It is more frequently lost than PTEN or Tumor protein 53 (p53) in ovarian cancer tissue microarray. - INPP4B knockdown led to a decrease in SGK3 activation. - INPP4B overexpression resulted in AKT - knockdown of INPP4B decreased growth in colon cancer xenografts in mice - INPP4B downregulates PTEN expression levels through dephosphorylation of PTEN - INPP4B loss increased activation of AKT - It is lost in primary prostate tumour samples - INPP4B expression suppressor invasive potential of prostate cancer cells Laryngeal Cancer 88,105 Promotes tumour resistance - Radiation induced expression of INPP4B in laryngeal cancer cell lines - De novo INPP4B expression increased resistance to both radiation and chemotherapy Melanoma 85 Tumour suppressor - INPP4B is present in primary samples but is lost in tumour tissue, - INPP4B overexpression prevents AKT activation. - INPP4B expression suppressed tumour size and burden occurring in xenograft mouse models 22
35 Melanoma, primary and metastatic 106 Oncogenic Driver, Upregulated - INPP4B expression does not affect AKT activation status - INPP4B is associated with activation of SGK3 and this causes enhanced cell growth in vitro Thyroid 70 Tumour suppressor - Inpp4b knockout causes progression of benign thyroid adenomas found in Pten +/- mice. - Specifically abrogate AKT signalling on endosomes Leukemia, acute myeloid 92,107 Biomarker of aggressive AML, overexpression phenotypes indicative of worse disease - Subset of patients with high levels of INPP4B survive less, respond less favorably to chemotherapy - Overexpression of INPP4B increases proliferation and chemoresistance Nasopharyngeal carcinoma 87 Tumour suppressor - Epigenetic inactivation of INPP4B in tumours and cell lines Bladder 86 Tumour suppressor - ERec expression is associated with high levels of INPP4B mrna - INPP4B knockdown increases AKT activation 23
36 1.6 Acute Myeloid Leukemia Overview Acute myeloid leukemia (AML) is an aggressive blood cell cancer identified as invasion primarily of the bone marrow and blood by irregularly differentiated cells of the haematopoietic hierarchy. Clonal expansion of these malignant and functionally immature myeloblasts or blasts hinders normal hematopoietic function leading to recurrent infections, bleeding or organ failure and rapid fatality if left untreated 108. AML is diagnosed by morphological assessment of primary tumour blasts in addition to expression of cell surface or cytoplasmic markers and the presence of cytogenetic abnormalities from the tumour 109. AML is the most common leukemia in adults with the median age of diagnosis being 67 years. The prognosis for AML worsens with age and chromosomal abnormalities commonly found in the elderly complicate treatment 109,110. In younger populations, AML does not account for a majority of cancer diagnoses (i.e. only less than 3% incidence in young persons), however in some estimates it is the leading cause of death in children as well as those under the age of Standard treatment for AML is composed of induction chemotherapy, most typically the 7+3 regimen in which 3 days of anthracycline (e.g. daunorubicin, idarubicin or mitoxantrone) simultaneous with 7 days of cytarabine (Ara-C) are given intravenously. These achieve complete remission (CR) in up to 60 to 80% of younger adults. There has been no other intervention that has consistently been shown to perform better Anthracycline antibiotics like daunorubicin cause cell death in various ways. Some of these include intercalation into DNA with subsequent induction of DNA damage along with interruption of synthesis and replication, as well as 24
37 interfering in the ability of DNA to unwind and the inhibition of topoisomerase II 115. Topisomerase II is an enzyme that actively relaxes supercoiled DNA and affects its topology. It introduces transient double stranded breaks as it is bound to DNA in an intermediate that allows other DNA duplexes to pass through 116. Topoisomerase poisons such as daunorubicin stabilize this intermediate complex and introduce permanent double stranded breaks 117, a process that turns on DNA damage response signalling and apoptosis 118. Despite the efficacy of these drugs, an overwhelming majority of patients relapse, as 40-70% do not achieve long term survival after treatment; indeed, 5 year survival is less than 50% in adults and even worse for the elderly 119,120. It is therefore necessary to know the changes in molecular signalling pathways that may account for the resistance to standard therapy often seen in AML The PI3K pathway in AML Signalling pathways that control proliferation and survival are often altered in AML, especially in the blood where cells undergo constant growth, renewal and differentiation. It is widely recognized that the haematopoietic system is organized as a hierarchy of cells all differentiating from a common progenitor, the haematopoietic stem cell (HSC). HSCs are a rare group of cells that are capable of self-renewal, and have the ability to give rise to all the mature cells found in the blood. It has been suggested that a similar paradigm exists in AML, where the leukemic stem cell (LSC) performs a similar function, and by constantly evading therapy perhaps due to inherent resistance mechanisms, they continuously contribute to a pool of leukemic blasts in a tumour that is often clonally derived 121. It follows then that aberrant signalling through multiple cellular pathways allows the maintenance and propagation of these cell types. 25
38 Both primary and cell line AML samples commonly display constitutive PI3K/AKT signal activation 122,123. These over amplified signals also predict a poor prognosis clinically 124. Like in many other cancers, characterizing and targeting the PI3K pathway in leukemias is of great importance. In some instances, leukemic cells are preferentially targeted by pharmacological inhibition of the PI3K pathway 125. Inhibitors that target multiple proteins in this pathway are more effective then single agents 126. This presents an appealing therapeutic opportunity for those with AML. As even constitutive activation of the further downstream mtorc1 can sometimes be found in all cases of AML, modulating this may be beneficial across multiple cohorts of patients. Despite this however, there are little to no reports of large groups of patients harbouring activating mutations in the catalytic isoforms of PI3K or AKT proteins 127. Through mutations in other proteins upstream of the PI3K and related proteins, there are multiple ways for this network continuously active in AML. For example, receptor tyrosine kinases that are frequently over activated in AML due to mutation (especially in cytogenetically normal AML), such as FLT3 and c-kit (tyrosine-protein kinase Kit or CD117; the former of which is mutated in up to 30% of AML and as well in other leukemias), lead to PI3K activation. This is exacerbated by the fact that not only do patients with FLT3 mutation have shorter time to relapse, their overall survival is among the worst for those with AML 128. FLT3 mutations may directly activate PI3K, but also indirectly through activation of the RAS signalling network, thus contributing further to constant oncogenic PI3K signalling. As a proof of principle, this is seen in vitro where mutant PI3K drives factor independent growth of haematopoietic cells, and in vivo, mutant PI3K can drive leukemic transformation in sublethally irradiated mice. Interestingly, mutant c-kit receptor had stronger transformative potential in vivo, suggesting additional signalling effects beyond PI3K. 26
39 Although to a lesser degree, mutations of the small GTPase family called RAS tell a similar story in some subtypes of AML. In cases with chromosomal rearrangements of Histonelysine N-methyltransferase 2A (KNMT2A, also known as mixed lineage leukemia or MLL), NRAS and KRAS mutations are both seen in up to a quarter of patients, and up to 50% had mutations in some type of protein that is part of this pathway 129. In other studies, similar mutations in components of this pathway were as high as 45%. Multiple RAS mutations were detected in the same leukemia, also at relapse 130. This potentially speaks to the importance of RAS-MAPK, as well PI3K signalling in the clonogenic potential of AML blasts. In MLL rearranged cells, mitogen activated protein kinase (MAPK) inhibition reduced cell survival, without exhibiting the same effect in normal control, suggesting this pathway may be selectively activated in MLL positive leukemia. Interestingly, a small subpopulation of therapy-resistant cells had sustained PI3K pathway activation, experimentally demonstrating an "escape route" for cells and delineating the complexity of the signalling pathophysiology involved 131. Since the manner in which the PI3K pathway is dysregulated in the pathogenesis of blood cancers is clearly distinct than in solid tumours, there is a great need to understand resistance mechanisms to conventional PI3K inhibition. 27
40 1.7 Autophagy Overview Autophagy (literally, "self-eating") is the term used to describe the various ways cellular cytoplasmic contents are delivered to lysosomes for their degradation. To date, three types of autophagy have been identified; they are microautophagy, macroautophagy and chaperonemediated autophagy 132 (Figure 5). Both macro- and microautophagy can be selective or nonselective, they differ however based on how autophagic contents reach the lysosome 133. Microautophagy is the direct engulfment of contents in the cytoplasm by the lysosome (or homologous proteins in other organsims, (e.g. vacuoles) 134. In this type of autophagy, the lysosome itself invaginates cytoplasmic cargo for degradation. Although this has not been studied extensively, it has been reported to occur in mammalian cells such as mouse/rat liver hepatocytes. Some hypothesize this lesser known type of autophagy might be required for basal turnover of long lived proteins 135. In both yeast and mammalian cells, microautophagy has been observed to be both constitutively active and stress induced via nutrient deprivation 136. Chaperone mediated autophagy (CMA) is only a selective form of autophagy wherein soluble cytosolic proteins labelled for degradation by specific peptide motifs are shuttled to the lysosome. This direct shuttling does not required the formation of additional vesicles, and proteins directly traverse the lysosomal membrane from the surrounding cytosol 137. These proteins are delivered by the heat shock protein hsc70 and then to the lysosome by binding to Lysosome-associated membrane protein 2 (LAMP2A) 137. Proteins degraded by CMA frequently have an amino acid sequence that is biochemically related to the Lys-Phe-Glu-Arg-Gln (KFERQ) sequence initially found in ribonuclease S peptide (RNase-S peptide). RNase-S 28
41 peptide was identified as a cleavage product of ribonuclease A (RNase A) protein that is selectively degraded by serum withdrawal in human fibroblasts 138. Thus, chaperone mediated autophagy is carried out by recognition of a motif similar to KFERQ in cytosolic proteins. It is estimated that 25-30% of cytosolic proteins have a similar sequence 139,140. The third and most well studied type of autophagy is referred to as macroautophagy Macroautophagy Macroautophagy (herein referred to as autophagy) is a cellular process by which bulk degradation of cytoplasmic contents occurs through their engulfment in double membraned vesicles termed autophagosomes. The subsequent fusion of autophagosomes with lysosomes (termed autolysosomes) ensures the degradation of cargo (Figure 5). Autophagy can degrade long-lived proteins, cytoplasmic aggregates, mitochondria, peroxisomes, ribosomes, macromolecules and even invasive pathogens 141. Starvation conditions can induce autophagy beyond a basal level 142, as autophagy is thought of as a cellular and energy recycling program, or in other words a way to conserve energy during times of cellular stress by degrading present cellular components into natural "building blocks". Current studies suggest that there are multiple sites at which autophagosomes may form, among which are the endoplasmic reticulum (ER), mitochondria and the plasma membrane. At these locations, pre-autophagosomal structures called phagophores or isolation membranes are where the budding of double membrane vesicles begin 143. Biogenesis of autophagosomes are tightly associated with the presence of PI(3)P at these pre-autophagosomal structures. Many proteins involved in autophagy initiation and maintenance contain protein domains that can bind PI(3)P with high affinity and thus selectively localize them to areas in the cell where 29
42 autophagosomes can form. One of these is the FYVE domain, the name arises from the first four proteins found to have this motif (Fab1p, YOTB, Vac1p and Early Endosome Antigen 1) 144. Cellular PI(3)P is mainly produced by Class III PI-3K (VPS34) and proteins that bind to PI(3)P are involved in membrane trafficking, for example, from endosomes to Golgi bodies to lysosomes. In one documented instance, double FYVE-domain containing protein 1 (DFCP1) binds PI(3)P in a VPS34 dependent manner and forms double membraned structures called omegasomes (i.e. they are physically shaped like the Greek letter omega, Ω 145 ) associated with the ER membrane and Golgi 144. Moreover, this is also where microtubule-associated protein light chain 3 A/B/C (MAP1LC3A/B/C or LC3 in humans, in yeast it is referred to as autophagy related protein 8, Atg8) positive structures can be seen emanating from these intracellular localizations 146. LC3 is the yeast homologue of Atg8 and is one of, if not the only, highly selective marker for mature autophagosomes (i.e. those that will soon be delivered to a lysosome) 147. After being translated, an unprocessed prolc3 is cleaved by Atg4 at the carboxy terminal to produce a cytosolic LC3-I. In a process mediated by the enzyme Atg7, LC3-I is conjugated or "lipidated" to a phosphatidylethanolamine (PE) moiety found on autophagosomal membranes, thus generating LC3-II 148. Since LC3-II is tethered and lipidated, it is readily distinguished from LC3-I on a SDS-PAGE gel due to differential mobility 147. Since there is a normally rapid turnover of autophagosomes by lysosomes 149, a need arises to inhibit this process to visualize autophagy. Autophagy inhibitors hydroxychloroquine or its derivative chloroquine and bafilomycin A1 are frequently used to achieve this. Chloroquine is a weak base that accumulates in lysosomes, thus preventing their acidification, and degradation of autophagic vesicles 150. Bafilomycin A1 achieves a similar result by inhibiting lysosomal H
43 ATPases necessary for acidification 151. As mentioned previously, the protein LC3 is highly selective for autophagy and is conjugated to the membranes of autophagosomes as they mature. The conjugated form is referred to as LC3-II and the soluble LC3-I, with the former being highly enriched on autophagosomes, and autophagy inhibitors cause accumulation of this marker 147. The most widely accepted technique to measure autophagosome generation is to measure the amount of LC3-II under blocked or non-blocked conditions and normalize this to actin level, as described by Klionsky and others 147. This method is therefore seen as a surrogate for autophagic flux, since these double membrane structures will eventually be delivered to the lysosome for degradation. Taken together, these findings are consistent with the notion that a) VPS34 and importantly PI(3)P are necessary for autophagy initiation b) one of the locales of autophagy initiation is the ER c) LC3 is an exclusive marker for mature autophagosomes that will eventually form autolysosomes. 31
44 NOTE: LC3-PE = LC3-II Figure 5. Overview of Autophagic Flux. Schematic of the 3 different types of autophagy. In chaperone-mediated autophagy (CMA), cytoplasmic contents are delivered straight to the lysosome. Proteins degraded by CMA have a peptide motif consisting of the residues KFERQ that localizes it to the lysosomal membrane associated LAMP2A, which mediates movement of cargo into the lysosome 137. In microautophagy, there is direct invagination of cytoplasmic contents by the lysosome 134. In macroautophagy, double membraned vesicles invaginate and enclose portions of the cytoplasm. These vesicles are termed autophagosomes, and they eventually fuse with the lysosome, forming an autolysosome. Pre-autophagosomal structures (PAS) are locations where the initiating events of macroautophagy occur 143. The phagophore is the location where the budding of double membrane vesicles occur, they are often referred to as omegasomes since they are similar in shape to the greek Ω 145. Shown below each step in macroautophagy is the names of the human genes responsible for mediating each specific step in the pathway. Eventually, degraded contents are released back into the cytoplasm for recycling and reuse. Figure adapted from Kaur and Debnath, Nature Reviews Molecular Cell Biology
45 1.7.3 The PI3K-mTOR Pathway and Autophagy A central node of control in the autophagic signalling pathway are the mtor complexes. Normally able to "sense" the availability of nutrients themselves, mtor complexes are also under direct regulation by AMP-activated protein kinase (AMPK). AMPK is sensitive to the ratio of cellular AMP:ATP and is activated by metabolic stresses 153. Under nutrient deficient conditions, AMPK can either promote activation of the negative mtorc1 regulator TSC2 or through direct inhibitory phosphorylation 154. When energy is abundant, mtor remains active and has inhibitory phosphorylation on the unc-51 like autophagy activating kinase 1 (ULK1) protein complex, which normally activates autophagy. This process is reversed and autophagy (even under nutrient rich conditions) is activated, when under the influence of rapamycin, a potent mtorc1 inhibitor 155. Another way mtor-mediated inhibition of autophagy may occur is when there is a removal of stimuli that activate mtor through the PI3K-AKT pathway. Since AKT is activated downstream of growth factors binding RTKs, a lack of growth factor signalling that subsequently decreases AKT activation would also decrease mtor activity, since it is a downstream target of AKT 156. There is evidence that both mtorc1 and 2 inhibit autophagy, and they both achieve this in different ways; whether AKT is activated as a result of mtorc2 upregulation or when mtorc1 is stimulated by AKT, the end result is autophagy inhibition 157. ULK1 and the closely related ULK2 proteins are normally activators of autophagy and knockdown of each or both protein complexes results in an inability to undergo normal autophagy 158. ULK1 phosphorylates key proteins involved in autophagy initiation. Following mtor inhibition and starvation, ULK1 can phosphorylate Beclin 1 (Atg6 in yeast, BECN1 in humans), enhancing the complex it forms with VPS34 and other proteins, as this complex is extremely important at isolation membranes where it accounts for the PI(3)P necessary for 33
46 autophagosome biogenesis 159. Conserved from yeast to humans, class III PI3K proteins (hvps34 in humans) use PI as a substrate to generate PI(3)P; VPS34 is also thought to be the main source of PI(3)P in cells 160. As mentioned previously, both PI(3)P and VPS34 are necessary for autophagy, and they are implicated in the earliest stages of this process 161. Pan PI3K inhibitors that target VPS34 such as 3-methyladenine, wortmannin and LY inhibit autophagy 162. Also, VPS34 knockout or mutation in vitro inhibits autophagy in multiple cell types 161,163, Autophagy and Cancer: a Double Edged Sword? Autophagy has frequently been implicated in the pathogenesis of neoplasms, and is thought to have context-specific roles in cancer maintenance and progression. Initially, autophagy was envisaged to have a tumour suppressive function in cancer. This is because proteins important for this process such as Beclin 1 (BECN1) are lost in cancer, sometimes up to 75% at the genetic level, in tumours of the breast, prostate and ovary. Interestingly, the genetic locus for BECN1 is adjacent to the known tumour suppressor breast cancer 1, early onset (BRCA1) 165. Becn1 null mice die prenatally and heterozygous mice frequently form tumours at random, indicating it is haploinsufficient 166, though this may be tissue dependent in some cases 167. Beclin 1 expression is also a predictor of clinical outcome in other cancers 168. In general, autophagy is important in some tissues or organ systems that may accumulate damaged mitochondria and protein aggregates. Impaired autophagy results in oxidative stresses, activation of DNA damage response or genomic instability and dysregulation of metabolism, all of which are implicated in cancer development and are of particular importance in degenerative diseases 169,170. In some cases, 34
47 blocking autophagy may inhibit differentiation, for example, as mature erythrocytes require the clearance of mitochondria and other organelles during development; a process thought to be achieved through autophagy. Ulk1 deletion in mice impairs their ability to remove mitochondria and ribosomes during development of red blood cells, albeit these knockout cells still have the ability to activate autophagy 171. This supports the idea that certain types of autophagy may be selective for some organelles or cargo (in this case, mitophagy or ribophagy), these however are nonetheless important in promoting critical events such as differentiation. Therefore, autophagy can be thought of as a process that "cleans up" products of malfunctioning processes in the cell, and when eliminated in a normal context, can have profound cell physiological consequences. This is consistent with the notion that cancer arises due to imbalances in normal maintenance of cellular functions. Indeed, genomic instability is thought to be the most prominent enabling characteristic of malignant cells that have acquired the "hallmarks of cancer" 172. On the other hand, a robust autophagy programme has been proposed to promote tumour formation. Cancer cells in some cases may undergo autophagy more than normal cells, possibly due to the increased demands in metabolism or biosynthesis, also they may encounter a lack of nutrients in hypoxic microenvironments and tumour niches 173. Expression of oncogenic mutant RAS upregulates basal autophagy 174 and certain tumours require autophagy for efficient growth under various conditions 175. Interestingly, almost all anticancer drugs in addition to radiation activate autophagy in some manner 150, thus suggesting resistant cancer cells are more effectively poised to activate autophagy, and thereby able to withstand more cytotoxic stresses. Since the manner by which autophagy affects cancer may be contextually or even temporally specific, 35
48 some have described an "autophagic switch" in cancer development. That is, some tumours may become dependent on autophagy after they have been formed, as a means of pro-survival 176. Chemotherapy resistance is also a significant hurdle needed to be overcome for optimal outcome of those treated with anticancer drugs. In addition to autophagy inhibitors being used in a myriad of clinical trials in cancer 150, autophagy inhibition has been frequently demonstrated to sensitize cells to various cancer therapies. For instance, combination of the autophagy inhibitor chloroquine with epirubicin decreased tumour survival and growth in breast cancer, and similarly with other chemotherapy drugs in melanoma, ovarian and colon cancer Autophagy and Leukemia Haematopoietic stem cells are in a state of constant quiescence when residing in the bone marrow 181. It is thought that autophagy is a crucial mechanism by which stem cells can maintain this phenotype, especially due to their long lifespan in some organisms 182. Ablation of the essential autophagy gene Atg7 (involved in post-translational processing of LC3 protein) in mice, results in a variety of blood-related complications. Indeed, haematopoietic stem and progenitor cells (HSPCs) lacking Atg7 were unable to form secondary colonies in colony-forming cell (CFC) assays, were unable to reconstitute the bone marrow of lethally irradiated mice and bone marrows from Atg7 -/- mice had an overall reduction of HSCs 183. Strikingly, the overall number of the more differentiated Lin - Sca-1 + c-kit + (LSK) progenitor cells are increased, albeit with defective autophagy and accumulation of dysfunctional mitochondria. The authors suggest this may account for the fact these mice have symptoms of a myeloproliferative disease, underlying the importance for autophagy in this model 183. The AKT-controlled transcription factor Forkhead box O3 (FOXO3A) transcribes genes that drive a pro-autophagy programme needed to survive 36
49 pro-apoptotic stimuli in HSCs 184. The same can be said for the master regulator of erythropoiesis, GATA-1, which transcribes a similar set of autophagy-related genes 185. Catalytic mtorc1 inhibitors can induce a protective autophagy program in AML, but autophagy blockers in combination with these inhibitors increase cytotoxicity. Thus, there is a promising avenue for synergistic roles for inhibiting autophagy and aberrant signalling pathways in leukemias with elevated PI3K/AKT signalling 186. Similarly, L-asparaginase (often used to treat acute lymphocytic leukemia) induces a cytoprotective autophagy in AML cells, thought to be as a result of mtorc1 inhibition 187. Owing to the theme of context-specificity, autophagy may play different roles in certain acute leukemias. In acute promyelocytic leukemia (APL) a bulk of tumours express the fusion protein promyelocytic leukemia gene (PML) - retinoic acid receptor α (RARα) caused by translocation t(15;17) (q22;q12). This fusion protein blocks the differentiation of myeloid cells by inhibiting transcription of genes necessary to facilitate their normal maturation. An overwhelming number of cases (up to 90%) can be cured with treatment by arsenic trioxide (As2O3) with all-trans retinoic acid (ATRA) 188,189. An interesting result of this therapy is that it has been shown to activate autophagy, and the PML-RARα fusion protein is degraded in an autophagy-dependent manner 190. This was also found to be exacerbated by mtorc1 inhibitor 191. Notably, this is not only limited to fusion protein found in APL; in chronic myeloid leukemia (CML), As2O3 induces the autophagic degradation of the BCR-ABL oncoprotein 192. It is now evident that autophagy plays multiple roles that are very likely context-dependent in cancer. Unfortunately, this contributes to the incredible amount of complexity and the difficulty involved in delineating the features of this pathway, it however provides great opportunity to discover targeted therapies that are effective for those with aggressive leukemias. 37
50 Figure 6. AKT activation results in indirect stimulation of mtor and inhibition of autophagy. An increase in mtor activity results in inhibition of the ULK complex of proteins necessary for autophagy stimulation. Shown here is two methods by which this may occur. A) AKT can directly block the inhibitor of Rheb proteins, TSC1/2, as normally activated Rheb can stimulate mtor. B) AMPK can feed into this pathway by direct inhibiton of TSC1/2, thus having opposite effects of AKT, and thereby initiating more autophagy. Bars indicate inhibitory inputs, arrows indicate stimulatory inputs. Figure adapted from Burman and Ktistakis, FEBS Letters
51 2. RATIONALE, AIMS AND HYPOTHESIS 2.1 Rationale and Aims Investigating mechanisms of INPP4B-mediated phenotypes in AML cells Given the unexpected association between INPP4B overexpression and poor disease outcome in AML, it was important to investigate the mechanisms mediating this phenotype. Our group discovered that INPP4B overexpression was associated with decreased overall survival, event free survival and poor response to induction therapy. We also demonstrated in vitro that INPP4B overexpressing cells recapitulate phenotypes representative of a more aggressive disease including increased proliferation, enhance colony forming potential and chemoresistance 91. These results presented a paradox because INPP4B had been clearly established as a tumour suppressor in other cancers, owing to its ability to regulate and prevent aberrant AKT activation. An important mechanistic question that arises from these data was whether these phenotypes can be explained by the lipid phosphatase activity of INPP4B or whether this is mediated by other phosphoinositide phosphatase-independent functions. Other groups have shed light on a potential mechanism for INPP4B overexpression in cancer. Gasser et al. demonstrated that INPP4B overexpression in breast cancer results in oncogenic PI3K signalling through SGK3 activation in vitro, but do not describe any patient outcomes associated with this upregulation 95. Despite this, the accumulation of PI(3)P is proposed to be a result of INPP4B upregulation, attributing functional consequence to lipid phosphatase activity of INPP4B. Similarily, two other studies in melanoma and colon cancer describe INPP4B overexpression mediating similar activation of SGK3. Guo and colleagues however also suggest this overexpression results in the dephosphorylation of PTEN, and hence, 39
52 sustained AKT activation because of PTEN inhibition. One common finding in all three studies is that INPP4B phosphatase activity is essential for these phenotypes 95,97. As a result, a central aim of this study was to explore phosphatase dependent phenotypes associated with INPP4B expression in our AML cell models INPP4B Overexpression and PI(3)P signalling: Implications for Autophagy The reports suggesting that INPP4B may activate SGK3 through PI(3)P accumulation 96,97 sheds light on unexplored considerations for INPP4B biology. In addition to the activation of SGK3, PI(3)P regulates a diverse array of biological processes such as cytokinesis 193, endosomal trafficking 194, phagocytosis 195 and autophagy 160, each of which can impact cancer. In particular, autophagy has long been considered to have multi-faceted roles in the generation and progression of cancers and it is a significant mechanism used by cancer cells to achieve resistance to chemotherapy 150.This has been demonstrated to be especially important in cancers such as AML where a majority of patients relapse after successful primary treatment regimens 113,176. There are currently no studies exploring the role of INPP4B in autophagy, however unpublished data from the Technical University of Munich hints at the involvement of ULK ULK1 is a lipid kinase essential for autophagy that phosphorylates VPS34 containing structures necessary for autophagy induction 159. The authors suggest there are predicted phosphorylation sites on INPP4B residues for ULK1. Interestingly, these sites are not found in its paralog, INPP4A. Moreover, they show that INPP4B and ULK1 interact in vitro, but do not show phosphorylation occurring under normal non-starvation conditions 196. It is possible that ULK1 40
53 phosphorylates proteins such as the Beclin 1-VPS34 complex necessary for autophagy, and even INPP4B, as a means to generate PI(3)P required for autophagic induction. Therefore, investigating the link between INPP4B and autophagy may explain the mechanisms behind some of the INPP4B overexpression-associated phenotypes demonstrated by our group and others. 2.2 Hypothesis We propose that INPP4B overexpression in AML promotes the activation of autophagy in a phosphatase dependent manner. 41
54 3. MATERIALS AND METHODS 3.1 Cell Culture OCI-AML2 and OCI-AML3 were cultured in αmem media, and NB-4 cells in RPMI 1640 media, both supplemented with 10% FBS and 100 units/ml penicillin as well as 100 units/ml streptomycin at 37 o C and 5% CO2. For the growth curve assay, cells were seeded on a 10 cm dish at a confluency of 1x10 3 or 1x10 4 cells/ml with counting once daily for a period of 6 days. Low serum conditions were defined as similar to normal media conditions as mentioned above except for the substitution of 10% FBS with 1% FBS for starvation purposes. 3.2 Lentivirus production psmal-puro, psmal-puro-flag-inpp4b wt (wild-type) and psmal-puro-flag- INPP4B mut -C824S (mutant) lentiviruses were produced by calcium phosphate transfection of HEK-293T cells with 2 nd generation lentiviral packaging and envelope plasmids (pspax2 and pci-vsvg) as described by the manufacturer (Life Technologies, Burlington, ON, Canada). Briefly, 6.4 μg of pspax2 and 3.6 μg of pci-vsvg along with 10 μg of target plasmid were transfected into HEK-293T cells at a confluency of 3.0x10 6 cells on a 10 cm plate. Media was changed 24 hours post-transfection. Viral particles in supernatants collected at 48 and 72 hours post-transfection were enriched with Lenti-X Concentrator according to manufacturer protocols (Clontech, Mountain View, CA, USA) or by centrifugation at 1500xg for 45 min at 4 C after overnight incubation with concentrator solution at 4 C. The concentrator solution consists of: 50% polyethylene glycol (PEG) 6000 and 0.3M NaCl. OCI-AML2, OCI-AML3 and NB4 cells were infected with 10X 42
55 concentrated lentivirus for 24 or 48 hours once or twice in the presence of 8 μg/ml protamine sulfate and selected with 2 μg/ml puromycin for 48 hours. 3.3 DNA plasmids Human cdna encoding wild-type INPP4B and C842S mutant INPP4B were cloned into psmal-puro (a kind gift from Dr. John Dick) using the PacI/XbaI restriction sites. The INPP4B C842S mutant cdna was generated by site directed mutagenesis using a Q5 Site Directed Mutagenesis Kit (New England Biolabs, Ipswich, MA, USA) according to manufacturer protocols. Site-directed mutagenesis primers were as follows: 5 -TTTCACCTGTAGTAAAAGT GC-3 (forward) and 5 -CGAATACCATTCAGTTTGC-3 (reverse). Plasmid DNA was purified using a Qiagen Maxiprep kit according to manufacturer's protocol (QIAGEN Inc., Hilden, Germany). 3.4 Methylcellulose Colony Formation Cell (CFC) Assay 900 μl of OCI-AML2 cells (control, INPP4B wt and INPP4B mut ) at a concentration of 1x10 3 cells/ml were mixed with 1.2 ml of 2.1% (w/v) methylcellulose and 900 μl of fetal bovine serum (FBS). 3 ml of this mixture was plated in triplicate in 3.5 cm plates with grids for counting. Pictures were taken of colonies and counted 9 days after plating and subsequent incubation at 37 o C and 5% CO2 with the EVOS XL Core Imaging System. 3.5 Phosphatase Assay INPP4B immunoprecipitated from 500 μg of lysates generated from the overexpression of INPP4B in the OCI-AML2 cell line (control or endogenous, INPP4B wt and INPP4B mut ). Isolated protein was combined with purified 100 μm dic8-ptdins(3,4)p2 (Echelon Biosciences, Salt Lake City, Utah) in 25 mm Tris-HCl (ph 7.5), 140 mm NaCl, 1 mm DTT at 37 C for 1 hour (see 43
56 manufacturer protocols for details). INPP4A protein provided by the manufacturer (Echelon Biosciences, Salt Lake City, Utah, USA) was used as a positive control, the same reaction with no enzyme was used as a negative control. Free inorganic phosphate release was measured with Pi-Glo: A Universal Bioluminescent Phosphatase Assay (gifted via Promega Corp.). 3.6 Immunoblotting Cell lysates were generated by spinning down cells in normal media and washing with PBS and subsequent spin at 4 o C. Cells were then resuspended in the appropriate volume RIPA buffer with protease inhibitors (Roche) and the following phosphatase inhibitors: 50 mm betaglycerol-phosphate (BGP), 1 mm Sodium Orthovanadate (activated chemical, Na3VO4), 4 mm Sodium Fluoride (NaF). Cells in RIPA buffer were then flash frozen using liquid N2. Cells were then lysed at 4 o C for 30 min and then centrifuged at 12,000xg for 20 min. Protein containing lysate was then mixed with 6X Laemmli buffer and ran on SDS-PAGE gel. The recipe for RIPA buffer is as follows: 10 mm Tris-Cl (ph 8.0), 1 mm EDTA, 1% Trition X-100, 0.1% sodium deoxycholate, 0.1% SDS and 140 mm NaCl. Western blotting was performed using the following commercial antibodies: LC3B (#3868), INPP4B (#8450 OR #14543), beta-actin (#4967S), GAPDH (#2118) and Anti-rabbit IgG, HRP-linked (#7074) from Cell Signaling Technology (CST, Danvers, Massachusetts, USA). All antibodies were incubated at a concentration of 1:1000 in 5% BSA in TBS-T buffer overnight at 4 o C unless otherwise stated. 3.7 Cyto-ID assay OCI-AML3 cells were collected and washed with PBS containing calcium or 1X assay buffer provided by the manufacturer (VWR, Radnor, Pennsylvania, USA). Cells were stained in 44
57 a solution containing 100 μl of PBS with 5% FBS and 100 μl of 1X Cyto-ID assay buffer. 1 ml of 1X Cyto-ID buffer is composed of: 1 μl Cyto-ID reagent, 50 μl FBS, and 949 μl PBS or 1X assay buffer. After 1 hour incubation at 37 o C in the dark, cells were pelleted and washed with 1X assay buffer or PBS and then ran through the flow cytometer. Live cells were determined by forward scatter and side scatter gating and green fluorescence was measured on the FITC channel (FL1). 3.8 Generation of Immortalized MEFs Low passage (P3-P5) wild-type and Inpp4b -/- C57BL/6J 58 primary MEFs were generated by infection with SV40 Large-T antigen retrovirus generated by retroviral transfection in HEK293T cells. After 48 hours infection in retroviral media and protamine sulfate (described similarly in lentiviral preparations), cells were selected in 75 μg/ml hygromycin for 48 hours Genotyping Genomic DNA was extracted from MEF cells using protocol outlined as previously described 198, without the use of phenol/chloroform/isoamyl alcohol. Primers used for genotyping were as follows: Wild-type forward: 5 GCTTCTGATAAAACATGGG 3 Wild type reverse: 5 TGGGCACATTTATAAGCCTTC 3 Mutant forward: 5 GCTTCTGATAAAACATGGG 3 Mutant reverse: 5 TGTTTTAAAAGCCTTGCTAAGTGTC 3 45
58 3.10 Statistics All P-values were determined to be significant if they were <0.05. Unless otherwise stated, all P-values were derived using a two-tailed, unpaired Student s t-test. *P<0.05, ** P <0.01, *** P < A one-way parametric analysis of variance (ANOVA) was used to determine the significance value for the daunorubicin dose-response curves. 46
59 4. RESULTS 4.1 Generation of a phosphatase-null INPP4B mutant (INPP4B mut ) protein INPP4B catalyzes the dephosphorylation of PI(3,4)P2 into PI(3)P through the action of a dual specificity-phosphatase (CX5R) consensus domain located near its carboxy terminal (Figure 3). In INPP4B, this is characterized by a CKSAKDR amino acid sequence starting at amino acid 842. In order to generate a catalytically inactive mutant, the cysteine (C) residue 842 was replaced by serine (S) to generate the C842S mutant through site directed mutagenesis of thymine (T)-2524 to adenine (A), as described previously (Figure 7) 64. To investigate whether the phosphatase function of INPP4B was necessary for the cellular phenotypes we observed previously in INPP4B wt overexpressing AML cell lines, lentiviral plasmids encoding catalytically active (psmal-puro-flag-inpp4b wt ) and inactive (psmalpuro-flag-inpp4b mut ) proteins or vector control (psmal-empty) were transduced in the OCI- AML2 line (Figure 8A). The presence of wild-type or mutant INPP4B did not change the basal AKT phosphorylation level in any of these cell lines. This was seen previously in wild-type cells by flow cytometric staining for p-akt (Ser473, both OCI-AML2 and OCI-AML3) in INPP4B WT cells. For phosphatase assays, INPP4B protein (isolated via immunoprecipitation with an anti-inpp4b antibody, see methods) was incubated with purified PI(3,4)P2 and phosphatase activity was measured via luminescence generated by a chemical reaction with liberated phosphate groups. Compared to vector control (i.e. endogenous INPP4B) and mutant, immunoprecipitates from INPP4B wt overexpressing OCI-AML2 cells had 1.6 and 2.0 fold greater 4-phosphatase activity, respectively (Figure 8B). Thus, these results indicate that our engineered 47
60 INPP4B mut protein is functionally inactive and suitable to perform further experiments to address the phosphatase dependency of INPP4B-overexpression phenotypes. 48
61 Mutant: Wild-type: Figure 7. Sequencing Data Encoding INPP4B wt and INPP4B mut phosphatase domains. Raw cdna sequencing data from plasmids encoding wild-type and null INPP4B proteins. The catalytic domain in INPP4B is characterized by a residue sequence of 7 amino acids: CKSAKDR. In other similar phosphatases, it is characterized by 5 amino acids surrounded by a cysteine and arginine (CX5R) 199. In the top-half of the panel is the sequencing data from the plasmid psmal-puro-flag-inpp4b C824S (mutant) and the bottom-half psmal-puro-flag-inpp4b wt (wild-type). The mutant is characterized by a cysteine to serine mutation, C842S, as described previously for INPP4B
62 Figure 8. Characterization of OCI-AML2 cells expressing vector control and INPP4B wt and INPP4B mut proteins. (A) Western blot characterizing expression of exogenous INPP4B protein and basal levels of AKT and p-akt protein in OCI-AML2 cells. C842S mutant was generated by site directed mutagenesis. (B) Phosphatase assay measuring catalytic activity of INPP4B in the OCI-AML2 cell line. Briefly, purified INPP4B protein was incubated with PI(3,4)P2 and the release of free phosphate was measured. A greater luminescence means more liberation of phosphate groups from PI(3,4)P2 by INPP4B. P-values were derived using the Student s t-test. *P<0.05, ** P <0.01, *** P < This figure was adapted from Dzneladze et al., 2015, Leukemia
63 4.2 INPP4B mut OCI-AML2 cells do not exhibit phenotypes observed in INPP4B wt overexpressing cells To determine whether the phenotypes caused by INPP4B wt overexpression are phosphatase dependent, we repeated previous experiments with the INPP4B mut overexpressing OCI-AML2 cell line. AML cells overexpressing INPP4B wt have faster growth rates compared to control cells; INPP4B mut cells on the other hand showed an intermediate growth response (Figure 9A). Similarly, INPP4B mut cells displayed only a partial decease viability after growth in low serum conditions when compared to INPP4B wt and vector control cells after 96 hours (Figure 9B). In all the aforementioned experiments, viability was determined by Trypan Blue exclusion staining. We previously determined that OCI-AML2 and OCI-AML3 show increased survival through diminished Annexin V staining when compared to control under normal growth conditions and negligible cell cycle differences 91. This suggests they do not proliferate quicker but rather expression of wild-type INPP4B confers a survival advantage. These data demonstrate that mutant INPP4B may confer partial growth phenotypes in liquid culture conditions. In contrast, colony formation potential of AML cell lines colony forming cell (CFC) assays in methylcellulose was enhanced by overexpression of INPP4B wt, however cells overexpressing INPP4B mut were no different than that of vector control cells (Figure 10). Similar to CFC assays, the resistance conferred by INPP4B wt overexpression appears to be completely dependent upon the phosphatase function of INPP4B. Dose response experiments demonstrated that the EC50 for daunorubicin (DNR) of INPP4B wt overexpressing cells was 63.5 nm whereas it was 20.2 and 23.7 nm for control and INPP4B mut, respectively (Figure 11A). Similarly, no differences were observed in the viability of control and INPP4B mut cells compared 51
64 to INPP4B wt expressing cells over 4 days in the presence of 10 nm or 50 nm DNR (Figure 11B). Viability was measured using the trypan blue exclusion assay. Overall, the cellular phenotypes observed upon overexpression of INPP4B wt in AML cell lines were generally phosphatase-dependent. However, partially enhanced viability and proliferative potential was observed upon expression of INPP4B mut in AML cell lines, suggesting the potential of context-linked phosphatase dependency. 52
65 A B Figure 9. INPP4B wt cells proliferate more rapidly than control or mutant cells in normal and low serum conditions. (A) Cells expressing exogenous wild-type INPP4B protein were able to grow in normal media faster than control or mutant expressing cells for a period of 5 days. (B) INPP4B wt cells show greater viability after being cultured in low serum (1% FBS) conditions at 96 hours. Error bars are +/- S.E.M. This figure was adapted from Dzneladze et al., 2015, Leukemia
66 Figure 10. INPP4B wt cells form colonies preferentially in vitro. In methylcellulose media, wild-type overexpressing INPP4B OCI-AML2 cells form more colonies than control or mutant cells. Shown are representative images of colonies formed from each cell line in 3 independent experiments. Error bars are +/- S.E.M. All P-values were derived using the Student s t-test. *P<0.05, ** P <0.01, *** P < This figure was adapted from Dzneladze et al., 2015, Leukemia 91 54
67 A B Figure 11. INPP4B wt cells are more resistant to daunorubicin in vitro. (A) INPP4B overexpression conferred chemoresistance to OCI-AML2 cells in cuture that was not seen in mutant or control cell lines. Shown is a dose response curve in which increase doses of daunorubicin were used to assay for drug sensitivity. Overall, wild type cells had approximately 3-fold greater EC50 than either control or mutant cell lines. (B) Using 2 specific doses of daunorubicin, 10 and 50 nm, INPP4B wt cells were more resistant to a time course of drug treatment over a period of 4 days. Mutant and control cells showed little to no difference in response over this time period. This figure was adapted from Dzneladze et al., 2015, Leukemia 91 55
68 4.3 INPP4B wt Overexpressing Cells Have Increased Staining of Autophagosomes in an Untreated Condition Since INPP4B overexpression generally demonstrated phosphatase dependent phenotypes, we reasoned that signalling as a result of PI(3)P overproduction may be activated in INPP4B wt overexpressing AML cell lines. Since PI(3)P is a potent activator of autophagy and sites of autophagosome biogenesis are enriched with PI(3)P 144, we sought to measure autophagy activation in INPP4B overexpressing cells. For this we first utilized a green-fluorescent dye that selectively stains autophagic vesicles ranging from autophagosomes to autophagolysosomes, called Cyto-ID. Cyto-ID stains live cells and can be used to monitor autophagy without the need of cell permeabilization 200. Using flow cytometry, we observed that INPP4B wt OCI-AML3 cells displayed greater staining as measured by the increase in green fluorescence compared to vector control cells (Figure 12A and B) In total, INPP4B wt cells displayed 20% greater staining versus control, indicating an increase in the amount of autophagic vesicles at a basal, untreated or unstimulated setting. These data provide the first evidence that overexpression of INPP4B may promote the activation of autophagy. Nevertheless, Cyto-ID alone is not a reliable measure of autophagy induction thus more reliable assays are required. 56
69 Fold Change Mean Fluoresence A B Control INPP4B Figure 12. INPP4B expression in OCI-AML3 cells leads to a higher level of autophagosomes in an unstimulated condition. (A) In an untreated condition, OCI- AML3 cells expressing exogenous wild-type INPP4B protein had a higher level of autophagosome staining measured by flow cytometry using the green fluorescent dye Cyto-ID, which is specific for membranes of autophagosomes. Data shown is of 4 independent experiments. Each line connecting a different shape represents one experiment in which both control and INPP4B wt cells were stained with Cyto-ID. In each case, there is a fold change in the difference in staining intensity, depicted by the line connecting any two shapes. Overall, wild-type overexpressing cells had approximately 20% more staining than control. (B) A representative figure of 4 experiments showing the shift in fluorescence for entire population of stained wild-type or control cells. Cells were gated for the live population only. 57
70 4.4 INPP4B wt cells demonstrate increased autophagosome accumulation in the presence of inhibitors of autolysosomal acidification As mentioned previously, there is a need to inhibit autophagy to visualize it and measure it effectively. This is because autophagy normally has rapid turnover, and is thus difficult to measure a steady state condition. To determine whether INPP4B wt overexpressing cells had increased levels of autophagy, we inhibited autolysosomal acidification and observed alterations in autophagosome accumulation through Western blot detection of LC3B-II protein. Overexpression of INPP4B in the OCI-AML3 cell-line resulted in a 2-fold increase in LC3B-II protein under inhibition via chloroquine (Figure 13A and B). Notably, even in the absence of chloroquine we were able to visualize LCB3-I and II proteins in this cell line (Figure 13A). This finding corroborates our observations with INPP4B wt OCI-AML3 cells stained with Cyto-ID, where there is an increase in the basal level of autophagosomes. When measured by LC3B-II protein by western blot, we observed 1.6 fold LC3B-II in INPP4B wt vs control (without chloroquine; Figure 13B). In the OCI-AML2 cell line, overexpression of INPP4B wt results in an increased accumulation of autophagosomes, as measured by levels of LC3B-II protein by Western blot (Figure 14A). There was a 4-fold increase in the amount of LC3B-II in the presence of chloroquine in INPP4B wt OCI-AML2 cells compared to control (Figure 14B). Without chloroquine, there is little to no detection of LC3B-II, consistent with previous reports that basal level of this protein is cell-line dependent (Figure 14A)
71 A B Figure 13. OCI-AML3 INPP4B wt have a greater propensity for autophagosome biogenesis. (A) Immunoblot characterizing the amount of LC3 accumulated in OCI- AML3 cells. In the presence of the autophagy inhibitor chloroquine, INPP4B wt OCI- AML3 cells have more accumulation of the mature autophagosome marker, LC3-II at 24 hours in normal media, as judged by immunoblotting. Shown is a representative image of 3 experiments. 5 μm chloroquine was used over a period of 24 hours. (B) Densitometric analysis showing relative levels of LC3-II protein in AML3 cells with respect to β-actin. Error bars are +/- S.E.M. Shown are the average of 3 independent experiments. All P-values were derived using the Student s t-test. *P<0.05, ** P <0.01, *** P <
72 A B Figure 14. Expression of INPP4B in OCI-AML2 increases amount of autophagosome accumulation. (A) Immunoblot characterizing the amount of LC3 accumulated in OCI-AML2 cells. In the presence of the autophagy inhibitor chloroquine, INPP4B wt OCI-AML2 cells have more accumulation of the mature autophagosome marker, LC3-II at 24 hours in normal media, as judged by immunoblotting. Shown is a representative image of 3 experiments. 25 μm chloroquine was used over a period of 24 hours. (B) Densitometric analysis showing relative levels of LC3-II protein in AML cells with respect to β-actin. Error bars are +/- S.E.M. Shown are the average of 3 independent experiments. 60
73 4.5 INPP4B wt NB4 cells demonstrate increased autophagosome accumulation when treated with chloroquine Treatment of cells with increasing levels of chloroquine has been previously demonstrated to simultaneously increase LC3 accumulation 201. To determine whether cell lines overexpressing INPP4B showed dose dependent differences in LC3 quantity, NB4 cells overexpressing INPP4B wt and control constructs were cultured in the presence of 5 and 10 μm chloroquine for 24 hours. Interestingly, there was an expected increase in LC3B-II accumulation (6.5 fold) in INPP4B wt NB4 cells treated with 5 μm chloroquine (Figure 15). In contrast, there were no changes in LC3IIB in control vs. INPP4B wt NB4 cells treated with 10 μm of chloroquine (Figure 15). This suggests there may be threshold effect with 10 μm chloroquine at 24 hours in NB4 cells. Figure 15. Dose dependent effect of chloroquine in INPP4B wt NB4 cells. Immunoblot characterizing more LC3B-II protein in the presence of chloroquine, this shows that wild-type expressing INPP4B NB4 cells had a greater accumulation of autophagosomes after 24 hours in culture, however only at a 5 μm dose, 6.5 fold higher. At a 10 μm dose, the effect was diminished and not seen. 61
74 4.6 Inpp4b -/- MEFs have a reduced ability to undergo autophagy Using Inpp4b knockout mouse embryonic fibroblasts (MEFs) obtained from the Vacher lab 75, we sought to determine whether Inpp4b is necessary for autophagy. In a basal, untreated condition, Inpp4b null MEFs have lower levels of LC3B-II protein compared to wild type MEF (Figure 16A). To determine whether these Inpp4b knockout cells had less accumulation of autophagic vesicles over time, we treated them with the autophagy inhibitor bafilomycin A1 at a concentration of 25 nm for up to 3 hours (Figure 16B). We noticed that surprisingly, over time wild-type MEFs had a much greater ability to accumulate autophagosomes in the presence of bafilomycin A1, as measured by the amount of LC3B-II at all timepoints. It is of note that time course experiments are better indicator of autophagic flux according to guidelines 147, since this indicates a constant delivery of autophagosomes to the lysosome. These data therefore suggest that knockout of Inpp4b causes a severe deficiency in the ability of MEFs to undergo autophagy. 62
75 A B Figure 16. Inpp4b -/- MEFs have a reduction in their ability to accumulate autophagosomes in a basal state. (A) Wild-type and Inpp4b null mouse embryonic fibroblasts were lysed and total LC3 protein was detected. In normal untreated conditions, Inpp4b -/- cells had decreased level of basal autophagosomes. Shown is a representative immunoblot of 3 independent experiments. (B) In the presence of the autophagy inhibitor bafilomycin A1 (25 nm), Inpp4b -/- cells had a considerable reduction in the accumulation of autophagosome at 1.5 or 3 hours in culture. Over time, accumulation of autophagosomes in the presence of bafilomycin A1 occurred in both cell lines, but to a much lesser extent in Inpp4b null MEFs. Consistent with previous results, they also had less basal amount of autophagosomes in an untreated setting (0 hours). 63
76 5. DISCUSSION 5.1 Summary of Results The primary objective of this study was to investigate the molecular consequences of INPP4B overexpression in AML. My project was designed to provide insight into novel cell signalling coordinated by INPP4B. In brief, we observed that INPP4B overexpression-mediated phenotypes in AML cells are greatly diminished or absent with the introduction of a phosphatase-null, mutant INPP4B protein suggesting that INPP4B is functioning in a phosphatase dependent manner. It was also observed that overexpression of INPP4B leads to increased autophagosome accumulation, an indicator that INPP4B promotes autophagic flux, presumably as a result of increased cellular PI(3)P production. Initially, I thought it was important to show that the phenotypes we observed upon exogenous INPP4B overexpression were mediated by INPP4B phosphatase function. For this reason I generated a phosphatase dead mutant version of INPP4B protein, by changing the DNA sequence coding for the cysteine at residue number 842 to serine (C842S; INPP4B mut ) (Figures 7 and 8). For these experiments I overexpressed INPP4B wt and INPP4B mut in the OCI-AML2 cell line, a well-established human AML-derived cell line commonly used to study biological properties of AML 202. Using a phosphatase assay, I demonstrated that the INPP4B mut protein was catalytically inactive, as compared to INPP4B wt protein in OCI-AML2 (Figure 8). Next, we performed a battery of experiments on INPP4B wt and INPP4B mut OCI-AML2 cells to investigate the role of INPP4B phosphatase function in observed AML phenotypes (Figures 8-11). In all tests performed, INPP4B mut presented phenotypes that were either similar to control or only partially recapitulated the INPP4B wt phenotype. In summary, colony formation in 64
77 methylcellulose (Figure 10) and DNR resistance (Figure 11) conferred by INPP4B wt overexpression were completely phosphatase dependent, whereas INPP4B may have some cell growth and viability effects (Figure 9) which are phosphatase-independent. Since autophagy is normally a rapid process with high turnover, visualizing it remains difficult. We therefore used agents that inhibit the lysosomal acidification of autophagosomes to permit the investigation of autophagosomal proteins and measure if their accumulation is altered in cells with INPP4B overexpression. Using AML cell lines (OCI-AML2, OCI-AML4, NB4) overexpressing INPP4B wt, we observed that INPP4B promotes autophagy. When blocking autophagy using inhibitors such as chloroquine in any cell line, there is usually an increase of the autophagosome-specific marker LC3B-II. We noticed that despite this, there was even greater increase of LC3B-II in OCI-AML2, OCI-AML3 and NB4 INPP4B wt cells with the addition of chloroquine (Figures 13-15); although the dose used to achieve this effect is different across cell lines. This is similar to previous findings that suggest different levels of chloroquine result in varying amounts of LC3 accumulation 201. Notably, INPP4B expression alone increases the amount of autophagosomes without stimulation or treatment in OCI-AML3 cells (Figure 12 and 13) and MEFs (Figure 16A), since in these cells it is possible to visualize this to a limited extent without pharmacological inhibition. In Inpp4b knockout MEFs, we observed a marked increase in the basal level of autophagosomes without stimulation or blocking. With the addition of bafilomycin, there was a greater accumulation over time of autophagosomes in the presence of 25 nm bafilomycin (Figure 16B), an indicator of increased autophagic flux. 65
78 5.2 INPP4B Phosphatase Activity Despite its importance in many solid cancers and some leukemias 203,204, the phosphoinositide 3-phosphatase PTEN does not play important roles in acute leukemias such as AML. This suggests that hyperactivation of the PI3K/AKT pathway may be achieved by other means specifically in cancers such as AML. INPP4B is a lipid phosphatase that dephosphorylates the phosphoinositide PI(3,4)P2 to generate the mono-phosphorylated PI(3)P. Until recently, it has been thought that INPP4B acts to halt AKT activation by depleting one of its membrane anchors, hence diminishing its membrane recruitment and subsequent activation. While this paradigm is true for many cancers and biological contexts, it was not described in blood cancers such as AML. Like our study published in Leukemia, Rijal and colleagues observed that INPP4B was upregulated in AML patients and this was associated with poor disease outcome. Based on expression from bone marrow patients measured via mass spectrometry, high levels of INPP4B mrna were correlated with decreased overall survival (OS). Overexpression of INPP4B protein in patient AML blasts was also associated with poor leukemia free survival (LFS) and OS. Finally, they also found that higher levels of INPP4B correlated with inferior response to induction therapy. This demonstrated there is likely a functional consequence for this surprising overexpression of INPP4B gene and protein 92. We sought to determine whether this phenomenon is true at the cellular level. The study by Rijal et al. also mainly confirmed our findings from in vitro experiments, as INPP4B mediated chemoresistance to multiple drugs and knockdown of INPP4B sensitized these cells to treatment. Interestingly, in both studies, overexpression of INPP4B in AML cell lines did not alter p-akt status, as would be expected based on the canonical tumour suppressor INPP4B function, further 66
79 suggesting an alternative mechanism for INPP4B function in this tissue or cell type 205. In xenograft experiments, INPP4B overexpressing AML cells were transplanted into sublethally irradiated NSG mice. Mice overexpressing INPP4B wt were more resistant to Ara-C treatment and died quicker from leukemia than did control or INPP4B mut expressing cells. These and our findings suggest that INPP4B upregulation in AML is functionally relevant both in a cellular context, where can it promote the cellular phenotypes and chemotherapy responses associated with more aggressive AML, and in a physiological context, where it causes chemoresistance in leukemia xenografts and finally, as a clinical tool, since it is an effective biomarker of poor prognosis AML. What differs in the findings by Rijal and colleagues with respect to our study is their discovery that almost of all these phenotypes are phosphatase-independent. That is, a catalytically inactive mutant protein (engineered as a similar cysteine to alanine mutation, C842A) recapitulated all of the phenotypes observed in cell lines or mice overexpressing wildtype INPP4B. Notably, they showed that INPP4B protein is also catalytically active in AML patient samples. Of concern, this directly contrasts the findings of our study that show that INPP4B phenotypes are completely (or partially) phosphatase-dependent. It is possible that there are phosphoinositide phosphatase-independent functions of INPP4B but these are to date, mechanistically unfounded. Notably, it has been recently reported that INPP4B may exhibit protein tyrosine phosphatase activity 64, suggesting it may modulate protein targets through dephosphorylation. Guo et al. have suggested this INPP4B-mediated protein phosphatase activity could promote cancer progression and activation 97, however the protein substrate targets INPP4B may act upon are not clearly identified, especially in the context of AML. 67
80 The lack of evidence of INPP4B controlling AKT phosphorylation in AML corroborates the possibility that other INPP4B functions may be at play. We hypothesized that INPP4B is indeed catalytically active as a PI(3,4)P2 4-phosphatase, thus more INPP4B activity will generate more of its product, PI(3)P. PI(3)P in turn drives specific signalling and cellular processes. This is not without precedent, as it has been demonstrated that INPP4B upregulation only in certain subtypes of breast cancer and melanoma drives SGK3 signalling, presumably through the local accumulation of PI(3)P. Regardless of the likely consequence of INPP4B overexpression, it is important to consider that it hasn t been well characterized whether the cellular pool of PI(3)P that comes from INPP4B has biological relevance, since Class III PI3K has been previously thought to be the main source of this phosphoinositide 160. This further validates a contextspecific theme for INPP4B biology, because this would be a novel situation in which INPP4B is preferentially upregulated in blood cancers, as opposed to mainly being lost or inactivated in solid tumours. 68
81 5.3 INPP4B and Autophagy Autophagy is a cellular degradation pathway wherein cytoplasmic contents are delivered to a lysosome for destruction and eventually, recycling back into the cell. Although autophagy has been proposed to have numerous roles in cancer, there is evidence that it is a resistance mechanism employed by many tumours to evade therapies and cytotoxic insults 150. Herein we showed that the phenotypes we observed upon ectopic INPP4B overexpression were mediated by the phosphatase activity of INPP4B. This led us to the hypothesis that INPP4B overexpression could lead to the accumulation of cellular PI(3)P. Since PI(3)P is required for autophagy, we assumed that INPP4B may contribute to the signalling that promotes this process. By showing that INPP4B overexpression leads to accumulation of autophagosomes (Figures 12-15), this would provide evidence to suggest that PI(3)P generated downstream of INPP4B may facilitate autophagy. As mentioned previously, autophagy normally has rapid turnover and the presence of the autophagosome-associated protein LC3 I or II is variable across cell lines, especially without the use of autophagy inhibitor. I demonstrated that INPP4B WT showed little to no difference in LC3 levels without the use of chloroquine but in the presence of autophagy inhibition there was a noticeable difference in the amount of LC3-II accumulation. This corroborates the idea that autophagy needs to be visualized over time (in this case, 24 hours in the presence of chloroquine) to appreciate the extent of autophagosome generation or accumulation 147. Further work still needs to be done to determine if autophagosomes generated in this scenario in fact mean there is more autophagic flux; that is, do these vesicles actually invaginate more cytoplasmic content, whether selectively or not, and is this subsequently cleared by the lysosome. 69
82 Furthermore, our study of Inpp4b -/- MEFs indicate that lack of INPP4B protein may have profound effects on LC3-II accumulation. In basal conditions, INPP4B-null MEF cells had compromised ability to accumulate autophagosomes, as measured by western blot of LC3B-II levels (Figure 16A). We also saw that autophagic flux was markedly reduced in knock out cells, by using bafilomycin to inhibit autophagosomal degradation and observed the accumulation of autophagosomes over time (Figure 16B). Previously, it was reported that MEF cells deficient in VPS34 can still undergo appreciable levels of autophagy 206, a puzzling finding given the documented indispensability of VPS34 for autophagy in other studies. Therefore, further studies of Inpp4b -/- MEF are necessary and warranted. In addition to this, it is interesting to note that autophagy is involved in some manner in each of the aberrant phenotypes observed in INPP4B wt overexpressing AML cells. Links to autophagy and cell proliferation, nutrient deprivation, chemoresistance and colony formation are documented 207. Whether increased autophagy is commonly observed across all AML cell lines upon the introduction or expression of INPP4B remains to be seen. Though this link is not particularly evident at this time, there is a convincing rationale to explore this avenue because autophagy is a process that can be targeted quite effectively with pharmacological intervention in a wide range of cancers
83 5.4 Conclusions To date, INPP4B has been proposed to have different roles in multiple cancers. It remains to be seen what mechanism best describes which function is preferred in a given context. Two independent studies have presented findings that point to INPP4B overexpression being representative of a more aggressive disease in AML. We propose that INPP4B promotes signalling in a manner consistent with its primary role as a lipid phosphatase in AML cell line models. Indeed, the catalytic function of INPP4B is critical for the cellular phenotypes including colony formation and chemoresistance. We hypothesized that INPP4B was capable of activating autophagy in AML cells; as measured by different tools, we have found this hypothesis to be true. In summary, this study sheds light on the cellular and molecular consequences of INPP4B and serves as a platform for future studies that intend to uncover the complexities of phosphoinositide signalling and INPP4B biology. 5.5 Future Directions It remains to be seen whether INPP4B function can be sufficient for leukemia formation, or just as a facilitator for worse disease. Mouse models in which INPP4B is deleted would provide a platform to study its direct effects on cancer generation and progression. A leukemia is not likely to be caused by a single genetic event, as this suggests INPP4B may act in concert with other known oncogenes. Furthermore, it is paramount to understand the contribution of INPP4B to the levels of cellular phosphoinositides in AML. Importantly, what is the effect on cellular PI(3,4)P2 and PI(3)P when it is lost or gained? Are the conventional sources of both lipids, Class II and III PI3Ks respectively, altered in similar disease states? 71
84 If there is in fact increases in phosphoinositides such as PI(3)P, is downstream signalling as a result controlled spatially in the cell? Since PI(3)P is formed upstream of autophagosome formation, there is rationale to investigate whether INPP4B co-localizes with preautophagosomal structures. INPP4B has been shown to be specifically enriched on endosomes 70, and in some cases endosomes can feed into the same pathway by which lysosomes degrade contents of autophagosomes 208. There are a number of proteins involved in autophagosome biogenesis that could be used as markers for potential co-localization experiments with INPP4B. There is limited data to show that our C842S mutant has defective autophagy (data not shown) but this requires further investigation under the same conditions for which there was a difference in INPP4B wt cells compared to control. Beyond an AML context, there is a larger question as to what is the "switch" between a tumour suppressive versus oncogenic activator or initiator roles for INPP4B. It remains unclear if AKT activation is changed with overexpression of INPP4B in AML cell lines with respect to a starvation or stimulation scenario. It is possible that some cellular contexts are more "primed" to lean one way or the other. In a cancer cell that has tight regulation of AKT, that is when PTEN is not deleted or mutated in a tumour, INPP4B regulation of AKT may not be significant in the pathogenesis and maintenance of disease. The reverse is potentially true as well, cells that otherwise do not upregulate or change levels of PI(3)P producing proteins (for example, Class II or III PI3Ks) may induce INPP4B to compensate this. Another aspect of these findings is whether or not they contribute to resistance mechanisms. To answer these questions, it may be required to block autophagy in the presence or absence of chemotherapeutic agents. Modulation of autophagy has clear benefit as inhibitors 72
85 are frequent agents used in clinical trials. Being selective, or "personalizing" contexts in which these drugs may be more efficacious is of great therapeutic value. 73
86 6. REFERENCES 1. Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, (2006). 2. Agranoff, B. W., Roy, M. & Brady, R. O. The Enzymatic Synthesis of Inositol Phosphatide. J. Biol. Chem. 233, (1958). 3. Balla, T. Phosphoinositides: tiny lipids with giant impact on cell regulation. Physiol. Rev. 93, (2013). 4. Lemmon, M. A. Membrane recognition by phospholipid-binding domains. Nat. Rev. Mol. Cell Biol. 9, (2008). 5. Lemmon, M. A. & Ferguson, K. M. Signal-dependent membrane targeting by pleckstrin homology (PH) domains. Biochem. J. 350 Pt 1, 1 18 (2000). 6. Bunney, T. D. & Katan, M. Phosphoinositide signalling in cancer: beyond PI3K and PTEN. Nat. Rev. Cancer 10, (2010). 7. Mayinger, P. Phosphoinositides and vesicular membrane traffic. Biochim. Biophys. Acta 1821, (2012). 8. Redondo-Muñoz, J., Josefa Rodríguez, M., Silió, V., Pérez-García, V., María Valpuesta, J. & Carrera, A. C. Phosphoinositide 3-kinase beta controls replication factor C assembly and function. Nucleic Acids Res. 41, (2013). 9. Kumar, A., Fernandez-Capetillo, O., Fernadez-Capetillo, O. & Carrera, A. C. Nuclear phosphoinositide 3-kinase beta controls double-strand break DNA repair. Proc. Natl. Acad. Sci. U. S. A. 107, (2010). 10. Marqués, M., Kumar, A., Poveda, A. M., Zuluaga, S., Hernández, C., Jackson, S., Pasero, P. & Carrera, A. C. Specific function of phosphoinositide 3-kinase beta in the control of DNA replication. Proc. Natl. Acad. Sci. U. S. A. 106, (2009). 11. Yin, Y. & Shen, W. H. PTEN: a new guardian of the genome. Oncogene 27, (2008). 12. Mejillano, M., Yamamoto, M., Rozelle, A. L., Sun, H. Q., Wang, X. & Yin, H. L. Regulation of apoptosis by phosphatidylinositol 4,5-bisphosphate inhibition of caspases, and caspase inactivation of phosphatidylinositol phosphate 5-kinases. J. Biol. Chem. 276, (2001). 13. Yusuf, I., Zhu, X., Kharas, M. G., Chen, J. & Fruman, D. A. Optimal B-cell proliferation requires phosphoinositide 3-kinase-dependent inactivation of FOXO transcription factors. Blood 104, (2004). 14. Falkenburger, B. H., Jensen, J. B., Dickson, E. J., Suh, B.-C. & Hille, B. Phosphoinositides: lipid regulators of membrane proteins. J. Physiol. 588, (2010). 74
87 15. Pendaries, C., Tronchère, H., Plantavid, M. & Payrastre, B. Phosphoinositide signaling disorders in human diseases. FEBS Lett. 546, (2003). 16. Mak, L. H. in Encyclopedia of Biophysics (Springer Berlin Heidelberg, 2013). doi: / _ Le Roy, C. & Wrana, J. L. Clathrin- and non-clathrin-mediated endocytic regulation of cell signalling. Nat. Rev. Mol. Cell Biol. 6, (2005). 18. Vivanco, I. & Sawyers, C. L. The phosphatidylinositol 3-Kinase AKT pathway in human cancer. Nat. Rev. Cancer 2, (2002). 19. VANHAESEBROECK, B. & ALESSI, D. R. The PI3K PDK1 connection: more than just a road to PKB. Biochem. J. 346, (2000). 20. Engelman, J. a, Luo, J. & Cantley, L. C. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat. Rev. Genet. 7, (2006). 21. Gassama-Diagne, A., Yu, W., ter Beest, M., Martin-Belmonte, F., Kierbel, A., Engel, J. & Mostov, K. Phosphatidylinositol-3,4,5-trisphosphate regulates the formation of the basolateral plasma membrane in epithelial cells. Nat. Cell Biol. 8, (2006). 22. Gual, P., Le Marchand-Brustel, Y. & Tanti, J.-F. Positive and negative regulation of insulin signaling through IRS-1 phosphorylation. Biochimie 87, (2005). 23. Jean, S. & Kiger, A. A. Classes of phosphoinositide 3-kinases at a glance. J. Cell Sci. 127, (2014). 24. Scheid, M. P., Marignani, P. A. & Woodgett, J. R. Multiple phosphoinositide 3-kinasedependent steps in activation of protein kinase B. Mol. Cell. Biol. 22, (2002). 25. Waugh, C., Sinclair, L., Finlay, D., Bayascas, J. R. & Cantrell, D. Phosphoinositide (3,4,5)-triphosphate binding to phosphoinositide-dependent kinase 1 regulates a protein kinase B/Akt signaling threshold that dictates T-cell migration, not proliferation. Mol. Cell. Biol. 29, (2009). 26. Ma, K., Cheung, S. M., Marshall, A. J. & Duronio, V. PI(3,4,5)P3 and PI(3,4)P2 levels correlate with PKB/akt phosphorylation at Thr308 and Ser473, respectively; PI(3,4)P2 levels determine PKB activity. Cell. Signal. 20, (2008). 27. Alessi, D. R., James, S. R., Downes, C. P., Holmes, A. B., Gaffney, P. R., Reese, C. B. & Cohen, P. Characterization of a 3-phosphoinositide-dependent protein kinase which phosphorylates and activates protein kinase Balpha. Curr. Biol. 7, (1997). 28. Sarbassov, D. D., Guertin, D. A., Ali, S. M. & Sabatini, D. M. Phosphorylation and regulation of Akt/PKB by the rictor-mtor complex. Science 307, (2005). 29. Vadlakonda, L., Dash, A., Pasupuleti, M., Anil Kumar, K. & Reddanna, P. The Paradox of Akt-mTOR Interactions. Front. Oncol. 3, 165 (2013). 30. Zhang, X., Tang, N., Hadden, T. J. & Rishi, A. K. Akt, FoxO and regulation of apoptosis. Biochim. Biophys. Acta 1813, (2011). 75
88 31. Kandel, E. S., Skeen, J., Majewski, N., Di Cristofano, A., Pandolfi, P. P., Feliciano, C. S., Gartel, A. & Hay, N. Activation of Akt/protein kinase B overcomes a G(2)/m cell cycle checkpoint induced by DNA damage. Mol. Cell. Biol. 22, (2002). 32. Carracedo, A. & Pandolfi, P. P. The PTEN-PI3K pathway: of feedbacks and cross-talks. Oncogene 27, (2008). 33. Manning, B. D. & Cantley, L. C. AKT/PKB signaling: navigating downstream. Cell 129, (2007). 34. Altomare, D. A. & Testa, J. R. Perturbations of the AKT signaling pathway in human cancer (2005). doi: /sj.onc Pópulo, H., JMl, L. & Soares, P. The mtor signalling pathway in human cancer. Int. J. Mol. Sci. 13, (2012). 36. Yasui, M., Matsuoka, S. & Ueda, M. PTEN hopping on the cell membrane is regulated via a positively-charged C2 domain. PLoS Comput. Biol. 10, e (2014). 37. Liaw, D., Marsh, D. J., Li, J., Dahia, P. L., Wang, S. I., Zheng, Z., Bose, S., Call, K. M., Tsou, H. C., Peacocke, M., Eng, C. & Parsons, R. Germline mutations of the PTEN gene in Cowden disease, an inherited breast and thyroid cancer syndrome. Nat. Genet. 16, 64 7 (1997). 38. Hobert, J. A. & Eng, C. PTEN hamartoma tumor syndrome: an overview. Genet. Med. 11, (2009). 39. Butler, M. G., Dasouki, M. J., Zhou, X.-P., Talebizadeh, Z., Brown, M., Takahashi, T. N., Miles, J. H., Wang, C. H., Stratton, R., Pilarski, R. & Eng, C. Subset of individuals with autism spectrum disorders and extreme macrocephaly associated with germline PTEN tumour suppressor gene mutations. J. Med. Genet. 42, (2005). 40. Salmena, L., Carracedo, A. & Pandolfi, P. P. Tenets of PTEN tumor suppression. Cell 133, (2008). 41. Sly, L. M., Rauh, M. J., Kalesnikoff, J., Büchse, T. & Krystal, G. SHIP, SHIP2, and PTEN activities are regulated in vivo by modulation of their protein levels: SHIP is upregulated in macrophages and mast cells by lipopolysaccharide. Exp. Hematol. 31, (2003). 42. Schurmans, S., Carrió, R., Behrends, J., Pouillon, V., Merino, J. & Clément, S. The mouse SHIP2 (Inppl1) gene: complementary DNA, genomic structure, promoter analysis, and gene expression in the embryo and adult mouse. Genomics 62, (1999). 43. Scheid, M. P., Huber, M., Damen, J. E., Hughes, M., Kang, V., Neilsen, P., Prestwich, G. D., Krystal, G. & Duronio, V. Phosphatidylinositol (3,4,5)P3 is essential but not sufficient for protein kinase B (PKB) activation; phosphatidylinositol (3,4)P2 is required for PKB phosphorylation at Ser-473: studies using cells from SH2-containing inositol-5- phosphatase knockout mice. J. Biol. Chem. 277, (2002). 44. Sharrard, R. M. & Maitland, N. J. Regulation of Protein Kinase B activity by PTEN and SHIP2 in human prostate-derived cell lines. Cell. Signal. 19, (2007). 76
89 45. Carver, D. J., Aman, M. J. & Ravichandran, K. S. SHIP inhibits Akt activation in B cells through regulation of Akt membrane localization. Blood 96, (2000). 46. Costinean, S., Sandhu, S. K., Pedersen, I. M., Tili, E., Trotta, R., Perrotti, D., Ciarlariello, D., Neviani, P., Harb, J., Kauffman, L. R., Shidham, A. & Croce, C. M. Src homology 2 domain-containing inositol-5-phosphatase and CCAAT enhancer-binding protein beta are targeted by mir-155 in B cells of Emicro-MiR-155 transgenic mice. Blood 114, (2009). 47. Lo, T. C. T., Barnhill, L. M., Kim, Y., Nakae, E. A., Yu, A. L. & Diccianni, M. B. Inactivation of SHIP1 in T-cell acute lymphoblastic leukemia due to mutation and extensive alternative splicing. Leuk. Res. 33, (2009). 48. Ooms, L. M., Binge, L. C., Davies, E. M., Rahman, P., Conway, J. R. W., Gurung, R., Ferguson, D. T., Papa, A., Fedele, C. G., Vieusseux, J. L., Chai, R. C., Koentgen, F., Price, J. T., Tiganis, T., Timpson, P., McLean, C. A. & Mitchell, C. A. The Inositol Polyphosphate 5-Phosphatase PIPP Regulates AKT1-Dependent Breast Cancer Growth and Metastasis. Cancer Cell 28, (2015). 49. Inhorn, R. C., Bansal, V. S. & Majerus, P. W. Pathway for inositol 1,3,4-trisphosphate and 1,4-bisphosphate metabolism. Proc. Natl. Acad. Sci. U. S. A. 84, (1987). 50. Bansal, V. S., Inhorn, R. C. & Majerus, P. W. The metabolism of inositol 1,3,4- trisphosphate to inositol 1,3-bisphosphate. J. Biol. Chem. 262, (1987). 51. Bansal, V. S., Caldwell, K. K. & Majerus, P. W. The Isolation and Characterization of Inositol Polyphosphate 4-Phosphatase. J. Biol. Chem. 265, (1990). 52. Norris, F. A. & Majerus, P. W. Hydrolysis of phosphatidylinositol 3,4-bisphosphate by inositol polyphosphate 4-phosphatase isolated by affinity elution chromatography. J. Biol. Chem. 269, (1994). 53. Norris, F. A., Atkins, R. & Majerus, P. The cdna Cloning and Characterization of Inositol Polyphosphate 4-Phosphatase Type II. EVIDENCE FOR CONSERVED ALTERNATIVE SPLICING IN THE 4-PHOSPHATASE FAMILY. J. Biol. Chem. 272, (1997). 54. Norris, F. A., Auethavekiat, V. & Majerus, P. The isolation and characterization of cdna encoding human and rat brain inositol polyphosphate 4-phosphatase. J. Biol. Chem. 272, (1995). 55. Pruitt, K., Brown, G., Tatusova, T. & Maglott, D. The Reference Sequence (RefSeq) Database. (2012). 56. Joseph, R. E., Walker, J. & Norris, F. A. Assignment of the inositol polyphosphate 4- phosphatase type I gene (INPP4A) to human chromosome band 2q11.2 by in situ hybridization. Cytogenet. Cell Genet. 87, (1999). 57. Fedele, C. G., Ooms, L. M., Ho, M., Vieusseux, J., O Toole, S. a, Millar, E. K., Lopez- Knowles, E., Sriratana, A., Gurung, R., Baglietto, L., Giles, G. G., Bailey, C. G., Rasko, J. E. J., Shields, B. J., Price, J. T., Majerus, P. W., Sutherland, R. L., Tiganis, T., McLean, C. a, et al. Inositol polyphosphate 4-phosphatase II regulates PI3K/Akt signaling and is lost 77
90 in human basal-like breast cancers. Proc. Natl. Acad. Sci. U. S. A. 107, (2010). 58. Ferron, M. & Vacher, J. Characterization of the murine Inpp4b gene and identification of a novel isoform. Gene 376, (2006). 59. Hakim, S., Bertucci, M. C., Conduit, S. E., Vuong, D. L. & Mitchell, C. A. Inositol polyphosphate phosphatases in human disease. Curr. Top. Microbiol. Immunol. 362, (2012). 60. Yates, A., Akanni, W., Amode, M. R., Barrell, D., Billis, K., Carvalho-Silva, D., Cummins, C., Clapham, P., Fitzgerald, S., Gil, L., Girón, C. G., Gordon, L., Hourlier, T., Hunt, S. E., Janacek, S. H., Johnson, N., Juettemann, T., Keenan, S., Lavidas, I., et al. Ensembl Nucleic Acids Res. 44, D710 6 (2016). 61. Cho, W. & Stahelin, R. V. Membrane binding and subcellular targeting of C2 domains. Biochim. Biophys. Acta - Mol. Cell Biol. Lipids 1761, (2006). 62. Liu, Y., Cheney, M. D., Gaudet, J. J., Chruszcz, M., Lukasik, S. M., Sugiyama, D., Lary, J., Cole, J., Dauter, Z., Minor, W., Speck, N. A. & Bushweller, J. H. The tetramer structure of the Nervy homology two domain, NHR2, is critical for AML1/ETO s activity. Cancer Cell 9, (2006). 63. Mukhopadhyay, R. & Rosen, B. P. The phosphatase C(X)5R motif is required for catalytic activity of the Saccharomyces cerevisiae Acr2p arsenate reductase. J. Biol. Chem. 276, (2001). 64. Lopez, S. M., Hodgson, M. C., Packianathan, C., Bingol-ozakpinar, O., Uras, F., Rosen, B. P. & Agoulnik, I. U. Determinants of the tumor suppressor INPP4B protein and lipid phosphatase activities. Biochem. Biophys. Res. Commun. 440, (2013). 65. Blanchetot, C. Substrate-trapping techniques in the identification of cellular PTP targets. Methods 35, 66. Franke, T. F., Kaplan, D. R., Cantley, L. C. & Tokert, A. Direct Regulation of the Akt Proto-Oncogene Product by Phosphatidylinositol-3, 4- bisphosphate Author ( s ): Thomas F. Franke, David R. Kaplan, Lewis C. Cantley and Alex Toker Published by : American Association for the Advancement of Science Stable. 275, (2016). 67. Marshall, A. J., Krahn, A. K., Ma, K., Duronio, V. & Hou, S. TAPP1 and TAPP2 Are Targets of Phosphatidylinositol 3-Kinase Signaling in B Cells: Sustained Plasma Membrane Recruitment Triggered by the B-Cell Antigen Receptor. Mol. Cell. Biol. 22, (2002). 68. Gewinner, C., Wang, Z. C., Richardson, A., Teruya-Feldstein, J., Etemadmoghadam, D., Bowtell, D., Barretina, J., Lin, W. M., Rameh, L., Salmena, L., Pandolfi, P. P. & Cantley, L. C. Evidence that inositol polyphosphate 4-phosphatase type II is a tumor suppressor that inhibits PI3K signaling. Cancer Cell 16, (2009). 69. Salmena, L., Shaw, P., Fans, I., McLaughlin, Rosen, B., Risch, H., Mitchell, C., Sun, P., Narod, S. A. & Kotsopoulos, J. Prognostic value of INPP4B protein immunohistochemistry in ovarian cancer. Eur. J. Gynaecol. Oncol. 36, (2015). 78
91 70. Chew, C. L., Lunardi, A., Gulluni, F., Ruan, D. T., Chen, M., Salmena, L., Nishino, M., Papa, A., Ng, C., Fung, J., Clohessy, J. G., Sasaki, J., Sasaki, T., Bronson, R. T., Hirsch, E. & Pandolfi, P. P. In vivo role of INPP4B in tumor and metastasis suppression through regulation of PI3K/AKT signaling at endosomes. Cancer Discov (2015). doi: / cd Stjernström, A., Karlsson, C., Fernandez, O. J., Söderkvist, P., Karlsson, M. G. & Thunell, L. K. Alterations of INPP4B, PIK3CA and pakt of the PI3K pathway are associated with squamous cell carcinoma of the lung. Cancer Med. 3, (2014). 72. Sasaki, J., Kofuji, S., Itoh, R., Momiyama, T., Takayama, K., Murakami, H., Chida, S., Tsuya, Y., Takasuga, S., Eguchi, S., Asanuma, K., Horie, Y., Miura, K., Davies, E. M., Mitchell, C., Yamazaki, M., Hirai, H., Takenawa, T., Suzuki, A., et al. The PtdIns(3,4)P(2) phosphatase INPP4A is a suppressor of excitotoxic neuronal death. Nature 465, (2010). 73. Sachs, A. J., David, S. a, Haider, N. B. & Nystuen, A. M. Patterned neuroprotection in the Inpp4a(wbl) mutant mouse cerebellum correlates with the expression of Eaat4. PLoS One 4, e8270 (2009). 74. Ivetac, I., Munday, A. D., Kisseleva, M. V., Zhang, X.-M., Luff, S., Tiganis, T., Whisstock, J. C., Rowe, T., Majerus, P. W. & Mitchell, C. A. The Type I Inositol Polyphosphate 4-Phosphatase Generates and Terminates Phosphoinositide 3-Kinase Signals on Endosomes and the Plasma Membrane. Mol. Biol. Cell 16, (2005). 75. Ferron, M., Boudiffa, M., Arsenault, M., Rached, M., Pata, M., Giroux, S., Elfassihi, L., Kisseleva, M., Majerus, P. W., Rousseau, F., Vacher, J., Asagiri, M., Sato, K., Usami, T., Ochi, S., Nishina, H., Yoshida, H., Morita, I., Wagner, E. F., et al. Inositol polyphosphate 4-phosphatase B as a regulator of bone mass in mice and humans. Cell Metab. 14, (2011). 76. Lemcke, S., Müller, S., Möller, S., Schillert, A., Ziegler, A., Cepok-Kauffeld, S., Comabella, M., Montalban, X., Rülicke, T., Nandakumar, K. S., Hemmer, B., Holmdahl, R., Pahnke, J., Ibrahim, S. M., Fernandez, V., Valls-Sole, J., Relova, J. L., Raguer, N., Miralles, F., et al. Nerve conduction velocity is regulated by the inositol polyphosphate-4- phosphatase II gene. Am. J. Pathol. 184, (2014). 77. Ji, L., Kim, N.-H., Huh, S.-O. & Rhee, and H. J. Depletion of Inositol Polyphosphate 4- Phosphatase II Suppresses Callosal Axon Formation in the Developing Mice. Mol. Cells 39, (2016). 78. Westbrook, T. F., Martin, E. S., Schlabach, M. R., Leng, Y., Liang, A. C., Feng, B., Zhao, J. J., Roberts, T. M., Mandel, G., Hannon, G. J., Depinho, R. a, Chin, L. & Elledge, S. J. A genetic screen for candidate tumor suppressors identifies REST. Cell 121, (2005). 79. Barnache, S., Le Scolan, E., Kosmider, O., Denis, N. & Moreau-Gachelin, F. Phosphatidylinositol 4-phosphatase type II is an erythropoietin-responsive gene. Oncogene 25, (2006). 80. Naylor, T. L., Greshock, J., Wang, Y., Colligon, T., Yu, Q. C., Clemmer, V., Zaks, T. Z. & Weber, B. L. High resolution genomic analysis of sporadic breast cancer using array- 79
92 based comparative genomic hybridization. Breast Cancer Res. 7, R (2005). 81. TCGA. Comprehensive molecular portraits of human breast tumours. Nature 490, (2012). 82. Hodgson, M. C., Shao, L., Frolov, A., Li, R., Peterson, L. E., Ayala, G., Ittmann, M. M., Weigel, N. L. & Agoulnik, I. U. Decreased expression and androgen regulation of the tumor suppressor gene INPP4B in prostate cancer. Cancer Res. 71, (2011). 83. Rynkiewicz, N. K., Fedele, C. G., Chiam, K., Gupta, R., Kench, J. G., Ooms, L. M., McLean, C. a, Giles, G. G., Horvath, L. G. & Mitchell, C. a. INPP4B is highly expressed in prostate intermediate cells and its loss of expression in prostate carcinoma predicts for recurrence and poor long term survival. Prostate (2014). doi: /pros Hodgson, M. C., Deryugina, E. I., Suarez, E., Lopez, S. M., Lin, D., Xue, H., Gorlov, I. P., Wang, Y., Agoulnik, I. U., Thurairaja, R., McFarlane, J., Traill, Z., Persad, R., Morgan, T., Lange, P., Porter, M., Lin, D., Ellis, W., Gallaher, I., et al. INPP4B suppresses prostate cancer cell invasion. Cell Commun. Signal. 12, 61 (2014). 85. Perez-Lorenzo, R., Gill, K. Z., Shen, C.-H., Zhao, F. X., Zheng, B., Schulze, H.-J., Silvers, D. N., Brunner, G., Horst, B. A., Balch, C. M., Gershenwald, J. E., Soong, S. J., al., et, Cantley, L. C., Chin, L., Garraway, L. A., Fisher, D. E., Clement, S., Krause, U., et al. A Tumor Suppressor Function for the Lipid Phosphatase INPP4B in Melanocytic Neoplasms. J. Invest. Dermatol. 134, (2014). 86. Hsu, I., Yeh, C.-R., Slavin, S., Miyamoto, H., Netto, G., Tsai, Y.-C., Muyan, M., Wu, X.- R., Yeh, S., Hsu, I., Yeh, C.-R., Slavin, S., Miyamoto, H., Netto, G., Tsai, Y.-C., Muyan, M., Wu, X.-R. & Yeh, S. Estrogen receptor alpha prevents bladder cancer via INPP4B inhibited akt pathway in vitro and in vivo. Oncotarget 5, (2014). 87. Yuen, J. W.-F., Chung, G. T.-Y., Lun, S. W.-M., Cheung, C. C.-M., To, K.-F., Lo, K.-W., Manning, B., Cantley, B., Engelman, J., Liu, P., Cheng, H., Roberts, T., Zhao, J., Bertucci, M., Mitchell, C., Gewinner, C., Wang, Z., Richardson, A., Teruya-Feldstein, J., et al. Epigenetic Inactivation of Inositol polyphosphate 4-phosphatase B (INPP4B), a Regulator of PI3K/AKT Signaling Pathway in EBV-Associated Nasopharyngeal Carcinoma. PLoS One 9, e (2014). 88. Kim, J.-S., Yun, H. S., Um, H.-D., Park, J. K., Lee, K.-H., Kang, C.-M., Lee, S.-J. & Hwang, S.-G. Identification of inositol polyphosphate 4-phosphatase type II as a novel tumor resistance biomarker in human laryngeal cancer HEp-2 cells. Cancer Biol. Ther. 13, (2012). 89. Min, J. W., Kim, K. Il, Kim, H.-A., Kim, E.-K., Noh, W. C., Jeon, H. B., Cho, D.-H., Oh, J. S., Park, I.-C., Hwang, S.-G. & Kim, J.-S. INPP4B-mediated tumor resistance is associated with modulation of glucose metabolism via hexokinase 2 regulation in laryngeal cancer cells. Biochem. Biophys. Res. Commun. 2 7 (2013). doi: /j.bbrc Ross, M. E., Zhou, X., Song, G., Shurtleff, S. A., Girtman, K., Williams, W. K., Liu, H.- C., Mahfouz, R., Raimondi, S. C., Lenny, N., Patel, A. & Downing, J. R. Classification of pediatric acute lymphoblastic leukemia by gene expression profiling. Blood 102,
93 (2003). 91. Dzneladze, I., He, R., Woolley, J. F., Son, M. H., Sharobim, M. H., Greenberg, S. A., Gabra, M., Langlois, C., Rashid, A., Hakem, A., Ibrahimova, N., Arruda, A., Löwenberg, B., Valk, P. J. M., Minden, M. D. & Salmena, L. INPP4B overexpression is associated with poor clinical outcome and therapy resistance in acute myeloid leukemia. Leukemia 29, (2015). 92. Rijal, S., Fleming, S., Cummings, N., Rynkiewicz, N. K., Ooms, L. M., Nguyen, N.-Y. N., Teh, T.-C., Avery, S., McManus, J. F., Papenfuss, A. T., McLean, C., Guthridge, M. A., Mitchell, C. A. & Wei, A. H. Inositol polyphosphate 4-phosphatase II (INPP4B) is associated with chemoresistance and poor outcome in AML. Blood 125, (2015). 93. Vasudevan, K. M., Barbie, D. A., Davies, M. A., Rabinovsky, R., McNear, C. J., Kim, J. J., Hennessy, B. T., Tseng, H., Pochanard, P., Kim, S. Y., Dunn, I. F., Schinzel, A. C., Sandy, P., Hoersch, S., Sheng, Q., Gupta, P. B., Boehm, J. S., Reiling, J. H., Silver, S., et al. AKT-independent signaling downstream of oncogenic PIK3CA mutations in human cancer. Cancer Cell 16, (2009). 94. Tessier, M. & Woodgett, J. R. Serum and glucocorticoid-regulated protein kinases: Variations on a theme. J. Cell. Biochem. 98, (2006). 95. Gasser, J. A., Inuzuka, H., Lau, A. W., Wei, W., Beroukhim, R. & Toker, A. SGK3 Mediates INPP4B-Dependent PI3K Signaling in Breast Cancer. Mol. Cell 1 13 (2014). doi: /j.molcel Gasser, J. A., Inuzuka, H., Lau, A. W., Wei, W., Beroukhim, R. & Toker, A. SGK3 Mediates INPP4B-Dependent PI3K Signaling in Breast Cancer. Mol. Cell 56, (2014). 97. Guo, S. T., Chi, M. N., Yang, R. H., Guo, X. Y., Zan, L. K., Wang, C. Y., Xi, Y. F., Jin, L., Croft, A., Tseng, H.-Y., Yan, X. G., Farrelly, M., Wang, F. H., Lai, F., Wang, J. F., Li, Y. P., Ackland, S., Scott, R., Agoulnik, I. U., et al. INPP4B is an oncogenic regulator in human colon cancer. Oncogene 1 13 (2015). doi: /onc Ross, A. & Gericke, A. Phosphorylation keeps PTEN phosphatase closed for business. Proc. Natl. Acad. Sci. U. S. A. 106, (2009). 99. Vazquez, F., Ramaswamy, S., Nakamura, N. & Sellers, W. R. Phosphorylation of the PTEN Tail Regulates Protein Stability and Function. Mol. Cell. Biol. 20, (2000) Fragoso, R. & Barata, J. T. Kinases, tails and more: Regulation of PTEN function by phosphorylation. Methods 77, (2015) Chi, M. N., Guo, S. T., Wilmott, J. S., Guo, X. Y., Yan, X. G., Wang, C. Y., Liu, X. Y., Jin, L., Tseng, H.-Y., Liu, T., Croft, A., Hondermarck, H., Scolyer, R. A., Jiang, C. C., Zhang, X. D., Chi, M. N., Tang Guo, S., Wilmott, J. S., Yun Guo, X., et al. INPP4B is upregulated and functions as an oncogenic driver through SGK3 in a subset of melanomas. Oncotarget 6, (2015) Siggs, O. M., Smart, N. G. & Beutler, B. Record for woolly, updated May 13, at 81
94 <mutagenetix.utsouthwestern.edu> 103. Hodgson, M. C., Deryugina, E. I., Suarez, E., Lopez, S. M., Lin, D., Xue, H., Gorlov, I. P., Wang, Y. & Agoulnik, I. U. INPP4B suppresses prostate cancer cell invasion. Cell Commun. Signal. 12, 61 (2014) Rynkiewicz, N. K. INPP4B is highly expressed in prostate intermediate cells and its loss of expression in prostate carcinoma predicts for recurrence and poor long term survival. 75, 105. Min, J. W., Kim, K. Il, Kim, H.-A., Kim, E.-K., Noh, W. C., Jeon, H. B., Cho, D.-H., Oh, J. S., Park, I.-C., Hwang, S.-G. & Kim, J.-S. INPP4B-mediated tumor resistance is associated with modulation of glucose metabolism via hexokinase 2 regulation in laryngeal cancer cells. Biochem. Biophys. Res. Commun. 440, (2013) Chi, M. N., Guo, S. T., Wilmott, J. S., Guo, X. Y., Yan, X. G., Wang, C. Y., Ying Liu, X., Jin, L., Tseng, H.-Y., Liu, T., Croft, A., Hondermarck, H., Scolyer, R. A., Jiang, C. C. & Zhang, X. D. INPP4B is upregulated and functions as an oncogenic driver through SGK3 in a subset of melanomas. Oncotarget 6, (2015) Dzneladze, I., He, R., Woolley, J. F., Son, M. H., Sharobim, M. H., Greenberg, S. A., Gabra, M., Langlois, C., Rashid, A., Hakem, A., Ibrahimova, N., Arruda, A., Löwenberg, B., Valk, P. J. M., Minden, M. D. & Salmena, L. INPP4B overexpression is associated with poor clinical outcome and therapy resistance in acute myeloid leukemia. Leukemia 29, (2015) Shipley, J. L. & Butera, J. N. Acute myelogenous leukemia. Exp. Hematol. 37, (2009) Estey, E. & Döhner, H. Acute myeloid leukaemia. Lancet 368, (2006) Jemal, A., Siegel, R., Ward, E., Murray, T., Xu, J. & Thun, M. J. Cancer statistics, CA. Cancer J. Clin. 57, Deschler, B. & Lübbert, M. Acute myeloid leukemia: Epidemiology and etiology. Cancer 107, (2006) Döhner, H., Estey, E. H. E., Amadori, S., Appelbaum, F. R. F. R., Büchner, T., Burnett, A. K. a. K., Dombret, H., Fenaux, P., Grimwade, D., Larson, R. a. R. A., Lo-Coco, F., Naoe, T., Niederwieser, D., Ossenkoppele, G. J., Sanz, M. A., Sierra, J., Tallman, M. S., Löwenberg, B., Bloomfield, C. D., et al. Diagnosis and management of acute myeloid leukemia in adults: recommendations from an international expert panel, on behalf of the European LeukemiaNet. Blood 115, (2010) Estey, E. & Döhner, H. Acute myeloid leukaemia. Lancet 368, (2006) Löwenberg, B., Griffin, J. D. & Tallman, M. S. Acute myeloid leukemia and acute promyelocytic leukemia. Hematology Am. Soc. Hematol. Educ. Program (2003). at < Gewirtz, D. A. A critical evaluation of the mechanisms of action proposed for the antitumor effects of the anthracycline antibiotics adriamycin and daunorubicin. Biochem. 82
95 Pharmacol. 57, (1999) Burden, D. A. & Osheroff, N. Mechanism of action of eukaryotic topoisomerase II and drugs targeted to the enzyme. Biochim. Biophys. Acta - Gene Struct. Expr. 1400, (1998) Willmore, E., de Caux, S., Sunter, N. J., Tilby, M. J., Jackson, G. H., Austin, C. A. & Durkacz, B. W. A novel DNA-dependent protein kinase inhibitor, NU7026, potentiates the cytotoxicity of topoisomerase II poisons used in the treatment of leukemia. Blood 103, (2004) Kaina, B. DNA damage-triggered apoptosis: critical role of DNA repair, double-strand breaks, cell proliferation and signaling. Biochem. Pharmacol. 66, (2003) Savani, B. N., Mielke, S., Reddy, N., Goodman, S., Jagasia, M. & Rezvani, K. Management of relapse after allo-sct for AML and the role of second transplantation. Bone Marrow Transplant. 44, (2009) Kell, J. Emerging treatments in acute myeloid leukaemia. Expert Opin. Emerg. Drugs 9, (2004) Lapidot, T., Sirard, C., Vormoor, J., Murdoch, B., Hoang, T., Caceres-Cortes, J., Minden, M., Paterson, B., Caligiuri, M. A. & Dick, J. E. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367, (1994) Grandage, V. L., Gale, R. E., Linch, D. C. & Khwaja, A. PI3-kinase/Akt is constitutively active in primary acute myeloid leukaemia cells and regulates survival and chemoresistance via NF-kappaB, Mapkinase and p53 pathways. Leukemia 19, (2005) Xu, Q., Simpson, S.-E., Scialla, T. J., Bagg, A. & Carroll, M. Survival of acute myeloid leukemia cells requires PI3 kinase activation. Blood 102, (2003) Kornblau, S. M., Womble, M., Qiu, Y. H., Jackson, C. E., Chen, W., Konopleva, M., Estey, E. H. & Andreeff, M. Simultaneous activation of multiple signal transduction pathways confers poor prognosis in acute myelogenous leukemia. Blood 108, (2006) Zhao, S., Konopleva, M., Cabreira-Hansen, M., Xie, Z., Hu, W., Milella, M., Estrov, Z., Mills, G. B. & Andreeff, M. Inhibition of phosphatidylinositol 3-kinase dephosphorylates BAD and promotes apoptosis in myeloid leukemias. Leukemia 18, (2004) Sandhöfer, N., Metzeler, K. H., Rothenberg, M., Herold, T., Tiedt, S., Groiß, V., Carlet, M., Walter, G., Hinrichsen, T., Wachter, O., Grunert, M., Schneider, S., Subklewe, M., Dufour, a, Fröhling, S., Klein, H.-G., Hiddemann, W., Jeremias, I. & Spiekermann, K. Dual PI3K/mTOR inhibition shows antileukemic activity in MLL-rearranged acute myeloid leukemia. Leukemia 1 11 (2014). doi: /leu Park, S., Chapuis, N., Tamburini, J., Bardet, V., Cornillet-Lefebvre, P., Willems, L., Green, A., Mayeux, P., Lacombe, C. & Bouscary, D. Role of the PI3K/AKT and mtor signaling pathways in acute myeloid leukemia. Haematologica 95, (2010). 83
96 128. Levis, M. FLT3 mutations in acute myeloid leukemia: what is the best approach in 2013? Hematology Am. Soc. Hematol. Educ. Program 2013, (2013) Grossmann, V., Schnittger, S., Poetzinger, F., Kohlmann, A., Stiel, A., Eder, C., Fasan, A., Kern, W., Haferlach, T. & Haferlach, C. High incidence of RAS signalling pathway mutations in MLL-rearranged acute myeloid leukemia. Leukemia 27, (2013) Lavallée, V.-P., Baccelli, I., Krosl, J., Wilhelm, B., Barabé, F., Gendron, P., Boucher, G., Lemieux, S., Marinier, A., Meloche, S., Hébert, J. & Sauvageau, G. The transcriptomic landscape and directed chemical interrogation of MLL-rearranged acute myeloid leukemias. Nat. Genet. 47, (2015) Kampen, K., Ter Elst, A., Mahmud, H., Scherpen, F., Diks, S., Peppelenbosch, M., De Haas, V., Guryev, V. & De Bont, E. Insights in dynamic kinome reprogramming as a consequence of MEK inhibition in MLL-rearranged AML. Leukemia 28, (2013) Levine, B. & Kroemer, G. Autophagy in the Pathogenesis of Disease. Cell 132, (2008) Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, (2008) Li, W. W., Li, J. & Bao, J. K. Microautophagy: Lesser-known self-eating. Cell. Mol. Life Sci. 69, (2012) Mortimore, G. E., Lardeux, B. R. & Adams, C. E. Regulation of microautophagy and basal protein turnover in rat liver. Effects of short-term starvation. J. Biol. Chem. 263, (1988) Mijaljica, D., Prescott, M. & Devenish, R. J. Microautophagy in mammalian cells: Revisiting a 40-year-old conundrum. Autophagy 7, (2011) Knecht, E. & Salvador, N. Chaperone-Mediated Autophagy. (Landes Bioscience). at < Dice, J. F., Chiang, H. L., Spencer, E. P. & Backer, J. M. Regulation of catabolism of microinjected ribonuclease A. Identification of residues 7-11 as the essential pentapeptide. J. Biol. Chem. 261, (1986) Bejarano, E. & Cuervo, A. M. Chaperone-Mediated Autophagy. 34, (Landes Bioscience, 2010) Fred Dice, J. Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem. Sci. 15, (1990) Boya, P., Reggiori, F. & Codogno, P. Emerging regulation and functions of autophagy. Nat. Cell Biol. 15, (2013) Li, L., Chen, Y. & Gibson, S. B. Starvation-induced autophagy is regulated by mitochondrial reactive oxygen species leading to AMPK activation. Cell. Signal. 25, (2013) Tooze, S. A. & Yoshimori, T. The origin of the autophagosomal membrane. Nat. Cell 84
97 Biol. 12, (2010) Burman, C. & Ktistakis, N. T. Regulation of autophagy by phosphatidylinositol 3- phosphate. FEBS Lett. 584, (2010) Roberts, R. & Ktistakis, N. T. Omegasomes: PI3P platforms that manufacture autophagosomes. Essays Biochem. 55, (2013) Axe, E. L., Walker, S. A., Manifava, M., Chandra, P., Roderick, H. L., Habermann, A., Griffiths, G. & Ktistakis, N. T. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 182, (2008) Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A, Adachi H, Adams CM, Adams PD, Adeli K, Adhihetty PJ, Adler SG, Agam G, Agarwal R, Aghi MK, Agnello M, Agostinis P, Aguilar PV, Aguirre-Ghiso J, Airoldi EM, Ait-Si- Ali S, Akemat, Z. S. Guidelines for use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, (2016) Barth, S., Glick, D. & Macleod, K. F. Autophagy: Assays and artifacts. J. Pathol. 221, (2010) Mizushima, N., Yoshimori, T. & Levine, B. Methods in mammalian autophagy research. Cell 140, (2010) Thorburn, A., Thamm, D. H. & Gustafson, D. L. Autophagy and Cancer Therapy. Mol. Pharmacol. 85, (2014) Juhasz, G. Interpretation of bafilomycin, ph neutralizing or protease inhibitor treatments in autophagic flux experiments: Novel considerations. Autophagy 8, (2012) Kaur, J. & Debnath, J. Autophagy at the crossroads of catabolism and anabolism. Nat. Rev. Mol. Cell Biol. 16, (2015) Hardie, D. G. AMP-activated/SNF1 protein kinases: conserved guardians of cellular energy. Nat. Rev. Mol. Cell Biol. 8, (2007) Gwinn, D. M., Shackelford, D. B., Egan, D. F., Mihaylova, M. M., Mery, A., Vasquez, D. S., Turk, B. E. & Shaw, R. J. AMPK Phosphorylation of Raptor Mediates a Metabolic Checkpoint. Mol. Cell 30, (2008) Kim, J., Kundu, M., Viollet, B. & Guan, K.-L. AMPK and mtor regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, (2011) Vander Haar, E., Lee, S.-I., Bandhakavi, S., Griffin, T. J. & Kim, D.-H. Insulin signalling to mtor mediated by the Akt/PKB substrate PRAS40. Nat. Cell Biol. 9, (2007) Jung, C. H., Ro, S.-H., Cao, J., Otto, N. M. & Kim, D.-H. mtor regulation of autophagy. FEBS Lett. 584, (2010) Chan, E. Y. W., Kir, S. & Tooze, S. A. sirna screening of the kinome identifies ULK1 as a multidomain modulator of autophagy. J. Biol. Chem. 282, (2007). 85
98 159. Russell, R. C., Tian, Y., Yuan, H., Park, H. W., Chang, Y.-Y., Kim, J., Kim, H., Neufeld, T. P., Dillin, A. & Guan, K.-L. ULK1 induces autophagy by phosphorylating Beclin-1 and activating VPS34 lipid kinase. Nat. Cell Biol. 15, (2013) Burman, C. & Ktistakis, N. T. Regulation of autophagy by phosphatidylinositol 3- phosphate. FEBS Lett. 584, (2010) Simonsen, A. & Tooze, S. A. Coordination of membrane events during autophagy by multiple class III PI3-kinase complexes. J. Cell Biol. 186, (2009) Blommaart, E. F., Krause, U., Schellens, J. P., Vreeling-Sindelárová, H. & Meijer, A. J. The phosphatidylinositol 3-kinase inhibitors wortmannin and LY inhibit autophagy in isolated rat hepatocytes. Eur. J. Biochem. 243, (1997) Obara, K., Noda, T., Niimi, K. & Ohsumi, Y. Transport of phosphatidylinositol 3- phosphate into the vacuole via autophagic membranes in Saccharomyces cerevisiae. Genes Cells 13, (2008) Juhász, G., Hill, J. H., Yan, Y., Sass, M., Baehrecke, E. H., Backer, J. M. & Neufeld, T. P. The class III PI(3)K Vps34 promotes autophagy and endocytosis but not TOR signaling in Drosophila. J. Cell Biol. 181, (2008) Aita, V. M., Liang, X. H., Murty, V. V. V. S., Pincus, D. L., Yu, W., Cayanis, E., Kalachikov, S., Gilliam, T. C. & Levine, B. Cloning and Genomic Organization of Beclin 1, a Candidate Tumor Suppressor Gene on Chromosome 17q21. Genomics 59, (1999) Yue, Z., Jin, S., Yang, C., Levine, A. J. & Heintz, N. Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc. Natl. Acad. Sci. U. S. A. 100, (2003) Takamura, A., Komatsu, M., Hara, T., Sakamoto, A., Kishi, C., Waguri, S., Eishi, Y., Hino, O., Tanaka, K. & Mizushima, N. Autophagy-deficient mice develop multiple liver tumors. Genes Dev. 25, (2011) Chen, Y., Lu, Y., Lu, C. & Zhang, L. Beclin-1 expression is a predictor of clinical outcome in patients with esophageal squamous cell carcinoma and correlated to hypoxiainducible factor (HIF)-1alpha expression. Pathol. Oncol. Res. 15, (2009) Rabinowitz, J. D. & White, E. Autophagy and metabolism. Science 330, (2010) Karantza-Wadsworth, V., Patel, S., Kravchuk, O., Chen, G., Mathew, R., Jin, S. & White, E. Autophagy mitigates metabolic stress and genome damage in mammary tumorigenesis. Genes Dev. 21, (2007) Kundu, M., Lindsten, T., Yang, C.-Y., Wu, J., Zhao, F., Zhang, J., Selak, M. A., Ney, P. A. & Thompson, C. B. Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood 112, (2008) Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: The next generation. Cell 144, (2011). 86
99 173. White, E. The Role for Autophagy in Cancer. 125, (2015) Guo, J. Y., Chen, H.-Y., Mathew, R., Fan, J., Strohecker, A. M., Karsli-Uzunbas, G., Kamphorst, J. J., Chen, G., Lemons, J. M. S., Karantza, V., Coller, H. A., Dipaola, R. S., Gelinas, C., Rabinowitz, J. D. & White, E. Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev. 25, (2011) Yang, S., Wang, X., Contino, G., Liesa, M., Sahin, E., Ying, H., Bause, A., Li, Y., Stomme, J. M., Dell Antonio, G., Mautner, J., Tonon, G., Haigis, M., Shirihai, O. S., Doglioni, C., Bardeesy, N. & Kimmelman, A. C. Pancreatic cancers require autophagy for tumor growth. Genes Dev. 25, (2011) Sehgal, A. R., Konig, H., Johnson, D. E., Tang, D., Amaravadi, R. K., Boyiadzis, M. & Lotze, M. T. You eat what you are: autophagy inhibition as a therapeutic strategy in leukemia. Leukemia 29, (2015) Chittaranjan, S., Bortnik, S., Dragowska, W. H., Xu, J., Abeysundara, N., Leung, A., Go, N. E., DeVorkin, L., Weppler, S. A., Gelmon, K., Yapp, D. T., Bally, M. B. & Gorski, S. M. Autophagy inhibition augments the anticancer effects of epirubicin treatment in anthracycline-sensitive and -resistant triple-negative breast cancer. Clin. Cancer Res. 20, (2014) Selvakumaran, M., Amaravadi, R. K., Vasilevskaya, I. A. & O Dwyer, P. J. Autophagy inhibition sensitizes colon cancer cells to antiangiogenic and cytotoxic therapy. Clin. Cancer Res. 19, (2013) Ma, X.-H., Piao, S.-F., Dey, S., Mcafee, Q., Karakousis, G., Villanueva, J., Hart, L. S., Levi, S., Hu, J., Zhang, G., Lazova, R., Klump, V., Pawelek, J. M., Xu, X., Xu, W., Schuchter, L. M., Davies, M. A., Herlyn, M., Winkler, J., et al. Targeting ER stress induced autophagy overcomes BRAF inhibitor resistance in melanoma. J. Clin. Invest. 124, (2014) Wang, J. & Wu, G. S. Role of Autophagy in Cisplatin Resistance in Ovarian Cancer Cells. J. Biol. Chem. 289, (2014) Wilson, A., Laurenti, E., Oser, G., van der Wath, R. C., Blanco-Bose, W., Jaworski, M., Offner, S., Dunant, C. F., Eshkind, L., Bockamp, E., Lió, P., Macdonald, H. R., Trumpp, A., Adolfsson, J., Borge, O. J., Bryder, D., Theilgaard-Monch, K., Astrand-Grundstrom, I., Sitnicka, E., et al. Hematopoietic stem cells reversibly switch from dormancy to selfrenewal during homeostasis and repair. Cell 135, (2008) Guan, J., Simon, A. K., Prescott, M., Menendez, J. A., Wang, F., Wang, C., Wolvetang, E., Vazquez-martin, A., Simon, A. K., Prescott, M., Menendez, J. A., Wang, F., Wang, C., Wolvetang, E., Vazquez-martin, A., Zhang, J., Guan, J., Simon, A. K., Prescott, M., et al. Autophagy in stem cells. 8627, (2015) Mortensen, M., Soilleux, E. J., Djordjevic, G., Tripp, R., Lutteropp, M., Sadighi-Akha, E., Stranks, A. J., Glanville, J., Knight, S., W. Jacobsen, S.-E., Kranc, K. R. & Simon, A. K. The autophagy protein Atg7 is essential for hematopoietic stem cell maintenance. J. Exp. Med. 208, (2011). 87
100 184. Warr, M. R., Binnewies, M., Flach, J., Reynaud, D., Garg, T., Malhotra, R., Debnath, J. & Passegué, E. FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature 494, (2013) Kang, Y.-A., Sanalkumar, R., O Geen, H., Linnemann, A. K., Chang, C.-J., Bouhassira, E. E., Farnham, P. J., Keles, S. & Bresnick, E. H. Autophagy driven by a master regulator of hematopoiesis. Mol. Cell. Biol. 32, (2012) Willems, L., Chapuis, N., Puissant, A., Maciel, T. T., Green, A. S., Jacque, N., Vignon, C., Park, S., Guichard, S., Herault, O., Fricot, A., Hermine, O., Moura, I. C., Auberger, P., Ifrah, N., Dreyfus, F., Bonnet, D., Lacombe, C., Mayeux, P., et al. The dual mtorc1 and mtorc2 inhibitor AZD8055 has anti-tumor activity in acute myeloid leukemia. Leukemia 26, (2012) Willems, L., Jacque, N., Jacquel, A., Neveux, N., Maciel, T. T., Schmitt, A., Poulain, L., Green, A. S., Uzunov, M., Kosmider, O., Moura, I. C., Auberger, P., Ifrah, N., Bardet, V., Chapuis, N., Lacombe, C., Mayeux, P., Tamburini, J., Bouscary, D., et al. Inhibiting glutamine uptake represents an attractive new strategy for treating acute myeloid leukemia Inhibiting glutamine uptake represents an attractive new strategy for treating acute myeloid leukemia. 122, (2013) de Thé, H. & Chen, Z. Acute promyelocytic leukaemia: novel insights into the mechanisms of cure. Nat. Rev. Cancer 10, (2010) Shen, Z.-X., Shi, Z.-Z., Fang, J., Gu, B.-W., Li, J.-M., Zhu, Y.-M., Shi, J.-Y., Zheng, P.- Z., Yan, H., Liu, Y.-F., Chen, Y., Shen, Y., Wu, W., Tang, W., Waxman, S., De Thé, H., Wang, Z.-Y., Chen, S.-J. & Chen, Z. All-trans retinoic acid/as2o3 combination yields a high quality remission and survival in newly diagnosed acute promyelocytic leukemia. Proc. Natl. Acad. Sci. U. S. A. 101, (2004) Isakson, P., Bjørås, M., Bøe, S. O. & Simonsen, A. Autophagy contributes to therapyinduced degradation of the PML/RARA oncoprotein. Blood 116, (2010) Wang, Z., Cao, L., Kang, R., Yang, M., Liu, L., Zhao, Y., Yu, Y., Xie, M., Yin, X., Livesey, K. M. & Tang, D. Autophagy regulates myeloid cell differentiation by p62/sqstm1-mediated degradation of PML-RARα oncoprotein. Autophagy 7, (2011) Goussetis, D. J., Gounaris, E., Wu, E. J., Vakana, E., Sharma, B., Bogyo, M., Altman, J. K. & Platanias, L. C. Autophagic degradation of the BCR-ABL oncoprotein and generation of antileukemic responses by arsenic trioxide. Blood 120, (2012) Sagona, A. P., Nezis, I. P., Pedersen, N. M., Liestøl, K., Poulton, J., Rusten, T. E., Skotheim, R. I., Raiborg, C. & Stenmark, H. PtdIns(3)P controls cytokinesis through KIF13A-mediated recruitment of FYVE-CENT to the midbody. Nat. Cell Biol. 12, (2010) Jean, S., Cox, S., Schmidt, E. J., Robinson, F. L. & Kiger, A. Sbf/MTMR13 coordinates PI(3)P and Rab21 regulation in endocytic control of cellular remodeling. Mol. Biol. Cell 23, (2012). 88
101 195. Martinez, J., Malireddi, R. K. S., Lu, Q., Cunha, L. D., Pelletier, S., Gingras, S., Orchard, R., Guan, J.-L., Tan, H., Peng, J., Kanneganti, T.-D., Virgin, H. W. & Green, D. R. Molecular characterization of LC3-associated phagocytosis reveals distinct roles for Rubicon, NOX2 and autophagy proteins. Nat. Cell Biol. 17, (2015) Mack, H. I. D. Dissertation: Regulation of Mammalian Autophagy by Unc-51-Like Kinase 1. (TECHNISCHE UNIVERSITÄT MÜNCHEN, 2011, 2011) Sun, H. & Taneja, R. Analysis of Transformation and Tumorigenicity Using Mouse Embryonic Fibroblast Cells. Cancer Genomics Proteomics Methods Protoc. 383, Strauss, W. M. & Strauss, W. M. in Current Protocols in Molecular Biology (John Wiley & Sons, Inc., 2001). doi: / mb0202s Agoulnik, I. U., Hodgson, M. C., Bowden, W. A., Ittmann, M. M., Agoulnik, I. U., Hodgson, M. C., Bowden, W. A. & Ittmann, M. M. INPP4B: the New Kid on the PI3K Block. Oncotarget 2, (2011) Chan, L. L.-Y., Shen, D., Wilkinson, A. R., Patton, W., Lai, N., Chan, E., Kuksin, D., Lin, B. & Qiu, J. A novel image-based cytometry method for autophagy detection in living cells. Autophagy 8, (2012) Amaravadi, R. K., Yu, D., Lum, J. J., Bui, T., Christophorou, M. A., Evan, G. I., Thomas- Tikhonenko, A., Thompson, C. B., Kanzawa, T., Paglin, S., Bursch, W., Kondo, Y., Kanzawa, T., Sawaya, R., Kondo, S., Yu, L., Qu, X., Kiffin, R., Christian, C., et al. Autophagy inhibition enhances therapy-induced apoptosis in a Myc-induced model of lymphoma. J. Clin. Invest. 117, (2007) Tiacci, E., Spanhol-Rosseto, a, Martelli, M. P., Pasqualucci, L., Quentmeier, H., Grossmann, V., Drexler, H. G. & Falini, B. The NPM1 wild-type OCI-AML2 and the NPM1-mutated OCI-AML3 cell lines carry DNMT3A mutations. Leukemia 26, (2012) Peng, C., Chen, Y., Yang, Z., Zhang, H., Osterby, L., Rosmarin, A. G., Dc, W., Peng, C., Chen, Y., Yang, Z., Zhang, H., Osterby, L., Rosmarin, A. G. & Li, S. leukemias in mice PTEN is a tumor suppressor in CML stem cells and BCR-ABL induced leukemias in mice. 115, (2013) Gutierrez, A., Sanda, T., Grebliunaite, R., Carracedo, A., Salmena, L., Ahn, Y., Dahlberg, S., Neuberg, D., Moreau, L. a, Winter, S. S., Larson, R., Zhang, J., Protopopov, A., Chin, L., Pandolfi, P. P., Silverman, L. B., Hunger, S. P., Sallan, S. E. & Look, a T. Brief report High frequency of PTEN, PI3K, and AKT abnormalities in T-cell acute lymphoblastic leukemia. Science (80-. ). 114, (2009) Woolley, J. F., Dzneladze, I. & Salmena, L. Phosphoinositide signaling in cancer: INPP4B Akt(s) out. Trends Mol. Med. 21, (2015) Devereaux, K., Dall Armi, C., Alcazar-Roman, A., Ogasawara, Y., Zhou, X., Wang, F., Yamamoto, A., de Camilli, P. & Di Paolo, G. Regulation of Mammalian Autophagy by Class II and III PI 3-Kinases through PI3P Synthesis. PLoS One 8, (2013) Mathew, R., Karantza-Wadsworth, V. & White, E. Role of autophagy in cancer. Nat. Rev. 89
102 Cancer 7, (2007) Klionsky, D. J., Eskelinen, E. L. & Deretic, V. Autophagosomes, phagosomes, autolysosomes, phagolysosomes, autophagolysosomes... Wait, I m confused. Autophagy 10, (2014). 90
103 APPENDICES Appendix 1. INPP4B high AML patients have lower CR rates and shorter survival. (Adapted from Dzneladze et al. 2015, Leukemia 91 ). (A) INPP4B expression is aligned with patient response to therapy. (B-D) Response to therapy and test of significance. CR = complete response, NR = no response (E-H) Kaplan-Meier plots for INPP4B high patient OS. 91
104 Appendix 2. INPP4B high constitutes a significant hazard in total and CN-AML (Adapted from Dzneladze et al. 2015, Leukemia 91 ). (A-B) OS and EFS of patients with INPP4B High or FLT3-ITD versus all patients. (C) Forest plot of log hazard rates for OS in AML patients within OCI-PMH data sets. (D) ROC analysis of EFS for potential biomarkers in AML vs. INPP4B (E-H) Comparison of INPP4B High and INPP4B Low OS and EFS in cytogenetically normal AML or FLT3-ITD AMLversus total patients in OCI- PMH dataset. 92
105 Appendix 3. Ectopic overexpression of INPP4B in AML cells leads to increased colony-forming potential and proliferation. (Adapted from Dzneladze et al. 2015, Leukemia 91 ). (A-B) Ectopic overexpression of INPP4B in OCI-AML2 and OCI-AML3 cell lines at the mrna and protein level. (C-D) Colony formation for INPP4B wt and control OCI-AML2 and OCI-AML3 cells in methylcellulose. (E-F) Proliferation assay and Annexin V staining of INPP4B wt and control cells OCI-AML2 and OCI-AML3 cells. (G-H) Low serum growth assay of INPP4B wt and control cells OCI-AML2 and OCI- AML3 cells. Viability is measured by trypan blue cell counts. All P-values were derived using the Student s t-test. *P<0.05, ** P <0.01, *** P <
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy se
 A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Phospho-AKT Sampler Kit
 Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Lecture: CHAPTER 13 Signal Transduction Pathways
 Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Signal Transduction Pathways. Part 2
 Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
609G: Concepts of Cancer Genetics and Treatments (3 credits)
 Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Growth and Differentiation Phosphorylation Sampler Kit
 Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
Acute myeloid leukemia. M. Kaźmierczak 2016
 Acute myeloid leukemia M. Kaźmierczak 2016 Acute myeloid leukemia Malignant clonal disorder of immature hematopoietic cells characterized by clonal proliferation of abnormal blast cells and impaired production
Acute myeloid leukemia M. Kaźmierczak 2016 Acute myeloid leukemia Malignant clonal disorder of immature hematopoietic cells characterized by clonal proliferation of abnormal blast cells and impaired production
Chapter 11: Enzyme Catalysis
 Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
oncogenes-and- tumour-suppressor-genes)
 Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
MCB*4010 Midterm Exam / Winter 2008
 MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Contents. Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX
 Contents Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX 1 General Aspects of Signal Transduction and Cancer Therapy 1 1.1 General Principles of Signal Transduction
Contents Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX 1 General Aspects of Signal Transduction and Cancer Therapy 1 1.1 General Principles of Signal Transduction
number Done by Corrected by Doctor Maha Shomaf
 number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
Corporate Medical Policy. Policy Effective February 23, 2018
 Corporate Medical Policy Genetic Testing for FLT3, NPM1 and CEBPA Mutations in Acute File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_flt3_npm1_and_cebpa_mutations_in_acute_myeloid_leukemia
Corporate Medical Policy Genetic Testing for FLT3, NPM1 and CEBPA Mutations in Acute File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_flt3_npm1_and_cebpa_mutations_in_acute_myeloid_leukemia
Signal Transduction I
 Signal Transduction I Prof. Tianhua Zhou Department of Cell Biology Zhejiang University School of Medicine Office hours by appointment tzhou@zju.edu.cn Signal transduction: Key contents for signal transduction:
Signal Transduction I Prof. Tianhua Zhou Department of Cell Biology Zhejiang University School of Medicine Office hours by appointment tzhou@zju.edu.cn Signal transduction: Key contents for signal transduction:
The PI3K/AKT axis. Dr. Lucio Crinò Medical Oncology Division Azienda Ospedaliera-Perugia. Introduction
 The PI3K/AKT axis Dr. Lucio Crinò Medical Oncology Division Azienda Ospedaliera-Perugia Introduction Phosphoinositide 3-kinase (PI3K) pathway are a family of lipid kinases discovered in 1980s. They have
The PI3K/AKT axis Dr. Lucio Crinò Medical Oncology Division Azienda Ospedaliera-Perugia Introduction Phosphoinositide 3-kinase (PI3K) pathway are a family of lipid kinases discovered in 1980s. They have
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapt 15: Molecular Genetics of Cell Cycle and Cancer
 Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Chapter 20. Cell - Cell Signaling: Hormones and Receptors. Three general types of extracellular signaling. endocrine signaling. paracrine signaling
 Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Regulation of PI3K effector signalling in cancer by the phosphoinositide phosphatases
 Review Article Regulation of PI3K effector signalling in cancer by the phosphoinositide phosphatases Samuel J. Rodgers, Daniel T. Ferguson, Christina A. Mitchell and Lisa M. Ooms Cancer Program, Monash
Review Article Regulation of PI3K effector signalling in cancer by the phosphoinositide phosphatases Samuel J. Rodgers, Daniel T. Ferguson, Christina A. Mitchell and Lisa M. Ooms Cancer Program, Monash
Regulation of cell function by intracellular signaling
 Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Molecular Markers in Acute Leukemia. Dr Muhd Zanapiah Zakaria Hospital Ampang
 Molecular Markers in Acute Leukemia Dr Muhd Zanapiah Zakaria Hospital Ampang Molecular Markers Useful at diagnosis Classify groups and prognosis Development of more specific therapies Application of risk-adjusted
Molecular Markers in Acute Leukemia Dr Muhd Zanapiah Zakaria Hospital Ampang Molecular Markers Useful at diagnosis Classify groups and prognosis Development of more specific therapies Application of risk-adjusted
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Cellular Signaling Pathways. Signaling Overview
 Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
THE HALLMARKS OF CANCER
 THE HALLMARKS OF CANCER ONCOGENES - Most of the oncogenes were first identified in retroviruses: EGFR (ErbB), Src, Ras, Myc, PI3K and others (slightly more than 30) - Mutated cellular genes incorporated
THE HALLMARKS OF CANCER ONCOGENES - Most of the oncogenes were first identified in retroviruses: EGFR (ErbB), Src, Ras, Myc, PI3K and others (slightly more than 30) - Mutated cellular genes incorporated
Can we classify cancer using cell signaling?
 Can we classify cancer using cell signaling? Central hypotheses (big ideas) Alterations to signaling genes would cause leukemic cells to react in an inappropriate or sensitized manner to environmental
Can we classify cancer using cell signaling? Central hypotheses (big ideas) Alterations to signaling genes would cause leukemic cells to react in an inappropriate or sensitized manner to environmental
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
What causes cancer? Physical factors (radiation, ionization) Chemical factors (carcinogens) Biological factors (virus, bacteria, parasite)
 Oncogenes What causes cancer? Chemical factors (carcinogens) Physical factors (radiation, ionization) Biological factors (virus, bacteria, parasite) DNA Mutation or damage Oncogenes Tumor suppressor genes
Oncogenes What causes cancer? Chemical factors (carcinogens) Physical factors (radiation, ionization) Biological factors (virus, bacteria, parasite) DNA Mutation or damage Oncogenes Tumor suppressor genes
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Problem Set 8 Key 1 of 8
 7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
VIII Curso Internacional del PIRRECV. Some molecular mechanisms of cancer
 VIII Curso Internacional del PIRRECV Some molecular mechanisms of cancer Laboratorio de Comunicaciones Celulares, Centro FONDAP Estudios Moleculares de la Celula (CEMC), ICBM, Facultad de Medicina, Universidad
VIII Curso Internacional del PIRRECV Some molecular mechanisms of cancer Laboratorio de Comunicaciones Celulares, Centro FONDAP Estudios Moleculares de la Celula (CEMC), ICBM, Facultad de Medicina, Universidad
Signal Transduction: Information Metabolism. Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire
 Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Insulin Resistance. Biol 405 Molecular Medicine
 Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Mechanisms of Hormone Action
 Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Determination Differentiation. determinated precursor specialized cell
 Biology of Cancer -Developmental Biology: Determination and Differentiation -Cell Cycle Regulation -Tumor genes: Proto-Oncogenes, Tumor supressor genes -Tumor-Progression -Example for Tumor-Progression:
Biology of Cancer -Developmental Biology: Determination and Differentiation -Cell Cycle Regulation -Tumor genes: Proto-Oncogenes, Tumor supressor genes -Tumor-Progression -Example for Tumor-Progression:
Objectives. Morphology and IHC. Flow and Cyto FISH. Testing for Heme Malignancies 3/20/2013
 Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
Oncogenes. Dr. S Hosseini-Asl
 Oncogenes Dr. S Hosseini-Asl An oncogene is a mutated form of a normal cellular gene called a proto-oncogene that contributes to the development of a cancer. Proto-oncogenes typically regulate cell growth
Oncogenes Dr. S Hosseini-Asl An oncogene is a mutated form of a normal cellular gene called a proto-oncogene that contributes to the development of a cancer. Proto-oncogenes typically regulate cell growth
1. Activated receptor tyrosine kinases (RTKs) phosphorylates themselves
 Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
The Tissue Engineer s Toolkit
 The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
By the name of Allah
 By the name of Allah Receptors function and signal transduction ( Hormones and receptors Types) We were talking about receptors of the neurotransmitters; we have 2 types of receptors: 1- Ionotropic receptors
By the name of Allah Receptors function and signal transduction ( Hormones and receptors Types) We were talking about receptors of the neurotransmitters; we have 2 types of receptors: 1- Ionotropic receptors
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Introduction! Introduction! Introduction! Chem Lecture 10 Signal Transduction & Sensory Systems Part 2
 Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 2 Questions of the Day: How does the hormone insulin trigger the uptake of glucose in the cells that it targets. Introduction! Signal transduction
Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 2 Questions of the Day: How does the hormone insulin trigger the uptake of glucose in the cells that it targets. Introduction! Signal transduction
Signal-Transduction Cascades - 2. The Phosphoinositide Cascade
 Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Diabetes Mellitus and Breast Cancer
 Masur K, Thévenod F, Zänker KS (eds): Diabetes and Cancer. Epidemiological Evidence and Molecular Links. Front Diabetes. Basel, Karger, 2008, vol 19, pp 97 113 Diabetes Mellitus and Breast Cancer Ido Wolf
Masur K, Thévenod F, Zänker KS (eds): Diabetes and Cancer. Epidemiological Evidence and Molecular Links. Front Diabetes. Basel, Karger, 2008, vol 19, pp 97 113 Diabetes Mellitus and Breast Cancer Ido Wolf
Chem Lecture 10 Signal Transduction
 Chem 452 - Lecture 10 Signal Transduction 111130 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Chem 452 - Lecture 10 Signal Transduction 111130 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Done By : WESSEN ADNAN BUTHAINAH AL-MASAEED
 Done By : WESSEN ADNAN BUTHAINAH AL-MASAEED Acute Myeloid Leukemia Firstly we ll start with this introduction then enter the title of the lecture, so be ready and let s begin by the name of Allah : We
Done By : WESSEN ADNAN BUTHAINAH AL-MASAEED Acute Myeloid Leukemia Firstly we ll start with this introduction then enter the title of the lecture, so be ready and let s begin by the name of Allah : We
Principles of cell signaling Lecture 4
 Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
Life Science 1A Final Exam. January 19, 2006
 ame: TF: Section Time Life Science 1A Final Exam January 19, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use non-erasable
ame: TF: Section Time Life Science 1A Final Exam January 19, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use non-erasable
Sarah Jaar Marah Al-Darawsheh
 22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Receptors Families. Assistant Prof. Dr. Najlaa Saadi PhD Pharmacology Faculty of Pharmacy University of Philadelphia
 Receptors Families Assistant Prof. Dr. Najlaa Saadi PhD Pharmacology Faculty of Pharmacy University of Philadelphia Receptor Families 1. Ligand-gated ion channels 2. G protein coupled receptors 3. Enzyme-linked
Receptors Families Assistant Prof. Dr. Najlaa Saadi PhD Pharmacology Faculty of Pharmacy University of Philadelphia Receptor Families 1. Ligand-gated ion channels 2. G protein coupled receptors 3. Enzyme-linked
Regulation of Enzymatic Activity. Lesson 4
 Regulation of Enzymatic Activity Lesson 4 Regulation of Enzymatic Activity no real regulation: - regulation of enzyme expression and turnover - control of enzyme trafficking - supply of cofactors real
Regulation of Enzymatic Activity Lesson 4 Regulation of Enzymatic Activity no real regulation: - regulation of enzyme expression and turnover - control of enzyme trafficking - supply of cofactors real
What would you observe if you fused a G1 cell with a S cell? A. Mitotic and pulverized chromosomes. B. Mitotic and compact G1 chromosomes.
 What would you observe if you fused a G1 cell with a S cell? A. Mitotic and pulverized chromosomes. B. Mitotic and compact G1 chromosomes. C. Mostly non-compact G1 chromosomes. D. Compact G1 and G2 chromosomes.
What would you observe if you fused a G1 cell with a S cell? A. Mitotic and pulverized chromosomes. B. Mitotic and compact G1 chromosomes. C. Mostly non-compact G1 chromosomes. D. Compact G1 and G2 chromosomes.
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Biochemistry of Carcinogenesis. Lecture # 35 Alexander N. Koval
 Biochemistry of Carcinogenesis Lecture # 35 Alexander N. Koval What is Cancer? The term "cancer" refers to a group of diseases in which cells grow and spread unrestrained throughout the body. It is difficult
Biochemistry of Carcinogenesis Lecture # 35 Alexander N. Koval What is Cancer? The term "cancer" refers to a group of diseases in which cells grow and spread unrestrained throughout the body. It is difficult
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
LESSON 3.2 WORKBOOK. How do normal cells become cancer cells? Workbook Lesson 3.2
 For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
Biol403 MAP kinase signalling
 Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Introduction. Cancer Biology. Tumor-suppressor genes. Proto-oncogenes. DNA stability genes. Mechanisms of carcinogenesis.
 Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Molecular Hematopathology Leukemias I. January 14, 2005
 Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Complexity DNA. Genome RNA. Transcriptome. Protein. Proteome. Metabolites. Metabolome
 DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
Revision. camp pathway
 االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
Cancer. Questions about cancer. What is cancer? What causes unregulated cell growth? What regulates cell growth? What causes DNA damage?
 Questions about cancer What is cancer? Cancer Gil McVean, Department of Statistics, Oxford What causes unregulated cell growth? What regulates cell growth? What causes DNA damage? What are the steps in
Questions about cancer What is cancer? Cancer Gil McVean, Department of Statistics, Oxford What causes unregulated cell growth? What regulates cell growth? What causes DNA damage? What are the steps in
Signaling Through Immune System Receptors (Ch. 7)
 Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia. Dec 14 & 19, 2006 Prof. Erin O Shea Prof. Dan Kahne
 Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne 1 Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne 1 Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis
 Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Chapter 11. Cell Communication
 Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
T cell maturation. T-cell Maturation. What allows T cell maturation?
 T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
T-cell Maturation What allows T cell maturation? Direct contact with thymic epithelial cells Influence of thymic hormones Growth factors (cytokines, CSF) T cell maturation T cell progenitor DN DP SP 2ry
The elements of G protein-coupled receptor systems
 The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
Activation of cellular proto-oncogenes to oncogenes. How was active Ras identified?
 Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
NFκB What is it and What s the deal with radicals?
 The Virtual Free Radical School NFκB What is it and What s the deal with radicals? Emily Ho, Ph.D Linus Pauling Institute Scientist Department of Nutrition and Food Management Oregon State University 117
The Virtual Free Radical School NFκB What is it and What s the deal with radicals? Emily Ho, Ph.D Linus Pauling Institute Scientist Department of Nutrition and Food Management Oregon State University 117
BL-8040: BEST-IN-CLASS CXCR4 ANTAGONIST FOR TREATMENT OF ONCOLOGICAL MALIGNANCIES. Overview and Mechanism of Action Dr.
 BL-8040: BEST-IN-CLASS CXCR4 ANTAGONIST FOR TREATMENT OF ONCOLOGICAL MALIGNANCIES Overview and Mechanism of Action Dr. Leah Klapper, CSO 88 BL-8040: Novel CXCR4 Antagonist For Hematological Cancers Indications:
BL-8040: BEST-IN-CLASS CXCR4 ANTAGONIST FOR TREATMENT OF ONCOLOGICAL MALIGNANCIES Overview and Mechanism of Action Dr. Leah Klapper, CSO 88 BL-8040: Novel CXCR4 Antagonist For Hematological Cancers Indications:
General aspects of this review - specific examples were addressed in class.
 General aspects of this review - specific examples were addressed in class. 1 Exam 1 Lecture 2: Discussed intracellular killing mechanisms Important maturation steps Rapid development into a microbicidal
General aspects of this review - specific examples were addressed in class. 1 Exam 1 Lecture 2: Discussed intracellular killing mechanisms Important maturation steps Rapid development into a microbicidal
Transformation of Normal HMECs (Human Mammary Epithelial Cells) into Metastatic Breast Cancer Cells: Introduction - The Broad Picture:
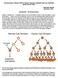 Transformation of Normal HMECs (Human Mammary Epithelial Cells) into Metastatic Breast Cancer Cells: Introduction - The Broad Picture: Spandana Baruah December, 2016 Cancer is defined as: «A disease caused
Transformation of Normal HMECs (Human Mammary Epithelial Cells) into Metastatic Breast Cancer Cells: Introduction - The Broad Picture: Spandana Baruah December, 2016 Cancer is defined as: «A disease caused
HEMATOLOGIC MALIGNANCIES BIOLOGY
 HEMATOLOGIC MALIGNANCIES BIOLOGY Failure of terminal differentiation Failure of differentiated cells to undergo apoptosis Failure to control growth Neoplastic stem cell FAILURE OF TERMINAL DIFFERENTIATION
HEMATOLOGIC MALIGNANCIES BIOLOGY Failure of terminal differentiation Failure of differentiated cells to undergo apoptosis Failure to control growth Neoplastic stem cell FAILURE OF TERMINAL DIFFERENTIATION
Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber
 Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber jweber@dom.wustl.edu Oncogenes & Cancer DNA Tumor Viruses Simian Virus 40 p300 prb p53 Large T Antigen Human Adenovirus p300 E1A
Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber jweber@dom.wustl.edu Oncogenes & Cancer DNA Tumor Viruses Simian Virus 40 p300 prb p53 Large T Antigen Human Adenovirus p300 E1A
Functional Limitations
 Regulation of the Cell Cycle Chapter 12 Pg. 228 245 Functional Limitations Various factors determine whether and when a cell divides. Two functional limitations for cell size limit growth or influence
Regulation of the Cell Cycle Chapter 12 Pg. 228 245 Functional Limitations Various factors determine whether and when a cell divides. Two functional limitations for cell size limit growth or influence
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia
 Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Biosignals, Chapter 8, rearranged, Part I
 Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
CHAPTER VII CONCLUDING REMARKS AND FUTURE DIRECTION. Androgen deprivation therapy is the most used treatment of de novo or recurrent
 CHAPTER VII CONCLUDING REMARKS AND FUTURE DIRECTION Stathmin in Prostate Cancer Development and Progression Androgen deprivation therapy is the most used treatment of de novo or recurrent metastatic PCa.
CHAPTER VII CONCLUDING REMARKS AND FUTURE DIRECTION Stathmin in Prostate Cancer Development and Progression Androgen deprivation therapy is the most used treatment of de novo or recurrent metastatic PCa.
