Changes in Regulation of Cell Cell Adhesion during Tumor Transformation
|
|
|
- Todd Walton
- 6 years ago
- Views:
Transcription
1 ISSN , Biochemistry (Moscow), 2008, Vol. 73, No. 7, pp Pleiades Publishing, Ltd., Original Russian Text N. A. Gloushankova, 2008, published in Biokhimiya, 2008, Vol. 73, No. 7, pp REVIEW Changes in Regulation of Cell Cell Adhesion during Tumor Transformation N. A. Gloushankova Blokhin Cancer Research Center, Russian Academy of Medical Sciences, Kashirskoe Shosse 24, Moscow, Russia; fax: (495) ; Received December 19, 2007 Abstract Cadherin-mediated cell cell adhesion defines the integrity of most tissues. Cell cell adherens junctions are dynamic structures whose functional state is regulated by interactions of cadherin with β-catenin, p120, and actin cytoskeleton structures. Small GTPases of the Rho family and GTPase Rap1 play the central role in the formation and maintenance of cell cell adhesion. Aberrant activation of signaling pathways, transcriptional repression of the E-cadherin gene, ectopic expression of N-cadherin, and disturbances in regulation of adhesive and transcriptional functions of β-catenin stimulate tumor progression. DOI: /S X Key words: cell cell adherens junctions, cadherins, actin cytoskeleton, epithelial-mesenchymal transformation CELL CELL ADHERENS JUNCTIONS Cell cell interactions play the most important role in embryonic development, differentiation, and tissue architecture regulation. Disruption of cell cell adhesion during carcinogenesis is the basis for invasion and metastasis of tumor cells [1-5]. The main molecules of cell cell adhesion are cadherins, transmembrane proteins of the classical cadherin family, which form in the presence of Ca 2+ cell cell adherens junctions (AJ) associated via the cytoplasmic plaque proteins with actin microfilaments [6]. Cadherins provide for mechanical cell cell adhesion and regulate cell shape, segregation, migration, proliferation, and differentiation [7, 8]. In epithelial cells E-cadherin is expressed, whereas AJ in cells of other tissue types are formed by cadherins N, P, R, VE, etc. Cadherins are synthesized as precursors that undergo several posttranslational modifications including proteolytic cleavage. In particular, E-cadherin is transformed from a 135 kd precursor into a 120 kd molecule with N-terminal Asp135. Correct cleavage of precursors is necessary for construction of adhesive dimers [9]. Abbreviations: AJ) cell cell adherens junctions; EGF) epidermal growth factor; EMT) epithelial-mesenchymal transformation; FGF) fibroblast growth factor; GEFs) guanine nucleotide exchange factors; HGF/SF) hepatocyte growth factor/scatter factor; PDGF) platelet-derived growth factor. The cadherin molecule incorporated into plasma membrane has extracellular, transmembrane, and intracellular regions. The extracellular region of classical cadherins consists of five domains (EC1-EC5), each of which is formed by 110 amino acids. There are four Ca 2+ -binding sites between EC domains [10]. It is now assumed that trans-interactions of cadherin molecules are caused by binding of EC1 domains of adjacent cells via reciprocal interaction of Trp156 from one EC1 domain with a hydrophobic pocket of another EC1 domain [9, 11]. The cytoplasmic domain of cadherins is highly conserved and includes the membrane-adjacent site for cadherin binding to p120-catenin (p120) and C-terminal site for binding β-catenin, which regulate cell cell adhesion [12]. Cadherins bind via β-catenin, plakoglobin, and α- catenin to actin filaments, which stabilize the structure of AJ [13, 14]. The interaction of E-cadherin βcatenin α-catenin actin filaments in the region of cell cell contacts is dynamic [15, 16]. DYNAMICS OF AJ FORMATION The process of AJ formation can be arbitrarily divided into the following stages: mechanically weak transinteractions of individual cadherin receptors bound by their cytoplasmic domains with β-catenin molecules; contact stabilization by formation of cadherin clusters after recruiting the cytoplasmic plaque proteins and bind- 742
2 REGULATION OF CELL CELL ADHESION DURING TUMOR TRANSFORMATION 743 ing of actin filaments; contact maturation due to further actin accumulation in the contact region. The interactions of cadherin with actin are of no importance in the first steps of intercellular interaction, but are extremely important for contact stabilization [17]. Adhesive interactions, supported by the Ca 2+ -independent binding of Ig-like nectins of adjacent cells play an essential role in stabilization of initial cadherin contacts [18]. Formation of cell cell contacts is accompanied by structural rearrangements of actin cytoskeleton. These rearrangements are different in epithelial cells and fibroblasts and are defined by common organization of the actin cytoskeleton in the cells of two tissue types. In particular, when AJ of epithelial cells are formed, marginal actin bundle in the contact zone dissociates as the result of local breaks and ark-like bundles are formed at the lateral free cell edges [19, 20]. Further actin accumulation and formation of adhesion belts along the cell perimeter takes place during maturation of epithelial contacts. Formation of cell cell contact of epitheliocytes is accompanied by inhibition of pseudopodial activity (contact paralysis) in the contact region and at free edges of the cells. Fibroblast-like cells form AJ of different spatial organization from AJ of epitheliocytes. AJ of fibroblasts are most often formed by N-cadherin, are located in the region of overlap of the adjacent cell lamellas, and are characterized by radial organization. Such contacts are perpendicular to the contact border and are associated with short straight actin bundles [21, 22]. Formation and maintenance of radial contacts is defined by contractility of associated microfilament bundles, because inhibitors of actin-myosin contractility cause their disappearance. In fibroblasts, formation of AJ is not accompanied by inhibition of pseudopodial activity: lateral lamellae are formed at the free cell edges of the colliding cells, which may result in change in cell motility direction after their contact [22]. In the course of embryonic development as well as during metastatic tumor progression, the so-called epithelial-mesenchymal transformation (EMT) takes place, which results in transformation of epithelial cells into fibroblast-like ones. The effect of phorbol ether (TPA) on epithelial cells in culture can be used as a model for investigation of early EMT steps. Already 1-2 h later there begins destruction of the actin cytoskeleton typical of epitheliocytes, i.e. the disappearance of circular microfilament bundles accompanied by radical rearrangement of cell cell interactions. In the course of cell cell interaction, the cells form E-cadherin-containing AJ that have not tangential but radial organization characteristic of fibroblasts. In this case, pseudopodial activity is not stabilized after the AJ formation [23]. Actin polymerization de novo plays an essential role in formation of stable AJ [24-26]. Experiments with exogenous G-actin incorporation have shown that the labeled globular actin is accumulated in the region of newly formed AJ of epitheliocytes and is co-localized with E-cadherin within 5 min after transfer of the cells from the low-calcium medium into the high-calcium medium (calcium concentration ~1.8 mm) [27]. Actin polymerization begins from filament nucleation and following elongation. The protein complex Arp2/3 consisting of seven polypeptides plays the key role in nucleation of the branched network of actin filaments [28]. The available data show that the Arp2/3 complex is involved in formation of cell cell contacts [26, 29]. The Arp2/3 activators N-WASP, WAVE2, and cortactin were shown to be necessary for the assembly of stable AJ [27, 30, 31]. These proteins are co-localized with cadherins in the regions of new AJ. Introduction into cells of a dominantly negative cortactin, lacking the F-actin-binding site, or unable to bind Arp2/3 or inhibition of the cortactin gene expression by a small interfering RNA (sirna) results in inhibition of new AJ formation and destruction of already existing contacts [25]. The N-WASP inhibitor wiskostatin blocks the assembly of E-cadherin-containing AJ of epitheliocytes [27]. Inhibition of WAVE2 expression upon introduction of specific sirna also resulted in distortion of actin recruitment and AJ disorganization [31]. Formins, able to nucleate polymerization of linear actin filaments and provide for their elongation, also contribute to formation of new actin structures in the region of cell cell contacts [32]. Thus, the α-catenin-binding FMN1 is necessary for AJ formation and maintenance in keratinocytes [33]. Recently it has been also shown that another member of the formin family Dial, activated by the small GTPase Rho, is involved in AJ formation in epitheliocytes [34]. The formin-stimulated mechanism of actin filament growth is based on the ability of the formin dimer to bind in a stepwise manner to each next attached actin monomer using the FH2 domain. In this case, the filament plus-end appears to be protected against binding to capping proteins, whereas the formin dimer is rapidly translocated along the growing filament [35-37]. The formin domain FH1 is able to bind profilin, which can also stimulate filament growth due to efficient recruitment of actin molecules [35]. INVOLVEMENT OF SMALL GTPases OF Rho FAMILY IN AJ FORMATION There are now numerous data concerning the involvement in AJ formation of small GTPases of Rho family (Rho, Rac, and Cdc42), regulating intracellular dynamics of actin cytoskeleton. It was shown that expression of dominantly negative mutants of RhoA, Rac1, and Cdc42 genes as well as injection into cells of negatively dominant forms of their proteins disturbed formation of E-cadherin-containing contacts of MDCK epitheliocytes and keratinocytes [38, 39]. It was found that GTPases Rac1 and Cdc42 were recruited into zones of N- and E-
3 744 GLOUSHANKOVA cadherin-dependent cell cell adhesion [40-42]. GTPase Rac1, activated via phosphatidylinositol-3-kinase (PI3K), nectins, or via IQGAP1 [18, 43, 44] and regulating the Arp2/3-mediated actin polymerization, is of essential significance for AJ formation. Rac1 is activated via the lipid product of PI3K phosphatidylinositol (3,4,5)-triphosphate (PIP3). PIP3 interacts directly with GEFs (P-Rex1, SWAP-70, Vav1, Sos1, and evidently Tiam1) activating Rac1 [44]. Rac1, in turn, is able to stimulate via proteins of the WASP family (N-WASP and WAVE2) the Arp2/3-mediated actin assembly in the zone of cell cell contact. Since WAVE2 can form a supramolecular complex with Rac via IRSp53, it is assumed that the Rac WAVE2 Arp2/3 pathway plays the leading role in the assembly of actin structures that stabilize cell cell contacts [45]. Tiam1, a Rac1-specific GEF, plays an essential role in the regulation of assembly of AJ. Tiam1 was initially described as the invasive phenotype inductor of lymphoma cells. In epithelial cells, Tiam1 is accumulated in cell cell contacts. The expression of Tiam1 in Ras-transformed epitheliocytes stimulates normalization of transformed phenotype and restoration of cell cell adhesion. It was also shown that the effect of exogenous Tiam1 in epithelial cells depends on the cell substrate. Tiam1 activates motility of epitheliocytes grown on collagen and, in contrast, it stabilizes cell cell contacts of epitheliocytes grown on fibronectin or laminin [46]. There are also data concerning the involvement of IQGAP1 in AJ stabilization. IQGAP1 contains binding sites for actin, calmodulin, the myosin light chain kinase, Rac1/Cdc42, Rap1, β-catenin, E-cadherin, and APC [47]. It is assumed that IQGAP1 inhibits GTP hydrolysis and stabilizes Rac1 and Cdc42 in the active state, and in this case IQGAP1 is an effector of Rac1 and Cdc42. Rac1 and Cdc42 inhibit the interaction of IQGAP1 with β- catenin, which stabilizes contacts [43]. Although there are data concerning the most important role of activated GTPases of Rho family in AJ assembly, relative contribution of different pathways, regulated by these GTPases, to cell cell adhesion and assembly of actin structures in the contact zone is still unclear. CELL CELL ADHESION REGULATOR Rap1 The small GTPase Rap1 was first described as a suppressor of Ras-transformation [48]. The involvement of Rap1 in regulation of cell cell contacts in mammals was found during studies of GEF for Rap1 DOCK4, inactivating mutations of which were detected in some lines obtained from human tumors. Human osteosarcoma cells that lost DOCK4 did not form AJ. In this case, introduction of DOCK4 or an active Rap1 restored formation of AJ and decreased metastatic activity of tumor cells [49]. The reappearance of AJ was also observed upon introduction of active Rap1 into Ras-transformed MDCK epitheliocytes. Besides, endogenous Rap1 activation blocks HGF/SF-stimulated cell dissemination and AJ disassembly [50]. Rap1 is now considered to be the key regulator of E- and VE-cadherin-mediated adhesion. Two models have been proposed for the participation of Rap1 in AJ assembly. According to the first model, in the initial steps of cell cell contact formation trans-interaction of nectins activates the src-tyrosine kinase that phosphorylates C3G (GEF for Rap1) and recruits it onto the membrane, which results in the activation of Rap1 [51]. Another GEF for Rap1, PDZ-GEF1, is able to bind β-catenin and MAGI-1 and MAGI-2 proteins and can be involved in AJ maturation [50, 52]. It was shown that activation of small GTPases Rac1 and Cdc42 upon AJ formation depends on Rap1 activity [53, 54]. It is supposed that GEFs of small GTPases Rac1 and Cdc42 (Vav, Tiam1, FRG), regulating actin polymerization in the region of cell cell contacts and thus stimulating contact maturation, are activated by Rap1 [54]. According to the second model, Rap1 activation by nectins recruits afadin/af6 that binds to p120, which, in turn, inhibits E-cadherin endocytosis [55]. It is also supposed that Rap1 plays an important role in coordinated disassembly of cadherincontaining AJ and formation of cell matrix focal contacts upon induction of EMT [56]. TUMOR SUPPRESSOR E-CADHERIN It is now generally accepted that E-cadherin is the main suppressor of epithelial tumor invasion. Decrease or disappearance of E-cadherin expression is described in many human carcinomas [57, 58]. A single mutation of the E-cadherin gene can be responsible for transformation of adenoma to carcinoma. Expression of exogenous E-cadherin in cells of transformed epithelial lines significantly reduces their invasive potential and restores normal phenotype [59]. There are data concerning the inhibitory effect of the E-cadherin homophilic transinteractions on cell proliferation via interaction of β- catenin, bound to E-cadherin at the plasma membrane, with epidermal growth factor receptor (EGFR). Interaction of cell cell adhesion molecules with EGFR results in inhibition of EGFR phosphorylation at Tyr845, which, in turn, decreases activation of the ERK-independent signaling pathway via STAT5b activation [60-62]. The enhanced activity of receptor tyrosine kinases characteristic of many tumors can decrease due to binding of the growth factor receptor to the extracellular domain of E-cadherin [63]. In the course of tumor progression, epithelial cells undergo EMT during which cells acquire the fibroblastlike phenotype, the ability for directed migration, and dissociate from each other. These events are the basis for
4 REGULATION OF CELL CELL ADHESION DURING TUMOR TRANSFORMATION 745 invasion and metastasis of malignant cells. The decrease of E-cadherin expression leading to destruction of cell cell contacts plays the most important role in EMT [64]. Inhibition of E-cadherin expression in tumors is achieved in three ways: by transcriptional repression or mutation of E-cadherin gene CDH1, as well as by hypermethylation of CpG islets of the gene promoter [65-69]. Histone H3 deacetylation of CpG islets was also found in some lines of epithelial tumors [70]. Transcriptional repression of the E-cadherin gene CDH1 is the key regulator of EMT and is most often involved in carcinoma progression. CDH1 transcription repressors belong to three families: i) Snail (SNAIL1, SNAIL2 (SLUG), SNAIL3); ii) ZEB1 (DeltaEF1)/ ZEB2 (SIP1), and iii) TWIST/E47 factors bhlh interacting with the E-cadherin gene promoter [71-73]. The Snail family proteins regulate EMT in embryogenesis upon mesoderm formation, gastrulation, and neural crest formation. SNAIL1 overexpression in epithelial cells also stimulates EMT, which is revealed in acquirement by cells of ability for directed migration and for invasiveness, which was detected during investigation of cell invasion through a gel consisting of collagen IV [71]. It has been shown that in many human carcinomas, in particular, in gastric tumors, liver, colon, and ovary carcinomas, and breast cancer SNAIL1 expression correlates with inhibition of E-cadherin expression [74, 75]. SNAIL1 protein belongs to the family of transcription repressors containing four zinc fingers at the conserved C-terminus, which bind the latter by E2-boxes located near the site of CDH1 gene transcription initiation [76, 77]. SNAIL1 repressor activity is defined by the SNAG domain at the N-terminus of the molecule and is regulated by the central domain, whose phosphorylation by GSK3β kinase alters the subcellular localization, stability, and activity of SNAIL1. Cooperation with LOXL2 stabilizes SNAIL1 by regulation of its binding to GSK3β [78]. SNAIL2 expression is detected in breast cancer, carcinomas of ovaries, colon, and in melanomas. Expression of SNAIL1 and SNAIL2 often correlates with poor clinical prognosis [78]. Multiple signaling cascades including the FGF, PDGF, Wnt, EGF, HGF/SF, Ras-MAPK, Ras/PI3K/ AKT, TGFβ, and Hedgehog-GLI1 pathways (involved in EMT upon oncogene activation), hypoxia, and microenvironment regulate activity of the CDH1 transcription repressors [79]. Transcription repressors ZEB1/ZEB2 were detected in gastric tumors, carcinomas of, pancreas, bladder, and ovaries. The TWIST factor is revealed in ductal breast cancer, bladder and prostate tumors, hepatocarcinomas, and melanomas. TWIST is now considered as a factor playing a key role in early stages of metastasis [78, 79]. TWIST also induces N-cadherin expression in prostate carcinoma cells [80]. Co-expression and sequential activation of transcription repressors Snail, ZEB1/ZEB2, and TWIST/E47 can be observed in epithelial tumors [72]. Binding sites of Snail and other repressors of the E-cadherin gene are overlapping. Comparative analysis of the affinity to E1/2- boxes of CDH1 repressors SNAIL1, SNAIL2, and E47 has shown that in the presence in a cell of all three repressors their hierarchies by the extent of affinity and by prevalent contribution to repressor effect are observed. Different repressors can be involved in CDH1 repression, specific for different types of tumors, or certain stages of tumor progression. Transcription repressors modulate expression of the E-cadherin gene and of many different genes of epithelial cells. In particular, they decrease expression of the tight junction proteins occludin and claudin, desmosome protein plakophilin-3, cytokeratins 17/18, integrins α3/α4, and actin-binding protein gelsolin; they also change spectrum of catenin p120 isoforms [78, 81]. Recent studies of functioning of CDH1 transcription repressors have shown that these proteins, contributing to EMT, are able to regulate expression of many genes by influencing tumor cell proliferation and survival, by inhibition of apoptosis, and by stimulation of multidrug resistance, which is important for tumor progression [78]. ROLE OF p120 IN CADHERIN STABILIZATION ON MEMBRANE Along with other adhesion plaque molecules, p120 plays an important role in cell cell adhesion by participation in cadherin clustering and endocytosis, and in N- cadherin transport onto the plasma membrane [82]. It has been shown in recent works that p120 is a key regulator of AJ stability. It was shown that p120 stabilizes the macromolecular adhesion complex by prevention of transmembrane cadherin internalization and thus by regulation of intracellular cadherin turnover [55]. The first direct indication that p120 is a central molecule regulating the adhesion function of cadherin was reported in analysis of the SW48 carcinoma cell line containing a mutation in the p120 gene. It appeared that in the absence of p120, transformed cells are not able to form compact colonies and cannot spread on the substrate covered with chimeric protein formed by the E-cadherin ectodomain bound to Fc-fragment of IgG. In these cells the amount of E-cadherin dramatically decreased, whereas the level of E-cadherin mrna did not change. The loss of p120 caused internalization of membrane cadherin and its delivery to lysosomes with following proteolytic degradation. In this case, expression of exogenous p120 increased intracellular level of E-cadherin and restored cell cell adhesion and cell spreading on the substrate covered with chimeric protein [83]. Two models of the E-cadherin endocytosis regulation with involvement of p120 are now being discussed. In the first model, p120 is considered as a capping molecule
5 746 GLOUSHANKOVA connected with the cadherin tail and preventing the interaction of cadherin with the endocytosis machine. An alternative model suggests that p120 influences the stability of E-cadherin in AJ via interaction with small GTPases Rho, Rac, and Cdc42 [55]. An important role in ligation of E-cadherin molecules belongs to the cell cell adhesion molecule nectin, which is involved in the assembly of a supramolecular complex by p120 recruitment onto the cytoplasmic tail of E-cadherin due to activation of small GTPase Rap1 and interaction with the actin-binding protein afadin [84]. It is still not clear which events serve as a launching moment for p120 dissociation from the E-cadherin cytoplasmic tail and following endocytosis. It is supposed that E-cadherin endocytosis begins after its phosphorylation on tyrosine, binding by E3-ubiquitin-ligase Hakai, and ubiquitin binding [85]. The E-cadherin phosphorylation and binding to Hakai can be prevented by p120 [55]. Recently it has also been shown that casein kinase 1 phosphorylates Ser846 of the E-cadherin cytoplasmic tail, thus promoting its internalization and weakening of cell cell adhesion [86]. Cadherin endocytosis is triggered in response to growth factors, change in the blood vessel endothelium permeability caused by the growth factor VEGF due to alteration of VE-cadherin barrier function, and in response to oncogenic transformation [87-91]. The changes in cell cell adhesion might be mediated by p120 phosphorylation. Thus, HGF/SF, EGF, and PDGF phosphorylate p120 via protein kinase Src, which results in destabilization of the E-cadherin/catenin complex [92-94]. The FGFR1 or met-receptors in the case of binding, respectively, to FGF or HGF/SF, are co-internalized with E-cadherin [87, 89]. Overexpression of MDM2, a negative regulator of tumor suppressor p53, contributes to the development of invasive phenotype. Recently it has been shown that E-cadherin is a substrate for MDM2, and upon its binding to the latter the processes of E-cadherin ubiquitination and endocytosis are triggered. Dominantly negative dynamin mutants, blocking endocytosis, disturbed the interaction of E-cadherin with MDM2 and decreased migration and invasive activities of MCF-7 carcinoma cells expressing exogenous MDM2 [95]. The decrease in p120 expression or complete disappearance of p120, regulating E-cadherin endocytosis, is found in many human tumors; in some tumors, a relationship between decreased levels of E-cadherin and p120 was detected [96]. Inhibition of p120 expression, change in intracellular protein localization (transfer into the cytoplasm and nucleus), appearance of the long isoforms of p120 typical of high-motility fibroblast-like cells, and the p120 binding to microtubules and Kaiso transcription factor might contribute to tumor progression. Inhibition of cell cell adhesion and enhancement of migration ability are the main result of changes in regulation of E-cadherin endocytosis in transformed cells [63]. ADHESIVE AND TRANSCRIPTIONAL FUNCTIONS OF β-catenin The E-cadherin β-catenin complex is formed in endoplasmic reticulum and then is transported onto the plasma membrane [97]. In the case of AJ formation, β- catenin provides for link between cadherin and α-catenin and plays an important role in the maintenance of contact stability [13]. Tyrosine phosphatase PTP1B of adhesion complexes maintains β-catenin in its dephosphorylated state [98]. The distortion of β-catenin adhesive function is first of all caused by phosphorylation of tyrosines in positions 142 and 654, which results in disassembly of E- cadherin-containing complexes in the membrane. Phosphorylation of β-catenin at Tyr654 by c-src disturbs its binding to E-cadherin and destroys cell cell adhesion [99, 100]. Phosphorylation of β-catenin was described upon Ras activation, and it is also characteristic of effects of some growth factors. Thus, EGF induces β-catenin and plakoglobin phosphorylation, resulting in breaking of E-cadherin binding to actin cytoskeleton and contact disassembly [60]. EGF receptors are overexpressed in many carcinomas, which may contribute to EMT due to β- catenin phosphorylation [63]. β-catenin is also the most important component of cellular signaling pathways, in particular, of the Wnt-signaling pathway. In a complex with transcription factors TCF/LEF, β-catenin activates transcription of many genes, in particular of myc, cyclin D1 gene, FGF18, and FGF20 [ ]. An increasing amount of recent data show that adhesive and transcriptional functions of β- catenin are tightly connected and are involved in changes in tumor cell morphology and motility [107, 108]. Phosphorylation at tyrosine influences both adhesive and transcriptional functions of β-catenin: phosphorylation at Tyr142 in the case of BCL9-2 overexpression not only results in β-catenin release from AJ with subsequent EMT, but also enhances nuclear transcriptional activity of β-catenin [109]. β-catenin significantly contributes to transformation via the Wnt-signaling pathway that begins from receptors of the Frizzled family. In the absence of signals β-catenin is recruited by a complex consisting of APC protein, axin, GSK3β kinases, and casein kinase 1. Phosphorylation of the β-catenin N-terminus at serine threonine in this complex serves as a signal for its ubiquitination and subsequent proteosomal degradation [ ]. Canonical Wnt-signals induce the Frizzled Dishevelled complex interaction with LRP5/6 co-receptors, which results in axin recruitment to the plasma membrane, of β-catenin degradation, and its transfer to the nucleus in a complex with transcription factors TCF/LEF [113, 114]. It has recently been found that proteins BCL9 and BCL9-2 of the Legless family play an important role in the transcriptional function of β-catenin by binding the latter to co-activators
6 REGULATION OF CELL CELL ADHESION DURING TUMOR TRANSFORMATION 747 PYGO1/2, which is necessary for translocation of β- catenin into the nucleus [115, 116]. BCL9-2 overexpression induces malignant transformation of epithelial cells in vitro and is detected in human colon carcinomas [109, 117]. However, even in the case of the Wnt-signaling pathway activation β-catenin can be sequestered by E-cadherin and excluded from transcriptional regulation [118]. APC, CTNNB1, and AXIN mutations, activating the canonical Wnt-pathway, are detected in human colorectal carcinomas, hepatoblastomas, and melanomas [ ]. Mutations of APC are responsible for development of familial adenomatous polyposis coli and are detected in 85% of cases of sporadic colorectal cancer [122]. It has also been shown that mutations of APC via GEF of small Rac GTPase, Asef, contribute to the enhanced migration activity of the colon carcinoma cells [123]. The serine threonine kinase GSK3β, involved in degradation of both β-catenin and the cadherin transcription repressor SNAIL1 [124], is also a key component of the FGF signaling pathway playing a noticeable role in control of tumor cell proliferation, differentiation, migration, and survival. The activation of the FGF signaling pathway in which Akt phosphorylates GSK3β at Ser9 results in down-regulation of GSK3β activity with subsequent increase in SNAIL1 level. SNAIL1 represses E-cadherin expression, which results in release of β- catenin from adhesion contacts and its translocation into the nucleus [ ]. Since FGF18 and FGF20 are effector genes of the Wnt-signaling pathway, activation of the latter upon transformation triggers the FGF signaling system as well [128, 129]. Thus, multiple pathways of deregulated adhesive and transcriptional functions of β-catenin play an important role in different stages of carcinogenesis. PARTICIPATION OF N-CADHERIN IN CELL CELL ADHESION: ITS ROLE IN TUMOR CELL INVASION AND METASTASIS Adherens junctions in nervous and connective tissues, myocardium, bone, and cartilage are formed by N- cadherin [130]. Measuring of the strength that need be applied for separation of cells in contact has shown that N-cadherin-containing cell cell contacts are significantly weaker than those involving E-cadherin [17]. These differences can be determined either by cadherin binding to different p120 isoforms or by the extent of p120 phosphorylation in different adherens junctions [130]. In the case of formation of N-cadherin-containing contacts, the contribution of specific Fer-kinase is of great importance. Upon binding of N-cadherin molecules of adjacent cells, Rac-dependent cortactin recruitment into the region of cell cell contacts and cortactin phosphorylation by Fer-kinase resulting in adhesion enhancement occurs [131]. At the same time, data continue to appear showing that N-cadherin plays a decisive role in invasion and metastasis of epithelial tumor cells by stimulation of their migration. In many human carcinomas, including breast cancer, thyroid, bladder, and prostate carcinomas, aberrant expression of N-cadherin has been detected along with inhibition of E-cadherin expression. Re-expression of N-cadherin, found during embryogenesis in melanocytes only at the stage of neural crest formation, is also described in metastatic melanomas [132]. It was also shown in vitro that N-cadherin expression by epithelial cells, independently of the E-cadherin expression level, causes EMT and stimulates cell migration activity [133]. Exogenous N-cadherin expression in MCF-7 breast cancer cells resulted in metastases to lymph nodes, liver, and lungs after injection of these cells into nude mice [134]. It can be supposed that the proinvasive effect of N-cadherin expression in epithelial tumors is explained by the N-cadherin-mediated tumor cell interactions with stromal fibroblasts that, according to recent data, are involved in collective invasion of carcinoma cells [135]. It has been also shown that N-cadherin interacts with FGF receptors. This inhibits their internalization and results in MAPK-cascade activation and expression of metalloproteases [133, 136]. Recent data also show that in HT-1080 cells N-cadherin/β-catenin form via NHERF a complex with receptors of PDGF that modulates actin cytoskeleton and contributes to cell motility [137]. Thus, aberrant expression of N-cadherin in carcinomas can be involved in EMT during invasion. The data show that AJ are of key significance for the tissue architecture. The formation, stabilization, and maturation of AJ are first of all defined by their close interaction with the actin cytoskeleton structures as well as by small GTPases regulating trans-interactions of cadherin molecules, their endocytosis, and actin dynamics in the region of cell cell adhesion. In connection with widely described disruption of cell cell adhesion at different stages of carcinogenesis, in particular in the case of EMT, studies of molecular events accompanying oncogene activation and resulting in rearrangements of AJ organization up to their complete disappearance become most important. Investigation of such rearrangements, changes in regulation of adhesive and transcriptional functions of the cell cell adhesion molecules, and their relationships with changes in cytoskeleton structures during malignization will lead to better understanding of events that are the basis of neoplastic transformation. REFERENCES 1. Willert, K., and Jones, K. A. (2007) Genes Dev., 20, Perez-Moreno, M., and Fuchs, E. (2006) Dev. Cell, 11,
7 748 GLOUSHANKOVA 3. Katoh, M., and Katoh, M. (2006) Cancer Biol. Ther., 5, Erez, N., Bershadsky, A., and Geiger, B. (2005) Eur. J. Cell Biol., 84, Polakis, P. (2007) Curr. Opin. Genet. Dev., 17, Takeichi, M. (1995) Curr. Opin. Cell. Biol., 7, Gumbiner, B. M. (2000) J. Cell Biol., 148, Gooding, J. M., Yap, K. L., and Ikura, M. (2004) BioEssays, 26, Troyanovsky, S. (2005) Eur. J. Cell. Biol., 84, Nagafuchi, A. (2001) Curr. Opin. Cell. Biol., 13, Shapiro, L., Fannon, A. M., Kwong, P. D., Thompson, A., Lehmann, M. S., Grubel, G., Legrand, J. F., Als-Nielsen, J., Colman, D. R., and Hendrickson, W. A. (1995) Nature, 374, Kobielak, A., and Fuchs, E. (2004) Nat. Rev. Mol. Cell. Biol., 5, Kemler, R. (1993) Trends Genet., 9, Rimm, D. L., Koslov, E. R., Kebriaei, P., Cianci, C. D., and Morrow, J. S. (1995) Proc. Natl. Acad. Sci. USA, 92, Drees, F., Pokutta, S., Yamada, S., Nelson, W. J., and Weis, W. I. (2005) Cell, 123, Yamada, S., Pokutta, S., Drees, F., Weis, W. I., and Nelson, W. J. (2005) Cell, 123, Chu, Y. S., Thomas, W. A., Eder, O., Pincet, F., Perez, E., Thiery, J. P., and Dufour, S. (2004) J. Cell Biol., 167, Sakisaka, T., Ikeda, W., Ogita, H., Fujita, N., and Takai, Y. (2007) Curr. Opin. Cell. Biol., 19, Gloushankova, N. A., Alieva, N. O., Krendel, M. F., Bonder, E. M., Feder, H. H., Vasiliev, J. M., and Gelfand, I. M. (1997) Proc. Natl. Acad. Sci. USA, 94, Krendel, M. F., and Bonder, E. M. (1999) Cell Motil. Cytoskeleton, 43, Yonemura, S., Itoh, M., Nagafuchi, A., and Tsukita, S. (1995) J. Cell Sci., 108, Gloushankova, N. A., Krendel, M. F., Alieva, N. O., Bonder, E. M., Feder, H. H., Vasiliev, J. M., and Gelfand, I. M. (1998) Proc. Natl. Acad. Sci. USA, 95, Krendel, M. F., Gloushankova, N. A., Alieva, N. O., Bonder, E. M., Feder, H. H., Vasiliev, J. M., and Gelfand, I. M. (1999) Proc. Natl. Acad. Sci. USA, 96, Vasioukhin, V., Bauer, C., Yin, M., and Fuchs, E. (2000) Cell, 100, Helwani, F. M., Kovacs, E. M., Paterson, A. D., Verma, S., Ali, R. G., Fanning, A. S., Weed, S. A., and Yap, A. S. (2004) J. Cell Biol., 164, Verma, S., Shewan, A. M., Scott, J. A., Helwani, F. M., den Elzen, N. R., Miki, H., Takenawa, T., and Yap, A. S. (2004) J. Biol. Chem., 279, Ivanov, A. I., Hunt, D., Utech, M., Nusrat, A., and Parkos, C. A. (2005) Mol. Biol. Cell, 16, Higgs, H. N., and Pollard, T. D. (2001) Annu. Rev. Biochem., 70, Kovacs, E. M., Goodwin, M., Ali, R. G., Paterson, A. D., and Yap, A. S. (2002) Curr. Biol., 12, El Sayegh, T. Y., Arora, P. D., Laschinger, C. A., Lee, W., Morrison, C., Overall, C. M., Kapus, A., and McCulloch, C. A. (2004) J. Cell Sci., 117, Yamazaki, D., Oikawa, T., and Takenawa, T. (2007) J. Cell Sci., 120, Goode, B. L., and Eck, M. J. (2007) Annu. Rev. Biochem., 76, Kobielak, A., Pasolli, H. A., and Fuchs, E. (2004) Nat. Cell Biol., 6, Carramusa, L., Ballestrem, C., Zilberman, Y., and Bershadsky, A. D. (2007) J. Cell Sci., 120, Zigmond, S. H. (2004) Curr. Opin. Cell Biol., 16, Xu, Y., Moseley, J. B., Sagot, I., Poy, F., Pellman, D., Goode, B. L., and Eck, M. J. (2004) Cell, 116, Higashida, C., Miyoshi, T., Fujita, A., Oceguera-Yanez, F., Monypenny, J., Andou, Y., Narumiya, S., and Watanabe, N. (2004) Science, 303, Takaishi, K., Sasaki, T., Kotani, H., Nishioka, H., and Takai, Y. (1997) J. Cell Biol., 139, Braga, V. M. M., Machesky, L. M., Hall, A., and Hotchin, N. A. (1997) J. Cell Biol., 137, Kim, S. H., Li, Z., and Sacks, D. B. (2000) J. Biol. Chem., 275, Kovacs, E. M., Ali, R. G., McCormack, A. J., and Yap, A. S. (2002) J. Biol. Chem., 277, Ehrlich, J. S., Hansen, M. D., and Nelson, W. J. (2002) Dev. Cell, 3, Fukata, M., and Kaibuchi, K. (2001) Nat. Rev. Mol. Cell. Biol., 2, Welch, H. C., Coadwell, W. J., Stephens, L. R., and Hawkins, P. T. (2003) FEBS Lett., 546, Stradal, T. E., Rottner, K., Disanza, A., Confalonieri, S., Innocenti, M., and Scita, G. (2004) Trends Cell Biol., 14, Price, L. S., and Collard, J. G. (2001) Semin. Cancer Biol., 11, Noritake, J., Watanabe, T., Sato, K., Wang, S., and Kaibuchi, K. (2005) J. Cell Sci., 118, Kitayama, H., Sugimoto, Y., Matsuzaki, T., Ikawa, Y., and Noda, M. (1989) Cell, 56, Yajnik, V., Paulding, C., Sordella, R., McClatchey, A. I., Saito, M., Wahrer, D. C., Reynolds, P., Bell, D. W., Lake, R., van den Heuvel, S., Settleman, J., and Haber, D. A. (2003) Cell, 112, Kooistra, M. R., Dube, N., and Bos, J. L. (2007) J. Cell Sci., 120, Fukuyama, T., Ogita, H., Kawakatsu, T., Fukuhara, T., Yamada, T., Sato, T., Shimizu, K., Nakamura, T., Matsuda, M., and Takai, Y. (2005) J. Biol. Chem., 280, Kawajiri, A., Itoh, N., Fukata, M., Nakagawa, M., Yamaga, M., Iwamatsu, A., and Kaibuchi, K. (2000) Biochem. Biophys. Res. Commun., 273, Hogan, C., Serpente, N., Cogram, P., Hosking, C. R., Bialucha, C. U., Feller, S. M., Braga, V. M., Birchmeier, W., and Fujita, Y. (2004) Mol. Cell. Biol., 24, Fukuyama, T., Ogita, H., Kawakatsu, T., Inagaki, M., and Takai, Y. (2006) Oncogene, 25, Xiao, K., Oas, R. G., Chiasson, C. M., and Kowalczyk, A. P. (2007) Biochim. Biophys. Acta, 1773, Retta, S. F., Balzac, F., and Avolio, M. (2006) Eur. J. Cell Biol., 85, Hirohashi, S. (1998) Am. J. Pathol., 153, Berx, G., and van Roy, F. (2001) Breast Cancer Res., 3, Cavallaro, U., and Christofori, G. (2004) Nat. Rev. Cancer, 4,
8 REGULATION OF CELL CELL ADHESION DURING TUMOR TRANSFORMATION Hoschuetzky, H., Aberle, H., and Kemler, R. (1994) J. Cell Biol., 127, Boerner, J. L., Biscardi, J. S., Silva, C. M., and Parsons, S. J. (2005) Mol. Carcinog., 44, Perrais, M., Chen, X., Perez-Moreno, M., and Gumbiner, B. M. (2007) Mol. Biol. Cell, 18, Van Hengel, J., and van Roy, F. (2007) Biochim. Biophys. Acta, 1773, Thiery, J. P. (2002) Nat. Rev. Cancer, 2, Yoshiura, K., Kanai, Y., Ochiai, A., Shimoyama, Y., Sugimura, T., and Hirohashi, S. (1995) Proc. Natl. Acad. Sci. USA, 92, Berx, G., Becker, K. F., Hofler, H., and van Roy, F. (1998) Hum. Mutat., 12, Hajra, K. M., Ji, X., and Fearon, E. R. (1999) Oncogene, 18, Di Croce, L., and Pelicci, P. G. (2003) Eur. J. Cancer, 39, Peinado, H., Portillo, F., and Cano, A. (2004) Int. J. Dev. Biol., 48, Koizume, S., Tachibana, K., Sekiya, T., Hirohashi, S., and Shiraishi, M. (2002) Nucleic Acids Res., 30, Cano, A., Perez-Moreno, M. A., Rodrigo, I., Locasciop, A., Blanco, M. J., del Barrio, M. G., Portillo, F., and Nieto, M. A. (2000) Nat. Cell Biol., 2, Comijn, J., Berx, G., Vermassen, P., Verschueren, K., van Grunsven, L., Bruyneel, E., Mareel, M., Huylebroeck, D., and van Roy, F. (2001) Mol. Cell, 7, Bolos, V., Peinado, H., Perez-Moreno, M. A., Fraga, M. F., Esteller, M., and Cano, A. (2003) J. Cell Sci., 116, Blanco, M. J., Moreno-Bueno, G., Sarrio, D., Locascio, A., Cano, A., Palacios, J., and Nieto, M. A. (2002) Oncogene, 21, Jiao, W., Miyazaki, K., and Kitajima, Y. (2002) Br. J. Cancer, 86, Batlle, E., Sancho, E., Franci, C., Dominguez, D., Monfar, M., Baulida, J., and Garcia de Herreros, A. (2000) Nat. Cell Biol., 2, Barrallo-Gimeno, A., and Nieto, M. A. (2005) Development, 132, Peinado, H., Olmeda, D., and Cano, A. (2007) Nat. Rev. Cancer, 7, Larue, L., and Bellacosa, A. (2005) Oncogene, 24, Alexander, N. R., Tran, N. L., Rekapally, H., Summers, C. E., Glackin, C., and Heimark, R. L. (2006) Cancer Res., 66, De Craene, B., Gilbert, B., Stove, C., Bruyneel, E., van Roy, F., and Berx, G. (2005) Cancer Res., 65, Reynolds, A. B. (2007) Biochim. Biophys. Acta, 1773, Ireton, R. C., Davis, M. A., van Hengel, J., Mariner, D. J., Barnes, K., Thoreson, M. A., Anastasiadis, P. Z., Matrisian, L., Bundy, L. M., Sealy, L., Gilbert, B., van Roy, F., and Reynolds, A. B. (2002) J. Cell Biol., 159, Hoshino, T., Sakisaka, T., Baba, T., Yamada, T., Kimura, T., and Takai, Y. (2005) J. Biol. Chem., 280, Fujita, Y., Krause, G., Scheffner, M., Zechner, D., Leddy, H. E., Behrens, J., Sommer, T., and Birchmeier, W. (2002) Nat. Cell Biol., 4, Dupre-Crochet, S., Figueroa, A., Hogan, C., Ferber, E. C., Bialucha, C. U., Adams, J., Richardson, E. C., and Fujita, Y. (2007) Mol. Cell. Biol., 27, Kamei, T., Matozaki, T., Sakisaka, T., Kodama, A., Yokoyama, S., Peng, Y. F., Nakano, K., Takaishi, K., and Takai, Y. (1999) Oncogene, 18, Xiao, K., Allison, D. F., Buckley, K. M., Kottke, M. D., Vincent, P. A., Faundez, V., and Kowalczyk, A. P. (2003) J. Cell Biol., 163, Bryant, D. M., Wylie, F. G., and Stow, J. L. (2005) Mol. Biol. Cell, 16, Kimura, T., Sakisaka, T., Baba, T., Yamada, T., and Takai, Y. (2006) J. Biol. Chem., 281, Suzuki, K., and Takahashi, K. (2006) Biochem. Biophys. Res. Commun., 349, Downing, J. R., and Reynolds, A. B. (1991) Oncogene, 6, Palacios, F., Tushir, J. S., Fujita, Y., and D Souza-Schorey, C. (2005) Mol. Cell. Biol., 25, Mariner, D. J., Anastasiadis, P., Keilhack, H., Bohmer, F. D., Wang, J., and Reynolds, A. B. (2001) J. Biol. Chem., 276, Yang, J. Y., Zong, C. S., Xia, W., Wei, Y., Ali-Seyed, M., Li, Z., Broglio, K., Berry, D. A., and Hung, M. C. (2006) Mol. Cell. Biol., 26, Thoreson, M. A., and Reynolds, A. B. (2002) Differentiation, 70, Chen, Y. T., Stewart, D. B., and Nelson, W. J. (1999) J. Cell Biol., 144, Lilien, J., and Balsamo, J. (2005) Curr. Opin. Cell Biol., 17, Roura, S., Miravet, S., Piedra, J., Garcia de Herreros, A., and Dunach, M. (1999) J. Biol. Chem., 274, Piedra, J., Miravet, S., Castano, J., Palmer, H. G., Heisterkamp, N., Garcia de Herreros, A., and Dunach, M. (2003) Mol. Cell. Biol., 23, Behrens, J., von Kries, J. P., Kuhl, M., Bruhn, L., Wedlich, D., Grosschedl, R., and Birchmeier, W. (1996) Nature, 382, Huber, O., Korn, R., McLaughlin, J., Ohsugi, M., Herrmann, B. G., and Kemler, R. (1996) Mech. Dev., 59, He, T. C., Sparks, A. B., Rago, C., Hermeking, H., Zawel, L., da Costa, L. T., Morin, P. J., Vogelstein, B., and Kinzler, K. W. (1998) Science, 281, Shtutman, M., Zhurinsky, J., Simcha, I., Albanese, C., D Amico, M., Pestell, R., and Ben-Ze ev, A. (1999) Proc. Natl. Acad. Sci. USA, 96, Tetsu, O., and McCormick, F. (1999) Nature, 398, Shimokawa, T., Furukawa, Y., Sakai, M., Li, M., Miwa, N., Lin, Y. M., and Nakamura, Y. (2003) Cancer Res., 63, Gavert, N., and Ben-Ze ev, A. (2007) J. Cell. Biochem., 102, Brembeck, F. H., Rosario, M., and Birchmeier, W. (2006) Curr. Opin. Genet. Dev., 16, Brembeck, F. H., Schwarz-Romond, T., Bakkers, J., Wilhelm, S., Hammerschmidt, M., and Birchmeier, W. (2004) Genes Dev., 18, Behrens, J., Jerchow, B. A., Wurtele, M., Grimm, J., Asbrand, C., Wirtz, R., Kuhl, M., Wedlich, D., and Birchmeier, W. (1998) Science, 280, Kishida, S., Yamamoto, H., Ikeda, S., Kishida, M., Sakamoto, I., Koyama, S., and Kikuchi, A. (1998) J. Biol. Chem., 273,
9 750 GLOUSHANKOVA 112. Liu, C., Li, Y., Semenov, M., Han, C., Baeg, G. H., Tan, Y., Zhang, Z., Lin, X., and He, X. (2002) Cell, 108, Pinson, K. I., Brennan, J., Monkley, S., Avery, B. J., and Skarnes, W. C. (2000) Nature, 407, Tolwinski, N. S., Wehrli, M., Rives, A., Erdeniz, N., DiNardo, S., and Wieschaus, E. (2003) Dev. Cell, 4, Kramps, T., Peter, O., Brunner, E., Nellen, D., Froesch, B., Chatterjee, S., Murone, M., Zullig, S., and Basler, K. (2002) Cell, 109, Townsley, F. M., Cliffe, A., and Bienz, M. (2004) Nat. Cell Biol., 6, Adachi, S., Jigami, T., Yasui, T., Nakano, T., Ohwada, S., Omori, Y., Sugano, S., Ohkawara, B., Shibuya, H., Nakamura, T., and Akiyama, T. (2004) Cancer Res., 64, Herzig, M., Savarese, F., Novatchkova, M., Semb, H., and Christofori, G. (2007) Oncogene, 26, Morin, P. J., Sparks, A. B., Korinek, V., Barker, N., Clevers, H., Vogelstein, B., and Kinzler, K. W. (1997) Science, 275, De La Coste, A., Romagnolo, B., Billuart, P., Renard, C. A., Buendia, M. A., Soubrane, O., Fabre, M., Chelly, J., Beldjord, C., Kahn, A., and Perret, C. (1998) Proc. Natl. Acad. Sci. USA, 95, Satoh, S., Daigo, Y., Furukawa, Y., Kato, T., Miwa, N., Nishiwaki, T., Kawasoe, T., Ishiguro, H., Fujita, M., Tokino, T., Sasaki, Y., Imaoka, S., Murata, M., Shimano, T., Yamaoka, Y., and Nakamura, Y. (2000) Nat. Genet., 24, Groden, J., Thliveris, A., Samowitz, W., Carlson, M., Gelbert, L., Albertsen, H., Joslyn, G., Stevens, J., Spirio, L., and Robertson, M. (1991) Cell, 66, Kawasaki, Y., Sato, R., and Akiyama, T. (2003) Nat. Cell Biol., 5, Zhou, B. P., Deng, J., Xia, W., Xu, J., Li, Y. M., Gunduz, M., and Hung, M. C. (2004) Nat. Cell Biol., 6, Frame, S., and Cohen, P. (2001) Biochem. J., 359, Ciruna, B., and Sleeman, J. (2006) Dev. Cell, 1, Thiery, J. P., and Sleeman, J. (2006) Nat. Rev. Mol. Cell. Biol., 7, Chamorro, M. N., Schwartz, D. R., Vonica, A., Brivanlou, A. H., Cho, K. R., and Varmus, H. E. (2005) EMBO J., 24, Katoh, M., and Katoh, M. (2006) Int. J. Oncol., 29, El Sayegh, T. Y., Kapus, A., and McCulloch, C. A. (2007) FEBS Lett., 581, El Sayegh, T. Y., Arora, P. D., Fan, L., Laschinger, C. A., Lee, W., Greer, P. A., Christopher, A., McCulloch, C. A., and Kapus, A. (2004) Mol. Biol. Cell, 16, Derycke, L. D. M., and Bracke, M. E. (2004) Int. J. Dev., 48, Nieman, M. T., Prudoff, R. S., Johnson, K. R., and Wheelock, M. J. (1999) J. Cell Biol., 147, Hazan, R. B., Phillips, G. R., Qiao, R. F., Norton, L., and Aaronson, S. A. (2000) J. Cell Biol., 148, Gaggioli, C., Hooper, S., Hidalgo-Carcedo, C., Grosse, R., Marshall, J. F., Harrington, K., and Sahai, E. (2007) Nat. Cell. Biol., 9, Suyama, K., Shapiro, I., Guttman, M., and Hazan, R. B. (2002) Cancer Cell, 2, Theisen, C. S., Wahl, J. K., III, Johnson, K. R., and Wheelock, M. J. (2007) Mol. Biol. Cell, 18,
Cell Polarity and Cancer
 Cell Polarity and Cancer Pr Jean-Paul Borg Email: jean-paul.borg@inserm.fr Features of malignant cells Steps in Malignant Progression Cell polarity, cell adhesion, morphogenesis and tumorigenesis pathways
Cell Polarity and Cancer Pr Jean-Paul Borg Email: jean-paul.borg@inserm.fr Features of malignant cells Steps in Malignant Progression Cell polarity, cell adhesion, morphogenesis and tumorigenesis pathways
VIII Curso Internacional del PIRRECV. Some molecular mechanisms of cancer
 VIII Curso Internacional del PIRRECV Some molecular mechanisms of cancer Laboratorio de Comunicaciones Celulares, Centro FONDAP Estudios Moleculares de la Celula (CEMC), ICBM, Facultad de Medicina, Universidad
VIII Curso Internacional del PIRRECV Some molecular mechanisms of cancer Laboratorio de Comunicaciones Celulares, Centro FONDAP Estudios Moleculares de la Celula (CEMC), ICBM, Facultad de Medicina, Universidad
Cell Cell Communication
 IBS 8102 Cell, Molecular, and Developmental Biology Cell Cell Communication January 29, 2008 Communicate What? Why do cells communicate? To govern or modify each other for the benefit of the organism differentiate
IBS 8102 Cell, Molecular, and Developmental Biology Cell Cell Communication January 29, 2008 Communicate What? Why do cells communicate? To govern or modify each other for the benefit of the organism differentiate
Cell Cell Communication
 IBS 8102 Cell, Molecular, and Developmental Biology Cell Cell Communication January 29, 2008 Communicate What? Why do cells communicate? To govern or modify each other for the benefit of the organism differentiate
IBS 8102 Cell, Molecular, and Developmental Biology Cell Cell Communication January 29, 2008 Communicate What? Why do cells communicate? To govern or modify each other for the benefit of the organism differentiate
Cell Motility. Junfeng Ji ( 纪俊峰 ), PhD
 Cell Motility Junfeng Ji ( 纪俊峰 ), PhD Professor Laboratory of Somatic Reprogramming and Stem Cell Aging Center for Stem Cell and Tissue Engineering School of Medicine Zhejiang University Email: jijunfeng@zju.edu.cnedu
Cell Motility Junfeng Ji ( 纪俊峰 ), PhD Professor Laboratory of Somatic Reprogramming and Stem Cell Aging Center for Stem Cell and Tissue Engineering School of Medicine Zhejiang University Email: jijunfeng@zju.edu.cnedu
Rearrangements of the Actin Cytoskeleton and E-Cadherin Based Adherens Junctions Caused by Neoplasic Transformation Change Cell Cell Interactions
 Rearrangements of the Actin Cytoskeleton and E-Cadherin Based Adherens Junctions Caused by Neoplasic Transformation Change Cell Cell Interactions Dmitry V. Ayollo., Irina Y. Zhitnyak., Jury M. Vasiliev,
Rearrangements of the Actin Cytoskeleton and E-Cadherin Based Adherens Junctions Caused by Neoplasic Transformation Change Cell Cell Interactions Dmitry V. Ayollo., Irina Y. Zhitnyak., Jury M. Vasiliev,
MCB Topic 19 Regulation of Actin Assembly- Prof. David Rivier
 MCB 252 -Topic 19 Regulation of Actin Assembly- Prof. David Rivier MCB 252 Spring 2017 MCB 252 Cell Biology Topic 19 Regulation of Actin Assembly Reading: Lodish 17.2-17.3, 17.7 MCB 252 Actin Cytoskeleton
MCB 252 -Topic 19 Regulation of Actin Assembly- Prof. David Rivier MCB 252 Spring 2017 MCB 252 Cell Biology Topic 19 Regulation of Actin Assembly Reading: Lodish 17.2-17.3, 17.7 MCB 252 Actin Cytoskeleton
TITLE: Mechanism of Cadherin Switching in Breast Carcinoma
 AD Award Number: W81XWH-04-1-0339 TITLE: Mechanism of Cadherin Switching in Breast Carcinoma PRINCIPAL INVESTIGATOR: David I. Bellovin CONTRACTING ORGANIZATION: Beth Israel Deaconess Medical Center, Incorporated
AD Award Number: W81XWH-04-1-0339 TITLE: Mechanism of Cadherin Switching in Breast Carcinoma PRINCIPAL INVESTIGATOR: David I. Bellovin CONTRACTING ORGANIZATION: Beth Israel Deaconess Medical Center, Incorporated
Development of Carcinoma Pathways
 The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
Tumor microenvironment Interactions and Lung Cancer Invasiveness. Pulmonary Grand Rounds Philippe Montgrain, M.D.
 Tumor microenvironment Interactions and Lung Cancer Invasiveness Pulmonary Grand Rounds Philippe Montgrain, M.D. February 26, 2009 Objectives Review epithelial mesenchymal transition (EMT), and its implications
Tumor microenvironment Interactions and Lung Cancer Invasiveness Pulmonary Grand Rounds Philippe Montgrain, M.D. February 26, 2009 Objectives Review epithelial mesenchymal transition (EMT), and its implications
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
In vitro scratch assay: method for analysis of cell migration in vitro labeled fluorodeoxyglucose (FDG)
 In vitro scratch assay: method for analysis of cell migration in vitro labeled fluorodeoxyglucose (FDG) 1 Dr Saeb Aliwaini 13/11/2015 Migration in vivo Primary tumors are responsible for only about 10%
In vitro scratch assay: method for analysis of cell migration in vitro labeled fluorodeoxyglucose (FDG) 1 Dr Saeb Aliwaini 13/11/2015 Migration in vivo Primary tumors are responsible for only about 10%
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
CMB621: Cytoskeleton. Also known as How the cell plays with LEGOs to ensure order, not chaos, is temporally and spatially achieved
 CMB621: Cytoskeleton Also known as How the cell plays with LEGOs to ensure order, not chaos, is temporally and spatially achieved Lecture(s) Overview Lecture 1: What is the cytoskeleton? Membrane interaction
CMB621: Cytoskeleton Also known as How the cell plays with LEGOs to ensure order, not chaos, is temporally and spatially achieved Lecture(s) Overview Lecture 1: What is the cytoskeleton? Membrane interaction
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
THE HALLMARKS OF CANCER
 THE HALLMARKS OF CANCER ONCOGENES - Most of the oncogenes were first identified in retroviruses: EGFR (ErbB), Src, Ras, Myc, PI3K and others (slightly more than 30) - Mutated cellular genes incorporated
THE HALLMARKS OF CANCER ONCOGENES - Most of the oncogenes were first identified in retroviruses: EGFR (ErbB), Src, Ras, Myc, PI3K and others (slightly more than 30) - Mutated cellular genes incorporated
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
Roles of transcriptional factor Snail and adhesion factor E-cadherin in clear cell renal cell carcinoma
 EXPERIMENTAL AND THERAPEUTIC MEDICINE 6: 1489-1493, 2013 Roles of transcriptional factor Snail and adhesion factor E-cadherin in clear cell renal cell carcinoma JINQUAN CAI Department of Urology, Fuzhou
EXPERIMENTAL AND THERAPEUTIC MEDICINE 6: 1489-1493, 2013 Roles of transcriptional factor Snail and adhesion factor E-cadherin in clear cell renal cell carcinoma JINQUAN CAI Department of Urology, Fuzhou
Biol403 MAP kinase signalling
 Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
1. The metastatic cascade. 3. Pathologic features of metastasis. 4. Therapeutic ramifications. Which malignant cells will metastasize?
 1. The metastatic cascade 3. Pathologic features of metastasis 4. Therapeutic ramifications Sir James Paget (1814-1899) British Surgeon/ Pathologist Paget s disease of Paget s disease of the nipple (intraductal
1. The metastatic cascade 3. Pathologic features of metastasis 4. Therapeutic ramifications Sir James Paget (1814-1899) British Surgeon/ Pathologist Paget s disease of Paget s disease of the nipple (intraductal
PIP 2 + PIP 2. MCB3m CS5 Membrane cytoskeletal Interactions S.K.Maciver Feb, 2001
 MB3m S5 Membrane cytoskeletal Interactions S.K.Maciver Feb, 2001 Membrane ytoskeleton Interactions S5 The cytoskeleton, especially the actin cytoskeleton interacts with cell membranes. The cell cortex
MB3m S5 Membrane cytoskeletal Interactions S.K.Maciver Feb, 2001 Membrane ytoskeleton Interactions S5 The cytoskeleton, especially the actin cytoskeleton interacts with cell membranes. The cell cortex
Transcriptional regulation of cadherins during development and carcinogenesis
 Int. J. Dev. Biol. 48: 365-375 (2004) Transcriptional regulation of cadherins during development and carcinogenesis HÉCTOR PEINADO, FRANCISCO PORTILLO and AMPARO CANO* Department of Biochemistry, Faculty
Int. J. Dev. Biol. 48: 365-375 (2004) Transcriptional regulation of cadherins during development and carcinogenesis HÉCTOR PEINADO, FRANCISCO PORTILLO and AMPARO CANO* Department of Biochemistry, Faculty
1.The metastatic cascade. 2.Pathologic features of metastasis. 3.Therapeutic ramifications
 Metastasis 1.The metastatic cascade 2.Pathologic features of metastasis 3.Therapeutic ramifications Sir James Paget (1814-1899) British Surgeon/ Pathologist Paget s disease of bone Paget s disease of the
Metastasis 1.The metastatic cascade 2.Pathologic features of metastasis 3.Therapeutic ramifications Sir James Paget (1814-1899) British Surgeon/ Pathologist Paget s disease of bone Paget s disease of the
ABL TYROSINE KINASES MEDIATE INTERCELLULAR ADHESION. Nicole L. Zandy. Department of Pharmacology and Cancer Biology Duke University.
 ABL TYROSINE KINASES MEDIATE INTERCELLULAR ADHESION by Nicole L. Zandy Department of Pharmacology and Cancer Biology Duke University Date: Approved: Ann Marie Pendergast, Supervisor Christopher Counter
ABL TYROSINE KINASES MEDIATE INTERCELLULAR ADHESION by Nicole L. Zandy Department of Pharmacology and Cancer Biology Duke University Date: Approved: Ann Marie Pendergast, Supervisor Christopher Counter
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Animal Tissue Culture SQG 3242 Biology of Cultured Cells. Dr. Siti Pauliena Mohd Bohari
 Animal Tissue Culture SQG 3242 Biology of Cultured Cells Dr. Siti Pauliena Mohd Bohari The Culture Environment Changes of Cell s microenvironment needed that favor the spreading, migration, and proliferation
Animal Tissue Culture SQG 3242 Biology of Cultured Cells Dr. Siti Pauliena Mohd Bohari The Culture Environment Changes of Cell s microenvironment needed that favor the spreading, migration, and proliferation
Biochemistry of Carcinogenesis. Lecture # 35 Alexander N. Koval
 Biochemistry of Carcinogenesis Lecture # 35 Alexander N. Koval What is Cancer? The term "cancer" refers to a group of diseases in which cells grow and spread unrestrained throughout the body. It is difficult
Biochemistry of Carcinogenesis Lecture # 35 Alexander N. Koval What is Cancer? The term "cancer" refers to a group of diseases in which cells grow and spread unrestrained throughout the body. It is difficult
CHAPTER VII CONCLUDING REMARKS AND FUTURE DIRECTION. Androgen deprivation therapy is the most used treatment of de novo or recurrent
 CHAPTER VII CONCLUDING REMARKS AND FUTURE DIRECTION Stathmin in Prostate Cancer Development and Progression Androgen deprivation therapy is the most used treatment of de novo or recurrent metastatic PCa.
CHAPTER VII CONCLUDING REMARKS AND FUTURE DIRECTION Stathmin in Prostate Cancer Development and Progression Androgen deprivation therapy is the most used treatment of de novo or recurrent metastatic PCa.
Actin structure. Actin a highly conserved gene. Molecular Cell Biology
 Harvey Lodish Arnold Berk Paul Matsudaira Chris A. Kaiser Monty Krieger Matthew P. Scott Lawrence Zipursky James Darnell Molecular Cell Biology Fifth Edition Chapter 19: Cytoskeleton I: Microfilaments
Harvey Lodish Arnold Berk Paul Matsudaira Chris A. Kaiser Monty Krieger Matthew P. Scott Lawrence Zipursky James Darnell Molecular Cell Biology Fifth Edition Chapter 19: Cytoskeleton I: Microfilaments
Cancer Biology Course. Invasion and Metastasis
 Cancer Biology Course Invasion and Metastasis 2016 Lu-Hai Wang NHRI Cancer metastasis Major problem: main reason for killing cancer patients, without it cancer can be cured or controlled. Challenging questions:
Cancer Biology Course Invasion and Metastasis 2016 Lu-Hai Wang NHRI Cancer metastasis Major problem: main reason for killing cancer patients, without it cancer can be cured or controlled. Challenging questions:
Molecular biology :- Cancer genetics lecture 11
 Molecular biology :- Cancer genetics lecture 11 -We have talked about 2 group of genes that is involved in cellular transformation : proto-oncogenes and tumour suppressor genes, and it isn t enough to
Molecular biology :- Cancer genetics lecture 11 -We have talked about 2 group of genes that is involved in cellular transformation : proto-oncogenes and tumour suppressor genes, and it isn t enough to
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Cell signaling. How do cells receive and respond to signals from their surroundings?
 Cell signaling How do cells receive and respond to signals from their surroundings? Prokaryotes and unicellular eukaryotes are largely independent and autonomous. In multicellular organisms there is a
Cell signaling How do cells receive and respond to signals from their surroundings? Prokaryotes and unicellular eukaryotes are largely independent and autonomous. In multicellular organisms there is a
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Negative Regulation of c-myc Oncogenic Activity Through the Tumor Suppressor PP2A-B56α
 Negative Regulation of c-myc Oncogenic Activity Through the Tumor Suppressor PP2A-B56α Mahnaz Janghorban, PhD Dr. Rosalie Sears lab 2/8/2015 Zanjan University Content 1. Background (keywords: c-myc, PP2A,
Negative Regulation of c-myc Oncogenic Activity Through the Tumor Suppressor PP2A-B56α Mahnaz Janghorban, PhD Dr. Rosalie Sears lab 2/8/2015 Zanjan University Content 1. Background (keywords: c-myc, PP2A,
Cytoskeleton and cell communication
 Cytoskeleton and cell communication Actin monomer has subdomains 1-4. A simplified cartoon is at right. ATP binds, along with Mg ++, within a deep cleft between subdomains 2 & 4. G-actin (globular actin),
Cytoskeleton and cell communication Actin monomer has subdomains 1-4. A simplified cartoon is at right. ATP binds, along with Mg ++, within a deep cleft between subdomains 2 & 4. G-actin (globular actin),
1. Activated receptor tyrosine kinases (RTKs) phosphorylates themselves
 Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Discovery and Optimization of Inhibitors of STAT3 Activation for the Treatment of Squamous Cell Carcinoma of the Head and Neck
 Discovery and ptimization of Inhibitors of STAT3 Activation for the Treatment of Squamous Cell Carcinoma of the Head and Neck Feng Zhang Wipf Group Research Topic Seminar 02-09-2013 1 Feng Zhang @ Wipf
Discovery and ptimization of Inhibitors of STAT3 Activation for the Treatment of Squamous Cell Carcinoma of the Head and Neck Feng Zhang Wipf Group Research Topic Seminar 02-09-2013 1 Feng Zhang @ Wipf
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
The Angiopoietin Axis in Cancer
 Ang2 Ang1 The Angiopoietin Axis in Cancer Tie2 An Overview: The Angiopoietin Axis Plays an Essential Role in the Regulation of Tumor Angiogenesis Growth of a tumor beyond a limiting size is dependent upon
Ang2 Ang1 The Angiopoietin Axis in Cancer Tie2 An Overview: The Angiopoietin Axis Plays an Essential Role in the Regulation of Tumor Angiogenesis Growth of a tumor beyond a limiting size is dependent upon
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
The cytoskeleton and cell movement. (Actin microfilaments)
 The cytoskeleton and cell movement (Actin microfilaments) What is the cytoskeleton? A dynamic network of protein filaments extending throughout the cytoplasm Three types: microfilaments (actin), microtubules
The cytoskeleton and cell movement (Actin microfilaments) What is the cytoskeleton? A dynamic network of protein filaments extending throughout the cytoplasm Three types: microfilaments (actin), microtubules
Impact of NM23-M1 "knock-out" on metastatic potential of hepatocellular carcinoma: a transgenic approach
 Impact of NM23-M1 "knock-out" on metastatic potential of hepatocellular carcinoma: a transgenic approach NM23-M1 "knock- out" mice (without NDPK A) Experimental models of hepatocarcinogenesis Chemical
Impact of NM23-M1 "knock-out" on metastatic potential of hepatocellular carcinoma: a transgenic approach NM23-M1 "knock- out" mice (without NDPK A) Experimental models of hepatocarcinogenesis Chemical
Plasticity of Tumor Cell Migration: Acquisition of New Properties or Return to the Past?
 ISSN 0006-2979, Biochemistry (Moscow), 2014, Vol. 79, No. 9, pp. 947-963. Pleiades Publishing, Ltd., 2014. Published in Russian in Biokhimiya, 2014, Vol. 79, No. 9, pp. 1169-1187. REVIEW Plasticity of
ISSN 0006-2979, Biochemistry (Moscow), 2014, Vol. 79, No. 9, pp. 947-963. Pleiades Publishing, Ltd., 2014. Published in Russian in Biokhimiya, 2014, Vol. 79, No. 9, pp. 1169-1187. REVIEW Plasticity of
PCB 3023 Exam 4 - Form A First and Last Name
 PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
The Hallmarks of Cancer
 The Hallmarks of Cancer Theresa L. Hodin, Ph.D. Clinical Research Services Theresa.Hodin@RoswellPark.org Hippocrates Cancer surgery, circa 1689 Cancer Surgery Today 1971: Nixon declares War on Cancer
The Hallmarks of Cancer Theresa L. Hodin, Ph.D. Clinical Research Services Theresa.Hodin@RoswellPark.org Hippocrates Cancer surgery, circa 1689 Cancer Surgery Today 1971: Nixon declares War on Cancer
CADHERINS AS MODULATORS OF CELLULAR PHENOTYPE
 Annu. Rev. Cell Dev. Biol. 2003. 19:207 35 doi: 10.1146/annurev.cellbio.19.011102.111135 Copyright c 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on June 20,
Annu. Rev. Cell Dev. Biol. 2003. 19:207 35 doi: 10.1146/annurev.cellbio.19.011102.111135 Copyright c 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on June 20,
Neoplasia 18 lecture 8. Dr Heyam Awad MD, FRCPath
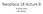 Neoplasia 18 lecture 8 Dr Heyam Awad MD, FRCPath ILOS 1. understand the angiogenic switch in tumors and factors that stimulate and inhibit angiogenesis. 2. list the steps important for tumor metastasis
Neoplasia 18 lecture 8 Dr Heyam Awad MD, FRCPath ILOS 1. understand the angiogenic switch in tumors and factors that stimulate and inhibit angiogenesis. 2. list the steps important for tumor metastasis
CELL BIOLOGY - CLUTCH CH CELL JUNCTIONS AND TISSUES.
 !! www.clutchprep.com CONCEPT: CELL-CELL ADHESION Cells must be able to bind and interact with nearby cells in order to have functional and strong tissues Cells can in two main ways - Homophilic interactions
!! www.clutchprep.com CONCEPT: CELL-CELL ADHESION Cells must be able to bind and interact with nearby cells in order to have functional and strong tissues Cells can in two main ways - Homophilic interactions
Wnt signaling. Ramray Bhat.
 Wnt signaling Ramray Bhat ramray@mrdg.iisc.ernet.in Starting with animal biology and viral infections The discovery of certain laboratory murine strains that were highly susceptible to mammary gland cancer.
Wnt signaling Ramray Bhat ramray@mrdg.iisc.ernet.in Starting with animal biology and viral infections The discovery of certain laboratory murine strains that were highly susceptible to mammary gland cancer.
Genetics and Cancer Ch 20
 Genetics and Cancer Ch 20 Cancer is genetic Hereditary cancers Predisposition genes Ex. some forms of colon cancer Sporadic cancers ~90% of cancers Descendants of cancerous cells all cancerous (clonal)
Genetics and Cancer Ch 20 Cancer is genetic Hereditary cancers Predisposition genes Ex. some forms of colon cancer Sporadic cancers ~90% of cancers Descendants of cancerous cells all cancerous (clonal)
The Tissue Engineer s Toolkit
 The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Intercellular indirect communication
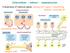 Intercellular indirect communication transmission of chemical signals: sending cell signal transmitting tissue hormone medium receiving cell hormone intercellular fluid blood neurocrine neurotransmitter
Intercellular indirect communication transmission of chemical signals: sending cell signal transmitting tissue hormone medium receiving cell hormone intercellular fluid blood neurocrine neurotransmitter
Contents. Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX
 Contents Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX 1 General Aspects of Signal Transduction and Cancer Therapy 1 1.1 General Principles of Signal Transduction
Contents Preface XV Acknowledgments XXI List of Abbreviations XXIII About the Companion Website XXIX 1 General Aspects of Signal Transduction and Cancer Therapy 1 1.1 General Principles of Signal Transduction
G-Protein-Coupled Receptors
 Cellular Signalling Cells must be ready to respond to essential signals in their environment. These are often chemicals in the extracellular fluid (ECF) from distant locations in a multicellular organism
Cellular Signalling Cells must be ready to respond to essential signals in their environment. These are often chemicals in the extracellular fluid (ECF) from distant locations in a multicellular organism
oncogenes-and- tumour-suppressor-genes)
 Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Advanced Cell Biology. Lecture 36
 Advanced Cell Biology. Lecture 36 Alexey Shipunov Minot State University May 3, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 36 May 3, 2013 1 / 43 Outline Questions and answers Cellular communities
Advanced Cell Biology. Lecture 36 Alexey Shipunov Minot State University May 3, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 36 May 3, 2013 1 / 43 Outline Questions and answers Cellular communities
A RENAISSANCE FOR SRC
 A RENAISSANCE FOR SRC Timothy J. Yeatman The c-src non-receptor tyrosine kinase is overexpressed and activated in a large number of human malignancies and has been linked to the development of cancer and
A RENAISSANCE FOR SRC Timothy J. Yeatman The c-src non-receptor tyrosine kinase is overexpressed and activated in a large number of human malignancies and has been linked to the development of cancer and
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Karyotype analysis reveals transloction of chromosome 22 to 9 in CML chronic myelogenous leukemia has fusion protein Bcr-Abl
 Chapt. 18 Cancer Molecular Biology of Cancer Student Learning Outcomes: Describe cancer diseases in which cells no longer respond Describe how cancers come from genomic mutations (inherited or somatic)
Chapt. 18 Cancer Molecular Biology of Cancer Student Learning Outcomes: Describe cancer diseases in which cells no longer respond Describe how cancers come from genomic mutations (inherited or somatic)
Angiostasis and Angiogenesis Regulated by Angiopoietin1-Tie2 Receptor System
 Japan-Mexico Workshop on Pharmacology and Nanobiology Feb. 25, 2009; Universidad Nacional Autönoma de Mëxico, Mexico City Angiostasis and Angiogenesis Regulated by Angiopoietin1-Tie2 Receptor System Shigetomo
Japan-Mexico Workshop on Pharmacology and Nanobiology Feb. 25, 2009; Universidad Nacional Autönoma de Mëxico, Mexico City Angiostasis and Angiogenesis Regulated by Angiopoietin1-Tie2 Receptor System Shigetomo
Chapter 20. Cell - Cell Signaling: Hormones and Receptors. Three general types of extracellular signaling. endocrine signaling. paracrine signaling
 Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Growth and Differentiation Phosphorylation Sampler Kit
 Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
α-catenin in the adherens junction complex
 α-catenin in the adherens junction complex and the possible implications in cancer M domain of αe-catenin Utrecht, 23-12-2011 M.F. Pustjens 3019462 Supervisor: P.W.B. Derksen, PhD Department of Pathology
α-catenin in the adherens junction complex and the possible implications in cancer M domain of αe-catenin Utrecht, 23-12-2011 M.F. Pustjens 3019462 Supervisor: P.W.B. Derksen, PhD Department of Pathology
Fundamental research on breast cancer in Belgium. Rosita Winkler
 Fundamental research on breast cancer in Belgium Rosita Winkler Medline search for «breast cancer» and Belgium limits: english, posted in the last 5 years. Result: 484 papers - fundamental / clinical -
Fundamental research on breast cancer in Belgium Rosita Winkler Medline search for «breast cancer» and Belgium limits: english, posted in the last 5 years. Result: 484 papers - fundamental / clinical -
Neoplasia 18 lecture 6. Dr Heyam Awad MD, FRCPath
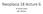 Neoplasia 18 lecture 6 Dr Heyam Awad MD, FRCPath ILOS 1. understand the role of TGF beta, contact inhibition and APC in tumorigenesis. 2. implement the above knowledge in understanding histopathology reports.
Neoplasia 18 lecture 6 Dr Heyam Awad MD, FRCPath ILOS 1. understand the role of TGF beta, contact inhibition and APC in tumorigenesis. 2. implement the above knowledge in understanding histopathology reports.
BL 424 Test pts name Multiple choice have one choice each and are worth 3 points.
 BL 424 Test 3 2010 150 pts name Multiple choice have one choice each and are worth 3 points. 1. The plasma membrane functions as a a. selective barrier to the passage of molecules. b. sensor through which
BL 424 Test 3 2010 150 pts name Multiple choice have one choice each and are worth 3 points. 1. The plasma membrane functions as a a. selective barrier to the passage of molecules. b. sensor through which
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis
 Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
supplementary information
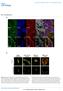 DOI: 10.1038/ncb2133 Figure S1 Actomyosin organisation in human squamous cell carcinoma. (a) Three examples of actomyosin organisation around the edges of squamous cell carcinoma biopsies are shown. Myosin
DOI: 10.1038/ncb2133 Figure S1 Actomyosin organisation in human squamous cell carcinoma. (a) Three examples of actomyosin organisation around the edges of squamous cell carcinoma biopsies are shown. Myosin
Correlation of Snail expression with histological grade and lymph node status in breast carcinomas
 (2002) 21, 3241 ± 3246 ã 2002 Nature Publishing Group All rights reserved 0950 ± 9232/02 $25.00 www.nature.com/onc SHORT REPORTS Correlation of Snail expression with histological grade and lymph node status
(2002) 21, 3241 ± 3246 ã 2002 Nature Publishing Group All rights reserved 0950 ± 9232/02 $25.00 www.nature.com/onc SHORT REPORTS Correlation of Snail expression with histological grade and lymph node status
Dr Mahmood S Choudhery, PhD, Postdoc (USA) Assistant Professor Tissue Engineering and Regenerative Medicine King Edward Medical University/Mayo
 Integration of Cells into Tissues Dr Mahmood S Choudhery, PhD, Postdoc (USA) Assistant Professor Tissue Engineering and Regenerative Medicine King Edward Medical University/Mayo Hospital Lahore 1. How
Integration of Cells into Tissues Dr Mahmood S Choudhery, PhD, Postdoc (USA) Assistant Professor Tissue Engineering and Regenerative Medicine King Edward Medical University/Mayo Hospital Lahore 1. How
Supplementary Figure 1: Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points
 A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
A. B. 8 4 Supplementary Figure : Func8onal Network Analysis of Kinases Significantly Modulated by MERS CoV Infec8on and Conserved Across All Time Points Examined. A) Venn diagram analysis of kinases significantly
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
EMT: Epithelial Mesenchimal Transition
 EMT: Epithelial Mesenchimal Transition A phenotypic change that is characteristic of some developing tissues and certain forms of cancer. This is a multistep, key process in embryonic development and metastasis
EMT: Epithelial Mesenchimal Transition A phenotypic change that is characteristic of some developing tissues and certain forms of cancer. This is a multistep, key process in embryonic development and metastasis
Src-INACTIVE / Src-INACTIVE
 Biology 169 -- Exam 1 February 2003 Answer each question, noting carefully the instructions for each. Repeat- Read the instructions for each question before answering!!! Be as specific as possible in each
Biology 169 -- Exam 1 February 2003 Answer each question, noting carefully the instructions for each. Repeat- Read the instructions for each question before answering!!! Be as specific as possible in each
Cancer genetics
 Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
number Done by Corrected by Doctor Maha Shomaf
 number 21 Done by Ahmad Rawajbeh Corrected by Omar Sami Doctor Maha Shomaf Ability to Invade and Metastasize The metastatic cascade can be subdivided into two phases: 1-invasion of ECM and vascular dissemination:
number 21 Done by Ahmad Rawajbeh Corrected by Omar Sami Doctor Maha Shomaf Ability to Invade and Metastasize The metastatic cascade can be subdivided into two phases: 1-invasion of ECM and vascular dissemination:
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#5:(October-18-2010) Cancer and Signals Outline 1 2 Outline 1 2 Cancer is a disease of malfunctioning cells. Cell Lineage: Adult
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#5:(October-18-2010) Cancer and Signals Outline 1 2 Outline 1 2 Cancer is a disease of malfunctioning cells. Cell Lineage: Adult
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Principles of cell signaling Lecture 4
 Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
B eta catenin was first identified as a protein associated
 347 ORIGINAL ARTICLE Nuclear b catenin as a potential prognostic and diagnostic marker in patients with colorectal cancer from Hong Kong S C C Wong, E S F Lo, A K C Chan, K C Lee, W L Hsiao... See end
347 ORIGINAL ARTICLE Nuclear b catenin as a potential prognostic and diagnostic marker in patients with colorectal cancer from Hong Kong S C C Wong, E S F Lo, A K C Chan, K C Lee, W L Hsiao... See end
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) All of the following are synthesized along various sites of the endoplasmic reticulum
Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) All of the following are synthesized along various sites of the endoplasmic reticulum
Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber
 Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber jweber@dom.wustl.edu Oncogenes & Cancer DNA Tumor Viruses Simian Virus 40 p300 prb p53 Large T Antigen Human Adenovirus p300 E1A
Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber jweber@dom.wustl.edu Oncogenes & Cancer DNA Tumor Viruses Simian Virus 40 p300 prb p53 Large T Antigen Human Adenovirus p300 E1A
number Done by Corrected by Doctor Maha Shomaf
 number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. 2 3 4 DOI: 10.1038/NMAT4893 EGFR and HER2 activate rigidity sensing only on rigid matrices Mayur Saxena 1,*, Shuaimin Liu 2,*, Bo Yang 3, Cynthia Hajal
In the format provided by the authors and unedited. 2 3 4 DOI: 10.1038/NMAT4893 EGFR and HER2 activate rigidity sensing only on rigid matrices Mayur Saxena 1,*, Shuaimin Liu 2,*, Bo Yang 3, Cynthia Hajal
Top 10 Contributions on Biomedical Sciences: 2nd Edition. St Vincent s Hospital, The University of Melbourne, Australia 2
 Chapter 06 Cancer Progression: The Impact of Cytoskeletal Molecules Sam L Francis 1 and Juliana Antonipillai 2 * 1 St Vincent s Hospital, The University of Melbourne, Australia 2 School of Health and Biomedical
Chapter 06 Cancer Progression: The Impact of Cytoskeletal Molecules Sam L Francis 1 and Juliana Antonipillai 2 * 1 St Vincent s Hospital, The University of Melbourne, Australia 2 School of Health and Biomedical
Introduction. Cancer Biology. Tumor-suppressor genes. Proto-oncogenes. DNA stability genes. Mechanisms of carcinogenesis.
 Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Tala Saleh. Ahmad Attari. Mamoun Ahram
 23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
A class of genes that normally suppress cell proliferation. p53 and Rb..ect. suppressor gene products can release cells. hyperproliferation.
 Tumor Suppressor Genes A class of genes that normally suppress cell proliferation. p53 and Rb..ect Mutations that inactivate the tumor suppressor gene products can release cells from growth suppression
Tumor Suppressor Genes A class of genes that normally suppress cell proliferation. p53 and Rb..ect Mutations that inactivate the tumor suppressor gene products can release cells from growth suppression
Computational Biology I LSM5191
 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Crosstalk between Adiponectin and IGF-IR in breast cancer. Prof. Young Jin Suh Department of Surgery The Catholic University of Korea
 Crosstalk between Adiponectin and IGF-IR in breast cancer Prof. Young Jin Suh Department of Surgery The Catholic University of Korea Obesity Chronic, multifactorial disorder Hypertrophy and hyperplasia
Crosstalk between Adiponectin and IGF-IR in breast cancer Prof. Young Jin Suh Department of Surgery The Catholic University of Korea Obesity Chronic, multifactorial disorder Hypertrophy and hyperplasia
Disorders of Cell Growth & Neoplasia. Lecture 4 Molecular basis of cancer
 General Pathology VPM 152 Disorders of Cell Growth & Neoplasia Lecture 4 Molecular basis of cancer Enrique Aburto Apr 2010 Skin tumor in a 10-year-old Rottweiler. Considering the external appearance and
General Pathology VPM 152 Disorders of Cell Growth & Neoplasia Lecture 4 Molecular basis of cancer Enrique Aburto Apr 2010 Skin tumor in a 10-year-old Rottweiler. Considering the external appearance and
Enzymes Part III: regulation II. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
