Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
|
|
|
- Eileen Lewis
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs. Single or multiple mirna precursor sequences arranged in tandem were placed downstream of a GFP gene to monitor expression driven by a CMV promoter in a lentiviral vector. Sensors harboring four repeats of the mirna target sequence were cloned in the 3 UTR of a RFP gene expressed from an UBC promoter to report on mirna activity. The constructs were delivered by lentiviruses to HEK293T cells and then analyzed for GFP and RFP expression using flow cytometry. b, Repression of RFP reporter activity by mirna expression. Lentiviral constructs harboring a mirna, the cognate sensor, or both were introduced into HEK293T cells. c, Combinatorial mirna expression constructs effectively repressed RFP reporters containing the cognate mirna sensors. Lentiviral constructs harboring two-wise or three-wise mirna combinations, with or without the cognate sensors, were introduced into HEK293T cells. d, Limited cross-reactivity between mirnas and non-cognate sensors. Lentiviral constructs harboring mirnas paired with different sensors were delivered into HEK293T cells. The percentages of RFP-positive cells within GFP-positive cell populations were determined by flow cytometry. Data represent mean ± s.d. (n = 3).
2 Supplementary Figure 2 Efficient Lentiviral Delivery of a Dual-Fluorescent Protein Reporter Construct in Human Cells. a, Strategy for testing lentiviral delivery of a dual-fluorescent protein reporter construct in human cells. Lentiviruses generated for vectors containing a GFP gene expressed from a CMV or UBC promoter, or a single vector encoding RFP and GFP genes under the UBC and CMV promoters, respectively, were delivered to HEK293T cells for analysis of GFP and RFP expression. b, c, Lentiviral delivery and expression of the dual-fluorescent protein reporter construct in human cells. (b) Fluorescent microscopy revealed that RFP and GFP were expressed in UBCp-RFP-CMVp-GFP virus-infected cells, whereas only GFP was expressed in cells infected with UBCp- GFP and CMVp-GFP lentiviruses. Scale bar denotes 400 μm. (c) Flow cytometry was used to measure cell populations positive for RFP and GFP fluorescence. Over 97 percent of UBCp-RFP-CMVp-GFP virus-infected HEK293T cells were positive for both RFP and GFP, and similar percentages of UBCp-GFP or CMVp-GFP virus-infected cells were GFP-positive.
3 Supplementary Figure 3 Identification of the Exponential Phase During PCR for CombiGEM Barcode Amplification. a, b, Procedures for identifying the transition point from exponential to linear phase during PCR for CombiGEM barcode amplification. The one-wise mirna vector library pooled-assembled in E. coli (a) and the genomic DNA isolated from human breast cancer cells (MCF7) infected with the two-wise library (b) were used as templates in replicate PCR reactions, and the barcodes representing each mirna combinations were amplified using primers targeting the sequences located outside the barcode region. PCR products were collected from the reactions stopped at cycles between 10 to 20 (a) or 19 to 28 (b), and were then diluted as templates for quantitative PCR reactions. The mean difference of threshold cycle (Ct) between cycles was determined. Error bars indicate s.d. from triplicates. Primer efficiencies were estimated to be 102% (a) and 100% (b) respectively. PCR cycle numbers highlighted in magenta (a) and green (b) were used in unbiased barcode amplification for subsequent Illumina sequencing. c, d, Agarose gel analyses of the amplified PCR products with indicated cycle numbers from (a) and (b) are shown in (c) and (d), respectively.
4 Supplementary Figure 4 High Reproducibility of Barcode Quantitation in Biological Replicates for Combinatorial mirna Screens. a, b, Scatter plots showing high correlation between barcode representations (log 2 number of normalized barcode counts) between two biological replicates for both docetaxel (25 nm)-treated or vehicle-treated OVCAR8-ADR cells infected with the two-wise (a) or threewise (b) mirna combinatorial libraries. R is Pearson correlation coefficient.
5 Supplementary Figure 5 Consistent Fold Changes of Barcodes among Same mirna Combinations Arranged in Different Orders in the Expression Constructs. a-c, The coefficient of variation (CV; defined as s.d./mean of the fold changes of normalized barcode counts for docetaxel (25 nm)- treated versus four-day vehicle-treated (a, b) and four-day versus one-day cultured cells (c)) was determined for the same two-wise (a) or three-wise (b, c) combination arranged in different orders (i.e., [A,B] vs. [B,A] for two-wise combinations; [A,B,C], [A,C,B], [B,A,C], [B,C,A], [C,A,B], and [C,B,A] for three-wise combinations). 92% of two-wise mirna combinations had a CV of <0.2 in the drugsensitivity screen (a), while 95% and 98% of three-wise combinations showed a CV of <0.2 in the drug-sensitivity (b) and cellproliferation (c) screens respectively.
6 Supplementary Figure 6 Docetaxel Dose-Response Curves for the OVCAR8 Cell Line and the Docetaxel-Resistant OVCAR8-ADR Cell Line. OVCAR8 cells and OVCAR8-ADR cells (OVCAR8 s docetaxel-resistant derivative) cells were treated with docetaxel at indicated doses for three days and subjected to the MTT assay. Cell viabilities were compared to their respective no drug controls. The OVCAR8-ADR cell line has a ~3-fold higher IC 50 than the parental OVCAR8 cell line. Data represent mean ± s.d. (n = 3).
7 Supplementary Figure 7 Log 2 Fold-Changes in Barcode Representation Between Biological Replicates for All Individual Combinations in the Pooled Screens. a, b (upper panel), Log 2 fold-changes in biological replicate 1 is plotted against replicate 2 for mean values of normalized barcode counts for docetaxel (25 nm)-treated versus vehicle-treated OVCAR8-ADR cells (a), and for relative cell viability at day 4 versus day 1 (b), for each three-wise combination. a, b (lower panel), Distributions of differences in the log 2 fold change between two biological replicates at a bin size of 0.1 are shown. A majority of three-wise combinations (88-90%) had <0.3 log 2 fold-change differences. R is the Pearson correlation coefficient.
8 Supplementary Figure 8 Three-dimensional Plots Depicting the Docetaxel-Sensitizing (a) and Proliferation-Modulating (b) Effects of Three-Wise mirna Combinations. The log 2 ratios of the normalized barcode counts for docetaxel-treated versus four-day vehicle-treated OVCAR8-ADR cells (a) or fourday versus one-day cultured cells (b) were determined for all three-wise mirna combinations, and were presented as colored bubbles. mirna combinations with drug-resistance and drug-sensitization effects (a) have log 2 ratios >0 and <0, respectively, and those with pro-proliferation and anti-proliferation effects (b) have log 2 ratios >0 and <0, respectively. Each two-dimensional plane was arranged in the same hierarchically clustered order as in (Fig. 5a-c), and the additional third mirna element is labeled. All log 2 ratios shown were determined from the mean of two biological replicates.
9 Supplementary Figure 9 Definitions of Genetic Interactions (GIs) in This Study. Synergistic or buffering interactions have positive and negative GI scores, respectively, as described for case 1 to 7 in this figure. Positive and negative phenotypes have fold-changes of normalized barcode reads of >1 and <1 respectively, while no phenotype corresponds to a fold-change = 1. For mirnas [A] and [B] with individual phenotypes A and B, the expected phenotype for the twowise combination [A,B] is ( A + B - 1) according to the additive model. Deviation was calculated by subtracting the expected phenotype from observed phenotype (i.e., Observed phenotype Expected phenotype).
10 Supplementary Figure 10 GI Scores Between Biological Replicates for All Individual Combinations in the Pooled Screens. a-c (upper panel), GI scores in biological replicate 1 are plotted against replicate 2 for docetaxel (25 nm)-treated versus vehicle-treated OVCAR8-ADR cells for each two-wise combination (a) and three-wise mirna combination (b) respectively, and for relative cell viability at day 4 versus day 1 for each three-wise combination (c). a-c (lower panel), Distributions of differences in the GI scores between two biological replicates at a bin size of 0.1 are shown. A majority of combinations (80-93%) had GI score differences of <0.2.
11 Supplementary Figure 11 Synergistic Interactions between the mir-16-1/15a Cluster, mir-128b, and the let-7e/mir-99b Cluster Modulate Cell Proliferation Phenotypes. a, GI scores for a given three-wise mirna combination [A,B,C] are plotted and compared to other combinations that harbor two of the same mirnas (in the first two positions) and every other mirna member in our library (denoted as X in the third position). GI scores of each three-wise mirna combination were calculated as described in Online Methods, and represent the interaction between the additional third mirna and the two-wise mirna combinations. GI scores were determined for the three possible permutations (i.e., [A,B,C], [B,C,A], and [A,C,B], see Online Methods). In this example, A, B, and C represent the mir-16-1/15a cluster, mir-128b, and the let-7e/mir-99b cluster respectively, and X represents all 39 library members. b-d, GI maps based on cell-proliferation phenotypes for all three-wise mirna combinations harboring the mir-16-1/15a cluster, mir-128b, and/or the let-7e/mir-99b cluster are displayed in the same order as for the GI map in Fig. 5a for easy comparison, with the additional third mirna element in the three-wise combination being labeled above each GI map. The combinations for which no GIs were measured are indicated in gray. The combinations with GI at a Q-value of <0.003 are boxed.
12 Supplementary Figure 12 Three-Wise mirna Combinations Display Distinct Docetaxel Sensitivity and Anti-Proliferation Phenotypes. Fold changes of normalized barcode counts for docetaxel (25 nm)-treated versus vehicle-treated OVCAR8-ADR cells (y-axis) and also for four-day versus one-day cultured cells (x-axis) were plotted for all three-wise mirna combinations. Each data point represents the mean of two biological replicates.
13 Supplementary Figure 13 Combinatorial Expression of the mir-16-1/15a Cluster, mir-128b, and the let-7e/mir-99b Cluster Inhibits Colony Formation by Viable OVCAR8-ADR Cells. a, b, ~10,000 OVCAR8-ADR cells infected with each indicated mirna combinations were treated with 25 nm of docetaxel for three days, and were cultured for another eleven days. Cells were stained with crystal violet to visualize colony formation for quantification. Representative images are shown in (a). The number of colonies for each sample was determined (b). The maximum number of discrete colonies that could be reliably counted was ~500 per well, and thus, samples with more than 500 colonies are presented as >500 colonies. Data represent mean ± s.d. (n = 3; *P < 0.05).
14 Supplementary Figure 14 High Consistency between Pooled Screens and Validation Data for Individual Hits. For each two-wise and three-wise mirna combination, the fold-change in the normalized barcode count for docetaxel (25 nm)-treated versus vehicle-treated OVCAR8-ADR cells, or four-day versus one-day cultured cells, obtained from the pooled screening data ( Screen phenotype ) was plotted against its relative cell viability compared to vector control determined from the individual drug-sensitivity or cell-proliferation assays, respectively ( Validation phenotype ) (R = 0.899). Data for the screening data are the mean of two biological replicates while the individual validation data represent the mean of three independent experiments. R is the Pearson correlation coefficient.
15 Supplementary Figure 15 Combinatorial Expression of the mir-16/15a Cluster, mir-128b, and/or the let-7e/mir-99b Cluster Reduce mrna Levels of Targeted Genes in OVCAR8-ADR Cells.
16 a, qrt-pcr quantification of relative mrna levels in OVCAR8-ADR cells expressing the mir-16/15a cluster or co-expressing the mir- 16/15a cluster, mir-128b, and the let-7e/mir-99b cluster. Measured mrna levels were normalized to GAPDH mrna levels, and data represent mean ± s.d. (n = 3). mrnas that were predicted or validated to contain conserved sites matching the seed region of the corresponding mirnas using TargetScan and mirtarbase are shaded in orange in the table below the graph. Significant differences in the mrna levels of CCND1, CCND3, CCNE1 and CHEK1 in cells expressing the mir-16/15a cluster (orange #; P < 0.05) or coexpressing the mir-16/15a cluster, mir-128b, and the let-7e/mir-99b cluster (purple #; P < 0.05) were determined by comparison with vector-control-infected cells. The asterisk (*P < 0.05) represents a significant difference in mrna levels between cells expressing the mir-16/15a cluster versus co-expressing the mir-16/15a cluster, mir-128b, and the let-7e/mir-99b cluster. b, Relative mrna levels of CDC14B in cells expressing various combinations of the mir-16/15a cluster, mir-128b, and/or the let-7e/mir-99b cluster. The mrna level of CDC14B was significantly reduced in cells co-expressing the let-7e/mir-99b cluster and mir-128b, or the triple-combination expressing the mir-16/15a cluster, mir-128b, and the let-7e/mir-99b cluster. Data represent mean ± s.d. (n = 9; *P < 0.05). c, Summary diagram illustrating the potential roles of the mir-16/15a cluster, mir-128b, and the let-7e/mir-99b cluster in regulating the mrna levels of multiple downstream targets. These genes may represent targets for future investigation into mechanisms that can modulate docetaxel resistance and/or proliferation phenotypes in OVCAR8-ADR cells.
SUPPLEMENTARY FIGURES AND TABLES
 SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
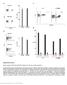 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Supplementary Information and Figure legends
 Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Information and Figure legends Table S1. Primers for quantitative RT-PCR Target Sequence (5 -> 3 ) Target Sequence (5 -> 3 ) DAB2IP F:TGGACGATGTGCTCTATGCC R:GGATGGTGATGGTTTGGTAG Snail F:CCTCCCTGTCAGATGAGGAC
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1: Experimental design. DISCOVERY PHASE VALIDATION PHASE (N = 88) (N = 20) Healthy = 20. Healthy = 6. Endometriosis = 33
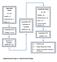 DISCOVERY PHASE (N = 20) Healthy = 6 Endometriosis = 7 EAOC = 7 Quantitative PCR (mirnas = 1113) a Quantitative PCR Verification of Candidate mirnas (N = 24) VALIDATION PHASE (N = 88) Healthy = 20 Endometriosis
DISCOVERY PHASE (N = 20) Healthy = 6 Endometriosis = 7 EAOC = 7 Quantitative PCR (mirnas = 1113) a Quantitative PCR Verification of Candidate mirnas (N = 24) VALIDATION PHASE (N = 88) Healthy = 20 Endometriosis
Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent Protein.
 prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
SUPPLEMENTARY MATERIAL
 SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
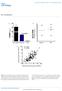 DOI: 10.1038/ncb2210 b. ICAM1 ng ml -1 P = 0.0001 Small RNA (15-30nts) ng ml -1 Cell Lysate Exosome HDL Plasma HDL Normal Human HDL mirnas R = 0.45 P < 0.0001 Normal Human Exosome mirnas Figure S1. Characterization
DOI: 10.1038/ncb2210 b. ICAM1 ng ml -1 P = 0.0001 Small RNA (15-30nts) ng ml -1 Cell Lysate Exosome HDL Plasma HDL Normal Human HDL mirnas R = 0.45 P < 0.0001 Normal Human Exosome mirnas Figure S1. Characterization
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Material
 Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Material Summary: The supplementary information includes 1 table (Table S1) and 4 figures (Figure S1 to S4). Supplementary Figure Legends Figure S1 RTL-bearing nude mouse model. (A) Tumor
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supplementary Information
 Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier
 Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting
 mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Functional genomics reveal that the serine synthesis pathway is essential in breast cancer
 Functional genomics reveal that the serine synthesis pathway is essential in breast cancer Results Presented by Stacey Lin Lloyd Lab http://www.amsbio.com/expression-ready-lentiviral-particles.aspx Overview
Functional genomics reveal that the serine synthesis pathway is essential in breast cancer Results Presented by Stacey Lin Lloyd Lab http://www.amsbio.com/expression-ready-lentiviral-particles.aspx Overview
Gene expression scaled by distance to the genome replication site. Ying et al. Supporting Information. Contents. Supplementary note I p.
 Gene expression scaled by distance to the genome replication site Ying et al Supporting Information Contents Supplementary note I p. 2 Supplementary note II p. 3-5 Supplementary note III p. 5-7 Supplementary
Gene expression scaled by distance to the genome replication site Ying et al Supporting Information Contents Supplementary note I p. 2 Supplementary note II p. 3-5 Supplementary note III p. 5-7 Supplementary
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
SUPPLEMENTARY INFORMATION
 -. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
-. -. SUPPLEMENTARY INFORMATION DOI: 1.1/ncb86 a WAT-1 WAT- BAT-1 BAT- sk-muscle-1 sk-muscle- mir-133b mir-133a mir-6 mir-378 mir-1 mir-85 mir-378 mir-6a mir-18 mir-133a mir- mir- mir-341 mir-196a mir-17
Figure 1. Possible role of oncogene activation, receptor, G-protein mutation, or tumor
 Figures Part of introduction Figure 1. Possible role of oncogene activation, receptor, G-protein mutation, or tumor supressor gene deletion in the induction of thyroid carcinoma. ( by James A Fagin, M.D.)
Figures Part of introduction Figure 1. Possible role of oncogene activation, receptor, G-protein mutation, or tumor supressor gene deletion in the induction of thyroid carcinoma. ( by James A Fagin, M.D.)
Electron micrograph of phosphotungstanic acid-stained exosomes derived from murine
 1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplemental Figures:
 Supplemental Figures: Figure 1: Intracellular distribution of VWF by electron microscopy in human endothelial cells. a) Immunogold labeling of LC3 demonstrating an LC3-positive autophagosome (white arrow)
Supplemental Figures: Figure 1: Intracellular distribution of VWF by electron microscopy in human endothelial cells. a) Immunogold labeling of LC3 demonstrating an LC3-positive autophagosome (white arrow)
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
Nature Medicine: doi: /nm.2109
 HIV 1 Infects Multipotent Progenitor Cells Causing Cell Death and Establishing Latent Cellular Reservoirs Christoph C. Carter, Adewunmi Onafuwa Nuga, Lucy A. M c Namara, James Riddell IV, Dale Bixby, Michael
HIV 1 Infects Multipotent Progenitor Cells Causing Cell Death and Establishing Latent Cellular Reservoirs Christoph C. Carter, Adewunmi Onafuwa Nuga, Lucy A. M c Namara, James Riddell IV, Dale Bixby, Michael
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1
 Supplementary Figure 1 14 12 SEM4C PLXN2 8 SEM4C C 3 Cancer Cell Non Cancer Cell Expression 1 8 6 6 4 log2 ratio Expression 2 1 4 2 2 p value.1 D Supplementary Figure 1. Expression of Sema4C and Plexin2
Supplementary Figure 1 14 12 SEM4C PLXN2 8 SEM4C C 3 Cancer Cell Non Cancer Cell Expression 1 8 6 6 4 log2 ratio Expression 2 1 4 2 2 p value.1 D Supplementary Figure 1. Expression of Sema4C and Plexin2
EXOTESTTM. ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids
 DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
<10. IL-1β IL-6 TNF + _ TGF-β + IL-23
 3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
3 ns 25 ns 2 IL-17 (pg/ml) 15 1 ns ns 5 IL-1β IL-6 TNF
Retro-X qrt-pcr Titration Kit User Manual
 Takara Bio USA Retro-X qrt-pcr Titration Kit User Manual Cat. No. 631453 PT3952-1 (030218) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support: techus@takarabio.com United States/Canada
Takara Bio USA Retro-X qrt-pcr Titration Kit User Manual Cat. No. 631453 PT3952-1 (030218) 1290 Terra Bella Avenue, Mountain View, CA 94043, USA U.S. Technical Support: techus@takarabio.com United States/Canada
Supplementary Information. Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase
 Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion
 Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Supplementary Figure 1. Basal level EGFR across a panel of ESCC lines. Immunoblots demonstrate the expression of phosphorylated and total EGFR as
 Supplementary Figure 1. Basal level EGFR across a panel of ESCC lines. Immunoblots demonstrate the expression of phosphorylated and total EGFR as well as their downstream effectors across a panel of ESCC
Supplementary Figure 1. Basal level EGFR across a panel of ESCC lines. Immunoblots demonstrate the expression of phosphorylated and total EGFR as well as their downstream effectors across a panel of ESCC
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
SUPPLEMENTAY FIGURES AND TABLES
 SUPPLEMENTAY FIGURES AND TABLES Supplementary Figure S1: Validation of mir expression by quantitative real-time PCR and time course studies on mir- 29a-3p and mir-210-3p. A. The graphs illustrate the expression
SUPPLEMENTAY FIGURES AND TABLES Supplementary Figure S1: Validation of mir expression by quantitative real-time PCR and time course studies on mir- 29a-3p and mir-210-3p. A. The graphs illustrate the expression
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Differential expression of mirnas from the pri-mir-17-92a locus.
 Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
GFP/Iba1/GFAP. Brain. Liver. Kidney. Lung. Hoechst/Iba1/TLR9!
 Supplementary information a +KA Relative expression d! Tlr9 5!! 5! NSC Neuron Astrocyte Microglia! 5! Tlr7!!!! NSC Neuron Astrocyte! GFP/Sβ/! Iba/Hoechst Microglia e Hoechst/Iba/TLR9! GFP/Iba/GFAP f Brain
Supplementary information a +KA Relative expression d! Tlr9 5!! 5! NSC Neuron Astrocyte Microglia! 5! Tlr7!!!! NSC Neuron Astrocyte! GFP/Sβ/! Iba/Hoechst Microglia e Hoechst/Iba/TLR9! GFP/Iba/GFAP f Brain
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the
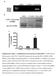 Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
NC bp. b 1481 bp
 Kcna3 NC 11 178 p 1481 p 346 p c *** *** d relative expression..4.3.2.1 CD4 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna Kcna6 Kcna7 Kcnn4 relative expression..4.3.2.1 CD8 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna
Kcna3 NC 11 178 p 1481 p 346 p c *** *** d relative expression..4.3.2.1 CD4 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna Kcna6 Kcna7 Kcnn4 relative expression..4.3.2.1 CD8 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information. Novel lentiviral vectors with mutated reverse transcriptase for mrna delivery of TALE nucleases
 Supplementary Information Novel lentiviral vectors with mutated reverse transcriptase for mrna delivery of TALE nucleases Ulrike Mock 1, Kristoffer Riecken 1, Belinda Berdien 1, Waseem Qasim 2, Emma Chan
Supplementary Information Novel lentiviral vectors with mutated reverse transcriptase for mrna delivery of TALE nucleases Ulrike Mock 1, Kristoffer Riecken 1, Belinda Berdien 1, Waseem Qasim 2, Emma Chan
Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse
 Supplemental figure legends Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse hippocampus targets GFAP-positive but not NeuN-positive cells. (A) Stereotaxic
Supplemental figure legends Supplemental Figure 1. Intracranial transduction of a modified ptomo lentiviral vector in the mouse hippocampus targets GFAP-positive but not NeuN-positive cells. (A) Stereotaxic
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures
 Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
Type of file: PDF Size of file: 0 KB Title of file for HTML: Supplementary Information Description: Supplementary Figures Supplementary Figure 1 mir-128-3p is highly expressed in chemoresistant, metastatic
Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2
 Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2 activity. Upper panel: Representative histograms for FACS
Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2 activity. Upper panel: Representative histograms for FACS
Spatially resolved multiparametric single cell analysis. Technical Journal Club 19th September 2017 Christina Müller (Group Speck)
 Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
Protein SD Units (P-value) Cluster order
 SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
SUPPLEMENTAL TABLE AND FIGURES Table S1. Signature Phosphoproteome of CD22 E12 Transgenic Mouse BPL Cells. T-test vs. Other Protein SD Units (P-value) Cluster order ATPase (Ab-16) 1.41 0.000880 1 mtor
Supplementary Figure 1 The plasmids used for the GPCR-CRISPR ChaCha and the GPCR-CRISPR Tango systems. (a) The plasmids for the GPCR-CRISPR ChaCha
 Supplementary Figure 1 The plasmids used for the GPCR-CRISPR ChaCha and the GPCR-CRISPR Tango systems. (a) The plasmids for the GPCR-CRISPR ChaCha system. The dcas9-vpr effector is fused to the C-terminus
Supplementary Figure 1 The plasmids used for the GPCR-CRISPR ChaCha and the GPCR-CRISPR Tango systems. (a) The plasmids for the GPCR-CRISPR ChaCha system. The dcas9-vpr effector is fused to the C-terminus
Supplementary Figure S1. Generation of LSL-EZH2 conditional transgenic mice.
 Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
Downstream Col1A locus S P P P EP Genotyping with P1, P2 frt PGKneopA + frt hygro-pa Targeting vector Genotyping with P3, P4 P1 pcag-flpe P2 P3 P4 frt SApA CAG LSL PGKATG frt hygro-pa C. D. E. ormal KRAS
(A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and a
 Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary figure legends Supplementary Figure 1. Expression of Shh signaling components in a panel of gastric cancer. (A) RT-PCR for components of the Shh/Gli pathway in normal fetus cell (MRC-5) and
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Supplementary Figure 1. BMS enhances human T cell activation in vitro in a
 Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Supplementary Figure 1. BMS98662 enhances human T cell activation in vitro in a concentration-dependent manner. Jurkat T cells were activated with anti-cd3 and anti-cd28 antibody in the presence of titrated
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
Supplemental Information. Derivation of Human Trophoblast Stem Cells
 Cell Stem Cell, Volume 22 Supplemental Information Derivation of Human Trophoblast Stem Cells Hiroaki Okae, Hidehiro Toh, Tetsuya Sato, Hitoshi Hiura, Sota Takahashi, Kenjiro Shirane, Yuka Kabayama, Mikita
Cell Stem Cell, Volume 22 Supplemental Information Derivation of Human Trophoblast Stem Cells Hiroaki Okae, Hidehiro Toh, Tetsuya Sato, Hitoshi Hiura, Sota Takahashi, Kenjiro Shirane, Yuka Kabayama, Mikita
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
Supplementary Materials
 Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
Supplementary Information. Tissue-wide immunity against Leishmania. through collective production of nitric oxide
 Supplementary Information Tissue-wide immunity against Leishmania through collective production of nitric oxide Romain Olekhnovitch, Bernhard Ryffel, Andreas J. Müller and Philippe Bousso Supplementary
Supplementary Information Tissue-wide immunity against Leishmania through collective production of nitric oxide Romain Olekhnovitch, Bernhard Ryffel, Andreas J. Müller and Philippe Bousso Supplementary
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Pre-made Reporter Lentivirus for MAPK/ERK Signal Pathway
 Pre-made Reporter for MAPK/ERK Signal Pathway Cat# Product Name Amounts LVP957-P or: LVP957-P-PBS SRE-GFP (Puro) LVP958-P or: LVP958-P-PBS SRE-RFP (Puro) LVP959-P or: LVP959-P-PBS SRE-Luc (Puro) LVP960-P
Pre-made Reporter for MAPK/ERK Signal Pathway Cat# Product Name Amounts LVP957-P or: LVP957-P-PBS SRE-GFP (Puro) LVP958-P or: LVP958-P-PBS SRE-RFP (Puro) LVP959-P or: LVP959-P-PBS SRE-Luc (Puro) LVP960-P
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
4. Th1-related gene expression in infected versus mock-infected controls from Fig. 2 with gene annotation.
 List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
Cellecta Overview. Started Operations in 2007 Headquarters: Mountain View, CA
 Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Pre-made Reporter Lentivirus for JAK-STAT Signaling Pathway
 Pre-made Reporter for JAK-STAT Signaling Pathway Cat# Product Name Amounts LVP937-P or: LVP937-P-PBS ISRE-GFP (Puro) LVP938-P or: LVP938-P-PBS ISRE-RFP (Puro) LVP939-P or: LVP939-P-PBS ISRE-Luc (Puro)
Pre-made Reporter for JAK-STAT Signaling Pathway Cat# Product Name Amounts LVP937-P or: LVP937-P-PBS ISRE-GFP (Puro) LVP938-P or: LVP938-P-PBS ISRE-RFP (Puro) LVP939-P or: LVP939-P-PBS ISRE-Luc (Puro)
Supplementary Fig. 1: Ex vivo tetramer enrichment with anti-c-myc beads
 Supplementary Fig. 1: Ex vivo tetramer enrichment with anti-c-myc beads Representative example of comparative ex vivo tetramer enrichment performed in three independent experiments with either conventional
Supplementary Fig. 1: Ex vivo tetramer enrichment with anti-c-myc beads Representative example of comparative ex vivo tetramer enrichment performed in three independent experiments with either conventional
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
A263 A352 A204. Pan CK. pstat STAT3 pstat3 STAT3 pstat3. Columns Columns 1-6 Positive control. Omentum. Rectosigmoid A195.
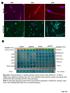 pstat3 75 Pan CK A A263 A352 A24 B Columns 1-6 Positive control A195 A22 A24 A183 Rectal Nodule STAT3 pstat3 STAT3 pstat3 Columns 7-12 Omentum Rectosigmoid Left Ovary Right Ovary Omentum Uterus Uterus
pstat3 75 Pan CK A A263 A352 A24 B Columns 1-6 Positive control A195 A22 A24 A183 Rectal Nodule STAT3 pstat3 STAT3 pstat3 Columns 7-12 Omentum Rectosigmoid Left Ovary Right Ovary Omentum Uterus Uterus
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities
 Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary methods:
 Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
activation with anti-cd3/cd28 beads and 3d following transduction. Supplemental Figure 2 shows
 Supplemental Data Supplemental Figure 1 compares CXCR4 expression in untreated CD8 + T cells, following activation with anti-cd3/cd28 beads and 3d following transduction. Supplemental Figure 2 shows the
Supplemental Data Supplemental Figure 1 compares CXCR4 expression in untreated CD8 + T cells, following activation with anti-cd3/cd28 beads and 3d following transduction. Supplemental Figure 2 shows the
Conditional and reversible disruption of essential herpesvirus protein functions
 nature methods Conditional and reversible disruption of essential herpesvirus protein functions Mandy Glaß, Andreas Busche, Karen Wagner, Martin Messerle & Eva Maria Borst Supplementary figures and text:
nature methods Conditional and reversible disruption of essential herpesvirus protein functions Mandy Glaß, Andreas Busche, Karen Wagner, Martin Messerle & Eva Maria Borst Supplementary figures and text:
SUPPORTING INFORMATION
 SUPPORTING INFORMATION SUPPLEMENTARY FIGURE LEGENDS Fig. S1. Separation of non-dissolved nanoparticles. Tests were conducted on the separation of non-dissolved nanoparticles added in excess to BEGM (A)
SUPPORTING INFORMATION SUPPLEMENTARY FIGURE LEGENDS Fig. S1. Separation of non-dissolved nanoparticles. Tests were conducted on the separation of non-dissolved nanoparticles added in excess to BEGM (A)
Nature Immunology: doi: /ni.3631
 Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane. (a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above
mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons
 Supplemental Material mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons Antoine de Chevigny, Nathalie Coré, Philipp Follert, Marion Gaudin, Pascal
Supplemental Material mir-7a regulation of Pax6 in neural stem cells controls the spatial origin of forebrain dopaminergic neurons Antoine de Chevigny, Nathalie Coré, Philipp Follert, Marion Gaudin, Pascal
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Drd1a-Cre driven ChR2 expression in the SCN. (a) Low-magnification image of a representative Drd1a-ChR2 coronal brain section (n = 2) showing endogenous tdtomato fluorescence (magenta).
Supplementary Figure 1 Drd1a-Cre driven ChR2 expression in the SCN. (a) Low-magnification image of a representative Drd1a-ChR2 coronal brain section (n = 2) showing endogenous tdtomato fluorescence (magenta).
Santulli G. et al. A microrna-based strategy to suppress restenosis while preserving endothelial function
 ONLINE DATA SUPPLEMENTS Santulli G. et al. A microrna-based strategy to suppress restenosis while preserving endothelial function Supplementary Figures Figure S1 Effect of Ad-p27-126TS on the expression
ONLINE DATA SUPPLEMENTS Santulli G. et al. A microrna-based strategy to suppress restenosis while preserving endothelial function Supplementary Figures Figure S1 Effect of Ad-p27-126TS on the expression
Influenza virus exploits tunneling nanotubes for cell-to-cell spread
 Supplementary Information Influenza virus exploits tunneling nanotubes for cell-to-cell spread Amrita Kumar 1, Jin Hyang Kim 1, Priya Ranjan 1, Maureen G. Metcalfe 2, Weiping Cao 1, Margarita Mishina 1,
Supplementary Information Influenza virus exploits tunneling nanotubes for cell-to-cell spread Amrita Kumar 1, Jin Hyang Kim 1, Priya Ranjan 1, Maureen G. Metcalfe 2, Weiping Cao 1, Margarita Mishina 1,
Concentration Estimation from Flow Cytometry Exosome Data Protocol
 Concentration Estimation from Flow Cytometry Exosome Data Protocol 1. STANDARD CURVE Create a standard curve for the target exosome by plotting the mean fluorescence (y axis) against the protein concentration
Concentration Estimation from Flow Cytometry Exosome Data Protocol 1. STANDARD CURVE Create a standard curve for the target exosome by plotting the mean fluorescence (y axis) against the protein concentration
Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas
 Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas Yu Guo 1,2,4, Katherine Alexander 1, Andrew G Clark 1, Andrew Grimson 1 and Haiyuan Yu 2,3* 1 Department
Integrated network analysis reveals distinct regulatory roles of transcription factors and micrornas Yu Guo 1,2,4, Katherine Alexander 1, Andrew G Clark 1, Andrew Grimson 1 and Haiyuan Yu 2,3* 1 Department
