Lecture 10. Eukaryotic gene regulation: chromatin remodelling
|
|
|
- Elfrieda Wells
- 5 years ago
- Views:
Transcription
1 Lecture 10 Eukaryotic gene regulation: chromatin remodelling
2 Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional activators: DNA binding, transactivation Role of chromatin: Acetylation & deacetylation of histones Histone methylation, demethylation, phosphorylation etc., Histone code DNA methylation, epigenetic code
3 Thus far, we have studied chromatin remodelling involving post translational modifications of histones and DNA methylation leading to regulation of gene expression ATP-dependent remodelling of chromatin
4 Chromatin remodelling complex ATP ADP + Pi Displacement of nucleosomes during chromatin remodelling leading to the generation of a nucleosome-free region in the chromatin
5 In addition to nucleosome displacement, ATP-dependent chromatin remodelling may involve: Nucleosome sliding Histone actamers may slide along DNA, thereby exposing specific DNA sequences for interaction with other proteins
6 Adjusting spacing between histone octamers so that certain DNA sequences are exposed
7 Why is ATP required for moving nucleosomes around? Bending DNA around nucleosomes depends on DNA sequence to some extent (Some sequences bend more easily than others) Nucleosomes do have strong affinity for DNA and even have sequence preferences This preference may be exploited in nature to organize a chromatin landscape energetically conducive or not conducive for gene expression But that landscape has to be changed to suit gene expression requirements and this requires energy
8 The first ATP-dependent chromatin remodelling complex discovered: yeast SWI/SNF complex (SWItch/Sucrose NonFermentable) SWI/SNF complex was originally identified genetically as a positive regulator of HO and SUC2 genes: -SWI -HO-gene encodes an endonuclease that is involved in mating type switching in yeast -SNF -Sucrose non-fermentor: SUC2 involved in sucrose- ermentation Several subunits: SWI1, SWI2, SWI3, SNF5, SNF6, SNF11, TFG3/ANC1, SWP73 etc molecules of Swi/Snf per cell) 2 Mega daltons The SWI/SNF complex is needed for transcriptional activation bytranscription factors such as GAL4. In S. cerevisiae, <5% of the total genes require Swi/Snf
9 Swi/Snf binds DNA and nucleosomes in an ATP-dependent manner with high affinity (Kd = nm range). ~1000 ATP molecules are consumed per minute. After binding to DNA, it starts an ATP-dependent remodelling of chromating by displacement of nucleosomes
10 The key component of a chromatin remodelling complex is the ATPase, which provides energy for remodelling by ATP hydrolysis SWI2 is a DNA-stimulated ATPase.
11 Chromatin remodeling complexes are classified based on protein motifs found in addition to the ATPase domain, or on how the ATPase domain itself is structured This classification is purely structural, designed to make it easier for us to sort them all out The classificationi s not necessarily based on functional criteria
12 SWI2/SNF2 ATPase SUPERFAMILY Originally isolated genetically in Saccharomyces cerevisiae as mutants in mating type switching and sucrose non-fermenting functions SWI2/SNF2 subfamily ISWI subfamily CHD/Mi2 subfamily Ino80 subfamily NURF- Nucleosome remodeling factor CHRAC - Chromatin accessibility complex ACF - ATP-dependent chromatin assembly and remodeling factor NURF- Nucleosome remodeling factor is invovled in Transcriptional activation of homeotic genes
13 Different requirements of various nucleosome remodeling complexes SWI/SNF Maximal activity with DNA alone Mi-2 Requires nucleosomal structure Does not require histone tails ISWI Partially activated by DNA Further stimulated by nucleosomes Requires histone H4 tails
14 Nucleosome repositioning ISWI moves nucleosome from central to terminal position CHRAC (contains ISWI) facilitates movement from terminal to central position
15 BRAHMA ATPase Associates with acetylated histones
16 Mechanisms of ATP remodelling Twist defect diffusion model In this model, small local alterations from mean DNA twist ( defects ) propagate around the nucleosome Bulge propagation model In the bulge diffusion model, unwinding of DNA from the histone octamer at the entry/exit of the nucleosome and subsequent rebinding of more distal sequences to the same histone contact points would establish nucleosomal particles harbouring excess DNA. Subsequent migration of the bulge around the nucleosomal superhelix would result in the nucleosome apparently increasing bp Cis vs trans Langst & Becker 2001 Journal of Cell Science
17 Eviction- displacement of histones Dimers of H2A and H2B can be rather easily exchanged in and out of nucleosome Entire histone octamers, including H3 and H4, can also be displaced or exchanged under certain circumstances Variant histone incorporation Many variant forms of histones exist, different from the classical core histones, are often expressed in a cell cycle-specific manner and these are incorporated into chromatin in a DNA replication-independent manner. For example, a chromatin remodelling complex known as SWR1 promotes the exchange of H2A from the nucleosome with a histone variant called H2AZ DNA Repair Certain Chromatin remodelling complexes such as the INO80 complex is involved in DNA repair
18 Swi/Snf recruitment by transcription factors Just as certain transcription factors recruit histone modifying enzymes such as HATs, HDACS, HMTs, DNMTs etc., certain gene-specific activators (GCN4, GAL4-VP16 etc.) recruit Swi/Snfcomplex directly Swi/Snf Purified Swi/Snf complex stimulates binding of GAL4-AH to nucleosomal DNA fold
19 Altering chromatin structure and gene expression by a combination of histone modifications and ATP-dependent chromatin remodelling Targeted acetylation of histone H3 by SAGA stabilizes its binding and that of a targeted SWI/ SNF chromatin-remodeling complex. This requires the bromodomains of Gcn5 and Swi2, respectively, which bind to acetyl-lysine. It appears that different promoters may use different combinations of histone modifiers and chromatin remodellers. There is no general rule that is common to all promoters. Certain activators/co-activator complexes contain components that recognize a specific histone modification as well as those that can remodel chromatin For example, the HURF complex contains BPTF, which can recognize methylated lysine 4 of Histone H3 as well as SNF2L ATPase
20 Chromatin remodelling may depend on nucleotide sequence of promoters Certain promoters are more susceptible or sensitive to chromatin remodelling than others. For ex., AT-rich regions are often nucleosome-free and therefore promoters containing such regions are better primed for transcription activation Transcription factor binding sites are often present in nucleosome-free regions and therefore does not require nucleosome displacement. It is often found that nucleosome density at promoter regions is typically lower than that in the coding region
21 Rotational positioning of nucleosomes A 5 bp sliding of nucleosome can expose a transcription factor binding site In case of the MMTV LTR, 6 partly palindromic sites, each bound by one dimer of glucocorticoid receptor (GR) is present. It also has two adjascent sirtes for binding of NFI and OCTI. Both GR and another TF called NFI cannot bind to MMTV LTR simultaneously on a linear DNA template but they can bind together on a chromatin template. In the absence of hormone, the GR can only bind to two of its four binding sites. In order to bind to the remaining two sites, a change in rotational positioning on the nucleosome is required. When hormone bound GR bind to MMTV LTR chromatin template, it results in recruitment of SWI-SNF complex, leading to dissociation of histone H1 and exposure of binding sites for NF1 + OCTI leading to recruitment of TFIID and ultimately transcriptional activation. Infact, NFI binding to chromatin template can be footprinted after hormone addition but not before.
22 Interactions between transcription factors and remodelling complexes provides important insights into their mechanism of action. For ex., in case of the TF Swi5p, an activator of the HO locus in yeast, It enters the nucleus at the end of mitosis and bind to the HO promoter And recruits SWI/SNF. Swi5p is then released, leaving SMI/SNF at the promoter. Thus, the function of the TF is only to recruit a remodelling complex To the promoter
23 Several genetic diseases are associated with chromatin remodeling Huang et al Curr Opin Genet Dev
24 Fry & Peterson 2001 Curr Biol 11 R Kingston & Narlikar 1999 G&D Wagner 2003 Curr Opin Plant Biol Narlikar et al Cell Langst & Becker 2001 JCS
25 SUMMARY Chromatin remodelling complexes are ATPases that form large multisubunit chromatin remodeling complexes. These complexes use the energy of ATP to remodel nucleosomal DNA. They are associated with activation of chromatin but they can silence as well
26 Cryderman et al. (1999) NAR 27(16): 3364 Euchromatin Heterochromatin
Transcription and chromatin. General Transcription Factors + Promoter-specific factors + Co-activators
 Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Gene Expression DNA RNA. Protein. Metabolites, stress, environment
 Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Stacie Bulfer Proteins Research Proposal April 27, Background
 Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Chromatin Structure & Gene activity part 2
 Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling
 Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
On the way of revealing coactivator complexes cross talk during transcriptional activation
 DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
Control of nuclear receptor function by local chromatin structure
 MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
5. Heterochromatin allows for silencing of large domains of the genome and utilizes a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
6. Heterochromatin allows for silencing of large domains of the genome and appears to utilize a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Epigenetics: The Future of Psychology & Neuroscience. Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1
 Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS
 R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
9/3/2009 DNA i DNA n euk euk yotes Organizatio Organ izatio n of o f gen ge e n tic Locati t on: In n ucleu e s material mater in e ial
 DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
ubiquitinylation succinylation butyrylation hydroxylation crotonylation
 Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Host cell activation
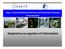 Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Organization of genetic material in eukaryotes
 Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Where Splicing Joins Chromatin And Transcription. 9/11/2012 Dario Balestra
 Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
The Role of Protein Domains in Cell Signaling
 The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
Supplemental Figure 1: Asymmetric chromatin maturation leads to epigenetic asymmetries on sister chromatids.
 Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Watson-Crick Model of B-DNA
 Reading: Ch8; 285-290 Ch24; 963-978 Problems: Ch8 (text); 9 Ch8 (study-guide: facts); 3 Ch24 (text); 5,7,9,10,14,16 Ch24 (study-guide: applying); 1 Ch24 (study-guide: facts); 1,2,4 NEXT Reading: Ch1; 29-34
Reading: Ch8; 285-290 Ch24; 963-978 Problems: Ch8 (text); 9 Ch8 (study-guide: facts); 3 Ch24 (text); 5,7,9,10,14,16 Ch24 (study-guide: applying); 1 Ch24 (study-guide: facts); 1,2,4 NEXT Reading: Ch1; 29-34
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Histone octamer assembly
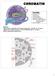 CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Conservation and Expression of SNF2 Proteins in Chondrus crispus
 Conservation and Expression of SNF2 Proteins in Chondrus crispus By Ethan Caudell A thesis submitted to the Faculty of East Carolina University in partial fulfillment of the requirements for the degree
Conservation and Expression of SNF2 Proteins in Chondrus crispus By Ethan Caudell A thesis submitted to the Faculty of East Carolina University in partial fulfillment of the requirements for the degree
BIO 5099: Molecular Biology for Computer Scientists (et al)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Chromatin Remodelling Edited by Danuta Radzioch Contributors Shiaw-Yih Lin, Lili Gong, Edward Wang, Luigi Bagella, Ire
 CHROMATIN REMODELLING Edited by Danuta Radzioch Chromatin Remodelling http://dx.doi.org/10.5772/50815 Edited by Danuta Radzioch Contributors Shiaw-Yih Lin, Lili Gong, Edward Wang, Luigi Bagella, Irene
CHROMATIN REMODELLING Edited by Danuta Radzioch Chromatin Remodelling http://dx.doi.org/10.5772/50815 Edited by Danuta Radzioch Contributors Shiaw-Yih Lin, Lili Gong, Edward Wang, Luigi Bagella, Irene
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
Structure and function of active chromatin and DNase I hypersensitive sites
 MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
Transcriptional and Epigenetic Mechanisms of Addiction
 Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin remodelling in Pol I and III transcription. Erica Cavellán
 Chromatin remodelling in Pol I and III transcription Erica Cavellán Stockholm 2006 Erica Cavellán ISBN 91-7155-229-4 (page 1-65) PrintCenter, Stockholm, 2006 Abstract Compaction of chromosomes in the eukaryotic
Chromatin remodelling in Pol I and III transcription Erica Cavellán Stockholm 2006 Erica Cavellán ISBN 91-7155-229-4 (page 1-65) PrintCenter, Stockholm, 2006 Abstract Compaction of chromosomes in the eukaryotic
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Gene Regulation. Bacteria. Chapter 18: Regulation of Gene Expression
 Chapter 18: Regulation of Gene Expression A Biology 2013 1 Gene Regulation rokaryotes and eukaryotes alter their gene expression in response to their changing environment In multicellular eukaryotes, gene
Chapter 18: Regulation of Gene Expression A Biology 2013 1 Gene Regulation rokaryotes and eukaryotes alter their gene expression in response to their changing environment In multicellular eukaryotes, gene
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being A Eukaryote. Eukaryotic Cells. Basic eukaryotes have:
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
Chapter 11 How Genes Are Controlled
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Genetics. Instructor: Dr. Jihad Abdallah Transcription of DNA
 Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Are you the way you are because of the
 EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
How Cells Divide. Chapter 10
 How Cells Divide Chapter 10 Bacterial Cell Division Bacteria divide by binary fission. -the single, circular bacterial chromosome is replicated -replication begins at the origin of replication and proceeds
How Cells Divide Chapter 10 Bacterial Cell Division Bacteria divide by binary fission. -the single, circular bacterial chromosome is replicated -replication begins at the origin of replication and proceeds
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors
 Pilar Blancafort Associate Professor School of Anatomy, Physiology and Human Biology UWA! Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors The University
Pilar Blancafort Associate Professor School of Anatomy, Physiology and Human Biology UWA! Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors The University
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Gene Regulation Part 2
 Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Role for ADA/GCN5 Products in Antagonizing Chromatin- Mediated Transcriptional Repression
 MOLECULAR AND CELLULAR BIOLOGY, Nov. 1997, p. 6212 6222 Vol. 17, No. 11 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Role for ADA/GCN5 Products in Antagonizing Chromatin- Mediated
MOLECULAR AND CELLULAR BIOLOGY, Nov. 1997, p. 6212 6222 Vol. 17, No. 11 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Role for ADA/GCN5 Products in Antagonizing Chromatin- Mediated
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation gene PHR1
 Published online 3 February 2011 Nucleic Acids Research, 2011, Vol. 39, No. 10 4151 4165 doi:10.1093/nar/gkr040 The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation
Published online 3 February 2011 Nucleic Acids Research, 2011, Vol. 39, No. 10 4151 4165 doi:10.1093/nar/gkr040 The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME
 UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Computational Biology I LSM5191
 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Gene Regulation - 4. One view of the Lactose Operon
 Gene Regulation - 1 Regulating Genes We have been discussing the structure of DNA and that the information stored in DNA is used to direct protein synthesis. We've studied how RNA molecules are used to
Gene Regulation - 1 Regulating Genes We have been discussing the structure of DNA and that the information stored in DNA is used to direct protein synthesis. We've studied how RNA molecules are used to
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Regulation of Gene Expression
 18 Regulation of Gene Expression KEY CONCEPTS Figure 18.1 What regulates the precise pattern of gene expression in the developing wing of a fly embryo? 18.1 Bacteria often respond to environmental change
18 Regulation of Gene Expression KEY CONCEPTS Figure 18.1 What regulates the precise pattern of gene expression in the developing wing of a fly embryo? 18.1 Bacteria often respond to environmental change
Transcription Through Chromatin - Dynamic Organization of Genes
 GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
Identifying the Combinatorial Effects of Histone Modifications by Association Rule Mining in Yeast
 Evolutionary Bioinformatics Original Research Open Access Full open access to this and thousands of other papers at http://www.la-press.com. Identifying the Combinatorial Effects of Histone Modifications
Evolutionary Bioinformatics Original Research Open Access Full open access to this and thousands of other papers at http://www.la-press.com. Identifying the Combinatorial Effects of Histone Modifications
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN
 Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
THE HISTONE CODE AT DNA BREAKS: A GUIDE TO REPAIR? Haico van Attikum and Susan M. Gasser
 THE HISTONE CODE AT DNA BREAKS: A GUIDE TO REPAIR? Haico van Attikum and Susan M. Gasser Abstract Chromatin modifications are important for all cellular processes that involve DNA, including transcription,
THE HISTONE CODE AT DNA BREAKS: A GUIDE TO REPAIR? Haico van Attikum and Susan M. Gasser Abstract Chromatin modifications are important for all cellular processes that involve DNA, including transcription,
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Post-translational modifications of proteins in gene regulation under hypoxic conditions
 203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
'''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Five'Levels'of'Organiza-on' Molecular'
 '''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Anggraini'Barlian,' Iriawa-' Tjandra'Anggraeni' SITH4ITB' Five'Levels'of'Organiza-on' Molecular'
'''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Anggraini'Barlian,' Iriawa-' Tjandra'Anggraeni' SITH4ITB' Five'Levels'of'Organiza-on' Molecular'
Cells. Variation and Function of Cells
 Cells Variation and Function of Cells Cell Theory states that: 1. All living things are made of cells 2. Cells are the basic unit of structure and function in living things 3. New cells are produced from
Cells Variation and Function of Cells Cell Theory states that: 1. All living things are made of cells 2. Cells are the basic unit of structure and function in living things 3. New cells are produced from
The HIV life cycle. integration. virus production. entry. transcription. reverse transcription. nuclear import
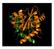 The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
AP Biology
 Tour of the Cell (1) 2007-2008 Types of cells Prokaryote bacteria cells - no organelles - organelles Eukaryote animal cells Eukaryote plant cells Cell Size Why organelles? Specialized structures - specialized
Tour of the Cell (1) 2007-2008 Types of cells Prokaryote bacteria cells - no organelles - organelles Eukaryote animal cells Eukaryote plant cells Cell Size Why organelles? Specialized structures - specialized
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Epigenetics DNA methylation. Biosciences 741: Genomics Fall, 2013 Week 13. DNA Methylation
 Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Chromatin remodeling by ATP-dependent molecular machines
 Chromatin remodeling by ATP-dependent molecular machines Alexandra Lusser and James T. Kadonaga* Summary The eukaryotic genome is packaged into a periodic nucleoprotein structure termed chromatin. The
Chromatin remodeling by ATP-dependent molecular machines Alexandra Lusser and James T. Kadonaga* Summary The eukaryotic genome is packaged into a periodic nucleoprotein structure termed chromatin. The
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Regulation of DNA replication-coupled histone gene expression
 /, 2017, Vol. 8, (No. 55), pp: 95005-95022 Regulation of DNA replication-coupled histone gene expression Qianyun Mei 1,*, Junhua Huang 1,*, Wanping Chen 1,2, Jie Tang 1, Chen Xu 1, Qi Yu 1, Ying Cheng
/, 2017, Vol. 8, (No. 55), pp: 95005-95022 Regulation of DNA replication-coupled histone gene expression Qianyun Mei 1,*, Junhua Huang 1,*, Wanping Chen 1,2, Jie Tang 1, Chen Xu 1, Qi Yu 1, Ying Cheng
p53 and Apoptosis: Master Guardian and Executioner Part 2
 p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome
 The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
MCB*4010 Midterm Exam / Winter 2008
 MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Nuclear Architecture and Chromatin Dynamics
 International symposium on Nuclear Architecture and Chromatin Dynamics & 2 nd meeting of the Asian Forum of Chromosome and Chromatin Biology November 26-29, 2008 Centre for Cellular and Molecular Biology,
International symposium on Nuclear Architecture and Chromatin Dynamics & 2 nd meeting of the Asian Forum of Chromosome and Chromatin Biology November 26-29, 2008 Centre for Cellular and Molecular Biology,
Chapter 11 How Genes Are Controlled
 Chapter 11 How Genes Are Controlled PowerPoint Lectures Campbell Biology: Concepts & Connections, Eighth Edition REECE TAYLOR SIMON DICKEY HOGAN Lecture by Edward J. Zalisko Introduction Well-preserved
Chapter 11 How Genes Are Controlled PowerPoint Lectures Campbell Biology: Concepts & Connections, Eighth Edition REECE TAYLOR SIMON DICKEY HOGAN Lecture by Edward J. Zalisko Introduction Well-preserved
