Received 3 January 2005/Accepted 6 January 2005
|
|
|
- Amie Perkins
- 5 years ago
- Views:
Transcription
1 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, July 2005, p Vol. 71, No /05/$ doi: /aem Copyright 2005, American Society for Microbiology. All Rights Reserved. Similarities and Specificities of Fungal Keratinolytic Proteases: Comparison of Keratinases of Paecilomyces marquandii and Doratomyces microsporus to Some Known Proteases Helena Gradišar, 1 * Jožica Friedrich, 1 Igor Križaj, 2 and Roman Jerala 1 Laboratory of Biotechnology, National Institute of Chemistry, Hajdrihova 19, Ljubljana 1000, Slovenia, 1 and Department of Biochemistry and Molecular Biology, Jožef Stefan Institute, Jamova 39, Ljubljana 1000, Slovenia 2 Received 3 January 2005/Accepted 6 January 2005 Based on previous screening for keratinolytic nonpathogenic fungi, Paecilomyces marquandii and Doratomyces microsporus were selected for production of potent keratinases. The enzymes were purified and their main biochemical characteristics were determined (molecular masses, optimal temperature and ph for keratinolytic activity, N-terminal amino acid sequences). Studies of substrate specificity revealed that skin constituents, such as the stratum corneum, and appendages such as nail but not hair, feather, and wool were efficiently hydrolyzed by the P. marquandii keratinase and about 40% less by the D. microsporus keratinase. Hydrolysis of keratin could be increased by the presence of reducing agents. The catalytic properties of the keratinases were studied and compared to those of some known commercial proteases. The profile of the oxidized insulin B-chain digestion revealed that both keratinases, like proteinase K but not subtilisin, trypsin, or elastase, possess broad cleavage specificity with a preference for aromatic and nonpolar amino acid residues at the P-1 position. Kinetic studies were performed on a synthetic substrate, succinyl-ala-ala-pro-phe-p-nitroanilide. The keratinase of P. marquandii exhibited the lowest K m among microbial keratinases reported in the literature, and its catalytic efficiency was high in comparison to that of D. microsporus keratinase and proteinase K. All three keratinolytic enzymes, the keratinases of P. marquandii and D. microsporus as well as proteinase K, were significantly more active on keratin than subtilisin, trypsin, elastase, chymotrypsin, or collagenase. Keratins are the most abundant proteins in epithelial cells of vertebrates and represent the major constituents of skin and its appendages such as nail, hair, feather, and wool. The protein chains are packed tightly either in -chain ( -keratins) or in -sheet ( -keratins) structures. Keratins belong to the superfamily of intermediate filament proteins. Their high degree of cross-linking by disulfide bonds, hydrophobic interactions, and hydrogen bonds stabilizes keratin filament structure (11). Therefore, keratinous material is water insoluble and extremely resistant to degradation by proteolytic enzymes such as trypsin, pepsin, and papain. A group of proteolytic enzymes which are able to hydrolyze insoluble keratins more efficiently than other proteases are called keratinases (29). They are produced by some insects and mostly by microorganisms. The best studied are keratinases from the dermatophytic genera Microsporum (26, 37) and Trichophyton (30, 38) as well as from bacteria of the genera Bacillus (4, 14, 20, 24, 34, 35) and Streptomyces (2, 3, 27). There are relatively few reports on characterization of the keratinases from nondermatophytic fungi (5, 7, 12, 25, 32, 33). Most keratinases have some common characteristics despite their different origins. They belong mainly to the extracellular serine proteases, with the exception of keratinases from yeasts, which belong to the aspartic proteases (23, 28). The molecular masses of the enzymes range from 20 kda to 60 kda. They are mostly active in alkaline environments, with optimal activity at temperatures up to 50 C. Thermostable keratinases with optimal temperatures of around 85 C and a higher molecular mass have been reported (5, 8, 31). The potential use of keratinases is in different applications where keratins should be hydrolyzed, such as the leather and detergent industries, textiles, waste bioconversion, medicine, and cosmetics for drug delivery through nails and degradation of keratinized skin. In our previous studies potent keratinase producers, among them Aspergillus flavus (9) and Doratomyces microsporus (12), were selected out of 300 nonpathogenic fungi which are preferred for biotechnological applications. The keratinases were produced by cultivating strains in optimized media under submerged aerobic conditions. The enzymes were isolated and purified and their main biochemical characteristics were determined. A keratinase produced by the fungus Paecilomyces marquandii promised to be still more active than the previously studied keratinolytic enzymes. In the present report, we investigated its characteristics. We examined in detail the catalytic properties as well as the specificity of the P. marquandii and D. microsporus keratinases for both natural and synthetic substrates. A comparison of the enzyme properties with those of some commercial proteases is presented. Our aim was to elucidate specific characteristics which distinguish keratinases from nonkeratinolytic proteases. * Corresponding author. Mailing address: Laboratory of Biotechnology, National Institute of Chemistry, Hajdrihova 19, Ljubljana 1000, Slovenia. Phone: Fax: helena.gradisar@ki.si. MATERIALS AND METHODS Production and purification of keratinases. Fungal strains Paecilomyces marquandii (MZKI B639) and Doratomyces microsporus (MZKI B399) were used as producers of the keratinolytic enzymes. Both strains have been isolated and 3420
2 VOL. 71, 2005 FUNGAL KERATINOLYTIC PROTEASES 3421 identified in the Laboratory of Botany, Faculty of Medicine and Pharmacy, University of Franche-Comté, Besançon, France, and are deposited in the Culture Collection of the National Institute of Chemistry (MZKI), Ljubljana, Slovenia. The fungi are maintained on potato dextrose agar (Fluka) slants. For keratinase production, strains of P. marquandii and D. microsporus were cultivated in shaken flasks. Each strain was grown in a liquid medium containing soy flour as an enzyme inducer and (g/liter): KH 2 PO 4, 1.5; K 2 HPO 4, 1.0; MgSO 4 7H 2 O, 0.2; CaCl 2 2H 2 O, 0.2; NaCl, 0.2; ZnSO 4 7H 2 O, 0.002; peptone, 0.4; malt extract, 1.0; glucose, 1.0; soy flour, 5.0; and glycerol, 2 ml/liter. The ph of the medium was adjusted to 6, and aliquots of 100 ml were distributed into 500-ml Erlenmayer flasks and autoclaved. After inoculation with a spore suspension (10 6 spores/ml medium), the fermentation was carried out at 30 C and 100 rpm on a rotary shaker until the keratinolytic activity reached its maximum, i.e., for 5 days with P. marquandii and for 4 days with D. microsporus. Crude enzymes were prepared by concentration of the broth filtrate (PLCC 5K membrane, Minitan system, Millipore, Bedford, MA) followed by overnight dialysis against deionized water and subsequent lyophilization. For enzyme purification, the lyophilized powder was dissolved in 50 mm phosphate buffer (ph 7.0) containing 1 M ammonium sulfate and the enzyme solution was applied onto a preparative column for hydrophobic interaction chromatography (HiPrep 16/10 Phenyl FF, Pharmacia), previously equilibrated with the same buffer. The enzyme was eluted by stepwise decreasing salt concentrations from 1 M to 0 M at a flow rate of 3.0 ml/min. The keratinolytic active fraction was pooled and concentrated by ultrafiltration (YM10 membrane, Amicon). Subsequently the enzyme solution was applied on a gel filtration column (Superose 12 HR 10/30, Pharmacia) equilibrated with a 50 mm phosphate buffer (ph 7.5) containing 0.2 M NaCl. The purified enzyme was stored in aliquots at 20 C. Soluble protein determination. The concentration of soluble protein was measured at 562 nm by the bicinchoninic acid assay according to the manufacturer s instructions (Sigma). The protein concentration was determined by using a calibration curve that was established with known concentrations of bovine serum albumin. The purity of the keratinases and their molecular masses were determined by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) according to the method of Laemmli (19). The MiniProtean II system was used according to the manufacturer s instructions (Bio-Rad). The proteins were separated on a 12% gel and stained with a 0.1% (wt/vol) solution of Coomassie brilliant blue R (Sigma). Low-molecular-mass markers (Dalton mark VII-L, Sigma) were used as protein standards for determination of molecular masses. N-terminal amino acid sequencing of keratinases and sequence analysis. The N-terminal sequences of the purified keratinases were determined by automated Edman degradation using a pulsed-liquid sequence analyzer (Procise 492A protein sequencing system, Applied Biosystems). Samples were purified by reversedphase high-performance liquid chromatography (RP-HPLC), dried on a Speed- Vac concentrator (Savant), dissolved in 20% (vol/vol) acetonitrile/water containing 0.001% (vol/vol) trifluoroacetic acid (TFA), and sequenced. The sequences obtained were compared to sequences in the TrEMBL/Swissprot database using BLASTp algorithm ( Determination of keratinase activity. Keratinolytic activity was examined using keratin from the stratum corneum of the human sole as a substrate. The preparation of keratin powder was described previously (12). Briefly, scrapings of human sole were defatted, well rinsed, dried, ground to a powder, and sifted through a 0.2-mm screen. The previously described activity assay (12) was slightly modified and performed as follows: the reaction mixture comprised 4.0 ml of 28 mm Tris-HCl buffer (ph 8.0), 1.0 ml of the enzyme solution, and 20 mg of the keratin powder. Incubation was carried out in a water bath at 45 C for 30 min with constant agitation of 160 rpm. The enzyme reaction was terminated by the addition of 2.0 ml 10% (wt/vol) trichloroacetic acid (TCA) and then allowed to stand at 4 C for 30 min. After centrifugation (10,000 rpm, 15 min) in a cooled centrifuge (Sorvall), the absorbance of the supernatant was measured spectrophotometrically at 280 nm (spectrophotometer from Beckman). The control was treated the same way except that TCA was added before the incubation. One unit of keratinolytic activity was defined as an increase of corrected A 280 for under the conditions described. The data presented are mean values of three parallel determinations. Further, more native keratins (keratin from human nail and hair, porcine nail, chicken feather, sheep wool, and commercial bovine keratin [ICN Biomedicals Inc.]) and nonkeratinous fibrillar proteins (collagen and elastin) as well as bovine serum albumin and casein were used as substrates to examine keratinase activity on different native proteins. The preparation of keratinous materials was described in our previous publication (12). The incubation procedure was the same as described for keratinolytic activity determination. The extent of proteolysis TABLE 1. Effect of protease inhibitors and additives on activity of P. marquandii and D. microsporus keratinases Compound Concn Keratinase of P. marquandii Residual activity (%) Keratinase of D. microsporus Control PMSF 1 mm 0 0 EDTA 5 mm Iodoacetamide 0.05 mm DTT 0.5 mm mm mm ME 5 mm mm SDS 0.1% % DMSO 1% % % Isopropanol 1% was determined spectrophotometrically at 280 nm by measuring of TCA-soluble peptides released from the substrate during the incubation. In addition, the keratinase activity assay was used to compare the activities of selected proteases (Sigma). The enzymes tested were keratinases from P. marquandii and D. microsporus, chymotrypsin, collagenase, elastase, proteinase K, subtilisin, and trypsin. Keratin of the stratum corneum and casein were chosen as model substrates for enzyme activity determination. Stock solutions in concentration of 1 mg/ml were prepared in 28 mm Tris-HCl buffer (ph 8.0) for each enzyme. They were diluted adequately with the same buffer and then the substrate was added. Incubation and determination conditions were the same as described above. Effects of ph, temperature, proteinase inhibitors, organic solvents, detergents, and reducing agents on keratinolytic activity. The effect of ph on the keratinolytic activity of the keratinases was assayed at 45 C using 28 mm buffers of various ph values: citrate buffer (ph 4 to 6), phosphate buffer (ph 6 to 8), Tris-HCl (ph 7 to 9), and glycine-naoh (ph 9 to 11). The optimum temperature for the enzyme activity was determined by performing the enzyme reaction at various temperatures between 20 C and 80 C at ph 8.0 (28 mm Tris-HCl). All other conditions were the same as described in the keratinolytic activity assay. The effects of protease inhibitors (EDTA, phenylmethylsulfonyl fluoride [PMSF], and iodoacetamide) and other chemicals, dithiothreitol (DTT), -mercaptoethanol ( -ME), sodium dodecyl sulfate (SDS), dimethyl sulfoxide (DMSO), and isopropanol, on keratinolytic activity were determined (for concentrations, see Table 1). The purified keratinases were preincubated with each compound in 28 mm Tris-HCl buffer (ph 8.0) at 30 C for 10 min. The control was preincubated without the compound. Keratin of the stratum corneum was then added to each preparation, and the residual keratinolytic activity was measured at 45 C as described above for determination of keratinase activity. Proteolytic specificity of keratinases. The proteolytic specificity was determined by measuring the ability of the keratinases to hydrolyze selected synthetic substrates and the oxidized B-chain of insulin. The synthetic substrates used in this study (all purchased from Bachem) were prepared as stock solutions in DMSO: 200 mm N-succinyl-Ala-Ala-Ala-pNA (AAA), Ac-Tyr-OEt (ATEE), Bz-Arg-pNA HCl (L-BAPA), 100 mm N-succinyl-Ala-Ala-Pro-Phe-pNA (AAPF), FA-Leu-Gly-Pro-Ala-OH (FALGPA), or in isopropanol (100 mm Bz-Tyr-pNA). The enzymatic reactions were carried out in 28 mm Tris-HCl buffer (ph 8.0) at 45 C. The substrate concentration was 1 mm, while the keratinase was added at an amount adequate to be able to follow the initial reaction velocity. The hydrolysis of peptides was monitored spectrophotometrically at 237 nm for ATEE, at 324 nm for FALGPA, and at 405 nm for p-na peptides. Then the best substrate was chosen for kinetic studies of keratinase activity. At least five concentrations of the chosen synthetic peptide were assayed under the same conditions. The final concentration of organic solvent in the reaction mixture never exceeded 5% (vol/vol). The hydrolysis was followed continuously and the initial velocities were determined. The values of Michaelis- Menten constant (K m ), maximal velocity (V max ), and catalytic constant (k cat ) were calculated by nonlinear regression using Michaelis-Menten equation. To determine the hydrolysis sites, each keratinase (0.2 g) was incubated with
3 3422 GRADIŠAR ET AL. APPL. ENVIRON. MICROBIOL. the oxidized insulin B-chain (0.1% [wt/vol]) in 28 mm Tris-HCl buffer (ph 8.0), and the final volume was 100 l. The reaction mixture was incubated at 30 C for 20 min and 12 h. The reaction was stopped by adding 1 volume of 5% (vol/vol) acetonitrile in 0.05% (vol/vol) TFA. An aliquot of 100 l of the sample was applied on RP-HPLC column C18 (Chromospher, HPLC system; Knauer). The peptides were eluted with an increasing gradient of acetonitrile from 5% to 95% (vol/vol) in 0.05% (vol/vol) TFA. The detection was performed at 215 nm. The molecular masses of the peptide products were determined by electrospray ionization mass spectroscopy (ESI-MS). The cleavage sites were defined by using a computer program, FindPept ( which enables peptide identification after protein cleavage on the basis of the experimentally determined size of the products. RESULTS AND DISCUSSION FIG. 1. SDS-PAGE of the purified keratinases of P. marquandii (P.m.) and D. microsporus (D.m.). The positions of low-molecularmass markers (st.) from Sigma are shown. The gel (12%) was stained with Coomassie brilliant blue. Keratinases which are able to degrade extremely resistant keratins are mostly isolated from pathogenic dermatophytes or bacteria. We searched for prospective keratinase producers among the nonpathogenic filamentous fungi. The most outstanding keratinolytic activity was found in keratinases from Paecilomyces marquandii, Doratomyces microsporus, and Aspergillus flavus strains. Due to the reputation of A. flavus as the potential producer of aflatoxins, we have focused our research on the other two strains. Production and purification of keratinases. Fungal strains of D. microsporus and P. marquandii were cultivated in a submerged fermentation. Keratinolytic activity of the culture filtrate appeared after 40 h for both strains and increased until reached its maximum, i.e., 49.5 U/ml after 95 h for D. microsporus culture filtrate and U/ml after 110 h for that of P. marquandii. The culture filtrates were collected and then concentrated, dialyzed, and lyophilized; 0.25 g of the crude keratinase from D. microsporus and 0.4 g of that of P. marquandii were obtained from 1 liter of fermentation broth. The keratinases were purified, as described in the Materials and Methods section, 3.8-fold to a specific activity of 1,005 U/mg and 4.9-fold to a specific activity of 326 U/mg, respectively. By SDS-PAGE, a single protein band was obtained in each case (Fig. 1). Characterization of keratinases. The basic biochemical characteristics of D. microsporus keratinase were described in our previous report (12). In this study, the molecular mass of the keratinase of P. marquandii was estimated to be 33 kda by both SDS-PAGE and gel filtration on Superose 12. The enzyme seemed to be a monomer. The molecular masses of the purified keratinases, 33 kda for P. marquandii, 30 kda for D. microsporus (12), and 22 kda for A. flavus keratinase (not published), fit into the range between 20 kda and 60 kda reported for other keratinases. All three keratinases are serine proteases, as are the majority of keratinases. The keratinolytic activity of the keratinase of P. marquandii on stratum corneum keratin was detected in a broad range of ph values (ph 6 to 11), with the maximum activity at ph 8 (Fig. 2A). The optimal ph of keratinolytic activity for both the P. marquandii and D. microsporus enzymes is around ph 8, while the keratinase of A. flavus has its optimum at ph 11, which is very high among the keratinases reported before. The P. marquandii keratinase is special for its temperature optimum, which was determined to be 60 to 65 C (Fig. 2B). Other reported keratinases, except thermostable ones, were mostly active up to 50 C. Since enzyme assays were performed at 45 C, a slightly high temperature, the enzyme stability of keratinases at that temperature was examined. The enzyme solution of each keratinase (0.06 mg/ml) was incubated at 45 C. Every 5 min the residual activity was determined spectrophotometrically by following hydrolysis of 1 mm AAPF under the conditions described for activity measurement in Materials and Methods. The reaction rate decreased for 50% in 15 min with D. microsporus keratinase and in 150 min with P. marquandii keratinase, so the P. marquandii keratinase was stable 10 times longer than that of D. microsporus. SDS-PAGE showed complete degradation of the enzyme and appearance of smaller peptides due to enzyme autolysis (data not shown). The addition of the inhibitor PMSF prevents autodegradation of keratinases. Since stabilization of the native conformation with calcium ions is commonly observed with extracellular enzymes, the effect of Ca 2 ions on the stability of D. microsporus keratinase was studied in our previous work. It was shown that Ca 2 ions at 1 mm concentration did not significantly influence enzyme stability (15). However, autolysis did not represent a problem when other proteins were present as a substrate. If the stability of the P. marquandii keratinase was compared to the data for keratinases from the literature, only a keratinase from Streptomyces pactum (2) and protease D-1 from Stenotrophomonas sp. (39) proved to be similarly stable, in addition to keratinolytic enzymes of thermophilic microorganisms (5, 31). We have analyzed the effect of protease inhibitors and reducing agents on enzyme activity (Table 1). Both purified keratinases were totally inhibited by the serine protease inhibitor PMSF and partially by EDTA (around 30%), while iodoacetamide did not cause any inhibition. The maximal increase of enzyme activity on stratum corneum keratin was observed by the presence of the reducing agent DTT. The activity was
4 VOL. 71, 2005 FUNGAL KERATINOLYTIC PROTEASES 3423 FIG. 2. ph (A) and temperature (B) optima for the keratinase of P. marquandii. Œ, citrate buffer; F, phosphate buffer; Œ, Tris-HCl;, glycine-naoh buffer. increased twofold for the P. marquandii keratinase and threefold for the D. microsporus keratinase. The reducing agent -ME only slightly increased the activity of the P. marquandii keratinase but significantly increased the D. microsporus keratinase activity. Since DTT and -ME are known to cleave disulfide bridges, an influence either on enzyme or on keratin substrate was possible. Thus, the enzyme activity was measured with 1 mm AAPF, which does not contain disulfide bridges. The activity on the synthetic substrate remained unchanged in the presence of DTT and -ME for both keratinases (data not shown). The result confirmed that the reducing agents acted on the keratin substrate and not on the enzymes. The influence of some additives which were present in some assays was also measured (Table 1). DMSO did not influence the activity of keratinases up to a concentration of 5%, while isopropanol decreased the activity about 13% at a concentration of 1%. SDS inhibited both enzymes at a concentration of just 0.1%. Thirteen-amino-acid-residue sequences of the N termini of the purified keratinases from P. marquandii and D. microsporus were determined, and a sequence similarity search was performed in the SwissProt/TrEMBL database. The comparisons between determined sequences and 13-residue sequences of the N termini of various proteases are presented in Table 2. The sequence of the P. marquandii keratinase and that of the TABLE 2. Alignment of the N-terminal sequences of P. marquandii and D. microsporus keratinases with N-terminal sequences of similar proteases a Enzyme b Sequence Keratinase of P. marquandii...altqqpgapwglg Serine protease of P. lilacinus (Q01471)...AYTQQPGAPWGLG Keratinase of D. microsporus...atvtqnnapwglg Alkaline proteinase of A. chrysogenum (P29118)...ALVTQNGA WGLG Proteinase K (P06873)... AAQTNAPWGLA Subtilisin (SUBSCL)...ALA QTV PYGIP a Underlined amino acid residues are identical to corresponding residues in the P. marquandii sequence. b The SwissProt/TrEMBL data base number is in parentheses. D. microsporus keratinase vary by five amino acid residues. The highest similarity with the sequence of the P. marquandii enzyme, 12 out of 13 residues identical, was found in the sequence of a serine protease from another species of the same genus, Paecilomyces lilacinus, while the sequence most homologous to the D. microsporus enzyme, 10 out of 13 residues identical, was found at the N terminus of a serine protease from Acremonium chrysogenum. Among the proteases from Table 2, only proteinase K from Tritirachium album is described as keratinolytic. Proteolytic specificity of keratinases. The peptide cleavage specificity of the enzymes was tested using synthetic substrates and oxidized insulin B-chain. A variety of synthetic oligopeptides were examined as a substrate for keratinases. The most favored substrate for both enzymes was AAPF; cleavage of the substrate AAA was 100-fold lower than cleavage of AAPF. The keratinases also possessed an esterase activity, as they were able to hydrolyze ATEE. The other substrates, FALGPA, L-BAPA, and Bz-Tyr-pNA, were not hydrolyzed (not shown). This experiment demonstrated that keratinases preferentially cleave the peptide bond at hydrophobic aromatic (AAPF) and aliphatic (AAA) amino acids at the P-1 site of synthetic oligopeptides. Several researchers also found AAPF to be the best substrate for their keratinase (6, 17, 26, 32, 38). Therefore, the tetrapeptide AAPF was chosen to determine kinetic parameters. Besides the two keratinases, proteinase K was also tested for comparison. The initial velocities of hydrolysis for different AAPF concentrations in the range from 0.1 mm to 5.0 mm were determined. Then, kinetic parameters were calculated using the molar absorption coefficient for p-nitroaniline determined under our experimental conditions (ε 8,800 l mol 1 cm 1 ). The results are summarized in Table 3. The highest affinity towards the substrate AAPF was exhibited by the keratinase of P. marquandii. Its catalytic efficiency (k cat /K m ) of 241 mm 1 s 1 was higher than those of the D. microsporus keratinase and of proteinase K, 9 mm 1 s 1 and 95 mm 1 s 1, respectively. Our measurements on AAPF showed that the kinetic constants for the P. marquandii keratinase are comparable to the constants
5 3424 GRADIŠAR ET AL. APPL. ENVIRON. MICROBIOL. TABLE 3. Kinetic parameters for hydrolysis of AAPF with P. marquandii and D. microsporus keratinases and with proteinase K Enzyme K m (mm) V max (10 7 mmol/s) k cat (s 1 ) k cat /K m (mm 1 s 1 ) Keratinase of P. marquandii Keratinase of D. microsporus Proteinase K given in the literature for other microbial keratinases (2, 3, 6, 26, 32). Among them, the Michaelis-Menten constant K m for AAPF was the lowest for the keratinase of P. marquandii. Further, the proteolytic specificity of both keratinases was examined by hydrolysis of the oxidized insulin B-chain. The cleavage sites were compared with the cleavage sites of some other proteases (Fig. 3). The two fungal keratinases exhibited very broad specificity, with preferential selectivity for hydrophobic and aromatic amino acids on the P-1 site for Leu (L), Val (V), Tyr (Y), and Phe (F). The profile of cleavage sites for the keratinase of D. microsporus was more similar to that of proteinase K, while the keratinase of P. marquandii revealed a smaller number of cleavage sites. However, a large number of cleavage sites for the keratinases of P. marquandii and D. microsporus as well as for proteinase K in the oxidized B-chain of insulin distinguished these enzymes from subtilisin and especially from trypsin and elastase. Substrate specificity of keratinases. To investigate the keratinase substrate specificity, we prepared different keratinous substrates from natural human and animal keratinous materials. Skin keratins, called soft keratins, possess up to 10% of cysteine residues. Hard keratins, which are constituents of skin appendages, possess more than 15% of cysteine residues and are more resistant to proteolysis than soft keratins. The activity of the P. marquandii and D. microsporus keratinases on keratins and on other proteins is presented in Fig. 4. Among keratins, the soft keratins of the stratum corneum were preferably hydrolyzed. The keratinases also showed the ability to degrade commercial bovine keratin and human and porcine nail, but to a much lesser degree. Human hair, sheep wool, and chicken feathers were not hydrolyzed in short incubation times of 30 min. With prolonged incubation of up to 24 h, the D. microsporus (10) and P. marquandii keratinases also hydrolyzed hair and wool keratin (data not shown). Thus, FIG. 4. Hydrolysis of different keratinous and nonkeratinous substrates by keratinases of P. marquandii and D. microsporus after 30 min of incubation. The activity of D. microsporus keratinase on stratum corneum was defined as 100%. keratinases are able to hydrolyze -keratins from skin, nail, and hair but not -keratins from chicken feathers. It was shown that the presence of reducing agents stimulated enzyme hydrolysis of keratin. At a 1 mm concentration of DTT, the activity of the D. microsporus keratinase was three times higher and that of the P. marquandii keratinase was two times higher than the activity of the enzymes without DTT addition. The stimulating effect of reducing agents has been reported by many authors (16, 18, 20, 22, 36). They agreed that reducing agents reduced disulfide bonds of keratinous filaments and allowed access of the enzymes to the substrate for proteolytic attack. The amounts of soluble products formed during casein hydrolysis were comparable to the product amount after stratum corneum hydrolysis, whereas the globular protein bovine serum albumin was not hydrolyzed to such a high level. The keratinases of P. marquandii and D. microsporus as well FIG. 3. Cleavage sites on oxidized insulin B-chain for the keratinases of P. marquandii and D. microsporus. For comparison, the cleavage sites for proteinase K (14) as well as subtilisin, trypsin, and elastase (1) are presented. Thick arrow, major cleavage site, 20 min of hydrolysis; thin arrow, minor cleavage site, 12 h of hydrolysis.
6 VOL. 71, 2005 FUNGAL KERATINOLYTIC PROTEASES 3425 FIG. 5. Amount of soluble products during hydrolysis of (A) keratin and (B) casein by keratinases and selected proteases. as keratinases reported by other authors, with one single exception (5), in addition to keratin hydrolyzed the nonkeratinous substrates casein and bovine serum albumin. Interestingly, elastin and collagen are also fibrillar constituents of skin, but only slight activity of the tested keratinases on these substrates was detected. The same observations were reported for the Streptomyces albidoflavus (3) and Stenotrophomonas sp. keratinases (39). However, on the majority of substrates tested, the keratinase of P. marquandii proved to be about twice as active as the D. microsporus keratinase. A quantitative comparison to other keratinases reported in the literature is practically impossible because keratinous substrates were not identical and methods of activity measurements and definitions of keratinolytic units are not unified. Hydrolysis of model substrates with keratinases and other proteases. We have chosen two model substrates (keratin of the stratum corneum and casein) and compared the keratinase activities on these substrates with the activities of some proteases of fungal (proteinase K), bacterial (collagenase and subtilisin), and animal (chymotrypsin, elastase, and trypsin) origin. Proteinase K is a keratinolytic protease with a wellknown broad specificity. Except for the collagenase, which is a metalloprotease, all other enzymes are serine-type proteases, as are the two keratinases studied. The keratinolytic and caseinolytic activities of the proteolytic enzymes are shown in Fig. 5. TABLE 4. Ratio of velocities of keratin to casein hydrolysis as a criterion of enzyme specificity for keratinous substrates Enzyme Velocity (U/mg/min) Keratin Casein Ratio, keratin/casein Proteinase K Keratinase of P. marquandii Keratinase of D. microsporus Elastase Subtilisin Trypsin Chymotrypsin Collagenase As expected, the P. marquandii and D. microsporus keratinases, as well as proteinase K, hydrolyzed keratin more extensively than the other proteases. The keratinase of P. marquandii was about 20% and the keratinase of D. microsporus was about 50% less active on stratum corneum keratin than was proteinase K. However, all the other enzymes tested were only slightly active or not active at all, as was the case with the collagenase. When caseinolytic activity was measured, the keratinases and proteinase K were closely followed by elastase, chymotrypsin, and subtilisin. The ratio between the velocity of keratin hydrolysis and the velocity of casein hydrolysis for each enzyme was a criterion for the enzyme specificity for keratinous substrates (Table 4). The keratinase of P. marquandii and proteinase K revealed the highest specificity, with a ratio of around 0.7, and the specificity of the D. microsporus keratinase was lower, with a ratio of However, with the other enzymes the values were lower than 0.3. The value for trypsin was 0.42 due to the weak hydrolysis of casein and not due to the significant degradation of keratin. A ratio higher than 0.5 for the keratinases and proteinase K confirms that these enzymes degrade keratins more efficiently than other proteases do. Similar results were reported by Cheng et al. (4) for the bacterial keratinase of Bacillus licheniformis. Generally, the keratinases reported in the literature were 5- to 20-fold more active on keratinous substrates than other proteases (2, 3, 21, 39). In summary, the keratinases are enzymes which catalyze the degradation of keratins. All of them are proteases, but not all proteases hydrolyze keratins. Taking into account that in keratins a high percentage of the molecule represents hydrophobic and aromatic amino acids (approximately 50%) (13), it could be concluded that keratinases are successful in hydrolysis of keratinous materials due to the specific amino acid composition of keratins as well as to their broad specificity. ACKNOWLEDGMENTS This work was partially supported by the Ministry of Higher Education, Science and Technology of Slovenia. We thank J. P. Chamount from the Faculty of Medicine and Pharmacy in Besançon, France, for the donation of fungal strains. We also
7 3426 GRADIŠAR ET AL. APPL. ENVIRON. MICROBIOL. thank B. Kralj from the Jožef Stefan Institute in Ljubljana, Slovenia, for ESI-MS analyses. REFERENCES 1. Beynon, R. J., and J. S. Bond Proteolytic enzymes, a practical approach. Oxford University Press, Oxford, United Kingdom. 2. Böckle, B., B. Galunsky, and R. Müller Characterization of a keratinolytic serine proteinase from Streptomyces pactum DSM Appl. Environ. Microbiol. 61: Bressollier, P., F. Letourneau, M. Urdaci, and B. Verneuil Purification and characterization of a keratinolytic serine proteinase from Streptomyces albidoflavus. Appl. Environ. Microbiol. 65: Cheng, S. W., H. M. Hu, S. W. Shen, H. Takagi, M. Asano, and Y. C. Tsai Production and characterization of keratinase of a feather-degrading Bacillus licheniformis PWD-1. Biosci. Biotechnol. Biochem. 59: Dozie, I. N. S., C. N. Okeke, and N. C. Unaeze A thermostable, alkaline-active, keratinolytic proteinase from Chrysosporium keratinophilum. World J. Microbiol. Biotechnol. 10: Evans, K. L., J. Crowder, and E. S. Miller Subtilisins of Bacillus spp. hydrolyze keratin and allow growth on feathers. Can. J. Microbiol. 46: Farag, A. M., and M. A. Hassan Purification, characterization and immobilization of a keratinase from Aspergillus oryzae. Enzyme Microb. Technol. 34: Friedrich, A., and G. Antranikian Keratin degradation by Fervidobacterium pennavorans, a novel thermophilic anaerobic species of the order Thermotogales. Appl. Environ. Microbiol. 62: Friedrich, J., H. Gradišar, D. Mandin, and J. P. Chaumont Screening fungi for synthesis of keratinolytic enzymes. Lett. Appl. Microbiol. 28: Friedrich, J., and S. Kern Hydrolysis of native proteins by keratinolytic protease of Doratomyces microsporus. J. Mol. Catal. B Enzymol. 21: Fuchs, E Keratins and the skin. Annu. Rev. Cell Dev. 11: Gradišar, H., S. Kern, and J. Friedrich Keratinase of Doratomyces microsporus. Appl. Microbiol. Biotechnol. 53: Gregg, K., S. D. Wilson, D. A. D. Parry, and G. E. Rogers A comparison of genomic coding sequences for feather and scale keratins: Structural and evolutionary implications. EMBO J. 3: Han, X. Q., and S. Damodaran Purification and characterization of Protease Q: a detergent- and urea-stable serine endopeptidase from Bacillus pumilus. J. Agric. Food Chem. 46: Hublin, A., H. Gradišar, J. Friedrich, and D. Vasić-Raćki Stability and stabilization of Doratomyces microsporus keratinase. Biocatal. Biotransform. 20: Kunert, J Effect of reducing agents on proteolytic and keratinolytic activity of enzymes of Microsporum gypseum. Mycoses 35: Kunert, J., and E. Kasafirek Preliminary characterization of extracellular proteolytic enzymes of dermatophytes by chromogenic substrates. J. Med. Vet. Mycol. 26: Kunert, J., and Z. Stransky Thiosulfate production from cystine by keratinolytic prokaryote Streptomyces fradiae. Arch. Microbiol. 150: Laemmli, U. K Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: Lee, H., D. B. Suh, J. H. Hwang, and H. J. Suh Characterization of a keratinolytic metalloprotease from Bacillus sp. SCB-3. Appl. Biochem. Biotechnol. 97: Letourneau, F., V. Soussotte, P. Bressollier, P. Branland, and B. Verneuil Keratinolytic activity of Streptomyces sp. S.K1-02: a new isolated strain. Lett. Appl. Microbiol. 26: Lin, X., C. G. Lee, E. S. Casale, and J. C. H. Shih Purification and characterization of a keratinase from a feather-degrading Bacillus licheniformis strain. Appl. Environ. Microbiol. 58: Lin, X., J. Tang, G. Koelsch, M. Monod, and S. Foundling Recombinant Canditropsin, an extracellular aspartic protease from yeast Candida tropicalis. J. Biol. Chem. 268: Lin, X., D. W. Kelemen, E. S. Miller, and J. C. H. Shih Nucleotide sequence and expression of kera, the gene encoding a keratinolytic protease of Bacillus licheniformis PWD-1. Appl. Environ. Microbiol. 61: Malviya, H. K., R. C. Rajak, and S. K. Hasija Purification and partial characterization of two extracellular keratinases of Scopulariopsis brevicaulis. Mycopathologia 119: Mignon, B., M. Swinnen, J. P. Bouchara, M. Hofinger, A. Nikkles, G. Pierard, C. Gerday, and B. Losson Purification and characterization of a 31.5 kda keratinolytic subtilizin-like serine protease from Microsporum canis and evidence of its secretion in naturally infected cats. Med. Mycol. 36: Mukhopadhyay, R. P., and A. L. Chandra Keratinase of a streptomycete. Indian J. Exp. Biol. 28: Negi, M., R. Tsuboi, T. Matsui, and H. Ogawa Isolation and characterization of proteinase from Candida albicans: substrate specificity. J. Investig. Dermatol. 83: Onifade, A. A A review: potentials for biotechnological applications of keratin-degrading microorganisms and their enzymes for nutritional improvement of feathers and other keratins as livestock feed resources. Bioresour. Technol. 66: Qin, L. M., S. Dekio, and J. Jidoi Some biochemical characteristics of a partially purified extracellular keratinase from Trichophyton schoenleinii. Zentralbl. Bakteriol. 277: Riessen, S., and G. Antranikian Isolation of Thermoanaerobacter keratinophilus sp. nov., a novel thermophilic, anaerobic bacterium with keratinolytic activity. Extremophiles 5: Rojanavanich, V., T. Yoshiike, R. Tsuboi, K. Takamori, and H. Ogawa Purification and characterization of an extracellular proteinase from Hendersonula toruloidea. Infect. Immun. 58: Santos, R. M. D. B., A. A. P. Firmino, C.M. de Sá, and C. R. Felix Keratinolytic activity of Aspergillus fumigatus fresenius. Curr. Microbiol. 33: Suh, H. J., and H. K. Lee Characterization of a keratinolytic serine protease from Bacillus subtilis KS-1. J. Protein Chem. 20: Suntornsuk, W., and L. Suntornsuk Feather degradation by Bacillus sp. FK 46 in submerged cultivation. Bioresour. Technol. 86: Takami, H., F. Nakamura, R. Aono, and K. Hirokoshi Degradation of human hair by a thermostable alkaline proteinase from Bacillus sp. no. AH 101. Biosci. Biotechnol. Biochem. 56: Takiuchi, I., Y. Sei, H. Takagi, and M. Negi Partial characterization of the extracellular keratinase from Microsporum canis. Sabouraudia: J. Med. Vet. Mycol. 22: Tsuboi, R., I. J. Ko, K. Takamori, and H. Ogawa Isolation of a keratinolytic proteinase from Trichophyton mentagrophytes with enzymatic activity at acidic ph. Infect. Immun. 57: Yamamura, S., Y. Morita, Q. Hasan, K. Yokoyama, and E. Tamiya Keratin degradation. A cooperative action of two enzymes from Stenotrophomonas sp. Biochem. Biophys. Res. Commun. 294:
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Evaluation of Protein Hydrolysate from Chrome-Tanned Leather Waste in Keratinase Production by Bacillus Subtilis Atcc 6633
 Evaluation of Protein Hydrolysate from Chrome-Tanned Leather Waste in Keratinase Production by Bacillus Subtilis Atcc 6633 Bilge Hilal ÇADIRCI 1 *, Ahmet ASLAN 2, İhsan YAŞA 1, Hüseyin Ata KARAVANA 2,
Evaluation of Protein Hydrolysate from Chrome-Tanned Leather Waste in Keratinase Production by Bacillus Subtilis Atcc 6633 Bilge Hilal ÇADIRCI 1 *, Ahmet ASLAN 2, İhsan YAŞA 1, Hüseyin Ata KARAVANA 2,
Pelagia Research Library
 Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 211, 1 (3):124-129 ISSN: 2248 9215 Production of Alkaline Protease by Bacillus subtilis (MTCC7312) using Submerged
Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 211, 1 (3):124-129 ISSN: 2248 9215 Production of Alkaline Protease by Bacillus subtilis (MTCC7312) using Submerged
Scholars Research Library. Purification and characterization of neutral protease enzyme from Bacillus Subtilis
 Journal of Microbiology and Biotechnology Research Scholars Research Library J. Microbiol. Biotech. Res., 2012, 2 (4):612-618 (http://scholarsresearchlibrary.com/archive.html) Purification and characterization
Journal of Microbiology and Biotechnology Research Scholars Research Library J. Microbiol. Biotech. Res., 2012, 2 (4):612-618 (http://scholarsresearchlibrary.com/archive.html) Purification and characterization
INTERNATIONAL JOURNAL OF ENGINEERING SCIENCES & RESEARCH TECHNOLOGY
 [Ravish, 2(2): Feb., 2013] ISSN: 2277-9655 IJESRT INTERNATIONAL JOURNAL OF ENGINEERING SCIENCES & RESEARCH TECHNOLOGY Isolation And Characterization Of Proteolytic Bacteria And Its Protease Himani Ravish
[Ravish, 2(2): Feb., 2013] ISSN: 2277-9655 IJESRT INTERNATIONAL JOURNAL OF ENGINEERING SCIENCES & RESEARCH TECHNOLOGY Isolation And Characterization Of Proteolytic Bacteria And Its Protease Himani Ravish
Preliminary Characterization of Keratinolytic Enzyme of Aspergillus flavus K-03 and Its Potential in Biodegradation of Keratin Wastes
 Mycobiology 31(4): 209-213 (2003) Copyright 2003 by The Korean Society of Mycology Preliminary Characterization of Keratinolytic Enzyme of Aspergillus flavus K-03 and Its Potential in Biodegradation of
Mycobiology 31(4): 209-213 (2003) Copyright 2003 by The Korean Society of Mycology Preliminary Characterization of Keratinolytic Enzyme of Aspergillus flavus K-03 and Its Potential in Biodegradation of
BIODEGRADATION OF KERATIGENOUS WASTES BY DERMATOPHYTIC FUNGI WITH REFERENCE TO TRICOPHYTON RUBRUM
 BIODEGRADATION OF KERATIGENOUS WASTES BY DERMATOPHYTIC FUNGI WITH REFERENCE TO TRICOPHYTON RUBRUM * R.A. Shah, S. Khan and R. Shaikh The Institute of Science, 15 Madame Cama Road, Mumbai 400 032, India
BIODEGRADATION OF KERATIGENOUS WASTES BY DERMATOPHYTIC FUNGI WITH REFERENCE TO TRICOPHYTON RUBRUM * R.A. Shah, S. Khan and R. Shaikh The Institute of Science, 15 Madame Cama Road, Mumbai 400 032, India
Production and Preliminary Characterization of Alkaline Protease from Aspergillus flavus and Aspergillus terreus
 ISSN: 0973-4945; CODEN ECJHAO E- Chemistry http://www.e-journals.net 2010, 7(2), 479-482 Production and Preliminary Characterization of Alkaline Protease from Aspergillus flavus and Aspergillus terreus
ISSN: 0973-4945; CODEN ECJHAO E- Chemistry http://www.e-journals.net 2010, 7(2), 479-482 Production and Preliminary Characterization of Alkaline Protease from Aspergillus flavus and Aspergillus terreus
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Aspergillus foetidus BY AQUEOUS TWO PHASE
 33 CHAPTER 3 PARTIAL PURIFICATION OF TANNASE FROM Aspergillus foetidus BY AQUEOUS TWO PHASE EXTRACTION AND ITS CHARACTERIZATION 3.1 INTRODUCTION Partial purification of proteins in general and tannase
33 CHAPTER 3 PARTIAL PURIFICATION OF TANNASE FROM Aspergillus foetidus BY AQUEOUS TWO PHASE EXTRACTION AND ITS CHARACTERIZATION 3.1 INTRODUCTION Partial purification of proteins in general and tannase
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Chapter PURIFICATION OF ALKALINE PROTEASES
 Chapter PURIFICATION OF ALKALINE PROTEASES E /xtracellular alkaline proteases produced by Bacillus sp. K 25 and bacillus pumilus K 242, were purified and the homogeneity was examined by electrophoresis.
Chapter PURIFICATION OF ALKALINE PROTEASES E /xtracellular alkaline proteases produced by Bacillus sp. K 25 and bacillus pumilus K 242, were purified and the homogeneity was examined by electrophoresis.
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
SYNOPSIS STUDIES ON THE PREPARATION AND CHARACTERISATION OF PROTEIN HYDROLYSATES FROM GROUNDNUT AND SOYBEAN ISOLATES
 1 SYNOPSIS STUDIES ON THE PREPARATION AND CHARACTERISATION OF PROTEIN HYDROLYSATES FROM GROUNDNUT AND SOYBEAN ISOLATES Proteins are important in food processing and food product development, as they are
1 SYNOPSIS STUDIES ON THE PREPARATION AND CHARACTERISATION OF PROTEIN HYDROLYSATES FROM GROUNDNUT AND SOYBEAN ISOLATES Proteins are important in food processing and food product development, as they are
Production and Optimization of Protease from Aspergillus niger and Bacillus subtilis using Response Surface Methodology
 IOSR Journal of Biotechnology and Biochemistry (IOSR-JBB) ISSN: 2455-264X, Volume 2, Issue 7 (Nov. Dec. 2016), PP 01-07 Production and Optimization of Protease from Aspergillus niger and Bacillus subtilis
IOSR Journal of Biotechnology and Biochemistry (IOSR-JBB) ISSN: 2455-264X, Volume 2, Issue 7 (Nov. Dec. 2016), PP 01-07 Production and Optimization of Protease from Aspergillus niger and Bacillus subtilis
Degradation of Chicken Feathers by Proteus vulgaris And Micrococcus luteus
 ISSN 0976 3333 Available Online at www.ijpba.info International Journal of Pharmaceutical & Biological Archives 2013; 4(2): 366-370 ORIGINAL RESEARCH ARTICLE Degradation of Chicken Feathers by Proteus
ISSN 0976 3333 Available Online at www.ijpba.info International Journal of Pharmaceutical & Biological Archives 2013; 4(2): 366-370 ORIGINAL RESEARCH ARTICLE Degradation of Chicken Feathers by Proteus
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
EXTRACTION OF THERMO-STABLE ALPHA AMYLASE FROM FERMENTED WHEAT BRAN
 BIOLOGIA 2001, 47 (1&2), PP 47 52 ISSN 0006 3096 EXTRACTION OF THERMO-STABLE ALPHA AMYLASE FROM FERMENTED WHEAT BRAN *HAMAD ASHRAF, IKRAM UL HAQ, AND JAVED IQBAL Biotechnology Research Laboratory, Department
BIOLOGIA 2001, 47 (1&2), PP 47 52 ISSN 0006 3096 EXTRACTION OF THERMO-STABLE ALPHA AMYLASE FROM FERMENTED WHEAT BRAN *HAMAD ASHRAF, IKRAM UL HAQ, AND JAVED IQBAL Biotechnology Research Laboratory, Department
MAXIMIZATION OF PRODUCTION OF PROTEIN HYDROLYSATES BY USING IMMOBILIZED PAPAIN
 Int. J. Chem. Sci.: 7(4), 2009, 2624-2632 MAXIMIZATION OF PRODUCTION OF PROTEIN HYDROLYSATES BY USING IMMOBILIZED PAPAIN T. SATHISH a and N. Y. S. MURTHY * Department of Biotechnology, Malla Reddy Engineering
Int. J. Chem. Sci.: 7(4), 2009, 2624-2632 MAXIMIZATION OF PRODUCTION OF PROTEIN HYDROLYSATES BY USING IMMOBILIZED PAPAIN T. SATHISH a and N. Y. S. MURTHY * Department of Biotechnology, Malla Reddy Engineering
Degradation of chicken feathers by Leuconostoc sp. and Pseudomonas microphilus
 Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 2012, 2 (2):358-362 Degradation of chicken feathers by Leuconostoc sp. and Pseudomonas microphilus ISSN: 2248
Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 2012, 2 (2):358-362 Degradation of chicken feathers by Leuconostoc sp. and Pseudomonas microphilus ISSN: 2248
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Keratinase Activity of Some Hyphomycetous Fungi from Dropped Off Chicken Feathers
 ISSN 0976 3333 Available Online at www.ijpba.info International Journal of Pharmaceutical & Biological Archives 2011; 2(6):1745-1750 ORIGINAL RESEARCH ARTICLE Keratinase Activity of Some Hyphomycetous
ISSN 0976 3333 Available Online at www.ijpba.info International Journal of Pharmaceutical & Biological Archives 2011; 2(6):1745-1750 ORIGINAL RESEARCH ARTICLE Keratinase Activity of Some Hyphomycetous
Effect of ph on the production of protease by Fusarium oxysporum using agroindustrial waste
 Biotechnological Communication Biosci. Biotech. Res. Comm. 8(1): 78-83 (2015) Effect of ph on the production of protease by Fusarium oxysporum using agroindustrial waste Rupali R. Deshmukh and N. N. Vidhale*
Biotechnological Communication Biosci. Biotech. Res. Comm. 8(1): 78-83 (2015) Effect of ph on the production of protease by Fusarium oxysporum using agroindustrial waste Rupali R. Deshmukh and N. N. Vidhale*
Purification and Characterization of a Keratinase from a
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, OCt. 1992, p. 3271-3275 0099-2240/92/103271-05$02.00/0 Copyright 1992, American Society for Microbiology Vol. 58, No. 10 Purification and Characterization of a Keratinase
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, OCt. 1992, p. 3271-3275 0099-2240/92/103271-05$02.00/0 Copyright 1992, American Society for Microbiology Vol. 58, No. 10 Purification and Characterization of a Keratinase
PROTAZYME AK TABLETS
 www.megazyme.com ASSAY OF endo-protease using PROTAZYME AK TABLETS T-PRAK 05/16 Megazyme International Ireland 2016 SUBSTRATE: The substrate employed is Azurine-crosslinked casein (AZCL-casein). This substrate
www.megazyme.com ASSAY OF endo-protease using PROTAZYME AK TABLETS T-PRAK 05/16 Megazyme International Ireland 2016 SUBSTRATE: The substrate employed is Azurine-crosslinked casein (AZCL-casein). This substrate
International Journal of Research in Biological Sciences
 Available online at http://www.urpjournals.com International Journal of Research in Biological Sciences Universal Research Publications. All rights reserved ISSN 2249 9687 Research Article Production and
Available online at http://www.urpjournals.com International Journal of Research in Biological Sciences Universal Research Publications. All rights reserved ISSN 2249 9687 Research Article Production and
Communication. Identification of Methionine N -Acetyltransferase from Saccharomyces cerevisiae
 Communication THE JOURNAL OP BIOLOGICAL CHEMISTRY Vol. 265, No. 7, Issue of March 5, pp. 3603-3606,lSSO 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U. S. A. Identification
Communication THE JOURNAL OP BIOLOGICAL CHEMISTRY Vol. 265, No. 7, Issue of March 5, pp. 3603-3606,lSSO 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U. S. A. Identification
Chapter 3. Structure of Enzymes. Enzyme Engineering
 Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Chapter 10. Regulatory Strategy
 Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard. Product Number: AD0014
 TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard Product Number: AD0014 INTRODUCTION: Iodoacetamido-activated
TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard Product Number: AD0014 INTRODUCTION: Iodoacetamido-activated
PRO G max Probiotic fermented soybean meal Benefits of PRO G max
 PRO G max Probiotic fermented soybean meal Benefits of PRO G max Probiotic bacteria > 10 10 CFU/kg High protein with low molecular weight protein approaching small peptides enhancing digestion and absorption
PRO G max Probiotic fermented soybean meal Benefits of PRO G max Probiotic bacteria > 10 10 CFU/kg High protein with low molecular weight protein approaching small peptides enhancing digestion and absorption
Principles of Biotechnology INDUSTRIAL BIOTECHNOLOGY WEEKS 8+9
 Principles of Biotechnology INDUSTRIAL BIOTECHNOLOGY WEEKS 8+9 Industrial Microbiology Industrial Microorganisms and Product formation involved: 1- Use microorganisms to produce valuable commercial product
Principles of Biotechnology INDUSTRIAL BIOTECHNOLOGY WEEKS 8+9 Industrial Microbiology Industrial Microorganisms and Product formation involved: 1- Use microorganisms to produce valuable commercial product
Complete Amino Acid Sequence of the Protease Inhibitor BWI-4a from Buckwheat Seeds
 Biochemistry (Moscow), Vol. 65, No. 10, 2000, pp. 1140-1144. Translated from Biokhimiya, Vol. 65, No. 10, 2000, pp. 1347-1352. Original Russian Text Copyright 2000 by Belozersky, Dunaevsky, Musolyamov,
Biochemistry (Moscow), Vol. 65, No. 10, 2000, pp. 1140-1144. Translated from Biokhimiya, Vol. 65, No. 10, 2000, pp. 1347-1352. Original Russian Text Copyright 2000 by Belozersky, Dunaevsky, Musolyamov,
Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples:
 Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Broad Spectrum Protease Inhibitor Cocktail
 Broad Spectrum Protease Inhibitor Cocktail Catalog number: AR1182 Boster s Broad Spectrum Protease Inhibitor Cocktail is a complex of various protease inhibitors, which has been tested for inhibiting proteases
Broad Spectrum Protease Inhibitor Cocktail Catalog number: AR1182 Boster s Broad Spectrum Protease Inhibitor Cocktail is a complex of various protease inhibitors, which has been tested for inhibiting proteases
Screening of bacteria producing amylase and its immobilization: a selective approach By Debasish Mondal
 Screening of bacteria producing amylase and its immobilization: a selective approach By Debasish Mondal Article Summary (In short - What is your article about Just 2 or 3 lines) Category: Bacillus sp produce
Screening of bacteria producing amylase and its immobilization: a selective approach By Debasish Mondal Article Summary (In short - What is your article about Just 2 or 3 lines) Category: Bacillus sp produce
Journal of Chemical and Pharmaceutical Research, 2017, 9(6): Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2017, 9(6):160-164 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 ABSTRACT Physicochemical Characterization of Purified
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2017, 9(6):160-164 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 ABSTRACT Physicochemical Characterization of Purified
Media Optimization Studies for Enhanced Production of Serratiopeptidase
 Media Optimization Studies for Enhanced Production of Serratiopeptidase from Bacillus Licheniformis (NCIM ) Manasi J. Wagdarikar*, Anagha M. Joshi, Amir A. Shaikh SCES s Indira College of Pharmacy, Tathawade,
Media Optimization Studies for Enhanced Production of Serratiopeptidase from Bacillus Licheniformis (NCIM ) Manasi J. Wagdarikar*, Anagha M. Joshi, Amir A. Shaikh SCES s Indira College of Pharmacy, Tathawade,
Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
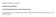 Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Collagenase Assay Kit
 Collagenase Assay Kit Catalog # 31 and 32 For Research Use Only - Not Human or Therapeutic Use INTRODUCTION Collagenases are members of the matrix metalloproteinase (MMP) family and degrade collagen types
Collagenase Assay Kit Catalog # 31 and 32 For Research Use Only - Not Human or Therapeutic Use INTRODUCTION Collagenases are members of the matrix metalloproteinase (MMP) family and degrade collagen types
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Gentilucci, Amino Acids, Peptides, and Proteins. Peptides and proteins are polymers of amino acids linked together by amide bonds CH 3
 Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Suk Hoo Yoon Korea Food Research Institute 1/42
 Development of Phospholipases to Produce Structured Phospholipids Suk Hoo Yoon Korea Food Research Institute 1/42 Phospholipase D H H C O R R Z Fatty acyl chain -H Phosphatidic acid (PA) R O C H O - -CH
Development of Phospholipases to Produce Structured Phospholipids Suk Hoo Yoon Korea Food Research Institute 1/42 Phospholipase D H H C O R R Z Fatty acyl chain -H Phosphatidic acid (PA) R O C H O - -CH
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Collagenase Assay Kit
 Collagenase Assay Kit Catalog # 31 and 32 For Research Use Only - Not Human or Therapeutic Use INTRODUCTION The collagenases are members of the matrix metalloproteinase (MMP) family and degrade collagen
Collagenase Assay Kit Catalog # 31 and 32 For Research Use Only - Not Human or Therapeutic Use INTRODUCTION The collagenases are members of the matrix metalloproteinase (MMP) family and degrade collagen
Midterm 1 Last, First
 Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
1. INTRODUCTION. Exploration of Keratinolytic Actinobacteria for the Bioconversion of Poultry Feather. Mr. SUBHASISH SAHA 1
 1. INTRODUCTION Feather waste is generated in large quantities as a by-product of commercial poultry processing. Feather consists of pure keratin protein. Keratin in its native state is not degradable
1. INTRODUCTION Feather waste is generated in large quantities as a by-product of commercial poultry processing. Feather consists of pure keratin protein. Keratin in its native state is not degradable
Screening of Nutritional Parameters for the Production of Protease from Aspergillus Oryzae
 ISSN: 0973-4945; CODEN ECJHAO E-Journal of Chemistry http://www.e-journals.net Vol. 4, No. 2, pp 208-215, April 2007 Screening of Nutritional Parameters for the Production of Protease from Aspergillus
ISSN: 0973-4945; CODEN ECJHAO E-Journal of Chemistry http://www.e-journals.net Vol. 4, No. 2, pp 208-215, April 2007 Screening of Nutritional Parameters for the Production of Protease from Aspergillus
SUMMARY AND CONCLUSION
 SUMMARY AND CONCLUSION A potential lipase producing marine fungus was selected among 14 lipase producers isolated from seawater and sediments of South Indian coastal environments which was identified as
SUMMARY AND CONCLUSION A potential lipase producing marine fungus was selected among 14 lipase producers isolated from seawater and sediments of South Indian coastal environments which was identified as
Antifungal Properties of Cranberry Juice
 APPLIED MICROBIOLOGY, OCt. 1968, p. 1524-1527 Copyright @ 1968 American Society for Microbiology Vol. 16, No. 10 Printed in U.S.A. Antifungal Properties of Cranberry Juice JACOB H. SWARTZ AND THEODORE
APPLIED MICROBIOLOGY, OCt. 1968, p. 1524-1527 Copyright @ 1968 American Society for Microbiology Vol. 16, No. 10 Printed in U.S.A. Antifungal Properties of Cranberry Juice JACOB H. SWARTZ AND THEODORE
SUPPLEMENTARY INFORMATION. Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was
 SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
About the Kits...2 Description 2 Components 3 Storage 3. Factors That Influence Factor Xa Activity... 4
 Novagen User Protocol TB205 Rev. C 0107 1 of 9 Factor Xa Kits Table of Contents About the Kits...2 Description 2 Components 3 Storage 3 Factors That Influence Factor Xa Activity... 4 Factor Xa Cleavage...5
Novagen User Protocol TB205 Rev. C 0107 1 of 9 Factor Xa Kits Table of Contents About the Kits...2 Description 2 Components 3 Storage 3 Factors That Influence Factor Xa Activity... 4 Factor Xa Cleavage...5
Substrate Specificity and Salt Inhibition of Five Proteinases Isolated from the Pyloric Caeca and Stomach of Sardine
 Agric. Biol. Chem., 46 (6), 1565~1569, 1982 1565 Substrate Specificity and Salt Inhibition of Five Proteinases Isolated from the Pyloric Caeca and Stomach of Sardine Minoru Noda, Thanh Vo Van, Isao Kusakabe
Agric. Biol. Chem., 46 (6), 1565~1569, 1982 1565 Substrate Specificity and Salt Inhibition of Five Proteinases Isolated from the Pyloric Caeca and Stomach of Sardine Minoru Noda, Thanh Vo Van, Isao Kusakabe
GLYCATION OF PROTEINS IN ESCHERICHIA COLI: EFFECT OF NUTRIENT BROTH INGREDIENTS ON GLYCATION
 Industry GLYCATION OF PROTEINS IN ESCHERICHIA COLI: EFFECT OF NUTRIENT BROTH INGREDIENTS ON GLYCATION R. Dimitrova, R. Mironova, I. Ivanov Institute of Molecular biology, Bulgarian Academy of Sciences
Industry GLYCATION OF PROTEINS IN ESCHERICHIA COLI: EFFECT OF NUTRIENT BROTH INGREDIENTS ON GLYCATION R. Dimitrova, R. Mironova, I. Ivanov Institute of Molecular biology, Bulgarian Academy of Sciences
Preliminary studies of cellulase production by Acinetobacter anitratus and Branhamella sp.
 frican Journal of iotechnology Vol. 6 (1), pp. 28-33, 4 January 27 vailable online at http://www.academicjournals.org/j ISSN 1684 5315 27 cademic Journals Full Length Research Paper Preliminary studies
frican Journal of iotechnology Vol. 6 (1), pp. 28-33, 4 January 27 vailable online at http://www.academicjournals.org/j ISSN 1684 5315 27 cademic Journals Full Length Research Paper Preliminary studies
Proteins. Amino acids, structure and function. The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka
 Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
DELFIA Eu-DTPA ITC Chelate & Europium Standard
 AD0026P-3 (en) 1 DELFIA Eu-DTPA ITC Chelate & AD0021 Europium Standard For Research Use Only INTRODUCTION DELFIA Eu-DTPA ITC Chelate is optimized for the europium labelling of proteins and peptides for
AD0026P-3 (en) 1 DELFIA Eu-DTPA ITC Chelate & AD0021 Europium Standard For Research Use Only INTRODUCTION DELFIA Eu-DTPA ITC Chelate is optimized for the europium labelling of proteins and peptides for
CHAPTER 9: CATALYTIC STRATEGIES. Chess vs Enzymes King vs Substrate
 CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
FUTA Journal of Research in Sciences, 2016 (1): 34-45
 FUTA Journal of Research in Sciences, 2016 (1): 34-45 KERATINOLYTIC ACTIVITIES OF Aspergillus flavus and Alternaria tenuissima ASSOCIATED WITH BIODEGRADATION OF SELECTED ANIMAL WASTES B.J* Akinyele, and
FUTA Journal of Research in Sciences, 2016 (1): 34-45 KERATINOLYTIC ACTIVITIES OF Aspergillus flavus and Alternaria tenuissima ASSOCIATED WITH BIODEGRADATION OF SELECTED ANIMAL WASTES B.J* Akinyele, and
LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade
 AD0017P-4 (en) 1 LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade INTRODUCTION Fluorescent isothiocyanato-activated (ITC-activated) Eu-W1024 chelate is optimized for labelling proteins
AD0017P-4 (en) 1 LANCE Eu-W1024 ITC Chelate & Europium Standard AD0013 Development grade INTRODUCTION Fluorescent isothiocyanato-activated (ITC-activated) Eu-W1024 chelate is optimized for labelling proteins
Caution: For Laboratory Use. A product for research purposes only. Eu-W1024 ITC Chelate & Europium Standard. Product Number: AD0013
 TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1024 ITC Chelate & Europium Standard Product Number: AD0013 INTRODUCTION: Fluorescent isothiocyanato-activated
TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1024 ITC Chelate & Europium Standard Product Number: AD0013 INTRODUCTION: Fluorescent isothiocyanato-activated
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
National Standard of the People s Republic of China. National food safety standard. Determination of pantothenic acid in foods for infants and
 National Standard of the People s Republic of China GB 5413.17 2010 National food safety standard Determination of pantothenic acid in foods for infants and young children, milk and milk products Issued
National Standard of the People s Republic of China GB 5413.17 2010 National food safety standard Determination of pantothenic acid in foods for infants and young children, milk and milk products Issued
Manja Henze, Dorothee Merker and Lothar Elling. 1. Characteristics of the Recombinant β-glycosidase from Pyrococcus
 S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
UV Tracer TM Maleimide NHS ester
 UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Extraction and Characterization of Protease From Senesced Leaves of Papaya (Carica Papaya) and It s Application
 International Journal of Genetic Engineering and Biotechnology. ISSN 0974 3073 Volume 5, Number 1 (2014), pp. 29-34 International Research Publication House http://www.irphouse.com Extraction and Characterization
International Journal of Genetic Engineering and Biotechnology. ISSN 0974 3073 Volume 5, Number 1 (2014), pp. 29-34 International Research Publication House http://www.irphouse.com Extraction and Characterization
PROTEINS. Building blocks, structure and function. Aim: You will have a clear picture of protein construction and their general properties
 PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
Operating Instructions
 Poroszyme Immobilized Trypsin Cartridge Operating Instructions Your Poroszyme Cartridge is Unique! Applied Biosystems Poroszyme Immobilized Trypsin cartridges perform online tryptic digests in a flow-through
Poroszyme Immobilized Trypsin Cartridge Operating Instructions Your Poroszyme Cartridge is Unique! Applied Biosystems Poroszyme Immobilized Trypsin cartridges perform online tryptic digests in a flow-through
Production of Thermostable and Ca +2 Independent α-amylases from Halphilic Bacteria
 International Journal of Biotechnology and Biochemistry ISSN 0973-2691 Volume 12, Number 2 (2016) pp. 153-159 Research India Publications http://www.ripublication.com Production of Thermostable and Ca
International Journal of Biotechnology and Biochemistry ISSN 0973-2691 Volume 12, Number 2 (2016) pp. 153-159 Research India Publications http://www.ripublication.com Production of Thermostable and Ca
Acid Protease Production by Fungi Used in Soy3bean Food
 APPLIED MICROBIOLOGY, May 1974, p. 906-911 Copyright 1974 American Society for Microbiology Vol. 27, No. 5 Printed in U.S.A. Acid Protease Production by Fungi Used in Soy3bean Food Fermentation HWA L.
APPLIED MICROBIOLOGY, May 1974, p. 906-911 Copyright 1974 American Society for Microbiology Vol. 27, No. 5 Printed in U.S.A. Acid Protease Production by Fungi Used in Soy3bean Food Fermentation HWA L.
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
The Structure and Func.on of Macromolecules Proteins GRU1L6
 The Structure and Func.on of Macromolecules Proteins GRU1L6 Proteins Proteins Most structurally & functionally diverse group Function: involved in almost everything enzymes (pepsin, DNA polymerase) structure
The Structure and Func.on of Macromolecules Proteins GRU1L6 Proteins Proteins Most structurally & functionally diverse group Function: involved in almost everything enzymes (pepsin, DNA polymerase) structure
Purification and Characterization of Amidase from Paracoccus
 Purification and Characterization of Amidase from Paracoccus sp. SKG: Utilization of amidase inhibited whole cells for bioconversion of acrylonitrile to acrylamide By Prof. T. B. Karegoudar Department
Purification and Characterization of Amidase from Paracoccus sp. SKG: Utilization of amidase inhibited whole cells for bioconversion of acrylonitrile to acrylamide By Prof. T. B. Karegoudar Department
Purification and Characterization of Iso-Ribonucleases from a Novel Thermophilic Fungus
 Int. J. Mol. Sci. 2014, 15, 944-957; doi:10.3390/ijms15010944 Article OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Purification and Characterization
Int. J. Mol. Sci. 2014, 15, 944-957; doi:10.3390/ijms15010944 Article OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Purification and Characterization
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/1/e1500678/dc1 Supplementary Materials for Chemical synthesis of erythropoietin glycoforms for insights into the relationship between glycosylation pattern and
advances.sciencemag.org/cgi/content/full/2/1/e1500678/dc1 Supplementary Materials for Chemical synthesis of erythropoietin glycoforms for insights into the relationship between glycosylation pattern and
DELFIA Tb-N1 DTA Chelate & Terbium Standard
 AD0029P-1 (en) 1 DELFIA Tb-N1 DTA Chelate & AD0012 Terbium Standard For Research Use Only INTRODUCTION DELFIA Tb-N1 DTA Chelate is optimized for the terbium labeling of proteins and peptides for use in
AD0029P-1 (en) 1 DELFIA Tb-N1 DTA Chelate & AD0012 Terbium Standard For Research Use Only INTRODUCTION DELFIA Tb-N1 DTA Chelate is optimized for the terbium labeling of proteins and peptides for use in
SensoLyte Generic MMP Assay Kit *Colorimetric*
 SensoLyte Generic MMP Assay Kit *Colorimetric* Revision#1.2 Catalog # Kit Size Last updated: May2017 AS-72095 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect MMP activity
SensoLyte Generic MMP Assay Kit *Colorimetric* Revision#1.2 Catalog # Kit Size Last updated: May2017 AS-72095 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect MMP activity
Extracting DNA from cheek cells: a classroom experiment for Year 7 upwards
 Extracting DNA from cheek cells: a classroom experiment for Year 7 upwards Dr Kathryn Scott Research Administrator, Zitzmann Group, Department of Biochemistry Lecturer in Biochemistry, Christ Church Extracting
Extracting DNA from cheek cells: a classroom experiment for Year 7 upwards Dr Kathryn Scott Research Administrator, Zitzmann Group, Department of Biochemistry Lecturer in Biochemistry, Christ Church Extracting
Studies on Glucose Isomerase from a Streptomyces Species
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Oct. 1976, P. 489-493 Copyright ) 1976 American Society for Microbiology Vol. 32, No. 4 Printed in U.S.A. Studies on Glucose Isomerase from a Streptomyces Species
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Oct. 1976, P. 489-493 Copyright ) 1976 American Society for Microbiology Vol. 32, No. 4 Printed in U.S.A. Studies on Glucose Isomerase from a Streptomyces Species
DELFIA Tb-DTPA ITC Chelate & Terbium Standard
 AD0035P-2 (en) 1 DELFIA Tb-DTPA ITC Chelate & AD0029 Terbium Standard For Research Use Only INTRODUCTION DELFIA Tb-DTPA ITC Chelate is optimized for the terbium labelling of proteins and peptides for use
AD0035P-2 (en) 1 DELFIA Tb-DTPA ITC Chelate & AD0029 Terbium Standard For Research Use Only INTRODUCTION DELFIA Tb-DTPA ITC Chelate is optimized for the terbium labelling of proteins and peptides for use
Communication MULTIPLE FORMS OF ACID PHOSPHATASE PRODUCED BY ASPERGJLL US OR YZAE YONEKICHI SAKURAI AND HIDEO SHIOTA
 Short Communication J. Gen. Appl. Microbiol., 16, 335-339 (1970) MULTIPLE FORMS OF ACID PHOSPHATASE PRODUCED BY ASPERGJLL US OR YZAE YONEKICHI SAKURAI AND HIDEO SHIOTA Faculty of Agriculture, Iwate University,
Short Communication J. Gen. Appl. Microbiol., 16, 335-339 (1970) MULTIPLE FORMS OF ACID PHOSPHATASE PRODUCED BY ASPERGJLL US OR YZAE YONEKICHI SAKURAI AND HIDEO SHIOTA Faculty of Agriculture, Iwate University,
Name: Chem 351 Exam 1
 Use the pk a values below for all problems requiring pk a s: α-amino groups = 9.4 α-carboxyl groups = 2.3 Sidechain ionizable groups: Cysteine = 8.5 Tyrosine = 10 Lysine = 10.5 Arginine = 12 Histidine
Use the pk a values below for all problems requiring pk a s: α-amino groups = 9.4 α-carboxyl groups = 2.3 Sidechain ionizable groups: Cysteine = 8.5 Tyrosine = 10 Lysine = 10.5 Arginine = 12 Histidine
Protein Classification based upon Biological functions
 PROTEINS (a) The light produced by fireflies is the result of a reaction involving the protein luciferin and ATP, catalyzed by the enzyme luciferase. (b) Erythrocytes contain large amounts of the oxygen-transporting
PROTEINS (a) The light produced by fireflies is the result of a reaction involving the protein luciferin and ATP, catalyzed by the enzyme luciferase. (b) Erythrocytes contain large amounts of the oxygen-transporting
6. C-type cytochrome, soluble and membrane protein
 185 6. C-type cytochrome, soluble and membrane protein analysis of Rhodobacter sp SW2 and Rhodopseudomonas palustris TIE-1 ABSTRACT The ability to grown on Fe(II) is thought to be a primitive metabolism
185 6. C-type cytochrome, soluble and membrane protein analysis of Rhodobacter sp SW2 and Rhodopseudomonas palustris TIE-1 ABSTRACT The ability to grown on Fe(II) is thought to be a primitive metabolism
EFFECT OF SOME AMINO ACIDS ON THE GROWTH AND L-GLUTAMIC ACID FERMENTATION BY AN AUXOTROPHIC MUTANT Micrococcus glutamicus AB 100.
 S. Ganguly et. al. / International Journal on Pharmaceutical and Biomedical Research (IJPBR) Vol. 2(1), 2011, 21-25 EFFECT OF SOME AMINO ACIDS ON THE GROWTH AND L-GLUTAMIC ACID FERMENTATION BY AN AUXOTROPHIC
S. Ganguly et. al. / International Journal on Pharmaceutical and Biomedical Research (IJPBR) Vol. 2(1), 2011, 21-25 EFFECT OF SOME AMINO ACIDS ON THE GROWTH AND L-GLUTAMIC ACID FERMENTATION BY AN AUXOTROPHIC
Tannase Production By Aspergillus niger
 ISSN: 0973-4945; CODEN ECJHAO E- Chemistry http://www.e-journals.net Vol. 4, No. 2, pp 192-198, April 2007 Tannase Production By Aspergillus niger N. LOKESWARI* and K. JAYA RAJU Center for Biotechnology,
ISSN: 0973-4945; CODEN ECJHAO E- Chemistry http://www.e-journals.net Vol. 4, No. 2, pp 192-198, April 2007 Tannase Production By Aspergillus niger N. LOKESWARI* and K. JAYA RAJU Center for Biotechnology,
Trypsin Digestion Mix
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
Electronic Supplementary Information. Table of Contents
 Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S]
![v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S] v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S]](/thumbs/81/84404059.jpg) Exam 3 Spring 2017 Dr. Stone 8:00 Name There are 100 possible points on this exam. -ΔG / RT v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S] h rate forward = k forward [reactants]
Exam 3 Spring 2017 Dr. Stone 8:00 Name There are 100 possible points on this exam. -ΔG / RT v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S] h rate forward = k forward [reactants]
Enzymes for Flavour Development in Dairy Substrates. Presented by: Blanca Camarasa Senior Business Manager
 Enzymes for Flavour Development in Dairy Substrates Presented by: Blanca Camarasa Senior Business Manager Cheese composition Cheese consists of proteins and fat from milk, usually the milk of cows, goat,
Enzymes for Flavour Development in Dairy Substrates Presented by: Blanca Camarasa Senior Business Manager Cheese composition Cheese consists of proteins and fat from milk, usually the milk of cows, goat,
Statistical Optimization of Keratinase Production from Marine Fungus
 RESEARCH ARTICLE OPEN ACCESS Statistical Optimization of Keratinase Production from Marine Fungus S.Satya lakshmi*, G.Girija Shankar, T.Prabhakar, T.Satish, Pharmaceutical Biotechnology Division, College
RESEARCH ARTICLE OPEN ACCESS Statistical Optimization of Keratinase Production from Marine Fungus S.Satya lakshmi*, G.Girija Shankar, T.Prabhakar, T.Satish, Pharmaceutical Biotechnology Division, College
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/ MCQs. Location : 102, 105, 106, 301, 302
 FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
Enzyme activity Page 1 of 8
 Enzyme activity Page 1 of 8 Contains: - protease - L-glutaminase - Leucine aminopeptidase - sucrase - amylase - cellulase - lipase - catalase - carboxypeptidase Protease Assay Based on: Frankena J., van
Enzyme activity Page 1 of 8 Contains: - protease - L-glutaminase - Leucine aminopeptidase - sucrase - amylase - cellulase - lipase - catalase - carboxypeptidase Protease Assay Based on: Frankena J., van
Int.J.Curr.Microbiol.App.Sci (2015) 4(9):
 ISSN: 2319-7706 Volume 4 Number 9 (2015) pp. 535-548 http://www.ijcmas.com Original Research Article Screening and Selection of Fungus for Keratinase Production by Solid State Fermentation and Optimization
ISSN: 2319-7706 Volume 4 Number 9 (2015) pp. 535-548 http://www.ijcmas.com Original Research Article Screening and Selection of Fungus for Keratinase Production by Solid State Fermentation and Optimization
CHAPTER 3 Amino Acids, Peptides, Proteins
 CHAPTER 3 Amino Acids, Peptides, Proteins Learning goals: Structure and naming of amino acids Structure and properties of peptides Ionization behavior of amino acids and peptides Methods to characterize
CHAPTER 3 Amino Acids, Peptides, Proteins Learning goals: Structure and naming of amino acids Structure and properties of peptides Ionization behavior of amino acids and peptides Methods to characterize
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
N-Glycosidase F Deglycosylation Kit
 For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
