SDS-Assisted Protein Transport Through Solid-State Nanopores
|
|
|
- Frank Simmons
- 5 years ago
- Views:
Transcription
1 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department of Bionanoscience, Kavli Institute of Nanoscience, Delft University of Technology, van der Maasweg 9, 2629 HZ Delft, The Netherlands 2 Department of Physics, University of Illinois at Urbana; Champaign, Urbana, Illinois 61801, United States Table of Contents Table S1: Summary of MD simulations of nanopore systems Figure S1. MD simulation of folded titin translocation... 3 Figure S2. Simulated ionic current blockades produced by translocation of molecular species through a 6 nm diameter nanopore... 5 Figure S3. Simulated ionic current blockades in a 10 nm-diameter nanopore... 6 Note S1. Estimation of the critical micelle concentration of SDS using dynamic light scattering (DLS)... 7 Figure S4. DLS measurements of micelle hydrodynamic radius at different SDS concentrations Note S2. Tryptophan fluorescence bulk assay... 8 Figure S5. Tryptophan fluorescence bulk measurements of β amylase at different SDS concentrations... 8 Note S3. Characterization of SDS micelles in nanopores... 9 Figure S6. Translocation of SDS micelles... 9 Figure S7. Translocation of native and SDS-unfolded titin Figure S8. Simulated ionic current blockades produced by a stretched peptide chain Captions to animations of MD trajectories
2 Table S1: Summary of MD simulations of nanopore systems. Nanopore and membrane dimensions System 1: Membrane cross section: 15 nm x 15 nm Membrane thickness: 6 nm Inner pore diameter: 6 nm Outer pore diameter: 7.5 nm System 2: Membrane cross section: 15 nm x 15 nm Membrane thickness: 10 nm Inner pore diameter: 10 nm Outer pore diameter: 13 nm Biomolecules SDS micelle (70 SDS molecules) Number of atoms 322, 601 Folded titin, conf , 898 Folded titin, conf ,624 SDS/titin, conf. 1 (76 SDS molecules) SDS/titin, conf. 2 (76 SDS molecules) SDS/titin dimer, conf. 1 (104 SDS molecules) SDS/titin dimer, conf. 2 (104 SDS molecules) SDS/β-amylase, conf. 1 SDS/β-amylase, conf. 2 SDS/β-amylase, conf. 3 SDS micelle (70 SDS molecules) 340, , , ,979 Transmembrane bias (mv) Simulation time (ns) / / , , , , Folded titin 291, Folded β-amylase 290, SDS/β-amylase, conf. 2 SDS/β-amylase, orient. 1 SDS/β-amylase, orient. 2 SDS/β-amylase, orient , , , ,
3 Figure S1. MD simulation of folded titin translocation a His-tag SsrA-tag +500 mv Z -500 mv b c d of CoM (nm) Z coord. (na) I SsrA-tag (--RSAANDE NYALAAA) membrane boundaries Simulation time (ns) Open pore, +500 mv Open pore, -500 mv I(nA) Simulation time (ns) His-tag (--MMRGSHHH HHHGLVPRGS) Open pore, +500 mv Simulation time (ns) waters of No Simulation time (ns) 3
4 Figure S1. MD simulation of folded titin translocation. (a) Molecular structure of the folded titin monomer. To reproduce the constructs used in experiment, SsrA and His tags were added to the C and N terminals of the protein, respectively. The blue and red arrows define the initial orientation of the protein with respect to the z-axis of our setup (shown in black), which is also the direction of a positive transmembrane bias. (b-d) MD simulation of folded titin translocation through a 6 nm diameter nanopore (System 1). (b) The z coordinate of the titin s center of mass as a function of simulation time. The horizontal black lines indicate the location of the membrane. The color and the style (solid/dashed) of the lines indicate the initial orientation of the protein and the direction of the transmembrane bias, respectively, defined in panels a. In one simulation (gray), the bias was switched during the MD simulation. Each translocation traces shows a 1-ns running average of 4.8 ps sampled data. (c) The nanopore ionic current recorded in the simulations of folded titin translocation. Each ionic current trace shows a 5 ns running average of 4.8 ps sampled instantaneous current. The inset on the right shows a zoomed-in view of the ionic current trace for the fragment of the MD trajectory where protein translocation was observed (black rectangle in the main plot). In the inset, the ionic current trace shows 1 ns running average of 4.8 ps sampled instantaneous currents. (d) The total number of water molecules that passed through the nanopore from the beginning of the applied bias simulation. In this panel, positive values indicate net water flux in the direction of the z axis. The direction of the water flux is correlated with the sign of the transmembrane bias, indicating an electro-osmotic flow produced by a negatively charged nanopore surface. 4
5 Figure S2. Simulated ionic current blockades produced by translocation of molecular species through a 6 nm diameter nanopore Figure S2. Simulated ionic current blockades produced by translocation of molecular species through a 6 nm diameter nanopore. Left column illustrates typical microscopic conformations observed during nanopore translocation simulations; the black arrow indicated the direction of the positive transmembrane bias. The conformation of a protein is depicted as a trace of the protein backbone; SDS molecules are shown as molecular bonds; water and ions are not shown for clarity. Right columns shows the ionic current traces recorded from the translocation simulations and the open pore current levels. Because of the periodic boundary conditions employed in our MD simulations, individual ionic current traces feature multiple translocation events. Ionic current traces from independent simulations are delineated by the // mark. Black circles and horizontal black bars indicate the average blockade current of individual blockade events and the duration of each event, respectively. The blockade events were defined by the reduction of the nanopore current below 75% of the open pore value. 5
6 Figure S3. Simulated ionic current blockades in a 10 nm-diameter nanopore a SDS micelle Folded titin Folded β-amylase β-amylase / SDS Z 10 nm 500 mv b β-amylase / SDS, orientation 1 β-amylase / SDS, orientation 2 β-amylase / SDS, orientation 3 t = 0 ns t = 47 ns t = 0 ns t = 48 ns t = 0 ns t = 44 ns c ΔG* (ns) SDS micelle Folded titin Folded β-amylase β-amylase / SDS β-amylase / SDS orientation 1 β-amylase / SDS orientation 2 β-amylase / SDS orientation 3 Figure S3. Simulated ionic current blockades in a 10 nm-diameter nanopore. (a) Molecular configurations used for the simulation of the ionic current blockades. The center of mass of each molecule or molecular assembly was restrained to the center of the nanopore. Water and ions are not shown for clarify. (b) Additional simulations of -amylase/sds blockade currents. The initial conformations of the -amylase/sds assemblies were takes from the translocation simulations performed using the 6 nmdiameter nanopore (System 1). In each 48-ns simulation, the center of mass of the - amylase/sds assembly was restrained to the center of the nanopore. (c) The average conductance blockade amplitudes. To enable direct comparison with experiment, the conductance blockades computed from MD simulations were multiplied by the ratio of the experimental and simulation bulk conductivities of 0.4 KCl (3.6/4.8). The average blockade conductance amplitudes were each computed from a 48 ns MD trajectory. 6
7 Note S1. Estimation of the critical micelle concentration of SDS using dynamic light scattering (DLS) It is well known that SDS critical micelle concentration is dependent on the ionic strength of the solution. The formation of micelles depends on the interplay between the attractive forces driven by hydrophobic interactions of the hydrocarbon tails and the repulsive electrostatic interaction between the charged heads. At high ionic concentrations, charges are screened and the critical micelle concentration is reduced. To determine the CMC of SDS in the buffer used for our experiments, we used DLS to determine at which concentrations we notice the presence and absence of SDS micelles. DLS measurements were performed using a Malvern Instuments Zetasizer Nano ZS. Samples containing different SDS concentration were prepared in a buffer containing 0.4M NaCl, 10mM TRIS and 1mM ETDA ph 7.5. The hydrodynamic radius of each sample was measured and is presented in Figure S4. For SDS concentrations higher than 0.05% micelles were observed with the expected hydrodynamic radius. No micelles were observed bellow 0.01% SDS. These results indicate a CMC for SDS in the range of 0.01% %. Figure S4. DLS measurements of micelle hydrodynamic radius at different SDS concentrations. Figure S4. DLS measurements of micelle hydrodynamic radius at different SDS concentrations. The presence of micelles above 0.05% SDS and their absence below 0.01% SDS marks the region where the CMC is located. 7
8 Note S2. Tryptophan fluorescence bulk assay Tryptophan fluorescence is routinely used to study protein conformational changes. Tryptophan fluorescence emission is highly dependent on the nature of its local environment, which can be used to infer the conformation of a protein. In a native or folded protein, most tryptophan residues are either partially or fully buried in the core of the protein due to their hydrophobic nature. Once the protein is unfolded with denaturing agents such as SDS or urea, tryptophan residues become partially or fully exposed and their fluorescent emission shifts. Here we used this principle to study the effect of SDS at different concentrations on one of our protein substrates. All our measurements were done in a solution containing 0.4 M NaCl, 10mM Tris ph 7.5, 1mM ETDA, 1mM of DTT and different concentrations of SDS as shown in the following figure. Fluorescent measurements were done in a Cary eclipse fluorescence spectrophotometer with an excitation wavelength of 295nm and emission collected from 300nm 450nm. The buffer background was measured and subtracted from each protein measurement. Figure S5. Tryptophan fluorescence bulk measurements of β amylase at different SDS concentrations Figure S5. Tryptophan fluorescence bulk measurements of β amylase at different SDS concentrations. A clear conformational change is observed in the protein even at SDS concentrations bellow CMC. 8
9 Note S3. Characterization of SDS micelles in nanopores We performed control experiments to study the translocation of SDS micelles through solid-state nanopores in the absence of proteins. The experimental set up and a typical current trace are shown in Figure S6a and Figure S6b respectively. The scatter plot in Figure S6c shows the amplitude vs. dwell time for both SDS micelles alone and SDS-treated proteins. The conductance blockade histogram is presented in Figure S6d. Figure S6. Translocation of SDS micelles Figure S6. Translocation of SDS micelles (a) Schematic representation of the nanopore control experiment. (b) Example of a current trace measured showing the translocation of SDS micelles. (c) Dwell time vs. conductance blockade scatter plot for SDS-micelles and SDS-treated protein. (d) Conductance blockade histogram of the translocation of micelles. 9
10 Figure S7. Translocation of native and SDS-unfolded titin Figure S7. Translocation of native and SDS-unfolded titin. (a) Scatter plots of dwell time vs. conductance blockade for titin, SDS-titin and SDS micelles. (b) Histograms of the amplidute of the blockade for native titin (left), SDS-titin at an SDS concentration above CMC (middle) and SDS-titin bellow CMC (right). 10
11 Figure S8. Simulated ionic current blockades produced by a stretched peptide chain. Figure S8. Simulated ionic current blockades produced by a stretched peptide chain. (a) Molecular configurations featuring a 60 amino acid fragment of β-amylase (residues 10 to 70) with (Systems P1 and P3) and without (Systems P2 and P4) attached SDS molecules threaded through either the 6 nm diameter nanopore (Systems P1 and P2) or the 10 nm diameter nanopore (Systems P3 and P4). The initial configuration of the peptide was taken from the simulation of the β-amylase/sds assembly, orientation 3 at t = 0 ns (see Figure S3). The center of mass of the peptide chain/sds assembly (Systems P1 and P3) or only of the peptide chain (Systems P2 and P4) was aligned with the geometrical center of each nanopore. During MD simulations of the ionic current, the backbone atoms of the peptide chain were harmonically restrained to their initial coordinates; SDS molecules were not restrained. At the end of the 60 ns simulations of Systems P1 and P3 under a 500 mv transmembrane bias, 20 and 1 SDS molecules detached from the peptide chain, respectively. (b) The average conductance blockade amplitudes. To enable direct comparison with experiment, the conductance blockades computed from MD simulations were multiplied by the ratio of the experimental and simulation bulk conductivities of 0.4 KCl (3.6/4.8). The average blockade conductance amplitudes for each system were computed from a 60 ns MD trajectory. 11
12 Captions to animations of MD trajectories. Movie 1. Self-assembly of a titin-sds complex. The movie illustrates a 300 ns explicit solvent all-atom MD simulation. The protein backbone is shown in blue, SDS molecules in cyan and red, water and 0.4 M NaCl are now shown. A smoothing filter was applied to visually reduce thermal motion of the molecules. The initial configuration was prepared by combining an unfolded conformation of a protein with 120 randomly distributed SDS molecules. The systems were simulated at 373 K for the first 200 ns. Following that, the systems were cooled down to 300 K in eight simulation runs, decreasing the system s temperature by 10 K between each run. Following that, the SDS-protein systems were simulated for another 50 ns at 300 K. Movie 2. MD simulation of folded titin translocation. The movie illustrates a 60 ns explicit solvent all-atom MD simulation carried out under a 500 mv bias directed from the bottom to the top of the simulation system. The nanopore is shown as a cutaway green molecular surface, the protein (red) is shown in a cartoon representation, water and 0.4 M NaCl are now shown. During the simulation, the center of mass of the protein was restrained to remain at the symmetry axis of the nanopore. Movie 3. MD simulation of SDS-titin complex translocation. The movie illustrates a 100 ns explicit solvent all-atom MD simulation carried out under a 250 mv bias directed from the bottom to the top of the simulation system. The nanopore is shown as a cutaway green molecular surface; the protein conformation is depicted as a trace of the protein backbone (magenta). The SDS molecules are shown using the molecular bonds representation: Carbon, sulfur and oxygen atoms are shown in cyan, yellow and red, respectively, hydrogen atoms are not shown. Water and 0.4 M NaCl are now shown as well. During the simulation, the center of mass of the SDS-protein complex was restrained to remain at the symmetry axis of the nanopore. A smoothing filter was applied to visually reduce thermal motion of the molecules. Because of the periodic boundary conditions employed in our MD simulations, the same SDS-protein complex is seen to permeation through the nanopore multiple times. Flickering of the molecules is produced by crossings between different periodic images of the simulation unit cell. Movie 4. MD simulation of SDS-titin dimer complex translocation. The movie illustrates a 50 ns explicit solvent all-atom MD simulation carried out under a 250 mv bias directed from the bottom to the top of the simulation system. The nanopore is shown as a cutaway green molecular surface; the protein conformation is depicted as a trace of the protein backbone (light blue). The SDS molecules are shown using the molecular bonds representation: Carbon, sulfur and oxygen atoms are shown in cyan, yellow and red, respectively, hydrogen atoms are not shown. Water and 0.4 M NaCl are now shown as well. During the simulation, the center of mass of the SDS-protein complex was restrained to remain at the symmetry axis of the nanopore. A smoothing filter was applied to visually reduce thermal motion of the molecules. Because of the periodic boundary conditions employed in our MD simulations, the same SDS-protein complex is seen to permeation through the nanopore multiple times. Flickering of the molecules is produced by crossings between different periodic images of the simulation unit cell. Movie 5. MD simulation of SDS-beta amylase complex translocation. The movie illustrates a 150 ns explicit solvent all-atom MD simulation carried out under a 500 mv 12
13 bias directed from the bottom to the top of the simulation system. The nanopore is shown as a cutaway green molecular surface; the protein conformation is depicted as a trace of the protein backbone (magenta). The SDS molecules are shown using the molecular bonds representation: Carbon, sulfur and oxygen atoms are shown in cyan, yellow and red, respectively, hydrogen atoms are not shown. Water and 0.4 M NaCl are now shown as well. During the simulation, the center of mass of the SDS-protein complex was restrained to remain at the symmetry axis of the nanopore. A smoothing filter was applied to visually reduce thermal motion of the molecules. Because of the periodic boundary conditions employed in our MD simulations, the same SDS-protein complex is seen to permeation through the nanopore multiple times. Flickering of the molecules is produced by crossings between different periodic images of the simulation unit cell. Movie 6. MD simulation of SDS micelle translocation. The movie illustrates a 85 ns explicit solvent all-atom MD simulation carried out under a 250 mv bias directed from the bottom to the top of the simulation system. The nanopore is shown as a cutaway green molecular surface. The SDS molecules are shown using the molecular bonds representation: Carbon, sulfur and oxygen atoms are shown in cyan, yellow and red, respectively, hydrogen atoms are not shown. Water and 0.4 M NaCl are now shown as well. During the simulation, the center of mass of the SDS micelle was restrained to remain at the symmetry axis of the nanopore. A smoothing filter was applied to visually reduce thermal motion of the molecules. Because of the periodic boundary conditions employed in our MD simulations, the micelle is seen to permeation through the nanopore multiple times. Flickering of the molecules is produced by crossings between different periodic images of the simulation unit cell. 13
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Biology 2E- Zimmer Protein structure- amino acid kit
 Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Nature Biotechnology: doi: /nbt.3828
 Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supporting Information. Electrophoretic Deformation of Individual Transfer. RNA Molecules Reveals Their Identity
 Supporting Information Electrophoretic Deformation of Individual Transfer RNA Molecules Reveals Their Identity Robert Y. Henley a, Brian Alan Ashcroft f, Ian Farrell c, Barry S. Cooperman d, Stuart M.
Supporting Information Electrophoretic Deformation of Individual Transfer RNA Molecules Reveals Their Identity Robert Y. Henley a, Brian Alan Ashcroft f, Ian Farrell c, Barry S. Cooperman d, Stuart M.
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Lecture 10 More about proteins
 Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
SAM Teacher s Guide Four Levels of Protein Structure
 SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation
 Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Proteins. (b) Protein Structure and Conformational Change
 Proteins (b) Protein Structure and Conformational Change Protein Structure and Conformational Change Proteins contain the elements carbon (C), hydrogen (H), oxygen (O2) and nitrogen (N2) Some may also
Proteins (b) Protein Structure and Conformational Change Protein Structure and Conformational Change Proteins contain the elements carbon (C), hydrogen (H), oxygen (O2) and nitrogen (N2) Some may also
HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total. The ph at which a peptide has no net charge is its isoelectric point.
 BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
Paper No. 01. Paper Title: Food Chemistry. Module-16: Protein Structure & Denaturation
 Paper No. 01 Paper Title: Food Chemistry Module-16: Protein Structure & Denaturation The order of amino acids in a protein molecule is genetically determined. This primary sequence of amino acids must
Paper No. 01 Paper Title: Food Chemistry Module-16: Protein Structure & Denaturation The order of amino acids in a protein molecule is genetically determined. This primary sequence of amino acids must
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
Life Sciences 1a. Practice Problems 4
 Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Guided Inquiry Skills Lab. Additional Lab 1 Making Models of Macromolecules. Problem. Introduction. Skills Focus. Materials.
 Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Supplementary Figure 1. Overview of steps in the construction of photosynthetic protocellular systems
 Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
BIOPHYSICS II. By Prof. Xiang Yang Liu Department of Physics,
 BIOPHYSICS II By Prof. Xiang Yang Liu Department of Physics, NUS 1 Hydrogen bond and the stability of macromolecular structure Membrane Model Amphiphilic molecule self-assembly at the surface and din the
BIOPHYSICS II By Prof. Xiang Yang Liu Department of Physics, NUS 1 Hydrogen bond and the stability of macromolecular structure Membrane Model Amphiphilic molecule self-assembly at the surface and din the
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Term Definition Example Amino Acids
 Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
MBB 694:407, 115:511. Please use BLOCK CAPITAL letters like this --- A, B, C, D, E. Not lowercase!
 MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
Biological Membranes. Lipid Membranes. Bilayer Permeability. Common Features of Biological Membranes. A highly selective permeability barrier
 Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
The main biological functions of the many varied types of lipids include: energy storage protection insulation regulation of physiological processes
 Big Idea In the biological sciences, a dehydration synthesis (condensation reaction) is typically defined as a chemical reaction that involves the loss of water from the reacting molecules. This reaction
Big Idea In the biological sciences, a dehydration synthesis (condensation reaction) is typically defined as a chemical reaction that involves the loss of water from the reacting molecules. This reaction
Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids
 Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
H 2 O. Liquid, solid, and vapor coexist in the same environment
 Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Activities for the α-helix / β-sheet Construction Kit
 Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Ch5: Macromolecules. Proteins
 Ch5: Macromolecules Proteins Essential Knowledge 4.A.1 The subcomponents of biological molecules and their sequence determine the properties of that molecule A. Structure and function of polymers are derived
Ch5: Macromolecules Proteins Essential Knowledge 4.A.1 The subcomponents of biological molecules and their sequence determine the properties of that molecule A. Structure and function of polymers are derived
Understand how protein is formed by amino acids
 Identify between fibrous and globular proteins Understand how protein is formed by amino acids Describe the structure of proteins using specific examples Functions of proteins Fibrous proteins Globular
Identify between fibrous and globular proteins Understand how protein is formed by amino acids Describe the structure of proteins using specific examples Functions of proteins Fibrous proteins Globular
Bio 12 Chapter 2 Test Review
 Bio 12 Chapter 2 Test Review 1.Know the difference between ionic and covalent bonds In order to complete outer shells in electrons bonds can be Ionic; one atom donates or receives electrons Covalent; atoms
Bio 12 Chapter 2 Test Review 1.Know the difference between ionic and covalent bonds In order to complete outer shells in electrons bonds can be Ionic; one atom donates or receives electrons Covalent; atoms
Q1: Circle the best correct answer: (15 marks)
 Q1: Circle the best correct answer: (15 marks) 1. Which one of the following incorrectly pairs an amino acid with a valid chemical characteristic a. Glycine, is chiral b. Tyrosine and tryptophan; at neutral
Q1: Circle the best correct answer: (15 marks) 1. Which one of the following incorrectly pairs an amino acid with a valid chemical characteristic a. Glycine, is chiral b. Tyrosine and tryptophan; at neutral
Microtubule Teardrop Patterns
 Supporting Information Microtubule Teardrop Patterns Kosuke Okeyoshi 1, Ryuzo Kawamura 1, Ryo Yoshida 2, and Yoshihito Osada 1 * 1 RIKEN Advanced Science Institute, 2-1 Hirosawa, Wako-shi, Saitama 351-0198,
Supporting Information Microtubule Teardrop Patterns Kosuke Okeyoshi 1, Ryuzo Kawamura 1, Ryo Yoshida 2, and Yoshihito Osada 1 * 1 RIKEN Advanced Science Institute, 2-1 Hirosawa, Wako-shi, Saitama 351-0198,
BIOCHEMISTRY 460 FIRST HOUR EXAMINATION FORM A (yellow) ANSWER KEY February 11, 2008
 WRITE YOUR AND I.D. NUMBER LEGIBLY ON EVERY PAGE PAGES WILL BE SEPARATED FOR GRADING! CHECK TO BE SURE YOU HAVE 6 PAGES, (print): ANSWERS INCLUDING COVER PAGE. I swear/affirm that I have neither given
WRITE YOUR AND I.D. NUMBER LEGIBLY ON EVERY PAGE PAGES WILL BE SEPARATED FOR GRADING! CHECK TO BE SURE YOU HAVE 6 PAGES, (print): ANSWERS INCLUDING COVER PAGE. I swear/affirm that I have neither given
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
PHAR3316 Pharmacy biochemistry Exam #2 Fall 2010 KEY
 1. How many protons is(are) lost when the amino acid Asparagine is titrated from its fully protonated state to a fully deprotonated state? A. 0 B. 1 * C. 2 D. 3 E. none Correct Answer: C (this question
1. How many protons is(are) lost when the amino acid Asparagine is titrated from its fully protonated state to a fully deprotonated state? A. 0 B. 1 * C. 2 D. 3 E. none Correct Answer: C (this question
The building blocks of life.
 The building blocks of life. The 4 Major Organic Biomolecules The large molecules (biomolecules OR polymers) are formed when smaller building blocks (monomers) bond covalently. via anabolism Small molecules
The building blocks of life. The 4 Major Organic Biomolecules The large molecules (biomolecules OR polymers) are formed when smaller building blocks (monomers) bond covalently. via anabolism Small molecules
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Unit 2 Biomolecules NGSS
 Unit 2 Biomolecules NGSS Content Area: Science Course(s): Biology CP, Biology Honors, STEM Biology Honors Time Period: October Length: Approximate Blocks TBD Status: Published Transfer Skills Biological
Unit 2 Biomolecules NGSS Content Area: Science Course(s): Biology CP, Biology Honors, STEM Biology Honors Time Period: October Length: Approximate Blocks TBD Status: Published Transfer Skills Biological
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
SRTUCTURE OF PROTEINS DR. A. TARAB DEPT. OF BIOCHEMISTRY HKMU
 SRTUCTURE OF PROTEINS DR. A. TARAB DEPT. OF BIOCHEMISTRY HKMU I. OVERVIEW The twenty amino acids commonly found in proteins are joined together by peptide bonds The linear sequence of the linked amino
SRTUCTURE OF PROTEINS DR. A. TARAB DEPT. OF BIOCHEMISTRY HKMU I. OVERVIEW The twenty amino acids commonly found in proteins are joined together by peptide bonds The linear sequence of the linked amino
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
OCR (A) Biology A-level
 OCR (A) Biology A-level Topic 2.2: Biological molecules Notes Water Water is a very important molecule which is a major component of cells, for instance: Water is a polar molecule due to uneven distribution
OCR (A) Biology A-level Topic 2.2: Biological molecules Notes Water Water is a very important molecule which is a major component of cells, for instance: Water is a polar molecule due to uneven distribution
Gold nanocrystals at DPPC bilayer. Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad
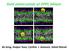 Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects.
 Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects. Manuel J. Louwerse and Evert Jan Baerends Theoretical Chemistry, Vrije Universiteit Amsterdam, De Boelelaan
Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects. Manuel J. Louwerse and Evert Jan Baerends Theoretical Chemistry, Vrije Universiteit Amsterdam, De Boelelaan
Membrane Structure and Membrane Transport of Small Molecules. Assist. Prof. Pinar Tulay Faculty of Medicine
 Membrane Structure and Membrane Transport of Small Molecules Assist. Prof. Pinar Tulay Faculty of Medicine Introduction Cell membranes define compartments of different compositions. Membranes are composed
Membrane Structure and Membrane Transport of Small Molecules Assist. Prof. Pinar Tulay Faculty of Medicine Introduction Cell membranes define compartments of different compositions. Membranes are composed
Qualitative test of protein-lab2
 1- Qualitative chemical reactions of amino acid protein functional groups: Certain functional groups in proteins can react to produce characteristically colored products. The color intensity of the product
1- Qualitative chemical reactions of amino acid protein functional groups: Certain functional groups in proteins can react to produce characteristically colored products. The color intensity of the product
CLASS SET. Modeling Life s Important Compounds. AP Biology
 Modeling Life s Important Compounds AP Biology CLASS SET OBJECTIVES: Upon completion of this activity, you will be able to: Explain the connection between the sequence and the subcomponents of a biological
Modeling Life s Important Compounds AP Biology CLASS SET OBJECTIVES: Upon completion of this activity, you will be able to: Explain the connection between the sequence and the subcomponents of a biological
Elements & Macromolecules in Organisms
 Name: Period: Date: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
Name: Period: Date: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Save My Exams! The Home of Revision For more awesome GCSE and A level resources, visit us at Lipids.
 Lipids Mark Scheme Level Subject Exam Board Topic Sub-Topic Booklet International A Level Biology Edexcel Molecules, Transport and Health Lipids Mark Scheme Time Allowed: 63 minutes Score: /52 Percentage:
Lipids Mark Scheme Level Subject Exam Board Topic Sub-Topic Booklet International A Level Biology Edexcel Molecules, Transport and Health Lipids Mark Scheme Time Allowed: 63 minutes Score: /52 Percentage:
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
The building blocks of life.
 The building blocks of life. All the functions of the cell are based on chemical reactions. the building blocks of organisms BIOMOLECULE MONOMER POLYMER carbohydrate monosaccharide polysaccharide lipid
The building blocks of life. All the functions of the cell are based on chemical reactions. the building blocks of organisms BIOMOLECULE MONOMER POLYMER carbohydrate monosaccharide polysaccharide lipid
Supplementary Fig. 1.
 Supplementary Fig. 1. (a,b,e,f) SEM and (c,d,g,h) TEM images of (a-d) TiO 2 mesocrystals and (e-h) NiO mesocrystals. The scale bars in the panel c, d, g, and h are 500, 2, 50, and 5 nm, respectively. SAED
Supplementary Fig. 1. (a,b,e,f) SEM and (c,d,g,h) TEM images of (a-d) TiO 2 mesocrystals and (e-h) NiO mesocrystals. The scale bars in the panel c, d, g, and h are 500, 2, 50, and 5 nm, respectively. SAED
Derived copy of Proteins *
 OpenStax-CNX module: m65486 1 Derived copy of Proteins * Martha Smith-Caldas Christopher Herren Based on Proteins by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
OpenStax-CNX module: m65486 1 Derived copy of Proteins * Martha Smith-Caldas Christopher Herren Based on Proteins by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
The Interaction between Lipid Bilayers and Biological Membranes. Chapter 18
 The Interaction between Lipid Bilayers and Biological Membranes Chapter 18 Introduction Membrane & Phospholipid Bilayer Structure Membrane Lipid bilayer 2 Introduction Forces Acting between Surfaces in
The Interaction between Lipid Bilayers and Biological Membranes Chapter 18 Introduction Membrane & Phospholipid Bilayer Structure Membrane Lipid bilayer 2 Introduction Forces Acting between Surfaces in
Molecular modeling of the ion channel-like nanotube structure of amyloid β-peptide
 Chinese Science Bulletin 2007 SCIENCE IN CHINA PRESS Springer Molecular modeling of the ion channel-like nanotube structure of amyloid β-peptide JIAO Yong & YANG Pin Key Laboratory of Chemical Biology
Chinese Science Bulletin 2007 SCIENCE IN CHINA PRESS Springer Molecular modeling of the ion channel-like nanotube structure of amyloid β-peptide JIAO Yong & YANG Pin Key Laboratory of Chemical Biology
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Biology Chapter 2 Review
 Biology Chapter 2 Review Vocabulary: Define the following words on a separate piece of paper. Element Compound Ion Ionic Bond Covalent Bond Molecule Hydrogen Bon Cohesion Adhesion Solution Solute Solvent
Biology Chapter 2 Review Vocabulary: Define the following words on a separate piece of paper. Element Compound Ion Ionic Bond Covalent Bond Molecule Hydrogen Bon Cohesion Adhesion Solution Solute Solvent
Proteins and their structure
 Proteins and their structure Proteins are the most abundant biological macromolecules, occurring in all cells and all parts of cells. Proteins also occur in great variety; thousands of different kinds,
Proteins and their structure Proteins are the most abundant biological macromolecules, occurring in all cells and all parts of cells. Proteins also occur in great variety; thousands of different kinds,
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Surfactants. The Basic Theory. Surfactants (or surface active agents ): are organic compounds with at least one lyophilic. Paints and Adhesives
 Surfactants Surfactants (or surface active agents ): are organic compounds with at least one lyophilic ( solvent-loving ) group and one lyophobic ( solvent-fearing ) group in the molecule. In the simplest
Surfactants Surfactants (or surface active agents ): are organic compounds with at least one lyophilic ( solvent-loving ) group and one lyophobic ( solvent-fearing ) group in the molecule. In the simplest
and hydrophilic and how they relate to solubility.
 o o o and hydrophilic and how they relate to solubility. o o o o o o o o Page 1: Introduction Page 2: 1. Hydrocarbons are referred to as organic molecules with a "backbone." Take a snapshot of the hydrocarbon
o o o and hydrophilic and how they relate to solubility. o o o o o o o o Page 1: Introduction Page 2: 1. Hydrocarbons are referred to as organic molecules with a "backbone." Take a snapshot of the hydrocarbon
Structure of proteins
 Structure of proteins Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Structure of proteins The 20 a.a commonly found
Structure of proteins Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Structure of proteins The 20 a.a commonly found
Transport through biological membranes. Christine Carrington Biochemistry Unit Apr 2010
 Transport through biological membranes Christine Carrington Biochemistry Unit Apr 2010 Biological membranes Membranes control the structures and environments of the compartments they define and thereby
Transport through biological membranes Christine Carrington Biochemistry Unit Apr 2010 Biological membranes Membranes control the structures and environments of the compartments they define and thereby
Amino Acids - Building Blocks of Proteins
 Amino Acids - Building Blocks of Proteins Introduction Proteins are more than an important part of your diet. Proteins are complex molecular machines that are involved in nearly all of your cellular functions.
Amino Acids - Building Blocks of Proteins Introduction Proteins are more than an important part of your diet. Proteins are complex molecular machines that are involved in nearly all of your cellular functions.
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Minicircle topoisomer generation. a, Generation of supercoiled topoisomers. Minicircles were nicked with the sequence-specific nicking endonuclease, Nb.BbvCI.
Supplementary Figures Supplementary Figure 1 Minicircle topoisomer generation. a, Generation of supercoiled topoisomers. Minicircles were nicked with the sequence-specific nicking endonuclease, Nb.BbvCI.
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Translation Activity Guide
 Translation Activity Guide Student Handout β-globin Translation Translation occurs in the cytoplasm of the cell and is defined as the synthesis of a protein (polypeptide) using information encoded in an
Translation Activity Guide Student Handout β-globin Translation Translation occurs in the cytoplasm of the cell and is defined as the synthesis of a protein (polypeptide) using information encoded in an
Biomolecule Stations
 AP Biology Biomolecule Stations Names Per. In this two-day activity, you will move through several different stations and learn about the four macromolecules in the biological world. Day 1: Modeling Carbohydrates
AP Biology Biomolecule Stations Names Per. In this two-day activity, you will move through several different stations and learn about the four macromolecules in the biological world. Day 1: Modeling Carbohydrates
Protein Structure and Function
 Protein Structure and Function Protein Structure Classification of Proteins Based on Components Simple proteins - Proteins containing only polypeptides Conjugated proteins - Proteins containing nonpolypeptide
Protein Structure and Function Protein Structure Classification of Proteins Based on Components Simple proteins - Proteins containing only polypeptides Conjugated proteins - Proteins containing nonpolypeptide
Biological Molecules B Lipids, Proteins and Enzymes. Triglycerides. Glycerol
 Glycerol www.biologymicro.wordpress.com Biological Molecules B Lipids, Proteins and Enzymes Lipids - Lipids are fats/oils and are present in all cells- they have different properties for different functions
Glycerol www.biologymicro.wordpress.com Biological Molecules B Lipids, Proteins and Enzymes Lipids - Lipids are fats/oils and are present in all cells- they have different properties for different functions
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Biological Molecules
 The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
The Chemical Building Blocks of Life. Chapter 3
 The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
2.2 Properties of Water
 2.2 Properties of Water I. Water s unique properties allow life to exist on Earth. A. Life depends on hydrogen bonds in water. B. Water is a polar molecule. 1. Polar molecules have slightly charged regions
2.2 Properties of Water I. Water s unique properties allow life to exist on Earth. A. Life depends on hydrogen bonds in water. B. Water is a polar molecule. 1. Polar molecules have slightly charged regions
Review of Biochemistry
 Review of Biochemistry Chemical bond Functional Groups Amino Acid Protein Structure and Function Proteins are polymers of amino acids. Each amino acids in a protein contains a amino group, - NH 2,
Review of Biochemistry Chemical bond Functional Groups Amino Acid Protein Structure and Function Proteins are polymers of amino acids. Each amino acids in a protein contains a amino group, - NH 2,
2.1.1 Biological Molecules
 2.1.1 Biological Molecules Relevant Past Paper Questions Paper Question Specification point(s) tested 2013 January 4 parts c and d p r 2013 January 6 except part c j k m n o 2012 June 1 part ci d e f g
2.1.1 Biological Molecules Relevant Past Paper Questions Paper Question Specification point(s) tested 2013 January 4 parts c and d p r 2013 January 6 except part c j k m n o 2012 June 1 part ci d e f g
Cellular Neurophysiology I Membranes and Ion Channels
 Cellular Neurophysiology I Membranes and Ion Channels Reading: BCP Chapter 3 www.bioelectriclab All living cells maintain an electrical potential (voltage) across their membranes (V m ). Resting Potential
Cellular Neurophysiology I Membranes and Ion Channels Reading: BCP Chapter 3 www.bioelectriclab All living cells maintain an electrical potential (voltage) across their membranes (V m ). Resting Potential
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector
 kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
