Protein Correlation Profiles Identify Lipid Droplet Proteins with High Confidence* S
|
|
|
- Moses Jones
- 6 years ago
- Views:
Transcription
1 Protein Correlation Profiles Identify Lipid Droplet Proteins with High Confidence* S Research 213 by The American Society for Biochemistry and Molecular Biology, Inc. This paper is available on line at Natalie Krahmer, Maximiliane Hilger, Nora Kory, Florian Wilfling, Gabriele Stoehr, Matthias Mann, Robert V. Farese, Jr. **, and Tobias C. Walther Lipid droplets (LDs) are important organelles in energy metabolism and lipid storage. Their cores are composed of neutral lipids that form a hydrophobic phase and are surrounded by a phospholipid monolayer that harbors specific proteins. Most well-established LD proteins perform important functions, particularly in cellular lipid metabolism. Morphological studies show LDs in close proximity to and interacting with membrane-bound cellular organelles, including the endoplasmic reticulum, mitochondria, peroxisomes, and endosomes. Because of these close associations, it is difficult to purify LDs to homogeneity. Consequently, the confident identification of bona fide LD proteins via proteomics has been challenging. Here, we report a methodology for LD protein identification based on mass spectrometry and protein correlation profiles. Using LD purification and quantitative, high-resolution mass spectrometry, we identified LD proteins by correlating their purification profiles to those of known LD proteins. Application of the protein correlation profile strategy to LDs isolated from Drosophila S2 cells led to the identification of 111 LD proteins in a cellular LD fraction in which 1481 proteins were detected. LD localization was confirmed in a subset of identified proteins via microscopy of the expressed proteins, thereby validating the approach. Among the identified LD proteins were both well-characterized LD proteins and proteins not previously known to be localized to LDs. Our method provides a high-confidence LD proteome of Drosophila cells and a novel approach that can be applied to identify LD proteins of other cell types and tissues. Molecular & Cellular Proteomics 12: 1.174/mcp.M , , 213. One of the most pressing issues in eukaryotic cell biology is the need to determine the dynamic composition of organelles that allow for the coordinated execution of many different From the Department of Cell Biology, Yale University School of Medicine, New Haven, Connecticut 652; Signal Transduction and Proteomics, Max Planck Institute of Biochemistry, D Martinsried, Germany; Gladstone Institute of Cardiovascular Disease, San Francisco, California 94158; **Department of Medicine, Department of Biochemistry and Biophysics, University of California, San Francisco, California Received May 4, 212, and in revised form, January 9, 213 Published, MCP Papers in Press, January 12, 213, DOI 1.174/ mcp.m simultaneous processes. Such knowledge of organelle composition provides the basis for understanding protein functions and their coordination in different organelles. Therefore, comprehensive and highly accurate inventories of organelle proteins are essential. Rapid advances in mass spectrometry (MS)-based 1 proteomics are revolutionizing cell biology (1 3). Subcellular fractionation combined with MS enables the systematic determination of organelle composition. Because the sensitivity of MS has increased, even less abundant proteins of an organelle can now be identified. However, the increased sensitivity also identifies more, and less abundant, contaminant proteins, resulting in longer proteome lists for an experiment and complicating the identification of genuine proteins of an organelle. This analytic problem is particularly apparent for lipid droplets (LDs). LD architecture differs from that of other organelles in that they have hydrophobic cores of neutral lipids, primarily triacylglycerol and sterol esters, rather than an aqueous lumen. The LD core is shielded from the cytosol by a phospholipid monolayer, into which specific proteins are embedded (4 6). The primary function of LDs is to store lipids as reservoirs of metabolic energy and membrane precursors. Additionally, LDs participate in host pathogen interactions (7) and play central roles in numerous physiological processes and linked pathologies. For example, obesity is characterized by the overaccumulation of LDs in tissues, and LD accumulation in macrophages is a hallmark of atherosclerosis (8, 9). Given the diverse distribution of LDs in cell types and processes, their protein composition likely varies among cell types and is dynamic. Knowledge of LD compositions in different cell types will lay the foundation for functional characterization of this organelle and could help to reveal novel mechanisms contributing to metabolic diseases. LD protein composition has been analyzed in over a dozen studies that employed different methodologies to study different cell types ranging in evolutionary complexity from yeast to human (1 21). These studies have provided long lists of putative LD proteins, but apart from the consistent identification of several LD marker proteins (e.g., perilipins), LD pro- 1 The abbreviations used are: ER, endoplasmic reticulum; LD, lipid droplet; MS, mass spectroscopy; PCP, protein correlation profile; SILAC, stable isotope labeling of amino acids in cell culture. Molecular & Cellular Proteomics
2 teomes have been plagued by the identification of contaminants that co-purify with LDs. This is due in part to the tight interactions of LDs with other organelles. Microscopy analyses revealed intimate connections between LDs and the endoplasmic reticulum (ER) and mitochondria (12, 22 26), and LDs can form contacts with peroxisomes and different types of endosomes (27, 28). Not surprisingly, therefore, ER and mitochondrial proteins are found abundantly in reported LD proteomes. To overcome these limitations of LD protein identification, we designed and applied a quantitative high-resolution MSbased proteomics strategy that employs protein correlation profiles (PCPs) to confidently identify LD proteins in Drosophila cells. PCPs provide a quantitative approach for determining the correlation profiles of purified proteins relative to known organelle markers (29). PCP studies have yielded some of the most reliable inventories of organelles (3 33), but such an approach has not previously been applied to LDs. EXPERIMENTAL PROCEDURES Drosophila S2 Cells Drosophila S2 cells were cultured in Schneider s Drosophila medium (Invitrogen) supplemented with 1% fetal bovine serum and antibiotics (1 unit/ml penicillin and 1 g/ml streptomycin) at 27 C as described elsewhere (34). For protein localization in S2 cells, expression vectors (actin promoter) were cloned with the Gateway system (Invitrogen) for the following proteins: Alg14-mCherry (CG638), Alg1-mCherry (CG1812), Alg2-mCherry (CG1291), Alg11-mCherry (CG1136), Alg1-mCherry (CG3276), mcherry-seipin (CG994), selenoprotein SELT-mCherry (CG3887), MBOAT-mCherry (CG5926), Lunapark-mCherry (CG8735), mcherry-steroid binding protein (CG966), mcherry-sec14 homolog (CG13848), lyso-pa acyltransferase (CG32699), mcherry-short chain dehydrogenase (CG264), lipase-mcherry (CG17292), mcherry-phospholipase D (CG7718), oxysterol binding protein-mcherry (CG1513), Tango 14-mCherry (CG4775), mcherry-fas associated factor 2 (CG1372), fatty acid transferase-mcherry (CG74), CGI-58-mCherry (CG1882), and mcherry-hsl (CG1155). Transfection of S2 cells was performed using Effectene reagent (Qiagen) according to manufacturer s instructions. Transfected cells were stained with 1 g/ml BODIPY493/53 (Invitrogen) for 1 min and imaged with a spinning-disk confocal microscope (TiLL imic CSU22, Andor) with a back-illuminated electron-multiplying charge-coupled device camera (ixonem 897, Andor) and a NA oil immersion objective (Olympus); 16-bit images were collected with Image iq (version 1.9, Andor), deconvoluted (Huygens, SVI), and cropped (ImageJ). LD Purification For LD purification, ten 75-cm 2 cell culture dishes of Drospohila S2 cells were used. Stable isotope labeling of amino acids in cell culture (SILAC) of the S2 cells was performed as previously described, and the labeling efficiency was 98% before the last passage (35). LD formation was induced by incubating confluent cells with 1 mm oleate complexed to BSA (36) overnight. LD formation was confirmed via microscopy. Cells were harvested, washed twice with ice-cold PBS, resuspended in 2 ml of buffer (2 mm Tris/HCL, ph 7.5, 2 mm magnesium diactetate, and Complete Protease Inhibitor (Roche)), and lysed on ice with a 2-ml tissue grinder (Kontes) with 2 strokes by hand. The resulting cell lysate was fractionated via three 1-h centrifugation steps at 3g, 2,g, and 1,g at 4 C in thin-wall polyallomer (2.2 ml, 11 mm 35 mm) tubes (Beckman Coulter) with an S-55S swinging bucket rotor in a Sorvall MTX 15 micro-ultracentrifuge (Thermo Scientific). The supernatant was adjusted to 1 M sucrose in buffer, and the resulting 2 ml fraction was layered under a 12-ml sucrose step-gradient with 2-ml steps (.75,.5, 5,.125, M) and centrifuged for 12 h at 2,g at 4 C in 13.2-ml thin-wall polyallomer tubes (Beckman Coulter) with a TH-641 swinging bucket rotor in a Sorvall WX8 ultracentrifuge (Thermo Scientific). The floating white LD fraction was collected with a tube slicer (Beckman Coulter), and the other 2-ml fractions were collected with a pipette. Six fractions of the gradient and the three pellet fractions of the differential centrifugation steps were analyzed via MS-based proteomics. Protein Digestion One hundred micrograms of each heavy SILAC sucrose gradient fraction were mixed with equal amounts of light SILAC LD standard. Combined samples were precipitated with 2 ml of ice-cold acetone at 4 C overnight. Precipitated proteins were collected via centrifugation, dried at room temperature, dissolved in 5 l 6 M urea, 2 M thiourea, and 1 mm Tris, and subjected to in-solution digestion. Protein samples were then reduced with 5 l 1 mm DTT for 45 min at room temperature, alkylated with 5 l 5.5 mm iodoacetamide for 3 min, and digested with 4 g endopeptidase Lys-C (Waco) for 3 h. The resulting peptide mixtures were diluted with 2 l 5mM ammonium bicarbonate and digested with 4 g sequencing-grade modified trypsin (Promega) per protein overnight at room temperature. Trypsin was inactivated via acidification with.5 l trifluoroacetic acid (ph 3), and subsequently 5 g of peptides was concentrated and desalted on reversed-phase C 18 STAGE tips (37, 38). Liquid Chromatography MS/MS Analysis Peptides were eluted from STAGE tips by 3 l buffer B (8% acetonitrile in.5% acetic acid) solution into a 96-well sample plate (Abgene), concentrated in a SpeedVac until the organic solvent was removed, and reconstituted with a one-to-one mix of buffer A (.5% acetic acid) and buffer A* (2% acetonitrile in.1% trifluoroacetic acid). Eluted peptides were analyzed using a nanoflow high-pressure liquid chromatography system (Agilent Technologies or Proxeon Biosystems) coupled on-line via a nanoelectrospray ion source (Proxeon Biosystems) to an LTQ-Orbitrap mass spectrometer (Thermo Scientific). Peptide samples were loaded onto a C 18 reversed-phase column (15 cm long, 75 m inner diameter, packed in-house with ReproSil-Pur C 18 -AQ 3- m resin) in buffer A (.5% acetic acid) with a flow rate of 5 nl/min and eluted with a linear gradient from 2% to 4% buffer B (8% acetonitrile and.5% acetic acid solution) at a flow rate of 25 nl/min over 2 h. Mass spectra were acquired in the positive ion mode, applying a data-dependent automatic switch between the acquisition of one Orbitrap survey scan (mass range of m/z 3 17) and MS/MS of the five most intense ions in the LTQ ( top 5 method). The target value in the LTQ-Orbitrap was 1,, for the survey scan at a resolution of 6, at m/z 4 with lock masses. Fragmentation in the LTQ was performed by means of collision-induced dissociation with a target value of 5 ions. The ion selection threshold was 1 counts. Selected sequenced ions were dynamically excluded for 9 s. MS Data Analysis Raw mass spectrometric data were analyzed with MaxQuant (version ) (39). The raw data and MaxQuant output files are available from the Tranche proteome repository with the following access codes: raw data: DgrMLzEv/6uaJdL QG1eq- TzvRMsAVU9N7VkL1urdh4cEd9s81j2LyAZtnIMVNaxi oxms8g- Exz8Cqwipojr8uY57EAAAAAAAASFg MaxQuant output files: ciebtexjn93219yd BeDHPxkP7inbSYed2mj34buWup2jD- Dy9B2i kip3u9nogsysn934kmxzasqabmdlwf/xkqqaaaaaaa- AHLQ Summary output file: 2LQWICE HalrmoOv5bTV- H2IY4kP2wx63wYf3jJ8 ecsepid8bdexjh8oidikuus2idpx9ia5- ZSAxZXJzhArn4XKRvlHsAAAAAAAABVg Molecular & Cellular Proteomics 12.5
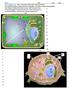
3 The peak list was searched against FlyBase (version 5.24), containing 15,16 proteins combined with common contaminants and concatenated with the reversed versions of all sequences using Andromeda, a search engine incorporated into the MaxQuant framework (4). Precursor and fragment ions were searched with a maximal initial mass deviation of up to 7 ppm or.5 Da, respectively. In the peptide search, trypsin, allowing for cleavage N-terminal to proline, was used as the enzyme specificity. Cysteine carbamidomethylation was selected as a fixed modification, and protein N-terminal acetylation and methionine oxidation were chosen as variable modifications. Depending on a priori knowledge about the number of Arg and Lys residues in the precursor ion determined by MaxQuant before the search, Arg1 and Lys8 were used as additional fixed or variable modifications. A maximum of two missed cleavages and three labeled amino acids were allowed. MaxQuant automatically quantified and normalized peptides and proteins based on the light SILAC LD standard. Quantification of SILAC pairs was performed by MaxQuant with standard settings using a minimum ratio count of 2. A false discovery rate of.1 was required for proteins and peptides with a minimum length of six amino acids. Hierarchical clustering and gene ontology annotation analysis were performed using Perseus software, version Soft clustering was performed using Mfuzz, a software package implemented in the statistical program language R (41). Lys Arg(L) Lys8 Arg1 (H) Proteoly c Digest LC- ESI MS MS/MS Data Analysis 5 Sample Mixing RESULTS Frac ons FIG. 1.Experimental scheme of quantitative analysis of an LD proteome via protein correlation profiling. LDs are purified from two populations of Drosophila S2 cells, one unlabeled (ArgLys) (purple) and one labeled with heavy amino acids Arg1 and Lys8 (green). After three steps of differential centrifugation (centrifugation pellets are fractions 9 7), cell lysates are further fractionated with a sucrose step gradient (fractions 6 1). LDs are large round structures in fraction 1. The partitioning of schematic sample proteins is indicated. For quantitative analysis, the top (LD-containing) fraction of the light sample is mixed with each of the six fractions of the sucrose gradient and the three pellet fractions of the heavy samples. Mixed fractions are de-lipidated, and proteins are precipitated. After insolution digestion of the proteins, samples are separately analyzed via liquid chromatography on-line coupled to high-resolution MS/MS. The heavy/light ratio (H/L) for the proteins of each fraction was normalized by dividing it by the highest ratio among the fractions, giving a value of to 1 (H/L) for each fraction. Profiles show values plotted against each fraction for a number of example proteins represented schematically. H/L A Protein Correlation Profile Strategy to Analyze the LD Proteome We used Drosophila S2 cells, for which abundant functional data on LD biology exist, to develop an LD PCP. The PCP strategy requires two components that are each labeled with different isotope-containing amino acids: a purified LD sample (the target organelle or LD standard ) and all fractions of a separate LD purification (Fig. 1). To obtain these fractions, we purified LDs from cells that were treated with oleate and metabolically labeled with heavy, non-radioactive isotope-containing lysine and arginine (SILAC (42)). Of several tested protocols, a combination of sequential differential centrifugations followed by a sucrose density gradient purification proved to be most effective. To reduce the possibility of contamination of the LD fraction by other organelles or vesicles of disrupted organelles during cell homogenization, we first used three differential centrifugation steps to efficiently remove nuclei, mitochondria, and cellular membranes (fractions 7 9). These centrifugation steps efficiently removed many heavy organelles from the LDs, optimizing our protocol for the analysis of LD proteins, but this procedure precludes the distinction of, for example, mitochondrial and nuclear proteins in the resulting protein correlation profile. In addition, these centrifugation steps decrease the protein complexity of the resulting samples, which enables more comprehensive MS analysis. Samples were then subjected to sucrose density gradient centrifugation, which is commonly used to isolate LDs (36). We modified this part of the purification protocol with a sucrose step gradient (fractions 6 1), instead of a continuous sucrose gradient, in order to increase the focusing of protein complexes from different organelles at the interphases of the density steps. To generate the LD standard for PCP analysis, we collected the LD fraction (fraction 1) from a similar purification scheme of oleate-loaded cells that were labeled with light isotope-containing lysine and arginine. The PCP method is based on spiking a light amino-acidlabeled LD standard in each of the heavy isotope-labeled fractions of the purification. Upon analysis, therefore, the ratio of heavy- and light-labeled versions of each protein in each sample reflects its abundance in this fraction relative to the Molecular & Cellular Proteomics
4 Fraction Proteins identified Protein SILAC ratio Protein SILAC ratio (%) TABLE I Identified proteins Peptides identified Razor peptides Unique peptides Proteins with 5 valid values , ,4 13,673 13, ,14 14,379 13, ,422 14,799 14, ,253 14,647 14, ,47 14,767 14, ,47 13,429 12, ,728 13,11 12, ,477 13,859 13, Total ,961 25,969 25, Table indicates the numbers of proteins identified in total and in each of the purification fractions of MS measurements before and after filtering for at least five valid values in all nine fractions, as well as the number and percentage of proteins identified with the SILAC ratio and the numbers of peptides, unique peptides, and razor peptides in each fraction. abundance in the LD fraction. Assembling these abundance ratios for each protein in all fractions yields a plot reflecting the purification and enrichment of each of the identified proteins (shown conceptually in Fig. 1). To determine the relative abundance of the proteins in each fraction, we subjected the mixed samples to liquid chromatography coupled on-line to high-resolution MS (Fig. 1). The resulting spectra were analyzed with the MaxQuant software suite (39, 4), leading to an overall identification of 273 proteins. For the LD fraction, this resulted in the identification of 1481 proteins. The number of identified proteins in the other fractions ranged from 1946 (fraction 8) to 2126 (fraction 9), and in each case, more than 99% were quantified with SILAC ratios (see Table I for proteomics results, supplemental Table S2 for complete proteomics results, and supplemental Fig. S1 for annotated spectra for all single peptide identifications). To assess the reproducibility of the LD PCP measurements, we compared the results of two different mass spectrometric analyses of the purification fractions mixed with the LD standard. Plotting the protein abundance ratios from the two technical replicates revealed a high correlation between ratios of peptide abundance in a fraction of the two different experiments (Fig. 2), indicating that our measurements were highly reproducible. Considering this high correlation, most of the variation in the ratios in fraction 1 (average heavy/light ratio (H/L) of S.D.) likely derives from variation in either biological samples or different preparations. Our analysis also revealed that the average ratio in fraction 1 did not center on 1 (Fig. 2A). This systematic error is likely due to the complications of measuring protein concentrations in an LD fraction containing large amounts of triacylglycerol. Importantly, because the same amount of LD standard was added to each fraction, the measured ratios of proteins to this standard were comparable across all measured samples. The use of the same internal standard in all measurements thus further adds to the robustness of the PCP approach. FIG. 2. High reproducibility of MS measurements for the LD PCP. The ratios of the heavy and light labeled versions of each protein from two independent measurements are plotted against each other for each fraction of the purification. The high correlation of the data points in the scatterplots indicates that differences in the amounts of proteins from light versus heavy labeled samples can be detected with high reproducibility. Pearson correlations and regression lines are shown in red. LD PCP Yields Distinct Purification Profiles for LDs and Other Organelles We next tested whether the LD PCP segregated organelles into distinct fractions. To compare the profiles of individual proteins, we set the maximum ratio obtained to 1 for each protein and normalized the ratios in the remaining fractions relative to this (H/L). We first analyzed the averaged profiles of known marker proteins for abundant organelles, including the nuclei, the ER, mitochondria, and LDs. The individual abundance profiles of the marker proteins are shown in supplemental Table S2 and are plotted in supplemental Fig. S2. Each of the organelle profiles was distinct from each of the others and from profiles of cytosolic proteins (Fig. 3A). Moreover, LDs peaked in the top fraction of the gradient centrifugation, as expected, whereas proteins of organelles characterized by higher density, such as the ER and 1118 Molecular & Cellular Proteomics 12.5
5 FIG. 3. Identification of LD proteins in S2 cells via LD PCP. A, averaged fractionation profiles for different organelle marker proteins. The average normalized ratios for marker proteins of the indicated cellular organelles, as determined via quantitative LC-MS/MS, are shown for fractions of an LD purification. Values are mean S.D. of five proteins. B, hierarchical clustering of H/L for proteins identified in fraction 1 of LC-MS/MS analysis of all fractions of a LD purification with Perseus (39). At least five of nine valid values per protein were required. The color code represents the protein ratio H/L normalized by the maximum value among the fractions. Gray indicates that the protein was not identified in that fraction. Clusters enriched for proteins of certain organelles are marked with white boxes. Cellular organelles whose proteins were enriched in a certain cluster are indicated on the right. C, soft clustering of proteins identified in fraction 1 of LC-MS/MS analysis of all fractions of an LD purification with Mfuzz in R (41). Proteins identified in the LD fraction that were identified in at least five of the nine fractions were analyzed. A normalized H/L was used. The cluster number C was set at 9, and cluster stability m Clusters that showed enrichment of proteins of a certain organelle or function are indicated. Proteins with a minimal membership value of.3 for LD cluster 1 are shown in supplemental Table S2. Golgi apparatus, peaked in the denser fractions. We cannot exclude the possibility that fragments of these organelles might float with LDs. However, as proteins of other organelles have their main abundance in other fractions, such contaminants are nonetheless efficiently subtracted from bona fide LD proteins. We next analyzed the PCP results for the LD fraction. Because MS analysis identifies the most abundant peaks in a spectrum, the lower abundance of some proteins can result in stochastic absences of measurements of their peptides in some fractions. To offset this potential problem, we filtered our data for proteins that were detected in at least five of the nine purification fractions. Of the 1481 proteins identified in the LD fraction, 1389 met these filtering criteria. To assess the relative behavior of different proteins with the LD PCP method, we applied hierarchical clustering of the filtered profiles (peptides in at least five fractions). This analysis revealed specific clusters with distinct purification profiles (Fig. 3B), as expected for proteins purifying as part of an organelle. This contrasts with a continuous spectrum of purification profiles, as would be expected if proteins distributed independently of one another. Analysis of the clusters by gene ontology showed that many of them had an enrichment of marker proteins of a predominant organelle (Fig. 3B). Among the proteins in the LD fraction, we found some that peaked in abundance in a different fraction and thus likely were part of a different organelle. Particularly, ER membrane proteins were abundant in the LD fraction (fraction 1), possibly because of the close association between ER and LDs, but they also had a major peak in fraction 8. In contrast, ER luminal proteins had a peak with cytosolic proteins (fractions 2 4), in addition to the ER membrane peak in fraction 8, likely indicating the release of luminal proteins into the cytosol during purification. The application of hierarchical clustering to LD PCP profiles has a limitation: the process assigns a protein to one organelle class, even if there are two different localizations of the protein (e.g. LDs and ER). To overcome this limitation, we analyzed our data via soft clustering, which generates clusters of typical profiles and assigns to each protein a membership value for each cluster (41). The membership value (a value between and 1) provides a measurement of the similarity of each protein profile to the core proteins of the cluster. Thus, it measures the likelihood that a protein belongs to a cluster. For example, the similarity of a protein s profile to the LD profile is expressed as an LD membership value for each protein, which is an indication of the likelihood that a particular protein is an LD protein (supplemental Table S3). LD PCP Reliably Identifies LD Proteins Soft clustering identified a profile ( cluster 1 ) that was typical of LD proteins (Fig. 3C, supplemental Table S3). Of the 1389 proteins identified in the LD fraction with more than five valid values in all fractions, only 111 showed an LD-specific purification profile with a membership value.3 for the LD cluster. This indicates that only 7.5% of the proteins identified in this fraction were candidates for bona fide LD proteins, corresponding to roughly.74% of all Drosophila genes. Soft clustering additionally identified groups of proteins representing other organ- Molecular & Cellular Proteomics
6 A B Brummer CG5295 CGI-58 CG1882 SELT CG3887 Short chain dehydrogenase CG264 C Faf2 CG1372 Fatp CG74 Prenyltransferase CG1778 Lipase CG17292 Oxysterol binding protein CG1513 LD LD+ER LD+other H/L H/L H/L H/L H/L H/L H/L H/L H/L FIG. 4. Localization of a subset of proteins found in the LD proteome as determined via fluorescent microscopy of expressed proteins. The indicated fluorescent mcherry-tagged enzymes were transiently expressed in S2 cells (left-hand panels, red) loaded with 1 mm oleate for 12 h. LDs were stained with BODIPY (middle panels, green). The overlays of the two channels and zoomed views of a representative LD section are shown (rightmost two panels). Bar 5 m (overview) or 1 m. Protein correlation profiles for the tagged proteins are shown in the right-hand panels in red. Dark gray lines indicate averaged PCPs for LD proteins, and bright gray lines indicate averaged protein correlation profiles for ER proteins (Fig. 3A). A, proteins localizing exclusively to LDs. B, proteins localizing to LDs and ER. C, proteins localizing to LDs and any other cellular compartment. elles (e.g. cluster 2 contains primarily proteins of the Golgi apparatus). To test whether the proteins identified by PCP as LD proteins were indeed LD proteins, we analyzed their subcellular distribution by means of fluorescence microscopy of expressed proteins in Drosophila S2 cells. Of the 111 candidates, we randomly picked 2 proteins (supplemental Table S4) for analysis. Confirming the PCP analysis, 18 of the 2 proteins (9%) showed a characteristic ring-like staining pattern around LDs (examples can be seen in Figs. 4A 4C). The other two proteins, encoded by CG514 (VAP33) and CG33113 (reticulon), showed fluorescent aggregates in most cells and a reticular ER pattern in a few cells. It is not clear whether the observed aggregation and localization to other organelles might be due to tagging and/or overexpression of these proteins, rather than a reflection of their normal subcellular distribution. Alternatively, these two proteins might not be genuine LD proteins. Moreover, considering that the over- 112 Molecular & Cellular Proteomics 12.5
7 A 81 S2 cell LD-PCP 6 Beller et al, Cermelli et al, 24 expression of fluorescently tagged proteins might lead to spill-over to other organelles, it is difficult to distinguish bona fide LD proteins from proteins with dual cellular localization. Therefore, PCP might be advantageous over microscopy for tagged proteins, as it measures the distribution of endogenous proteins. Of the remaining 18 proteins, 4 showed only LD-ring signal in fluorescent microscopy, and we classified these as having an LD-only localization (Fig. 4A, supplemental Fig. S3A). In contrast, nine other proteins contained an additional small peak in the fraction containing mostly ER proteins and also showed a reticular pattern, likely representing an ER signal, under fluorescence microscopy. We classified these proteins as LD ER (Fig. 4B, supplemental Fig. S3B). In these cases, tagged proteins might localize to both the ER and LDs, either because they have a dual localization or because the LD surface is saturated and cannot accommodate additional protein. Alternatively, LD localization in these cases might be due to the spill-over of overexpressed protein from the ER. The remaining five proteins showed a fluorescence signal around LDs, but also some signal that likely represented an organelle other than the ER, such as the plasma membrane (see Fig. 4C or supplemental Fig. S3C CG17292 for an example), and had small additional peaks in the LD PCP that did not overlap with the ER fraction. We classified those proteins as localized to LD other organelles. Comparing the LD PCP and Other LD Proteome Studies Several proteomics studies have analyzed the composition of LDs in different cell types and species (1, 11, 13, 16 19, 21). No data are available on the LD composition of Drosophila cell lines, although proteomic data were generated by two conventional MS-based proteomics from LDs purified from Drosophila larval fat body or embryos (1, 11). Relative to the other Drosophila studies, the sensitivity of our measurements was much greater (in total, 1481 proteins identified in the LD fraction, in comparison to 645 and 247 proteins). In contrast, however, our LD PCP strategy filtered out contaminants and thereby defined with high confidence a much smaller set of LD proteins (111). This suggests that more than 9% of the proteins typically identified in an LD fraction via MS are likely contaminants. Not surprisingly, therefore, the two LD proteomes of Drosophila fat body and larvae overlap by only 14.4% (112 of a total 78 proteins). In comparing our results with the previously published Drosophila studies, we found little overlap (Fig. 5A, Table II). For example, we found 1 and 24 proteins overlapping with those identified in the studies of Beller et al. and Cermelli et al., respectively (1, 11). Comparing the LD PCP Proteome with Genes Identified via Functional Screens We and others reported high-content genome-wide screens for LD phenotypes after depletion of each Drosophila protein from cell lines via RNAi (34, 43). In addition, screens in Drosophila larvae (44) and adult flies (45) identified genes that function in fat storage. We expected that some of the LD proteins we identified would perform important functions for normal LD morphology and/or fat storage. To identify such proteins, we analyzed the data from our LD PCP proteome for overlap with genes previously found in screens for LD-related phenotypes (34, 43 45). We found relatively little overlap between the data sets (Table III), showing that many proteins influencing LD morphology are not localized to the organelle. Reciprocally, many LD proteins function in cellular processes that are independent of LD morphology. Moreover, proteins playing crucial roles in LD function might have redundantly functioning homologs, precluding a strong LD phenotype when only one of the proteins is depleted. Our LD PCP identified groups of proteins participating in the biological processes that have not been previously linked to LDs (Fig. 5B). For example, we found a transcription factor, enzymes of lysine metabolism, and parts of the machinery 513 B N-glycan biosynthesis B-1,4N-acglucosaminyltransferase CG638 B-1,4N-acglucosaminyltransferase CG14512 B-1,4-mannosyltransferase CG1812 Mannosyl-oligosaccharide A-1,2-mannosidase CG42275 A-1,3/A-1,6-mannosyltransferase CG1291 A-1,2-mannosyltransferase CG1136 Faf2 CG1372 UBX CG842 Derlin CG198 HECT CG564 Tango14 CG6668 TG synthesis G-3-P-acyltransferase CG329 G-3-P-acyltransferase CG558 Lyso-PA-acyltransferase CG4729 Lipin CG879 Lipid droplet Phospholipid metabolism Nessy CG9566 Lyso-PC acyltransferase CG32699 CCT1 CG149 CCT2 CG1833 Lipolysis HSL CG1155 Lipase CG17292 Brummer CG5295 CGI-58 CG1882 FIG. 5.Comparison of the LD PCP with other proteomic studies and analysis for enriched pathways. A, comparison of the LD proteome with published Drosophila LD proteomes (1, 11). Venn diagram shows overlap between the three studies. B, overrepresentation of specific pathways among LD-protein-associated functions. Groups of proteins with related functions detected on LDs are shown. Molecular & Cellular Proteomics
8 TABLE II Overlap between LD proteomes Protein name/function CG Cermelli et al., 26 (11) 44DEa, fatty-acid ligase activity CG8732 Saccheropin dehydrogenase CG5167 Atlastin CG6668 Abhydrolase 1, CGI-58 CG1882 Saccheropin dehydrogenase CG264 Hormone sensitive lipase CG1155 Brummer CG5295 Amidase CG8839 Lipase CG9186 Glycosyltransferase CG1291 Amidase CG5112 Kugelkem CG5175 Toutatis CG1897 Prenyltransferase CG1778 Fatp CG74 Fatty acyl-coa reductase CG833 Vap-33 1 CG514 Rab8 CG8287 SNF4A, Loechrig, kinase CG17299 Rab7 CG5915 Beller et al., 26 (1) Reticulon CG33113 Saccheropin dehydrogenase CG5167 Saccheropin dehydrogenase CG264 Brummer CG5295 Lipase CG9186 Alpha esterase CG1112 Fatty acid desaturase CG5887 Cytochrome b5 CG214 Fatty-acid ligase activity CG3961 All three studies Saccheropin dehydrogenase CG5167 Saccheropin dehydrogenase CG264 Brummer CG5295 Lipase CG9186 Proteins identified in LD PCP and in one of the published Drosophila LD proteome studies. generating dolichol-linked glycans for protein glycosylation on LDs. To further test the reliability of the LD PCP and to assay for the localization of the N-glycan synthesis machinery, we fluorescently tagged all enzymes of dolichol-linked glycan synthesis indentified via MS and compared the localization of the tagged proteins with their profiles generated via PCP (Fig. 6A). In each of the tested cases, localization of the tagged enzymes via fluorescence microscopy correlated well with their assignment to LDs or the ER based on the correlation profiles of the LD PCP. Alg1, Alg2, and Alg14 had a clear LD profile in the LD PCP and exclusively localized to LDs via fluorescence microscopy. Alg11, showing a strong peak in the LD and a smaller peak in the ER fraction, showed dual localization between LDs and ER. In contrast, Alg1, which had an ER profile in the LD PCP, was present in a reticular pattern excluded from LDs, likely representing the ER. Alg3 and Alg9, which were not detected in the LD PCP, also localized in an ER pattern (data not shown). These findings show that specifically, enzymes catalyzing the early steps of glycan synthesis for N-linked protein modification localize to LDs. Furthermore, these data further confirm that our approach reliably distinguishes between LD and ER proteins. DISCUSSION In this paper, we describe a quantitative proteomics approach based on LD purification and MS that employs SILAC and PCP, similar to what has been employed for other organelles (29), to efficiently determine an LD proteome with a high degree of confidence. The combination of a careful purification scheme and PCP methodology enabled us to extract from the complete LD-containing fraction a set of just over 1 LD proteins with significant confidence. This approach overcomes the limitations of classical fractionation approaches that assign proteins to an organelle based solely on their presence in a purified organelle fraction, rather than co-enrichment with an organelle. Importantly, although our approach relies on cellular fractionation, which can yield false organelle protein identification as result of changes in the behavior of disrupted membranes, we were able to confirm almost all tested candidate proteins for LD localization via fluorescence microscopy, increasing our confidence in our methodology. Our results included some previously identified proteins, validating our approach, and the procedure excluded other proteins as contaminants and identified novel proteins that might function at the LDs. We focused our analysis on LDs in Drosophila S2 cells as proof of principle for the methodology. These cells provide a well-established system for studying LD cell biology and have been used in high-content, microscopy-based RNAi screens for LD phenotypes (34, 43). Moreover, the ease and efficiency of gene expression knockdown by RNAi in S2 cells allows future analyses of LD proteomes after the depletion of various cellular factors. Our LD PCP results yielded several expected proteins with important LD functions. For example, we found CCT1 and CCT2, the isoenzymes catalyzing the rate-limiting step of de novo phosphatidylcholine synthesis, which we had previously found to localize to LDs (46). Similarly, as expected, we identified Brummer, the Drosophila homolog of the major triglyceride lipase ATGL. In addition, we identified a number of proteins that were not previously known as LD proteins but which are functionally connected to LDs. For example, our LD PCP results identified lysophosphatidic acid acyltransferase, which was among 66 genes that resulted in increased larval fat mass when inactivated (44). In addition, the overlay of hit lists from genetic screens and our LD PCP identified several proteins not previously implicated in LD biology. For example, we identified SELT, a glycosyltransferase, which results in smaller, more dispersed LDs when depleted (34), and Mes2, a transcription factor highly expressed in fat body (47, 48) that 1122 Molecular & Cellular Proteomics 12.5
9 TABLE III LD proteins with LD phenotype Protein name CG Phenotype Study SelT-like protein CG3887 Moderately altered LD morphology in S2 cells; Guo et al., 28 (34) more, smaller, more dispersed LDs CCT1 CG149 Dramatically altered LD morphology in S2 cells; Guo et al., 28 (34) fewer, larger LDs CCT2 CG1833 Dramatically altered LD morphology in S2 cells; Guo et al., 28 (34) fewer, larger LDs Rab8 CG8287 Altered mean signal in automated analysis in S2 Guo et al., 28 (34) Brummer CG5295 Overstorage in S2 cells Beller et al., 28 (43) Mes2 CG111 Overstorage in S2 cells Beller et al., 28 (43) Lyso-PA acyltransferase CG32699 Increased fat storage in Drosophila larvae Reis et al., 21 (44) Fatp CG74 Increased fat storage in Drosophila larvae Reis et al., 21 (44) Cytochrome P45 6a2 CG9438 Reduced triacylglycerol levels in adult Drosophila Pospisilik et al., 21 (45) Alpha-esterase-7 CG1112 Reduced triacylglycerol levels in adult Drosophila Pospisilik et al., 21 (45) Myosin V CG2146 Elevated triacylglycerol levels in adult Drosophila Pospisilik et al., 21 (45) Proteins identified in the LD PCP whose deletion leads to an alteration in LD morphology or lipid metabolism found in different published screens. leads to triacylglycerol over-storage when depleted (43). The localization of Mes2 on LD suggests a signaling function in transmitting information from LDs to the nucleus. We also identified Rab8, a small GTPase involved in vesicular trafficking that was found in a genome-wide screen of genes involved in LD morphology (43). A comparison of our proteomes with conventional proteomic studies of Drosophila larval fat body or total larvae LDs (1, 11) showed little overlap between the identified proteins. This could be due to either variations between larval and S2 cell LD proteomes or a larger number of contaminants in the conventional proteomes than in the LD PCP. The latter possibility is supported by the presence of many proteins in the conventional proteome results that have been localized primarily to other organelles, such as ribosomal components or mitochondrial proteins. However, assessing a priori whether a protein is a contaminant based on its known function is challenging. This is highlighted by the unexpected finding that histones, with a well-defined nuclear function, are important LD proteins in developing Drosophila embryos (11). In addition to expected proteins, such as CCT and Brummer, we found a number of interesting proteins that had been reported in other studies. For example, the LD PCP and conventional proteomes found an as yet uncharacterized protein with homology to lipases (CG9186 (11)). Moreover, prenyltransferase and fatty acid desaturases were detected among other lipid metabolism enzymes by means of both LD PCP and conventional proteomics. Most likely, these enzymes are abundant and thus have been identified in a number of proteomic studies. Intriguingly, all three LD proteomics studies identified saccheropin dehydrogenase on LDs, suggesting a role for LDs in lysine metabolism. We also found several proteins that were previously not known to localize to LDs. Among them, CG17292 is particularly interesting, as it contains a lipase motif and is highly expressed in Drosophila oenocytes, a cell type that functions similarly to hepatocytes (49). This finding, together with data suggesting that the enzyme has retinyl esterase activity in vitro, 2 suggests a function for CG17292 in lipid metabolism. Unexpectedly, the enzymes catalyzing the early steps of the N-linked protein glycosylation pathway were found in the LD PCP, and we confirmed their localization via fluorescence microscopy. In this pathway, glycans are first sequentially added to a dolichol lipid moiety. This process is thought to occur on the cytosolic side of the ER. The resulting glycolipid then flips across the membrane in a poorly characterized reaction to function as a substrate for oligosaccharyltransferase, which transfers the glycan to asparagine residues of ER-lumen exposed proteins. Here, we found specifically that the enzymes that build the glycan tree on dolichol were enriched on LDs (Fig. 6B). In addition, we detected the homolog of Rft1, proposed to be involved in flipping the dolichol-glycan across the ER bilayer (5), only in LD fraction 1, and not any other fraction, suggesting that it might also be enriched strongly on LDs. Previously, these enzymes were thought to catalyze their reactions on the cytoplasmic side of the ER (51). Interestingly, a previous study in yeast found Dpm1, the enzyme catalyzing the first step of glycan synthesis, on LDs (52). These data suggest a link between N-linked glycosylation and LD biology. Of note, dolichol isoprenoid sidechains are particularly extended lipids that might be more easily accommodated at LDs than at the ER bilayer membrane. In addition to proteins that we found to be exclusively enriched with LDs, our dataset also revealed proteins that have a dual localization. Such proteins have a pool localized on the LD surface, leading to an abundance peak in the LD fraction, and an additional pool acting at a different cellular site, leading to an additional peak in a different fraction of the purification. Examples of such proteins are the members of 2 N. Krahmer, T. C. Walter, M. Kumari, and R. Zechner, unpublished observation. Molecular & Cellular Proteomics
10 A Alg14 CG638 Alg1 CG1812 Alg2 CG1291 Alg11 CG1136 Alg1 CG3276 LD LD+ER ER H/L H/L H/L H/L H/L B Cytosol ALG1 ALG13/14 ALG7 ALG2 ALG2 ALG11 LD ALG11 RFT1 ALG3 ALG9 ER lumen ALG12 ALG9 STT3 OST ALG1 ALG8 ALG6 FIG. 6. Several N-glycan biosynthesis enzymes localize to LDs. A, the indicated mcherry-tagged proteins of the N-glycan synthesis pathway were expressed and analyzed as described for Fig. 4. PCPs of the indicated proteins are shown on the left. B, organization of the N-glycan biosynthesis on LDs and ER. the COPI machinery, the lysosomal NPC2 protein, and epsin, which thus might have dual localizations in the cell, with a pool of each protein localized to LDs. The identification of such proteins might aid in elucidating the processes that mediate the interaction between organelles and might shed light on the role of such junctions in lipid metabolism. The LD PCP methodology described herein can be easily adapted for the analysis of LD proteomes of a wide range of samples. Also, LD PCP can be combined with dynamic measurements (e.g. to determine changes in proteome composition after the treatment of cells with different stimuli). In such an experiment, one can include another LD fraction, with a third SILAC condition. Changes in the abundance ratio between heavy and medium versions of a protein would then indicate changes in the LD proteome. For determining LD proteomes in tissues, SILAC labeling of the sample is not easily feasible, although it has been achieved for whole animals (53, 54). Alternatively, for such cases an LD standard can be obtained from a closely related cell line. This approach, using a labeled hepatocyte cell line, was productive in the analysis of the phosphoproteomic response to liver insulin signaling (55). In cases when a single cell type does not reflect 1124 Molecular & Cellular Proteomics 12.5
11 the composition of the tissue cells well, a Super-SILAC approach, in which extracts of several cell lines are mixed to reflect the tissue proteome composition (56), can also be used to generate an LD standard for use in an LD PCP of tissue samples. Thus, the LD PCP is an approach with which one can confidently indentify LD proteins that are isolated from a wide range of sources, including cells and tissues, and studied under dynamic conditions. In summary, we have established a robust method with which to identify LD proteins based on the subtraction of contaminants using PCP combined with subcellular fractionation. By applying this LD PCP method to Drosophila cells, we have revealed expected LD proteins, but we also have identified novel LD proteins and uncovered potential links between LDs and other cellular processes, such as protein glycosylation. We expect that this methodology will be widely applicable to LD research and will help define the dynamic proteome of this important metabolic organelle in different tissues and metabolic states. Acknowledgments We thank Nadin Neuhauser and Nagarjuna Nagaraj for assistance in data processing and members of the Walther and Farese laboratories for critical comments on the manuscript, as well as Gary Howard for editorial assistance. * This work was supported by Grant No. R1 GM97194 to T.C.W. R.V.F. was supported by Grant No. RO1 GM99844 and the Gladstone Institutes. F.W. is a fellow of the Boehringer Ingelheim Fonds. S This article contains supplemental material. To whom correspondence should be addressed: Tobias Walther, Yale University School of Medicine, Department of Cell Biology, SHM-C 425C, 333 Cedar Street, New Haven, CT 652, tobias.walther@yale.edu. REFERENCES 1. Bensimon, A., Heck, A. J., and Aebersold, R. (212) Mass spectrometrybased proteomics and network biology. Annu. Rev. Biochem. 81, Yan, W., Aebersold, R., and Raines, E. W. (29) Evolution of organelleassociated protein profiling. J. Proteomics 72, Walther, T. C., and Mann, M. (21) Mass spectrometry-based proteomics in cell biology. J. Cell. Biol. 19, Farese, R. V., Jr., and Walther, T. C. (29) Lipid droplets finally get a little R-E-S-P-E-C-T. Cell 139, Fujimoto, T., and Parton, R. G. (211) Not just fat: the structure and function of the lipid droplet. Cold Spring Harb. Perspect. Biol. 3, a Brasaemle, D. L., and Wolins, N. E. (212) Packaging of fat: an evolving model of lipid droplet assembly and expansion. J. Biol. Chem. 287, Herker, E., and Ott, M. (211) Unique ties between hepatitis C virus replication and intracellular lipids. Trends Endocrinol. Metab. 22, Greenberg, A. S., Coleman, R. A., Kraemer, F. B., McManaman, J. L., Obin, M. S., Puri, V., Yan, Q. W., Miyoshi, H., and Mashek, D. G. (211) The role of lipid droplets in metabolic disease in rodents and humans. J. Clin. Invest. 121, Bozza, P. T., and Viola, J. P. (21) Lipid droplets in inflammation and cancer. Prostaglandins Leukot. Essent. Fatty Acids 82, Beller, M., Riedel, D., Jänsch, L., Dieterich, G., Wehland, J., Jäckle, H., and Kühnlein, R. (26) Characterization of the Drosophila lipid droplet subproteome. Mol. Cell. Proteomics 5, Cermelli, S., Guo, Y., Gross, S. P., and Welte, M. A. (26) The lipid-droplet proteome reveals that droplets are a protein-storage depot. Curr. Biol. 16, Pu, J., Ha, C. W., Zhang, S., Jung, J. P., Huh, W. K., and Liu, P. (211) Interactomic study on interaction between lipid droplets and mitochondria. Protein Cell 2, Brasaemle, D. L., Dolios, G., Shapiro, L., and Wang, R. (24) Proteomic analysis of proteins associated with lipid droplets of basal and lipolytically stimulated 3T3-L1 adipocytes. J. Biol. Chem. 279, Ding, Y., Wu, Y., Zeng, R., and Liao, K. (212) Proteomic profiling of lipid droplet-associated proteins in primary adipocytes of normal and obese mouse. Acta Biochim. Biophys. Sin. (Shanghai) 15. Larsson, S., Resjo, S., Gomez, M. F., James, P., and Holm, C. (212) Characterization of the lipid droplet proteome of a clonal insulin-producing beta-cell line (INS-1 832/13). J. Proteome Res. 11, Zhang, H., Wang, Y., Li, J., Yu, J., Pu, J., Li, L., Zhang, S., Peng, G., Yang, F., and Liu, P. (211) Proteome of skeletal muscle lipid droplet reveals association with mitochondria and apolipoprotein a-i. J. Proteome Res. 1, Blouin, C. M., Le Lay, S., Eberl, A., Kofeler, H. C., Guerrera, I. C., Klein, C., Le Liepvre, X., Lasnier, F., Bourron, O., Gautier, J. F., Ferre, P., Hajduch, E., and Dugail, I. (21) Lipid droplet analysis in caveolin-deficient adipocytes: alterations in surface phospholipid composition and maturation defects. J. Lipid Res. 51, Bartz, R., Zehmer, J. K., Zhu, M., Chen, Y., Serrero, G., Zhao, Y., and Liu, P. (27) Dynamic activity of lipid droplets: protein phosphorylation and GTP-mediated protein translocation. J. Proteome Res. 6, Liu, P., Ying, Y., Zhao, Y., Mundy, D. I., Zhu, M., and Anderson, R. G. (24) Chinese hamster ovary K2 cell lipid droplets appear to be metabolic organelles involved in membrane traffic. J. Biol. Chem. 279, Wu, C. C., Howell, K. E., Neville, M. C., Yates, J. R., III, and McManaman, J. L. (2) Proteomics reveal a link between the endoplasmic reticulum and lipid secretory mechanisms in mammary epithelial cells. Electrophoresis 21, Zhang, P., Na, H., Liu, Z., Zhang, S., Xue, P., Chen, Y., Pu, J., Peng, G., Huang, X., Yang, F., Xie, Z., Xu, T., Xu, P., Ou, G., Zhang, S. O., and Liu, P. (212) Proteomic study and marker protein identification of caenorhabditis elegans lipid droplets. Mol. Cell. Proteomics, 11, Blanchette-Mackie, E. J., and Scow, R. O. (1983) Movement of lipolytic products to mitochondria in brown adipose tissue of young rats: an electron microscope study. J. Lipid Res. 24, Shaw, C. S., Jones, D. A., and Wagenmakers, A. J. M. (28) Network distribution of mitochondria and lipid droplets in human muscle fibres. Histochem. Cell Biol. 129, Jacquier, N., Choudhary, V., Mari, M., Toulmay, A., Reggiori, F., and Schneiter, R. (211) Lipid droplets are functionally connected to the endoplasmic reticulum in Saccharomyces cerevisiae. J. Cell Sci. 124, Ozeki, S., Cheng, J., Tauchi-Sato, K., Hatano, N., Taniguchi, H., and Fujimoto, T. (25) Rab18 localizes to lipid droplets and induces their close apposition to the endoplasmic reticulum-derived membrane. J. Cell Sci. 118, Martin, S., Driessen, K., Nixon, S. J., Zerial, M., and Parton, R. G. (25) Regulated localization of Rab18 to lipid droplets: effects of lipolytic stimulation and inhibition of lipid droplet catabolism. J. Biol. Chem. 28, Liu, P., Bartz, R., Zehmer, J. K., Ying, Y. S., Zhu, M., Serrero, G., and Anderson, R. G. (27) Rab-regulated interaction of early endosomes with lipid droplets. Biochim. Biophys. Acta 1773, Binns, D., Januszewski, T., Chen, Y., Hill, J., Markin, V. S., Zhao, Y., Gilpin, C., Chapman, K. D., Anderson, R. G., and Goodman, J. M. (26) An intimate collaboration between peroxisomes and lipid bodies. J. Cell Biol. 173, Andersen, J. S., and Mann, M. (26) Organellar proteomics: turning inventories into insights. EMBO Rep. 7, Andersen, J. S., Lam, Y. W., Leung, A. K., Ong, S. E., Lyon, C. E., Lamond, A. I., and Mann, M. (25) Nucleolar proteome dynamics. Nature 433, Andersen, J. S., Lyon, C. E., Fox, A. H., Leung, A. K., Lam, Y. W., Steen, H., Mann, M., and Lamond, A. I. (22) Directed proteomic analysis of the human nucleolus. Curr. Biol. 12, Andersen, J. S., Wilkinson, C. J., Mayor, T., Mortensen, P., Nigg, E. A., and Mann, M. (23) Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, Molecular & Cellular Proteomics
Molecular Cell, Volume 46. Supplemental Information
 Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting
 Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Chapter 1 The Lipid Droplet: a Dynamic Organelle, not only Involved in the Storage and Turnover of Lipids
 Chapter 1 The Lipid Droplet: a Dynamic Organelle, not only Involved in the Storage and Turnover of Lipids Sven-Olof Olofsson, Pontus Boström, Jens Lagerstedt, Linda Andersson, Martin Adiels, Jeanna Perman,
Chapter 1 The Lipid Droplet: a Dynamic Organelle, not only Involved in the Storage and Turnover of Lipids Sven-Olof Olofsson, Pontus Boström, Jens Lagerstedt, Linda Andersson, Martin Adiels, Jeanna Perman,
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Regulation of Lipid Homeostasis: Lipid Droplets
 Regulation of Lipid Homeostasis: Lipid Droplets Bernd Helms Article Brand Recter Hendrik Mertens 1 The Basics I: FAs The Basics II: FA Activation 2 Basics III: TG-FA Interplay. Why? Adipocytes 3 Foam cells
Regulation of Lipid Homeostasis: Lipid Droplets Bernd Helms Article Brand Recter Hendrik Mertens 1 The Basics I: FAs The Basics II: FA Activation 2 Basics III: TG-FA Interplay. Why? Adipocytes 3 Foam cells
AP Biology Cells: Chapters 4 & 5
 AP Biology Cells: Chapters 4 & 5 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. The was the first unifying principle of biology. a. spontaneous generation
AP Biology Cells: Chapters 4 & 5 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. The was the first unifying principle of biology. a. spontaneous generation
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Introduction. Biochemistry: It is the chemistry of living things (matters).
 Introduction Biochemistry: It is the chemistry of living things (matters). Biochemistry provides fundamental understanding of the molecular basis for the function and malfunction of living things. Biochemistry
Introduction Biochemistry: It is the chemistry of living things (matters). Biochemistry provides fundamental understanding of the molecular basis for the function and malfunction of living things. Biochemistry
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Protein Trafficking in the Secretory and Endocytic Pathways
 Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Zool 3200: Cell Biology Exam 4 Part I 2/3/15
 Name: Key Trask Zool 3200: Cell Biology Exam 4 Part I 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Name: Key Trask Zool 3200: Cell Biology Exam 4 Part I 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
FOCUS SubCell. For the Enrichment of Subcellular Fractions. (Cat. # ) think proteins! think G-Biosciences
 169PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name FOCUS SubCell For the Enrichment of Subcellular Fractions (Cat. # 786 260) think
169PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name FOCUS SubCell For the Enrichment of Subcellular Fractions (Cat. # 786 260) think
LECTURE-15. itraq Clinical Applications HANDOUT. Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS
 LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Master class Biomolecular Sciences Molecular Cell Biology.
 Master class Biomolecular Sciences Molecular Cell Biology. 04-09-08: Paul van Bergen en Henegouwen. Clathrin-indep Endocytosis 11-09-08: Willem Stoorvogel. Endocytosis and MHC classii 18-09-08: X-track
Master class Biomolecular Sciences Molecular Cell Biology. 04-09-08: Paul van Bergen en Henegouwen. Clathrin-indep Endocytosis 11-09-08: Willem Stoorvogel. Endocytosis and MHC classii 18-09-08: X-track
OxiSelect Human Oxidized LDL ELISA Kit (OxPL-LDL Quantitation)
 Product Manual OxiSelect Human Oxidized LDL ELISA Kit (OxPL-LDL Quantitation) Catalog Number STA-358 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic
Product Manual OxiSelect Human Oxidized LDL ELISA Kit (OxPL-LDL Quantitation) Catalog Number STA-358 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic
Intracellular Compartments and Protein Sorting
 Intracellular Compartments and Protein Sorting Intracellular Compartments A eukaryotic cell is elaborately subdivided into functionally distinct, membrane-enclosed compartments. Each compartment, or organelle,
Intracellular Compartments and Protein Sorting Intracellular Compartments A eukaryotic cell is elaborately subdivided into functionally distinct, membrane-enclosed compartments. Each compartment, or organelle,
1. endoplasmic reticulum This is the location where N-linked oligosaccharide is initially synthesized and attached to glycoproteins.
 Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
endomembrane system internal membranes origins transport of proteins chapter 15 endomembrane system
 endo system chapter 15 internal s endo system functions as a coordinated unit divide cytoplasm into distinct compartments controls exocytosis and endocytosis movement of molecules which cannot pass through
endo system chapter 15 internal s endo system functions as a coordinated unit divide cytoplasm into distinct compartments controls exocytosis and endocytosis movement of molecules which cannot pass through
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
ab SREBP-2 Translocation Assay Kit (Cell-Based)
 ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Practice Exam 2 MCBII
 1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for
 Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Cell Cell
 Go to cellsalive.com. Select Interactive Cell Models: Plant and Animal. Fill in the information on Plant and Animal Organelles, then Click on Start the Animation Select Plant or Animal Cell below the box.
Go to cellsalive.com. Select Interactive Cell Models: Plant and Animal. Fill in the information on Plant and Animal Organelles, then Click on Start the Animation Select Plant or Animal Cell below the box.
Zool 3200: Cell Biology Exam 4 Part I 2/3/15
 Name: Trask Zool 3200: Cell Biology Exam 4 Part I 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Name: Trask Zool 3200: Cell Biology Exam 4 Part I 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
10/13/11. Cell Theory. Cell Structure
 Cell Structure Grade 12 Biology Cell Theory All organisms are composed of one or more cells. Cells are the smallest living units of all living organisms. Cells arise only by division of a previously existing
Cell Structure Grade 12 Biology Cell Theory All organisms are composed of one or more cells. Cells are the smallest living units of all living organisms. Cells arise only by division of a previously existing
2013 John Wiley & Sons, Inc. All rights reserved. PROTEIN SORTING. Lecture 10 BIOL 266/ Biology Department Concordia University. Dr. S.
 PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
Complexity DNA. Genome RNA. Transcriptome. Protein. Proteome. Metabolites. Metabolome
 DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
Problem Set #5 4/3/ Spring 02
 Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Cellular control of cholesterol. Peter Takizawa Department of Cell Biology
 Cellular control of cholesterol Peter Takizawa Department of Cell Biology Brief overview of cholesterol s biological role Regulation of cholesterol synthesis Dietary and cellular uptake of cholesterol
Cellular control of cholesterol Peter Takizawa Department of Cell Biology Brief overview of cholesterol s biological role Regulation of cholesterol synthesis Dietary and cellular uptake of cholesterol
Structure & Function of Cells
 Anatomy & Physiology 101-805 Unit 4 Structure & Function of Cells Paul Anderson 2011 Anatomy of a Generalised Cell Attached or bound ribosomes Cilia Cytosol Centriole Mitochondrion Rough endoplasmic reticulum
Anatomy & Physiology 101-805 Unit 4 Structure & Function of Cells Paul Anderson 2011 Anatomy of a Generalised Cell Attached or bound ribosomes Cilia Cytosol Centriole Mitochondrion Rough endoplasmic reticulum
Proteins: Proteomics & Protein-Protein Interactions Part I
 Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
1. This is the location where N-linked oligosaccharide is initially synthesized and attached to glycoproteins.
 Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Thursday, October 16 th
 Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
Human Carbamylated LDL ELISA Kit (CBL-LDL Quantitation)
 Product Manual Human Carbamylated LDL ELISA Kit (CBL-LDL Quantitation) Catalog Number MET-5032 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic
Product Manual Human Carbamylated LDL ELISA Kit (CBL-LDL Quantitation) Catalog Number MET-5032 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic
7.06 Cell Biology EXAM #3 April 24, 2003
 7.06 Spring 2003 Exam 3 Name 1 of 8 7.06 Cell Biology EXAM #3 April 24, 2003 This is an open book exam, and you are allowed access to books and notes. Please write your answers to the questions in the
7.06 Spring 2003 Exam 3 Name 1 of 8 7.06 Cell Biology EXAM #3 April 24, 2003 This is an open book exam, and you are allowed access to books and notes. Please write your answers to the questions in the
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Summary of Endomembrane-system
 Summary of Endomembrane-system 1. Endomembrane System: The structural and functional relationship organelles including ER,Golgi complex, lysosome, endosomes, secretory vesicles. 2. Membrane-bound structures
Summary of Endomembrane-system 1. Endomembrane System: The structural and functional relationship organelles including ER,Golgi complex, lysosome, endosomes, secretory vesicles. 2. Membrane-bound structures
Human Oxidized LDL ELISA Kit (MDA-LDL Quantitation), General
 Human Oxidized LDL ELISA Kit (MDA-LDL Quantitation), General For the detection and quantitation of human OxLDL in plasma, serum or other biological fluid samples Cat. No. KT-959 For Research Use Only.
Human Oxidized LDL ELISA Kit (MDA-LDL Quantitation), General For the detection and quantitation of human OxLDL in plasma, serum or other biological fluid samples Cat. No. KT-959 For Research Use Only.
Endoplasmic Reticulum
 Endoplasmic Reticulum What s ER? How is ER? Why is ER? definition description functions Nissl s bodies neurons Berg s bodies hepatocytes Organelle structure histocytochemical evidences Ergastoplasm pancreatic
Endoplasmic Reticulum What s ER? How is ER? Why is ER? definition description functions Nissl s bodies neurons Berg s bodies hepatocytes Organelle structure histocytochemical evidences Ergastoplasm pancreatic
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose.
 Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Mammalian Membrane Protein Extraction Kit
 Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine
 Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
Protein sorting (endoplasmic reticulum) Dr. Diala Abu-Hsasan School of Medicine dr.abuhassand@gmail.com An overview of cellular components Endoplasmic reticulum (ER) It is a network of membrane-enclosed
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
Cellular compartments
 Cellular compartments 1. Cellular compartments and their function 2. Evolution of cellular compartments 3. How to make a 3D model of cellular compartment 4. Cell organelles in the fluorescent microscope
Cellular compartments 1. Cellular compartments and their function 2. Evolution of cellular compartments 3. How to make a 3D model of cellular compartment 4. Cell organelles in the fluorescent microscope
Cell are made up of organelles. An ORGANELLE is a specialized subunit within a cell that has a specific function.
 Plant and Animal Cells The Cell Theory All living things are made up of one or more cells. All cells come from other cells. Organization of Living Things Cell are made up of organelles. An ORGANELLE is
Plant and Animal Cells The Cell Theory All living things are made up of one or more cells. All cells come from other cells. Organization of Living Things Cell are made up of organelles. An ORGANELLE is
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes
 Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
ab CytoPainter Golgi/ER Staining Kit
 ab139485 CytoPainter Golgi/ER Staining Kit Instructions for Use Designed to detect Golgi bodies and endoplasmic reticulum by microscopy This product is for research use only and is not intended for diagnostic
ab139485 CytoPainter Golgi/ER Staining Kit Instructions for Use Designed to detect Golgi bodies and endoplasmic reticulum by microscopy This product is for research use only and is not intended for diagnostic
Cell morphology. Cell organelles structure and function. Chapter 1: UNIT 1. Dr. Charushila Rukadikar
 UNIT 1 Cell morphology Cell organelles structure and function Chapter 1: Dr. Charushila Rukadikar Assistant Professor Department Of Physiology ZMCH, Dahod Physiology The science that is concerned with
UNIT 1 Cell morphology Cell organelles structure and function Chapter 1: Dr. Charushila Rukadikar Assistant Professor Department Of Physiology ZMCH, Dahod Physiology The science that is concerned with
Cell Structure & Function. Source:
 Cell Structure & Function Source: http://koning.ecsu.ctstateu.edu/cell/cell.html Definition of Cell A cell is the smallest unit that is capable of performing life functions. http://web.jjay.cuny.edu/~acarpi/nsc/images/cell.gif
Cell Structure & Function Source: http://koning.ecsu.ctstateu.edu/cell/cell.html Definition of Cell A cell is the smallest unit that is capable of performing life functions. http://web.jjay.cuny.edu/~acarpi/nsc/images/cell.gif
Homework Hanson section MCB Course, Fall 2014
 Homework Hanson section MCB Course, Fall 2014 (1) Antitrypsin, which inhibits certain proteases, is normally secreted into the bloodstream by liver cells. Antitrypsin is absent from the bloodstream of
Homework Hanson section MCB Course, Fall 2014 (1) Antitrypsin, which inhibits certain proteases, is normally secreted into the bloodstream by liver cells. Antitrypsin is absent from the bloodstream of
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Total Phosphatidic Acid Assay Kit
 Product Manual Total Phosphatidic Acid Assay Kit Catalog Number MET- 5019 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Phosphatidic Acid (PA) is a critical precursor
Product Manual Total Phosphatidic Acid Assay Kit Catalog Number MET- 5019 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Phosphatidic Acid (PA) is a critical precursor
BIOLOGY 111. CHAPTER 3: The Cell: The Fundamental Unit of Life
 BIOLOGY 111 CHAPTER 3: The Cell: The Fundamental Unit of Life The Cell: The Fundamental Unit of Life Learning Outcomes 3.1 Explain the similarities and differences between prokaryotic and eukaryotic cells
BIOLOGY 111 CHAPTER 3: The Cell: The Fundamental Unit of Life The Cell: The Fundamental Unit of Life Learning Outcomes 3.1 Explain the similarities and differences between prokaryotic and eukaryotic cells
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
O O H. Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION EXPERIMENTAL. LC /MS conditions
 Simplifying Qual/Quan Analysis in Discovery DMPK using UPLC and Xevo TQ MS Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION The determination of the drug metabolism
Simplifying Qual/Quan Analysis in Discovery DMPK using UPLC and Xevo TQ MS Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION The determination of the drug metabolism
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
CELLS.
 CELLS http://www.aimediaserver.com/studiodaily/harvard/harvard.swf INTERESTING FACTS The longest cells in the human body are the motor neurons. They can be up to 1.37 meters long and go from the spinal
CELLS http://www.aimediaserver.com/studiodaily/harvard/harvard.swf INTERESTING FACTS The longest cells in the human body are the motor neurons. They can be up to 1.37 meters long and go from the spinal
Trypsin Digestion Mix
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2008
 Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Chapter 1 Membrane Structure and Function
 Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Cellular Biochemistry
 Cellular Biochemistry Fall Semester 2013 Sept. 23 Benoit Kornmann Institute of Biochemistry Introduction to biological membranes General functions and properties Membrane lipids Physical properties Distribution/asymmetry
Cellular Biochemistry Fall Semester 2013 Sept. 23 Benoit Kornmann Institute of Biochemistry Introduction to biological membranes General functions and properties Membrane lipids Physical properties Distribution/asymmetry
CELL PARTS TYPICAL ANIMAL CELL
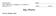 AP BIOLOGY CText Reference, Campbell v.8, Chapter 6 ACTIVITY1.12 NAME DATE HOUR CELL PARTS TYPICAL ANIMAL CELL ENDOMEMBRANE SYSTEM TYPICAL PLANT CELL QUESTIONS: 1. Write the name of the cell part in the
AP BIOLOGY CText Reference, Campbell v.8, Chapter 6 ACTIVITY1.12 NAME DATE HOUR CELL PARTS TYPICAL ANIMAL CELL ENDOMEMBRANE SYSTEM TYPICAL PLANT CELL QUESTIONS: 1. Write the name of the cell part in the
Cell Structure. Present in animal cell. Present in plant cell. Organelle. Function. strength, resist pressure created when water enters
 Cell Structure Though eukaryotic cells contain many organelles, it is important to know which are in plant cells, which are in animal cells and what their functions are. Organelle Present in plant cell
Cell Structure Though eukaryotic cells contain many organelles, it is important to know which are in plant cells, which are in animal cells and what their functions are. Organelle Present in plant cell
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
Supplementary Figure 1. Method development.
 Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
The Cell Organelles. Eukaryotic cell. The plasma membrane separates the cell from the environment. Plasma membrane: a cell s boundary
 Eukaryotic cell The Cell Organelles Enclosed by plasma membrane Subdivided into membrane bound compartments - organelles One of the organelles is membrane bound nucleus Cytoplasm contains supporting matrix
Eukaryotic cell The Cell Organelles Enclosed by plasma membrane Subdivided into membrane bound compartments - organelles One of the organelles is membrane bound nucleus Cytoplasm contains supporting matrix
04_polarity. The formation of synaptic vesicles
 Brefeldin prevents assembly of the coats required for budding Nocodazole disrupts microtubules Constitutive: coatomer-coated Selected: clathrin-coated The formation of synaptic vesicles Nerve cells (and
Brefeldin prevents assembly of the coats required for budding Nocodazole disrupts microtubules Constitutive: coatomer-coated Selected: clathrin-coated The formation of synaptic vesicles Nerve cells (and
Applying a Novel Glycan Tagging Reagent, RapiFluor-MS, and an Integrated UPLC-FLR/QTof MS System for Low Abundant N-Glycan Analysis
 Applying a Novel Glycan Tagging Reagent, RapiFluor-MS, and an Integrated UPLC-FLR/QTof MS System for Low Abundant N-Glycan Analysis Ying Qing Yu Waters Corporation, Milford, MA, USA APPLICATION BENEFITS
Applying a Novel Glycan Tagging Reagent, RapiFluor-MS, and an Integrated UPLC-FLR/QTof MS System for Low Abundant N-Glycan Analysis Ying Qing Yu Waters Corporation, Milford, MA, USA APPLICATION BENEFITS
The Study of Cells The diversity of the cells of the body The following figure shows the proportion of cell size of the variety of cells in the body
 Adapted from Martini Human Anatomy 7th ed. Chapter 2 Foundations: The Cell Introduction There are trillions of cells in the body Cells are the structural building blocks of all plants and animals Cells
Adapted from Martini Human Anatomy 7th ed. Chapter 2 Foundations: The Cell Introduction There are trillions of cells in the body Cells are the structural building blocks of all plants and animals Cells
Mitochondrial DNA Isolation Kit
 Mitochondrial DNA Isolation Kit Catalog Number KA0895 50 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
Mitochondrial DNA Isolation Kit Catalog Number KA0895 50 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
Don t Freak Out. Test on cell organelle on Friday!
 Cell Structure 1 Don t Freak Out Test on cell organelle on Friday! This test should be a buffer test and help raise your overall test score. All information will come from this week! 2 Cells Provide Compartments
Cell Structure 1 Don t Freak Out Test on cell organelle on Friday! This test should be a buffer test and help raise your overall test score. All information will come from this week! 2 Cells Provide Compartments
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
Cell Structure and Function. Biology 12 Unit 1 Cell Structure and Function Inquiry into Life pages and 68-69
 Cell Structure and Function Biology 12 Unit 1 Cell Structure and Function Inquiry into Life pages 45 59 and 68-69 Assignments for this Unit Pick up the notes/worksheet for this unit and the project There
Cell Structure and Function Biology 12 Unit 1 Cell Structure and Function Inquiry into Life pages 45 59 and 68-69 Assignments for this Unit Pick up the notes/worksheet for this unit and the project There
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Renata Schipp Medical Biology Department
 Renata Schipp Medical Biology Department Deffinition of cell The cell is the smallest structural and functional unit of all known living organisms The cell was discovered by Robert Hooke in 1665 and also
Renata Schipp Medical Biology Department Deffinition of cell The cell is the smallest structural and functional unit of all known living organisms The cell was discovered by Robert Hooke in 1665 and also
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry
 Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids
 Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids Using the QTRAP System with mtraq Reagents Karin A. Zemski-Berry 1, John M. Hevko 2, and Robert C. Murphy 1 1 Department
Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids Using the QTRAP System with mtraq Reagents Karin A. Zemski-Berry 1, John M. Hevko 2, and Robert C. Murphy 1 1 Department
In-Solution Digestion for proteomics
 In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin
 SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
PROTEIN TRAFFICKING. Dr. SARRAY Sameh, Ph.D
 PROTEIN TRAFFICKING Dr. SARRAY Sameh, Ph.D Overview Proteins are synthesized either on free ribosomes or on ribosomes bound to endoplasmic reticulum (RER). The synthesis of nuclear, mitochondrial and peroxisomal
PROTEIN TRAFFICKING Dr. SARRAY Sameh, Ph.D Overview Proteins are synthesized either on free ribosomes or on ribosomes bound to endoplasmic reticulum (RER). The synthesis of nuclear, mitochondrial and peroxisomal
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Supporting information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
/searchlist/6850.html Tour of the Cell 1
 http://www.studiodaily.com/main /searchlist/6850.html Tour of the Cell 1 2011-2012 Cytology: science/study of cells To view cells: Light microscopy resolving power: measure of clarity Electron microscopy
http://www.studiodaily.com/main /searchlist/6850.html Tour of the Cell 1 2011-2012 Cytology: science/study of cells To view cells: Light microscopy resolving power: measure of clarity Electron microscopy
