Cellular Biology. The Deubiquitylase USP2 Regulates the LDLR Pathway by Counteracting the E3-Ubiquitin Ligase IDOL
|
|
|
- Ellen Hill
- 5 years ago
- Views:
Transcription
1 Cellular Biology The Deubiquitylase USP2 Regulates the LDLR Pathway by Counteracting the E3-Ubiquitin Ligase IDOL Jessica Kristine Nelson,* Vincenzo Sorrentino,* Rossella Avagliano Trezza, Claire Heride, Sylvie Urbe, Ben Distel, Noam Zelcer Downloaded from by guest on July 13, 2018 Rationale: The low-density lipoprotein (LDL) receptor (LDLR) is a central determinant of circulating LDLcholesterol and as such subject to tight regulation. Recent studies and genetic evidence implicate the inducible degrader of the LDLR (IDOL) as a regulator of LDLR abundance and of circulating levels of LDL-cholesterol in humans. Acting as an E3-ubiquitin ligase, IDOL promotes ubiquitylation and subsequent lysosomal degradation of the LDLR. Consequently, inhibition of IDOL-mediated degradation of the LDLR represents a potential strategy to increase hepatic LDL-cholesterol clearance. Objective: To establish whether deubiquitylases counteract IDOL-mediated ubiquitylation and degradation of the LDLR. Methods and Results: Using a genetic screening approach, we identify the ubiquitin-specific protease 2 (USP2) as a post-transcriptional regulator of IDOL-mediated LDLR degradation. We demonstrate that both USP2 isoforms, USP2-69 and USP2-45, interact with IDOL and promote its deubiquitylation. IDOL deubiquitylation requires USP2 enzymatic activity and leads to a marked stabilization of IDOL protein. Paradoxically, this also markedly attenuates IDOL-mediated degradation of the LDLR and the ability of IDOL to limit LDL uptake into cells. Conversely, loss of USP2 reduces LDLR protein in an IDOL-dependent manner and limits LDL uptake. We identify a tri-partite complex encompassing IDOL, USP2, and LDLR and demonstrate that in this context USP2 promotes deubiquitylation of the LDLR and prevents its degradation. Conclusions: Our findings identify USP2 as a novel regulator of lipoprotein clearance owing to its ability to control ubiquitylation-dependent degradation of the LDLR by IDOL. (Circ Res. 2016;118: DOI: / CIRCRESAHA ) Key Words: E3 ubiquitin ligase LDL cholesterol LDL receptors MYLIP/IDOL ubiquitin USP2 The low-density lipoprotein (LDL) receptor (LDLR) is a major determinant of circulating levels of LDL. 1 Accordingly, mutations in this receptor are the leading cause for development of autosomal-dominant hypercholesterolemia, a disease characterized by reduced hepatic LDL clearance, elevated plasma cholesterol levels, and accelerated atherosclerosis. 2,3 Owing to its central role in lipoprotein metabolism, the level of the LDLR is subject to tight homeostatic regulation. Transcription of the LDLR is controlled by the sterol regulatory element-binding proteins, and it is increased when cellular cholesterol levels decline to allow increased endocytosis of cholesterol-rich LDL particles. 4,5 In This Issue, see p 367 Editorial, see p 371 Next to transcriptional regulation, post-transcriptional mechanisms are also emerging as important determinants of LDLR abundance. Among these, ubiquitylation of the LDLR plays an important role in controlling the intracellular trafficking of the receptor, 6,7 similar to the role this process plays for other membrane receptors. We have recently identified inducible degrader of the LDLR (IDOL) as an E3-ubiquitin ligase (E3) that specifically promotes ubiquitylation of the LDLR. 8 IDOL itself is subject to direct transcriptional regulation by the sterol-sensing transcription factors liver X receptors (LXR). Activation of LXR when cellular sterol levels increase leads to induction of IDOL expression and activity, and subsequent ubiquitylation of conserved residues in the short intracellular tail of the LDLR. In turn, ubiquitylation of the LDLR serves to mark the receptor for internalization and subsequent lysosomal degradation. 8 In particular, the plasma membrane pool of LDLR is highly sensitive to IDOL activity, and ubiquitylation leads to rapid removal of this receptor pool via an epsin-dependent but clathrin-independent endocytic Original received July 28, 2015; revision received December 8, 2015; accepted December 14, In November 2015, the average time from submission to first decision for all original research papers submitted to Circulation Research was days. From the Department of Medical Biochemistry, Academic Medical Center of the University of Amsterdam, Amsterdam, The Netherlands (J.K.N., V.S., R.A.T., B.D., N.Z.); and Department of Cellular and Molecular Physiology, University of Liverpool, Liverpool, United Kingdom (C.H., S.U.). *These authors contributed equally to this article. The online-only Data Supplement is available with this article at /-/DC1. Correspondence to Noam Zelcer, PhD, Department of Medical Biochemistry, Meibergdreef 15, 1105AZ Amsterdam, The Netherlands. n.zelcer@amc.uva.nl 2015 American Heart Association, Inc. Circulation Research is available at DOI: /CIRCRESAHA
2 Nelson et al USP2 Counteracts IDOL-Mediated LDLR Degradation 411 Downloaded from by guest on July 13, 2018 Nonstandard Abbreviations and Acronyms E3 IDOL LDLR LXR USP E3-ubiquitin ligase inducible degrader of the LDLR low-density lipoprotein receptor liver X receptors ubiquitin-specific protease route. 6 The role of IDOL in human lipoprotein metabolism is supported by genome-wide association studies that identify an association between genetic variation in the IDOL locus and circulating levels of LDL-cholesterol. 9,10 Furthermore, we recently described a rare loss-of-function IDOL variant in individuals with LDL-cholesterol. 11 Consistent with this, silencing of IDOL in nonhuman primates using an RNAi-based approach resulted in a reduction of circulating levels of LDLcholesterol, which was associated with increased levels of hepatic LDLR protein. 12 Thus, the LXR IDOL LDLR axis is a potent ubiquitylation-dependent mechanism to acutely limit lipoprotein-derived cholesterol uptake. 13,14 Similar to other post-translational modifications, ubiquitylation is reversible; a process dependent on the activity of deubiquitylases. 15 The human genome encodes 100 deubiquitylases, 79 of which are predicted to be active. 16,17 These deubiquitylases are classified into 5 distinct families, the largest being the ubiquitin-specific protease (USP) family. A major challenge in elucidating the cellular function and physiological roles of deubiquitylases is identification of their substrate specificity. 16,18 Adding to the complexity, recent studies indicate that in addition to removing or remodeling ubiquitin (chains) from ubiquitylated substrates, deubiquitylases may also act directly on E3s, thereby modifying the stability and activity of the latter. 19,20 Characterizing the deubiquitylase-e3 interactome thus represents an important step in understanding how E3s are themselves regulated by ubiquitylation. 17,21 Whether deubiquitylases play a role in cholesterol metabolism is largely unknown. Our work and that of Scotti et al 7 have recently shown that activity of the endosomal-sorting complex required for transport associated USP8 is required for IDOL-dependent degradation of ubiquitylated LDLR. 6 However, this most likely represents a nonselective deubiquitylation step used to salvage ubiquitin from ubiquitylated cargo before it entering multivesicular bodies. Given its potent LDLR-lowering activity, we hypothesized that the IDOL pathway may be itself subject to post-transcriptional regulation, in addition to transcriptional control by LXRs. Herein, we identify USP2 as a negative regulator of IDOL-dependent degradation of the LDLR, thereby implicating this deubiquitylase as a regulator of lipoprotein metabolism. Methods A brief description of the methods is provided below. Detailed description of the methods are available at the Online Data Supplement. Cell Culture and Transfections HEK293T, HeLa, Huh7, and HepG2 cells were obtained from the ATCC and maintained in DMEM supplemented with 10% fetal bovine serum at 37 C and 5% CO 2. A431 cells were a kind gift from Drs Vilja Pietiainen and Elina Ikonen (University of Helsinki). Wild-type and Idol ( / ) mouse embryonic fibroblasts (MEFs) were a kind gifts from Dr Peter Tontonoz (University of California at Los Angeles). DU145 were from Dr Koen van de Wetering (NKI, Amsterdam). Where indicated, cells were sterol depleted by culture in sterol-depletion medium (DMEM supplemented by 10% lipoprotein-deficient serum, 5 μg/ml simvastatin, and 100 μmol/l mevalonate) or treated with 1 μmol/l GW3965 to induce LXR signaling. HEK293T and HepG2 cells were transfected with the indicated amounts of plasmids using the JetPrime reagent, and transfection efficiency was monitored by cotransfection of a GFP expression plasmid. Plasmids and Expression Constructs Expression plasmids encoding IDOL, LDLR, USP8, very low-density lipoprotein receptor, GFP, Ubiquitin, and Epsin1 were previously reported. 6,8,22 Expression plasmids encoding USP2-69 and USP2-45 cdnas were generated with gateway-mediated recombination (Invitrogen). Unless otherwise indicated, USP2-69 is referred to as USP2. The activity disabling C276A 23,24 mutation in USP2 was introduced using the QuikChange site-directed mutagenesis kit. All plasmids used in this study experiments were isolated by CsCl 2 gradient centrifugation and their correctness verified by sequencing. Antibodies and Immunoblot Analysis Total cell lysates were prepared in radioimmunoprecipitation assay buffer (RIPA; 150 mmol/l NaCl, 1% Nonidet P-40, 0.1% sodium deoxycholate, 0.1% SDS, 100 mmol/l Tris-HCl, and ph 7.4) supplemented with protease inhibitors. Lysates were cleared by centrifugation at 4 C for 10 minutes at g. Samples were separated on NuPAGE BisTris gels and transferred to nitrocellulose. The primary antibodies used are listed in the extended Methods section in Online Data Supplement. Secondary HRP (horseradish peroxidase)-conjugated antibodies were used and visualized with enhanced chemiluminescence. All immunoblots are representative of at least 3 independent experiments. LDL Uptake Assay DyLight 488-labeled LDL was produced as previously described. 25 Briefly, HepG2, HeLa, or A431 cells were incubated in sterol-depletion medium for 16 hours before addition of LDL to increase LDLR abundance. To initiate LDL uptake, cells were incubated with 5 μg/ml DyLight488-labeled LDL in DMEM supplemented with 0.5% BSA at 37 C. Subsequently, cells were washed twice with PBS supplemented with excess sterols (PBS with 10% FCS), followed by an additional wash with PBS. Endocytosis of LDL was determined by lysing cells in RIPA buffer and quantification of the fluorescence signal on a Typhoon imager. Relative LDL uptake was calculated from the measure of fluorescence values corrected for specific uptake (cells treated with labeled LDL in the presence of an excess of nonlabeled LDL) and presented as mean±sd. Measurement of Cell Surface LDLR Surface LDLR density in the different cells was determined as previously reported. 6 Briefly, cells were treated as indicated, dissociated, and incubated in fluorescence-activated cell sorter blocking buffer for 30 minutes on ice. Subsequently, cells were stained with a R-Phycoerythrin-conjugated mouse antihuman LDLR antibody for 1 hour on ice. Cells were subsequently washed 3 with fluorescenceactivated cell sorter buffer and directly analyzed on a fluorescenceactivated cell sorter Calibur Flow Cytometer (BD Biosciences). Relative surface LDLR was calculated from geomean values after correction for background using FlowJo, and presented as mean±sd. sirna Transfection To knockdown USP2 expression, we used ON-TARGETplus Human USP2 SMARTpool (Dharmacon: cat#l ), or individual ON-TARGETplus Human USP2 sirnas (Dharmacon: cat#j ). As control, we used the ON-TARGETplus Non-targeting Pool (Dharmacon: cat#d ). A431and HeLa cells were transfected with sirnas (30 nmol/l) using the JetPrime reagent. HepG2 and DU145 cells were transfected similarly, but with the RNAiMAX reagent. The efficiency of USP2 silencing was assessed by quantitative polymerase chain reaction. Unless otherwise indicated, sirna transfection experiments were conducted for 48 hours.
3 412 Circulation Research February 5, 2016 Downloaded from by guest on July 13, 2018 RNA Isolation and Quantitative Polymerase Chain Reaction Total RNA was isolated from cells using TRIzol and 1 μg of total RNA was reverse transcribed using the iscript reverse transcription reagent. SYBR Green real-time quantitative polymerase chain reaction assays were performed on a Lightcycler 480 II apparatus. Oligonucleotide sequences are available on request. Yeast-2-Hybrid Screen Saccharomyces cerevisiae strains used in this study are PJ69-4a (trp1-901 leu2-3,112 ura3-52 his3-200 gal4d gal80d LYS2::GAL1-HIS3 GAL2-ADE2 met2::gal7-lacz) and PJ69-4α (trp1-901 leu2-3,112 ura3-52 his3-200 gal4d gal80d LYS2::GAL1-HIS3 GAL2-ADE2 met2::gal7-lacz). Cells were grown at 28 C in YPAD or in minimal glucose medium. PJ69-4α transformed with human IDOL constructs or LDLR intracellular tail (residues ) cloned into pact2- DEST, and PJ69-4a transformed with the bait constructs cloned into pgdbu-dest were selected on minimal agar plates lacking leucine or uracil, respectively. For mating, cells were precultured for 16 hours in selection medium, and subsequently of transformed PJ69-4a and PJ69-4α cells were mated in 2 YPAD for 24 hours with slow shaking. Diploids were selected on minimal glucose plates lacking both leucine and uracil. To reconfirm interactions, we used a spot assay. Briefly, diploid cells were incubated for 16 hours in minimal glucose medium lacking leucine and uracil, diluted to OD 600 of 0.2 and grown till an OD 600 of 1. Cells were then serially diluted so as to plate cells/spot on minimal glucose agar lacking leucine and uracil, with or without histidine, 10 to 20 mmol/l 3-aminotriazol, and adenine. Plates were incubated for 3 days before analysis. Statistical Analysis Data are presented as mean±sd. One-way ANOVA followed by the Bonferroni correction was performed for comparison of >2 groups. Student s t test was performed for comparison of 2 groups. P values of <0.05 were considered significant. Results USP2 Is a Novel-Binding Partner of IDOL The LDLR is a constitutive recycling receptor that rapidly shuttles between the plasma membrane and endosomal compartments. Internalization of the receptor is a clathrin-dependent process that requires adaptor proteins yet does not involve receptor ubiquitylation. 3 Next to the classical LDLR pathway, IDOL-stimulated ubiquitylation of the receptor promotes internalization of the receptor via a distinct, clathrin-independent endocytic route that requires activity of the endosomal-sorting complex required for transport associated deubiquitylase USP8. 6,7 As strategies to increase abundance of the LDLR form the cornerstone of current therapies of hypercholesterolemia, we hypothesized that other deubiquitylases might counteract IDOL-mediated ubiquitylation and degradation of the LDLR, and thus regulate LDL metabolism. To uncover deubiquitylases potentially involved in the regulation of the IDOL LDLR network, we used a biased yeast-2-hybrid approach aimed at identifying deubiquitylases interacting with IDOL, or alternatively with the intracellular tail of the LDLR. The latter contains the structural and functional requirements for internalization, sorting, and ubiquitylation of the LDLR. 3,22 Our prey library contained 61 deubiquitylases, representing 77% of known active mammalian deubiquitylases, previously used in a systematic deubiquitylase-e3 interaction screen (Online Table I). 20 This screen failed to identify any LDLR-tail-interacting deubiquitylases (Online Figure IA). However, the assay identified the deubiquitylase USP2-69 isoform (referred to further as USP2), as an interacting partner of the full-length IDOL protein (Online Table I). IDOL contains 2 functional regions, a FERM and a RING domain, which are joined by a short linker (Figure 1A). Using the yeast-2-hybrid assay, we mapped IDOL s F3 and linker subdomains as the minimal regions required for interacting with USP2 (Figure 1A and 1B; Online Figure IB). Importantly, the interaction with USP2 was also maintained with a severe IDOL RING mutant, IDOL C387A. This mutation disrupts the conserved RING structure and as a result this mutant is unable to promote LDLR degradation and to support IDOL autoubiquitylation, 8 suggesting that both functions are not essential for the physical interaction of USP2 and IDOL. Furthermore, this result also indicates that USP2 does not compete with binding of the E2-ligase to IDOL, an interaction that depends on an intact RING domain. 22,26 We next confirmed that this interaction also takes place in mammalian cells (HEK293T). Immunoprecipitation of transfected USP2 or IDOL (1:1 ratio) resulted in copurification of the corresponding binding partner (Figure 1C; Online Figure IIA). Furthermore, the interaction with IDOL was independent of USP2s catalytic activity because it was maintained in the USP2 C276A catalytic mutant (Online Figure IIB). In addition to USP2-69 identified in our yeast-2-hybrid screen, a shorter isoform of USP2, USP2-45, has been also described. 27 These 2 USP2 isoforms differ in their N-terminal region and tissue distribution, but share the common C-terminal catalytic domain (Online Figure IIC). Like the USP2-69 isoform, the lowermolecular weight isoform, USP2-45 interacted with IDOL as well (Figure 1D). This points to the catalytic C-terminal region of USP2 as the IDOL interacting region, which is the same region implicated in the interaction of USP2 with the E3 MDM2. 28 Taken together, these results identify USP2 as a novel-binding partner of IDOL and as a potential regulatory component of the IDOL LDLR axis. USP2 Deubiquitylates and Stabilizes IDOL To study the functional consequence of the USP2 IDOL interaction, we initially used a gain-of-function approach. Remarkably, we found that overexpression of USP2 markedly stabilized IDOL protein (Figure 2A). USP8 however failed to do so, supporting the notion that the antagonizing effects of USP8 on the LXR IDOL LDLR axis occur primarily at the level of the ubiquitylated receptor. The stabilization of wild-type IDOL was equivalent to that observed with the proteasome inhibitor MG132, and required USP2 catalytic activity, as it was not seen with USP2 C276A (Figure 2B). Thus, in spite of the preserved interaction of IDOL with both wild-type USP2 and USP2 C276A only the active form of USP2 could increase IDOL levels. This finding implies that the deubiquitylase activity of USP2, rather than steric hindrance is required for IDOL stabilization. We have previously reported that IDOL is a highly unstable protein, likely because of its high autoubiquitylation activity. 8 Accordingly, using cycloheximide to prevent protein synthesis we could determine that stabilization of IDOL by USP2 was a result of the extension of IDOL s half-life (Figure 2C; Online Figure III). The most parsimonious explanation for this observation is that IDOL is subject to USP2-mediated deubiquitylation, which
4 Nelson et al USP2 Counteracts IDOL-Mediated LDLR Degradation 413 Downloaded from by guest on July 13, 2018 Figure 1. The deubiquitylase ubiquitin-specific protease 2 (USP2) is a novel-binding partner of inducible degrader of the lowdensity lipoprotein receptor (IDOL). A, Representation of the IDOL USP2 interaction assay in yeast. IDOL contains an N-terminal FERM domain and a C-terminal RING domain separated by a short linker region. The FERM contains the 3 subdomains, F1, F2, and F3, found in other FERM-containing proteins. A semiquantitive score for the interaction of the IDOL constructs with USP2, based on the growth of the yeast colonies, is reported on the right (+++=strong interaction, =no interaction). B, The different IDOL fragments shown in (A) and USP2 were cloned into yeast-2-hybrid prey and bait plasmids and transformed into yeast. Haploid cells were mated, and diploid cells were serially diluted and plated on selective plates. An image from the first serial dilution (10 5 diploids plated) is shown. C and D, HEK293T cells were transfected with the indicated expression constructs for IDOL, USP2-69, and USP2-45 (IDOL:USP2 ratio of 1:1 3). Total cell lysates were immunoblotted or immunoprecipitated and analyzed as indicated. All immunoblots are representative of at least 3 independent experiments. prevents its degradation. To test this, we conducted experiments in conjunction with pharmacological blocking of proteasomal degradation. This treatment equalizes IDOL levels and promotes accumulation of ubiquitylated IDOL species. Under this condition, both USP2 isoforms reduced the degree of ubiquitylated IDOL (Figure 2D and 2E). Furthermore, consistent with the lack of IDOL stabilization by the catalytic mutant USP2 C276A or by USP8, we observed no changes in the degree of IDOL ubiquitylation by these deubiquitylases. If USP2 removes ubiquitin from IDOL and promotes its stability, the opposite can be expected if USP2 is silenced. Unfortunately, we cannot determine the endogenous level of IDOL in cells, as no commercial or homemade antibodies are able to detect it (data not shown). For that reason, we transfected IDOL into hepatocyte-like human HepG2 and Huh7 cells, and determined the level of heterologous IDOL protein after silencing USP2 expression. We observed that effective silencing of USP2 in these cells (55±6% reduction) reduced immunodetectable IDOL protein (Online Figure IVA and IVB), consistent with the notion that USP2 removes ubiquitin from IDOL to prevent its degradation. Thus, our experiments demonstrate that both gain and loss of USP2 function modulate the level of IDOL protein. USP2 Counteracts IDOL-Mediated Degradation of the LDLR In view of these findings, we predicted that USP2 would promote degradation of the LDLR because of increased IDOL abundance. However, unexpectedly we observed the opposite outcome. Despite massive stabilization of IDOL,
5 414 Circulation Research February 5, 2016 Downloaded from by guest on July 13, 2018 Figure 2. Ubiquitin-specific protease 2 (USP2) deubiquitylates inducible degrader of the low-density lipoprotein receptor (IDOL) and increases its stability. A, HEK293T cells were transfected with the indicated expression constructs for IDOL, USP2, and USP8 (at a IDOL:DUB ratio of 1:3) and total cell lysates were immunoblotted as indicated. B, HEK293T cells were transfected with the indicated expression constructs for IDOL, USP2 wild-type (WT), or the USP2 catalytic mutant. Forty-eight hours after transfection, cells were treated with 25 μmol/l MG132 for 6 hours and collected for analysis. Total cell lysates were immunoblotted as indicated. C, HEK293T cells were transfected with IDOL (black circle) or together with USP2 expression vectors at a 1:5 (gray circle) or 1:10 (white circle) ratio. Forty-eight hours after transfection, cells were incubated for the indicated times with 10 μg/ml cycloheximide (CHX), and cells harvested for immunoblots analysis (n=3, ***P<0.001). D and E, HEK293T cells were transfected with the indicated expression constructs for IDOL, USP8, USP2-69 WT or catalytic mutant, USP2-45 and Ubiquitin. Forty-eight hours after transfection, cells were treated with 25 μmol/l MG132 for 6 hours and collected for analysis. Total cell lysates were immunoblotted or immunoprecipitated and analyzed as indicated. All immunoblots are representative of at least 3 independent experiments. USP2 ( 69 and 45) prevented degradation of the LDLR, as well as of a second IDOL target, the very low-density lipoprotein receptor, 29 in a dose- and activity-dependent fashion (Figure 3A 3C; Online Figure VA). Because IDOL is a transcriptional target of LXR, 8 and LXR-induced degradation of the LDLR is IDOL-dependent, 30 we also evaluated whether USP2 can counteract the effect of LXR on the LDLR. In agreement with our findings, USP2 was also able to counteract LXR-induced LDLR degradation, even though expression of IDOL was induced (Online Figure VB and VC). We also considered the possibility that expression of USP2 is sensitive to the cellular sterol status. However, we found that neither LXR activation nor variation of the cellular sterol content altered USP2 expression (Online Figure VD). Functionally, antagonism of IDOL activity by USP2 counteracted the decrease in cellular LDL uptake imposed by IDOL (Figure 3D), positioning this deubiquitylase as a potential regulator of lipoprotein uptake. Silencing USP2 Promotes LDLR Degradation in an IDOL-Dependent Manner To understand how USP2 counters the IDOL LDLR nexus, we tested the consequence of silencing USP2 on the LDLR pathway in hepatic and nonhepatic cell lines. These cells express IDOL and are, therefore, suitable for studying the functional interaction between IDOL and USP2 (Online Figure VIA). Silencing of USP2 in these cells led to a marked reduction in LDLR protein without changing the level of LDLR mrna (Figure 4A and 4B; Online Figure VIB VID). The secreted protein PCSK9 promotes degradation of the LDLR, and it is expressed by HepG2 cells. 31 It was therefore conceivable that silencing USP2 increases PCSK9 expression, and that this, in turn, leads to LDLR degradation. However, this does not seem to be the case because in HepG2 cells, PCSK9 expression was not sensitive to the level of USP2 (Figure 4B). In contrast to the effects of silencing USP2 on the LDLR, another membrane receptor, the epidermal growth factor receptor (EGFR), remained either
6 Nelson et al USP2 Counteracts IDOL-Mediated LDLR Degradation 415 Downloaded from by guest on July 13, 2018 Figure 3. Ubiquitin-specific protease 2 (USP2) antagonizes the inducible degrader of the low-density lipoprotein (LDL) receptor (IDOL) mediated degradation of lipoprotein receptors. A and B, HEK293T cells were transfected with expression constructs encoding FLAG-IDOL, LDL receptor (LDLR) (A) or very low-density lipoprotein receptor (VLDLR)-HA; B), and MYC-USP2 wild-type (WT) or catalytic mutant as indicated. Forty-eight hours after transfection cells were collected and total cell lysates immunoblotted as indicated. C, HEK293T cells were transfected with the indicated LDLR, IDOL, and USP2 expression vectors, with increasing amounts of WT or mutant USP2. Forty-eight hours after transfection, cells were harvested for immunoblots analysis as shown (D). HEK293T cells were transfected with expression constructs encoding FLAG-IDOL, LDLR, and MYC-USP2 WT. Twenty-four hours after transfection, cells were incubated for 30 minutes with 5 μg/ml Dylight 488-labeled LDL. After extensive washing, internalized LDL was quantified by measuring fluorescence in cell lysates. LDL uptake in cells not expressing IDOL and USP2 was set to 100%. Each bar and error represent the average±sd (n=8, ***P<0.0001). All immunoblots are representative of at least 3 independent experiments. unchanged or even slightly increased after knockdown of USP2 in these cells (Figure 4A; Online Figure VIB). Unfortunately, we were unable to detect endogenous USP2 with commercially available antibodies in our experimental assays (not shown). Therefore, to establish the specificity of our USP2 sirna and to rule out off target effects, we tested 3 individual USP2 sir- NAs. All the 3 effectively knocked down expression of USP2 in HepG2 cells and decreased the protein levels of the LDLR (Online Figure VIIA and VIIB). Consistent with IDOL targeting an LDLR pool present at the plasma membrane, 6,7 we found that silencing USP2 resulted in prominent removal of LDLR from the cell surface (100±2.6 in control versus 30.4±4.9 in siusp2, n=8, P<0.0001; Figure 4C), and as a consequence attenuated LDL uptake in different cell types (Figure 4D). Having established that USP2 influences the level of LDLR protein, we now asked if this mechanism requires IDOL activity. To test this, we silenced Usp2 in wild-type or Idol ( / ) MEFs (Figure 4E; Online Figure VIE). Effective silencing of Usp2 in wildtype cells decreased the LDLR. In Idol ( / ) cells LDLR protein was elevated, as previously reported, 30 yet was insensitive to loss of Usp2. Collectively, these results demonstrate the functional importance of the USP2 IDOL nexus in physiological control of LDLR levels. IDOL, USP2, and the LDLR Form a Functional Tri- Partite Complex to Regulate LDLR Ubiquitylation and Degradation Our results demonstrate that USP2 can control LDLR protein abundance in an IDOL-dependent manner. Yet this raises a paradox; whereas USP2 dramatically stabilizes IDOL by promoting its deubiquitylation, it also inhibits IDOL-stimulated degradation of the LDLR. Conversely, silencing of USP2 decreases IDOL protein yet enhances degradation of the LDLR. To untangle this conundrum, we set out to test the requirements for interaction between IDOL, USP2, and the LDLR, and to determine the consequence of these interactions on ubiquitylation and degradation of the receptor. First, because USP2 can reverse IDOL-mediated degradation of the LDLR, we wanted to test whether elevating the level of USP2 influences ectopic LDLR levels independent of IDOL (Figure 5A). We introduced USP2 into cells at an increasing dose but found that this had no effect on LDLR protein, unlike what we observed in the presence of IDOL (Figure 3). This suggests that for USP2 to modulate LDLR levels, ubiquitylation of the receptor is required. In the yeast-2-hybrid screen, no interaction between USP2 and the intracellular tail of the LDLR was detected (Online Figure IA). Yet, in this assay it is likely that the
7 416 Circulation Research February 5, 2016 Downloaded from by guest on July 13, 2018 Figure 4. Silencing ubiquitin-specific protease 2 (USP2) promotes low-density lipoprotein receptor (LDLR) degradation in an inducible degrader of the LDL receptor (IDOL)-dependent manner. A, HepG2 and A431 cells were cultured for 16 hours in steroldepletion medium to induce LDLR. Subsequently, cells were transfected as indicated with control (nontargeting [NT]) or USP2 sirna. Seventy-two hours after transfection, cells were collected and analyzed by immunoblotting. B, HepG2 were cultured and treated as in A and analyzed by quantitative polymerase chain reaction (n=4, ***P<0.01). C, A431 cells were cultured and treated as in A, and LDLR at the cell surface was determined by fluorescence-activated cell sorter. D, HepG2, A431, and HeLa cells were cultured and treated as in A and B. Seventy-two hours after transfection, cells were incubated for 30 minutes with 5 μg/ml Dylight 488-labeled LDL. After extensive washing, internalized LDL was quantified by measuring fluorescence in cell lysates. LDL uptake in control cells was set to 100%. Each bar and error represent the average±sd (n=4, ***P<0.001). E, Wild-type (WT) and Idol ( / ) mouse embryonic fibroblasts (MEFs) were cultured and treated as in A and B. Seventy-two hours after transfection, cells were collected and analyzed by immunoblotting. All immunoblots are representative of at least 3 independent experiments. tail is not ubiquitylated because yeast do not encode an IDOL homolog, and in this assay the tail is presented out of its normal, membrane-linked context. Therefore, we tested whether USP2 interacts with the LDLR in mammalian cells, and found that USP2 could be pulled down with the LDLR, albeit weakly (Figure 5B, lane 4). We specifically chose to use the FERM domain in these experiments because the FERM domain binds USP2, and it is recruited to, and binds the LDLR at the plasma membrane, 22 but does not stimulate degradation of the receptor because of the lack of the RING domain. As expected, the FERM domain interacted with the LDLR (Figure 5B, lane 7). Yet when all the 3 were introduced into cells, we found that both USP2 and IDOL were readily coimmunoprecipitated with the LDLR (Figure 5B, lane 8). This result supports the idea that a tri-partite complex, containing IDOL, USP2, and the LDLR is formed at the plasma membrane. To capture this complex, we used overexpression of the endocytic adaptor molecule Epsin1, which acts in a dominant negative manner to block Epsin1-dependent endocytosis. We have previously used this approach to demonstrate that it effectively blocks IDOL-stimulated degradation of the LDLR. 6 This is because of Epsin1 blocking internalization of the LDLR following IDOL-mediated ubiquitylation, thus resulting in accumulation of ubiquitylated LDLR at the plasma membrane. As Figure 5. Inducible degrader of the low-density lipoprotein receptor (IDOL), ubiquitin-specific protease 2 (USP2), and the LDLR form a functional tri-partite complex to regulate LDL receptor (LDLR) ubiquitylation and degradation. A, B, and C, HEK293T cells were transfected with (A) LDLR and increased amounts of USP2-69 expression plasmids, (B) HA-LDLR, USP2-69, and IDOL FERM expression plasmids, and (C) LDLR, Epsin, IDOL, and USP2. Forty-eight hours after transfection, total cell lysates were immunoblotted or immunoprecipitated and analyzed as indicated. All immunoblots are representative of at least 3 independent experiments. WT indicates wild-type.
8 Nelson et al USP2 Counteracts IDOL-Mediated LDLR Degradation 417 Downloaded from by guest on July 13, 2018 shown, both Epsin1 and USP2 could block IDOL-dependent degradation of the receptor (Figure 5C, lanes 1 4). The use of dominant negative Epsin1 also increased ubiquitylated LDLR species, as expected. Yet when the deubiquitylase USP2 was also present, the level of ubiquitylated LDLR was markedly reduced (Figure 5C, lane 5). Collectively, these experiments support the notion that USP2 can counteract IDOL-dependent ubiquitylation of the LDLR, and prevent the receptor from being directed toward lysosomal degradation. Discussion The LDLR provides the main entry portal for LDL into the cell, and the mechanisms characterizing its transcriptional regulation and endocytic network have been largely described. 3,4 Recently, the ubiquitin-proteasome system (UPS) was recognized as a novel regulatory layer in the LDLR pathway through the discovery of ubiquitylation-dependent degradation of the LDLR, mediated by the E3-ubiquitin ligase IDOL. 8 IDOL stimulates clathrin-independent internalization and lysosomal degradation of the LDLR, a process requiring sorting of the ubiquitylated receptor by the endosomal-sorting complex required for transport system. 6,7,13 The main finding of this study is the identification of USP2 as a deubiquitylase counteracting IDOL activity, and as a consequence abundance of the LDLR. In view of the reversible nature of ubiquitylation, we reasoned that deubiquitylases counteracting IDOL activity might exist. To investigate this, we screened IDOL and the intracellular tail of the LDLR for potential interactions with a representative collection of human deubiquitylases. The screen with the LDLR intracellular tail failed to identify any LDLRinteracting deubiquitylases. This was somewhat surprising as previous studies, ours included, reported that USP8 is essential for IDOL-mediated degradation of the LDLR, 6,7 and USP8 was included in our deubiquitylase screen. This apparent incongruence is possibly because of the fact that the cytosolic LDLR tail is only recognized by USP8 when ubiquitylated and in the context of a biological membrane, and not in the native form that is likely the one prevalent in the yeast-2-hybrid assay. However, by screening the deubiquitylase library against IDOL, we identified USP2 as a novel IDOL-binding protein. Our biochemical and molecular-based findings collectively indicate that USP2 interacts with IDOL and significantly increases its lifespan. The increased stabilization of IDOL is a result of its USP2-dependent deubiquitylation, similar to what has been described for the USP2 MDM2 and USP2-MDMX complexes. 28,32 Regulation of E3 ligases by deubiquitylasemediated deubiquitylation has been previously reported and represents an emerging concept in E3 control. 17,19 21 Previous reports indicated that deubiquitylases stabilize E3s by limiting their autoubiquitylation. Consequently, deubiquitylation of E3s acts to enhance substrate ubiquitylation. In line with this, USP2 deubiquitylates and stabilizes MDM2 and ITCH, which results in increased ubiquitylation and degradation of their cognate substrates. 28,33 In contrast to this model, deubiquitylation of IDOL by USP2 leads to a paradoxical outcome in which increased levels of the E3 ligase are mirrored by decreased degradation of its targets (ie, the LDLR and very low-density lipoprotein receptor) and functionally leads to enhanced cellular LDL uptake. In line with this, silencing of USP2 accelerates LDLR degradation and decreases LDL uptake in an IDOL-dependent manner in a variety of cell types. Our results suggest that IDOL, USP2, and the LDLR form a tri-partite complex at the plasma membrane, and in that context ubiquitylation of the receptor is decreased. The most plausible scenario to explain this outcome is that IDOL-mediated ubiquitylation of the LDLR is counteracted by subsequent removal of ubiquitin chains from the LDLR by USP2. In this sequential scenario, USP2 is recruited by IDOL and acts directly on the receptor, reminiscent of the reported effect of USP2 on EGFR after ligand binding. 34 This would be consistent with the fact that the phenotype is IDOL dependent, and that USP2 is able to process diverse ubiquitin linkages. 35,36 Although we think this is the probable scenario, our experiments cannot formally rule out a more provocative possibility, namely that ubiquitylation of IDOL itself serves to both regulate its stability (ie, autoubiquitylation) and activity (ie, LDLR ubiquitylation). As such, by removing ubiquitin from IDOL, USP2 could counteract both processes. Although speculative, this form of regulation is not without precedent Both cis and trans ubiquitylation of E3 ligases has been reported, and we have observed that USP2 can remove ubiquitin chains from an IDOL mutant lacking autoubiquitylation activity (data not shown). The significance of this observation requires further investigation, yet it is possible that these modifications may allow a hierarchical E3-based network for fine tuning of cellular proteostasis. 19,40 The role of USP2 in regulating the IDOL LDLR pathway seems to be exclusively dependent on post-transcriptional events because USP2 expression is unaffected by perturbing the cellular sterol content or by pharmacologically activating LXR in different cell types. Therefore, identifying the signals, or post-transcriptional modifications on USP2 or IDOL that regulate their functional interaction will be an important goal in future studies. The USP2 gene is alternatively spliced into 3 isoforms, 27 all sharing an identical C-terminal catalytic core. The 2 major USP2 isoforms, USP2-69 and USP2-45, promote IDOL deubiquitylation and rescue LDLR degradation. The specific functions and regulation of these 2 distinct isoforms are not well understood. The USP2-69 isoform is induced by androgens both in human and in rat prostate cells 41 and by the AKT mtorc1 axis in hepatocellular carcinoma. 42 The mouse Usp2-45 isoform is regulated by the transcriptional coactivator PGC-1α, which couples the circadian and nutritional control in the liver in response to starvation. 43 The above-mentioned regulation suggests that expression of USP2 is sensitive to nutritional and circadian input. Given that cholesterol metabolism is subject to these cues as well, it will be important to investigate how USP2 is integrated with the IDOL LDLR axis in vivo. We speculate that the functional interaction between USP2 and IDOL, and the formation of a tri-partite complex with the LDLR, may be sensitive to post-transcriptional modifications, nutritional cues, or to the dynamic levels of these proteins. However, a serious limitation in experimentally testing the physiological role of the USP2 IDOL nexus is the species-dependent differences in hepatic IDOL activity and transcriptional regulation 12 ; whereas in human and in nonhuman primates, the IDOL pathway is highly active in
9 418 Circulation Research February 5, 2016 Downloaded from by guest on July 13, 2018 hepatocytes and subject to strong sterol-dependent transcriptional regulation, in rodent hepatocytes it is not. Accordingly, Idol ( / ) mice do not display a marked cholesterol phenotype or changes in hepatic LDLR content. We therefore anticipate that a functional USP2 IDOL interaction will be predominantly active in human hepatocytes. In that respect, the growing body of evidence pointing toward genetic variation in IDOL as a modifier of LDL cholesterol in humans, 9 11 warrants further studies to investigate the basic mechanisms governing IDOL activity and regulation by USP2. In summary, this study reports the identification of the deubiquitylase USP2 as an endogenous inhibitor of IDOL and a novel modulator of lipoprotein uptake owing to its ability to counteract IDOL-stimulated degradation of the LDLR (graphically summarized in Online Figure VIII). This finding further supports the notion that inhibition of IDOL activity would result in increased LDLR abundance and enhance LDL uptake, an approach that can be exploited as a therapeutic strategy to treat dyslipidemia and coronary artery disease. Acknowledgments We acknowledge members of the Zelcer group, Menno de Winter and Irith Koster for their comments and suggestions. Sources of Funding B. Distel is supported by a TOP grant ( ) from the Netherlands Organization of Scientific Research (NWO). N. Zelcer is an Established Investigator of the Dutch Heart Foundation (2013T111), and he is supported by a VIDI grant ( ) from the Netherlands Organization of Scientific Research (NWO), and by a European Research Council (ERC) Consolidator grant (617376) from the European Research Council. None. Disclosures References 1. Hobbs HH, Russell DW, Brown MS, Goldstein JL. The LDL receptor locus in familial hypercholesterolemia: mutational analysis of a membrane protein. Annu Rev Genet. 1990;24: doi: /annurev. ge Tolleshaug H, Goldstein JL, Schneider WJ, Brown MS. Posttranslational processing of the LDL receptor and its genetic disruption in familial hypercholesterolemia. Cell. 1982;30: Brown MS, Goldstein JL. A receptor-mediated pathway for cholesterol homeostasis. Science. 1986;232: Horton JD, Goldstein JL, Brown MS. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest. 2002;109: doi: /JCI Goldstein JL, Brown MS. The LDL receptor. Arterioscler Thromb Vasc Biol. 2009;29: doi: /ATVBAHA Sorrentino V, Nelson JK, Maspero E, Marques AR, Scheer L, Polo S, Zelcer N. The LXR-IDOL axis defines a clathrin-, caveolae-, and dynamin-independent endocytic route for LDLR internalization and lysosomal degradation. J Lipid Res. 2013;54: doi: /jlr.M Scotti E, Calamai M, Goulbourne CN, Zhang L, Hong C, Lin RR, Choi J, Pilch PF, Fong LG, Zou P, Ting AY, Pavone FS, Young SG, Tontonoz P. IDOL stimulates clathrin-independent endocytosis and multivesicular body-mediated lysosomal degradation of the low-density lipoprotein receptor. Mol Cell Biol. 2013;33: doi: /MCB Zelcer N, Hong C, Boyadjian R, Tontonoz P. LXR regulates cholesterol uptake through Idol-dependent ubiquitination of the LDL receptor. Science. 2009;325: doi: /science Teslovich TM, Musunuru K, Smith AV, et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature. 2010;466: doi: /nature Chasman DI, Paré G, Mora S, Hopewell JC, Peloso G, Clarke R, Cupples LA, Hamsten A, Kathiresan S, Mälarstig A, Ordovas JM, Ripatti S, Parker AN, Miletich JP, Ridker PM. Forty-three loci associated with plasma lipoprotein size, concentration, and cholesterol content in genome-wide analysis. PLoS Genet. 2009;5:e doi: /journal.pgen Sorrentino V, Fouchier SW, Motazacker MM, Nelson JK, Defesche JC, Dallinga-Thie GM, Kastelein JJ, Kees Hovingh G, Zelcer N. Identification of a loss-of-function inducible degrader of the low-density lipoprotein receptor variant in individuals with low circulating low-density lipoprotein. Eur Heart J. 2013;34: doi: /eurheartj/ehs Hong C, Marshall SM, McDaniel AL, et al. The LXR-Idol axis differentially regulates plasma LDL levels in primates and mice. Cell Metab. 2014;20: doi: /j.cmet Sorrentino V, Zelcer N. Post-transcriptional regulation of lipoprotein receptors by the E3-ubiquitin ligase inducible degrader of the low-density lipoprotein receptor. Curr Opin Lipidol. 2012;23: doi: / MOL.0b013e Sharpe LJ, Cook EC, Zelcer N, Brown AJ. The UPS and downs of cholesterol homeostasis. Trends Biochem Sci. 2014;39: doi: /j. tibs Reyes-Turcu FE, Ventii KH, Wilkinson KD. Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu Rev Biochem. 2009;78: doi: /annurev.biochem Sowa ME, Bennett EJ, Gygi SP, Harper JW. Defining the human deubiquitinating enzyme interaction landscape. Cell. 2009;138: doi: /j.cell Komander D, Clague MJ, Urbé S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 2009;10: doi: /nrm Ventii KH, Wilkinson KD. Protein partners of deubiquitinating enzymes. Biochem J. 2008;414: doi: /BJ de Bie P, Ciechanover A. Ubiquitination of E3 ligases: self-regulation of the ubiquitin system via proteolytic and non-proteolytic mechanisms. Cell Death Differ. 2011;18: doi: /cdd Hayes SD, Liu H, MacDonald E, Sanderson CM, Coulson JM, Clague MJ, Urbé S. Direct and indirect control of mitogen-activated protein kinase pathway-associated components, BRAP/IMP E3 ubiquitin ligase and CRAF/RAF1 kinase, by the deubiquitylating enzyme USP15. J Biol Chem. 2012;287: doi: /jbc.M Clague MJ, Coulson JM, Urbé S. Cellular functions of the DUBs. J Cell Sci. 2012;125: doi: /jcs Sorrentino V, Scheer L, Santos A, Reits E, Bleijlevens B, Zelcer N. Distinct functional domains contribute to degradation of the low density lipoprotein receptor (LDLR) by the E3 ubiquitin ligase inducible Degrader of the LDLR (IDOL). J Biol Chem. 2011;286: doi: / jbc.m Shan J, Zhao W, Gu W. Suppression of cancer cell growth by promoting cyclin D1 degradation. Mol Cell. 2009;36: doi: /j. molcel Oh KH, Yang SW, Park JM, Seol JH, Iemura S, Natsume T, Murata S, Tanaka K, Jeon YJ, Chung CH. Control of AIF-mediated cell death by antagonistic functions of CHIP ubiquitin E3 ligase and USP2 deubiquitinating enzyme. Cell Death Differ. 2011;18: doi: / cdd Motazacker MM, Pirruccello J, Huijgen R, Do R, Gabriel S, Peter J, Kuivenhoven JA, Defesche JC, Kastelein JJ, Hovingh GK, Zelcer N, Kathiresan S, Fouchier SW. Advances in genetics show the need for extending screening strategies for autosomal dominant hypercholesterolaemia. Eur Heart J. 2012;33: doi: /eurheartj/ehs Zhang L, Fairall L, Goult BT, Calkin AC, Hong C, Millard CJ, Tontonoz P, Schwabe JW. The IDOL-UBE2D complex mediates sterol-dependent degradation of the LDL receptor. Genes Dev. 2011;25: doi: /gad Gousseva N, Baker RT. Gene structure, alternate splicing, tissue distribution, cellular localization, and developmental expression pattern of mouse deubiquitinating enzyme isoforms Usp2-45 and Usp2-69. Gene Expr. 2003;11: Stevenson LF, Sparks A, Allende-Vega N, Xirodimas DP, Lane DP, Saville MK. The deubiquitinating enzyme USP2a regulates the p53 pathway by targeting Mdm2. EMBO J. 2007;26: doi: /sj.emboj Hong C, Duit S, Jalonen P, Out R, Scheer L, Sorrentino V, Boyadjian R, Rodenburg KW, Foley E, Korhonen L, Lindholm D, Nimpf J, van Berkel TJ, Tontonoz P, Zelcer N. The E3 ubiquitin ligase IDOL induces the degradation of the low density lipoprotein receptor family members VLDLR and ApoER2. J Biol Chem. 2010;285: doi: /jbc.M
10 Nelson et al USP2 Counteracts IDOL-Mediated LDLR Degradation 419 Downloaded from by guest on July 13, Scotti E, Hong C, Yoshinaga Y, Tu Y, Hu Y, Zelcer N, Boyadjian R, de Jong PJ, Young SG, Fong LG, Tontonoz P. Targeted disruption of the idol gene alters cellular regulation of the low-density lipoprotein receptor by sterols and liver x receptor agonists. Mol Cell Biol. 2011;31: doi: /MCB Seidah NG. PCSK9 as a therapeutic target of dyslipidemia. Expert Opin Ther Targets. 2009;13: doi: / Allende-Vega N, Sparks A, Lane DP, Saville MK. MdmX is a substrate for the deubiquitinating enzyme USP2a. Oncogene. 2010;29: doi: /onc Haimerl F, Erhardt A, Sass G, Tiegs G. Down-regulation of the de-ubiquitinating enzyme ubiquitin-specific protease 2 contributes to tumor necrosis factor-alpha-induced hepatocyte survival. J Biol Chem. 2009;284: doi: /jbc.M Liu Z, Zanata SM, Kim J, Peterson MA, Di Vizio D, Chirieac LR, Pyne S, Agostini M, Freeman MR, Loda M. The ubiquitin-specific protease USP2a prevents endocytosis-mediated EGFR degradation. Oncogene. 2013;32: doi: /onc Komander D, Reyes-Turcu F, Licchesi JD, Odenwaelder P, Wilkinson KD, Barford D. Molecular discrimination of structurally equivalent Lys 63-linked and linear polyubiquitin chains. EMBO Rep. 2009;10: doi: /embor Virdee S, Ye Y, Nguyen DP, Komander D, Chin JW. Engineered diubiquitin synthesis reveals Lys29-isopeptide specificity of an OTU deubiquitinase. Nat Chem Biol. 2010;6: doi: / nchembio Zaaroor-Regev D, de Bie P, Scheffner M, Noy T, Shemer R, Heled M, Stein I, Pikarsky E, Ciechanover A. Regulation of the polycomb protein What Is Known? The E3-ubiquitin ligase inducible degrader of the low-density lipoprotein (LDL) receptor (IDOL) promotes ubiquitylation and subsequent degradation of the low-density lipoprotein receptor (LDLR), and thereby modulates LDL metabolism. Common and rare genetic variation in IDOL associates with the circulating level of LDL-cholesterol in humans. What New Information Does This Article Contribute? The deubiquitylase ubiquitin-specific protease 2 (USP2) interacts with IDOL. USP2 deubiquitylates IDOL leading to its stabilization, which paradoxically attenuates degradation of the LDLR by IDOL. USP2, IDOL, and the LDLR form a tri-partite complex in which USP2 can deubiquitylate the LDLR and prevent its degradation. Novelty and Significance Ring1B by self-ubiquitination or by E6-AP may have implications to the pathogenesis of Angelman syndrome. Proc Natl Acad Sci U S A. 2010;107: doi: /pnas Herman-Bachinsky Y, Ryoo HD, Ciechanover A, Gonen H. Regulation of the Drosophila ubiquitin ligase DIAP1 is mediated via several distinct ubiquitin system pathways. Cell Death Differ. 2007;14: doi: /sj.cdd Silke J, Kratina T, Chu D, Ekert PG, Day CL, Pakusch M, Huang DC, Vaux DL. Determination of cell survival by RING-mediated regulation of inhibitor of apoptosis (IAP) protein abundance. Proc Natl Acad Sci U S A. 2005;102: doi: /pnas Weissman AM, Shabek N, Ciechanover A. The predator becomes the prey: regulating the ubiquitin system by ubiquitylation and degradation. Nat Rev Mol Cell Biol. 2011;12: doi: /nrm Graner E, Tang D, Rossi S, Baron A, Migita T, Weinstein LJ, Lechpammer M, Huesken D, Zimmermann J, Signoretti S, Loda M. The isopeptidase USP2a regulates the stability of fatty acid synthase in prostate cancer. Cancer Cell. 2004;5: Calvisi DF, Wang C, Ho C, Ladu S, Lee SA, Mattu S, Destefanis G, Delogu S, Zimmermann A, Ericsson J, Brozzetti S, Staniscia T, Chen X, Dombrowski F, Evert M. Increased lipogenesis, induced by AKTmTORC1-RPS6 signaling, promotes development of human hepatocellular carcinoma. Gastroenterology. 2011;140: doi: /j. gastro Molusky MM, Li S, Ma D, Yu L, Lin JD. Ubiquitin-specific protease 2 regulates hepatic gluconeogenesis and diurnal glucose metabolism through 11β-hydroxysteroid dehydrogenase 1. Diabetes. 2012;61: doi: /db The E3 ubiquitin ligase IDOL promotes ubiquitylation and subsequent degradation of the LDLR. As such, IDOL activity represents an acute mechanism to shut down uptake of LDL-derived cholesterol into cells. Yet IDOL also promotes its own ubiquitylation and degradation and is, therefore, a highly unstable protein. Using a genetic approach, we identified the deubiquitylase USP2 as an IDOL-interacting partner. We found that USP2 deubiquitylates IDOL resulting in increased IDOL stability. However, despite elevated levels of IDOL degradation of the LDLR is paradoxically attenuated. Conversely, silencing of USP2 expression enhances IDOL-dependent degradation of the LDLR. Underlying these observations, we demonstrate that IDOL, USP2, and the LDLR can form a tri-partite complex in which USP2 can deubiquitylate the LDLR. As such, our study identifies USP2 as a new regulator of LDL metabolism owing to its ability to counteract IDOL-mediated degradation of the LDLR. Furthermore, this finding emphasizes the important role of the ubiquitin-proteasomal system in lipoprotein metabolism and cardiovascular disease.
11 Downloaded from by guest on July 13, 2018 The Deubiquitylase USP2 Regulates the LDLR Pathway by Counteracting the E3-Ubiquitin Ligase IDOL Jessica Kristine Nelson, Vincenzo Sorrentino, Rossella Avagliano Trezza, Claire Heride, Sylvie Urbe, Ben Distel and Noam Zelcer Circ Res. 2016;118: ; originally published online December 14, 2015; doi: /CIRCRESAHA Circulation Research is published by the American Heart Association, 7272 Greenville Avenue, Dallas, TX Copyright 2015 American Heart Association, Inc. All rights reserved. Print ISSN: Online ISSN: The online version of this article, along with updated information and services, is located on the World Wide Web at: Data Supplement (unedited) at: Permissions: Requests for permissions to reproduce figures, tables, or portions of articles originally published in Circulation Research can be obtained via RightsLink, a service of the Copyright Clearance Center, not the Editorial Office. Once the online version of the published article for which permission is being requested is located, click Request Permissions in the middle column of the Web page under Services. Further information about this process is available in the Permissions and Rights Question and Answer document. Reprints: Information about reprints can be found online at: Subscriptions: Information about subscribing to Circulation Research is online at:
12 The deubiquitylase USP2 regulates the LDLR pathway by counteracting the E3-ubiquitin ligase IDOL Jessica Kristine Nelson, MSc 1,3, Vincenzo Sorrentino, PhD 1,3, Rossella Avagliano Trezza, MSc 1, Claire Heride, PhD 2, Sylvie Urbe, PhD 2, Ben Distel, PhD 1, and Noam Zelcer, PhD 1,4 1 Department of Medical Biochemistry, Academic Medical Center of the University of Amsterdam, Amsterdam, 1105AZ, The Netherlands, 2 Department of Cellular and Molecular Physiology, University of Liverpool, Liverpool, L69 3BX, United Kingdom SUPPLEMENTAL MATERIAL 1
13 Materials and methods Reagents Simvastatin and the proteasome inhibitor MG-132 were from Calbiochem. Lipoprotein deficient serum (LPDS) was prepared following standard procedures and confirmed to contain no lipoproteins (not shown). GW3965, and Bafilomycin A1 were from Sigma. Cell culture and transfections HEK293T, HeLa, Huh7, and HepG2 cells were obtained from the ATCC. Cells were maintained in DMEM supplemented with 10% FBS at 37 C and 5% CO 2. A431 cells were a kind gift from Drs. Vilja Pietiainen and Elina Ikonen (University of Helsinki). Wildtype and Idol (-/-) MEFs were a kind gift from Dr. Peter Tontonoz (University of California at Los Angeles). DU145 were from Dr. Koen van de Wetering (NKI, Amsterdam). Where indicated, cells were sterol-depleted by culture in sterol-depletion medium (DMEM supplemented by 10% LPDS, 5 μg/ml simvastatin, and 100 μm mevalonate) or treated with 1 μm GW3965 to induce LXR signaling. HEK293T and HepG2 cells were transfected with the indicated amounts of plasmids using the JetPrime reagent. In transfection experiments the efficiency of transfection was monitored by co-transfection of an expression plasmid for GFP and was consistently >85% in HEK293T and HepG2 cells. Plasmids and expression constructs Expression plasmids encoding IDOL, IDOL-FERM, LDLR, USP8, VLDLR, GFP, Ubiquitin, and Epsin1 were previously reported 1-3. Expression plasmids encoding USP2-69 and USP2-45 cdnas were generated with gateway-mediated recombination (Invitrogen). In experiments where only expression constructs for USP2-69 are used, USP2-69 is indicated just as USP2. The activity disabling C276A 4,5 mutation in USP2 was introduced using the QuikChange site-directed mutagenesis kit. All plasmids used in this study experiments were isolated by CsCl 2 gradient centrifugation and their correctness verified by sequencing. Antibodies and immunoblot Analysis Total cell lysates were prepared in RIPA buffer (150 mm NaCl, 1% Nonidet P-40, 0.1% sodium deoxycholate, 0.1% SDS, 100 mm Tris-HCl, ph 7.4) supplemented with protease inhibitors. Lysates were cleared by centrifugation at 4 C for 10 min at 10,000 g. Protein concentration was determined using the Bradford assay with BSA as reference. Samples (10 40μg) were separated on NuPAGE BisTris gels and transferred to nitrocellulose. Membranes were probed with the following antibodies: LDLR (Epitomics, clone EP1553Y 1:4000), tubulin (Sigma, clone DM1A, 1:5000), FLAG-HRP (Sigma, clone M2, 1:1000), GFP (affinity purified rabbit polyclonal anti-gfp was from Dr. Mireille Riedinger, UCLA, 1:5000, or Santa Cruz, clone B-2, 1:3000), MYC (Cell Signaling, clone 9B11, 1:4000), HA (Covance, clone 16B12, 1:10000), Ubiquitin (ENZO, clone FK2, 1:1000), Epsin1 (rabbit polyclonal serum raised against amino acids of rat Epsin1, 1:5), His 6 (Rockland, 1:1000), EGFR (affinity purified rabbit polyclonal anti-egfr sera was from Dr. Simona Polo, 1:40,000). Secondary HRP-conjugated antibodies (Zymed Laboratories Inc.) were used and visualized with chemiluminescence on a Fuji LAS4000 (GE Healthcare). All immunoblots are representative of at least three independent experiments. LDL uptake assay DyLight 488-labeled LDL was produced as previously described 6. Briefly, HepG2, HeLa, or A431 cells were incubated in sterol-depletion medium for 16 h prior to addition of LDL to increase LDLR abundance. To initiate LDL uptake, cells were washed twice with PBS and 2
14 incubated with 5 μg/ml DyLight488-labeled LDL in DMEM supplemented with 0.5% BSA for the indicated time at 37 C. Subsequently, cells were washed twice with PBS supplemented with excess sterols (PBS with 10% FCS), followed by an additional wash with PBS. Endocytosis of LDL was measured by lysing cells in RIPA buffer and quantification of the fluorescence signal on a Typhoon imager. Relative LDL uptake was calculated from the measure of fluorescence values corrected for specific uptake (cells treated with labeled LDL in the presence of an excess of non-labeled LDL) and presented as mean ± SD. Measurement of cell-surface LDLR Cells were treated as indicated, dissociated with TrypLE Express and incubated in FACS blocking buffer (FACS buffer containing 2% goat serum) for 30 min on ice. Subsequently, 100,000 cells were stained in 50μL FACS buffer with a R-Phycoerythrin (PE)-conjugated mouse anti-human LDLR antibody (R&D Systems, #FAB2148P, 1:40) for 1 h on ice. Cells were subsequently washed three times with FACS buffer and directly analyzed on a FACSCalibur Flow Cytometer (BD Biosciences). Viable cells were gated and 10,000 events per condition acquired. Data were analyzed using the CellQuest software package. Relative surface LDLR was calculated from geomean values corrected for background (non-stained cells), and presented as mean ± SD. sirna transfection To knockdown USP2 expression we used the ON-TARGETplus Human USP2 SMARTpool (Dharmacon: cat#l ), or the individual ON-TARGETplus Human USP2 sirnas (Dharmacon: cat#j , J , J ). As control we used the ON- TARGETplus Non-targeting Pool (Dharmacon: cat#d ). A431and HeLa cells were transfected with control or USP2 sirnas (final concentration 30 nm) using the JetPrime reagent. HepG2 and DU145 cells were transfected similarly, but with the RNAiMAX reagent. The efficiency of USP2 silencing was assessed by quantitative PCR. Unless otherwise indicated, sirna transfection experiments were conducted for 48 hours. RNA isolation and quantitative PCR Total RNA was isolated from cells using TRIzol and one microgram of total RNA was reverse-transcribed using the iscript reverse transcription reagent. SYBR Green real-time quantitative PCR assays were performed on a Lightcycler 480 II apparatus. Oligonucleotide sequences are available upon request. Yeast two-hybrid screen S. cerevisiae strains used in this study are PJ69-4a (trp1-901 leu2-3,112 ura3-52 his3-200 gal4d gal80d LYS2::GAL1-HIS3 GAL2-ADE2 met2::gal7-lacz) and PJ69-4α (trp1-901 leu2-3,112 ura3-52 his3-200 gal4d gal80d LYS2::GAL1-HIS3 GAL2-ADE2 met2::gal7- lacz). Cells were grown at 28 0 C in YPAD (2% glucose, 2% peptone, 1% yeast extract and 4 mg/ml adenine) or in minimal glucose medium containing 0.67% Yeast Nitrogen Base (YNB) without amino acids, 2% glucose and amino acids as needed (2 mg/ml uracil, 3 mg/ml leucine, 2 mg/ml tryptophane, 2 mg/ml histidine, 4 mg/ml adenine, 2 mg/ml methionine). The yeast-two-hybrid screening was performed as following. PJ69-4α transformed with human IDOL constructs or LDLR intracellular tail (residues ) cloned into pact2-dest, and PJ69-4a transformed with the bait constructs cloned into pgdbu-dest were selected on minimal agar plates lacking leucine or uracil, respectively. For mating, cells were precultured for 16h in selection medium, and subsequently 2x10 6 of transformed PJ69-4a and PJ69-4α cells (1:1 ratio) were mated in 2xYPAD (4% glucose, 4% peptone, 2% yeast extract and 8 mg/ml adenine) for 24 hrs with slow shaking (50 rpm). Diploids were selected on minimal glucose plates lacking both leucine and uracil. To reconfirm interactions we used a spot assay. Briefly, diploid cells were incubated for 16 hrs in minimal glucose medium lacking leucine and uracil, diluted to OD 600 of 0.2 and grown till an OD 600 of 1. Cells were 3
15 then washed with sterile water and serially diluted so as to plate cells/spot on minimal glucose agar lacking leucine and uracil, and with or without histidine, mm 3- aminotriazol (3-AT), and adenine. Plates were incubated for 3 days before analysis. Statistical Analysis Data are presented as mean ± SD. One-way ANOVA followed by the Bonferroni correction was performed for comparison of more than two groups. Student s t-test was performed for comparison of two groups. P values of <0.05 were considered significant. References 1. Zelcer N, Hong C, Boyadjian R, Tontonoz P. LXR regulates cholesterol uptake through Idoldependent ubiquitination of the LDL receptor. Science. 2009;325: Sorrentino V, Nelson JK, Maspero E, Marques ARA, Scheer L, Polo S, Zelcer N. The LXR- IDOL axis defines a clathrin-, caveolae-, and dynamin-independent endocytic route for LDLR internalization and lysosomal degradation. J Lipid Res. 2013;54: Sorrentino V, Scheer L, Santos A, Reits E, Bleijlevens B, Zelcer N. Distinct functional domains contribute to degradation of the low density lipoprotein receptor (LDLR) by the E3 ubiquitin ligase inducible Degrader of the LDLR (IDOL). J Biol Chem. 2011;286: Shan J, Zhao W, Gu W. Suppression of cancer cell growth by promoting cyclin D1 degradation. Molecular Cell. 2009;36: Oh KH, Yang SW, Park JM, Seol JH, Iemura S, Natsume T, Murata S, Tanaka K, Jeon YJ, Chung CH. Control of AIF-mediated cell death by antagonistic functions of CHIP ubiquitin E3 ligase and USP2 deubiquitinating enzyme. Cell Death Differ. 2011;18: Motazacker MM, Pirruccello J, Huijgen R, et al. Advances in genetics show the need for extending screening strategies for autosomal dominant hypercholesterolaemia. Eur Heart J. 2012;33:
16 Online figure legends Online figure I. Summary of IDOL-DUB yeast-two-hybrid screen. Y2H assay for (A) USP2-LDLR IC and (B) IDOL-USP2. The corresponding IDOL constructs that were used are illustrated in Fig. 1A. The intracellular tail of the LDLR or the different IDOL fragments and USP2 were cloned into yeast-two-hybrid prey and bait plasmids, respectively, and transformed into yeast. Haploid cells were mated, and diploid cells were serially diluted and plated on selective plates. The assay was performed as described in the Methods section, and decreasing amounts of cells are spotted and selected as indicated. Online figure II. The DUB USP2 is a novel binding partner of IDOL. (A,B) HEK293T cells were transfected with 1:1 ratio of the indicated constructs for IDOL and the USP2 wildtype or catalytic mutant. Total cell lysates were immunoblotted or immunoprecipitated and analyzed as indicated. (C) Schematic representation of UPS2 functional domains. USP2, also known as USP2-69, has a N-terminal extension, involved in contacts with other proteins, and a C-terminal catalytic domain. Mutation of the key cysteine C276 residue in this domain impairs USP2 s DUB activity. USP2-45 is a shorter isoform of USP2, with a partial truncation of its N-terminal domain, but intact catalytic domain. This isoform also interacts with IDOL, as shown in Fig. 1D. Online figure III. USP2 deubiquitylates and stabilizes IDOL. HEK293T cells were transfected with the indicated IDOL and USP2 expression vectors, with a ratio of IDOL:USP2 of 1:5 (left panel) or 1:10 (right panel). 48 hours after transfection, cells were incubated for the indicated times with 10 μg/ml cycloheximide (CHX), and cells harvested for immunoblots and kinetic analysis as shown in Fig. 2C. Online figure IV. Silencing USP2 reduces the level of ectopic IDOL protein. (A) HepG2 and (B) Huh7 cells were co-transfected with IDOL and GFP expression plasmids, and either Control (Non-targeting, NT) or USP2 sirna. 48 hours later total cell lysates were collected and immunoblotted as indicated. A representative immunoblot of three independent experiments is shown. (A,B) the intensity of IDOL protein was quantified and normalized to that of GFP. The relative IDOL protein level is plotted. Each bar and error represent the mean ± SD (N=3). * p < 0.05 Online figure V. USP2 attenuates IDOL and LXR-induced degradation of the LDLR. (A) HEK293T cells were transfected with LDLR and with IDOL and USP2-45 (1:7 ratio). 48 hours after transfection, cells were collected and immunoblotted as indicated. (B,C) HeLa cells were transfected with the indicated USP2 expression vector. 48 hours after transfection, cells were treated with 1 μm GW3965 (GW) for 4 hours to activate the LXR pathway and subsequently harvested for (B) immunoblots and (C) qpcr analysis as shown (n=4, *** p< 0.001). Expression of USP2 does not affect LXR activation, as shown by the increased expression of IDOL and ABCA1. (D) HepG2 cells were cultured in complete medium or in sterol-depletion medium for 16 hours, and subsequently treated with vehicle or 1 μm GW3965 (GW) for 4 hours. The mrna levels of USP2, LDLR and IDOL were analyzed as indicated. Expression of USP2 is not affected by LXR activation or by variations in the cellular sterol content (n=5, *** p< ). Online figure VI. USP2 silencing decreases LDLR protein levels in different cell lines. (A) Relative IDOL expression in the indicated cell lines was determined by qpcr. Each bar and error represent the mean ± SD of three independent experiments (B) HeLa and DU145 cells were transfected as indicated with 30 nm of control (Non-targeting, NT) or USP2 sirna. 48 hours after transfection cells were shifted to sterol-depletion medium for an additional 24 hours after which total cell lysates were collected and analyzed by immunoblotting. (C,D) A431 and DU145 cells were cultured and treated as in (B) and 5
17 analyzed by qpcr to assess LDLR, IDOL and USP2 mrna levels (n=4, *p<0.05). (E) Wildtype and Idol (-/-) MEFs were cultured and treated as in Fig. 4E. 72 hours after transfection, cells were collected and analyzed by qpcr as indicated (n=4, *** p<0.0001). All immunoblots are representative of at least three independent experiments. Online figure VII. USP2 silencing decreases LDLR protein levels (A,B) The specificity of USP2 silencing in HepG2 cells was tested by transfecting cells as indicated with 30 nm of a control (Non-targeting, NT), three individual USP2 sirnas, or with a pooled set of USP2 sirnas. 48 hours after transfection cells were shifted to sterol depletion medium for an additional 24 hours after which cells were collected and total cell lysates were analyzed by immunoblotting (B) or by qpcr (A) to assess the efficiency of USP2 mrna silencing (n=3, ***p<0.001). Online figure VIII. Graphical scheme for the IDOL-USP2 nexus in regulation of the LDLR. (A) IDOL expression is induced by activation of the sterol-sensing nuclear receptors LXRs. This leads to enhanced ubiquitylation of the LDLR, which targets the receptor for lysosomal degradation. In parallel, auto-ubiquitylation of IDOL leads to its proteasomal degradation. (B) USP2 is recruited together with IDOL to the intracellular tail of the LDLR. There, USP2 can counteract IDOL-mediated ubiquitylation of the LDLR by removing these chains from the receptor. 6
18 Online table I PREY BAIT Diploids IDOL IDOL Ring IDOL IDOL Ring UCHL1 OTUB2 USP11 EIF3S3 + USP28 USP19 USP48B USP38 OTUD6A * CEZANNE ATXN3L EIF3S5 UCHL3 BAP1 USP12 USP4 USP30 USP20 USP49 USP39 OTUD5 VCPIP1 JOSD1 PSMD7 UCHL5 USP15 USP13 * * USP5 USP30 USP41B BRCC3 OTUD4 JOSD2 PSMD14 BAP1 AMSH * * USP14 USP6 USP32 + USP44 USP54 YOD1 CSN5 USP7 UBC13 USP25 * USP1 USP45 USP33 TNFAIP3 OTUB1 AMSH-LP CSN6 USP8 FLJ14981 USP26 USP USP46 USP16 OTUD6B USP36 Online table I: Identification of USP2 as a novel IDOL-binding partner by the Y2H assay The full length IDOL, or its RING domain, was screened against a library of human DUBs, comprehensive of the listed enzymes. The assay was performed as described in the Methods section. False positives are indicated by the asterisk, and they represent DUBs whose coexpression in the yeast system with the empty pact2-dest vector resulted in selective cell growth even in the absence of IDOL or LDLR. Only yeast diploids co-expressing full length IDOL and USP2-69 showed marked growth on both selective media (+++), and were considered for validation analysis. A similar screen was performed using the LDLR intracellular tail, however it did not identify any potential LDLR-interacting DUBs.
19 Online figure I A His+ His- 10mM 3AT His- 20mM 3AT His- Ade- Spotted cells: B BAIT USP2 PREY pact2 His+ His- His- 10mM 3AT His- 20mM 3AT Ade- USP2 IDOL USP2 IDOL C837A USP2 RING USP2 FERM USP2 ΔF1 USP2 ΔF2 USP2 USP2 F3+linker F3 - pact2 - IDOL - IDOL C837A - RING - FERM - ΔF1 - ΔF2 - - F3+linker F3 Spotted cells:
20 Online figure II A IDOL: B IP: MYC (IDOL) USP2: IB: FLAG-USP2 IB: MYC-IDOL IgG IP: FLAG (USP2) USP2: IDOL: IB: MYC-IDOL IB: FLAG-USP2 IgG MYC-IDOL 49 MYC-IDOL 49 Input FLAG-USP2 TUBULIN 64 Input FLAG-USP2 TUBULIN 64 GFP GFP C N-terminal extension Isopeptidase domain C276A USP2-69 USP2-45
21 Online figure III IDOL WT :USP2 1: CHX (min): IDOL:USP2 1: MYC-USP2 64 MYC-USP2 64 FLAG-IDOL WT 49 FLAG-IDOL WT 49 FLAG-IDOL WT TUBULIN 49 FLAG-IDOL WT TUBULIN 49 GFP GFP
22 Online figure IV A +sirna: FLAG-IDOL TUBULIN GFP HepG2 NT USP2 49 Relative IDOL protein sint siusp2 * B HUH7 +sirna: sint USP2 FLAG-IDOL TUBULIN GFP 49 Relative IDOL protein sint siusp2 **
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
Regulation of cholesterol metabolism: An IDOL-dependent pathway to degrade the LDL-receptor Sorrentino, V.
 UvA-DARE (Digital Academic Repository) Regulation of cholesterol metabolism: An IDOL-dependent pathway to degrade the LDL-receptor Sorrentino, V. Link to publication Citation for published version (APA):
UvA-DARE (Digital Academic Repository) Regulation of cholesterol metabolism: An IDOL-dependent pathway to degrade the LDL-receptor Sorrentino, V. Link to publication Citation for published version (APA):
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication
 Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
Cholesterol is essential for mammalian cells because it plays
 Brief Review Feedback Regulation of Cholesterol Uptake by the LXR IDOL LDLR Axis Li Zhang, Karen Reue, Loren G. Fong, Stephen G. Young, Peter Tontonoz Abstract Inducible degrader of the low-density lipoprotein
Brief Review Feedback Regulation of Cholesterol Uptake by the LXR IDOL LDLR Axis Li Zhang, Karen Reue, Loren G. Fong, Stephen G. Young, Peter Tontonoz Abstract Inducible degrader of the low-density lipoprotein
PCSK9 and its Role in LDL Receptor Regulation Muscat, Oman - 9 February 2019
 PCSK9 and its Role in LDL Receptor Regulation Muscat, Oman - 9 February 2019 Professor Gilles Lambert, PhD LaboratoireInserm U1188 Universitéde la Réunion Faculté de Médecine Saint Denis de la Réunion,
PCSK9 and its Role in LDL Receptor Regulation Muscat, Oman - 9 February 2019 Professor Gilles Lambert, PhD LaboratoireInserm U1188 Universitéde la Réunion Faculté de Médecine Saint Denis de la Réunion,
Post-transcriptional regulation of lipoprotein receptors by the E3-ubiquitin ligase inducible degrader of the low-density lipoprotein receptor
 REVIEW C URRENT OPINION Post-transcriptional regulation of lipoprotein receptors by the E3-ubiquitin ligase inducible degrader of the low-density lipoprotein receptor Vincenzo Sorrentino and Noam Zelcer
REVIEW C URRENT OPINION Post-transcriptional regulation of lipoprotein receptors by the E3-ubiquitin ligase inducible degrader of the low-density lipoprotein receptor Vincenzo Sorrentino and Noam Zelcer
LDL Uptake Cell-Based Assay Kit
 LDL Uptake Cell-Based Assay Kit Catalog Number KA1327 100 assays Version: 07 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
LDL Uptake Cell-Based Assay Kit Catalog Number KA1327 100 assays Version: 07 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
The clathrin adaptor Numb regulates intestinal cholesterol. absorption through dynamic interaction with NPC1L1
 The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
supplementary information
 DOI: 10.1038/ncb2153 Figure S1 Ectopic expression of HAUSP up-regulates REST protein. (a) Immunoblotting showed that ectopic expression of HAUSP increased REST protein levels in ENStemA NPCs. (b) Immunofluorescent
DOI: 10.1038/ncb2153 Figure S1 Ectopic expression of HAUSP up-regulates REST protein. (a) Immunoblotting showed that ectopic expression of HAUSP increased REST protein levels in ENStemA NPCs. (b) Immunofluorescent
Study of different types of ubiquitination
 Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
TITLE: Effect of MUC1 Expression on EGFR Endocytosis and Degradation in Human Breast Cancer Cell Lines
 AD AWARD NUMBER: W81XWH-06-1-0464 TITLE: Effect of MUC1 Expression on EGFR Endocytosis and Degradation in Human Breast Cancer Cell Lines PRINCIPAL INVESTIGATOR: Rachid M El Bejjani CONTRACTING ORGANIZATION:
AD AWARD NUMBER: W81XWH-06-1-0464 TITLE: Effect of MUC1 Expression on EGFR Endocytosis and Degradation in Human Breast Cancer Cell Lines PRINCIPAL INVESTIGATOR: Rachid M El Bejjani CONTRACTING ORGANIZATION:
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna
 Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Both K63 and K48 ubiquitin linkages signal lysosomal degradation of the LDL receptor
 Author s Choice Both K63 and K48 ubiquitin linkages signal lysosomal degradation of the LDL receptor Li Zhang, * Ming Xu, Elena Scotti, * Zhijian J. Chen,, Peter Tontonoz 1, * Howard Hughes Medical Institute
Author s Choice Both K63 and K48 ubiquitin linkages signal lysosomal degradation of the LDL receptor Li Zhang, * Ming Xu, Elena Scotti, * Zhijian J. Chen,, Peter Tontonoz 1, * Howard Hughes Medical Institute
Nature Structural & Molecular Biology: doi: /nsmb.3218
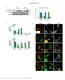 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
Product Manual. Human LDLR ELISA Kit. Catalog Number. FOR RESEARCH USE ONLY Not for use in diagnostic procedures
 Product Manual Human LDLR ELISA Kit Catalog Number STA-386 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Cholesterol is an essential component of cellular membranes,
Product Manual Human LDLR ELISA Kit Catalog Number STA-386 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Cholesterol is an essential component of cellular membranes,
LDL Uptake Cell-Based Assay Kit
 LDL Uptake Cell-Based Assay Kit Item No. 10011125 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
LDL Uptake Cell-Based Assay Kit Item No. 10011125 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson
 Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson General principles Properties of lysosomes Delivery of enzymes to lysosomes Endocytic uptake clathrin, others Endocytic pathways recycling vs.
Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson General principles Properties of lysosomes Delivery of enzymes to lysosomes Endocytic uptake clathrin, others Endocytic pathways recycling vs.
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
CYLD Negatively Regulates Transforming Growth Factor-β Signaling via Deubiquitinating Akt
 Supplementary Information CYLD Negatively Regulates Transforming Growth Factor-β Signaling via Deubiquitinating Akt Jae Hyang Lim, Hirofumi Jono, Kensei Komatsu, Chang-Hoon Woo, Jiyun Lee, Masanori Miyata,
Supplementary Information CYLD Negatively Regulates Transforming Growth Factor-β Signaling via Deubiquitinating Akt Jae Hyang Lim, Hirofumi Jono, Kensei Komatsu, Chang-Hoon Woo, Jiyun Lee, Masanori Miyata,
Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved
 1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
Supplementary Figure 1
 Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
Supplementary Figure 1 YAP negatively regulates IFN- signaling. (a) Immunoblot analysis of Yap knockdown efficiency with sh-yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN- -Luc and PRDs
PCSK9 RNAi Therapeutics. Kevin FitzGerald
 PCSK9 RNAi Therapeutics Kevin FitzGerald Presenter Disclosure Information PSCK9 RNAi Therapeutics The following relationships exist related to this presentation:» Kevin Fitzgerald and Alnylam team members:
PCSK9 RNAi Therapeutics Kevin FitzGerald Presenter Disclosure Information PSCK9 RNAi Therapeutics The following relationships exist related to this presentation:» Kevin Fitzgerald and Alnylam team members:
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Metabolism and Atherogenic Properties of LDL
 Metabolism and Atherogenic Properties of LDL Manfredi Rizzo, MD, PhD Associate Professor of Internal Medicine Faculty of Medicine, University of Palermo, Italy & Affiliate Associate Professor of Internal
Metabolism and Atherogenic Properties of LDL Manfredi Rizzo, MD, PhD Associate Professor of Internal Medicine Faculty of Medicine, University of Palermo, Italy & Affiliate Associate Professor of Internal
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer. Application Note
 Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Song et al. identified FBXW7 as a new positive regulator for RIG-Itriggered type I IFN signaling pathway. The authors observed
Reviewers' comments: Reviewer #1 (Remarks to the Author): In this manuscript, Song et al. identified FBXW7 as a new positive regulator for RIG-Itriggered type I IFN signaling pathway. The authors observed
Supplementary Figure 1. Spatial distribution of LRP5 and β-catenin in intact cardiomyocytes. (a) and (b) Immunofluorescence staining of endogenous
 Supplementary Figure 1. Spatial distribution of LRP5 and β-catenin in intact cardiomyocytes. (a) and (b) Immunofluorescence staining of endogenous LRP5 in intact adult mouse ventricular myocytes (AMVMs)
Supplementary Figure 1. Spatial distribution of LRP5 and β-catenin in intact cardiomyocytes. (a) and (b) Immunofluorescence staining of endogenous LRP5 in intact adult mouse ventricular myocytes (AMVMs)
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Myelin suppresses axon regeneration by PIR-B/SHPmediated inhibition of Trk activity
 Manuscript EMBO-2010-76298 Myelin suppresses axon regeneration by PIR-B/SHPmediated inhibition of Trk activity Yuki Fujita, Shota Endo, Toshiyuki Takai and Toshihide Yamashita Corresponding author: Toshihide
Manuscript EMBO-2010-76298 Myelin suppresses axon regeneration by PIR-B/SHPmediated inhibition of Trk activity Yuki Fujita, Shota Endo, Toshiyuki Takai and Toshihide Yamashita Corresponding author: Toshihide
TITLE: The Role of hcdc4 as a Tumor Suppressor Gene in Genomic Instability Underlying Prostate Cancer
 AD Award Number: TITLE: The Role of hcdc4 as a Tumor Suppressor Gene in Genomic Instability Underlying Prostate Cancer PRINCIPAL INVESTIGATOR: Audrey van Drogen, Ph.D. CONTRACTING ORGANIZATION: Sidney
AD Award Number: TITLE: The Role of hcdc4 as a Tumor Suppressor Gene in Genomic Instability Underlying Prostate Cancer PRINCIPAL INVESTIGATOR: Audrey van Drogen, Ph.D. CONTRACTING ORGANIZATION: Sidney
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Jewell et al., http://www.jcb.org/cgi/content/full/jcb.201007176/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. IR Munc18c association is independent of IRS-1. (A and
Supplemental material Jewell et al., http://www.jcb.org/cgi/content/full/jcb.201007176/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. IR Munc18c association is independent of IRS-1. (A and
Part-4. Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death
 Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Supplemental information
 Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
THE ROLE OF ALTERED CALCIUM AND mtor SIGNALING IN THE PATHOGENESIS OF CYSTINOSIS
 Research Foundation, 18 month progress report THE ROLE OF ALTERED CALCIUM AND mtor SIGNALING IN THE PATHOGENESIS OF CYSTINOSIS Ekaterina Ivanova, doctoral student Elena Levtchenko, MD, PhD, PI Antonella
Research Foundation, 18 month progress report THE ROLE OF ALTERED CALCIUM AND mtor SIGNALING IN THE PATHOGENESIS OF CYSTINOSIS Ekaterina Ivanova, doctoral student Elena Levtchenko, MD, PhD, PI Antonella
CKIP-1 couples Smurf1 ubiquitin ligase with Rpt6 subunit of proteasome to promote substrate degradation
 Manuscript EMBO-2012-36312 CKIP-1 couples Smurf1 ubiquitin ligase with Rpt6 subunit of proteasome to promote substrate degradation Yifang Wang, Jing Nie, Yiwu Wang, Luo Zhang, Guichun Xing, Ping Xie, Fuchu
Manuscript EMBO-2012-36312 CKIP-1 couples Smurf1 ubiquitin ligase with Rpt6 subunit of proteasome to promote substrate degradation Yifang Wang, Jing Nie, Yiwu Wang, Luo Zhang, Guichun Xing, Ping Xie, Fuchu
Supplemental Materials. STK16 regulates actin dynamics to control Golgi organization and cell cycle
 Supplemental Materials STK16 regulates actin dynamics to control Golgi organization and cell cycle Juanjuan Liu 1,2,3, Xingxing Yang 1,3, Binhua Li 1, Junjun Wang 1,2, Wenchao Wang 1, Jing Liu 1, Qingsong
Supplemental Materials STK16 regulates actin dynamics to control Golgi organization and cell cycle Juanjuan Liu 1,2,3, Xingxing Yang 1,3, Binhua Li 1, Junjun Wang 1,2, Wenchao Wang 1, Jing Liu 1, Qingsong
NFIL3/E4BP4 controls Type 2 T helper cell cytokine expression
 Manuscript EMBO-2010-75657 NFIL3/E4BP4 controls Type 2 T helper cell cytokine expression Masaki Kashiwada, Suzanne L. Cassel, John D. Colgan and Paul B. Rothman Corresponding author: Paul Rothman, University
Manuscript EMBO-2010-75657 NFIL3/E4BP4 controls Type 2 T helper cell cytokine expression Masaki Kashiwada, Suzanne L. Cassel, John D. Colgan and Paul B. Rothman Corresponding author: Paul Rothman, University
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION
 Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.
 Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
DOI: 10.1038/ncb2566 Figure S1 CDKL5 protein expression pattern and localization in mouse brain. (a) Multiple-tissue western blot from a postnatal day (P) 21 mouse probed with an antibody against CDKL5.
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene
 Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene YUELIN ZHANG, WEIHUA FAN, MARK KINKEMA, XIN LI, AND
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/9/439/ra78/dc1 Supplementary Materials for Small heterodimer partner mediates liver X receptor (LXR) dependent suppression of inflammatory signaling by promoting
www.sciencesignaling.org/cgi/content/full/9/439/ra78/dc1 Supplementary Materials for Small heterodimer partner mediates liver X receptor (LXR) dependent suppression of inflammatory signaling by promoting
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were
 Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Supplementary Figure 1 CD4 + T cells from PKC-θ null mice are defective in NF-κB activation during T cell receptor signaling. CD4 + T cells were isolated from wild type (PKC-θ- WT) or PKC-θ null (PKC-θ-KO)
Characterization of the role of EGF-A of low density lipoprotein receptor in PCSK9 binding
 Characterization of the role of EGF-A of low density lipoprotein receptor in PCSK9 binding Hong-mei Gu, 1 Ayinuer Adijiang, 1 Matthew Mah, and Da-wei Zhang 2 Departments of Pediatrics and Biochemistry,
Characterization of the role of EGF-A of low density lipoprotein receptor in PCSK9 binding Hong-mei Gu, 1 Ayinuer Adijiang, 1 Matthew Mah, and Da-wei Zhang 2 Departments of Pediatrics and Biochemistry,
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplementary Materials
 Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
FOR REVIEW. BMB Reports - Manuscript Submission. Manuscript Draft. Manuscript Number: BMB
 BMB Reports - Manuscript Submission Manuscript Draft Manuscript Number: BMB-18-095 Title: Insulin Receptor Substrate 2:A Bridge between Hippo and AKT Pathways Article Type: Perspective (Invited Only) Keywords:
BMB Reports - Manuscript Submission Manuscript Draft Manuscript Number: BMB-18-095 Title: Insulin Receptor Substrate 2:A Bridge between Hippo and AKT Pathways Article Type: Perspective (Invited Only) Keywords:
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10643 Supplementary Table 1. Identification of hecw-1 coding polymorphisms at amino acid positions 322 and 325 in 162 strains of C. elegans. WWW.NATURE.COM/NATURE 1 Supplementary Figure
doi:10.1038/nature10643 Supplementary Table 1. Identification of hecw-1 coding polymorphisms at amino acid positions 322 and 325 in 162 strains of C. elegans. WWW.NATURE.COM/NATURE 1 Supplementary Figure
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Loss of Calreticulin Uncovers a Critical Role for Calcium in Regulating Cellular Lipid Homeostasis
 SUPPLEMENTARY MATERIAL Loss of Calreticulin Uncovers a Critical Role for Calcium in Regulating Cellular Lipid Homeostasis Wen-An Wang 1, Wen-Xin Liu 1, Serpen Durnaoglu 2, Sun-Kyung Lee 2, Jihong Lian
SUPPLEMENTARY MATERIAL Loss of Calreticulin Uncovers a Critical Role for Calcium in Regulating Cellular Lipid Homeostasis Wen-An Wang 1, Wen-Xin Liu 1, Serpen Durnaoglu 2, Sun-Kyung Lee 2, Jihong Lian
From Biology to Therapy The biology of PCSK9 in humans Just LDL-cholesterol or more? May 24th. Dr. Gilles Lambert
 Dr. Gilles Lambert Associate Professor in Cell Biology University of Nantes Medical School Group Leader, Laboratory of Nutrition and Metabolism, University Hospital of Nantes From Biology to Therapy The
Dr. Gilles Lambert Associate Professor in Cell Biology University of Nantes Medical School Group Leader, Laboratory of Nutrition and Metabolism, University Hospital of Nantes From Biology to Therapy The
SUPPLEMENTARY INFORMATION
 Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
Alternative splicing. Biosciences 741: Genomics Fall, 2013 Week 6
 Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin
 SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): Bempedoic acid (ETC-1002) is an ATP-citrate lyase (ACL) inhibitor that in clinical studies has been shown to decrease plasma LDL cholesterol apparently
Reviewers' comments: Reviewer #1 (Remarks to the Author): Bempedoic acid (ETC-1002) is an ATP-citrate lyase (ACL) inhibitor that in clinical studies has been shown to decrease plasma LDL cholesterol apparently
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy
 Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
Oncolytic Adenovirus Complexes Coated with Lipids and Calcium Phosphate for Cancer Gene Therapy Jianhua Chen, Pei Gao, Sujing Yuan, Rongxin Li, Aimin Ni, Liang Chu, Li Ding, Ying Sun, Xin-Yuan Liu, Yourong
genome edited transient transfection, CMV promoter
 Supplementary Figure 1. In the absence of new protein translation, overexpressed caveolin-1-gfp is degraded faster than caveolin-1-gfp expressed from the endogenous caveolin 1 locus % loss of total caveolin-1-gfp
Supplementary Figure 1. In the absence of new protein translation, overexpressed caveolin-1-gfp is degraded faster than caveolin-1-gfp expressed from the endogenous caveolin 1 locus % loss of total caveolin-1-gfp
MCB130 Midterm. GSI s Name:
 1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
Figure S1. Western blot analysis of clathrin RNA interference in human DCs Human immature DCs were transfected with 100 nm Clathrin SMARTpool or
 Figure S1. Western blot analysis of clathrin RNA interference in human DCs Human immature DCs were transfected with 100 nm Clathrin SMARTpool or control nontargeting sirnas. At 90 hr after transfection,
Figure S1. Western blot analysis of clathrin RNA interference in human DCs Human immature DCs were transfected with 100 nm Clathrin SMARTpool or control nontargeting sirnas. At 90 hr after transfection,
Problem Set 8 Key 1 of 8
 7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
Defective Hepatic Autophagy in Obesity Promotes ER Stress and Causes Insulin Resistance
 Cell Metabolism, Volume 11 Supplemental Information Defective Hepatic Autophagy in Obesity Promotes ER Stress and Causes Insulin Resistance Ling Yang, Ping Li, Suneng Fu, Ediz S. Calay, and Gökhan S. Hotamisligil
Cell Metabolism, Volume 11 Supplemental Information Defective Hepatic Autophagy in Obesity Promotes ER Stress and Causes Insulin Resistance Ling Yang, Ping Li, Suneng Fu, Ediz S. Calay, and Gökhan S. Hotamisligil
Supplemental information contains 7 movies and 4 supplemental Figures
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplemental information contains 7 movies and 4 supplemental Figures Movies: Movie 1. Single virus tracking of A4-mCherry-WR MV
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplemental information contains 7 movies and 4 supplemental Figures Movies: Movie 1. Single virus tracking of A4-mCherry-WR MV
Supplementary Figure S1 Supplementary Figure S2
 Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
supplementary information
 Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice
 Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice Fasted Refed GFP OGT GFP OGT Liver G6P (mmol/g) 0.03±0.01 0.04±0.02 0.60±0.04 0.42±0.10 A TGs (mg/g of liver) 20.08±5.17 16.29±0.8
Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice Fasted Refed GFP OGT GFP OGT Liver G6P (mmol/g) 0.03±0.01 0.04±0.02 0.60±0.04 0.42±0.10 A TGs (mg/g of liver) 20.08±5.17 16.29±0.8
A. List of selected proteins with high SILAC (H/L) ratios identified in mass
 Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Schwarz et al. Activity-Dependent Ubiquitination of GluA1 Mediates a Distinct AMPAR Endocytosis
 Schwarz et al Activity-Dependent Ubiquitination of GluA1 Mediates a Distinct AMPAR Endocytosis and Sorting Pathway Supplemental Data Supplemental Fie 1: AMPARs undergo activity-mediated ubiquitination
Schwarz et al Activity-Dependent Ubiquitination of GluA1 Mediates a Distinct AMPAR Endocytosis and Sorting Pathway Supplemental Data Supplemental Fie 1: AMPARs undergo activity-mediated ubiquitination
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. 2 3 4 DOI: 10.1038/NMAT4893 EGFR and HER2 activate rigidity sensing only on rigid matrices Mayur Saxena 1,*, Shuaimin Liu 2,*, Bo Yang 3, Cynthia Hajal
In the format provided by the authors and unedited. 2 3 4 DOI: 10.1038/NMAT4893 EGFR and HER2 activate rigidity sensing only on rigid matrices Mayur Saxena 1,*, Shuaimin Liu 2,*, Bo Yang 3, Cynthia Hajal
Many G protein-coupled receptors (GPCRs) 2 are rapidly endocytosed after agonist binding, but the pathway of postendocytic
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 282, NO. 40, pp. 29646 29657, October 5, 2007 2007 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Hepatocyte Growth
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 282, NO. 40, pp. 29646 29657, October 5, 2007 2007 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in the U.S.A. Hepatocyte Growth
ab E3 Ligase Auto- Ubiquitilylation Assay Kit
 ab139469 E3 Ligase Auto- Ubiquitilylation Assay Kit Instructions for Use For testing ubiquitin E3 ligase activity through assessment of their ability to undergo auto-ubiquitinylation This product is for
ab139469 E3 Ligase Auto- Ubiquitilylation Assay Kit Instructions for Use For testing ubiquitin E3 ligase activity through assessment of their ability to undergo auto-ubiquitinylation This product is for
Appendix. Table of Contents
 Appendix Table of Contents Appendix Figures Figure S1: Gp78 is not required for the degradation of mcherry-cl1 in Hela Cells. Figure S2: Indel formation in the MARCH6 sgrna targeted HeLa clones. Figure
Appendix Table of Contents Appendix Figures Figure S1: Gp78 is not required for the degradation of mcherry-cl1 in Hela Cells. Figure S2: Indel formation in the MARCH6 sgrna targeted HeLa clones. Figure
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Supporting Information
 Supporting Information Valkenburg et al. 10.1073/pnas.1403684111 SI Materials and Methods ELISA and Microneutralization. Sera were treated with Receptor Destroying Enzyme II (RDE II, Accurate) before ELISA
Supporting Information Valkenburg et al. 10.1073/pnas.1403684111 SI Materials and Methods ELISA and Microneutralization. Sera were treated with Receptor Destroying Enzyme II (RDE II, Accurate) before ELISA
ab LDL Uptake Assay Kit (Cell-Based)
 ab133127 LDL Uptake Assay Kit (Cell-Based) Instructions for Use For the detection of LDL uptake into cultured cells. This product is for research use only and is not intended for diagnostic use. Version
ab133127 LDL Uptake Assay Kit (Cell-Based) Instructions for Use For the detection of LDL uptake into cultured cells. This product is for research use only and is not intended for diagnostic use. Version
NAT10 regulates p53 activation through acetylating p53 at K120 and ubiquitinating Mdm2
 Article NAT10 regulates p53 activation through acetylating p53 at K120 and ubiquitinating Mdm2 Xiaofeng Liu 1,2, Yuqin Tan 1,2, Chunfeng Zhang 1,3, Ying Zhang 1,4, Liangliang Zhang 1,2, Pengwei Ren 1,2,
Article NAT10 regulates p53 activation through acetylating p53 at K120 and ubiquitinating Mdm2 Xiaofeng Liu 1,2, Yuqin Tan 1,2, Chunfeng Zhang 1,3, Ying Zhang 1,4, Liangliang Zhang 1,2, Pengwei Ren 1,2,
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth.
 Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
Supplementary Information Supplementary Fig. 1. Elevated Usp9x in melanoma and NRAS mutant melanoma cells are dependent on NRAS for 3D growth. a. Immunoblot for Usp9x protein in NRAS mutant melanoma cells
SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
