File name: Supplementary Information Description: Supplementary figures, supplementary tables and supplementary references.
|
|
|
- Frederica Griffin
- 6 years ago
- Views:
Transcription
1 Description Supplementary Files File name: Supplementary Information Description: Supplementary figures, supplementary tables and supplementary references. File name: Peer review file
2 Supplementary Figure 1. Secondary structure prediction two GTPase-like domains in the p190rhogap middle domain. Predicted secondary structure (HHpred) two predicted domains within the p190rhogap-a (human) middle domain (pg1 and pg2) compared to the assigned secondary structure H- Ras (: 5P21), RhoA (: 1FTN), Rap1B (: 3X1X), Rab8A (: 4LHW), (determined in DSSP 1 ). 1
3 Supplementary Figure 2. Structure analysis the p190rhogap pg1. a) Representative stereoview electron density maps for p190rhogap-a pg1 (top) and p190rhogap-b pg1 (bottom). Blue: 2Fobs-Fcalc at 1σ; Green: Fobs-Fcalc at +3σ; Red: Fobs-Fcalc at 3σ. Some residue numbers are indicated. b) Topology map p190rhogap pg1 (left) 2
4 compared to H-Ras (right; : 5P21). The overall topology is similar but some differences exist, particularly in the α2, β2 and Switch I elements. Figure generated using PowerPoint (Microst). c). Structure-based sequence alignment (DALI server) and secondary structure assignments (DSSP) for frog p190rhogap-a pg1, human p190rhogap-b pg1, and human H- Ras (: 5P21). modeled in the structure are in bold. Residue numbering for Xenopus laevis p190rhogap-a is depicted above and for H-Ras below. The figure was created using ALINE 2. 3
5 a b P-loop Switch I Switch II G1 G2 G3 G4 G5 p190-a pg2 GDPFS.AD CKSSHCGSNNSVL SYHSS LTDG PCS p190-b pg2 GDPFS.VD CSAAQAGQNNSLM SYHSS VTDS YSL Rab8A GDSGVGKT FNSTFISTIGIDF DTAGQ NKCD SAK Rab28A GDGASGKT FGKQYKQTIGLDF DIGGQ NKID SAK Rab26 GDSGVGKT FLAGFISTVGIDF DTAGQ NKVD SAK Rab21 GEGCVGKT FNDKHITTLGASF DTAGQ NKID SAK Rab5B GDVGAGKS FVEFQESTIGAAF DTAGQ NKSD SAK H-Ras GAGGVGKS FVDEYDPTIEDSY DTAGQ NKCD SAK Consensus GxxxxGKS...T... DxxGQ NKxD SAK Supplementary Figure 3. Homology-based sequence alignment p190rhogap pg2. a) Aligned secondary structure prediction (HHpred) human p190rhogap-a pg2 and secondary structure assignment for the top HHpred hit Rab8A (DSSP; : 4LHW, see Supplementary Table 1), are shown above the sequences. Residue numbering is included. G-motifs for Rab8A are underlined and labeled in red. b) Aligned G motifs from top HHpred homologous domains (Table S3) to pg2 from p190rhogap-a (human) showing that the predicted G motifs in pg2 are likely degraded. 4
6 Supplementary Figure 4. Conservation analysis p190rhogap-a pg1. a) Ribbon diagram p190rhogap-a pg1 with residues colored according to conservation scores from ConSurf analysis3 the alignment in c. Dark blue indicates high conservation, white indicates low or no conservation. b) Surface representation p190rhogap-a pg1 colored by sequence conservation. c) A manually curated alignment p190rhogap-a pg1 regions from 74 species. 5
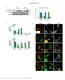
7 Supplementary Figure 5. Uncropped immunoblots from Figure 5 the main text. The molecular weight markers (Bio-Rad Precision Plus All Blue), which are visible in the 700 nm channel on the LI-COR Odyssey CLx Imaging system, are included in the images and labelled. 6
8 Supplementary Table 1. Top HHpred predicted homologous domains. pg1 Human p190rhogap-a pg1 (residues ) pg2 Human p190rhogap-a pg2 (residues ) 1 Rab8A 4LHW E Rab8A 4LHW E Rheb 3T5G E Rab28A 2HXS E Rap1B 3X1X E Rab26 2G6B E Rab5B 2EFE E Rab21 1Z E Ras3 4KU E Rab5B 2EFE E Rab5A 1R2Q E Rad 3Q E Rab23 1Z2A E Rab23 1Z2A E Rit1 4KLZ E Rit1 4KLZ E Rab28A 2HXS E Ras3 4KU E RalA 1U8Z E Rab5A 1R2Q E Human p190rhogap-b pg1 (residues ) Human p190rhogap-b pg2 (residues ) 1 Rheb 3SEA E RheB 3SEA E Rab 5UB E Centg 2IWR E M-Ras 3KKQ E Rab 5UB E Rap1b 3X1X E Ral-B 2KWI E Ras3 4KU E Ras-3 4KU E RhebL1 3OES E Rasl12 3C5C E Rem 3Q E RhebL1 3OES E Rheb 3T5G E Rasl12 3T5G E Rab2B 2A5J E EhRho1 3REG E Di-Ras1 2GF E Rho6 2REX E Drosophila p190rhogap pg1 (residues ) Drosophila p190rhogap pg2 (residues ) 1 Rheb 3SEA E Rab8A 4LHW E Rab 5UB E Rab28A 2HXS E M-Ras 3KKQ E Rab26 2G6B E Sponge p190rhogap pg1 (residues ) Sponge p190rhogap pg2 (residues ) 1 Rab8A 4LHW E Rab8A 4LHW E Rap1B 3X1X E Rab23 1Z2A E RalA 1U8Z E R-Ras 2FN E Uniprot IDs for Human (Homo sapiens) p190rhogap-a, Q9NRY4, and p190rhogap-b, Q Uniprot IDs for Drosophila (Drosophila melanogaster), Q9VX32. NCBI Reference sequence for Sponge (Amphimedon queenslandica), XP_
9 Supplementary Table 2. Structure-based comparison conserved G-motifs for selected small GTPases and pseudogtpases. P-loop Switch I Switch II G1 G2 G3 G4 G5 Consensus GxGxxGKS...T... DxxGQ NKxD SAK H-Ras GAGGVGKS FVDEYDPTIEDSY DTAGQ NKCD SAK Rad (RGK) GAPGVGKS P..EAEAAG.HTY DIWEQ NKSD SAA Rnd3 (Rnd) GDSQCGKT FPENYVPTVFENY DTSGS CKSD SAL AGAP2 GDARSGKS YQV-LEKTESEQY EEAGA TQDR CAT p190-a pg1 GRDG.LAR (deletion) PIEGN VNRR AST p190-b pg1 GKDG.LAQ (deletion) PVDAK ANQR PAG LIC GGTVDSQR RR(+10)FALGYT YTLTD QNAE TPS CENP-M GTEDALLQ (deletion) LAKSL TGAG DLE Red indicates residues that do not match the consensus. 8
10 Supplementary Table 3. Synthetic DNA sequences. p190rhogap-a pg1 (Gallus gallus) p190rhogap-a pg1 (Xenopus laevis) p190rhogap-a pg1 (Danio rerio) p190rhogap pg1 (Drosophila melanogaster) GGATCCGACCCGAACGTGGACCGTATCAACCTGGTTATTCTGG GCAAAGATGGCCTGGCTCGTGAACTGGCAAATGAAATCCGTGC TCTGTGTACCAACGATGACAAATATGTCATTGAAGGCAAAATG TACGAACTGTCCCTGCGTCCGATCGAGGGTAACGTCCGCCTGC CGGTGAATGCCTTTCAGACCCCGACGTTCCAACCGCATGGCTG CCTGTGTCTGTATAATAGCAAAGAAAGCCTGTCTTACGTGGTT GAAAGTATTGAAAAATCCGCCGCGGGTCGTCGCGAAAACCATC CGGCACACCTGCCGCTGACCCTGATGCTGGTTAATAAACGTGG TGATGCAGGCGGTGAAACGCTGCACAGTCTGGTGCAGCAAGGC CAGCAAATCGCTGGTAAACTGCAGTGCGTTTTTCTGGACCCGG CAGCTGCGGGCATTGCATATGGTCGTAGCGTTAACGAAAAACA GATCTCTCAAGTCCTGAAAGGTCTGCTGGATTCAAAACGCAAT CTGTCGCTGTGA GGATCCGACCCGAATGTTGACCGTATTAATCTGGTTATCCTGG GCCGTGATGGCCTGGCTCGTGAACTGGCAAATGAAATCCGTGC TCTGTGCACCAACGATGACAAATACGTGATCGATGGTAAAATG TACGAACTGTCACTGCGTCCGATCGAAGGCAACGTCCGCCTGC CGGTGAATAGCTTTCAGACCTCTACGTTCCAACCGCATGGTTG CCTGTGTCTGTATAACTCCAAAGAATCACTGTCGTACGTGGTT GAAAGTATTGAAAAATCCCGTGAAAGCTCTCTGGGCCGCAAAG AAAACCACCTGATTAATCTGCCGCTGACCCTGATCCTGGTTAA TCGTCGCGGTGATACCAGTTCCGAAACGGCACATAGCCTGATT CAGCACGGCCAGCAAGTTGCGTCTAAACTGCAATGTGTCTTTC TGGATGCCGCAAGCACCGGTCTGGGTTATGGTCGTAACATCAA CGAAAAACAGATCTCACTGATCCTGCGTGGTCTGCTGGAATCG AAACGCAACCTGAATCTGTGA GGATCCGACGCAAAAGTGGACCGTATCAACCTGGTCATTCTGG GTAAAGATGGCCTGGCTCGTGAACTGGCGAACGAAATCCGTGC CCTGTGTACCAACGATGACCATTATGTCCTGGAAGGTCGTATG TACGAACTGGTTCTGCGCCCGATTGAAGGCAACGTCCGTCTGC CGGTGAACAGCTTTCATACCTCGACGTTCACCCCGCACGGTTG CCTGTGTCTGTATAACAGTAAAGAAAGTCTGTCCTACGTGGTT GAAAGTATTGAAAAACTGCGCGAATCCACGCTGGGCCGTCGCG ATAATCACATGGCACAGCTGCCGCTGAGCCTGCTGCTGGTTTC TAAACGTGGTGTCGGCGGTCTGTCAGACATCGGCGGTGAAACC GCGCAGGCCCTGATTTCGCAAGGCCAGCAAGTGGCAATCAAAC TGCAGTGCAACTATCTGGAACCGAGCTCTCCGGGTACCGGTTA TGGTCGTAACGTGAACGAAAAACAGATCAACCAAGTTCTGAAA GCTCTGCTGGAAACCCGTCGCACGTCTACCTTTTGA GGATCCGGCTCGGACCGCACGCTGAACCTGCTGATTGTGGGCT CGGAACACCTGGCATCTGACCTGCTGAACGACATTCGCATCTG TACGGGTAGCAAAGGCGAATATATTTACGAAAACCAGACCTAT TACCTGAACTATCGTATCGCGAATGGCGATATGGAAGCGTTTA AAGCCATTGACGTCTATAGCTCTGGTCTGATCTGCGTGTACAG TAACCAGCAATCCTTCGAAACCCTGAAAGATAACCTGGAACGC ACGCTGCTGTGTAATCTGGAACTGGAAGACAAATTTGAAAATC TGCCGATTGTGCTGGTTTATCAGCCGCAAGATCTGAAAGAAAA CGAAGTTGAATACCTGCGTAATGAGGGTATGCGCCTGAGCGAA ATGCTGCATTGCGATTTCATCGACCATACGCAGAATCACCAAA AATACGTCTATGACATCCTGAACATCGTGATTCTGAGCCTGAA ACTGACCGAAATGAAATGA 9
11 Supplementary Table 4. Primer Sequences. pcdna Flagp190RhoGAP-A pg1 p190rhogap-a (residues ) pg2 p190rhogap-a (residues ) pg1 p190rhogap-b (residues ) pg2 p190rhogap-b (residues ) pcdna Flagp190RhoGAP-A ΔpG1-pG2 mutagenesis Forward Reverse Forward Reverse Forward Reverse Forward Reverse Forward Reverse Forward Reverse 5 -GCAGGATCCTATGATGATGGCAAGA-3 5 -CTGGAATTCTCACAGCGTGTGTTC-3 5 -GAAGGATCCGACCCCAATATTGAT-3 5 -CCTGAATTCCTACAGGTTTAAGTTGCG-3 5 -GAAGGATCCAACCTGGTTAGTTCT-3 5 -CCTGAATTCCTAAGCCACATTATCGTAC-3 5 -GAAGGATCCAGTACCAATATAGAT-3 5 -CCTGAATTCCTACACATCCAAATTGTG-3 5 -GAAGGATCCGTGAGCCCAATTCCT-3 5 -CCTGAATTCCTATGTATTATCAGACAA-3 5'-TAGGAATCAGAAGAACTCTTTGTCTTGCAGCACCACTG-3' 5'-CAGTGGTGCTGCAAGACAAAGAGTTCTTCTGATTCCTA-3' 10
12 Supplementary References 1. Kabsch, W. & Sander, C. Dictionary protein secondary structure: pattern recognition hydrogen-bonded and geometrical features. Biopolymers 22, (1983). 2. Bond, C.S. & Schuttelkopf, A.W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta crystallographica. Section D, Biological crystallography 65, (2009). 3. Landau, M. et al. ConSurf 2005: the projection evolutionary conservation scores residues on protein structures. Nucleic Acids Res 33, W (2005). 11
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supplementary Figure 1. CFTR protein structure and domain architecture.
 A Plasma Membrane NH ₂ COOH Supplementary Figure. CFT protein structure and domain architecture. (A) Open state CFT homology model, ribbon representation from Serohijos et al. 8 PNAS 5:356. CFT domains
A Plasma Membrane NH ₂ COOH Supplementary Figure. CFT protein structure and domain architecture. (A) Open state CFT homology model, ribbon representation from Serohijos et al. 8 PNAS 5:356. CFT domains
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
DOI: 10.1038/ncb3073 LATS2 Binding ability to SIAH2 Degradation by SIAH2 1-160 161-402 403-480 -/ 481-666 1-666 - - 667-1088 -/ 1-0 Supplementary Figure 1 Schematic drawing of LATS2 deletion mutants and
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Insights into the Giardia intestinalis Enolase and Human Plasminogen interaction
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1
 S U P P L E M E N TA R Y I N F O R M AT I O N DOI: 10.1038/ncb2896 Supplementary Figure 1 Supplementary Figure 1. Sequence alignment of TERB1 homologs in vertebrates. M. musculus TERB1 was derived from
S U P P L E M E N TA R Y I N F O R M AT I O N DOI: 10.1038/ncb2896 Supplementary Figure 1 Supplementary Figure 1. Sequence alignment of TERB1 homologs in vertebrates. M. musculus TERB1 was derived from
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
Lesson 5 Proteins Levels of Protein Structure
 Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1134389/dc1 Supporting Online Material for PI(3,4,5)P 3 and PI(4,5)P 2 Lipids Target Proteins with Polybasic Clusters to the Plasma Membrane Won Do Heo, Takanari Inoue,
www.sciencemag.org/cgi/content/full/1134389/dc1 Supporting Online Material for PI(3,4,5)P 3 and PI(4,5)P 2 Lipids Target Proteins with Polybasic Clusters to the Plasma Membrane Won Do Heo, Takanari Inoue,
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Nature Structural & Molecular Biology: doi: /nsmb.1933
 The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
WDR62 is associated with the spindle pole and mutated in human microcephaly
 WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Kirschner, 2005). Briefly, parallel MEF cultures were isolated from single littermate
 SUPPLEMENTAL MATERIALS AND METHODS Generation of MEFs, osteoblasts and cell culture Prkar1a -/- and control MEFs were generated and cultured as described (Nadella and Kirschner, 2005). Briefly, parallel
SUPPLEMENTAL MATERIALS AND METHODS Generation of MEFs, osteoblasts and cell culture Prkar1a -/- and control MEFs were generated and cultured as described (Nadella and Kirschner, 2005). Briefly, parallel
Supplementary Figure 1. Effect of cellular glycolysis on tumor cell exosome secretion. A549 cells were cultured in medium containing different
 Supplementary Figure 1. Effect of cellular glycolysis on tumor cell exosome secretion. A549 cells were cultured in medium containing different concentration of glucose (a) or treated the inhibitor of glycolysis
Supplementary Figure 1. Effect of cellular glycolysis on tumor cell exosome secretion. A549 cells were cultured in medium containing different concentration of glucose (a) or treated the inhibitor of glycolysis
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Choi YL, Soda M, Yamashita Y, et al. EML4-ALK mutations in
Protein Prediction 1 for Computational Biologists - Exercise. Exercise 1 - Introduction
 Protein Prediction 1 for Computational Biologists - Exercise Exercise 1 - Introduction Hi :) Maria Schelling: maria.schelling@tum.de PhD student @Rostlab Contact me via email if you have any questions/problems
Protein Prediction 1 for Computational Biologists - Exercise Exercise 1 - Introduction Hi :) Maria Schelling: maria.schelling@tum.de PhD student @Rostlab Contact me via email if you have any questions/problems
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Co-expressed Cyclin D variants cooperate to regulate proliferation of germline nuclei in. Accession number
 SUPPLEMENTARY MATERIAL Co-expressed Cyclin D variants cooperate to regulate proliferation of germline nuclei in a syncytium. Subramaniam et al. Table S1. Cyclin accession numbers Gene Human Cyclin D1a
SUPPLEMENTARY MATERIAL Co-expressed Cyclin D variants cooperate to regulate proliferation of germline nuclei in a syncytium. Subramaniam et al. Table S1. Cyclin accession numbers Gene Human Cyclin D1a
SUPPLEMENTARY MATERIAL S-1 INTREPID VARIANTS
 SUPPLEMENTARY MATERIAL S- INTREPID VARIANTS We define different INTREPID variants based on the positional conservation score cons(s, x) which is used to compute the importance score in Equation S-. IMP
SUPPLEMENTARY MATERIAL S- INTREPID VARIANTS We define different INTREPID variants based on the positional conservation score cons(s, x) which is used to compute the importance score in Equation S-. IMP
Project Manual Bio3055. Cholesterol Homeostasis: HMG-CoA Reductase
 Project Manual Bio3055 Cholesterol Homeostasis: HMG-CoA Reductase Bednarski 2003 Funded by HHMI Cholesterol Homeostasis: HMG-CoA Reductase Introduction: HMG-CoA Reductase is an enzyme in the cholesterol
Project Manual Bio3055 Cholesterol Homeostasis: HMG-CoA Reductase Bednarski 2003 Funded by HHMI Cholesterol Homeostasis: HMG-CoA Reductase Introduction: HMG-CoA Reductase is an enzyme in the cholesterol
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
SUPPLEMENTARY MATERIAL
 SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
SYPPLEMENTARY FIGURE LEGENDS SUPPLEMENTARY MATERIAL Figure S1. Phylogenic studies of the mir-183/96/182 cluster and 3 -UTR of Casp2. (A) Genomic arrangement of the mir-183/96/182 cluster in vertebrates.
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
doi:10.1038/nature12005 a S Ψ ΨΨ ΨΨ ΨΨ Ψ Ψ Ψ Ψ Ψ ΨΨ Ψ Ψ ΨΨΨΨΨΨ S1 (a.a. 1 751) S2 (a.a. 752 1353) S1 Fc S1 (a.a. 1-747) Fc b HCoV-EMC-S1-Fc SARS-CoV-S1-Fc HCoV-EMC-S1-Fc SARS-CoV-S1-Fc kda - 170 - - 130
Supplementary Information for Structural basis for the inhibition of Polo-like kinase 1
 Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
SpliceDB: database of canonical and non-canonical mammalian splice sites
 2001 Oxford University Press Nucleic Acids Research, 2001, Vol. 29, No. 1 255 259 SpliceDB: database of canonical and non-canonical mammalian splice sites M.Burset,I.A.Seledtsov 1 and V. V. Solovyev* The
2001 Oxford University Press Nucleic Acids Research, 2001, Vol. 29, No. 1 255 259 SpliceDB: database of canonical and non-canonical mammalian splice sites M.Burset,I.A.Seledtsov 1 and V. V. Solovyev* The
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Danish Research Institute of Translational Neuroscience DANDRITE, Nordic-EMBL Partnership
 Supplementary Information for Tuning of the Na,K-ATPase by the beta subunit Florian Hilbers 1,2,3, Wojciech Kopec 4, Toke Jost Isaksen 3,5, Thomas Hellesøe Holm 3,5, Karin Lykke- Hartmann 3,5,6, Poul Nissen
Supplementary Information for Tuning of the Na,K-ATPase by the beta subunit Florian Hilbers 1,2,3, Wojciech Kopec 4, Toke Jost Isaksen 3,5, Thomas Hellesøe Holm 3,5, Karin Lykke- Hartmann 3,5,6, Poul Nissen
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Supplementary figure legends
 Supplementary figure legends SUPPLEMENTRY FIGURE S1. Lentiviral construct. Schematic representation of the PCR fragment encompassing the genomic locus of mir-33a that was introduced in the lentiviral construct.
Supplementary figure legends SUPPLEMENTRY FIGURE S1. Lentiviral construct. Schematic representation of the PCR fragment encompassing the genomic locus of mir-33a that was introduced in the lentiviral construct.
Supplementary Figure 1: Additional metabolic parameters of obesity mouse models and controls. (a) Body weight, (b) blood glucose and (c) insulin
 Supplementary Figure 1: Additional metabolic parameters of obesity mouse models and controls. (a) Body weight, (b) blood glucose and (c) insulin resistance index of homeostatic model assessment (HOMA IR)
Supplementary Figure 1: Additional metabolic parameters of obesity mouse models and controls. (a) Body weight, (b) blood glucose and (c) insulin resistance index of homeostatic model assessment (HOMA IR)
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Identification of non-ser/thr-pro consensus motifs for Cdk1 and their roles in mitotic regulation of C2H2 zinc finger proteins and Ect2
 SUPPLEMENTARY INFORMATION Identification of non-ser/thr-pro consensus motifs for Cdk1 and their roles in mitotic regulation of C2H2 zinc finger proteins and Ect2 Kazuhiro Suzuki, Kosuke Sako, Kazuhiro
SUPPLEMENTARY INFORMATION Identification of non-ser/thr-pro consensus motifs for Cdk1 and their roles in mitotic regulation of C2H2 zinc finger proteins and Ect2 Kazuhiro Suzuki, Kosuke Sako, Kazuhiro
Homo Sapiens DCC Gene: In Silico Analysis to Understand the Affect of Mutations in Colon Cancer
 International Journal of Biotechnology and Biochemistry ISSN 0973-2691 Volume 6 Number 3 (2010) pp. 419 426 Research India Publications http://www.ripublication.com/ijbb.htm Homo Sapiens DCC Gene: In Silico
International Journal of Biotechnology and Biochemistry ISSN 0973-2691 Volume 6 Number 3 (2010) pp. 419 426 Research India Publications http://www.ripublication.com/ijbb.htm Homo Sapiens DCC Gene: In Silico
Data Mining in Bioinformatics Day 9: String & Text Mining in Bioinformatics
 Data Mining in Bioinformatics Day 9: String & Text Mining in Bioinformatics Karsten Borgwardt March 1 to March 12, 2010 Machine Learning & Computational Biology Research Group MPIs Tübingen Karsten Borgwardt:
Data Mining in Bioinformatics Day 9: String & Text Mining in Bioinformatics Karsten Borgwardt March 1 to March 12, 2010 Machine Learning & Computational Biology Research Group MPIs Tübingen Karsten Borgwardt:
a 0,8 Figure S1 8 h 12 h y = 0,036x + 0,2115 y = 0,0366x + 0,206 Labeling index Labeling index ctrl shrna Time (h) Time (h) ctrl shrna S G2 M G1
 (GFP+ BrdU+)/GFP+ Labeling index Labeling index Figure S a, b, y =,x +, y =,x +,,,,,,,, Time (h) - - Time (h) c d S G M G h M G S G M G S G h Time of BrdU injection after electroporation (h) M G S G M
(GFP+ BrdU+)/GFP+ Labeling index Labeling index Figure S a, b, y =,x +, y =,x +,,,,,,,, Time (h) - - Time (h) c d S G M G h M G S G M G S G h Time of BrdU injection after electroporation (h) M G S G M
Nature Neuroscience: doi: /nn Supplementary Figure 1. PICALM expression in brain capillary endothelium in human brain and in mouse brain.
 Supplementary Figure 1 PICALM expression in brain capillary endothelium in human brain and in mouse brain. a, Double immunostaining for PICALM (red, left) and lectin positive endothelial profiles (blue,
Supplementary Figure 1 PICALM expression in brain capillary endothelium in human brain and in mouse brain. a, Double immunostaining for PICALM (red, left) and lectin positive endothelial profiles (blue,
Changes to Biochemistry (4th ed.), 2nd Printing
 Changes to Biochemistry (4th ed.), 2nd Printing 1. p. vi, line 7. Change: hydrophobic to hydrophilic. Make the same change on the back cover. 2. p. xx, at the end of the Chap. 17 portion. Add the line:
Changes to Biochemistry (4th ed.), 2nd Printing 1. p. vi, line 7. Change: hydrophobic to hydrophilic. Make the same change on the back cover. 2. p. xx, at the end of the Chap. 17 portion. Add the line:
HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total. The ph at which a peptide has no net charge is its isoelectric point.
 BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects.
 Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects. Manuel J. Louwerse and Evert Jan Baerends Theoretical Chemistry, Vrije Universiteit Amsterdam, De Boelelaan
Supplementary Material to Oxidative properties of FeO2+: electronic structure and solvation effects. Manuel J. Louwerse and Evert Jan Baerends Theoretical Chemistry, Vrije Universiteit Amsterdam, De Boelelaan
Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ).
 Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
Supplementary Figure 1 Trim29 gene-targeting strategy. (a) Genotyping of wildtype mice (+/+), Trim29 heterozygous mice (+/ ) and homozygous mice ( / ). (b) Immunoblot analysis of TRIM29 in lung primary
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Aquaporins. Fatemeh Khalili Elizabeth Villa VMD Developers: John Stone Dan Wright John Eargle
 University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Beckman Institute Computational Biophysics Workshop Aquaporins Fatemeh Khalili Elizabeth Villa VMD
University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Beckman Institute Computational Biophysics Workshop Aquaporins Fatemeh Khalili Elizabeth Villa VMD
Nature Structural & Molecular Biology: doi: /nsmb.3218
 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Experimental design and workflow utilized to generate the WMG Protein Atlas.
 Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69
 Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
Analysis of protein modeling for envelope glycoprotein GP120 for HIV via bioinformatics approaches
 International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
SUPPLEMENTAL MATERIAL. UNC119 is required for G protein trafficking in sensory neurons
 1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons
 The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons Eli Koren, Galit Lev-Maor, Gil Ast * Department of Human Molecular Genetics, Sackler Faculty of Medicine, Tel Aviv University,
The Emergence of Alternative 39 and 59 Splice Site Exons from Constitutive Exons Eli Koren, Galit Lev-Maor, Gil Ast * Department of Human Molecular Genetics, Sackler Faculty of Medicine, Tel Aviv University,
PERFORMANCE MEASURES
 PERFORMANCE MEASURES Of predictive systems DATA TYPES Binary Data point Value A FALSE B TRUE C TRUE D FALSE E FALSE F TRUE G FALSE Real Value Data Point Value a 32.3 b.2 b 2. d. e 33 f.65 g 72.8 ACCURACY
PERFORMANCE MEASURES Of predictive systems DATA TYPES Binary Data point Value A FALSE B TRUE C TRUE D FALSE E FALSE F TRUE G FALSE Real Value Data Point Value a 32.3 b.2 b 2. d. e 33 f.65 g 72.8 ACCURACY
Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent Protein.
 prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
Project Manual Bio3055. Apoptosis: Superoxide Dismutase I
 Project Manual Bio3055 Apoptosis: Superoxide Dismutase I Bednarski 2003 Funded by HHMI Apoptosis: Superoxide Dismutase I Introduction: Apoptosis is another name for programmed cell death. It is a series
Project Manual Bio3055 Apoptosis: Superoxide Dismutase I Bednarski 2003 Funded by HHMI Apoptosis: Superoxide Dismutase I Introduction: Apoptosis is another name for programmed cell death. It is a series
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06173 SUPPLEMENTARY INFORMATION Contents of Supplementary Figures Supplementary Figure 1: Purification of mouse EHD2. Supplementary Figure 2: Analytical ultracentrifugation analysis.
doi: 10.1038/nature06173 SUPPLEMENTARY INFORMATION Contents of Supplementary Figures Supplementary Figure 1: Purification of mouse EHD2. Supplementary Figure 2: Analytical ultracentrifugation analysis.
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE
 Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE chromatin-associated proteins has been Quantitative Proteomic Analysis of Chromatin systems and reveal changes in chromatin
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE chromatin-associated proteins has been Quantitative Proteomic Analysis of Chromatin systems and reveal changes in chromatin
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Interaktionen von RNAs und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig May 5, 2017 RNA-protein binding vs. DNA-protein binding Is binding of proteins to RNA the same as binding of proteins to DNA? Will the same rules
Sonja Prohaska Computational EvoDevo Universitaet Leipzig May 5, 2017 RNA-protein binding vs. DNA-protein binding Is binding of proteins to RNA the same as binding of proteins to DNA? Will the same rules
Myers Psychology for AP* David G. Myers PowerPoint Presentation Slides by Kent Korek Germantown High School Worth Publishers, 2010
 Myers Psychology for AP* David G. Myers PowerPoint Presentation Slides by Kent Korek Germantown High School Worth Publishers, 2010 *AP is a trademark registered and/or owned by the College Board, which
Myers Psychology for AP* David G. Myers PowerPoint Presentation Slides by Kent Korek Germantown High School Worth Publishers, 2010 *AP is a trademark registered and/or owned by the College Board, which
Supplementary Information for. A cancer-associated BRCA2 mutation reveals masked nuclear. export signals controlling localization
 Supplementary Information for A cancer-associated BRCA2 mutation reveals masked nuclear export signals controlling localization Anand D Jeyasekharan 1, Yang Liu 1, Hiroyoshi Hattori 1,3, Venkat Pisupati
Supplementary Information for A cancer-associated BRCA2 mutation reveals masked nuclear export signals controlling localization Anand D Jeyasekharan 1, Yang Liu 1, Hiroyoshi Hattori 1,3, Venkat Pisupati
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Figure 1. Amino acid sequences of GodA and GodA*. Inserted. residues are colored red. Numbers indicate the position of each residue.
 Supplementary Figure 1. Amino acid sequences of GodA and GodA*. Inserted residues are colored red. Numbers indicate the position of each residue. Supplementary Figure 2. SDS-PAGE analysis of purified recombinant
Supplementary Figure 1. Amino acid sequences of GodA and GodA*. Inserted residues are colored red. Numbers indicate the position of each residue. Supplementary Figure 2. SDS-PAGE analysis of purified recombinant
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists
 Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists of: (i) the YPet domain (an enhanced YFP); (ii) the
Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists of: (i) the YPet domain (an enhanced YFP); (ii) the
For all of the following, you will have to use this website to determine the answers:
 For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Rapid Characterization of Allosteric Networks. with Ensemble Normal Mode Analysis
 Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
v Feature Stamping SMS 13.0 Tutorial Prerequisites Requirements Map Module Mesh Module Scatter Module Time minutes
 v. 13.0 SMS 13.0 Tutorial Objectives Learn how to use conceptual modeling techniques to create numerical models which incorporate flow control structures into existing bathymetry. The flow control structures
v. 13.0 SMS 13.0 Tutorial Objectives Learn how to use conceptual modeling techniques to create numerical models which incorporate flow control structures into existing bathymetry. The flow control structures
Supplementary Information. Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase
 Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
Journal: Nature Medicine Supplementary Information Induction of human pancreatic beta cell replication by inhibitors of dual specificity tyrosine regulated kinase 1,2 Peng Wang PhD, 1,2 Juan-Carlos Alvarez-Perez
obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and in a
 Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
SMPD 287 Spring 2015 Bioinformatics in Medical Product Development. Final Examination
 Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
Supplemental Information. An Atlas of b-glucuronidases. in the Human Intestinal Microbiome
 Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#10:(November-22-2010) Cancer and Signals 1 1 Micro-Environment Story How does the rest of our body influences the cancer cell?
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#10:(November-22-2010) Cancer and Signals 1 1 Micro-Environment Story How does the rest of our body influences the cancer cell?
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Ras and Cell Signaling Exercise
 Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
a) The statement is true for X = 400, but false for X = 300; b) The statement is true for X = 300, but false for X = 200;
 1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE
 Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Supplementary Figure 1. Rab27a-KD inhibits speed and persistence of HEp3 cells migrating in the chick CAM. (a) Western blot analysis of Rab27a
 Supplementary Figure 1. Rab27a-KD inhibits speed and persistence of HEp3 cells migrating in the chick CAM. (a) Western blot analysis of Rab27a expression in GFP-expressing HEp3 cells. (b) Representative
Supplementary Figure 1. Rab27a-KD inhibits speed and persistence of HEp3 cells migrating in the chick CAM. (a) Western blot analysis of Rab27a expression in GFP-expressing HEp3 cells. (b) Representative
Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A
 Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A stereo image of overall 2Fo-Fc electron density map of PI4KIIα contoured at 1.0 σ. b, 2FoFc electron density map (1.0 σ) around the
Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A stereo image of overall 2Fo-Fc electron density map of PI4KIIα contoured at 1.0 σ. b, 2FoFc electron density map (1.0 σ) around the
Predictive PP1Ca binding region in BIG3 : 1,228 1,232aa (-KAVSF-) HEK293T cells *** *** *** KPL-3C cells - E E2 treatment time (h)
 Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
Relative expression ERE-luciferase activity activity (pmole/min) activity (pmole/min) activity (pmole/min) activity (pmole/min) MCF-7 KPL-3C ZR--1 BT-474 T47D HCC15 KPL-1 HBC4 activity (pmole/min) a d
