Supplementary Materials for
|
|
|
- Virgil Lynch
- 5 years ago
- Views:
Transcription
1 advances.sciencemag.org/cgi/content/full/3/8/e /dc1 Supplementary Materials for Impaired DNA replication derepresses chromatin and generates a transgenerationally inherited epigenetic memory Adam Klosin, Kadri Reis, Cristina Hidalgo-Carcedo, Eduard Casas, Tanya Vavouri, Ben Lehner This PDF file includes: Published 16 August 2017, Sci. Adv. 3, e (2017) DOI: /sciadv fig. S1. Expression of the daf-21p::mcherry reporter in the progeny of animals treated with RNAi targeting different subunits of the DNA polymerase complex and its associated proteins. fig. S2. Increased transgene expression in div-1 mutants. fig. S3. Maternal div-1 deficiency results in elevated transgene expression in the offspring. fig. S4. Transgene up-regulation following pole-2 knockdown is suppressed in the mes-2;met-2;set-25 triple-mutant background. fig. S5. Impaired DNA replication reduces H3K27me3 levels on multiple loci. fig. S6. Global reduction of repressive histone marks and a gain of activating histone marks in late embryos. fig. S7. Knockdown of pole-2 results in reduction of H3K27me3 mark and increase in H3K4me3 in early embryonic chromatin. fig. S8. Reduction in H3K9me3 mark in mutant L1s detected by Western blot. fig. S9. Passage of the transgenic array through impaired replication for a single generation is sufficient to trigger a multigenerational effect. fig. S10. Quantification of H3K27me3 in interphase nuclei. table S1. List of genes whose knockdown results in upregulation of daf- 21p::mCherry transgene. table S2. C. elegans strains used in this study. table S3. Primers used in qpcr analyses. table S4. Transgenes tested for derepression with div-1(rnai).
2 Supplementary Figures and Tables fig. S1. Expression of the daf-21p::mcherry reporter in the progeny of animals treated with RNAi targeting different subunits of the DNA polymerase complex and its associated proteins. Quantification of mcherry fluorescence. L4 larvae carrying the daf-21p::mcherry transgene were fed with RNAi targeting the indicated genes for 24 hours (no upregulation in the parents was observed under these conditions). Offspring embryos were extracted, allowed to hatch overnight in M9 buffer and imaged under fluorescence microscope. Arrested embryos were not included in the quantification. Sample size: control, 110; rfc-1, 176; rfc-3, 129; Y47D3A.29, 11; rpa-2, 161; lrr-1, 151; pole-2, 221; div-1, 157; pole-1, 32. P-values: **** p< , ** p<0.01, * p<0.05, ns p> 0.05 (Wilcoxon rank test). Y-axes in log scale. Sample size indicated below each boxplot. All values are normalised to the median of the control.
3 A daf-21p::mcherry WT div-1 (or148) Relative daf-21p::mcherry Fluorescence **** WT B daf-21p::mcherry fig. S2. Increased transgene expression in div-1 mutants. a, Expression of daf- 21p::mCherry in mutant embryos synchronized at the 4-cell stage and allowed to develop for 6.5 hours at 20 C. The delay in development in was estimated to be approximately one hour based on embryo morphology. Sample size: WT, 43;, 35. b, Expression of daf-21p::mcherry in adult mutant div-1 (or148) worms is elevated in all tissues.
4 mcherry mcherry (1) (2) F1 mcherry(+/-) F1 div-1(or148/) mcherry(+/-) daf-21p::mcherry Fluorescence Relative to (1) **** (1) (2) mcherry mcherry (3) (4) F1 mcherry(+/-) F1 div-1(or148/) mcherry(+/-) daf-21p::mcherry Fluorescence Relative to (1) ns (3) (4) ; mcherry (5) (6) div-1(or148/) F1 mcherry (+/-) F1 ; mcherry div-1(or148/or148) mcherry (+/-) daf-21p::mcherry Fluorescence Relative to (1) ns (5) (6) Supplementary figure 3 fig. S3. Maternal div-1 deficiency results in elevated transgene expression in the offspring. The indicated crosses were performed in parallel and the resulting male cross-progeny imaged and mcherry expression quantified. Images show representative F1 animals with their genotype indicated. The genotype of the parents is given above each image. Animals were paired such that the daf-21p::mcherry transgene in the F1 worms is derived from the same population of parents. Sample size: (1) 51, (2) 57, (3) 48, (4) 45, (5) 50, (6) 54.
5 mes-2 (bn11); met-2 (n4256); set-25 (n5021) control (RNAi) pole-2 (RNAi) control (RNAi) pole-2 (RNAi) same contrast adjusted contrast control (RNAi) pole-2 (RNAi) Relative daf-21p::mcherry Fluorescence **** 4.3 x RNAi: control pole-2 adjusted contrast mes-2 (bn11); met-2 (n4256); set-25 (n5021) control (RNAi) pole-2 (RNAi) Relative daf-21p::mcherry Fluorescence mes-2 (bn11) met-2 (n4256) set-25 (n5021) **** 1.8 x RNAi: control pole-2 fig. S4. Transgene up-regulation following pole-2 knockdown is suppressed in the mes-2;met-2;set-25 triple-mutant background. Representative images and quantification of daf-21p::mcherry transgene expression in the hatched progeny derived from parents of the indicated genotypes fed for 24 hours with either control or pole-2 RNAi. Sample size: (control, 239, pole-2, 282), mes-2, met-2, set-25 (control, 153, pole- 2, 72).
6 fig. S5. Impaired DNA replication reduces H3K27me3 levels on multiple loci. H3K27me3 was quantified by ChIP-QPCR at the indicated loci using the primers listed in Supplementary table 3 in and div-1 mutant gravid adults. P-values calculated using two-sided T-test.
7 H3K27me3 H3K4me3 DAPI H3K9me3 DAPI DAPI DAPI H3K36me3 H3K27me3 Relative Fluorescence Intensity p= n: H3K9me3 Relative Fluorescence Intensity p= n: H3K4me3 Relative Fluorescence Intensity p= n: H3K36me3 Relative Fluorescence Intensity p= n: fig. S6. Global reduction of repressive histone marks and a gain of activating histone marks in late embryos. Representative images and quantification of histone modification levels in and nuclei of late embryos. Fold change relative to and p-values (two sided t-test) indicated for each comparison.
8 H3K4me3 H3K27me3 A DAPI H3K27me3 control pole-2 Relative Fluorescence Intensity H3K27me3 1.3x p= B DAPI control pole-2 Relative Fluorescence Intensity control pole-2 n: H3K4me3 1.6x p= control pole-2 n: fig. S7. Knockdown of pole-2 results in reduction of H3K27me3 mark and increase in H3K4me3 in early embryonic chromatin. Representative images and quantification of histone modification levels in control and pole-2 (RNAi) early embryos. Average of each embryo after subtracting the background is plotted. Fold change relative to control and p-values (two sided t-test) indicated for each comparison.
9 fig. S8. Reduction in H3K9me3 mark in mutant L1s detected by Western blot. Quantification of the western blot using anti-h3k9me3 antibody (ab8898) and anti-h3(ab1791). Samples are from synchronised and L1s.
10 F1 F2 F3 F4 div-1 (or148/or148) div-1 (or148/+) mcherry (+/-) mcherry (+/-) mcherry (+/-) mcherry (+/-) Relative Fluorescence Intensity p= 1.3e-10 p= 0 F5 F6 F1 F div-1 (or148/or148) 2.5 (+/+) RRY (+/-) (+/+) RRY (+/-) (+/+) RRY (+/+) (+/+) RRY (+/+) div-1 (or148/+) mcherry (+/-) mcherry (+/-) Relative Fluorescence Intensity 1.5 p= 1.3e-10 p= p= p= p= 9 (+/+) RRY (+/+) (+/+) RRY (+/+) F1 F3 F4 F5 F6 fig. S9. Passage of the transgenic array through impaired replication for a single generation is sufficient to trigger a multigenerational effect. Crossing scheme and quantification of transgene expression. Male worms homozygous for the daf- 21p::mCherry array where crossed to hermaphrodites carrying either a wild-type (denoted as div-1 (+)) or mutant allele. Expression was quantified in hermaphrodite F1 progeny of both crosses and the F1 males where crossed to wild-type hermaphrodites. Single F2 cross-progeny L4 hermaphrodites expressing mcherry where transferred to separate wells. Two days later, adults worms were removed and genotyped and F3 progeny of worms identified as wild-type were subsequently followed
11 and analyzed. From each well identified as wild-type, six F3 hermaphrodites at L4 stage were isolated and segregation of mcherry examined in their offspring to identify F3 animals that were homozygous for mcherry. Equal number of F4 progeny from each identified F3 mcherry (+) animal were pooled and used in the measurement of fluorescence intensity (at F4) and to generate subsequent generations. Expression in the F3 animals was compared based on fluorescence intensity in the pharynx to circumvent the high noise of expression observed in old animals. In F1, F4, F5 and F6 generations, expression was measured in whole animals (mean fluorescence intensity).
12 Relative Fluorescence Intensity H3K27me3 interphase 1.5 x p= fig. S10. Quantification of H3K27me3 in interphase nuclei. Only interphase nuclei in each embryo from Figure 3 were taken into account to calculate the H3K27me3 fluorescence intensity average that was plotted. Fold -change relative to and p-values (two sided t-test) are indicated.
13 table S1 (Part 1 of 3). List of genes whose knockdown results in upregulation of daf- 21p::mCherry transgene. Most affected generation indication of the generation at which the strongest effect of knockdown on transgene expression was observed, G0 in parental generation, G1 in offspring, G0, G1 equally in parents and offspring. Last column indicates overlap with following RNAi screens: T - Towbin et al. 2012, R - Robert et al. 2005, N - Nollen et al. 2004, L - Lamitina et al. 2006, G - Guisbert et al
14 table S1 (Part 2 of 3).
15 table S1 (Part 3 of 3). F58A4.5 clec>161 Collectin Other G0, G1 F58E2.9 srz$23& Serpentine receptor class Z Other G0 L H19N07.1 erfa>3 Ortholog of human GSPT2&GSPT1 Other G0 R, N K08D12.1 pbs$1& Protease subunt Other G0 N K08E4.1 spt$5& Transcription factor Other G0 L K12C11.2 smo$1& Ortholog of SUMO Other G0, G1 M28.5 phi$9& Small nuclear ribonucleoprotein Other G0 N M7.1 let$70& Class I E2 ubiquitination enzyme Other G0 R07G3.1 cdc$42& RHO GTPase Other G1 R12B2.5 mdt$15& Mediator subunit Other G0 G W01D2.2 nhr$61& DNA binding transcription factor Other G0 N W09G12.5 F38A1.8 Signal recognition particle receptor Other G0 ZK328.5 npp$10& Nuclear core complex Other G0 Supplementary(Table(1.(List(of(genes(whose(knockdown(result(in(upregulation(of(daf$ table S2. C. elegans strains used in this study. Strain name N2 EU538 MT17463 MT13293 SS186 CB1301 BCN1049 BCN1050 BCN6101 BCN6105 BCN6112 BCN6123 BCN6129 BCN6130 Genotype wild-type III set-25(n5021) III met-2(n4256) III mes-2(bn11) unc-4(e120)/mnc1 dpy-10(e128) unc- 52(e444) II unc-54(e1301) I crgis1004[daf-21p::gfp::unc- 54 3'UTR; unc-119(+)] crgis1002[daf-21p::mcherry::unc-54 3'UTR; unc-119(+)] III crgis1002 IV set-25(n5021) III crgis1002 IV met-2(n4256) III crgis1002 IV met-2(n4256) set-25(n5021) III crgis1002 IV mes-2(bn11) unc-4(e120)/mnc1 dpy-10(e128) unc- 52(e444)II crgis1002 IV mes-2(bn11) unc-4(e120)/mnc1 dpy-10(e128) unc- 52(e444)II met-2(n4256) set-25(n5021) III crgis1002 IV Supplementary!Table!2.!Strains!used!in!this!study.!!
16 table S3. Primers used in qpcr analyses. pdaf-21 fwd: GCAGCATCTTCTTCGTCCTC, pdaf-21 rev GAAAAATTGAGGGCAGGTGA mcherry fwd: AAGGGCGAGGAGGATAACAT mcherry rev: ACATGAACTGAGGGGACAGG ppmp-3 fwd: TGTTCACTCACAGCCAGCTC, ppmp-3 rev: ACCATCCCATTCAAACCAAA. pcdc-42 fwd: AGTTGTTTTGGCCATTTTGC, pcdc-42 rev: TGAAAACGAATTGCGAAACA. cdc-42 fwd: GCCTGAAATTTCGCATCATT, cdc-42 rev: TCCTTTGCCAACTTCTCTCC sra-25 fwd: ATCCCACTACAACCCCAGGT, sra-25 rev: GACTACCGTGCGGAAATCAT.! table S4. Transgenes tested for derepression with div-1(rnai).
Nature Genetics: doi: /ng Supplementary Figure 1. Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms.
 Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Supplementary Figure 1 hlrrk2 promotes CAP dependent protein translation.
 ` Supplementary Figure 1 hlrrk2 promotes CAP dependent protein translation. (a) Overexpression of hlrrk2 in HeLa cells enhances total protein synthesis in [35S] methionine/cysteine incorporation assays.
` Supplementary Figure 1 hlrrk2 promotes CAP dependent protein translation. (a) Overexpression of hlrrk2 in HeLa cells enhances total protein synthesis in [35S] methionine/cysteine incorporation assays.
Neuronal GPCR OCTR-1 regulates innate immunity by controlling protein synthesis in
 Supplementary Information Neuronal GPCR OCTR-1 regulates innate immunity by controlling protein synthesis in Caenorhabditis elegans Yiyong Liu 1, Durai Sellegounder 1, and Jingru Sun 1 * 1 Department of
Supplementary Information Neuronal GPCR OCTR-1 regulates innate immunity by controlling protein synthesis in Caenorhabditis elegans Yiyong Liu 1, Durai Sellegounder 1, and Jingru Sun 1 * 1 Department of
Supplementary Information
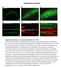 Supplementary Information Supplementary Figure 1: Luminal localization of CCM-3. (a) The CCM-3::GFP fusion protein localizes along the apical (luminal) surface of the pharynx (b) as well as the lumen of
Supplementary Information Supplementary Figure 1: Luminal localization of CCM-3. (a) The CCM-3::GFP fusion protein localizes along the apical (luminal) surface of the pharynx (b) as well as the lumen of
SUPPLEMENTARY FIGURE S1: nlp-22 is expressed in the RIA interneurons and is secreted. (a) An animal expressing both the RIA specific reporter
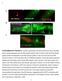 1 SUPPLEMENTARY FIGURE S1: nlp-22 is expressed in the RIA interneurons and is secreted. (a) An animal expressing both the RIA specific reporter Pglr-3:mCherry (red) and Pnlp-22:gfp (green) shows co-localization
1 SUPPLEMENTARY FIGURE S1: nlp-22 is expressed in the RIA interneurons and is secreted. (a) An animal expressing both the RIA specific reporter Pglr-3:mCherry (red) and Pnlp-22:gfp (green) shows co-localization
C. elegans Embryonic Development
 Autonomous Specification in Tunicate development Autonomous & Conditional Specification in C. elegans Embryonic Development Figure 8.36 Bilateral Symmetry in the Egg of the Tunicate Styela partita Fig.
Autonomous Specification in Tunicate development Autonomous & Conditional Specification in C. elegans Embryonic Development Figure 8.36 Bilateral Symmetry in the Egg of the Tunicate Styela partita Fig.
A role for WDR5 in TRA-1/Gli mediated transcriptional control of the sperm/oocyte switch in C. elegans
 A role for WDR5 in TRA-1/Gli mediated transcriptional control of the sperm/oocyte switch in C. elegans Tengguo Li, Emory University William G Kelly, Emory University Journal Title: Nucleic Acids Research
A role for WDR5 in TRA-1/Gli mediated transcriptional control of the sperm/oocyte switch in C. elegans Tengguo Li, Emory University William G Kelly, Emory University Journal Title: Nucleic Acids Research
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
Phenomena first observed in petunia
 Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
klp-18 (RNAi) Control. supplementary information. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach
![klp-18 (RNAi) Control. supplementary information. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach klp-18 (RNAi) Control. supplementary information. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach](/thumbs/91/104639484.jpg) DOI: 10.1038/ncb1891 A. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach embryos let hatch overnight transfer to RNAi plates; incubate 5 days at 15 C RNAi food L1 worms adult worms
DOI: 10.1038/ncb1891 A. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach embryos let hatch overnight transfer to RNAi plates; incubate 5 days at 15 C RNAi food L1 worms adult worms
A Role for Set1/MLL-Related Components in Epigenetic Regulation of the Caenorhabditis elegans Germ Line
 A Role for Set1/MLL-Related Components in Epigenetic Regulation of the Caenorhabditis elegans Germ Line Tengguo Li, Emory University William G Kelly, Emory University Journal Title: PLoS Genetics Volume:
A Role for Set1/MLL-Related Components in Epigenetic Regulation of the Caenorhabditis elegans Germ Line Tengguo Li, Emory University William G Kelly, Emory University Journal Title: PLoS Genetics Volume:
Title: Epigenetic mechanisms underlying maternal diabetes-associated risk of congenital heart disease
 1 Supplemental Materials 2 3 Title: Epigenetic mechanisms underlying maternal diabetes-associated risk of congenital heart disease 4 5 6 Authors: Madhumita Basu, 1 Jun-Yi Zhu, 2 Stephanie LaHaye 1,3, Uddalak
1 Supplemental Materials 2 3 Title: Epigenetic mechanisms underlying maternal diabetes-associated risk of congenital heart disease 4 5 6 Authors: Madhumita Basu, 1 Jun-Yi Zhu, 2 Stephanie LaHaye 1,3, Uddalak
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Generation and validation of mtef4-knockout mice.
 Supplementary Figure 1 Generation and validation of mtef4-knockout mice. (a) Alignment of EF4 (E. coli) with mouse, yeast and human EF4. (b) Domain structures of mouse mtef4 compared to those of EF4 (E.
Supplementary Figure 1 Generation and validation of mtef4-knockout mice. (a) Alignment of EF4 (E. coli) with mouse, yeast and human EF4. (b) Domain structures of mouse mtef4 compared to those of EF4 (E.
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Supplemental Figures
 Supplemental Figures Supplemental Figure 1. Fasting-dependent regulation of the SREBP ortholog SBP-1 and lipid homeostasis mediated by the SIRT1 ortholog SIR-2.1 in C. elegans. (A) Wild-type or sir-2.1(lof)
Supplemental Figures Supplemental Figure 1. Fasting-dependent regulation of the SREBP ortholog SBP-1 and lipid homeostasis mediated by the SIRT1 ortholog SIR-2.1 in C. elegans. (A) Wild-type or sir-2.1(lof)
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Programmed Cell Death (apoptosis)
 Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
Low Levels of p53 Protein and Chromatin Silencing of p53 Target Genes Repress Apoptosis in Drosophila Endocycling Cells
 Low Levels of p53 Protein and Chromatin Silencing of p53 Target Genes Repress Apoptosis in Drosophila Endocycling Cells Bingqing Zhang, Sonam Mehrotra, Wei Lun Ng, Brian R. Calvi* Department of Biology,
Low Levels of p53 Protein and Chromatin Silencing of p53 Target Genes Repress Apoptosis in Drosophila Endocycling Cells Bingqing Zhang, Sonam Mehrotra, Wei Lun Ng, Brian R. Calvi* Department of Biology,
Table S1. Total and mapped reads produced for each ChIP-seq sample
 Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Diabetic pdx1-mutant zebrafish show conserved responses to nutrient overload and anti-glycemic treatment
 Supplementary Information Diabetic pdx1-mutant zebrafish show conserved responses to nutrient overload and anti-glycemic treatment Robin A. Kimmel, Stefan Dobler, Nicole Schmitner, Tanja Walsen, Julia
Supplementary Information Diabetic pdx1-mutant zebrafish show conserved responses to nutrient overload and anti-glycemic treatment Robin A. Kimmel, Stefan Dobler, Nicole Schmitner, Tanja Walsen, Julia
Cell Cycle, Mitosis, and Microtubules. LS1A Final Exam Review Friday 1/12/07. Processes occurring during cell cycle
 Cell Cycle, Mitosis, and Microtubules LS1A Final Exam Review Friday 1/12/07 Processes occurring during cell cycle Replicate chromosomes Segregate chromosomes Cell divides Cell grows Cell Growth 1 The standard
Cell Cycle, Mitosis, and Microtubules LS1A Final Exam Review Friday 1/12/07 Processes occurring during cell cycle Replicate chromosomes Segregate chromosomes Cell divides Cell grows Cell Growth 1 The standard
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Nature Medicine: doi: /nm.4322
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
Transgenerational epigenetics in the germline cycle of Caenorhabditis elegans
 Transgenerational epigenetics in the germline cycle of Caenorhabditis elegans William G Kelly, Emory University Journal Title: Epigenetics and Chromatin Volume: Volume 7, Number 6 Publisher: BioMed Central
Transgenerational epigenetics in the germline cycle of Caenorhabditis elegans William G Kelly, Emory University Journal Title: Epigenetics and Chromatin Volume: Volume 7, Number 6 Publisher: BioMed Central
SUPPLEMENTARY INFORMATION
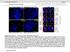 doi: 10.1038/nature05473 SUPPLEMENTARY FIGURES SUPPLEMENTARY INFORMATION Supplementary Figure 1: Association of Runx2 with mitotic chromosomes Mitotic chromosome spreads were prepared for a. Human Saos-2
doi: 10.1038/nature05473 SUPPLEMENTARY FIGURES SUPPLEMENTARY INFORMATION Supplementary Figure 1: Association of Runx2 with mitotic chromosomes Mitotic chromosome spreads were prepared for a. Human Saos-2
This is a published version of a paper published in PLoS genetics. Access to the published version may require subscription.
 Umeå University This is a published version of a paper published in PLoS genetics. Citation for the published paper: Holmqvist, P., Boija, A., Philip, P., Crona, F., Stenberg, P. et al. (2012) "Preferential
Umeå University This is a published version of a paper published in PLoS genetics. Citation for the published paper: Holmqvist, P., Boija, A., Philip, P., Crona, F., Stenberg, P. et al. (2012) "Preferential
Epigenetic Patterns Maintained in Early Caenorhabditis elegans Embryos Can Be Established by Gene Activity in the Parental Germ Cells
 Epigenetic Patterns Maintained in Early Caenorhabditis elegans Embryos Can Be Established by Gene Activity in the Parental Germ Cells Jackelyn K. Arico, Emory University David J Katz, Emory University
Epigenetic Patterns Maintained in Early Caenorhabditis elegans Embryos Can Be Established by Gene Activity in the Parental Germ Cells Jackelyn K. Arico, Emory University David J Katz, Emory University
Expanded View Figures
 PEX13 functions in selective autophagy Ming Y Lee et al Expanded View Figures Figure EV1. PEX13 is required for Sindbis virophagy. A, B Quantification of mcherry-capsid puncta per cell (A) and GFP-LC3
PEX13 functions in selective autophagy Ming Y Lee et al Expanded View Figures Figure EV1. PEX13 is required for Sindbis virophagy. A, B Quantification of mcherry-capsid puncta per cell (A) and GFP-LC3
Supplementary Figure 1. Gene schematics of hyls-1, gasr-8 and k10g6.4, and TEM analysis of TFs in WT and hyls-1 cilia. (a) Gene structure of hyls-1,
 Supplementary Figure 1. Gene schematics of hyls-1, gasr-8 and k10g6.4, and TEM analysis of TFs in WT and hyls-1 cilia. (a) Gene structure of hyls-1, gasr-8 and k10g6.4 based on WormBase (http://wormbase.org),
Supplementary Figure 1. Gene schematics of hyls-1, gasr-8 and k10g6.4, and TEM analysis of TFs in WT and hyls-1 cilia. (a) Gene structure of hyls-1, gasr-8 and k10g6.4 based on WormBase (http://wormbase.org),
BRaf V600E cooperates with Pten silencing to elicit metastatic melanoma (Nature Genetics Supplementary Information)
 BRaf V600E cooperates with Pten silencing to elicit metastatic melanoma (Nature Genetics Supplementary Information) David Dankort, David P. Curley, Robert A. Cartlidge, Betsy Nelson, Anthony N. Karnezis,
BRaf V600E cooperates with Pten silencing to elicit metastatic melanoma (Nature Genetics Supplementary Information) David Dankort, David P. Curley, Robert A. Cartlidge, Betsy Nelson, Anthony N. Karnezis,
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
SUPPLEMENTARY FIGURES 1 Supplementary Figure 1, Adult hippocampal QNPs and TAPs uniformly express REST a-b) Confocal images of adult hippocampal mouse sections showing GFAP (green), Sox2 (red), and REST
Supplementary Figure 1. mir124 does not change neuron morphology and synaptic
 Supplementary Figure 1. mir124 does not change neuron morphology and synaptic density. Hippocampal neurons were transfected with mir124 (containing DsRed) or DsRed as a control. 2 d after transfection,
Supplementary Figure 1. mir124 does not change neuron morphology and synaptic density. Hippocampal neurons were transfected with mir124 (containing DsRed) or DsRed as a control. 2 d after transfection,
Early Embryonic Development
 Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
oocytes were pooled for RT-PCR analysis. The number of PCR cycles was 35. Two
 Supplementary Fig. 1 a Kdm3a Kdm4b β-actin Oocyte Testis Oocyte Testis Oocyte Testis b 1.8 Relative expression.6.4.2 Kdm3a Kdm4b RT-PCR analysis of Kdm3a and Kdm4b expression in oocytes and testes. Twenty
Supplementary Fig. 1 a Kdm3a Kdm4b β-actin Oocyte Testis Oocyte Testis Oocyte Testis b 1.8 Relative expression.6.4.2 Kdm3a Kdm4b RT-PCR analysis of Kdm3a and Kdm4b expression in oocytes and testes. Twenty
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
advances.sciencemag.org/cgi/content/full/1/8/e1500296/dc1 Supplementary Materials for Transcriptional regulation of APOBEC3 antiviral immunity through the CBF- /RUNX axis This PDF file includes: Brett
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Left, western blot analysis of ISGylated proteins in Jurkat T cells treated with 1000U ml -1 IFN for 16h (IFN) or left untreated (CONT); right, western
SUPPLEMENTARY FIGURES Supplementary Figure 1. (A) Left, western blot analysis of ISGylated proteins in Jurkat T cells treated with 1000U ml -1 IFN for 16h (IFN) or left untreated (CONT); right, western
Concentration memory-dependent synaptic plasticity of a taste circuit regulates salt chemotaxis in Caenorhabditis elegans
 Supplementary Information for: Concentration memory-dependent synaptic plasticity of a taste circuit regulates salt chemotaxis in Caenorhabditis elegans Hirofumi Kunitomo, Hirofumi Sato, Ryo Iwata, Yohsuke
Supplementary Information for: Concentration memory-dependent synaptic plasticity of a taste circuit regulates salt chemotaxis in Caenorhabditis elegans Hirofumi Kunitomo, Hirofumi Sato, Ryo Iwata, Yohsuke
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
Supplementary information. The Light Intermediate Chain 2 Subpopulation of Dynein Regulates Mitotic. Spindle Orientation
 Supplementary information The Light Intermediate Chain 2 Subpopulation of Dynein Regulates Mitotic Spindle Orientation Running title: Dynein LICs distribute mitotic functions. Sagar Mahale a, d, *, Megha
Supplementary information The Light Intermediate Chain 2 Subpopulation of Dynein Regulates Mitotic Spindle Orientation Running title: Dynein LICs distribute mitotic functions. Sagar Mahale a, d, *, Megha
Soluble ADAM33 initiates airway remodeling to promote susceptibility for. Elizabeth R. Davies, Joanne F.C. Kelly, Peter H. Howarth, David I Wilson,
 Revised Suppl. Data: Soluble ADAM33 1 Soluble ADAM33 initiates airway remodeling to promote susceptibility for allergic asthma in early life Elizabeth R. Davies, Joanne F.C. Kelly, Peter H. Howarth, David
Revised Suppl. Data: Soluble ADAM33 1 Soluble ADAM33 initiates airway remodeling to promote susceptibility for allergic asthma in early life Elizabeth R. Davies, Joanne F.C. Kelly, Peter H. Howarth, David
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10643 Supplementary Table 1. Identification of hecw-1 coding polymorphisms at amino acid positions 322 and 325 in 162 strains of C. elegans. WWW.NATURE.COM/NATURE 1 Supplementary Figure
doi:10.1038/nature10643 Supplementary Table 1. Identification of hecw-1 coding polymorphisms at amino acid positions 322 and 325 in 162 strains of C. elegans. WWW.NATURE.COM/NATURE 1 Supplementary Figure
Supplementary Figure 1
 Supplementary Figure 1 Dox Cis Cam Pac 0 15 1 15 1 15 1 15 1 15 µmole/l Ub p53 Cytotoxic anticancer agents increase p53 levels but do not generally promote the accumulation of ubiquitinated. Western blots
Supplementary Figure 1 Dox Cis Cam Pac 0 15 1 15 1 15 1 15 1 15 µmole/l Ub p53 Cytotoxic anticancer agents increase p53 levels but do not generally promote the accumulation of ubiquitinated. Western blots
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells
 Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Reduced PRC2 function alters male germline epigenetic programming and paternal inheritance
 Stringer et al. BMC Biology (2018) 16:104 https://doi.org/10.1186/s12915-018-0569-5 RESEARCH ARTICLE Reduced PRC2 function alters male germline epigenetic programming and paternal inheritance Open Access
Stringer et al. BMC Biology (2018) 16:104 https://doi.org/10.1186/s12915-018-0569-5 RESEARCH ARTICLE Reduced PRC2 function alters male germline epigenetic programming and paternal inheritance Open Access
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated
 Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1 Transcription assay of nine ABA-responsive PP2C genes. Transcription assay of nine ABA-responsive PP2C genes. Total RNA was isolated from 7 day-old seedlings treated with or without
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.
 Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.5 and E13.5 prepared from uteri of dams and subsequently genotyped.
Supplementary Figure 1: Hsp60 / IEC mice are embryonically lethal (A) Light microscopic pictures show mouse embryos at developmental stage E12.5 and E13.5 prepared from uteri of dams and subsequently genotyped.
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE
 Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
BIO360 Quiz #1. September 14, Name five of the six Hallmarks of Cancer (not emerging hallmarks or enabling characteristics ): (5 points)
 Name: BIO360 Quiz #1 September 14, 2012 1. Name five of the six Hallmarks of Cancer (not emerging hallmarks or enabling characteristics ): (5 points) 2. The controversial hypothesis that only a small subset
Name: BIO360 Quiz #1 September 14, 2012 1. Name five of the six Hallmarks of Cancer (not emerging hallmarks or enabling characteristics ): (5 points) 2. The controversial hypothesis that only a small subset
Supplementary Figures
 J. Cell Sci. 128: doi:10.1242/jcs.173807: Supplementary Material Supplementary Figures Fig. S1 Fig. S1. Description and/or validation of reagents used. All panels show Drosophila tissues oriented with
J. Cell Sci. 128: doi:10.1242/jcs.173807: Supplementary Material Supplementary Figures Fig. S1 Fig. S1. Description and/or validation of reagents used. All panels show Drosophila tissues oriented with
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
MII. Supplement Figure 1. CapZ β2. Merge. 250ng. 500ng DIC. Merge. Journal of Cell Science Supplementary Material. GFP-CapZ β2 DNA
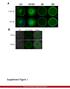 A GV GVBD MI DNA CapZ β2 CapZ β2 Merge B DIC GFP-CapZ β2 Merge CapZ β2-gfp 250ng 500ng Supplement Figure 1. MII A early MI late MI Control RNAi CapZαβ DNA Actin Tubulin B Phalloidin Intensity(A.U.) n=10
A GV GVBD MI DNA CapZ β2 CapZ β2 Merge B DIC GFP-CapZ β2 Merge CapZ β2-gfp 250ng 500ng Supplement Figure 1. MII A early MI late MI Control RNAi CapZαβ DNA Actin Tubulin B Phalloidin Intensity(A.U.) n=10
Article. An EDMD Mutation in C. elegans Lamin Blocks Muscle-Specific Gene Relocation and Compromises Muscle Integrity
 Current Biology 21, 163 1614, October 11, 211 ª211 Elsevier Ltd All rights reserved DOI 1.116/j.cub.211.8.3 An EDMD Mutation in C. elegans Lamin Blocks Muscle-Specific Gene Relocation and Compromises Muscle
Current Biology 21, 163 1614, October 11, 211 ª211 Elsevier Ltd All rights reserved DOI 1.116/j.cub.211.8.3 An EDMD Mutation in C. elegans Lamin Blocks Muscle-Specific Gene Relocation and Compromises Muscle
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Development Supplementary information. Supplementary Figures * * +/+ +/- -/- +/+ +/- -/-
 Development 144: doi:1.1242/dev.1473: Supplementary information Supplementary Figures A (f) FRT LoxP 2 3 4 B All Males Females I Ovary 1 (+) 77 bps (f) 78 bps (-) >13 bps (-) 2 4 (-) 424 bps M +/f +/-
Development 144: doi:1.1242/dev.1473: Supplementary information Supplementary Figures A (f) FRT LoxP 2 3 4 B All Males Females I Ovary 1 (+) 77 bps (f) 78 bps (-) >13 bps (-) 2 4 (-) 424 bps M +/f +/-
Relative activity (%) SC35M
 a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
Development of safe and immunogenic reassortant viruses with 5:3 genotype for live attenuated influenza vaccine
 Development of safe and immunogenic reassortant viruses with 5:3 genotype for live attenuated influenza vaccine Irina Isakova-Sivak, PhD Institute of Experimental Medicine, Saint Petersburg, Russia The
Development of safe and immunogenic reassortant viruses with 5:3 genotype for live attenuated influenza vaccine Irina Isakova-Sivak, PhD Institute of Experimental Medicine, Saint Petersburg, Russia The
Supplemental Figure 1. Western blot analysis indicated that MIF was detected in the fractions of
 Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
BIOL2005 WORKSHEET 2008
 BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
Today. Genomic Imprinting & X-Inactivation
 Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier
 Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
(A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC,
 Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure Legends Figure S1. Cell line characterization (A) Cells grown in monolayer were fixed and stained for surfactant protein-c (SPC, green) and co-stained with DAPI to visualize the nuclei.
Supplemental Figure 1: Asymmetric chromatin maturation leads to epigenetic asymmetries on sister chromatids.
 Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
marker. DAPI labels nuclei. Flies were 20 days old. Scale bar is 5 µm. Ctrl is
 Supplementary Figure 1. (a) Nos is detected in glial cells in both control and GFAP R79H transgenic flies (arrows), but not in deletion mutant Nos Δ15 animals. Repo is a glial cell marker. DAPI labels
Supplementary Figure 1. (a) Nos is detected in glial cells in both control and GFAP R79H transgenic flies (arrows), but not in deletion mutant Nos Δ15 animals. Repo is a glial cell marker. DAPI labels
EGFR shrna A: CCGGCGCAAGTGTAAGAAGTGCGAACTCGAGTTCGCACTTCTTACACTTGCG TTTTTG. EGFR shrna B: CCGGAGAATGTGGAATACCTAAGGCTCGAGCCTTAGGTATTCCACATTCTCTT TTTG
 Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
Supplementary Methods Sequence of oligonucleotides used for shrna targeting EGFR EGFR shrna were obtained from the Harvard RNAi consortium. The following oligonucleotides (forward primer) were used to
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent Protein.
 prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
prfp-vector RFP Exon1 Intron RFP Exon2 prfp-mir-124 mir-93/124 RFP Exon1 Intron RFP Exon2 Untransfected prfp-vector prfp-mir-93 prfp-mir-124 Supplementary Fig. 1. Delivery of mirnas via Red Fluorescent
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Sexual Reproduction and Genetics. Section 1. Meiosis
 Chromosomes and Chromosome Number! Human body cells have 46 chromosomes! Each parent contributes 23 chromosomes! Homologous chromosomes one of two paired chromosomes, one from each parent Chromosomes and
Chromosomes and Chromosome Number! Human body cells have 46 chromosomes! Each parent contributes 23 chromosomes! Homologous chromosomes one of two paired chromosomes, one from each parent Chromosomes and
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Alternative Splicing Regulation During C. elegans Development: Splicing Factors as Regulated Targets
 Alternative Splicing Regulation During C. elegans Development: Splicing Factors as Regulated Targets Sergio Barberan-Soler 1,2, Alan M. Zahler 1,2 * 1 Department of MCD Biology, University of California
Alternative Splicing Regulation During C. elegans Development: Splicing Factors as Regulated Targets Sergio Barberan-Soler 1,2, Alan M. Zahler 1,2 * 1 Department of MCD Biology, University of California
Carol Garvin, Richard Holdeman and Susan Strome
 Copyright 1998 by the Genetics Society of America The Phenotype of mes-2, mes-3, mes-4 and mes-6, Maternal-Effect Genes Required for Survival of the Germline in Caenorhabditis elegans, Is Sensitive to
Copyright 1998 by the Genetics Society of America The Phenotype of mes-2, mes-3, mes-4 and mes-6, Maternal-Effect Genes Required for Survival of the Germline in Caenorhabditis elegans, Is Sensitive to
ns ns hp761(lf); daf-28(lf) daf-28(gf) Figure S1
 A ns ns B 100 100 80 80 60 60 40 40 20 20 0 0% 0 0% hp761(lf); daf-28(lf) daf-28(gf) Figure S1 A 100 80 15 o C 22 o C 25 o C % Dauers 60 40 20 0 0% 0% * 0% 0% 0% Class I alleles Class II alleles daf-2(lf;ts)
A ns ns B 100 100 80 80 60 60 40 40 20 20 0 0% 0 0% hp761(lf); daf-28(lf) daf-28(gf) Figure S1 A 100 80 15 o C 22 o C 25 o C % Dauers 60 40 20 0 0% 0% * 0% 0% 0% Class I alleles Class II alleles daf-2(lf;ts)
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/308/ra4/dc1 Supplementary Materials for Antipsychotics Activate mtorc1-dependent Translation to Enhance Neuronal Morphological Complexity Heather Bowling, Guoan
www.sciencesignaling.org/cgi/content/full/7/308/ra4/dc1 Supplementary Materials for Antipsychotics Activate mtorc1-dependent Translation to Enhance Neuronal Morphological Complexity Heather Bowling, Guoan
Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion
 Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Supplementary Figure S1. Effects of UBL5 knockdown on cell cycle distribution and sister chromatid cohesion A. Representative examples of flow cytometry profiles of HeLa cells transfected with indicated
Histone acetyltransferase CBP-related H3K23 acetylation contributes to courtship learning in Drosophila
 Li et al. BMC Developmental Biology (2018) 18:20 https://doi.org/10.1186/s12861-018-0179-z RESEARCH ARTICLE Open Access Histone acetyltransferase CBP-related H3K23 acetylation contributes to courtship
Li et al. BMC Developmental Biology (2018) 18:20 https://doi.org/10.1186/s12861-018-0179-z RESEARCH ARTICLE Open Access Histone acetyltransferase CBP-related H3K23 acetylation contributes to courtship
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Differential expression of mirnas from the pri-mir-17-92a locus.
 Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 Differential expression of mirnas from the pri-mir-17-92a locus. (a) The mir-17-92a expression unit in the third intron of the host mir-17hg transcript. (b,c) Impact of knockdown
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was
 Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Paternal RNA contributions in the C. elegans zygote
 Manuscript EMBO-2014-88117 Paternal RNA contributions in the C. elegans zygote Marlon Stoeckius, Dominic Grün and Nikolaus Rajewsky Corresponding author: Nikolaus Rajewsky, Max Delbrueck Centrum fuer Molekulare
Manuscript EMBO-2014-88117 Paternal RNA contributions in the C. elegans zygote Marlon Stoeckius, Dominic Grün and Nikolaus Rajewsky Corresponding author: Nikolaus Rajewsky, Max Delbrueck Centrum fuer Molekulare
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
SUPPLEMENTARY INFORMATION
 DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
DOI: 1.138/ncb3355 a S1A8 + cells/ total.1.8.6.4.2 b S1A8/?-Actin c % T-cell proliferation 3 25 2 15 1 5 T cells Supplementary Figure 1 Inter-tumoral heterogeneity of MDSC accumulation in mammary tumor
Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v)
 SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
SUPPLEMENTARY MATERIAL AND METHODS Soft Agar Assay. For each cell pool, 100,000 cells were resuspended in 0.35% (w/v) top agar (LONZA, SeaKem LE Agarose cat.5004) and plated onto 0.5% (w/v) basal agar.
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC)
 Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Thursday 5 November EU-India PARTNERING EVENT Theme: Health RESEARCHER S NAME: Làszlò Tora RESEARCHER S ORGANISATION: Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) CNRS, INSERM,
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Supporting Information
 Supporting Information Fig. S1. Overexpression of Rpr causes progenitor cell death. (A) TUNEL assay of control intestines. No progenitor cell death could be observed, except that some ECs are undergoing
Supporting Information Fig. S1. Overexpression of Rpr causes progenitor cell death. (A) TUNEL assay of control intestines. No progenitor cell death could be observed, except that some ECs are undergoing
