Fine tuning of coenzyme specificity in family 2 aldo-keto reductases revealed by crystal structures of the Lys-274 fi Arg mutant of
|
|
|
- Mariah York
- 6 years ago
- Views:
Transcription
1 FEBS FEBS Letters 579 (2005) Fine tuning of coenzyme specificity in family 2 aldo-keto reductases revealed by crystal structures of the Lys-274 fi Arg mutant of Candida tenuis xylose reductase (AKR2B5) bound to NAD + and NADP + Stefan Leitgeb a,b, Barbara Petschacher a, David K. Wilson b, Bernd Nidetzky a, * a Institute of Biotechnology and Biochemical Engineering, Graz University of Technology, Petersgasse 12/I, A-8010 Graz, Austria b Section of Molecular and Cellular Biology, University of California, Davis, CA 95616, USA Received 5 November 2004; revised 10 December 2004; accepted 15 December 2004 Available online 11 January 2005 Edited by Hans Eklund Abstract Aldo-keto reductases of family 2 employ single site replacement Lys fi Arg to switch their cosubstrate preference from NADPH to NADH. X-ray crystal structures of Lys- 274 fi Arg mutant of Candida tenuis xylose reductase (AKR2B5) bound to NAD + and NADP + were determined at a resolution of 2.4 and 2.3 Å, respectively. Due to steric conflicts in the NADP + -bound form, the arginine side chain must rotate away from the position of the original lysine side chain, thereby disrupting a network of direct and water-mediated interactions between Glu-227, Lys-274 and the cofactor 2 0 -phosphate and 3 0 -hydroxy groups. Because anchoring contacts of its Glu-227 are lost, the coenzyme-enfolding loop that becomes ordered upon binding of NAD(P) + in the wild-type remains partly disordered in the NADP + -bound mutant. The results delineate a catalytic reaction profile for the mutant in comparison to wild-type. Ó 2005 Federation of European Biochemical Societies. Published by Elsevier B.V. All rights reserved. Keywords: Dual coenzyme specificity; Selectivity; Loop ordering; Xylose fermentation 1. Introduction There is a fundamental distinction between NADH and NADPH in metabolic pathways. NADH is the reductant in catabolic reactions, whereas NADPH generally provides reducing power for biosynthesis. Therefore, most nicotinamide cofactor-dependent enzymes are specific for either NADH or NADPH. Xylose reductase (XR) from the yeast Candida tenuis is a dual specific NADPH- and NADHdependent enzyme that converts D-xylose into xylitol [1]. The xylitol is oxidized to yield D-xylulose which in turn is phosphorylated and enters the pentose phosphate pathway (PPP) as D-xylulose 5-phosphate. The PPP is a main source of NADPH and supplies intermediates for energy production and biosynthesis [2]. The co-substrate selectivity of the XR probably reflects the capacity of the PPP to regenerate * Corresponding author. Fax: address: bernd.nidetzky@tugraz.at (B. Nidetzky). Abbreviations: AKR, aldo-keto reductase; XR, xylose reductase; CpXR; Candida parapsilosis XR; CtXR, Candida tenuis XR; PPP, pentose phosphate pathway; r.m.s., root mean square the NADPH consumed by XR and the extent to which carbon from xylose partitions between catabolism and anabolism in C. tenuis physiology. Based on sequence homology, C. tenuis XR (CtXR) has been classified into family 2 of the aldo-keto reductase (AKR) superfamily of proteins and enzymes and is systematically identified as AKR2B5 [3]. Family 2 is unusual among the 14 AKR families currently recognized because it includes dual specific NADPH/NADH-dependent reductases as well as NADPH-specific enzymes. Crystal structures of wild-type CtXR bound to NADP + and NAD + show that a conformational change of the side chain of Glu-227, to make strong interactions with the 2 0 -OH and 3 0 -OH groups when the cofactor 2 0 -phosphate group is lacking, is most critical for utilization of NADH [4,5]. Glu-227 of CtXR is widely conserved among the dual specific AKRs of family 2. The side chain of Lys-274 forms a strong hydrogen bond with the 2 0 -phosphate group and participates in a network of water-mediated contacts with Glu-227 and the cofactor 3 0 -hydroxyl group. In the NAD + -bound conformation, the side chain of Lys-274 rotates away slightly and is re-directed to hydrogen bond with solvent molecules. Recent results of mutational studies [6] corroborate the structure-derived notion that 20-fold tighter binding of NADPH than NADH results mainly from the interactions of Lys-274, which are specific to the NADP complex. Lys-274 is conserved in all members of family 2 except for XR from Candida parapsilosis which shows a Lys fi Arg replacement and is the only natural XR known to prefer NADH [7]. A K274R mutant of CtXR utilizes NADH and NADPH with the same specificity, whereas the wild-type prefers NADPH about 33-fold [6]. To provide a molecular basis of how coenzyme selectivity in family 2 AKRs is adjusted by the substitution of Lys-274 with Arg, we determined the X-ray crystal structures of the K274R mutant of CtXR bound to NADP + and NAD Materials and methods 2.1. Protein preparation and crystallization All chemicals used were reported previously [6]. The purified K274R mutant [6] was crystallized by using the hanging drop vapor diffusion method at 25 C and a drop consisting of 1 ll well solution and 1 ll protein solution (15 mg/ml) with 2.5 mm cofactor added. Crystals of the NAD + -bound holoenzyme were obtained using a well solution of 2.0 M ammonium sulfate and 100 mm sodium citrate, ph 6.2, in the presence of NADP + the well solution contained additional 160 mm /$30.00 Ó 2005 Federation of European Biochemical Societies. Published by Elsevier B.V. All rights reserved. doi: /j.febslet
2 764 S. Leitgeb et al. / FEBS Letters 579 (2005) sodium acetate. Crystals of the NAD + - and NADP + -bound enzymes took 2 and 4 months to grow, respectively, and had final dimensions of approximately mm Data collection, structure determination and refinement The crystals were flash-cooled in a 100 K nitrogen stream in a buffer containing 80% (v/v) of the well solution, 5 mm coenzyme and 20% (v/ v) glycerol as cryo-protectant. Data were collected at the Stanford Synchrotron Radiation Laboratory beamline 9-1 on a MAR Research 345 detector and were integrated and reduced using the programs DENZO and SCALEPACK [8]. The data collection statistics along with the respective space group and lattice constants are presented in Table 1. Since the mutant enzyme unit cells were isomorphous with the wildtype, rigid body refinement using the program CNS [9] commenced with the respective native structures stripped of NADP + or NAD + and water molecules as starting models (PDB-codes 1K8C and 1MI3). Before any refinement was performed, 5% of the observed reflections were flagged for calculation of the free R-factor. The structures were refined with CNS [9] and manually refitted into the electron density with the program O [10]. Many iterations of this cycle resulted in the final structures. Waters were automatically picked by CNS [9] and manually checked for electron density and appropriate hydrogen bonding. The statistics associated with the final round of refinement are shown in Table 1. Table 1 Statistics for the crystal structures of CtXR K274R mutant bound to NAD + and NADP + K274R-NAD + K274R-NADP + Data collection Space group C2 C2 Unit cell dimensions a (Å) b (Å) c (Å) b ( ) Overall temperature factors (Å 2 ) Monomer A Monomer B Monomer C Monomer D Ramachandran plot [20] Most favoured regions 90.3% 90.8% Additional allowed regions 9.5% 9.1% Generously allowed regions 0.2% Disallowed regions 0.1% Resolution range (Å) R merge (overall/high 0.080/ /0.347 resolution shell a ) Completeness (overall/high 96.2/ /98.8 resolution shell a ) (%) Mosaicity ( ) I/r(I) (overall/high resolution shell a ) 10.61/ /4.5 Refinement R cryst R free r.m.s. deviation from ideal bond length (Å) r.m.s. deviation from ideal bond angle ( ) a The high resolution shell is Å for the NAD + -bound and Å for the NADP + -bound structure Steady-state kinetic measurements Initial rates of NADP + - and NAD + -dependent oxidations of xylitol catalyzed by K274R were recorded spectrophotometrically (340 nm) at 25 C in 50 mm potassium phosphate buffer, ph 7.0. Xylitol was varied in seven concentrations between 48 and 2400 mm, and NAD + or NADP + was present at several constant concentrations between 1 and 300 lm. The concentration of K274R was approximately 3 lm in all assays. NADP + -dependent wild-type rates were obtained under identical conditions using similar reactant concentrations. Data fitting was performed as described elsewhere [6]. 3. Results and discussion 3.1. Kinetic consequences of replacing Lys-274 by arginine Kinetic parameters of K274R-catalyzed oxidation of xylitol by NAD + and NADP + are summarized in Table 2 along with previously determined kinetic parameters of K274R for the reduction of xylose by NADH and NADPH [6]. Table 2 also shows the relevant wild-type parameters. For a comparison of the wild-type and the mutant, the ratio of K i values which is a measure of the selectivity of binding is most informative. In the wild-type, apparent binding of NADPH and NADP + was about 19- and 39-fold tighter than binding of NADH and NAD +, respectively. Ratios of K i values for the K274R mutant were 1.33 (=K inadh /K inadph ) and 7.3 (=K inad+ / K inadp+ ). Therefore, replacement of Lys-274 by an arginine caused a change in NADP versus NAD binding selectivity for reduced and oxidized coenzymes by factors of 14 (=19/ 1.33) and 5.3 (=39/7.3), respectively. Comparison of catalytic efficiencies for reactions with NADP(H) and NAD(H), indicated in Table 2, shows that the wild-type preferred NADPH and NADP + in the directions of xylose reduction (33-fold) and xylitol oxidation (23-fold). The mutant displayed a slight preference for reactions with NADH (1.5-fold) and NAD + (2-fold). With the exception of k cat for NADPH-dependent reduction of xylose (discussed later), turnover numbers of K274R and wild-type were identical within experimental error. In the absence of a catalytic reaction profile for xylose reduction by CpXR, comparison of the literature data [7] and our results must be qualitative. It indicates however that the Lys- 274 fi Arg replacement induced kinetic properties in the mutant that most clearly distinguish the NADH-preferring CpXR from CtXR wild-type. In CpXR and K274R, apparent binding affinity for NADH seems to be gained partly at the expense of affinity for NADPH; and the K m for xylose in the NADPHlinked reaction is strongly increased, compared to CtXR Overall structures of K274R mutant The K274R mutant bound to NADP + or NAD + folds into the (b/a) 8 barrel previously described for wild-type CtXR [4,5]. There were four K274R molecules, two enzyme dimers, within the respective crystallographical asymmetric units. Numbering starts with 1 for the initiator methionine. Results are summarized in Table Structure of K274R mutant bound to NAD + The NAD + -bound structure of the mutant showed well-defined electron density around residues predicted from the protein and the NAD + coenzyme molecules. A total of 681 ordered waters were observed in the final structure of K274R but none was placed at a position possibly important for the enzyme function. The root mean square (r.m.s.) devia-
3 S. Leitgeb et al. / FEBS Letters 579 (2005) Table 2 Kinetic characterization of K274R mutant Oxidation Reduction NAD + NADP + NADH NADPH K ma (lm) 8 a (27) 4 a (9) 21 (38) 5 a (3) K ia (lm) 44 a (195) 6 (5 b ) 4 c (19) 3 d (1 d ) k cat (s 1 ) 0.93 (1.07) 0.91 (0.79) 8 (11) 26 (13) k cat /(K mb K ia ) e (s 1 mm 2 ) (0.026) (0.59) 37 (4) 24 (135) The standard deviations (S.D.) were <15%, unless indicated: a <25%; b <30%; c <60%; and d <40%. Values in parentheses are for the wild-type [1,6], except for the NADP + -dependent oxidation of xylitol (this work). k cat is the turnover number; K ia is the apparent coenzyme dissociation constant; and K ma and K mb are the Michaelis constants for coenzyme and substrate, respectively; e catalytic efficiency. tions between the Ca values of the models ranged from 0.23 to 0.30 Å, indicating that the four K274R monomers A D in the asymmetric unit were virtually identical. Since the overall temperature factor for molecule D was the lowest, measurements and conclusions in regard to specific atomic interactions were drawn from it. All of the non-hydrogen atoms in the NAD + were unambiguously observed in the final unbiased electron density map (Fig. 1A) with refined B factors for individual coenzyme atoms that ranged from 13 to 25 Å 2. Binding of the NAD + coenzyme to K274R occurred in a manner indistinguishable to that seen for the wild-type. Like Lys-274 of wild-type CtXR, Arg-274 does not engage in hydrogen bonding with NAD + (Fig. 2A). Its guanidinium group is P4.6 Å away from the adenosine 3 0 -hydroxyl. Mutation of Lys-274 did not cause significant structural changes in and around (60.25 Å) the active site. The significant kinetic consequences in the mutant regarding reactions with NADH and NAD + (see Table 2) must therefore reflect the ability of CtXR to accommodate the site-specific replacement through multiple conformational changes, which are too small to be resolved in a structural comparison of the wild-type and the mutant Structure of K274R mutant bound to NADP + Superpositioning of the models showed that the largest deviations occurred at the binding site of the adenosine moiety of the coenzyme. In particular, monomers A and C were different from monomers B and D. The final unbiased high-resolution electron density maps revealed that NAD + instead of NADP + was present in both monomers B and D. We have no way of Fig. 1. Unbiased F o F c electron densities contoured at 3.0 r of the cofactor adenosine moiety and Arg-274 in the NAD + -bound (A) and NADP + - bound mutant enzyme (B). The maps were derived from the refined models with the atoms of NAD(P) + and the side chain of Arg-274 removed. Figures were generated with MOLSCRIPT [16], BOBSCRIPT [17] and RASTER3D [18].
4 766 S. Leitgeb et al. / FEBS Letters 579 (2005) Fig. 2. Stereoviews of overlap of NAD + -bound structures (A) and NADP + -bound structures (B) of wild-type (yellow) and K274R (blue), illustrating the adenosine binding pocket. Hydrogen bonds are indicated as dashed lines and are shown for the mutant structures. The distances are given in Å, with wild-type values shown in parentheses. The overlays were made with the program EdPDB [19]. knowing the true source of NAD + in the experiment but conversion of NADP + may have occurred during the long time needed for crystallization. By contrast, the 2 0 -phosphate group of NADP + was unambiguously observed in structures of monomers A and C. The r.m.s. deviation of Ca values between these two monomers in the asymmetric unit was 0.21 Å. Of the two, monomer C had the lower overall temperature factor and therefore was chosen for further analysis. The final structure of CtXR K274R bound to NADP + contained 760 ordered water molecules. Important differences in interactions are noted when the adenosine moiety and its 2 0 -phosphate group is examined and compared to what is observed in structures of holo forms of the wild-type and the NAD + -bound structure of K274R. As in the wild-type apo enzyme [4], loop 7 was partly disordered in the K274R mutant bound to NADP +. In the wildtype, loop 7 becomes ordered upon binding of NADP + and is held in place by multiple interactions of Glu-227 and hydrogen bonds between Ser-224 and the juxtaposed Lys-25, which is contributed by a short turn connecting b-strand 1 and a- helix 1. The side chain of Glu-227 is part of a network of water-mediated contacts that is centered around the side chain of Lys-274 and also involves the 3 0 -hydroxy group of NADP +. The side chain of Lys-274 engages in a charged hydrogen bond (2.68 Å) with an oxygen of the 2 0 -phosphate group. All the interactions just discussed were disrupted in NADP + -bound K274R, clearly because of steric conflicts for the bulky arginine side chain. The conformation available to Arg-274 positions its guanidinium group 3.06 Å away from the 2 0 -phosphate (Fig. 2B). The distance between the hydroxyl of Ser-275 and the phosphate oxygen is longer in the NADP + -complex of K274R (2.91 Å) than in the same complex of the wild-type (2.72 Å). The backbone amide of Asn-276 is moved away slightly from its position in the wild-type, thereby increasing its original distance to the 2 0 -phosphate (2.71 Å) to a value of 2.85 Å. The distance between the phosphate oxygen and the side chain of Arg-280 is however shortened significantly in the mutant complex (2.66 Å) in comparison to the wild-type complex (2.75 Å). Summarizing, these observations suggest that the binding selectivity NADP + versus NAD + will be decreased in the mutant in comparison to the wild-type, in good agreement with the experimental ratios K inad+ /K inadp+ of 39 and 7.3 for wild-type and K274R, respectively. Interestingly, K inadp+ for K274R is not strongly increased, compared to the wild-type value Conformational changes involved in coenzyme binding and release Previous structures of wild-type CtXR [4,5] have shown that loop 7, which closes over the bound NAD(P) + in the holoenzyme, must undergo a conformational change to enable release of the coenzyme. The rearrangement of the loop from the closed to the open position has been correlated with a kinetically observed isomerization of the NAD + -bound enzyme, which is the main rate-limiting step of NADH-dependent reduction of xylose by wild-type CtXR [11]. Aside from the contacts made by Glu-227, the interactions between the side chains of Lys-25 and Ser-224 stabilize the closed conformation. In the NAD + -bound structure of the wild-type, the hydroxy group and backbone carbonyl oxygen of Ser-224 are
5 S. Leitgeb et al. / FEBS Letters 579 (2005) and 2.78 Å, respectively, away from the e-amino group of Lys-25. The corresponding distances in K274R bound to NAD + are similar and 2.91 and 2.66 Å, respectively. An overlay of the structures revealed the side chains of Lys-25 and Ser-224 to be almost superimposable in the two enzymes. These results suggest that the NAD + release rates should be similar for reactions catalyzed by K274R and the wild-type, in accordance with experimental turnover numbers for NADH-dependent reduction of xylose. The disorder seen in loop 7 in the NADP + -bound structure of K274R implies that the interactions between Lys-25 and Ser-224 are lost in this complex. The inability of K274R to anchor the coenzyme-enfolding loop in the closed position when NADP + is bound should lead to a faster net rate of NADP + release in the mutant, compared to wild-type. This explains the unusually high k cat for NADPH-dependent reduction of xylose by K274R. Also consistent with the proposed scenario, the turnover numbers for xylitol oxidation by wild-type and the mutant are identical, irrespective of whether NAD + or NADP + is used as a coenzyme. Because hydride transfer is the rate-limiting step of xylitol conversion by the wild-type [11], one does not expect that the k cat changes in that direction as a result of the mutation, considering that replacement Lys- 274 fi Arg affects mainly the steps leading to and from the binary complexes. Summarizing, this work provides a molecular basis of how the naturally occurring replacement Lys-274 fi Arg alters the coenzyme selectivity of family 2 AKRs towards improved acceptance of NADH, relative to NADPH. Lys-274 is positionally conserved across 5 of the current 14 AKR families, and several AKR structures outside family 2 confirm that the function of Lys-274 in hydrogen bonding with the co-substrate 2 0 -phosphate group is also conserved [12 15]. Therefore, this underscores the general relevance of our findings. The coordinates for the K274R mutant bound to NAD + and NADP + have been deposited into the Protein Data Bank under the ID codes 1YE4 and 1YE6, respectively. Acknowledgments: Financial support is acknowledged from the Austrian Science Funds (FWF project to B.N.), the National Institutes of Health (GM66135 to D.K.W.), and the Keck Foundation (D.K.W.). S.L. acknowledges support from a travel grant from Graz University of Technology. The data collection facilities at Stanford Synchrotron Radiation Laboratory are funded by the U.S. Department of Energy and the National Institutes of Health. We thank Dr. Ulrike Wagner (Department of Chemistry, University of Graz) for support with the preparation of the figures. References [1] Neuhauser, W., Haltrich, D., Kulbe, K.D. and Nidetzky, B. (1997) NAD(P)H-dependent aldose reductase from the xyloseassimilating yeast Candida tenuis. Isolation, characterization and biochemical properties of the enzyme. Biochem. J. 326, [2] Bruinenberg, P.M., de Bot, P.H.M., van Dijken, J.P. and Scheffers, W.A. (1983) The role of redox balances in the anaerobic fermentation of xylose by yeasts. Eur. J. Appl. Microbiol. Biotechnol. 18, [3] Jez, J.M. and Penning, T.M. (2001) The aldo-keto reductase (AKR) superfamily: an update. Chem. Biol. Interact , [4] Kavanagh, K.L., Klimacek, M., Nidetzky, B. and Wilson, D.K. (2002) The structure of apo and holo forms of xylose reductase, a dimeric aldo-keto reductase from Candida tenuis. Biochemistry 41, [5] Kavanagh, K.L., Klimacek, M., Nidetzky, B. and Wilson, D.K. (2003) Structure of xylose reductase bound to NAD + and the basis for single and dual co-substrate specificity in family 2 aldoketo reductases. Biochem. J. 373, [6] Petschacher, B., Leitgeb, S., Kavanagh, K.L., Wilson, D.K. and Nidetzky, B. (2005) The coenzyme specificity of Candida tenuis xylose reductase (AKR2B5) explored by site-directed mutagenesis and X-ray crystallography. Biochem. J. 385, [7] Lee, J.K., Koo, B.S. and Kim, S.Y. (2003) Cloning and characterization of the xyl1 gene, encoding an NADH-preferring xylose reductase from Candida parapsilosis, and its functional expression in Candida tropicalis. Appl. Environ. Microbiol. 69, [8] Otwinowski, Z. and Minor, W. (1997) Processing of X-ray diffraction data collected in oscillation model. Methods Enzymol. 276, [9] Brunger, A.T., et al. (1998) Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr., Part D 54, [10] Jones, T.A., Zou, J.Y., Cowan, S.W. and Kjeldgaard, M. (1991) Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr., Part A 47, [11] Nidetzky, B., Klimacek, M. and Mayr, P. (2001) Transientstate and steady-state kinetic studies of the mechanism of NADH-dependent aldehyde reduction catalyzed by xylose reductase from the yeast Candida tenuis. Biochemistry 40, [12] Ye, Q., Hyndman, D., Green, N.C., Li, L., Jia, Z. and Flynn, T.G. (2001) The crystal structure of an aldehyde reductase Y50F mutant NADP complex and its implications for substrate binding. Chem. Biol. Interact , [13] Wilson, D.K., Bohren, K.M., Gabbay, K.H. and Quiocho, F.A. (1992) An unlikely sugar substrate site in the 1.65 Å structure of the human aldose reductase holoenzyme implicated in diabetic complications. Science 257, [14] Penning, T.M., Bennett, M.J., Smith-Hoog, S., Schlegel, B.P., Jez, J.M. and Lewis, M. (1997) Structure and function of 3 alphahydroxysteroid dehydrogenase. Steroids 62, [15] Khurana, S., Powers, D.B., Anderson, S. and Blaber, M. (1998) Crystal structure of 2,5-diketo-D-gluconic acid reductase A complexed with NADPH at 2.1-Å resolution. Proc. Natl. Acad. Sci. USA 95, [16] Kraulis, P.J. (1991) MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, [17] Esnouf, R.M. (1997) An extensively modified version of Mol- Script that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model. 15, , [18] Merritt, E.A. and Bacon, D.J. (1997) Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, [19] Zhang, X.-J. and Matthews, B.W. (1995) EDPDB: a multifunctional tool for protein structure analysis. J. Appl. Crystallogr. 28, [20] Laskowski, R.A., MacArthur, M.W., Moss, D.S. and Thornton, J.M. (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26,
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Chemistry 135, First Exam. September 23, Chem 135, Exam 1 SID:
 Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Excerpt from J. Mol. Biol. (2002) 320, :
 Excerpt from J. Mol. Biol. (2002) 320, 1095 1108: Crystal Structure of the Ternary Complex of the Catalytic Domain of Human Phenylalanine Hydroxylase with Tetrahydrobiopterin and 3-(2-Thienyl)-L-alanine,
Excerpt from J. Mol. Biol. (2002) 320, 1095 1108: Crystal Structure of the Ternary Complex of the Catalytic Domain of Human Phenylalanine Hydroxylase with Tetrahydrobiopterin and 3-(2-Thienyl)-L-alanine,
Chemical Energy. Valencia College
 9 Pathways that Harvest Chemical Energy Valencia College 9 Pathways that Harvest Chemical Energy Chapter objectives: How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of
9 Pathways that Harvest Chemical Energy Valencia College 9 Pathways that Harvest Chemical Energy Chapter objectives: How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of
Cellular Respiration
 Cellular Respiration 1. To perform cell work, cells require energy. a. A cell does three main kinds of work: i. Mechanical work, such as the beating of cilia, contraction of muscle cells, and movement
Cellular Respiration 1. To perform cell work, cells require energy. a. A cell does three main kinds of work: i. Mechanical work, such as the beating of cilia, contraction of muscle cells, and movement
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Chapter 9: Cellular Respiration
 Chapter 9: Cellular Respiration To perform their many tasks, living cells require energy from outside sources. Energy stored in food utimately comes from the sun. Photosynthesis makes the raw materials
Chapter 9: Cellular Respiration To perform their many tasks, living cells require energy from outside sources. Energy stored in food utimately comes from the sun. Photosynthesis makes the raw materials
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
CELLULAR METABOLISM. Metabolic pathways can be linear, branched, cyclic or spiral
 CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Metabolism Energy Pathways Biosynthesis. Catabolism Anabolism Enzymes
 Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters
 Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters Chen-Hsiang Shen 1, Yuan-Fang Wang 1, Andrey Y. Kovalevsky 1, *, Robert W. Harrison 1,2 and Irene T.
Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters Chen-Hsiang Shen 1, Yuan-Fang Wang 1, Andrey Y. Kovalevsky 1, *, Robert W. Harrison 1,2 and Irene T.
Enzymes and Metabolism
 PowerPoint Lecture Slides prepared by Vince Austin, University of Kentucky Enzymes and Metabolism Human Anatomy & Physiology, Sixth Edition Elaine N. Marieb 1 Protein Macromolecules composed of combinations
PowerPoint Lecture Slides prepared by Vince Austin, University of Kentucky Enzymes and Metabolism Human Anatomy & Physiology, Sixth Edition Elaine N. Marieb 1 Protein Macromolecules composed of combinations
How Cells Harvest Energy. Chapter 7. Respiration
 How Cells Harvest Energy Chapter 7 Respiration Organisms classified on how they obtain energy: autotrophs: produce their own organic molecules through photosynthesis heterotrophs: live on organic compounds
How Cells Harvest Energy Chapter 7 Respiration Organisms classified on how they obtain energy: autotrophs: produce their own organic molecules through photosynthesis heterotrophs: live on organic compounds
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Respiration. Respiration. Respiration. How Cells Harvest Energy. Chapter 7
 How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
Metabolic engineering some basic considerations. Lecture 9
 Metabolic engineering some basic considerations Lecture 9 The 90ties: From fermentation to metabolic engineering Recruiting heterologous activities to perform directed genetic modifications of cell factories
Metabolic engineering some basic considerations Lecture 9 The 90ties: From fermentation to metabolic engineering Recruiting heterologous activities to perform directed genetic modifications of cell factories
7 Pathways That Harvest Chemical Energy
 7 Pathways That Harvest Chemical Energy Pathways That Harvest Chemical Energy How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of Glucose Metabolism? How Is Energy Harvested
7 Pathways That Harvest Chemical Energy Pathways That Harvest Chemical Energy How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of Glucose Metabolism? How Is Energy Harvested
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15
 Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15 AND STORAGE Berg, (Figures in red are for the 7th Edition) Tymoczko (Figures in Blue are for the 8th Edition) & Stryer] Two major questions
Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15 AND STORAGE Berg, (Figures in red are for the 7th Edition) Tymoczko (Figures in Blue are for the 8th Edition) & Stryer] Two major questions
Pentose Phosphate Pathway
 Pentose Phosphate Pathway An overview of the pathway, its regulation and relationship to glycolysis and other pathways. See chapter 15 of Fundamentals of Biochemisty: Life at the Molecular Level, 4 th
Pentose Phosphate Pathway An overview of the pathway, its regulation and relationship to glycolysis and other pathways. See chapter 15 of Fundamentals of Biochemisty: Life at the Molecular Level, 4 th
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation
 6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
Cellular Respiration
 Cellular I can describe cellular respiration Cellular respiration is a series of metabolic pathways releasing energy from a foodstuff e.g. glucose. This yields energy in the form of ATP adenosine P i P
Cellular I can describe cellular respiration Cellular respiration is a series of metabolic pathways releasing energy from a foodstuff e.g. glucose. This yields energy in the form of ATP adenosine P i P
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/ MCQs. Location : 102, 105, 106, 301, 302
 FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Introduction to Metabolism Cell Structure and Function
 Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
4. Which step shows a split of one molecule into two smaller molecules? a. 2. d. 5
 1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
1. Which of the following statements about NAD + is false? a. NAD + is reduced to NADH during both glycolysis and the citric acid cycle. b. NAD + has more chemical energy than NADH. c. NAD + is reduced
Crystal structure of fructose-1,6-bisphosphatase complexed with
 Proc. Natl. Acad. Sci. USA Vol. 91, pp. 12482-12486, December 1994 Biochemistry Crystal structure of fructose-1,6-bisphosphatase complexed with fructose 2,6-bisphosphate, AMP, and Zn2+ at 2.0-A resolution:
Proc. Natl. Acad. Sci. USA Vol. 91, pp. 12482-12486, December 1994 Biochemistry Crystal structure of fructose-1,6-bisphosphatase complexed with fructose 2,6-bisphosphate, AMP, and Zn2+ at 2.0-A resolution:
III. 6. Test. Respiració cel lular
 III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
The most common type of oxidative damage identified in
 8-Oxoguanine rearranges the active site of human topoisomerase I Diem-Thu Thieu Lesher*, Yves Pommier, Lance Stewart, and Matthew R. Redinbo* Departments of *Chemistry and Biochemistry and Biophysics,
8-Oxoguanine rearranges the active site of human topoisomerase I Diem-Thu Thieu Lesher*, Yves Pommier, Lance Stewart, and Matthew R. Redinbo* Departments of *Chemistry and Biochemistry and Biophysics,
Ch. 9 Cell Respiration. Title: Oct 15 3:24 PM (1 of 53)
 Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
7 Cellular Respiration and Fermentation
 CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY REECE 7 Cellular Respiration and Fermentation Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge, Simon Fraser University SECOND EDITION
2/25/2013. The Mechanism of Enzymatic Action
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 Chapter 5 Microbial Metabolism Catabolic and Anabolic Reactions Metabolism: The sum of the chemical reactions in an organism Catabolic and Anabolic Reactions Catabolism:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 Chapter 5 Microbial Metabolism Catabolic and Anabolic Reactions Metabolism: The sum of the chemical reactions in an organism Catabolic and Anabolic Reactions Catabolism:
Microbial Metabolism
 PowerPoint Lecture Slides for MICROBIOLOGY ROBERT W. BAUMAN Chapter 5 Microbial Metabolism Microbial Metabolism The sum total of chemical reactions that take place within cells (of an organism) Metabolic
PowerPoint Lecture Slides for MICROBIOLOGY ROBERT W. BAUMAN Chapter 5 Microbial Metabolism Microbial Metabolism The sum total of chemical reactions that take place within cells (of an organism) Metabolic
BIOLOGY - CLUTCH CH.9 - RESPIRATION.
 !! www.clutchprep.com CONCEPT: REDOX REACTIONS Redox reaction a chemical reaction that involves the transfer of electrons from one atom to another Oxidation loss of electrons Reduction gain of electrons
!! www.clutchprep.com CONCEPT: REDOX REACTIONS Redox reaction a chemical reaction that involves the transfer of electrons from one atom to another Oxidation loss of electrons Reduction gain of electrons
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached
 BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
Detection and Kinetic Properties of Alcohol Dehydrogenase in Dormant Corm of Crocus sativus L.
 Detection and Kinetic Properties of Alcohol Dehydrogenase in Dormant Corm of Crocus sativus L. Mahnaz Hadizadeh 1 and Ezzatollah Keyhani 1,2 1 Institute of Biochemistry and Biophysics, University of Tehran,
Detection and Kinetic Properties of Alcohol Dehydrogenase in Dormant Corm of Crocus sativus L. Mahnaz Hadizadeh 1 and Ezzatollah Keyhani 1,2 1 Institute of Biochemistry and Biophysics, University of Tehran,
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Lecture 11 - Biosynthesis of Amino Acids
 Lecture 11 - Biosynthesis of Amino Acids Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction Biosynthetic pathways for amino acids, nucleotides and lipids
Lecture 11 - Biosynthesis of Amino Acids Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction Biosynthetic pathways for amino acids, nucleotides and lipids
WHY IS THIS IMPORTANT?
 CHAPTER 3 ESSENTIALS OF METABOLISM WHY IS THIS IMPORTANT? It is important to have a basic understanding of metabolism because it governs the survival and growth of microorganisms The growth of microorganisms
CHAPTER 3 ESSENTIALS OF METABOLISM WHY IS THIS IMPORTANT? It is important to have a basic understanding of metabolism because it governs the survival and growth of microorganisms The growth of microorganisms
In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic
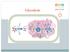 Glycolysis 1 In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic glycolysis. If this pyruvate is converted instead
Glycolysis 1 In glycolysis, glucose is converted to pyruvate. If the pyruvate is reduced to lactate, the pathway does not require O 2 and is called anaerobic glycolysis. If this pyruvate is converted instead
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Foundations in Microbiology Seventh Edition
 Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 8 An Introduction to Microbial Metabolism Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction
Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 8 An Introduction to Microbial Metabolism Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction
III. Metabolism - Gluconeogenesis
 Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism - Gluconeogenesis Carl & Gertrude Cori Slide 1 Carbohydrate Synthesis Lactate, pyruvate and glycerol are the important
Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism - Gluconeogenesis Carl & Gertrude Cori Slide 1 Carbohydrate Synthesis Lactate, pyruvate and glycerol are the important
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Chapter 8. Metabolism. Topics in lectures 15 and 16. Chemical foundations Catabolism Biosynthesis
 Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Enzymes
Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Enzymes
III. Metabolism Glucose Catabolism Part II
 Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism Glucose Catabolism Part II Slide 1 Metabolic Fates of NADH and Pyruvate Cartoon: Fate of pyruvate, the product of glycolysis.
Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism Glucose Catabolism Part II Slide 1 Metabolic Fates of NADH and Pyruvate Cartoon: Fate of pyruvate, the product of glycolysis.
BIOL 158: BIOLOGICAL CHEMISTRY II
 BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
Exercise 3. A Study of Enzyme Specificity. A Kinetic Analysis of Glucose Oxidase
 R e p r i n t e d f r o m G a l l i k S., C e l l B i o l o g y O L M P a g e 1 Exercise 3. A Study of Enzyme Specificity. A Kinetic Analysis of Glucose Oxidase A. Introduction Enzymes are one of the largest
R e p r i n t e d f r o m G a l l i k S., C e l l B i o l o g y O L M P a g e 1 Exercise 3. A Study of Enzyme Specificity. A Kinetic Analysis of Glucose Oxidase A. Introduction Enzymes are one of the largest
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM /19 BY: MOHAMAD FAHRURRAZI TOMPANG
 CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
CITRIC ACID CYCLE ERT106 BIOCHEMISTRY SEM 1 2018/19 BY: MOHAMAD FAHRURRAZI TOMPANG Chapter Outline (19-1) The central role of the citric acid cycle in metabolism (19-2) The overall pathway of the citric
3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP]
![3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP] 3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP]](/thumbs/72/66350985.jpg) 3.7 Cell respiration ( Chapter 9 in Campbell's book) 3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP] Organic compounds store
3.7 Cell respiration ( Chapter 9 in Campbell's book) 3.7.1 Define cell respiration [Cell respiration is the controlled release of energy from organic compounds in cells to form ATP] Organic compounds store
3.7 CELLULAR RESPIRATION. How are these two images related?
 3.7 CELLULAR RESPIRATION How are these two images related? CELLULAR RESPIRATION Cellular respiration is the process whereby the body converts the energy that we get from food (glucose) into an energy form
3.7 CELLULAR RESPIRATION How are these two images related? CELLULAR RESPIRATION Cellular respiration is the process whereby the body converts the energy that we get from food (glucose) into an energy form
Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life
 Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. The Metabolism of Microbes metabolism all chemical
Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. The Metabolism of Microbes metabolism all chemical
Oxidative phosphorylation & Photophosphorylation
 Oxidative phosphorylation & Photophosphorylation Oxidative phosphorylation is the last step in the formation of energy-yielding metabolism in aerobic organisms. All oxidative steps in the degradation of
Oxidative phosphorylation & Photophosphorylation Oxidative phosphorylation is the last step in the formation of energy-yielding metabolism in aerobic organisms. All oxidative steps in the degradation of
Chapter 8. Metabolism. Topics in lectures 15 and 16. Chemical foundations Catabolism Biosynthesis
 Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Chemical
Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Chemical
Independent Study Guide Metabolism I. Principles of metabolism (section 6.1) a. Cells must: (figure 6.1) i. Synthesize new components
 Independent Study Guide Metabolism I. Principles of metabolism (section 6.1) a. Cells must: (figure 6.1) i. Synthesize new components (anabolism/biosynthesis) ii. Harvest energy and convert it to a usable
Independent Study Guide Metabolism I. Principles of metabolism (section 6.1) a. Cells must: (figure 6.1) i. Synthesize new components (anabolism/biosynthesis) ii. Harvest energy and convert it to a usable
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Lab 5: Proteins and the small molecules that love them (AKA Computer Modeling with PyMol #2)
 Lab 5: Proteins and the small molecules that love them (AKA Computer Modeling with PyMol #2) Goals: The objective of this lab is to provide you with an understanding of: 1. Catalysis 2. Small molecule
Lab 5: Proteins and the small molecules that love them (AKA Computer Modeling with PyMol #2) Goals: The objective of this lab is to provide you with an understanding of: 1. Catalysis 2. Small molecule
OVERVIEW OF THE GLYCOLYTIC PATHWAY Glycolysis is considered one of the core metabolic pathways in nature for three primary reasons:
 Glycolysis 1 Supplemental Reading Key Concepts - Overview of the Glycolytic Pathway Glycolysis generates a small amount of ATP Preview of the ten enzyme-catalyzed reactions of glycolysis - Stage 1: ATP
Glycolysis 1 Supplemental Reading Key Concepts - Overview of the Glycolytic Pathway Glycolysis generates a small amount of ATP Preview of the ten enzyme-catalyzed reactions of glycolysis - Stage 1: ATP
Structure and functional analysis of the IGF-II/IGF2R interaction
 Structure and functional analysis of the IGF-II/IGF2R interaction James Brown 1, Carlie Delaine 2, Oliver J. Zaccheo 3, Christian Siebold 1, Robert J. Gilbert 1, Gijs van Boxel 1, Adam Denley 2, John C.
Structure and functional analysis of the IGF-II/IGF2R interaction James Brown 1, Carlie Delaine 2, Oliver J. Zaccheo 3, Christian Siebold 1, Robert J. Gilbert 1, Gijs van Boxel 1, Adam Denley 2, John C.
BY: RASAQ NURUDEEN OLAJIDE
 BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
Biochemical and Biophysical Research Communications 346 (2006) Crystal structure of rat carnitine palmitoyltransferase II (CPT-II)
 Biochemical and Biophysical Research Communications 346 (2006) 974 980 BBRC www.elsevier.com/locate/ybbrc Crystal structure of rat carnitine palmitoyltransferase II (CPT-II) Yu-Shan Hsiao a, Gerwald Jogl
Biochemical and Biophysical Research Communications 346 (2006) 974 980 BBRC www.elsevier.com/locate/ybbrc Crystal structure of rat carnitine palmitoyltransferase II (CPT-II) Yu-Shan Hsiao a, Gerwald Jogl
number Done by Corrected by Doctor Nayef Karadsheh
 number 17 Done by Abdulrahman Alhanbali Corrected by Lara Abdallat Doctor Nayef Karadsheh 1 P a g e Pentose Phosphate Pathway (PPP) Or Hexose Monophosphate Shunt In this lecture We will talk about the
number 17 Done by Abdulrahman Alhanbali Corrected by Lara Abdallat Doctor Nayef Karadsheh 1 P a g e Pentose Phosphate Pathway (PPP) Or Hexose Monophosphate Shunt In this lecture We will talk about the
Chemistry 2030 Introduction to Organic Chemistry Fall Semester 2012 Dr. Rainer Glaser
 Chemistry 2030 Introduction to Organic Chemistry Fall Semester 2012 Dr. Rainer Glaser Examination #5: The Final Lipids, Carbohydrates, Nucleobases & DNA. Monday, December 10, 2012, 3 5 pm. Name: Question
Chemistry 2030 Introduction to Organic Chemistry Fall Semester 2012 Dr. Rainer Glaser Examination #5: The Final Lipids, Carbohydrates, Nucleobases & DNA. Monday, December 10, 2012, 3 5 pm. Name: Question
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Cellular Respiration: Harvesting Chemical Energy CHAPTER 9
 Cellular Respiration: Harvesting Chemical Energy CHAPTER 9 9.1 Metabolic pathways that release energy are exergonic and considered catabolic pathways. Fermentation: partial degradation of sugars that occurs
Cellular Respiration: Harvesting Chemical Energy CHAPTER 9 9.1 Metabolic pathways that release energy are exergonic and considered catabolic pathways. Fermentation: partial degradation of sugars that occurs
BIOLOGY 101. CHAPTER 9: Cellular Respiration - Fermentation: Life is Work
 BIOLOGY 101 CHAPTER 9: Cellular Respiration - Fermentation: Life is Work An Introduction to Metabolism: Energy of Life 8.3 ATP powers cellular work by coupling exergonic reactions to endergonic reactions
BIOLOGY 101 CHAPTER 9: Cellular Respiration - Fermentation: Life is Work An Introduction to Metabolism: Energy of Life 8.3 ATP powers cellular work by coupling exergonic reactions to endergonic reactions
Chapter 9: Cellular Respiration: Harvesting Chemical Energy
 AP Biology Reading Guide Name: Date: Period Chapter 9: Cellular Respiration: Harvesting Chemical Energy Overview: Before getting involved with the details of cellular respiration and photosynthesis, take
AP Biology Reading Guide Name: Date: Period Chapter 9: Cellular Respiration: Harvesting Chemical Energy Overview: Before getting involved with the details of cellular respiration and photosynthesis, take
Metabolism. Metabolic pathways. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 11: Metabolic Pathways http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Metabolism Metabolism is the chemical change of
TCA CYCLE (Citric Acid Cycle)
 TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
Six Types of Enzyme Catalysts
 Six Types of Enzyme Catalysts Although a huge number of reactions occur in living systems, these reactions fall into only half a dozen types. The reactions are: 1. Oxidation and reduction. Enzymes that
Six Types of Enzyme Catalysts Although a huge number of reactions occur in living systems, these reactions fall into only half a dozen types. The reactions are: 1. Oxidation and reduction. Enzymes that
LAST LECTURE. 1. Things the course missed 2. Things you might be tested on
 LAST LECTURE 1. Things the course missed 2. Things you might be tested on Things I didn t tell you that might make more sense now that you have had BIS105 From Regulation of Cellular Metabolism by Protein
LAST LECTURE 1. Things the course missed 2. Things you might be tested on Things I didn t tell you that might make more sense now that you have had BIS105 From Regulation of Cellular Metabolism by Protein
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Microbiology AN INTRODUCTION
 TORTORA FUNKE CASE Microbiology AN INTRODUCTION EIGHTH EDITION B.E Pruitt & Jane J. Stein Chapter 5, part A Microbial Metabolism PowerPoint Lecture Slide Presentation prepared by Christine L. Case Microbial
TORTORA FUNKE CASE Microbiology AN INTRODUCTION EIGHTH EDITION B.E Pruitt & Jane J. Stein Chapter 5, part A Microbial Metabolism PowerPoint Lecture Slide Presentation prepared by Christine L. Case Microbial
Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis.
 ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
True or False: 1. Reactions are called endergonic if they occur spontaneously and release free energy.
 True or False: 1. Reactions are called endergonic if they occur spontaneously and release free energy. 2. Enzymes catalyze chemical reactions by lowering the activation energy 3. Biochemical pathways are
True or False: 1. Reactions are called endergonic if they occur spontaneously and release free energy. 2. Enzymes catalyze chemical reactions by lowering the activation energy 3. Biochemical pathways are
The building blocks of life.
 The building blocks of life. The 4 Major Organic Biomolecules The large molecules (biomolecules OR polymers) are formed when smaller building blocks (monomers) bond covalently. via anabolism Small molecules
The building blocks of life. The 4 Major Organic Biomolecules The large molecules (biomolecules OR polymers) are formed when smaller building blocks (monomers) bond covalently. via anabolism Small molecules
Cellular Pathways That Harvest Chemical Energy. Cellular Pathways That Harvest Chemical Energy. Cellular Pathways In General
 Cellular Pathways That Harvest Chemical Energy A. Obtaining Energy and Electrons from Glucose Lecture Series 12 Cellular Pathways That Harvest Chemical Energy B. An Overview: Releasing Energy from Glucose
Cellular Pathways That Harvest Chemical Energy A. Obtaining Energy and Electrons from Glucose Lecture Series 12 Cellular Pathways That Harvest Chemical Energy B. An Overview: Releasing Energy from Glucose
Metabolism. Chapter 8 Microbial Metabolism. Metabolic balancing act. Catabolism Anabolism Enzymes. Topics. Metabolism Energy Pathways Biosynthesis
 Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Carbohydrate Metabolism I
 Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Chapter 9 Overview. Aerobic Metabolism I: The Citric Acid Cycle. Live processes - series of oxidation-reduction reactions. Aerobic metabolism I
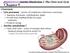 n n Chapter 9 Overview Aerobic Metabolism I: The Citric Acid Cycle Live processes - series of oxidation-reduction reactions Ingestion of proteins, carbohydrates, lipids Provide basic building blocks for
n n Chapter 9 Overview Aerobic Metabolism I: The Citric Acid Cycle Live processes - series of oxidation-reduction reactions Ingestion of proteins, carbohydrates, lipids Provide basic building blocks for
Cellular Respiration Harvesting Chemical Energy ATP
 Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
CryoProtX MD1 61. MD1-61 is presented as a 46 x 1.5 ml kit. To be used in conjunction with the Quick-Start Guide. Features of CryoProtX :
 CryoProtX MD1 61 Get better results from your crystals This kit is designed for the cryoprotection of protein crystals and is for use after the crystals have been grown. Get the most out of your protein
CryoProtX MD1 61 Get better results from your crystals This kit is designed for the cryoprotection of protein crystals and is for use after the crystals have been grown. Get the most out of your protein
The Binding Mode of by Electron Crystallography
 The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
Enzymes what are they?
 Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Microbial Metabolism. PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 5 Microbial Metabolism Big Picture: Metabolism Metabolism is the buildup and breakdown of nutrients
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 5 Microbial Metabolism Big Picture: Metabolism Metabolism is the buildup and breakdown of nutrients
