One Gene, Many Proteins. Applications of Mass Spectrometry to Proteomics. Why Proteomics? Raghothama Chaerkady, Ph.D.
|
|
|
- Emil Brown
- 6 years ago
- Views:
Transcription
1 Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins Gene mrna Protein Co/post-translational processing Gene Alternative splicing mrna 1 mrna 2 mrna 3 mrna 4 mrna 5 Protein 1 Protein 2 Protein 3 Protein 4 Protein 5 Protein 6... Protein i 1
2 Why Mass Spectrometry? Protein Characterization by Mass Spectrometry Very sensitive: Less than 1 pmol of protein is required for identification (5 ng of a 5 kda protein) Mixtures of proteins (hundreds to thousands of proteins) can be analyzed Proteins with a blocked N terminus can be identified Post-translational modifications such as phosphorylation can be identified and localized Quantitative mass spectrometry has numerous applications 2
3 Electrospray Tandem Mass Spectrometry (MS/MS) MS1 Scan - Detection of all the peptides in a mixture MS/MS - When one of the peptides is isolated and subjected to collision induced dissociation for sequencing Tandem mass spectrometry provides the actual sequence of the peptide Peptide Sequencing by MS/MS Select one peptide Collision Fragments containing Carboxyl-terminus Y 7 Y 6 Y 5 Y 4 Y 3 MS/MS Spectrum Relative Intensity (%) 3
4 Genomics and Proteomics Proteome 1 Biological changes (morphogenesis) Profiling of proteins Proteome n Proteome 1 Proteome 2 Proteome 4 Proteome 3 Proteome n mrna and Protein Correlation Generic mass spectrometry (MS)-based proteomics experiment. insight Nature 422,
5 Mass spectrometers used in proteome research. In reflector time-of-flight (TOF) instruments, the ions are accelerated to high kinetic energy and are separated along a flight tube as a result of their different velocities. The ions are turned around in a reflector, which compensates for slight differences in kinetic energy, and then impinge on a detector that amplifies and counts arriving ions. 5
6 b, The TOF-TOF instrument incorporates a collision cell between two TOF sections. Ions of one mass-to-charge ( m/ z) ratio are selected in the first TOF section, fragmented in the collision cell, and the masses of the fragments are separated in the second TOF section Quadrupole mass spectrometers select by time-varying electric fields between four rods, which permit a stable trajectory only for ions of a particular desired. Again, ions of a particular are selected in a first section (Q1), fragmented in a collision cell (q2), and the fragments separated in Q3. In the linear ion trap, ions are captured in a quadruple section, depicted by the red dot in Q3. They are then excited via resonant electric field and the fragments are scanned out, creating the tandem mass spectrum The quadrupole TOF instrument combines the front part of a triple quadruple instrument with a reflector TOF section for measuring the mass of the ions. 6
7 The (three-dimensional) ion trap captures the ions as in the case of the linear ion trap, fragments ions of a particular m/ z, and then scans out the fragments to generate the tandem mass spectrum. The FT-MS instrument also traps the ions, but does so with the help of strong magnetic fields. The figure shows the combination of FT-MS with the linear ion trap for efficient isolation, fragmentation and fragment detection in the FT- MS section. During ion detection, both the central electrode and deflector are maintained at very stable voltages so that no mass drift can take place. The outer electrode is split in half at z=, allowing the ion image current in the axial direction to be collected. The image current on each of half of the outer electrode is differentially amplified and then undergoes analog-to-digital conversion before processing using the fast Fourier transform algorithm. 7
8 MS/MS spectrum (sequencing) Micromass QTOF Finnigan LCQ Deca LTQ-Orbitrap XL 1D Liquid Chromatography Setup Peptide Mixture from pump Analytical column C 18 Emitter 8µm tip diverter valve Split line Manual 3-port valve for peak parking valve 8nL/min with valve open 8
9 A matchstick Needle with a very small opening and containing a reverse phase Column for separation Multidimensional Liquid Chromatography (MudPIT) Setup MS Liu, H. et al. (22) Offline Fractionation in the First Dimension Pump Manual Injector Auto Sampler Fraction Collector Reverse phase LCMS SCX Column UV Detector 9
10 Automated nanolc-ms/ms Purification of sample Very complex samples can be analyzed Identical peptides will elute in one peak (enhanced signal) Automated Many proteins can be identified in one run (-2, proteins) Coupling Liquid Chromatography to Tandem MS TIC: Total ion chromatogram Time (minutes) MS/MS spectrum MS/MS spectrum y 2 y V G C S M W Y W K A N T F Y T C F L G T S S L A G F K y y y 9 y y y 13 5 y y y y y y 12 y y y 7 y y y y y Quantitative Proteomics Based on difference in intensity By relative quantitation using MS 1
11 Quantitative Proteomics Based on difference in intensity 1D gels 2D gels Use of fluorescence-based methods 1D Gel-based Comparison EGF : IP : anti-ptyr 2D Gel-based Comparison Normal Cancer 11
12 2D Gels - Limitations Sample preparation - lot of optimization required Loading capacity limited Do not resolve very small (<1 kd) or large (> kd) proteins Do not resolve hydrophobic (e.g. membrane) proteins Issues with reproducibility Quantitative Proteomics Fluorescence-based quantitation DIGE (Difference in-gel electrophoresis) DIGE Samples to be labeled are labeled with Cy3 (green) and Cy5 (red) Samples are mixed and resolved by 2D gels Fluorescence measured and quantitated 12
13 2D-DIGE of Pancreatic Juice Normal Cy3 Cy3-Cy5 Cancer Cy5 Quantitative Proteomics By relative quantitation using MS in vitro labeling 18 O-labeling Peptide mass tagging (ICAT) in vivo labeling Labeling with stable isotope containing amino acids (SILAC) 18 O-labeling normal NC cancer Excise gel band Excise gel band In-gel digestion with H 16 2 O water In-gel digestion with H 18 2 O water Mix 1:1 MS-analysis 13
14 18 O-labeling Isotope-Coded Affinity Tag (ICAT) Structure of ICAT reagent 4 Stable Isotope Labeling in Cell Culture (SILAC) for Protein Quantitation Mammalian cell culture models are used for studying a number of biological processes In the SILAC approach, cells are grown continuously in media containing one or more stable isotopes (e.g. 13 C). All the proteins in the cells are heavier and can be used to mark a given state in mass spectrometric analysis 14
15 Time Course of Heavy Amino Acid Incorporation Advantages of the SILAC Method Simple In vivo Does not require any extra processing steps All proteins are uniformly labeled Complete and predictable incorporation Choice of labeled amino acids De novo sequencing can be performed General strategy for stable isotope labeling by amino acids 15
16 SILAC for Quantitation of Secreted Proteins 12 C 6 Labeled Lysine 13 C 6 Labeled Lysine Mix and run on SDS-PAGE Elute, Digest with Trypsin and Analyze by MS Relative Intensity Profile of Proteins Secreted by Adipocytes Day 1 Day 4 Day 1 Day 7 Day 1 Day 9 Mix Days 1+4 Mix Days 1+7 Mix Days 1+9 Ratio 1:.5 Fibronectin expression is downregulated Ratio 1:.4 Ratio 1: Ratio 1: Collagen alpha 3 expression is upregulated Ratio 1: Ratio 1:
17 Studying Dynamics Using SILAC 6Da 4Da VSHLLGINVTDFTR 5 x Intensity Biomarker Discovery Using Proteomics Ideal targets for biomarker Protein (differential expressed proteins) DNA (mutations, methylation) RNA (differential expressed genes) Biological specimen Tissue (whole tissue or isolated tumor cells) Pancreatic juice Serum Plasma Pancreatic juice 17
18 Physiological proteome of human pancreatic juice Collection of pancreatic juice (cancer) during surgery Run on 1D gel LC-MS/MS analysis Bioinformatic analysis of identified proteins Compare data with known microarray data Differential Proteomic Analysis of Pancreatic Cancer Secretome In-vivo labeling with both arginine and lysine Collect conditioned media and concentrate with centricon 3, Da MWCO Normalize and mix in 1:1 ratio Resolve proteins by 1D gel electrophoresis Excise bands and digest proteins by trypsin Identify proteins by nanolc-ms/ms (2x3 bands) Verify identified proteins (manually) Relative quantitation of ID proteins (manually) Quantitation of Secreted Proteins 18
19 Post-translational Modifications Peptides can have a number of modifications During database searching, a variable modification has to be specified otherwise, no hit Common PTMs are phosphorylation, acetylation, ubiquitination, glycosylation etc. Protein Phosphorylation One-third of all cellular proteins are phosphorylated at one time or another Phosphoamino acid content of a vertebrate cell: Serine - 9%; Threonine - 1%; Tyrosine -.5% Ser:Thr:Tyr - 18:2:1 Tyrosine phosphorylation is tightly regulated Identifying Activated Tyrosine Kinases in Lung cancer HBEC HBEC L858R HBEC Del Light K/D4 and R/13C6 K/13C6,15N2 and R/13C6,15N4 Mix lysate, enrich tyrosine phosphorylated proteins Elute, digest with trypsin, analyze by mass spectrometry 19
20 Anti-acetyl lysine peptide IP Cell lysate in urea buffer LC-MS/MS Trypsin digestion Sep-Pak C18 Reverse-phase purification Clean up peptides using C-18 tip Elution using 1% acetic acid Purified peptides Lyophilization Immunoaffinity Purification (Anti-acetyl lysine- Protein A agarose) GSK3 beta GSK3 beta 2
21 Why is Phosphorylation Analysis Difficult? Stoichiometry of phosphorylation is low Complete coverage of proteins is difficult to obtain Phosphorylated serine and threonine residues are labile whereas phosphotyrosines are more stable Phosphoserines and phosphothreonine residues can be subjected to a beta-elimination reaction but not phosphotyrosine residues Antibodies to enrich for serine and threonine phosphorylated proteins are not available Phosphopeptides are suppressed in a mass spectrum Immonium Ion of Phosphotyrosine as a Reporter Ion HO O P O OH HO O P O OH NH CH C O Phosphotyrosine + HN Immonium Ion ( Da) Phosphotyrosine-Specific Precursor Ion Scanning Bcr/Abl MS Scan Control p185 Bcr/Abl Phosphotyrosine-Specific Precursor Ion Scan * * * * 28 * IP: anti-ptyr
22 Large scale IP with an anti-pser/thr antibody Calyculin A: WB: Anti-pSer/Thr Lysates In Vivo Labeling with 32 P Calyculin A: - + IP: Anti-pSer/Thr Identification of Phosphorylated Ser/Thr residues 1RLpSPAPQLGP 19 23
23 MS-Based Identification of a 13 kda Protein in the EGF Receptor Signaling Pathway EGF : Silver stained gel IP : anti-ptyr Partial list of acetylated lysine containing peptides identified in an IP experiment Protein name Histone cluster 1, H4a Immunoglobulin superfamily, member 9 Hypothetical protein [gi ] RBP-binding zinc finger protein, c Putative homeodomain transcription factor 1 H2A histone family, member Z Block of proliferation 1 Peptide GLGKGGAKR GKPEVVSVVGR KGKPQVQDKVVK KNQLVQK LKKVENIK TKAVSR QELTKKLMPNCK Score
24 Assignment of the initiator methionine in a cdna fragment based on an N-terminal peptide >KIAA229 (118 residues) FRAGMENT SWGKGREGVVSPAGLGGALPGDGKFGSPSRLGCSLGEGVQRVAALGMGKEQ ELLRAARTGHLPAVEKLLSGKRLSSGFGGGGGGGSGGGGGGSGGGGGGLGS SSHPLSSLLSMWRGPNVNCVDSTGYTPLHHAALNGHHRRSSSSRSQDSAEGQ DGQVPEQFSGLLHGSSPVCEVGQDPFQLLCTAGQSHPDGSPQQGACHKASM QLEETGVHAPGASQPSALDQSKRVGYLTGLPTTNSRSHPETLTHTASPHPGGA EEGDRSGAR Assignment of the initiator methionine in a cdna fragment based on an N-terminal peptide CH 3 C H 2 N G K E Q L L R O >KIAA229 (118 residues) FRAGMENT SWGKGREGVVSPAGLGGALPGDGKFGSPSRLGCSLGEGVQRVAALGMGKEQ LLRAARTGHLPAVEKLLSGKRLSSGFGGGGGGGSGGGGGGSGGGGGGLGSS SHPLSSLLSMWRGPNVNCVDSTGYTPLHHAALNGHHRRSSSSRSQDSAEGQD GQVPEQFSGLLHGSSPVCEVGQDPFQLLCTAGQSHPDGSPQQGACHKASMQL EETGVHAPGASQPSALDQSKRVGYLTGLPTTNSRSHPETLTHTASPHPGGAEE GDRSGAR Lysine Modifications NH 2 CH 3 NH H 3 C CH 3 N H C CH 3 3 N + CH 3 O CH 3 C NH 2 CH 3 CH 3 CH 3 NH 2 CH C O OH NH 2 CH C O OH NH 2 CH C O OH NH 2 CH C O OH NH 2 CH C O OH Lysine Mono-methylated lysine Di-methylated lysine Tri-methylated lysine Acetylated lysine Mass (Da) Mass gain Da Da Da 42.1 Da 25
25 Acetylation vs Tri-methylation: a numbers game Tri-methylation: Da [H(6) C(3)] Acetylation: Da [H(2) C(2) O] Mass difference: Da Resolution of various mass spectrometers Normal ion trap Quadrupole-Time-of-Flight Orbitrap Zoom scan -scale is the same! <3, 8,-14, 3,-6, Approx. resolution in ppm A triple state SILAC experiment: qtof +2x4 Da + 2x4 Da (2x Lysine containing peptide)
26 A triple state SILAC experiment: Orbitrap molina_6dec7_udyan_silac_py-pep-ip #4485 RT: AV: 1 NL: 7.21E6 F: FTMS + p NSI Full ms [ ] Relative Abundance A triple state SILAC experiment: Orbitrap +2x4 Da + 2x4 Da (2x Lysine containing peptide) +6 Da +4 Da (Arginine containing peptide) Relative Abundance Identifying acetylation sites using Orbitrap Digest protein mixture into peptides Capture acetylated lysine containing peptides using anti-acetyl lysine antibodies Analyze on Orbitrap If the samples are labeled for quantitative proteomics, one can also obtain data on differential regulation of acetylation at individual lysine residues in this strategy 27
27 Peptide Capture using anti-acetyl lysine antibody Cell lysate in urea buffer LC-MS/MS Trypsin digestion Sep-Pak C18 Reverse-phase purification Clean up peptides using C-18 tip Elution using 1% acetic acid Purified peptides Lyophilization Immunoaffinity Purification (Anti-acetyl lysine- Protein A agarose) GGKGLGKGGAKR peptide from Histone Cluster 1, H4a: What are the PTMs? Measured mass: (3x Acetyl) (2x Acetyl. + 1x tri-met.) (1x Acetyl. + 2x tri-met.) (3x tri-met.) Answer: Acetylation at each of 3 lysine residues: GGK*GLGK*GGAK*R molina_6dec7_lys-acetyl-pep-ip2 # 33 RT: AV: 1 NL: 5.43E6 T: FTMS + p NSI Full ms [ ] (theoretical: ) Relative Abundance molina_6dec7_lys-acetyl-pep-ip2 # 31 RT: AV: 1 NL: 1.54E4 T: ITMS + c NSI d Full ms @cid35. [ ] Relative Abundance
28 Lysine Modifications Gly Gly Isopeptide bond NH Lysine residue NH 2 CH C O OH Di-glycyl lysine Mass gain Da Tryptic Digest of Ubiquitylated Peptide LIFAGKQLEDGR GG K GG Da Signature of a Ubiquitylated Peptide 12.7 Di-glycine Containing Peptide Immonium ion Non Di-glycine Containing Peptide
29 Genome Annotation - A case for proteomics-driven annotation of protein-coding regions Genome Annotation by Mass Spectrometry: What Can We Gain? Assigning start codons Proteins isoforms (alternative splicing, novel exons) Novel genes (proteins less than amino acids not predicted by programs) csnps Correction of incorrect gene predictions (~5% of the genes in human are predicted) Validation of gene predictions Transcripts and Proteins in Anopheles gambiae Proteins ~ 7 known Annotated genes: 15,189 (11,757 predicted) 7,84 unique to predictions by Celera 1,375 unique to predictions by Ensembl 5,974 common to both predictions Gomez et al. (Genome Biology: 6:R39, 25) 35, full-length enriched cdnas 3,7 genes of which only 65 are novel Krisventseva et al. (Genome Research, 25) 215,634 ESTs/cDNAs 7,961 clusters of which 3, are novel 3
30 Use of Ensembl Distributed Annotation System to Validate a Known Transcript Schematic representation of the systems biology paradigm involving proteomics. 31
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
4.2 RESULTS AND DISCUSSION
 phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Advances in Hybrid Mass Spectrometry
 The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Biological Mass spectrometry in Protein Chemistry
 Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
FOURIER TRANSFORM MASS SPECTROMETRY
 FOURIER TRANSFORM MASS SPECTROMETRY https://goo.gl/vx3ogw FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall,
FOURIER TRANSFORM MASS SPECTROMETRY https://goo.gl/vx3ogw FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall,
Proteomics/Peptidomics
 Proteomics/Peptidomics System biology tools and preclinical models for translational research in endometriosis, ESHRE Campus workshop, 4-5 September 2009 E. Waelkens Proteomics: What? Proteins Proteomics
Proteomics/Peptidomics System biology tools and preclinical models for translational research in endometriosis, ESHRE Campus workshop, 4-5 September 2009 E. Waelkens Proteomics: What? Proteins Proteomics
REDOX PROTEOMICS. Roman Zubarev.
 REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry
 Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
Proteins: Proteomics & Protein-Protein Interactions Part I
 Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses
 New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
New Instruments and Services
 New Instruments and Services http://planetorbitrap.com/orbitrap fusion Combining the best of quadrupole, Orbitrap, and ion trap mass analysis in a revolutionary Tribrid architecture, the Orbitrap Fusion
New Instruments and Services http://planetorbitrap.com/orbitrap fusion Combining the best of quadrupole, Orbitrap, and ion trap mass analysis in a revolutionary Tribrid architecture, the Orbitrap Fusion
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics
 New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
MALDI-TOF. Introduction. Schematic and Theory of MALDI
 MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MS/MS Scan Modes. Eötvös University, Budapest April 16, MS/MS Scan Modes. Árpád Somogyi. Product Ion Scan Select. Scan. Precursor Ion Scan Scan
 MS/MS Modes Árpád Somogyi Eötvös University, Budapest April 16, 2012 MS/MS Modes Product Ion Precursor Ion Neutral Loss Δ ed Reaction Monitoring (SRM) 1 modes in a triple quadrupole (QqQ) (one quadrupole
MS/MS Modes Árpád Somogyi Eötvös University, Budapest April 16, 2012 MS/MS Modes Product Ion Precursor Ion Neutral Loss Δ ed Reaction Monitoring (SRM) 1 modes in a triple quadrupole (QqQ) (one quadrupole
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE
 SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry
 Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Proteome analysis Cellular proteomics Plasma proteomics
 Proteome analysis Cellular proteomics Plasma proteomics Janne Lehtiö Maria Pernemalm Karolinska Biomics Center Karolinska Institutet / Karolinska University Hospital Janne Lehtio 2009-02-09 1 KBC Proteomics
Proteome analysis Cellular proteomics Plasma proteomics Janne Lehtiö Maria Pernemalm Karolinska Biomics Center Karolinska Institutet / Karolinska University Hospital Janne Lehtio 2009-02-09 1 KBC Proteomics
Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD
 Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD 4-7117 sbarnes@uab.edu Overview A mass spectrum Electrospray MS Analysis of intact proteins Molecular weight calculations
Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD 4-7117 sbarnes@uab.edu Overview A mass spectrum Electrospray MS Analysis of intact proteins Molecular weight calculations
Mass Spectrometry. - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications
 - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
- Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast
 Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Biomolecular Mass Spectrometry
 Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Characterization of an Unknown Compound Using the LTQ Orbitrap
 Characterization of an Unknown Compound Using the LTQ rbitrap Donald Daley, Russell Scammell, Argenta Discovery Limited, 8/9 Spire Green Centre, Flex Meadow, Harlow, Essex, CM19 5TR, UK bjectives unknown
Characterization of an Unknown Compound Using the LTQ rbitrap Donald Daley, Russell Scammell, Argenta Discovery Limited, 8/9 Spire Green Centre, Flex Meadow, Harlow, Essex, CM19 5TR, UK bjectives unknown
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
MASS SPECTROMETRY BASED METABOLOMICS. Pavel Aronov. ABRF2010 Metabolomics Research Group March 21, 2010
 MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
Introduction to Peptide Sequencing
 Introduction to Peptide equencing Quadrupole Ion Traps tructural Biophysics Course December 3, 2014 12/8/14 Introduction to Peptide equencing - athan Yates 1 Why are ion traps used to sequence peptides?
Introduction to Peptide equencing Quadrupole Ion Traps tructural Biophysics Course December 3, 2014 12/8/14 Introduction to Peptide equencing - athan Yates 1 Why are ion traps used to sequence peptides?
Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics
 Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
New Instruments and Services
 New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
Mass Spectrometry and Proteomics. Professor Xudong Yao Bioanalytical Chemistry Spring 2007
 Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
Mass Spectrometry and Proteomics Xudong Yao
 Mass Spectrometry and Proteomics Xudong Yao Dept of Chemistry University of Connecticut Storrs, CT April 19, 2005 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Gel or non-gel
Mass Spectrometry and Proteomics Xudong Yao Dept of Chemistry University of Connecticut Storrs, CT April 19, 2005 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Gel or non-gel
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g.
 Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
LC/QTOF Discovery of Previously Unreported Microcystins in Alberta Lake Waters
 LC/QTOF Discovery of Previously Unreported Microcystins in Alberta Lake Waters Ralph Hindle Vogon Laboratory Services Ltd. Cochrane, Alberta, Canada Xu Zhang David W. Kinniburgh Alberta Centre for Toxicology
LC/QTOF Discovery of Previously Unreported Microcystins in Alberta Lake Waters Ralph Hindle Vogon Laboratory Services Ltd. Cochrane, Alberta, Canada Xu Zhang David W. Kinniburgh Alberta Centre for Toxicology
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Protein Identification and Phosphorylation Site Determination by de novo sequencing using PepFrag TM MALDI-Sequencing kit
 Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Ion Source. Mass Analyzer. Detector. intensity. mass/charge
 Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose.
 Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Mass Spectrometry Infrastructure
 Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Quantification by Mass Spectrometry
 Quantification by Mass Spectrometry PC219 Lecture 5 30 April, 2010 sguan@cgl.ucsf.edu 1 Applications of Quantitative Proteomics Qualitative protein identification is NOT sufficient to describe a biological
Quantification by Mass Spectrometry PC219 Lecture 5 30 April, 2010 sguan@cgl.ucsf.edu 1 Applications of Quantitative Proteomics Qualitative protein identification is NOT sufficient to describe a biological
1. Sample Introduction to MS Systems:
 MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions
 Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions Application Note Targeted Proteomics Authors Linfeng Wu, Christine A. Miller, Jordy Hsiao, Te-wei Chu, Behrooz
Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions Application Note Targeted Proteomics Authors Linfeng Wu, Christine A. Miller, Jordy Hsiao, Te-wei Chu, Behrooz
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Quadrupole and Ion Trap Mass Analysers and an introduction to Resolution
 Quadrupole and Ion Trap Mass Analysers and an introduction to Resolution A simple definition of a Mass Spectrometer A Mass Spectrometer is an analytical instrument that can separate charged molecules according
Quadrupole and Ion Trap Mass Analysers and an introduction to Resolution A simple definition of a Mass Spectrometer A Mass Spectrometer is an analytical instrument that can separate charged molecules according
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
W.M. Keck Foundation Biotechnology Resource Laboratories. Erol Gulcicek Christopher Colangelo Terence Wu
 W.M. Keck Foundation Biotechnology Resource Laboratories Erol Gulcicek Christopher Colangelo Terence Wu http://info.med.yale.edu/nida-proteomics/ - Site is currently up with basic information - Overview
W.M. Keck Foundation Biotechnology Resource Laboratories Erol Gulcicek Christopher Colangelo Terence Wu http://info.med.yale.edu/nida-proteomics/ - Site is currently up with basic information - Overview
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform
 Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF
 Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF PUSHER TOF DETECTOR ZSPRAY TM Ion Source SAMPLING CONE SKIMMER RF HEXAPOLE RF HEXAPOLE QUADRUPOLE IN NARROW BANDPASS MODE
Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF PUSHER TOF DETECTOR ZSPRAY TM Ion Source SAMPLING CONE SKIMMER RF HEXAPOLE RF HEXAPOLE QUADRUPOLE IN NARROW BANDPASS MODE
Topics. Protein Bioinformatics ( ) How a Mass Spectrometers Measure Mass. Lecture 9: Quantitative Proteomics Tuesday, April 27, 2010
 Protein Bioinformatics (26.655) Lecture 9: Quantitative Proteomics Tuesday, April 27, 21 Robert N. Cole, Ph.D. Mass Spectrometry and Proteomics Facility Johns Hopkins School of Medicine 371 Broadway Research
Protein Bioinformatics (26.655) Lecture 9: Quantitative Proteomics Tuesday, April 27, 21 Robert N. Cole, Ph.D. Mass Spectrometry and Proteomics Facility Johns Hopkins School of Medicine 371 Broadway Research
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
LECTURE-15. itraq Clinical Applications HANDOUT. Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS
 LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
ABSTRACT. Catherine Fenselau, Professor, Department of Chemistry and Biochemistry
 ABSTRACT Title of Dissertation: ANALYSIS OF INTACT PROTEINS IN COMPLEX MIXTURES Avantika Dhabaria Doctor of Philosophy 2013 Directed By: Catherine Fenselau, Professor, Department of Chemistry and Biochemistry
ABSTRACT Title of Dissertation: ANALYSIS OF INTACT PROTEINS IN COMPLEX MIXTURES Avantika Dhabaria Doctor of Philosophy 2013 Directed By: Catherine Fenselau, Professor, Department of Chemistry and Biochemistry
Protein sequence mapping is commonly used to
 Reproducible Microwave-Assisted Acid Hydrolysis of Proteins Using a Household Microwave Oven and Its Combination with LC-ESI MS/MS for Mapping Protein Sequences and Modifications Nan Wang and Liang Li
Reproducible Microwave-Assisted Acid Hydrolysis of Proteins Using a Household Microwave Oven and Its Combination with LC-ESI MS/MS for Mapping Protein Sequences and Modifications Nan Wang and Liang Li
Automated Purification and Analytical Reinjection of a Small Molecule Drug, Probenecid, on a Gilson LC/MS Dual Function System
 Automated Purification and Analytical Reinjection of a Small Molecule Drug, Probenecid, on a Gilson LC/MS Dual Function System Keywords Introduction Application Note PHA0413 High Pressure Liquid Chromatography
Automated Purification and Analytical Reinjection of a Small Molecule Drug, Probenecid, on a Gilson LC/MS Dual Function System Keywords Introduction Application Note PHA0413 High Pressure Liquid Chromatography
for the Identification of Phosphorylated Peptides
 Application of a Data Dependent Neutral-Loss Experiment on the Finnigan LTQ for the Identification of Phosphorylated Peptides Gargi Choudhary Diane Cho Thermo Electron, San Jose, CA Abstracted from posters
Application of a Data Dependent Neutral-Loss Experiment on the Finnigan LTQ for the Identification of Phosphorylated Peptides Gargi Choudhary Diane Cho Thermo Electron, San Jose, CA Abstracted from posters
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
FOURIER TRANSFORM MASS SPECTROMETRY
 FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-12-1-0248 TITLE: Targeted Inhibition of Tyrosine Kinase-Mediated Epigenetic Alterations to Prevent Resurgence of Castration-Resistant Prostate Cancer PRINCIPAL INVESTIGATOR: Dr.
AD Award Number: W81XWH-12-1-0248 TITLE: Targeted Inhibition of Tyrosine Kinase-Mediated Epigenetic Alterations to Prevent Resurgence of Castration-Resistant Prostate Cancer PRINCIPAL INVESTIGATOR: Dr.
FOURIER TRANSFORM MASS SPECTROMETRY
 FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
More structural information with MS n
 PRODUCT SPECIFICATIONS The LTQ XL linear ion trap mass spectrometer More structural information with MS n The LTQ XL linear ion trap mass spectrometer delivers more structural information faster and with
PRODUCT SPECIFICATIONS The LTQ XL linear ion trap mass spectrometer More structural information with MS n The LTQ XL linear ion trap mass spectrometer delivers more structural information faster and with
Introduction to the Oligo HTCS Systems. Novatia, LLC
 Introduction to the Oligo HTCS Systems Novatia, LLC What is the Oligo HTCS and what can it do? Automated ESI/MS system that can analyze up to 14 oligonucleotide samples/day (>3/day with twin autosampler)
Introduction to the Oligo HTCS Systems Novatia, LLC What is the Oligo HTCS and what can it do? Automated ESI/MS system that can analyze up to 14 oligonucleotide samples/day (>3/day with twin autosampler)
Suppression of the Malignant Phenotype of Bladder Cancer Cells Investigations of the Proteome in Relation to Phenotype and Gene Expression
 Suppression of the Malignant Phenotype of Bladder Cancer Cells Investigations of the Proteome in Relation to Phenotype and Gene Expression Topics Modulation of Phenotype by Extracellular Matrix. Bioinformatics
Suppression of the Malignant Phenotype of Bladder Cancer Cells Investigations of the Proteome in Relation to Phenotype and Gene Expression Topics Modulation of Phenotype by Extracellular Matrix. Bioinformatics
Protein and peptide separations,
 A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
Mass Spectrometry at the Laboratory of Food Chemistry. Edwin Bakx Laboratory of Food Chemistry Wageningen University
 Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS
 Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Tandem mass spectrometry is becoming the
 Fragmentation of Phosphopeptides in an Ion Trap Mass Spectrometer Jon P. DeGnore and Jun Qin Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health,
Fragmentation of Phosphopeptides in an Ion Trap Mass Spectrometer Jon P. DeGnore and Jun Qin Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health,
Metabolomics: quantifying the phenotype
 Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES. Gilles Frache Materials Characterization Day October 14 th 2016
 LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Chemical Biology, Option II Mechanism Based Proteomic Tagging Case History CH1
 Proteome Wide Screening of Serine Protease Activity Proc Natl Acad Sci 1999, 97, 14694; Proteomics 2001, 1, 1067; Proc Natl Acad Sci 2002, 99, 10335; Biochemistry 2001, 40, 4005; J. Am. Chem. Soc., 2005,
Proteome Wide Screening of Serine Protease Activity Proc Natl Acad Sci 1999, 97, 14694; Proteomics 2001, 1, 1067; Proc Natl Acad Sci 2002, 99, 10335; Biochemistry 2001, 40, 4005; J. Am. Chem. Soc., 2005,
Biological Mass Spectrometry. April 30, 2014
 Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
MS/MS to Targeted Proteomics (MRM)
 MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
Nat Rev Mol Cell Biol (2006) 7, Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312,
 Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
The Immunoassay Guide to Successful Mass Spectrometry. Orr Sharpe Robinson Lab SUMS User Meeting October 29, 2013
 The Immunoassay Guide to Successful Mass Spectrometry Orr Sharpe Robinson Lab SUMS User Meeting October 29, 2013 What is it? Hey! Look at that! Something is reacting in here! I just wish I knew what it
The Immunoassay Guide to Successful Mass Spectrometry Orr Sharpe Robinson Lab SUMS User Meeting October 29, 2013 What is it? Hey! Look at that! Something is reacting in here! I just wish I knew what it
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Identification of Ginsenosides Using the SCIEX X500R QTOF System
 Identification of Ginsenosides Using the SCIEX X500R QTOF System Wang Sha, Cheng Haiyan, Liu Ting, Li Lijun, Jin Wenhai[Author] SCIEX, Pacific Applications Support Center (Beijing). China Background Ginseng
Identification of Ginsenosides Using the SCIEX X500R QTOF System Wang Sha, Cheng Haiyan, Liu Ting, Li Lijun, Jin Wenhai[Author] SCIEX, Pacific Applications Support Center (Beijing). China Background Ginseng
 a RT:. - 1.1 1 95 9 85 8 27.64 27.87 28.1 27.42 28.24 38.98 Low ph RPLC NL: 2.29E9 TIC MS TrypticBSA_Dig est_replicate1_ 2Mar18_Preci ous_18-1-6 75 7 65 48.75 65.8 Relative Abundance 6 55 5 45 28.54 48.98
a RT:. - 1.1 1 95 9 85 8 27.64 27.87 28.1 27.42 28.24 38.98 Low ph RPLC NL: 2.29E9 TIC MS TrypticBSA_Dig est_replicate1_ 2Mar18_Preci ous_18-1-6 75 7 65 48.75 65.8 Relative Abundance 6 55 5 45 28.54 48.98
PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX
 PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX E. Claude 1, E. Emekwue 2, M. Snel 1, T. McKenna 1 and J. Langridge 1 1: Waters Corporation, Manchester, UK 2:
PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX E. Claude 1, E. Emekwue 2, M. Snel 1, T. McKenna 1 and J. Langridge 1 1: Waters Corporation, Manchester, UK 2:
Time (min) Supplementary Figure 1: Gas decomposition products of irradiated DMC.
 200000 C 2 CH 3 CH 3 DMC 180000 160000 140000 Intensity 120000 100000 80000 60000 40000 C 2 H 6 CH 3 CH 2 CH 3 CH 3 CCH 3 EMC DEC 20000 C 3 H 8 HCCH 3 5 10 15 20 25 Time (min) Supplementary Figure 1: Gas
200000 C 2 CH 3 CH 3 DMC 180000 160000 140000 Intensity 120000 100000 80000 60000 40000 C 2 H 6 CH 3 CH 2 CH 3 CH 3 CCH 3 EMC DEC 20000 C 3 H 8 HCCH 3 5 10 15 20 25 Time (min) Supplementary Figure 1: Gas
The Detection of Allergens in Food Products with LC-MS
 The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Amadeo R. Fernández-Alba
 % of compounds % of compounds % of compounds % of compounds Amadeo R. Fernández-Alba LC-Orbitrap QExactive Focus Instrumental LOQ 1% 9% 8% 7% 6% 5% 4% 3% 2% 1% %.1 mg/g.2 mg/g Tomato.5 mg/g ddms2 Target
% of compounds % of compounds % of compounds % of compounds Amadeo R. Fernández-Alba LC-Orbitrap QExactive Focus Instrumental LOQ 1% 9% 8% 7% 6% 5% 4% 3% 2% 1% %.1 mg/g.2 mg/g Tomato.5 mg/g ddms2 Target
Nature Structural & Molecular Biology: doi: /nsmb.3218
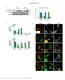 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Glycosylation analysis of blood plasma proteins
 Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
