Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids
|
|
|
- Jean Webb
- 5 years ago
- Views:
Transcription
1 Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of Physics, Faculty of Science, Kasetsart University, Bangkok, 10900, Thailand 2 Clarendon Laboratory, Department of Physics, University of Oxford, Oxford OX1 3PU, United Kingdom 3 Department of Mathematics and Computer Science & Institute for Complex Molecular Systems, Eindhoven University of Technology, MetaForum, 5600 MB Eindhoven, the Netherlands * Corresponding Authors J.W.: jirasak.w@ku.ac.th, and M.K.: mkarttu@tue.nl Figures and Tables Figure S1. The structures of lipid molecules 2 Figure S2. Snapshots of a 12-al micelle (A) and a 9-al micelle (B) viewed along the y- direction 3 Figure S3. The time evolution of the position in along the z-direction for the oxidized functional groups from the center of bilayer of 50 % oxidized lipids mixture bilayer 4 Figure S4. Water molecules were pulled across the leaflet by the aldehyde groups in the oxidized lipid tails 5 Figure S5. Definitions of the angles and lengths. 6 Figure S6. Lipid geometries. 7 Table S1. Compositions of the lipid bilayers used in this study.. 8 Table S2. The average area per lipid, bilayer thickness and volume per lipid Table S3. The average angles Table S4. The fitting parameters of the θ A distribution by using Boltzmann distribution function Table S5. The lengths of lipid tails (l 1 and l 2 ) and the distances between the tail ends (l 3 ). 12 Table S6. Summary of packing analysis.. 13
2 Figure S1. The structures of lipid molecules; grey, blue, red, purple and white balls represent united carbon, nitrogen, oxygen, phosphorus and hydrogen atoms, respectively.
3 Figure S2. Snapshots of a 12-al micelle (A) and a 9-al micelle (B) viewed along the y-direction. Lipid chains are shown in cyan and green for 12-al and 9-al, respectively, choline groups in orange, phosphate groups in yellow and oxygens in the sn-2 chain in red. Water molecules are not show for clarity.
4 Figure S3. The time evolution of the position in along the z-direction for the oxidized functional groups from the center of bilayer of 50 % oxidized lipids mixture bilayer. The black line represents the average position of the phosphate group in each of the two leaflets (the bilayer is centered at z=0 nm). The blue, red, green and purple lines show the positions of the functional groups in the oxidized tails of 13-tc, 9-tc, 12-al and 9-al in one leaflet, respectively.
5 Figure S4. Water molecules were pulled across the leaflet by the aldehyde groups in the oxidized lipid tails. Green & yellow: 9-al lipids in the upper and lower leaflets, respectively. White: PLPC. Green, yellow and white spheres: Phosphorus atoms on the different lipids. Red spheres: Oxygens in 9-al sn-2 tails. Blue & gray: Oxygens and hydrogens of water molecules, respectively.
6 Figure S5. Definitions of the angles and lengths. The interior angles of the lipid tails are represented by θ A, θ B and θ C. The tilt angle (θ N ) between the sn-1 and the bilayer normal is used to determine lipid orientation. l 1 and l 2 represent the lengths of the sn-1 and sn-2 lipid tails, respectively and l 3 is the distance between the ends of lipid tails.
7 Figure S6. Lipid geometries: (A) PLPC- cylinder, (B) peroxide lipids (13-tc, 9-tc) - cylinder, (C) aldehyde lipids (12-al, 9-al) - truncated cone and (D) PDPC - cylinder.
8 Table S1. Compositions of the lipid bilayers used in this study. All simulations were run for 1 µs. System Description Concentration of PLPC:Oxidized oxidized lipid (%) lipid molecules Final structure 1 Pure PLPC 0 128:0 Bilayer 2 PLPC + 12-al 50 64:64 Bilayer with a pore 3 PLPC + 12-al 75 32:96 Bilayer with a pore 4 Pure 12-al 100 0:128 Micelle 5 PLPC + 9-al 50 64:64 Bilayer with a pore 6 PLPC + 9-al 75 32:96 Micelle 7 Pure 9-al 100 0:128 Micelle 8 PLPC + 13-tc 50 64:64 Bilayer 9 PLPC + 13-tc 75 32:96 Bilayer 10 Pure 13-tc 100 0:128 Bilayer 11 PLPC + 9-tc 50 64:64 Bilayer 12 PLPC + 9-tc 75 32:96 Bilayer 13 Pure 9-tc 100 0:128 Bilayer 14 PLPC + PDPC 50 64:64 Bilayer 15 PLPC + PDPC 75 32:96 Bilayer 16 Pure PDPC 100 0:128 Bilayer
9 Table S2. The average area per lipid, bilayer thickness and volume per lipid. The results of % oxidized lipids concentration were obtained from previous study. 1,2 Systems Area per lipid (nm 2 ) Bilayer thickness (nm) Volume per lipid (nm 3 ) Pure PLPC 0.66 ± ± ± % 13-tc 0.66 ± ± ± % 13-tc 0.67 ± ± ± % 13-tc 0.69 ± ± ± % 13-tc 0.74 ± ± ± % 13-tc 0.79 ± ± ± 0.01 Pure 13-tc 0.80 ± ± ± % 9-tc 0.66 ± ± ± % 9-tc 0.67 ± ± ± % 9-tc 0.68 ± ± ± % 9-tc 0.71 ± ± ± % 9-tc 0.74 ± ± ± 0.00 Pure 9-tc 0.78 ± ± ± % 12-al 0.66 ± ± ± % 12-al 0.67 ± ± ± % 12-al 0.69 ± ± ± % 12-al 0.72 ± ± ± % 12-al 0.74 ± ± ± 0.01 Pure12-al % 9-al 0.66 ± ± ± % 9-al 0.67 ± ± ± % 9-al 0.68 ± ± ± % 9-al 0.71 ± ± ± % 9-al Pure 9-al % PDPC 0.66 ± ± ± % PDPC 0.65 ± ± ± 0.00 Pure PDPC 0.65 ± ± ± 0.00
10 Table S3. The average angles. Systems θ A θ B θ C θ N PLPC OXPL PLPC OXPL PLPC OXPL PLPC OXPL Pure PLPC 48 ± ± 1-30 ± 1-50% 13-tc 48 ± 1 63 ± ± 1 66 ± 0 54 ± 1 32 ± 0 30 ± 0 75% 13-tc 50 ± 2 60 ± ± 2 66 ± 1 56 ± 1 34 ± 1 34 ± 1 Pure 13-tc - 63 ± ± 2-56 ± 1-36 ± 1 50% 9-tc 50 ± 1 55 ± ± 1 66 ± 1 60 ± 1 32 ± 0 31 ± 0 75% 9-tc 51 ± 1 57 ± ± 1 66 ± 1 60 ± 0 33 ± 1 32 ± 0 Pure 9-tc - 56 ± ± 1-60 ± 1-33 ± 1 50% 12-al 52 ± 1 71 ± ± 0 41 ± 1 34 ± 0 34 ± 0 75% 12-al 53 ± 2 66 ± ± 2 45 ± 1 36 ± 2 35 ± 1 Pure12-al - 67 ± ± % 9-al 54 ± 1 72 ± ± 0 37 ± 2 36 ± 0 35 ± 1 75% 9-al 47 ± 1 73 ± ± 0 37 ± Pure 9-al - 74 ± ± % PDPC 50 ± 0 48 ± ± 0 46 ± 1 32 ± 0 31 ± 0 75% PDPC 52 ± 1 49 ± ± 0 46 ± 1 32 ± 0 32 ± 0 Pure PDPC - 50 ± ± 0-32 ± 1
11 Table S4. The fitting parameters of the θ A distribution by using Boltzmann distribution function, y = a 2 θ A 2 exp( θ A 2 2b 2). Systems a Parameters b Mean (θ A ) PLPC OXPL PLPC OXPL PLPC OXPL PLPC OXPL Pure PLPC % 13-tc % 13-tc Pure 13-tc % 9-tc % 9-tc Pure 9-tc % 12-al % 12-al Pure 12-al % 9-al % 9-al Pure 9-al % PDPC % PDPC Pure PDPC σ
12 Table S5. The lengths of lipid tails (l 1 and l 2 ) and the distances between the tail ends (l 3 ). Systems l 1 (nm) l 2 (nm) l 3 (nm) l 1 : l 2 l 2 : l 3 Pure PLPC 1.95 ± ± ± Pure 13-tc 1.88 ± ± ± Pure 9-tc 1.89 ± ± ± Pure 12-al 1.91 ± ± ± Pure 9-al 1.86 ± ± ± Pure PDPC 1.91 ± ± ±
13 Table S6. Summary of packing analysis The packing parameter of the PLPC and oxidized lipid systems were calculated as followed. (1) Bilayers (PLPC, 13-tc, 9-tc and PDPC): For lipid bilayers composed N lipid molecules, the average chain volume is v c = V/N where V is the total bilayer volume. a is the usual average area per lipid, and l c is the haft-bilayer (=monolayer) thickness. (2) Cylindrical micelles (12-al, 9-al): For cylindrical micelles composed of N lipid molecules, the hydrocarbon chain volume can be defined as v c = V/N = Lπr 2 /N and the average surface area of the lipid head group as a = 2πrL/N, where V is the total volume of micelle, and L and r are the length and the radius of the cylindrical micelle, respectively. The radius of a cylindrical micelle can be estimated from the density of the phosphorus atoms. In case of 12-al micelle r = 1.99 ± 0.15 nm and r = 1.98 ± 0.27 nm for 9-al micelle. The snapshot of cylindrical micelle (Figure S2) shown that the oxygen atoms in sn-2 chain are pointing to the micelle surface at the lipid head group region and formed stable hydrogen bond with water, suggested that the oxygen atom has co-surface area with lipid head group. Thus, the critical hydrocarbon chain l c can be estimated from the average tail length of l 1 which represents the length of non-polar chain with pointing to the center of micelle. Thus, the l c of 12-al and 9-al lipids in micelle are 1.91 ± 0.01 and 1.86 ± 0.16 nm, respectively. Systems v c a l c Packing Parameter S = v c /al c Pure PLPC Pure 13-tc Pure 9-tc Pure 12-al Pure 9-al Pure PDPC
14 References (1) Wong-ekkabut, J.; Xu, Z.; Triampo, W.; Tang, I. M.; Peter Tieleman, D.; Monticelli, L. Effect of Lipid Peroxidation on the Properties of Lipid Bilayers: A Molecular Dynamics Study. Biophys. J. 2007, 93, (2) Jarerattanachat, V.; Karttunen, M.; Wong-ekkabut, J. Molecular Dynamics Study of Oxidized Lipid Bilayers in NaCl Solution. J. Phys. Chem. B 2013, 117,
Massive oxidation of phospholipid membranes. leads to pore creation and bilayer. disintegration
 Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Gold nanocrystals at DPPC bilayer. Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad
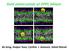 Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale
 Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
The main biological functions of the many varied types of lipids include: energy storage protection insulation regulation of physiological processes
 Big Idea In the biological sciences, a dehydration synthesis (condensation reaction) is typically defined as a chemical reaction that involves the loss of water from the reacting molecules. This reaction
Big Idea In the biological sciences, a dehydration synthesis (condensation reaction) is typically defined as a chemical reaction that involves the loss of water from the reacting molecules. This reaction
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
in-silico Design of Nanoparticles for Transdermal Drug Delivery Application
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
Supplementary Information: A Critical. Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers
 Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
WHAT IS A LIPID? OBJECTIVE The objective of this worksheet is to understand the structure and function of lipids
 WHAT IS A LIPID? OBJECTIVE The objective of this worksheet is to understand the structure and function of lipids PART A: Understanding Lipids Lipids are more commonly known as fats and include triglycerides,
WHAT IS A LIPID? OBJECTIVE The objective of this worksheet is to understand the structure and function of lipids PART A: Understanding Lipids Lipids are more commonly known as fats and include triglycerides,
3.1.3 Lipids. Source: AQA Spec
 alevelbiology.co.uk SPECIFICATION Triglycerides and phospholipids are two groups of lipid. Triglycerides are formed by the condensation of one molecule of glycerol and three molecules of fatty acid. A
alevelbiology.co.uk SPECIFICATION Triglycerides and phospholipids are two groups of lipid. Triglycerides are formed by the condensation of one molecule of glycerol and three molecules of fatty acid. A
How Cardiolipin Peroxidation Alters the Properties of the Inner Mitochondrial Membrane?
 https://helda.helsinki.fi How Cardiolipin Peroxidation Alters the Properties of the Inner Mitochondrial Membrane? Vähäheikkilä, Marjut 2018-08 Vähäheikkilä, M, Peltomaa, T, Róg, T, Vazdar, M, Pöyry, S
https://helda.helsinki.fi How Cardiolipin Peroxidation Alters the Properties of the Inner Mitochondrial Membrane? Vähäheikkilä, Marjut 2018-08 Vähäheikkilä, M, Peltomaa, T, Róg, T, Vazdar, M, Pöyry, S
C 6 H 12 O 6 + 6O 2 6CO 2 + 6H 2 O
 Sample Questions for the Individual Round Exam 1. Glucose is the most basic sugar involved in human metabolism. Its structure is provided below: a. The overall reaction of glucose metabolism is given below.
Sample Questions for the Individual Round Exam 1. Glucose is the most basic sugar involved in human metabolism. Its structure is provided below: a. The overall reaction of glucose metabolism is given below.
Biological Membranes. Lipid Membranes. Bilayer Permeability. Common Features of Biological Membranes. A highly selective permeability barrier
 Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and in a
 Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Life Sciences 1a. Practice Problems 4
 Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
H 2 O. Liquid, solid, and vapor coexist in the same environment
 Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics
 Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics Haneesh Kesari, Reza Banki, and Misty Davies Project Final Paper ME346 Stanford University December 15, 2005 1 Introduction Understanding
Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics Haneesh Kesari, Reza Banki, and Misty Davies Project Final Paper ME346 Stanford University December 15, 2005 1 Introduction Understanding
0.5 nm nm acyl tail region (hydrophobic) 1.5 nm. Hydrophobic repulsion organizes amphiphilic molecules: These scales are 5 10xk B T:
 Lecture 31: Biomembranes: The hydrophobic energy scale and membrane behavior 31.1 Reading for Lectures 30-32: PKT Chapter 11 (skip Ch. 10) Upshot of last lecture: Generic membrane lipid: Can be cylindrical
Lecture 31: Biomembranes: The hydrophobic energy scale and membrane behavior 31.1 Reading for Lectures 30-32: PKT Chapter 11 (skip Ch. 10) Upshot of last lecture: Generic membrane lipid: Can be cylindrical
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Chapter 12: Membranes. Voet & Voet: Pages
 Chapter 12: Membranes Voet & Voet: Pages 390-415 Slide 1 Membranes Essential components of all living cells (define boundry of cells) exclude toxic ions and compounds; accumulation of nutrients energy
Chapter 12: Membranes Voet & Voet: Pages 390-415 Slide 1 Membranes Essential components of all living cells (define boundry of cells) exclude toxic ions and compounds; accumulation of nutrients energy
We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field
 Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Physical Cell Biology Lecture 10: membranes elasticity and geometry. Hydrophobicity as an entropic effect
 Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
BIOMOLECULES. (AKA MACROMOLECULES) Name: Block:
 BIOMOLECULES (AKA MACROMOLECULES) Name: Block: BIOMOLECULES POGIL All living things share the same chemical building blocks and depend on chemical processes for survival. Life without carbon (C) would
BIOMOLECULES (AKA MACROMOLECULES) Name: Block: BIOMOLECULES POGIL All living things share the same chemical building blocks and depend on chemical processes for survival. Life without carbon (C) would
Chem 60 Takehome Test 2 Student Section
 Multiple choice: 1 point each. Mark only one answer for each question. 1. are composed primarily of carbon and hydrogen, but may also include oxygen, nitrogen, sulfur, phosphorus, and a few other elements.
Multiple choice: 1 point each. Mark only one answer for each question. 1. are composed primarily of carbon and hydrogen, but may also include oxygen, nitrogen, sulfur, phosphorus, and a few other elements.
SAM Teachers Guide Lipids and Carbohydrates
 SAM Teachers Guide Lipids and Carbohydrates Overview Students will explore the structure and function of two of the four major macromolecules, lipids and carbohydrates. They will look specifically at the
SAM Teachers Guide Lipids and Carbohydrates Overview Students will explore the structure and function of two of the four major macromolecules, lipids and carbohydrates. They will look specifically at the
2.3 Carbon-Based Molecules. KEY CONCEPT Carbon-based molecules are the foundation of life.
 KEY CONCEPT Carbon-based molecules are the foundation of life. Carbon atoms have unique bonding properties. Carbon forms covalent bonds with up to four other atoms, including other carbon atoms. Carbon-based
KEY CONCEPT Carbon-based molecules are the foundation of life. Carbon atoms have unique bonding properties. Carbon forms covalent bonds with up to four other atoms, including other carbon atoms. Carbon-based
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Role of Surface Ligands in Nanoparticle Permeation through a Model. Membrane: A Coarse grained Molecular Dynamics Simulations Study
 Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Ethanol Induces the Formation of Water-Permeable Defects in Model. Bilayers of Skin Lipids
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes
 Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Changes in a Phospholipid Bilayer Induced by the Hydrolysis of a Phospholipase A 2 Enzyme: A Molecular Dynamics Simulation Study
 Biophysical Journal Volume 80 February 2001 565 578 565 Changes in a Phospholipid Bilayer Induced by the Hydrolysis of a Phospholipase A 2 Enzyme: A Molecular Dynamics Simulation Study Marja T. Hyvönen,*
Biophysical Journal Volume 80 February 2001 565 578 565 Changes in a Phospholipid Bilayer Induced by the Hydrolysis of a Phospholipase A 2 Enzyme: A Molecular Dynamics Simulation Study Marja T. Hyvönen,*
Carbohydrates, Lipids, Proteins, and Nucleic Acids
 Carbohydrates, Lipids, Proteins, and Nucleic Acids Is it made of carbohydrates? Organic compounds composed of carbon, hydrogen, and oxygen in a 1:2:1 ratio. A carbohydrate with 6 carbon atoms would have
Carbohydrates, Lipids, Proteins, and Nucleic Acids Is it made of carbohydrates? Organic compounds composed of carbon, hydrogen, and oxygen in a 1:2:1 ratio. A carbohydrate with 6 carbon atoms would have
Elements & Macromolecules in Organisms
 Name: Period: Date: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
Name: Period: Date: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
Inorganic compounds: Usually do not contain carbon H 2 O Ca 3 (PO 4 ) 2 NaCl Carbon containing molecules not considered organic: CO 2
 Organic Chemistry The study of carbon-containing compounds and their properties. Biochemistry: Made by living things All contain the elements carbon and hydrogen Inorganic: Inorganic compounds: All other
Organic Chemistry The study of carbon-containing compounds and their properties. Biochemistry: Made by living things All contain the elements carbon and hydrogen Inorganic: Inorganic compounds: All other
Biology 2E- Zimmer Protein structure- amino acid kit
 Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation
 Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Supplementary information for Effects of Stretching Speed on. Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular
 Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Macromolecules Cut & Paste
 Macromolecules Cut & Paste Adapted from http://mrswords.weebly.com/uploads/1/5/2/4/15244382/ch_6-3_life_molecules_cut-out_lab.pdf INTRODUCTION Many of the molecules in living cells are so large that they
Macromolecules Cut & Paste Adapted from http://mrswords.weebly.com/uploads/1/5/2/4/15244382/ch_6-3_life_molecules_cut-out_lab.pdf INTRODUCTION Many of the molecules in living cells are so large that they
Modern Aspects of Colloid Science MICELLES
 Modern Aspects of Colloid Science MICELLES critical micelle concentration (CMC) micellar shape determination of critical micelle concentration purity of surfactants Krafft temperature micellar equilibria
Modern Aspects of Colloid Science MICELLES critical micelle concentration (CMC) micellar shape determination of critical micelle concentration purity of surfactants Krafft temperature micellar equilibria
Effect of Lipid Peroxidation on the Properties of Lipid Bilayers: A Molecular Dynamics Study
 Biophysical Journal Volume 93 December 2007 4225 4236 4225 Effect of Lipid Peroxidation on the Properties of Lipid Bilayers: A Molecular Dynamics Study Jirasak Wong-ekkabut,* y Zhitao Xu,* Wannapong Triampo,
Biophysical Journal Volume 93 December 2007 4225 4236 4225 Effect of Lipid Peroxidation on the Properties of Lipid Bilayers: A Molecular Dynamics Study Jirasak Wong-ekkabut,* y Zhitao Xu,* Wannapong Triampo,
Chapter 1 Membrane Structure and Function
 Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
2.3 Carbon-Based Molecules CARBON BASED MOLECULES
 CARBON BASED MOLECULES KEY CONCEPTS Carbon-based molecules are the foundation of life. Lipids are one class of organic molecules. This group includes fats, oils, waxes, and steroids. Lipids are made of
CARBON BASED MOLECULES KEY CONCEPTS Carbon-based molecules are the foundation of life. Lipids are one class of organic molecules. This group includes fats, oils, waxes, and steroids. Lipids are made of
B i o c h e m i s t r y N o t e s
 14 P a g e Carbon Hydrogen Nitrogen Oxygen Phosphorus Sulfur ~Major ~Found in all ~Found in most ~Found in all component of all organic organic molecules. molecules. ~Major structural atom in all organic
14 P a g e Carbon Hydrogen Nitrogen Oxygen Phosphorus Sulfur ~Major ~Found in all ~Found in most ~Found in all component of all organic organic molecules. molecules. ~Major structural atom in all organic
Macromolecule modeling lab
 Macromolecule modeling lab Goal: Use ball and stick models to build some of the macromolecules that make up cells also see how complex and large these macromolecules can be Directions to read before starting:
Macromolecule modeling lab Goal: Use ball and stick models to build some of the macromolecules that make up cells also see how complex and large these macromolecules can be Directions to read before starting:
Dissipative particle dynamics of tension-induced membrane fusion
 Molecular Simulation Vol. 35, No. 7, June 2009, 554 560 Dissipative particle dynamics of tension-induced membrane fusion Andrea Grafmüller a, Julian Shillcock b and Reinhard Lipowsky a * a Theory and Bio-Systems,
Molecular Simulation Vol. 35, No. 7, June 2009, 554 560 Dissipative particle dynamics of tension-induced membrane fusion Andrea Grafmüller a, Julian Shillcock b and Reinhard Lipowsky a * a Theory and Bio-Systems,
Biological Molecules
 Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
Lecture-3. Water and Phospholipid
 Lecture-3 Water and Phospholipid Life on earth began in water and evolved there for three billion years before spreading onto land. Although most of the water in liquid form, it is also in solid form and
Lecture-3 Water and Phospholipid Life on earth began in water and evolved there for three billion years before spreading onto land. Although most of the water in liquid form, it is also in solid form and
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
C H O N P S. Name : Color the Elements on the Periodic Table as listed below
 Name : Unit 9: and Changes in Digestion (if found, please return to Room 714) 7.6A Organic Compounds Identify that organic compounds contain CHONPS. Organic Compounds contain what? 6 1 8 7 15 16 C H O
Name : Unit 9: and Changes in Digestion (if found, please return to Room 714) 7.6A Organic Compounds Identify that organic compounds contain CHONPS. Organic Compounds contain what? 6 1 8 7 15 16 C H O
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Millicient A. Firestone, Amanda C. Wolf, and Sonke Seifert. Chris Bianchi 4/30/12
 Small-Angle X-ray Scattering of the Interaction of Poly(ethylene oxide)-b-poly(propylene oxide)-b- Poly(ethylene oxide)triblock Copolymers with Lipid Bilayers Millicient A. Firestone, Amanda C. Wolf, and
Small-Angle X-ray Scattering of the Interaction of Poly(ethylene oxide)-b-poly(propylene oxide)-b- Poly(ethylene oxide)triblock Copolymers with Lipid Bilayers Millicient A. Firestone, Amanda C. Wolf, and
The Chemical Building Blocks of Life. Chapter 3
 The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
Biological Molecules
 The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
Biology Chapter 2 Review
 Biology Chapter 2 Review Vocabulary: Define the following words on a separate piece of paper. Element Compound Ion Ionic Bond Covalent Bond Molecule Hydrogen Bon Cohesion Adhesion Solution Solute Solvent
Biology Chapter 2 Review Vocabulary: Define the following words on a separate piece of paper. Element Compound Ion Ionic Bond Covalent Bond Molecule Hydrogen Bon Cohesion Adhesion Solution Solute Solvent
BIOB111_CHBIO - Tutorial activity for Session 12
 BIOB111_CHBIO - Tutorial activity for Session 12 General topic for week 6 Session 12 Lipids Useful Links: 1. Animations on Cholesterol (its synthesis, lifestyle factors, LDL) http://www.wiley.com/college/boyer/0470003790/animations/cholesterol/cholesterol.htm
BIOB111_CHBIO - Tutorial activity for Session 12 General topic for week 6 Session 12 Lipids Useful Links: 1. Animations on Cholesterol (its synthesis, lifestyle factors, LDL) http://www.wiley.com/college/boyer/0470003790/animations/cholesterol/cholesterol.htm
Membranes & Membrane Proteins
 School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
Membrane Structure and Membrane Transport of Small Molecules. Assist. Prof. Pinar Tulay Faculty of Medicine
 Membrane Structure and Membrane Transport of Small Molecules Assist. Prof. Pinar Tulay Faculty of Medicine Introduction Cell membranes define compartments of different compositions. Membranes are composed
Membrane Structure and Membrane Transport of Small Molecules Assist. Prof. Pinar Tulay Faculty of Medicine Introduction Cell membranes define compartments of different compositions. Membranes are composed
and hydrophilic and how they relate to solubility.
 o o o and hydrophilic and how they relate to solubility. o o o o o o o o Page 1: Introduction Page 2: 1. Hydrocarbons are referred to as organic molecules with a "backbone." Take a snapshot of the hydrocarbon
o o o and hydrophilic and how they relate to solubility. o o o o o o o o Page 1: Introduction Page 2: 1. Hydrocarbons are referred to as organic molecules with a "backbone." Take a snapshot of the hydrocarbon
2
 1 2 What defines all living systems is the ability to generate a chemical environment inside the cell that is different from the extracellular one. The plasma membrane separates the inside of the cell
1 2 What defines all living systems is the ability to generate a chemical environment inside the cell that is different from the extracellular one. The plasma membrane separates the inside of the cell
Chemical Formulas. Chemical Formula CH 3 COCHCHOCHClCHNH Lewis Dot Structure
 Biochemistry . Chemical Formulas A chemical formula represents the chemical makeup of a compound. It shows the numbers and kinds of atoms present in a compound. It is a kind of shorthand that scientists
Biochemistry . Chemical Formulas A chemical formula represents the chemical makeup of a compound. It shows the numbers and kinds of atoms present in a compound. It is a kind of shorthand that scientists
Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers
 PUBLISHED IN: JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE, VOL. 2, PP. 40-43 (2005). Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers M. Kupiainen, E. Falck, S. Ollila, P.
PUBLISHED IN: JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE, VOL. 2, PP. 40-43 (2005). Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers M. Kupiainen, E. Falck, S. Ollila, P.
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner thomas.stockner@meduniwien.ac.at Molecular biology Molecular modelling Membranes environment Many cellular functions occur in or around membranes: energy
Simulationen von Lipidmembranen Thomas Stockner thomas.stockner@meduniwien.ac.at Molecular biology Molecular modelling Membranes environment Many cellular functions occur in or around membranes: energy
Carbon. Isomers. The Chemical Building Blocks of Life
 The Chemical Building Blocks of Life Carbon Chapter 3 Framework of biological molecules consists primarily of carbon bonded to Carbon O, N, S, P or H Can form up to 4 covalent bonds Hydrocarbons molecule
The Chemical Building Blocks of Life Carbon Chapter 3 Framework of biological molecules consists primarily of carbon bonded to Carbon O, N, S, P or H Can form up to 4 covalent bonds Hydrocarbons molecule
General Chemistry. Ch. 10
 General Chemistry Ch. 10 Essentials of Organic Chemistry Most biological important molecules are composed of organic compounds. These are mostly produced by biological systems. Organic molecules contain
General Chemistry Ch. 10 Essentials of Organic Chemistry Most biological important molecules are composed of organic compounds. These are mostly produced by biological systems. Organic molecules contain
Surfactants. The Basic Theory. Surfactants (or surface active agents ): are organic compounds with at least one lyophilic. Paints and Adhesives
 Surfactants Surfactants (or surface active agents ): are organic compounds with at least one lyophilic ( solvent-loving ) group and one lyophobic ( solvent-fearing ) group in the molecule. In the simplest
Surfactants Surfactants (or surface active agents ): are organic compounds with at least one lyophilic ( solvent-loving ) group and one lyophobic ( solvent-fearing ) group in the molecule. In the simplest
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Cells and Their Environment Chapter 8. Cell Membrane Section 1
 Cells and Their Environment Chapter 8 Cell Membrane Section 1 Homeostasis Key Idea: One way that a cell maintains homeostasis is by controlling the movement of substances across the cell membrane. Homeostasis
Cells and Their Environment Chapter 8 Cell Membrane Section 1 Homeostasis Key Idea: One way that a cell maintains homeostasis is by controlling the movement of substances across the cell membrane. Homeostasis
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
Chapter 3. Table of Contents. Section 1 Carbon Compounds. Section 2 Molecules of Life. Biochemistry
 Biochemistry Table of Contents Section 1 Carbon Compounds Section 2 Molecules of Life Section 1 Carbon Compounds Objectives Distinguish between organic and inorganic compounds. Explain the importance of
Biochemistry Table of Contents Section 1 Carbon Compounds Section 2 Molecules of Life Section 1 Carbon Compounds Objectives Distinguish between organic and inorganic compounds. Explain the importance of
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2011.151 Cell Entry of One-Dimensional Nanomaterials Occurs by Tip Recognition and Rotation Supplemental Information Xinghua Shi 1,4, Annette von dem Bussche
SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2011.151 Cell Entry of One-Dimensional Nanomaterials Occurs by Tip Recognition and Rotation Supplemental Information Xinghua Shi 1,4, Annette von dem Bussche
The Influence of Ceramide Tail Length on the. Structure of Bilayers Composed of Stratum. Corneum Lipids
 The Influence of Ceramide Tail Length on the Structure of Bilayers Composed of Stratum Corneum Lipids T. C. Moore, R. Hartkamp, C. R. Iacovella, A. L. Bunge, and C. M c Cabe Condensed Title: Stratum Corneum
The Influence of Ceramide Tail Length on the Structure of Bilayers Composed of Stratum Corneum Lipids T. C. Moore, R. Hartkamp, C. R. Iacovella, A. L. Bunge, and C. M c Cabe Condensed Title: Stratum Corneum
Supporting Information: Revisiting partition in. hydrated bilayer systems
 Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
A: All atom molecular simulation systems
 Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
Membrane Simulations in GROMACS Lipid bilayers and membrane proteins
 Membrane Simulations in GROMACS Lipid bilayers and membrane proteins Justin A. Lemkul September 13, 2013 jalemkul@outerbanks.umaryland.edu Objectives Learn the fundamentals of Self-consistent force fields
Membrane Simulations in GROMACS Lipid bilayers and membrane proteins Justin A. Lemkul September 13, 2013 jalemkul@outerbanks.umaryland.edu Objectives Learn the fundamentals of Self-consistent force fields
Classification, functions and structure
 Classification, functions and structure Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine and Pharmacy "Nicolae Testemitanu" Lipids
Classification, functions and structure Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine and Pharmacy "Nicolae Testemitanu" Lipids
Translocation of C 60 and Its Derivatives Across a Lipid Bilayer
 Translocation of C 60 and Its Derivatives Across a Lipid Bilayer Rui Qiao* NANO LETTERS 2007 Vol. 7, No. 3 614-619 Department of Mechanical Engineering, Clemson UniVersity, Clemson, South Carolina 29634
Translocation of C 60 and Its Derivatives Across a Lipid Bilayer Rui Qiao* NANO LETTERS 2007 Vol. 7, No. 3 614-619 Department of Mechanical Engineering, Clemson UniVersity, Clemson, South Carolina 29634
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Chemical Compounds in Cells
 Cell Structure and Function Name Date Class Cell Structure and Function Guided Reading and Study Chemical Compounds in Cells This section identifies the basic building blocks of cells. It also explains
Cell Structure and Function Name Date Class Cell Structure and Function Guided Reading and Study Chemical Compounds in Cells This section identifies the basic building blocks of cells. It also explains
Acta Crystallographica Section D
 Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: A conformational landscape for alginate secretion across the outer membrane of Pseudomonas aeruginosa
Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: A conformational landscape for alginate secretion across the outer membrane of Pseudomonas aeruginosa
Reading for lecture 6
 Reading for lecture 6 1. Lipids and Lipid Bilayers 2. Membrane Proteins Voet and Voet, Chapter 11 Alberts et al Chapter 6 Jones, R.A.L, Soft Condensed Matter 195pp Oxford University Press, ISBN 0-19-850590-6
Reading for lecture 6 1. Lipids and Lipid Bilayers 2. Membrane Proteins Voet and Voet, Chapter 11 Alberts et al Chapter 6 Jones, R.A.L, Soft Condensed Matter 195pp Oxford University Press, ISBN 0-19-850590-6
Cholesterol Modulates the Membrane Effects and Spatial Organization of Membrane-Penetrating Ligands for G-Protein Coupled Receptors
 12046 J. Phys. Chem. B 2010, 114, 12046 12057 Cholesterol Modulates the Membrane Effects and Spatial Organization of Membrane-Penetrating Ligands for G-Protein Coupled Receptors George Khelashvili,* Sayan
12046 J. Phys. Chem. B 2010, 114, 12046 12057 Cholesterol Modulates the Membrane Effects and Spatial Organization of Membrane-Penetrating Ligands for G-Protein Coupled Receptors George Khelashvili,* Sayan
Lipids are used to store and excess energy from extra carbohydrates in animals
 Lipids Lipids are a major source of energy used by cells, however lipids are more difficult for your body to break down. They produce nearly twice the amount of energy than proteins or carbohydrates. Lipids
Lipids Lipids are a major source of energy used by cells, however lipids are more difficult for your body to break down. They produce nearly twice the amount of energy than proteins or carbohydrates. Lipids
Guided Inquiry Skills Lab. Additional Lab 1 Making Models of Macromolecules. Problem. Introduction. Skills Focus. Materials.
 Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
ก Chirality Control on Lipid Nanotubule Morphology Investigated by Circular Dichroism 1, 1, 2, 2,,
 ก Chirality Control on Lipid Nanotubule Morphology Investigated by Circular Dichroism 1, 1, 2, 2,, ก 3, 1 Yuwathida Jantippana 1, Weerawat Intaratat 1, Wisit Singhsomroje 2, Sujint Wangsuya 2, Piboon Pantu
ก Chirality Control on Lipid Nanotubule Morphology Investigated by Circular Dichroism 1, 1, 2, 2,, ก 3, 1 Yuwathida Jantippana 1, Weerawat Intaratat 1, Wisit Singhsomroje 2, Sujint Wangsuya 2, Piboon Pantu
Aim: What are the molecules of life?
 Aim: What are the molecules of life? Do Now: List the elements & compounds cycled through ecosystems. Homework: Read pp. 59 63 P. 63 # 1,2,3,4,5 Vocabulary: Carbohydrate, lipid, protein, amino acid, nucleic
Aim: What are the molecules of life? Do Now: List the elements & compounds cycled through ecosystems. Homework: Read pp. 59 63 P. 63 # 1,2,3,4,5 Vocabulary: Carbohydrate, lipid, protein, amino acid, nucleic
2 3 Carbon Compounds (Macromolecules)
 2 3 Carbon Compounds (Macromolecules) Slide 1 of 37 Organic Chemistry Organic chemistry is the study of all compounds that contain bonds between carbon atoms. Slide 2 of 37 Carbon Living organisms are
2 3 Carbon Compounds (Macromolecules) Slide 1 of 37 Organic Chemistry Organic chemistry is the study of all compounds that contain bonds between carbon atoms. Slide 2 of 37 Carbon Living organisms are
Macromolecules Chapter 2.3
 Macromolecules Chapter 2.3 E.Q. What are the 4 main macromolecues found in living things and what are their functions? Carbon-Based Molecules Why is carbon called the building block of life? Carbon atoms
Macromolecules Chapter 2.3 E.Q. What are the 4 main macromolecues found in living things and what are their functions? Carbon-Based Molecules Why is carbon called the building block of life? Carbon atoms
Main Functions maintain homeostasis
 The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Bio 12 Chapter 2 Test Review
 Bio 12 Chapter 2 Test Review 1.Know the difference between ionic and covalent bonds In order to complete outer shells in electrons bonds can be Ionic; one atom donates or receives electrons Covalent; atoms
Bio 12 Chapter 2 Test Review 1.Know the difference between ionic and covalent bonds In order to complete outer shells in electrons bonds can be Ionic; one atom donates or receives electrons Covalent; atoms
Student Handout. green B 1. foam pieces. ) into a single unit to model the substrate in this reaction.
 Student Handout Introduction Enzymes are specialized proteins that catalyze or speed up chemical reactions within cells. The substance upon which an enzyme acts is called a substrate. Substrates are small
Student Handout Introduction Enzymes are specialized proteins that catalyze or speed up chemical reactions within cells. The substance upon which an enzyme acts is called a substrate. Substrates are small
SAM Teacher s Guide Four Levels of Protein Structure
 SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Activity Handout for Macromolecules - Station 1. Developed by Dr. Greg Perrier ACTIVITY MACROMOLECULES. Introduction
 Activity Handout for Macromolecules - Station 1 Developed by Dr. Greg Perrier ACTIVITY MACROMOLECULES Introduction In this exercise you will learn about the different functional groups that are added onto
Activity Handout for Macromolecules - Station 1 Developed by Dr. Greg Perrier ACTIVITY MACROMOLECULES Introduction In this exercise you will learn about the different functional groups that are added onto
GUTS Lecture Syllabus for Lipid Structure and Nomenclature
 GUTS Lecture Syllabus for Lipid Structure and Nomenclature For Questions or Assistance contact: Dr. Gwen Sancar, gsancar@ad.unc.edu Learning bjectives After completing the GUTS lecture and associated self-
GUTS Lecture Syllabus for Lipid Structure and Nomenclature For Questions or Assistance contact: Dr. Gwen Sancar, gsancar@ad.unc.edu Learning bjectives After completing the GUTS lecture and associated self-
