THERE is broad interest in genetic loci (called quantitative
|
|
|
- Camilla Avis Stewart
- 5 years ago
- Views:
Transcription
1 Copyright Ó 2006 by the Genetics Society of America DOI: /genetics The X Chromosome in Quantitative Trait Locus Mapping Karl W. Broman,*,1 Śaunak Sen, Sarah E. Owens,,2 Ani Manichaikul,* E. Michelle Southard-Smith,3 and Gary A. Churchill *Department of Biostatistics, Johns Hopkins University, Baltimore, Maryland 21205, Department of Epidemiology and Biostatistics, University of California, San Francisco, California 94107, Division of Genetic Medicine, Department of Medicine, Vanderbilt University School of Medicine, Nashville, Tennessee and The Jackson Laboratory, Bar Harbor, Maine Manuscript received May 22, 2006 Accepted for publication September 18, 2006 ABSTRACT The X chromosome requires special treatment in the mapping of quantitative trait loci (QTL). However, most QTL mapping methods, and most computer programs for QTL mapping, have focused exclusively on autosomal loci. We describe a method for appropriate treatment of the X chromosome for QTL mapping in experimental crosses. We address the important issue of formulating the null hypothesis of no linkage appropriately. If the X chromosome is treated like an autosome, a sex difference in the phenotype can lead to spurious linkage on the X chromosome. Further, the number of degrees of freedom for the linkage test may be different for the X chromosome than for autosomes, and so an X chromosome-specific significance threshold is required. To address this issue, we propose a general procedure to obtain chromosome-specific significance thresholds that controls the genomewide false positive rate at the desired level. We apply our methods to data on gut length in a large intercross of mice carrying the Sox10 Dom mutation, a model of Hirschsprung disease. We identified QTL contributing to variation in gut length on chromosomes 5 and 18. We found suggestive evidence of linkage to the X chromosome, which would be viewed as strong evidence of linkage if the X chromosome was treated as an autosome. Our methods have been implemented in the package R/qtl. THERE is broad interest in genetic loci (called quantitative trait loci, QTL) that contribute to variation in quantitative traits, and so numerous statistical methods and computer programs have been developed to map QTL. Virtually all of this work has focused exclusively on autosomal loci. However, the X chromosome displays special behavior and must be treated differently in QTL mapping. Often crosses are set up to avoid recombination on the X chromosome. When the X chromosome is recombining in a cross, there are several possible patterns, and each requires a different analysis. For example, in a backcross in which the X chromosome is segregating, males are hemizygous A or B, while females have genotype AA or AB. Thus, rather than comparing, as for the autosomes, the phenotypic means between the AA and AB genotype groups, the X chromosome requires a comparison of the phenotypic means across four genotypic groups. Both Rance et al. (1997) and Ahmadiyeh et al. (2003) described methods for the analysis of the X chromosome in the case of reciprocal F 2 intercrosses. However, 1 Corresponding author: Department of Biostatistics, Johns Hopkins University, 615 N. Wolfe St., Baltimore, MD kbroman@jhsph.edu 2 Present address: Department of Pediatrics, Vanderbilt University School of Medicine, Nashville, TN Author michelle.southard-smith@vanderbilt.edu (regarding data in article). a number of important cases, and the issue of appropriate significance thresholds for the X chromosome, remain to be addressed. Here, we describe methods for the treatment of the X chromosome in QTL mapping in experimental crosses that are suitable for routine use in QTL mapping. We develop our ideas in the context of rodent models, but our approach is general. We consider backcrosses and intercrosses derived from two inbred strains, A and B, and describe the necessary modifications to standard interval mapping. The most important modification concerns the formulation of the null hypothesis of no linkage to avoid spurious linkage to the X chromosome as a result of sex or cross-direction differences in the phenotype. Sex differences are observed in many phenotypes, and systematic phenotypic differences between reciprocal crosses may arise, for example, from parent-of-origin effects. If not taken into account, such systematic differences can lead to large LOD scores on the X chromosome even in the absence of X chromosome linkage. In addition, to account for the fact that the number of degrees of freedom for the linkage test on the X chromosome may be different from that on the autosomes, we propose a procedure to obtain an X chromosomespecific significance threshold while controlling the genomewide false positive rate at the desired level. To illustrate our approach, we study data on gut length in a large mouse intercross. The phenotype Genetics 174: (December 2006)
2 2152 K. W. Broman et al. Figure 1. The behavior of the X chromosome in a backcross. Circles and squares correspond to females and males, respectively. Open and hatched bars correspond to DNA from strains A and B, respectively. The small bar is the Y chromosome. exhibits a sex difference. Because of this, if the chromosome is treated as an autosome, there is a spurious inflation in the evidence for linkage to the X chromosome. METHODS X chromosome analysis: Central to the appropriate treatment of the X chromosome is an examination of its segregation behavior, which is markedly different from that of the autosomes. In particular, the behavior of the X chromosome depends on the direction of the cross, as well as the sex of the progeny. We enumerate the possibilities in Figures 1 and 2 for backcross and intercross populations. In Figure 1, the four possible backcrosses to a single strain are presented. For the backcrosses in Figure 1, c and d, in which the F 1 parent is male, the X chromosome is not subject to recombination; thus, we omit these crosses from further consideration. In Figure 1, a and b, in which the F 1 parent is female, the X chromosome does recombine and the order of the cross Figure 2. The behavior of the X chromosome in an intercross. Circles and squares correspond to females and males, respectively. Open and hatched bars correspond to DNA from strains A and B, respectively. The small bar is the Y chromosome. producing the F 1 parent is seen to have no impact on the behavior of the X chromosome in the backcross progeny. Backcrosses to the other strain would be similar, and we do not consider the combined analysis of backcrosses to each strain, as we view it to be outside the scope of this article. The combined analysis of multiple types of crosses is feasible on the basis of the principles we outline below. In Figure 2, the four possible intercrosses are presented. In all cases, F 2 progeny have a single X chromosome subject to recombination, and male F 2 progeny are, at any given locus, hemizygous A or B. In the cases that the F 1 male parent was derived from a cross A 3 B (with the A parent being female; Figure 2, a and b), the female F 2 progeny are either AA or AB. In the cases that the F 1 male parent was derived from a cross B 3 A (with the B parent being female; Figure 2, c and d), the female F 2 progeny are either BB or AB. Note that the direction of the cross giving the female F 1 parent does not affect the behavior of the X chromosome in the F 2 progeny. Thus, when we discuss the direction of the intercross, we consider only the direction of the cross that produced the male F 1 parent. The crosses in Figure 2, a and b, are
3 TABLE 1 Proposed contrasts for analysis of the X chromosome in standard crosses Cross Direction Sexes Contrasts Null hypothesis d.f. BC Both AA:AB:AY:BY Female:male 2 BC Females AA:AB Grand mean 1 BC Males AY:BY Grand mean 1 F 2 Both Both AA:ABf:ABr:BB:AY:BY Female forw:female rev:male 3 F 2 Both Females AA:ABf:ABr:BB Female forw:female rev 2 F 2 Both Males AY:BY Grand mean 1 F 2 One Both AA:AB:AY:BY Female:male 2 F 2 One Females AA:AB Grand mean 1 F 2 One Males AY:BY Grand mean 1 Forw, forward; rev, reverse. X Chromosome in QTL Mapping 2153 treated the same, and the crosses in Figure 2, c and d, are treated the same. We neglect the possibility of mitochondrial, Y chromosome, imprinting, or maternal effects; such parent-of-origin specific effects could be inspected through the inclusion of covariates that indicate the specific cross from which an animal was derived. Our goal is to adapt standard interval mapping (Lander and Botstein 1989) for the X chromosome. As with standard interval mapping, we survey all single- QTL models contributing to the phenotype by considering a grid of genomic positions. At each putative locus we construct a likelihood-ratio test for the hypothesis of no linkage to the X chromosome. The likelihood is maximized under the null and alternative hypothesis using the EM algorithm (Dempster et al. 1977), and the ratio is presented in base 10 logarithms to give a LOD score. The two key ingredients are appropriate formulation of the null and alternative hypotheses and calculation of the genotype probabilities at the putative locus under the null hypothesis. We discuss these, in turn, below. The choice of the null and alternative hypotheses requires careful thought. Since our goal is to develop a procedure for routine use in QTL mapping, our choice of genotype comparisons was based on the following basic principles. First, a sex or cross-direction difference in the phenotype should not lead to spurious linkage to the X chromosome. Second, the set of comparisons should be parsimonious but reasonable. Third, the null hypothesis must be nested within the alternative hypothesis. Our choices for all possible cases are presented in Table 1; we explain how we arrived at the choices through specific examples, below. Consider, for example, an intercross performed in one direction and including both sexes. Females then have X chromosome genotype AA or AB, while males are hemizygous A or B. Males that are hemizygous B should be treated separately from the AA or AB females, and so the null hypothesis must then allow for a sex difference in the phenotype, as otherwise the presence of such a sex difference would cause spurious linkage to the X chromosome. For the null hypothesis to be nested within the alternative, the alternative must allow separate phenotype averages for the genotype groups AA, AB, AY, and BY (see Table 1). We call this the contrast AA:AB:AY:BY. Note that there are 2 d.f. for this test of linkage, just as for the autosomes, as there are four mean parameters under the alternative and two under the null (the average phenotype for each of females and males). A somewhat more complex example is for the case that both directions of the intercross were performed, but only females were phenotyped. As the AA individuals come from one direction of the cross (which we call the forward direction) and the BB individuals come from the other direction of the cross (the reverse direction), a cross direction effect on the phenotype would cause spurious linkage to the X chromosome if the null hypothesis did not allow for that effect. But then, for the null hypothesis to be nested within the alternative, the AB individuals from the two cross directions must be allowed to be different. Thus we arrive at the contrasts AA:ABf:ABr:BB for the alternative and forward:reverse for the null. Here, again, the linkage test has 2 d.f. In the analogous case with males only, since both directions give rise to equal parts hemizygous A and hemizygous B individuals, we need not split the individuals according to the direction of the cross, as a crossdirection effect cannot cause spurious linkage to the X chromosome. In this case, the test for linkage has 1 d.f. In the most complex case, of an intercross with both directions and both sexes, all four types of females must be allowed to be separate, whereas the males from the two directions may be pooled, and so the simplest comparison includes the contrasts AA:ABf:ABr:BB:AY: BY, with the null hypothesis using the contrasts female forward:female reverse:male. Thus, the linkage test has 3 d.f. The actual statistical analysis is no different from what is done in standard interval mapping for autosomes. In the presence of a QTL, we first calculate, for individual i, the probability, p iq, that the individual falls into QTL
4 2154 K. W. Broman et al. genotype group q, for each of the genotype groups under consideration, given the available marker data. Calculation of these QTL genotype probabilities at the putative QTL, given the available multipoint marker genotype data, is most efficiently done via a hidden Markov model (Lander and Green 1987). This can be constructed to allow for the presence of genotyping errors (Lincoln and Lander 1992). As each backcross or intercross individual has a single X chromosome that was subject to recombination (see Figures 1 and 2), the calculations here are identical to those for an autosome in a backcross, and so nothing new is needed. We then assume that, given an individual s genotype, q, its phenotype follows a normal distribution with mean m q and common SD s. Maximum-likelihood estimates (MLEs) of the parameters m q and s are obtained via the EM algorithm (Dempster et al. 1977). The maximized likelihood is then Q P p i q iqfðy i j ˆm q ; ŝþ, where f is the normal density. Under the null hypothesis, individuals are grouped as described in Table 1, with individual i Q assigned to, say, group j i, and the likelihood is fðy i i j m ji ; sþ, for which MLEs may be obtained in closed form. A LOD score is calculated as the log (base 10) of the ratio of the maximized likelihoods under the alternative and the null hypotheses. Chromosome-specific thresholds: The usual approach for establishing statistical significance in a QTL genome scan is to calculate a single genomewide LOD threshold. At the 5% significance level, one calculates the 95th percentile of the distribution of the genomewide maximum LOD score, under the global null hypothesis that there are no QTL anywhere. This is best done via a permutation test (Churchill and Doerge 1994). The number of degrees of freedom of the linkage test on the X chromosome can be different from that on the autosomes (see Table 1). In the case that the degrees of freedom for the X chromosome are larger than those for autosomes, the null distribution of the LOD score on the X chromosome will be stochastically larger than that for the LOD score on an autosome. Additionally, in intercrosses, only one X chromosome is potentially recombinant (compared to both, for autosomes), leading to an effectively smaller genetic length relative to autosomes. Thus, if we apply a constant threshold, our tests on the X chromosome may be too liberal. One could assign chromosome-specific LOD thresholds, allowing for a chromosome-specific false positive rate of a i for chromosome i. We require, however, that the a i are chosen to maintain the desired genomewide significance level, a. Under the null hypothesis of no QTL and with the assumption of independent assortment of chromosomes, the LOD scores on separate chromosomes are independent, and so we must choose the a i so that a ¼ 1 Y i ð1 a i Þ: ð1þ Any choice of the a i satisfying Equation 1 will provide a genomewide false positive rate that is maintained at the desired level. For example, one could choose a 1 ¼ a and a i ¼ 0 for i 6¼ 1. A key issue, in choosing the a i, concerns the power to detect a QTL. In the preceding example, one would have high power to detect a QTL on chromosome 1, but no power to detect a QTL on any other chromosome. The usual approach, with a constant LOD threshold across the genome, provides high power to detect a QTL irrespective of its location: in the case of high and uniform marker density and the presence of a single autosomal QTL, the power to detect the QTL would be the same no matter where it resides. In tackling the problem of how to assign a LOD threshold for the X chromosome, then, one might choose to ensure that the power to detect a QTL on the X chromosome is the same as that for the autosomes. This is tricky, however, in that the nature of the effect of an X chromosome QTL can be markedly different from one on the autosomes, as the individuals have a different set of possible genotypes. Thus, we have chosen to take a different approach. It appears reasonable to choose the a i proportional to the genetic lengths of the chromosomes. Let L i denote the genetic length of chromosome i. We could choose a i ¼ kl i for some k, subject to the constraint in Equation 1. (It is interesting to note that the analytical results of Lander and Botstein 1989, for the dense map case, give a i ¼ a 1 bl i for a 6¼ 0.) The solutions, unfortunately, cannot be written in closed form, but one may take a i ¼ 1 ð1 aþ Li=L, where L ¼ P L i i, and get results that are indistinguishable in practice. And so the latter is the approach that we recommend. We recommend a constant LOD threshold for the autosomes and a separate threshold for the X chromosome. Taking L A to be the sum of the genetic lengths of the autosomes and L X to be the length of the X chromosome, we use a A ¼ 1 ð1 aþ LA=L and a X ¼ 1 ð1 aþ LX=L. In particular, in a permutation test to determine LOD thresholds, one would calculate, for permutation replicate j, LOD* ja as the maximum LOD score across all autosomes and LOD* jx as the maximum LOD score across the X chromosome. The LOD threshold for the autosomes would be the 1 a A quantile of the LOD* ja, and the LOD threshold for the X chromosome would be the 1 a X quantile of the LOD* jx. Typical values for these thresholds are shown in the application section (see Table 2). Genome-scan-adjusted P-values can be estimated from the permutation results as follows. For a putative QTL on an autosome, one would first calculate the proportion, call it p, of the LOD* ja that were greater or equal to the observed LOD score. The adjusted P-value would then be 1 ð1 pþ L=LA. Equations for a locus on the X chromosome are analogous, replacing the A s with X s.
5 TABLE 2 Genomewide 5% LOD thresholds Both sexes Males Females Common a Autosomes b X chromosome b a Thresholds derived in the standard way, constant for all chromosomes. b Thresholds derived allowing separate values for the autosomes and the X chromosome. X Chromosome in QTL Mapping 2155 Figure 3. The attained chromosome-specific false positive rates for autosomes with the use of a constant 5% genomewide LOD threshold (based on computer simulations), for a genome modeled after the mouse and having markers spaced at 10 cm (circles) or 1 cm (x s). The curve corresponds to the chromosome-specific false positive rates from the formula 1 0:95 Li=L. It is of some interest to compare, for autosomes, the chromosome-specific a i defined through a i ¼ 1 ð1 aþ Li=L to those that are attained with an assumption of a constant genomewide LOD threshold. We would be concerned if they were not similar. To investigate this, we simulated data on a backcross of 200 individuals, with a genome modeled after the mouse, markers spaced at either 10 or 1 cm, and phenotypes following a normal distribution and independent of marker genotypes (thus under the global null hypothesis of no QTL). We used 5,000,000 simulation replicates. For each replicate, we calculated the maximum LOD score for each autosome, say LOD ij for chromosome i in simulation replicate j. The genomewide maximum LOD for each replicate was calculated as M j ¼ max j LOD ij. The genomewide LOD threshold, call it T, was taken to be the 95th percentile of the M j. The chromosome-specific false positive rates were then taken as a i ¼ Prop(j: LOD ij. T). These are shown in Figure 3, along with the curve following the chromosome-specific thresholds chosen according to our recommended procedure, 1 ð1 aþ Li=L. The chromosome-specific false positive rates obtained with the assumption of a constant genomewide LOD threshold are somewhat higher than the procedure we have described for the smaller chromosomes and are somewhat lower for the larger chromosomes, but differ by no more than Thus, we conclude that the proposed procedure does not deviate too far from the use of a constant LOD threshold, but allows us to account for the difference in degrees of freedom between the autosomes and the X chromosome. One final detail concerning the permutation procedure deserves emphasis. Sex and cross-direction differences in the phenotype must be preserved in the permutations. This could be accomplished by a stratified permutation test: permuting the phenotypes relative to the genotypes separately within the four strata (males and females in each cross direction). We use a slightly different strategy: sex and cross direction are viewed as additional phenotypes, and the rows in the phenotype matrix are shuffled relative to the genotype data. Thus the sex (and cross direction) attached to a particular phenotype is preserved. Autosomal genotypes are permuted across all individuals. For the X chromosome data, all individuals have one chromosome subject to recombination and one nonrecombinant chromosome, and we permute the data for the recombinant chromosome across all individuals. This approach will give results equivalent to the stratified permutation test, provided that the pattern of missing genotype data is similar in the different strata. APPLICATION As an illustration of our methods, we consider data on gut length from the cross of Owens et al. (2005). The aim of this study was to identify modifiers of Sox10 Dom,a model of Hirschsprung disease. Lines of Sox10 Dom mice congenic on the C57BL/6J background were crossed with C3HeB/FeJ mice to generate B6 N15 C3Fe.Sox10 Dom (F 1 ) progeny. Intercrosses were performed by crossing male F 1 mice to wild-type B6C3Fe or C3FeB6 females as well as reciprocal intercrosses between female B6 N15 C3Fe.Sox10 Dom (F 1 ) mice and wild-type B6C3Fe or C3FeB6 males. From these crosses, 2210 F 2 mice were collected, but only the 1068 mice carrying the Sox10 Dom mutation were considered for the QTL analysis. To avoid the technicalities associated with the selection of individuals carrying the mutation we omitted chromosome 15, which carries this mutation, from our analysis. The estimated genetic length of the X chromosome was 78 cm; the total length of the autosomes (excluding chromosome 15) was 1250 cm. Individuals were typed at a set of 117 markers on the remaining chromosomes, including 6 markers on the X chromosome. A selective genotyping strategy was used, with the most extreme 30% of individuals, phenotypically,
6 2156 K. W. Broman et al. Figure 4. LOD curves for analysis of gut length with both sexes (a), males only (b), and females only (c). Dashed horizontal lines are plotted at the estimated genomewide 95% LOD thresholds, allowed to be separate for the autosomes and the X chromosome. genotyped at essentially all markers, and others genotyped at markers in regions showing an effect. Selective genotyping was based on the aganglionosis traits considered in Owens et al. (2005), but not considered here. Quantification of aganglionosis relies on measurement of total gut length from gastric sphincter to anus. The gut length phenotype was the focus of our analysis in this study. The F 2 mice comprised 563 males and 505 females. The gut length phenotype showed a clear sex difference, with average (SE) gut lengths of 16.6 (0.090) and 16.2 (0.095) cm in males and females, respectively. For the QTL analysis, sex was included as an additive covariate, and the two sexes were also considered separately. Genomewide 5% LOD thresholds, estimated from a permutation test with 100,000 replicates, are displayed in Table 2. When a common LOD threshold is used for the entire genome, the threshold is 3.45 (approximately), whether analysis concerns both sexes, males only, or females only. If a separate threshold is allowed for the X chromosome, the threshold for the autosomes changed very little. For the analysis of both sexes combined, the threshold for the X chromosome increased to 3.70, due to the fact that the linkage test for the X chromosome has 3 d.f. while for the autosomes there are 2 d.f. For the analysis of males alone, the X chromosome-specific threshold decreased to 2.67; in this case, the linkage test on the X chromosome has 1 d.f., while that for the autosomes had 2 d.f. For the analysis of the females alone, the X chromosomespecific threshold decreased to 3.17, in spite of the fact that the tests for each of the X chromosomes and the autosomes has 2 d.f. The decrease in the LOD threshold in this case may be due to the fact that each individual has just one recombinant X chromosome, but two recombinant autosomes, and so the effective number of independent statistical tests is smaller for the X chromosome than for an autosome of equivalent length. LOD curves for the gut length trait are displayed in Figure 4, with further details on the identified QTL shown in Table 3. In the analysis of both sexes, a strong QTL is seen on chromosome 5. A QTL near this position was previously observed to affect the extent of aganglionosis (Owens et al. 2005). The phenotype averages for each genotype group (Figure 5a), estimated by the multiple-imputation approach of Sen and Churchill (2001), indicate that the C3H allele is dominant and results in an increase in gut length. There is significant linkage to chromosome 18 in females but not in males. As shown in Figure 5b, the
7 X Chromosome in QTL Mapping 2157 TABLE 3 Estimated QTL positions, LOD scores, and genome-scan-adjusted P-values Both sexes Males Females Chr Pos (cm) LOD P a P b LOD P a P b LOD P a P b ,0.001, X Chr, chromosome; Pos, position. a Genome-scan-adjusted P-values calculated in the standard way, with a constant threshold for all chromosomes. b Genome-scan-adjusted P-values allowing separate thresholds for the autosomes and the X chromosome. C3H allele again results in an increase in gut length and appears to be recessive. Little effect is seen in males, but a test for a sex difference in the effect of this locus was not significant: the lack of evidence for linkage in males is not sufficient to conclude the lack of an effect. Potential linkage is seen on the X chromosome, and the support for an X chromosome QTL depends critically on whether an X chromosome-specific LOD threshold is used. When a constant threshold is used across the genome, the genome-scan-adjusted P-value for the putative X chromosome QTL is 6.5%, while if a separate threshold is used for the X chromosome, the P-value increases to 11%. [The latter P-value was obtained as follows: 0.68% of the permutations gave a maximum LOD score on the X chromosome equal to or greater than the observed LOD score of 2.32, and L/L X 17, and so the adjusted P-value was 1 ( ) ] Note that if the sex effect were not taken into account in the null hypothesis here, the LOD scores on the X chromosome would increase by 5.67, uniformly, resulting in greatly inflated evidence for X linkage. In the separate analyses of the two sexes, little evidence is seen for linkage to the X chromosome, but this may be due to the loss of power due to the halving of sample size. DISCUSSION Figure 5. Plot of average gut length (62 SE) as a function of sex and genotype at the inferred QTL on chromosomes 5, 18, and X. B and C denote the C57BL/6J and C3HeB/ FeJ alleles, respectively. Estimates were derived by multiple imputation. We have described an approach for the treatment of the X chromosome in QTL mapping, focusing on the development of a procedure for routine use that will avoid spurious linkage in the presence of sex or crossdirection differences in the phenotype. We consider all possible cases for a backcross or intercross, identifying appropriate contrasts (sets of genotype groups that are to be compared) and the necessary covariates that must be included in the null hypothesis to avoid spurious linkage. While we have focused on the use of standard interval mapping (Lander and Botstein 1989), our approach may be applied with other methods for dealing with missing genotype information, such as Haley Knott regression (Haley and Knott 1992) and multiple imputation (Sen and Churchill 2001). The extension to nonnormal models for the phenotype distribution given the QTL genotype is also straightforward. As the number of degrees of freedom for the linkage test on the X chromosome differs from that for the autosomes, it is important to apply separate LOD thresholds
8 2158 K. W. Broman et al. for evaluating significance of the X chromosome vs. the autosomes. We have described an approach for deriving such chromosome-specific thresholds in the context of a permutation test. We applied our methods to data on gut length in a large mouse intercross and identified QTL on chromosomes 5 and 18, plus a suggestion of a QTL on the X chromosome. Overly strong evidence for the X chromosome QTL would have been obtained had the sex difference in the phenotype not been taken into account. These loci may be relevant to gastroenterologists interested in short gut syndrome (Seashore et al. 1987; Kern et al. 1990), although it must be acknowledged that the QTL may affect gut length through body size; we made no attempt to account for body size in our analysis. It is important to point out that the precise estimation of the X chromosome-specific LOD threshold will require considerably more permutation replicates. A first-order Taylor expansion of the adjusted P-value, 1 ð1 pþ L=LX, indicates that its SE is, roughly, a factor L/L X larger than the SE of the unadjusted P-value. (Recall that L X is the genetic length of the X chromosome and L is the total genome length.) Further, we find that one must use roughly L/L X times more permutation replicates to get the same precision for an adjusted P-value for the X chromosome as one would typically need if a constant LOD threshold were used across the genome. (This result was confirmed via computer simulations, not shown.) For the application to the data of Owens et al. (2005), if one were to use 1000 permutation replicates for the autosomes, 17,000 replicates would be required for the X chromosome, since L/L X 17. As each replicate of the X chromosome requires a scan over L X /L of the genome, the total computational effort is about double the usual amount. If one sought a separate threshold for each chromosome, one would want, for chromosome i, L/L i times the usual number of permutation replicates, and, in the case of 20 chromosomes, the overall effort would be 20 times the usual amount. In our development we have neglected imprinting, maternal effects, the Y chromosome, and mitochondrial or cytoplasmic effects. Should these be deemed important for the phenotype of interest, modifications to our methods are necessary. These can be arrived at by careful consideration of Figures 1 and 2 and judicious use of the different cross options. We have focused on single-dimensional, single-qtl genome scans. Analysis of two-dimensional, two-qtl genome scans (Haley and Knott 1992; Sen and Churchill 2001) must also be modified for the case of the X chromosome. There are two key issues. First, in cases where the two QTL are located on the X chromosome,itmustbeacknowledgedthatindividualscanhave only two of the possible genotypes at each locus. For example, in an intercross in one direction but with both males and females, one considers the 4 genotype groups AA,AB,AY,andBY,ateachlocus,butofthe16two-locus genotypes, only 8 are observable. Care must be taken to avoid overparameterization. Second, in the case that one QTL is on the X chromosome and one is on an autosome, as the analysis of the X chromosome may require sex as a covariate in the null hypothesis, it must also be used for this portion of the two-dimensional, two-qtl scan. The X chromosome, while comprising only 5% of the genome, requires a doubling of effort for QTL mapping. The appropriate treatment of the X chromosome is not conceptually difficult, but it does require some rather tedious bookkeeping. We are not aware of QTL mapping software that treats the X chromosome appropriately, except for our own software, R/qtl (Broman et al. 2003), an add-on package to the R statistical software (Ihaka and Gentleman 1996). The methods we have described are implemented in R/qtl. The authors thank an anonymous reviewer for comments to improve the manuscript. This work was supported in part by National Institutes of Health grants GM (to K.W.B.), GM (to G.A.C.), NS43556 (to MS2), and DK60047 (to MS2) and by a National Science Foundation Graduate Research Fellowship (to A.M.). LITERATURE CITED Ahmadiyeh, N., G. A. Churchill, K. Shimomura, L. C. Solberg, J. S. Takahashi et al., 2003 X-linked and lineage-dependent inheritance of coping responses to stress. Mamm. Genome 14: Broman, K. W., H. Wu, Ś.Sen and G. A. Churchill, 2003 R/qtl: QTL mapping in experimental crosses. Bioinformatics 19: Churchill, G. A., and R. W. Doerge, 1994 Empirical threshold values for quantitative trait mapping. Genetics 138: Dempster, A. P., N. M. Laird and D. B. Rubin, 1977 Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. B 39: Haley, C. S., and S. A. Knott, 1992 A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity 69: Ihaka, R., and R. Gentleman, 1996 R: a language for data analysis and graphics. J. Comp. Graph. Stat. 5: Kern, I. B., A. Leece and T. Bohane, 1990 Congenital short gut, malrotation, and dysmotility of the small bowel. J. Pediatr. Gastroenterol. Nutr. 11: Lander, E. S., and D. Botstein, 1989 Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121: Lander, E. S., and P. Green, 1987 Construction of multilocus genetic linkage maps in humans. Proc. Natl. Acad. Sci. USA 84: Lincoln, S. E., and E. S. Lander, 1992 Systematic detection of errors in genetic linkage data. Genomics 14: Owens,S.E.,K.W.Broman,T.Wiltshire,J.B.Elmore,K.M.Bradley et al., 2005 Genome-wide linkage identifies novel modifier loci of aganglionosis in the Sox10 Dom model of Hirschsprung disease. Hum. Mol. Genet. 14: Rance, K. A., W. G. Hill and P. D. Keightley, 1997 Mapping quantitative trait loci for body weight on the X chromosome in mice. I. Analysis of a reciprocal F 2 population. Genet. Res. 70: Seashore, J. H., F. S. Collins, R. I. Markowitz and M. R. Seashore, 1987 Familial apple peel jejunal atresia: surgical, genetic, and radiographic aspects. Pediatrics 80: Sen, Ś., and G. A. Churchill, 2001 A statistical framework for quantitative trait mapping. Genetics 159: Communicating editor: M. S. McPeek
Complex Trait Genetics in Animal Models. Will Valdar Oxford University
 Complex Trait Genetics in Animal Models Will Valdar Oxford University Mapping Genes for Quantitative Traits in Outbred Mice Will Valdar Oxford University What s so great about mice? Share ~99% of genes
Complex Trait Genetics in Animal Models Will Valdar Oxford University Mapping Genes for Quantitative Traits in Outbred Mice Will Valdar Oxford University What s so great about mice? Share ~99% of genes
Genetics of extreme body size evolution in mice from Gough Island
 Genetics of extreme body size evolution in mice from Gough Island Karl Broman Biostatistics & Medical Informatics, UW Madison kbroman.org github.com/kbroman @kwbroman bit.ly/apstats2017 This is a collaboration
Genetics of extreme body size evolution in mice from Gough Island Karl Broman Biostatistics & Medical Informatics, UW Madison kbroman.org github.com/kbroman @kwbroman bit.ly/apstats2017 This is a collaboration
The Efficiency of Mapping of Quantitative Trait Loci using Cofactor Analysis
 The Efficiency of Mapping of Quantitative Trait Loci using Cofactor G. Sahana 1, D.J. de Koning 2, B. Guldbrandtsen 1, P. Sorensen 1 and M.S. Lund 1 1 Danish Institute of Agricultural Sciences, Department
The Efficiency of Mapping of Quantitative Trait Loci using Cofactor G. Sahana 1, D.J. de Koning 2, B. Guldbrandtsen 1, P. Sorensen 1 and M.S. Lund 1 1 Danish Institute of Agricultural Sciences, Department
Statistical Tests for X Chromosome Association Study. with Simulations. Jian Wang July 10, 2012
 Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
Received 5 April 2018; revised 31 July 2018; accepted 14 August 2018; published online 24 November 2018
 Journal of Genetics, Vol. 97, No. 5, December 2018, pp. 1413 1420 https://doi.org/10.1007/s12041-018-1040-7 Indian Academy of Sciences RESEARCH ARTICLE Quantitative trait loci that determine plasma insulin
Journal of Genetics, Vol. 97, No. 5, December 2018, pp. 1413 1420 https://doi.org/10.1007/s12041-018-1040-7 Indian Academy of Sciences RESEARCH ARTICLE Quantitative trait loci that determine plasma insulin
Lecture 6: Linkage analysis in medical genetics
 Lecture 6: Linkage analysis in medical genetics Magnus Dehli Vigeland NORBIS course, 8 th 12 th of January 2018, Oslo Approaches to genetic mapping of disease Multifactorial disease Monogenic disease Syke
Lecture 6: Linkage analysis in medical genetics Magnus Dehli Vigeland NORBIS course, 8 th 12 th of January 2018, Oslo Approaches to genetic mapping of disease Multifactorial disease Monogenic disease Syke
Name: PS#: Biol 3301 Midterm 1 Spring 2012
 Name: PS#: Biol 3301 Midterm 1 Spring 2012 Multiple Choice. Circle the single best answer. (4 pts each) 1. Which of the following changes in the DNA sequence of a gene will produce a new allele? a) base
Name: PS#: Biol 3301 Midterm 1 Spring 2012 Multiple Choice. Circle the single best answer. (4 pts each) 1. Which of the following changes in the DNA sequence of a gene will produce a new allele? a) base
GENETIC LINKAGE ANALYSIS
 Atlas of Genetics and Cytogenetics in Oncology and Haematology GENETIC LINKAGE ANALYSIS * I- Recombination fraction II- Definition of the "lod score" of a family III- Test for linkage IV- Estimation of
Atlas of Genetics and Cytogenetics in Oncology and Haematology GENETIC LINKAGE ANALYSIS * I- Recombination fraction II- Definition of the "lod score" of a family III- Test for linkage IV- Estimation of
Non-parametric methods for linkage analysis
 BIOSTT516 Statistical Methods in Genetic Epidemiology utumn 005 Non-parametric methods for linkage analysis To this point, we have discussed model-based linkage analyses. These require one to specify a
BIOSTT516 Statistical Methods in Genetic Epidemiology utumn 005 Non-parametric methods for linkage analysis To this point, we have discussed model-based linkage analyses. These require one to specify a
A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single
 8.3 A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single chromosome can alter their pattern of inheritance from those
8.3 A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single chromosome can alter their pattern of inheritance from those
Laboratory. Mendelian Genetics
 Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Laboratory 9 Mendelian Genetics Biology 171L FA17 Lab 9: Mendelian Genetics Student Learning Outcomes 1. Predict the phenotypic and genotypic ratios of a monohybrid cross. 2. Determine whether a gene is
Cancer develops after somatic mutations overcome the multiple
 Genetic variation in cancer predisposition: Mutational decay of a robust genetic control network Steven A. Frank* Department of Ecology and Evolutionary Biology, University of California, Irvine, CA 92697-2525
Genetic variation in cancer predisposition: Mutational decay of a robust genetic control network Steven A. Frank* Department of Ecology and Evolutionary Biology, University of California, Irvine, CA 92697-2525
Genetic resistance to diet-induced obesity in chromosome substitution strains of mice
 Mamm Genome (2010) 21:115 129 DOI 10.1007/s00335-010-9247-9 ORIGINAL CONTRIBUTIONS Genetic resistance to diet-induced obesity in chromosome substitution strains of mice Lindsay C. Burrage Annie E. Baskin-Hill
Mamm Genome (2010) 21:115 129 DOI 10.1007/s00335-010-9247-9 ORIGINAL CONTRIBUTIONS Genetic resistance to diet-induced obesity in chromosome substitution strains of mice Lindsay C. Burrage Annie E. Baskin-Hill
Mendelism: the Basic principles of Inheritance
 Chapter 3. Mendelism: the Basic principles of Inheritance 1. Mendel s Study of Heredity 2. Applications of Mendel s Principles 3. Formulating and Testing Genetic Hypothesis 4. Mendelian Principles in Human
Chapter 3. Mendelism: the Basic principles of Inheritance 1. Mendel s Study of Heredity 2. Applications of Mendel s Principles 3. Formulating and Testing Genetic Hypothesis 4. Mendelian Principles in Human
MBG* Animal Breeding Methods Fall Final Exam
 MBG*4030 - Animal Breeding Methods Fall 2007 - Final Exam 1 Problem Questions Mick Dundee used his financial resources to purchase the Now That s A Croc crocodile farm that had been operating for a number
MBG*4030 - Animal Breeding Methods Fall 2007 - Final Exam 1 Problem Questions Mick Dundee used his financial resources to purchase the Now That s A Croc crocodile farm that had been operating for a number
Ch 8 Practice Questions
 Ch 8 Practice Questions Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What fraction of offspring of the cross Aa Aa is homozygous for the dominant allele?
Ch 8 Practice Questions Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What fraction of offspring of the cross Aa Aa is homozygous for the dominant allele?
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits. Harold Snieder
 Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
Allelic penetrance approach as a tool to model two-locus interaction in complex binary traits
 ORIGINAL ARTICLE (2007) 99, 173 184 & 2007 Nature ublishing Group All rights reserved 0018-067X/07 $30.00 www.nature.com/hdy Allelic penetrance approach as a tool to model two-locus interaction in complex
ORIGINAL ARTICLE (2007) 99, 173 184 & 2007 Nature ublishing Group All rights reserved 0018-067X/07 $30.00 www.nature.com/hdy Allelic penetrance approach as a tool to model two-locus interaction in complex
What we mean more precisely is that this gene controls the difference in seed form between the round and wrinkled strains that Mendel worked with
 9/23/05 Mendel Revisited In typical genetical parlance the hereditary factor that determines the round/wrinkled seed difference as referred to as the gene for round or wrinkled seeds What we mean more
9/23/05 Mendel Revisited In typical genetical parlance the hereditary factor that determines the round/wrinkled seed difference as referred to as the gene for round or wrinkled seeds What we mean more
Introduction of Genome wide Complex Trait Analysis (GCTA) Presenter: Yue Ming Chen Location: Stat Gen Workshop Date: 6/7/2013
 Introduction of Genome wide Complex Trait Analysis (GCTA) resenter: ue Ming Chen Location: Stat Gen Workshop Date: 6/7/013 Outline Brief review of quantitative genetics Overview of GCTA Ideas Main functions
Introduction of Genome wide Complex Trait Analysis (GCTA) resenter: ue Ming Chen Location: Stat Gen Workshop Date: 6/7/013 Outline Brief review of quantitative genetics Overview of GCTA Ideas Main functions
The Determination of the Genetic Order and Genetic Map for the Eye Color, Wing Size, and Bristle Morphology in Drosophila melanogaster
 Kudlac 1 Kaitie Kudlac March 24, 2015 Professor Ma Genetics 356 The Determination of the Genetic Order and Genetic Map for the Eye Color, Wing Size, and Bristle Morphology in Drosophila melanogaster Abstract:
Kudlac 1 Kaitie Kudlac March 24, 2015 Professor Ma Genetics 356 The Determination of the Genetic Order and Genetic Map for the Eye Color, Wing Size, and Bristle Morphology in Drosophila melanogaster Abstract:
An Introduction to Quantitative Genetics I. Heather A Lawson Advanced Genetics Spring2018
 An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
Title: A robustness study of parametric and non-parametric tests in Model-Based Multifactor Dimensionality Reduction for epistasis detection
 Author's response to reviews Title: A robustness study of parametric and non-parametric tests in Model-Based Multifactor Dimensionality Reduction for epistasis detection Authors: Jestinah M Mahachie John
Author's response to reviews Title: A robustness study of parametric and non-parametric tests in Model-Based Multifactor Dimensionality Reduction for epistasis detection Authors: Jestinah M Mahachie John
Lab 5: Testing Hypotheses about Patterns of Inheritance
 Lab 5: Testing Hypotheses about Patterns of Inheritance How do we talk about genetic information? Each cell in living organisms contains DNA. DNA is made of nucleotide subunits arranged in very long strands.
Lab 5: Testing Hypotheses about Patterns of Inheritance How do we talk about genetic information? Each cell in living organisms contains DNA. DNA is made of nucleotide subunits arranged in very long strands.
Complex Traits Activity INSTRUCTION MANUAL. ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik
 Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0
 The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
Characterization of quantitative trait loci for growth and meat quality in a cross between commercial breeds of swine
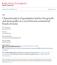 Animal Science Publications Animal Science 2004 Characterization of quantitative trait loci for growth and meat quality in a cross between commercial breeds of swine H. K. Thomsen Iowa State University
Animal Science Publications Animal Science 2004 Characterization of quantitative trait loci for growth and meat quality in a cross between commercial breeds of swine H. K. Thomsen Iowa State University
Analysis of single gene effects 1. Quantitative analysis of single gene effects. Gregory Carey, Barbara J. Bowers, Jeanne M.
 Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
Two-stage Methods to Implement and Analyze the Biomarker-guided Clinical Trail Designs in the Presence of Biomarker Misclassification
 RESEARCH HIGHLIGHT Two-stage Methods to Implement and Analyze the Biomarker-guided Clinical Trail Designs in the Presence of Biomarker Misclassification Yong Zang 1, Beibei Guo 2 1 Department of Mathematical
RESEARCH HIGHLIGHT Two-stage Methods to Implement and Analyze the Biomarker-guided Clinical Trail Designs in the Presence of Biomarker Misclassification Yong Zang 1, Beibei Guo 2 1 Department of Mathematical
Statistical power and significance testing in large-scale genetic studies
 STUDY DESIGNS Statistical power and significance testing in large-scale genetic studies Pak C. Sham 1 and Shaun M. Purcell 2,3 Abstract Significance testing was developed as an objective method for summarizing
STUDY DESIGNS Statistical power and significance testing in large-scale genetic studies Pak C. Sham 1 and Shaun M. Purcell 2,3 Abstract Significance testing was developed as an objective method for summarizing
Measuring Performance Of Physicians In The Diagnosis Of Endometriosis Using An Expectation-Maximization Algorithm
 Yale University EliScholar A Digital Platform for Scholarly Publishing at Yale Public Health Theses School of Public Health January 2014 Measuring Performance Of Physicians In The Diagnosis Of Endometriosis
Yale University EliScholar A Digital Platform for Scholarly Publishing at Yale Public Health Theses School of Public Health January 2014 Measuring Performance Of Physicians In The Diagnosis Of Endometriosis
Comments on Significance of candidate cancer genes as assessed by the CaMP score by Parmigiani et al.
 Comments on Significance of candidate cancer genes as assessed by the CaMP score by Parmigiani et al. Holger Höfling Gad Getz Robert Tibshirani June 26, 2007 1 Introduction Identifying genes that are involved
Comments on Significance of candidate cancer genes as assessed by the CaMP score by Parmigiani et al. Holger Höfling Gad Getz Robert Tibshirani June 26, 2007 1 Introduction Identifying genes that are involved
Mendelian Genetics: Patterns of Inheritance
 Mendelian Genetics: Patterns of Inheritance A Bit on Gregor Mendel Born to a poor farming family in what is now part of Czech Republic Attended Augustinian monastery (1843) Became an excellent teacher
Mendelian Genetics: Patterns of Inheritance A Bit on Gregor Mendel Born to a poor farming family in what is now part of Czech Republic Attended Augustinian monastery (1843) Became an excellent teacher
Inheritance of Aldehyde Oxidase in Drosophila melanogaster
 Inheritance of Aldehyde Oxidase in Drosophila melanogaster (adapted from Morgan, J. G. and V. Finnerty. 1991. Inheritance of aldehyde oxidase in Drosophilia melanogaster. Pages 33-47, in Tested studies
Inheritance of Aldehyde Oxidase in Drosophila melanogaster (adapted from Morgan, J. G. and V. Finnerty. 1991. Inheritance of aldehyde oxidase in Drosophilia melanogaster. Pages 33-47, in Tested studies
Mendelian Genetics using Fast Plants Report due Sept. 15/16. Readings: Mendelian genetics: Hartwell Chapter 2 pp , Chapter 5 pp
 1 Biology 423L Sept. 1/2 Mendelian Genetics using Fast Plants Report due Sept. 15/16. Readings: Mendelian genetics: Hartwell Chapter 2 pp. 13-27, Chapter 5 pp. 127-130. FASTPLANTS: Williams et al. (1986)
1 Biology 423L Sept. 1/2 Mendelian Genetics using Fast Plants Report due Sept. 15/16. Readings: Mendelian genetics: Hartwell Chapter 2 pp. 13-27, Chapter 5 pp. 127-130. FASTPLANTS: Williams et al. (1986)
The laws of Heredity. Allele: is the copy (or a version) of the gene that control the same characteristics.
 The laws of Heredity 1. Definition: Heredity: The passing of traits from parents to their offspring by means of the genes from the parents. Gene: Part or portion of a chromosome that carries genetic information
The laws of Heredity 1. Definition: Heredity: The passing of traits from parents to their offspring by means of the genes from the parents. Gene: Part or portion of a chromosome that carries genetic information
additive genetic component [d] = rded
![additive genetic component [d] = rded additive genetic component [d] = rded](/thumbs/92/108220661.jpg) Heredity (1976), 36 (1), 31-40 EFFECT OF GENE DISPERSION ON ESTIMATES OF COMPONENTS OF GENERATION MEANS AND VARIANCES N. E. M. JAYASEKARA* and J. L. JINKS Department of Genetics, University of Birmingham,
Heredity (1976), 36 (1), 31-40 EFFECT OF GENE DISPERSION ON ESTIMATES OF COMPONENTS OF GENERATION MEANS AND VARIANCES N. E. M. JAYASEKARA* and J. L. JINKS Department of Genetics, University of Birmingham,
Performing. linkage analysis using MERLIN
 Performing linkage analysis using MERLIN David Duffy Queensland Institute of Medical Research Brisbane, Australia Overview MERLIN and associated programs Error checking Parametric linkage analysis Nonparametric
Performing linkage analysis using MERLIN David Duffy Queensland Institute of Medical Research Brisbane, Australia Overview MERLIN and associated programs Error checking Parametric linkage analysis Nonparametric
A. Incorrect! Cells contain the units of genetic they are not the unit of heredity.
 MCAT Biology Problem Drill PS07: Mendelian Genetics Question No. 1 of 10 Question 1. The smallest unit of heredity is. Question #01 (A) Cell (B) Gene (C) Chromosome (D) Allele Cells contain the units of
MCAT Biology Problem Drill PS07: Mendelian Genetics Question No. 1 of 10 Question 1. The smallest unit of heredity is. Question #01 (A) Cell (B) Gene (C) Chromosome (D) Allele Cells contain the units of
Table S1. Trait summary for Northern Leaf Blight resistance in the maize nested association mapping (NAM) population and individual NAM families
 SUPPLEMENTAL MATERIAL Supplemental Tables: Table S1. Trait summary for Northern Leaf Blight resistance in the maize nested association mapping (NAM) population and individual NAM families "#$%&' (')*'%$+,-'
SUPPLEMENTAL MATERIAL Supplemental Tables: Table S1. Trait summary for Northern Leaf Blight resistance in the maize nested association mapping (NAM) population and individual NAM families "#$%&' (')*'%$+,-'
Single SNP/Gene Analysis. Typical Results of GWAS Analysis (Single SNP Approach) Typical Results of GWAS Analysis (Single SNP Approach)
 High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
Chapter 2. Linkage Analysis. JenniferH.BarrettandM.DawnTeare. Abstract. 1. Introduction
 Chapter 2 Linkage Analysis JenniferH.BarrettandM.DawnTeare Abstract Linkage analysis is used to map genetic loci using observations on relatives. It can be applied to both major gene disorders (parametric
Chapter 2 Linkage Analysis JenniferH.BarrettandM.DawnTeare Abstract Linkage analysis is used to map genetic loci using observations on relatives. It can be applied to both major gene disorders (parametric
HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007
 MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
BST227 Introduction to Statistical Genetics. Lecture 4: Introduction to linkage and association analysis
 BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
EPISTATIC effects are statistically defined as interactions
 Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.106.066571 Genomewide Analysis of Epistatic Effects for Quantitative Traits in Barley Shizhong Xu 1 and Zhenyu Jia Department of
Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.106.066571 Genomewide Analysis of Epistatic Effects for Quantitative Traits in Barley Shizhong Xu 1 and Zhenyu Jia Department of
Allowing for Missing Parents in Genetic Studies of Case-Parent Triads
 Am. J. Hum. Genet. 64:1186 1193, 1999 Allowing for Missing Parents in Genetic Studies of Case-Parent Triads C. R. Weinberg National Institute of Environmental Health Sciences, Research Triangle Park, NC
Am. J. Hum. Genet. 64:1186 1193, 1999 Allowing for Missing Parents in Genetic Studies of Case-Parent Triads C. R. Weinberg National Institute of Environmental Health Sciences, Research Triangle Park, NC
For more information about how to cite these materials visit
 Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
A search for quantitative trait loci exhibiting imprinting effects on mouse mandible size and shape
 (2008) 101, 518 526 & 2008 Macmillan Publishers Limited All rights reserved 0018-067X/08 $32.00 ORIGINAL ARTICLE www.nature.com/hdy A search for quantitative trait loci exhibiting imprinting effects on
(2008) 101, 518 526 & 2008 Macmillan Publishers Limited All rights reserved 0018-067X/08 $32.00 ORIGINAL ARTICLE www.nature.com/hdy A search for quantitative trait loci exhibiting imprinting effects on
Unit 7 Section 2 and 3
 Unit 7 Section 2 and 3 Evidence 12: Do you think food preferences are passed down from Parents to children, or does the environment play a role? Explain your answer. One of the most important outcomes
Unit 7 Section 2 and 3 Evidence 12: Do you think food preferences are passed down from Parents to children, or does the environment play a role? Explain your answer. One of the most important outcomes
A Unified Sampling Approach for Multipoint Analysis of Qualitative and Quantitative Traits in Sib Pairs
 Am. J. Hum. Genet. 66:1631 1641, 000 A Unified Sampling Approach for Multipoint Analysis of Qualitative and Quantitative Traits in Sib Pairs Kung-Yee Liang, 1 Chiung-Yu Huang, 1 and Terri H. Beaty Departments
Am. J. Hum. Genet. 66:1631 1641, 000 A Unified Sampling Approach for Multipoint Analysis of Qualitative and Quantitative Traits in Sib Pairs Kung-Yee Liang, 1 Chiung-Yu Huang, 1 and Terri H. Beaty Departments
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
A review of statistical methods in the analysis of data arising from observer reliability studies (Part 11) *
 A review of statistical methods in the analysis of data arising from observer reliability studies (Part 11) * by J. RICHARD LANDIS** and GARY G. KOCH** 4 Methods proposed for nominal and ordinal data Many
A review of statistical methods in the analysis of data arising from observer reliability studies (Part 11) * by J. RICHARD LANDIS** and GARY G. KOCH** 4 Methods proposed for nominal and ordinal data Many
The passing of traits from parents to offspring. The scientific study of the inheritance
 Inheritance The passing of traits from parents to offspring Genetics The scientific study of the inheritance Gregor Mendel -Father of modern genetics -Used peas to successfully identify the laws of heredity
Inheritance The passing of traits from parents to offspring Genetics The scientific study of the inheritance Gregor Mendel -Father of modern genetics -Used peas to successfully identify the laws of heredity
Diallel Analysis and its Applications in Plant Breeding
 Diallel Analysis and its Applications in Plant Breeding Madhu Choudhary*, Kana Ram Kumawat and Ravi Kumawat Department of Plant Breeding and Genetics, S.K.N. Agriculture University, Jobner-303329, Jaipur
Diallel Analysis and its Applications in Plant Breeding Madhu Choudhary*, Kana Ram Kumawat and Ravi Kumawat Department of Plant Breeding and Genetics, S.K.N. Agriculture University, Jobner-303329, Jaipur
Heredity and Meiosis AP BIOLOGY. Heredity. Slide 1 / 142 Slide 2 / 142. Slide 4 / 142. Slide 3 / 142. Slide 6 / 142. Slide 5 / 142.
 Slide 1 / 142 Slide 2 / 142 AP BIOLOGY Heredity Slide 3 / 142 Slide 4 / 142 Heredity Unit Topics Click on the topic to go to that section Heredity & Meiosis Law of Independent Assortment What Mendel Didn't
Slide 1 / 142 Slide 2 / 142 AP BIOLOGY Heredity Slide 3 / 142 Slide 4 / 142 Heredity Unit Topics Click on the topic to go to that section Heredity & Meiosis Law of Independent Assortment What Mendel Didn't
H. Thomsen, H. K. Lee, M. F. Rothschild, M. Malek and J. C. M. Dekkers
 Characterization of quantitative trait loci for growth and meat quality in a cross between commercial breeds of swine H. Thomsen, H. K. Lee, M. F. Rothschild, M. Malek and J. C. M. Dekkers J Anim Sci 2004.
Characterization of quantitative trait loci for growth and meat quality in a cross between commercial breeds of swine H. Thomsen, H. K. Lee, M. F. Rothschild, M. Malek and J. C. M. Dekkers J Anim Sci 2004.
Selection at one locus with many alleles, fertility selection, and sexual selection
 Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
GENETICS - CLUTCH CH.2 MENDEL'S LAWS OF INHERITANCE.
 !! www.clutchprep.com CONCEPT: MENDELS EXPERIMENTS AND LAWS Mendel s Experiments Gregor Mendel was an Austrian monk who studied Genetics using pea plants Mendel used pure lines meaning that all offspring
!! www.clutchprep.com CONCEPT: MENDELS EXPERIMENTS AND LAWS Mendel s Experiments Gregor Mendel was an Austrian monk who studied Genetics using pea plants Mendel used pure lines meaning that all offspring
MULTIFACTORIAL DISEASES. MG L-10 July 7 th 2014
 MULTIFACTORIAL DISEASES MG L-10 July 7 th 2014 Genetic Diseases Unifactorial Chromosomal Multifactorial AD Numerical AR Structural X-linked Microdeletions Mitochondrial Spectrum of Alterations in DNA Sequence
MULTIFACTORIAL DISEASES MG L-10 July 7 th 2014 Genetic Diseases Unifactorial Chromosomal Multifactorial AD Numerical AR Structural X-linked Microdeletions Mitochondrial Spectrum of Alterations in DNA Sequence
IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE.
 IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE By Siddharth Pratap Thesis Submitted to the Faculty of the Graduate School
IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE By Siddharth Pratap Thesis Submitted to the Faculty of the Graduate School
Tutorial on Genome-Wide Association Studies
 Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
A Comparison of Sample Size and Power in Case-Only Association Studies of Gene-Environment Interaction
 American Journal of Epidemiology ª The Author 2010. Published by Oxford University Press on behalf of the Johns Hopkins Bloomberg School of Public Health. This is an Open Access article distributed under
American Journal of Epidemiology ª The Author 2010. Published by Oxford University Press on behalf of the Johns Hopkins Bloomberg School of Public Health. This is an Open Access article distributed under
Take a look at the three adult bears shown in these photographs:
 Take a look at the three adult bears shown in these photographs: Which of these adult bears do you think is most likely to be the parent of the bear cubs shown in the photograph on the right? How did you
Take a look at the three adult bears shown in these photographs: Which of these adult bears do you think is most likely to be the parent of the bear cubs shown in the photograph on the right? How did you
I. Multiple Alleles. Chapter 5. Summary points. What pattern of inheritance is demonstrated in the following cross?
 Chapter 5 Extensions and Modifications of Basic Principles I. Multiple Alleles The ABO blood group has multiple alleles codominance and complete dominance. In codominance, both alleles are expressed simultaneously.
Chapter 5 Extensions and Modifications of Basic Principles I. Multiple Alleles The ABO blood group has multiple alleles codominance and complete dominance. In codominance, both alleles are expressed simultaneously.
Lec 02: Estimation & Hypothesis Testing in Animal Ecology
 Lec 02: Estimation & Hypothesis Testing in Animal Ecology Parameter Estimation from Samples Samples We typically observe systems incompletely, i.e., we sample according to a designed protocol. We then
Lec 02: Estimation & Hypothesis Testing in Animal Ecology Parameter Estimation from Samples Samples We typically observe systems incompletely, i.e., we sample according to a designed protocol. We then
MENDELIAN GENETICS. MENDEL RULE AND LAWS Please read and make sure you understand the following instructions and knowledge before you go on.
 MENDELIAN GENETICS Objectives Upon completion of this lab, students should: 1. Understand the principles and terms used in Mendelian genetics. 2. Know how to complete a Punnett square to estimate phenotypic
MENDELIAN GENETICS Objectives Upon completion of this lab, students should: 1. Understand the principles and terms used in Mendelian genetics. 2. Know how to complete a Punnett square to estimate phenotypic
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
Genomewide Linkage of Forced Mid-Expiratory Flow in Chronic Obstructive Pulmonary Disease
 ONLINE DATA SUPPLEMENT Genomewide Linkage of Forced Mid-Expiratory Flow in Chronic Obstructive Pulmonary Disease Dawn L. DeMeo, M.D., M.P.H.,Juan C. Celedón, M.D., Dr.P.H., Christoph Lange, John J. Reilly,
ONLINE DATA SUPPLEMENT Genomewide Linkage of Forced Mid-Expiratory Flow in Chronic Obstructive Pulmonary Disease Dawn L. DeMeo, M.D., M.P.H.,Juan C. Celedón, M.D., Dr.P.H., Christoph Lange, John J. Reilly,
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Chapter 6 Patterns of Inheritance
 Chapter 6 Patterns of Inheritance Genetics Explains and Predicts Inheritance Patterns Genetics can explain how these poodles look different. Section 10.1 Genetics Explains and Predicts Inheritance Patterns
Chapter 6 Patterns of Inheritance Genetics Explains and Predicts Inheritance Patterns Genetics can explain how these poodles look different. Section 10.1 Genetics Explains and Predicts Inheritance Patterns
Dan Koller, Ph.D. Medical and Molecular Genetics
 Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Pedigree Analysis Why do Pedigrees? Goals of Pedigree Analysis Basic Symbols More Symbols Y-Linked Inheritance
 Pedigree Analysis Why do Pedigrees? Punnett squares and chi-square tests work well for organisms that have large numbers of offspring and controlled mating, but humans are quite different: Small families.
Pedigree Analysis Why do Pedigrees? Punnett squares and chi-square tests work well for organisms that have large numbers of offspring and controlled mating, but humans are quite different: Small families.
PopGen4: Assortative mating
 opgen4: Assortative mating Introduction Although random mating is the most important system of mating in many natural populations, non-random mating can also be an important mating system in some populations.
opgen4: Assortative mating Introduction Although random mating is the most important system of mating in many natural populations, non-random mating can also be an important mating system in some populations.
Lecture 13: May 24, 2004
 Lecture 13: May 24, 2004 CH14: Mendel and the gene idea *particulate inheritance parents pass on discrete heritable units *gene- unit of inheritance which occupies a specific chromosomal location (locus)
Lecture 13: May 24, 2004 CH14: Mendel and the gene idea *particulate inheritance parents pass on discrete heritable units *gene- unit of inheritance which occupies a specific chromosomal location (locus)
25.1 QUANTITATIVE TRAITS
 CHAPTER OUTLINE 5.1 Quantitative Traits 5. Polygenic Inheritance 5.3 Heritability 5 QUANTITATIVE In this chapter, we will examine complex traits characteristics that are determined by several genes and
CHAPTER OUTLINE 5.1 Quantitative Traits 5. Polygenic Inheritance 5.3 Heritability 5 QUANTITATIVE In this chapter, we will examine complex traits characteristics that are determined by several genes and
MENDELIAN GENETICS. Law of Dominance: Law of Segregation: GAMETE FORMATION Parents and Possible Gametes: Gregory Mendel:
 MENDELIAN GENETICS Gregory Mendel: Heredity: Cross: X P1 Generation: F1 Generation: F2 Generation: Gametes: Dominant: Recessive: Genotype: Phenotype: Law of Dominance: Genes: Alleles: Law of Segregation:
MENDELIAN GENETICS Gregory Mendel: Heredity: Cross: X P1 Generation: F1 Generation: F2 Generation: Gametes: Dominant: Recessive: Genotype: Phenotype: Law of Dominance: Genes: Alleles: Law of Segregation:
Your DNA extractions! 10 kb
 Your DNA extractions! 10 kb Quantitative characters: polygenes and environment Most ecologically important quantitative traits (QTs) vary. Distributions are often unimodal and approximately normal. Offspring
Your DNA extractions! 10 kb Quantitative characters: polygenes and environment Most ecologically important quantitative traits (QTs) vary. Distributions are often unimodal and approximately normal. Offspring
Estimating genetic variation within families
 Estimating genetic variation within families Peter M. Visscher Queensland Institute of Medical Research Brisbane, Australia peter.visscher@qimr.edu.au 1 Overview Estimation of genetic parameters Variation
Estimating genetic variation within families Peter M. Visscher Queensland Institute of Medical Research Brisbane, Australia peter.visscher@qimr.edu.au 1 Overview Estimation of genetic parameters Variation
Does Mendel s work suggest that this is the only gene in the pea genome that can affect this particular trait?
 Mongenic Traits, Probability and Independent Assortment Genetical Jargon Demystified In typical genetical parlance the hereditary factor that determines the round/wrinkled seed difference as referred to
Mongenic Traits, Probability and Independent Assortment Genetical Jargon Demystified In typical genetical parlance the hereditary factor that determines the round/wrinkled seed difference as referred to
GENETICS - NOTES-
 GENETICS - NOTES- Warm Up Exercise Using your previous knowledge of genetics, determine what maternal genotype would most likely yield offspring with such characteristics. Use the genotype that you came
GENETICS - NOTES- Warm Up Exercise Using your previous knowledge of genetics, determine what maternal genotype would most likely yield offspring with such characteristics. Use the genotype that you came
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
Chapter 4 PEDIGREE ANALYSIS IN HUMAN GENETICS
 Chapter 4 PEDIGREE ANALYSIS IN HUMAN GENETICS Chapter Summary In order to study the transmission of human genetic traits to the next generation, a different method of operation had to be adopted. Instead
Chapter 4 PEDIGREE ANALYSIS IN HUMAN GENETICS Chapter Summary In order to study the transmission of human genetic traits to the next generation, a different method of operation had to be adopted. Instead
By Mir Mohammed Abbas II PCMB 'A' CHAPTER CONCEPT NOTES
 Chapter Notes- Genetics By Mir Mohammed Abbas II PCMB 'A' 1 CHAPTER CONCEPT NOTES Relationship between genes and chromosome of diploid organism and the terms used to describe them Know the terms Terms
Chapter Notes- Genetics By Mir Mohammed Abbas II PCMB 'A' 1 CHAPTER CONCEPT NOTES Relationship between genes and chromosome of diploid organism and the terms used to describe them Know the terms Terms
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
PRINCIPLE OF INHERITANCE AND
 29 CHAPTER 5 PRINCIPLE OF INHERITANCE AND VARIATION MULTIPLE-CHOICE QUESTIONS 1. All genes located on the same chromosome: a. Form different groups depending upon their relative distance b. Form one linkage
29 CHAPTER 5 PRINCIPLE OF INHERITANCE AND VARIATION MULTIPLE-CHOICE QUESTIONS 1. All genes located on the same chromosome: a. Form different groups depending upon their relative distance b. Form one linkage
WHAT S IN THIS LECTURE?
 What is meant by the term monogenic? WHAT S IN THIS LECTURE? WHAT S MENDEL S PRINCIPLE OF SEGREGATION? What s probability got to do with this? WHAT S MENDEL S PRINCIPLE OF INDEPENDENT ASSORTMENT? 1 FROM
What is meant by the term monogenic? WHAT S IN THIS LECTURE? WHAT S MENDEL S PRINCIPLE OF SEGREGATION? What s probability got to do with this? WHAT S MENDEL S PRINCIPLE OF INDEPENDENT ASSORTMENT? 1 FROM
Human Molecular Genetics Prof. S. Ganesh Department of Biological Sciences and Bioengineering Indian Institute of Technology, Kanpur
 Human Molecular Genetics Prof. S. Ganesh Department of Biological Sciences and Bioengineering Indian Institute of Technology, Kanpur Module - 02 Lecture - 06 Let us test your understanding of Pedigree
Human Molecular Genetics Prof. S. Ganesh Department of Biological Sciences and Bioengineering Indian Institute of Technology, Kanpur Module - 02 Lecture - 06 Let us test your understanding of Pedigree
Class XII Chapter 5 Principles of Inheritance and Variation Biology
 Question 1: Mention the advantages of selecting pea plant for experiment by Mendel. Mendel selected pea plants to carry out his study on the inheritance of characters from parents to offspring. He selected
Question 1: Mention the advantages of selecting pea plant for experiment by Mendel. Mendel selected pea plants to carry out his study on the inheritance of characters from parents to offspring. He selected
Non-Mendelian inheritance
 Non-Mendelian inheritance Focus on Human Disorders Peter K. Rogan, Ph.D. Laboratory of Human Molecular Genetics Children s Mercy Hospital Schools of Medicine & Computer Science and Engineering University
Non-Mendelian inheritance Focus on Human Disorders Peter K. Rogan, Ph.D. Laboratory of Human Molecular Genetics Children s Mercy Hospital Schools of Medicine & Computer Science and Engineering University
B-4.7 Summarize the chromosome theory of inheritance and relate that theory to Gregor Mendel s principles of genetics
 B-4.7 Summarize the chromosome theory of inheritance and relate that theory to Gregor Mendel s principles of genetics The Chromosome theory of inheritance is a basic principle in biology that states genes
B-4.7 Summarize the chromosome theory of inheritance and relate that theory to Gregor Mendel s principles of genetics The Chromosome theory of inheritance is a basic principle in biology that states genes
Adjusting for mode of administration effect in surveys using mailed questionnaire and telephone interview data
 Adjusting for mode of administration effect in surveys using mailed questionnaire and telephone interview data Karl Bang Christensen National Institute of Occupational Health, Denmark Helene Feveille National
Adjusting for mode of administration effect in surveys using mailed questionnaire and telephone interview data Karl Bang Christensen National Institute of Occupational Health, Denmark Helene Feveille National
Imaging Genetics: Heritability, Linkage & Association
 Imaging Genetics: Heritability, Linkage & Association David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July 17, 2011 Memory Activation & APOE ε4 Risk
Imaging Genetics: Heritability, Linkage & Association David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July 17, 2011 Memory Activation & APOE ε4 Risk
Lecture 1 Mendelian Inheritance
 Genes Mendelian Inheritance Lecture 1 Mendelian Inheritance Jurg Ott Gregor Mendel, monk in a monastery in Brünn (now Brno in Czech Republic): Breeding experiments with the garden pea: Flower color and
Genes Mendelian Inheritance Lecture 1 Mendelian Inheritance Jurg Ott Gregor Mendel, monk in a monastery in Brünn (now Brno in Czech Republic): Breeding experiments with the garden pea: Flower color and
Replacing IBS with IBD: The MLS Method. Biostatistics 666 Lecture 15
 Replacing IBS with IBD: The MLS Method Biostatistics 666 Lecture 5 Previous Lecture Analysis of Affected Relative Pairs Test for Increased Sharing at Marker Expected Amount of IBS Sharing Previous Lecture:
Replacing IBS with IBD: The MLS Method Biostatistics 666 Lecture 5 Previous Lecture Analysis of Affected Relative Pairs Test for Increased Sharing at Marker Expected Amount of IBS Sharing Previous Lecture:
BIOLOGY - CLUTCH CH.15 - CHROMOSOMAL THEORY OF INHERITANCE
 !! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
!! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
The domesticated brain: genetics of brain mass and brain structure in an avian species
 The domesticated brain: genetics of brain mass and brain structure in an avian species Henriksen, R. 1, Johnsson, M. 1, Andersson, L. 2, Jensen, P. 1, Wright, D. 1 1 AVIAN Behavioural Genomics and Physiology
The domesticated brain: genetics of brain mass and brain structure in an avian species Henriksen, R. 1, Johnsson, M. 1, Andersson, L. 2, Jensen, P. 1, Wright, D. 1 1 AVIAN Behavioural Genomics and Physiology
1042SCG Genetics & Evolutionary Biology Semester Summary
 1042SCG Genetics & Evolutionary Biology Semester Summary Griffith University, Nathan Campus Semester 1, 2014 Topics include: - Mendelian Genetics - Eukaryotic & Prokaryotic Genes - Sex Chromosomes - Variations
1042SCG Genetics & Evolutionary Biology Semester Summary Griffith University, Nathan Campus Semester 1, 2014 Topics include: - Mendelian Genetics - Eukaryotic & Prokaryotic Genes - Sex Chromosomes - Variations
Principles of Inheritance and Variation
 Principles of Inheritance and Variation Question 1: Mention the advantages of selecting pea plant for experiment by Mendel. Answer Mendel selected pea plants to carry out his study on the inheritance of
Principles of Inheritance and Variation Question 1: Mention the advantages of selecting pea plant for experiment by Mendel. Answer Mendel selected pea plants to carry out his study on the inheritance of
Chapter 1: Explaining Behavior
 Chapter 1: Explaining Behavior GOAL OF SCIENCE is to generate explanations for various puzzling natural phenomenon. - Generate general laws of behavior (psychology) RESEARCH: principle method for acquiring
Chapter 1: Explaining Behavior GOAL OF SCIENCE is to generate explanations for various puzzling natural phenomenon. - Generate general laws of behavior (psychology) RESEARCH: principle method for acquiring
Systems of Mating: Systems of Mating:
 8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
