A PER/TIM/DBT Interval Timer for Drosophila s Circadian Clock
|
|
|
- Emil McCoy
- 6 years ago
- Views:
Transcription
1 A PER/TIM/DBT Interval Timer for Drosophila s Circadian Clock L. SAEZ,* P. MEYER, AND M.W. YOUNG* *Laboratory of Genetics, The Rockefeller University, New York, New York 10021; Department of Microbiology, College of Physician & Surgeons, Columbia University, New York, New York Circadian rhythms in Drosophila are supported by a negative feedback loop, in which PERIOD (PER) and Timeless (TIM) shut down their own transcription as they translocate once a day from the cytoplasm of clock-containing cells to the nucleus. Period length is partially determined by an interval of cytoplasmic retention of the TIM and PER proteins. To study this process, we examined PER/TIM/Doubletime (DBT) physical interactions and nuclear translocation by imaging individual cultured Drosophila cells. Using live cell video microscopy and green fluorescent protein (GFP) tags, we observed dynamic patterns of stability and localization for DBT, PER, and TIM that resembled those previously found in vivo. These studies suggest that a cytoplasmic interval timer regulates nuclear translocation of these proteins. The cultured cell assay provides a potent system to study interactions among new and known genes involved in the generation of circadian behavior. INTRODUCTION That single genes could make a substantial contribution to circadian rhythms was first demonstrated by Konopka and Benzer (1971). Upon screening mutagenized Drosophila for changes in patterns of daily eclosion, they identified three mutations affecting a single X-chromosomal locus, period (per). One mutant, per 0, was arrhythmic, whereas per-short (per S ) and per-long (per L ) mutants were rhythmic with 19-hour and 29-hour periods, respectively. Locomotor activity was also affected, and each per mutation produced equivalent effects on eclosion and locomotor activity rhythms. More than a decade passed before the molecular identity of period was established (Bargiello and Young 1984; Reddy et al. 1984; Jackson et al. 1986; Citri et al. 1987). Yet, the primary sequence of the encoded protein provided little information about its function in circadian rhythm generation. In further work, the spatial expression of per was found to be relatively widespread, localized to the nucleus and cytoplasm of several tissues (Siwicki et al. 1988; Saez and Young 1988), and was not limited to the head, which was known to be the site of per s regulation of circadian locomotor behavior (Konopka et al. 1983). Most importantly, PER abundance in photoreceptor cells varied, with lowest levels observed in the middle of the day and highest levels in the middle of the night (Siwicki et al. 1988). This observation prompted an examination of the temporal expression of per mrna. RNA abundance cycled with a circadian rhythm in fly heads, with specifically altered periods in the mutants (Hardin et al. 1990). TIMELESS AND TEMPORAL DELAYS IN THE CIRCADIAN CLOCK Because high levels of PER protein were correlated with times of low per expression, it was proposed that PER proteins negatively regulate per RNA accumulation (Hardin et al. 1990). A basis for oscillating feedback, and two new classes of molecular cycling (affecting protein stability and subcellular localization), emerged with the discovery of a second-chromosome clock gene called timeless (tim) (Sehgal et al. 1994; Vosshall et al. 1994). The tim null mutation (tim 0 ) eliminated per RNA and protein oscillations in addition to causing arrhythmic eclosion and locomotor activity (Sehgal et al. 1994). Like per, tim mrna and protein were found to oscillate in a circadian fashion and these oscillations were abolished in per 0 and tim 0 mutants (Sehgal et al. 1995). Although the peak of tim mrna accumulation was slightly earlier than that of per, PER and TIM proteins accumulated with the same phase, indicating a link between these two genes (Sehgal et al. 1995). Subsequently it was shown that the PER and TIM proteins physically interact in vitro, in yeast and Drosophila cells (Gekakis et al. 1995; Saez and Young 1996). Interdependent functions of per and tim were also indicated by the finding that PER and TIM coexpression was required for the nuclear accumulation of either protein, and that PER stability was dependent on TIM (Vosshall et al. 1994; Price et al. 1995; Hunter-Ensor et al. 1996; Myers et al. 1996). Thus, with the discovery of tim and the regulated nuclear accumulation of PER and TIM (see also Curtin et al. 1995), a model was proposed that incorporated the concept of a specific temporal delay to explain how negative feedback could generate oscillations in gene expression. This delay was originally thought to be determined by the slow kinetics of association of PER and TIM in the cytoplasm, such that once dimerized, PER/TIM complexes would enter the nucleus and negatively regulate their own expression (cf. Sehgal et al. 1995). The isolation of several additional clock genes has helped to refine this mechanism (for review, see Young and Kay 2001; Hardin 2006). For example, some of the more recently identified genes regulate the phosphorylation or dephosphorylation of PER and TIM, affecting PER/TIM subcellular localization and stability (Harms et al. 2003; Bae and Edery 2006). Cold Spring Harbor Symposia on Quantitative Biology, Volume LXXII Cold Spring Harbor Laboratory Press
2 70 SAEZ, MEYER, AND YOUNG S2 CELLS AND NUCLEAR TRANSLOCATION OF PER AND TIM To better understand the regulation of PER/TIM interactions and their nuclear translocation, we began a new phase of work employing cultured Drosophila cells (Saez and Young 1996). Subsequently, this cell line (S2; Schneider s line 2), has become an important tool for investigating the function of Drosophila s circadian clock using transcription assays (Darlington et al. 1998), immunoprecipitation assays (Ceriani et al. 1999), degradation assays (Naidoo et al. 1999), and assays of phophorylation and dephosphorylation (cf. Ko et al. 2002; Sathyanarayanan et al. 2004). S2 cells were originally derived from dissociated embryos 35 years ago (Schneider 1972) and do not have a functional circadian clock. They express doubletime (dbt), casein kinase 2 (CK2), and cycle (cyc) (Nawathean et al. 2005; L. Saez, unpubl.) but lack expression of other known clock genes such as Clock (Clk) (Darlington et al. 1998), per, and tim (see Fig. 1) (Saez and Young 1996). Ectopic expression of Clock induces the expression of tim, and sometimes per in individual S2 cell lines (see Fig. 1) (McDonald and Rosbash 2001; Lim et al. 2007). Initially, we established immunocytochemically in fixed S2 cells that PER and TIM must be coexpressed to achieve efficient nuclear localization of either protein. Nuclear accumulation of both proteins was found to be complete approximately 6 hours after their initial synthesis (Saez and Young 1996). Expression of PER alone led to its accumulation in the cytoplasm with a half-life of approximately 10 hours as determined by pulse-chase experiments (see Fig. 1). TIM Figure 1. Response of endogenous clock genes in S2 cells to transient expression of Clock. (A) S2 cells were transfected with a plasmid containing a heat shock (HS) promoter fused to Clock cdna; 48 hours after transfection, cells were heat-shocked for 30 minutes at 37 C, and an equal amount of the culture was collected at the times (hr) indicated. HS-tim and HS-per are controls from S2 cells expressing heat-shock-induced PER or TIM. Following protein extraction and SDS-PAGE, proteins were transferred to a membrane and probed with anti-per (top) or anti-tim antibodies (bottom). (B) Half-life of PER in S2 cells. S2 cells transfected with HS-per were incubated in Schneider M3 media (Schneider 1972) without methionine and cysteine for 1 hour. Cells were heat-shocked for 30 minutes at 37 C, allowed to recover for 30 minutes at 25 C, and resuspended in media containing [ 35 S]methionine-cysteine for 30 minutes. Cells were collected by centrifugation and resuspended in complete M3 media (Sigma-Aldrich), and equal amounts of the growing culture were taken every 2 hours thereafter. 35 S-labeled PER was immunoprecipitated from each sample and subjected to PAGE. Theamountof 35 S-labeled PER recovered from each sample was expressed as the percentage remaining from the initial time point. The graph represents the average values obtained from four experiments. shuttles between the nucleus and cytoplasm with most cells showing cytoplasmic rather than nuclear accumulation (Ashmore et al. 2003; Meyer et al. 2006). Amino acid sequences that mediate PER/TIM interaction and cytoplasmic localization of PER and TIM (CLD; cytoplasmic localization domains), were also identified (Saez and Young 1996). Although these early studies reproduced elements of the temporal delay seen in nuclear translocation in vivo, and the inter-dependence of PER and TIM to achieve the delay, they gave little information about the kinetics of the PER/TIM interaction and did not provide the necessary resolution to establish a mechanism for regulated nuclear entry. PER AND TIM INTERACTION DOES NOT INDUCE NUCLEAR TRANSLOCATION To individually trace PER and TIM movements in the cytoplasm and to assess the role of PER/TIM interaction in nuclear translocation, we developed a single-cell, live imaging assay in S2 cells (Meyer et al. 2006). We individually or jointly expressed in a single cell, using a heat shock promoter, PER and TIM tagged at their carboxyl termini with cyan fluorescent protein (CFP) and yellow fluorescent protein (YFP), respectively. Initially, we followed PER-CFP for approximately 10 hours, when independently expressed in S2 cells. PER- CFP remained in the cytoplasm for the duration of the assay. Independent expression of TIM-YFP resulted in mainly cytoplasmic accumulation, with about 10% of the cells showing nuclear YFP (Meyer et al. 2006). These results are consistent with our previous observations using fixed S2 cells. We also confirmed that leptomycin-b abolishes nuclear export of TIM in S2 cells (Ashmore et al. 2003). However, our findings with PER differed from certain studies of subcellular localization that coexpressed Clk and per in S2 cells (Chang and Reppert 2003; Nawathean and Rosbash 2004). As previously discussed (Meyer and Young 2007), it seems likely that endogenous TIM that is produced in response to coexpressed Clk would influence PER nuclear accumulation in such studies. We next measured the onset of nuclear accumulation of PER and TIM when coexpressed in the same cell. Figure 2 shows time-lapse images of PER-CFP and TIM-YFP that were collected over several hours following heat shock induction of both proteins. The images show that PER and TIM were evenly distributed in the cytoplasm for a period of about 4 hours, followed by a gradual shift of both proteins to discrete cytoplasmic foci surrounding the nucleus. The onset of nuclear accumulation of PER and TIM occurred abruptly (e.g., ~340 minutes after heat shock induction for TIM in this cell), with the phase of nuclear accumulation lasting less than 1 hour (Fig. 2) (Meyer et al. 2006). Surprisingly, the onset of nuclear translocation proved to be largely independent of PER and TIM levels in this system (Meyer et al. 2006; Meyer and Young 2007). Even in the presence of leptomycin-b, TIM, which otherwise would accumulate in the nucleus, remained for approximately 5.5 hours in the cytoplasm in PER-TIM-expressing cells, presumably due to a rapid association with PER that retains TIM in the cytoplasm. The treatment with leptomycin-b did not affect the onset
3 PER/TIM/DBT INTERVAL TIMER 71 Figure 2. Nuclear accumulation profiles of PER-CFP and TIM- YFP in a single S2 cell. (Top two panels) Time-lapse images of a cell expressing PER-CFP and TIM-YFP. Starting at the top left, images were taken every 20 minutes, with the first image collected 180 minutes after heat shock. The last image (bottom right) was collected 400 minutes after heat shock. Arrows indicate the cytoplasmic foci whose formation preceded nuclear entry. The graph below these images shows the nuclear accumulation profiles (fluorescence measurements) of PER-CFP (thick line) andtim-yfp(thin line). In this example, the onset of nuclear accumulation of PER was advanced compared to TIM by about 25 minutes. (Redrawn, with permission, from Meyer et al.2006[ AAAS].) of nuclear entry in PER + TIM experiments (Meyer et al. 2006). These results indicate that nuclear translocation is not simply initiated by PER/TIM interaction, as corroborated by FRET analysis (see below). PER AND TIM NUCLEAR ACCUMULATION PROFILES ARE DIFFERENT Another intriguing observation came from a study of the individual patterns of nuclear accumulation of PER and TIM. A detailed analysis showed that nuclear accumulation profiles can be separated into three categories: In about 30% of the cells, PER and TIM have similar onsets and profiles of nuclear accumulation; in about 40% of the cells, PER and TIM had the same onsets of nuclear entry, but PER was transferred to nuclei more rapidly than TIM; in the remaining (~30%) cells, profiles of PER and TIM nuclear accumulation were similar, but the PER- CFP onset was advanced with respect to TIM (cf. Fig. 2). Thus, in approximately 70% of the cells analyzed, PER accumulated in the nucleus ahead of TIM (Meyer et al. 2006). This condition is similar to that previously observed in lateral neurons (pacemaker cells) of larval and adult brains, in which PER was detected in the nucleus before TIM (Shafer et al. 2002, 2004; Ashmore et al. 2003). These observations indicate that PER and TIM enter the nucleus independently. FRET ANALYSIS OF PER AND TIM INTERACTION Tagging PER and TIM with derivatives of GFP allowed us to develop a fluorescence resonant energy transfer (FRET) assay to measure PER/TIM interactions in a single cell. FRET is an indication of close proximity of two proteins, as intermolecular distances of less than 10 nm are needed for resonant energy transfer from one fluorophore to another (Forster 1948). In our S2 cell experiments, maximum levels of FRET were observed without a measurable delay following PER-CFP and TIM-YFP induction (see Fig. 3), suggesting rapid formation of PER/TIM complexes. Levels of FRET remained constant until the onset of PER and TIM nuclear translocation. A rapid decline in FRET was observed as the proteins moved to the nucleus. The interval over which FRET declined matched that of the independently measured nuclear translocation of PER and TIM in the same cell. Together with the divergent profiles of nuclear accumulation of PER and TIM, the loss of FRET suggests that following a prolonged physical association in the cytoplasm, PER and TIM dissociate and enter the nucleus independently. PER L MUTATION DELAYS NUCLEAR ENTRY WITHOUT AFFECTING THE PROFILE OF PER/TIM BINDING As the molecular behaviors of PER and TIM in S2 cells retained many properties previously observed in vivo, we sought a further test of this correlation by examining the behavior of proteins produced by two clock mutants, per L and tim UL, that generate long-period behavioral rhythms by different mechanisms. The mutation in per L is a singleamino-acid change (V242A) within the PAS (PER- ARNT-SM) domain (Baylies et al. 1987) that delays Figure 3. Profiles of FRET and nuclear translocation associated with PER + /TIM and PER L /TIM expression. (Top and bottom panels) Whole-cell FRET (thin lines) and nuclear accumulation (thick lines) profiles of PER-CFP and PER L -CFP, respectively, in five (top) and four (bottom) S2 cells expressing TIM-YFP. The decline in FRET levels coincides with the time of nuclear entry at approximately 330 minutes for PER-CFP (PER + )and about 500 minutes for PER L -CFP (PER L ). The bottom panel also shows the level of whole-cell FRET beginning 30 minutes after induction. For each panel, thick and thin lines of the same color represent measurements of the same cell. (Redrawn, with permission, from Meyer et al [ AAAS].
4 72 SAEZ, MEYER, AND YOUNG nuclear translocation of PER and TIM by 4 hours in pacemaker cells of the brain (Curtin et al. 1995), causing a 28- hour behavioral rhythm (Konopka and Benzer 1971). tim UL is a single-amino-acid substitution that delays the turnover of PER and TIM in the nucleus, giving a 33-hour circadian period without affecting the timing of nuclear translocation (Rothenfluh et al. 2000). The onset of nuclear entry for PER/TIM UL was quite similar to the onset observed for PER/TIM, indicating that tim UL does not affect nuclear entry in vivo or in S2 cells (Meyer et al. 2006). The per L mutation, on the other hand, could faithfully reproduce in S2 cells the increased delay observed in vivo, shifting the onset of nuclear accumulation of PER and TIM to about 8.5 hours (Fig. 3) (Meyer et al. 2006). Thus, a single-amino-acid change in PER appears to affect the function of an interval timer that regulates cytoplasmic retention of PER and TIM (Meyer et al. 2006). FRET, in cells expressing the wild-type and the mutant proteins, showed comparable rates of emergence and decay, indicating that association and dissociation kinetics of the PER/TIM complex are not substantially affected in the mutants. Because dissociation of the PER/TIM complex appears to be required for nuclear entry, it is possible that PER L alters a step needed to trigger dissociation from the PER L /TIM complex, delaying nuclear entry of both proteins. DBT, A NUCLEAR PROTEIN, IS RETAINED BY PER IN THE CYTOPLASM AND AFFECTS PER STABILITY We next focused our attention on DBT, another clock protein that associates with PER and the PER/TIM complex (Kloss et al. 1998, 2001). dbt is expressed throughout the circadian cycle. Nevertheless, in lateral neurons and in photoreceptor cells, DBT is nuclear in per and tim null mutants and shows a circadian subcellular distribution, cycling between the cytoplasm and nucleus in a fashion that parallels the behavior of PER accumulation in wild-type fly heads (Kloss et al. 2001). In vivo, DBT physically interacts with PER but is only found associated with TIM in PER/TIM complexes (Kloss et al. 2001). DBT-dependent phosphorylation destabilizes PER, promoting PER s degradation by the ubiquitin-proteosome degradation pathway (Price et al. 1998; Ko et al. 2002). TIM promotes PER stability, apparently by suppressing PER s phosphorylation by DBT (Kloss et al. 1998; Price et al. 1998; Ko et al. 2002). Thus, DBT-dependent phosphorylation is believed to affect the timing of nuclear entry in part by regulating the stability of PER. In turn, PER influences the subcellular distribution of DBT through formation of DBT/PER and DBT/PER/TIM complexes (Kloss et al. 2001). To study DBT in cell culture, we tagged it with YFP and induced DBT-YFP expression using a heat shock promoter. Consistent with observations made in vivo with per 0 mutants, DBT-YFP in S2 cells localized exclusively to the nucleus when expressed alone (Fig. 4A) or in the presence of TIM-CFP (Fig. 4B, top). Coexpresion of DBT-YFP and PER-CFP immediately relocalized DBT from the nucleus to the cytoplasm, presumably due to a Figure 4. Doubletime and PER interact in S2 cells. (A) Subcellular localization of DBT in S2 cells. DBT-YFP in S2 cells accumulates in the nucleus immediately following induction. (B) DBT interacts with PER and not with TIM. Images show cells expressing TIM-CFP and DBT-YFP (top) and PER- CFP and DBT-YFP (bottom). Approximately 3 hours after induction, TIM-CFP remains in the cytoplasm and DBT localizes to the nucleus. Coexpression of PER and DBT localized both proteins to the cytoplasm. (C) Time-lapse images of a single S2 cell expressing PER-CFP and DBT-YFP. As previously observed in vivo, in the absence of TIM, DBT promotes PER degradation. Images shown were taken every 32 minutes. physical association with cytoplasmic PER (see Fig. 4B, bottom). Because DBT affects the stability of PER, we collected time series data from these S2 cells over several hours (Fig. 4C). PER and DBT accumulated in the cytoplasm for approximately 180 minutes following their heat shock induction. Thereafter (~30 minutes), a progressive and rapid degradation of PER was observed with a relocalization of DBT to the nucleus. Because the degradation of PER did not occur immediately after its induction, and DBT was initially confined to the cytoplasm with PER, binding and a progressive phosphorylation of PER by DBT may be required for PER degradation. Endogenous DBT, which is found at low levels in S2 cells, did not have a detectable effect on the stability of PER in our experiments: PER, when expressed alone in S2 cells, was stable for at least 10 hours after induction, and dbt RNA interference (RNAi) did not alter the pattern of PER stability or the kinetics of nuclear entry in PER/TIM-expressing S2 cells (data not shown). Although progressive phosphorylation and degradation of PER were observed in our studies when PER was constitutively expressed and newly formed DBT was added through an inducible promoter, it was also shown by other investigators that coexpression of PER and DBT by a constitutive promoter can give constant low levels of PER that is highly phosphorylated (Ko et al. 2002; Ko and Edery 2005). These observations indicate that DBT affects PER stability and that patterns of DBT localization can be influenced by PER in S2 cells as previously observed in vivo. PER AND DBT ENTER THE NUCLEUS IN TIMELESS-EXPRESSING S2 CELLS To see whether TIM will protect PER from DBTdependent degradation, we coexpressed PER-CFP and DBT-YFP in TIM-expressing cells. Figure 5 shows time-
5 PER/TIM/DBT INTERVAL TIMER 73 Figure 5. Coordinated nuclear accumulation of PER, TIM, and DBT. S2 cells expressing TIM from a heat shock promoter were cotransfected with HS-per-cfp and HS-dbt-yfp; 48 hours after transfection, cells were heat-shocked at 37 C for 30 minutes and imaged for 464 minutes. Time-lapse images of a single cell expressing PER-CFP and DBT-YFP show that both proteins enter the nucleus with an onset similar to that previously seen for PER and TIM. Arrowheads indicate the formation of cytoplasmic PER/DBT foci before nuclear accumulation. lapse images of PER-CFP and DBT-YFP accumulation in these cells. Destabilization of PER by DBT was suppressed by TIM expression, because PER was observed to be stable in these cells for more than 8 hours, a period sufficient for complete DBT-dependent degradation of PER in the absence of TIM. As in the case of PER and TIM coexpression, DBT and PER progressively accumulated in the cytoplasm, forming discrete foci that presumably contain PER-CFP, DBT-YFP, and TIM (see small arrows in Fig. 5). The kinetics of nuclear accumulation of PER and DBT in these TIM-expressing cells has not been studied in detail but appears to be similar to that of PER and TIM, occurring at about 5 6 hours after induction. These results indicate that S2 cells recapitulate many of the in vivo behaviors of PER, TIM, and DBT. A summary of PER/TIM/DBT interactions and functions is indicated in the model in Figure 6. Further work will be required to determine patterns of PER degradation, whether PER and DBT independently enter nuclei, and profiles of FRET in cells coexpressing the three proteins. THE INTERVAL TIMER Although PER and TIM binding is not sufficient to trigger nuclear entry, physical association appears to be a prerequisite for their timed nuclear accumulation. The formation of the PER/TIM complex could, perhaps, allow subsequent posttranslational modifications or the binding of factors that determine the specific duration of its cytoplasmic phase. The formation of cytoplasmic foci is especially intriguing as these are not observed when PER, or TIM, or DBT is expressed alone. The formation of these foci always precedes the onset of nuclear accumulation and they disappear as nuclear accumulation proceeds. The foci represent the final cytoplasmic phase of PER/TIM/DBT accumulation. Their contents may be further explored by a program of protein tagging that includes the remaining Figure 6. Model for regulated nuclear accumulation of PER, DBT, and TIM in S2 cells. Wild-type PER and TIM proteins associate with little or no delay and thereafter remain in the cytoplasm for about 5.5 hours. During this time, PER and TIM progressively associate in distinct foci, which are lost as PER and TIM dissociate and move to the nucleus. In the nucleus, PER, DBT, and possibly TIM, associate with Clock/Cycle complexes, inhibiting transcription of Clock/cycle-dependent genes including per and tim (see also Kim et al. 2007; Nawathean et al. 2007). The duration of the PER/TIM complex in the cytoplasm is affected by PER L which delays the onset of nuclear accumulation by about 3 hours. A similar delay is observed in vivo (Curtin et al. 1995) and is thought to generate the abnormal (28 hours) period length of per L flies. In addition to these interactions, DBT association leads to phosphorylation and destabilization of PER in the absence of TIM. Our studies have not determined whether DBT and PER move to the nucleus independently or as a complex. clock proteins, as well as proteins known to be involved in either cytosolic trafficking or transport to the nucleus. CONCLUSIONS We have shown that a simple nonoscillating cell culture system can be used to study the molecular features of a key step in the circadian clockworks. This single-cell assay not only recapitulates many of the in vivo behaviors of PER and TIM and their mutants, but it has also provided a revised model of the Drosophila circadian clock. ACKNOWLEDGMENTS We are grateful to A. North (Director, The Rockefeller University Bio-Imaging Resource Center) for interest, advice, and helpful discussion. This work was supported by the National Institutes of Health grant GM to M.W.Y. REFERENCES Ashmore L.J., Sathyanarayanan S., Silvestre D.W., Emerson M.M., Schotland P., and Sehgal A Novel insights into the regulation of the timeless protein. J. Neurosci. 23: Bae K. and Edery I Regulating a circadian clock s period, phase and amplitude by phosphorylation: Insights from Drosophila. J. Biochem. 140: 609. Bargiello T.A. and Young M.W Molecular genetics of a biological clock in Drosophila. Proc. Natl. Acad. Sci. 81: 2142.
6 74 SAEZ, MEYER, AND YOUNG Baylies M.K., Bargiello T.A., Jackson F.R., and Young M.W Changes in abundance or structure of the per gene product can alter periodicity of the Drosophila clock. Nature 326: 390. Ceriani M.F., Darlington T.K., Staknis D., Mas P., Petti A.A., Weitz C.J., and Kay S.A Light-dependent sequestration of TIMELESS by CRYPTOCHROME. Science 285: 506. Chang D.C. and Reppert S.M A novel C-terminal domain of Drosophila PERIOD inhibits dclock:cycle-mediated transcription. Curr. Biol. 13: 758. Citri Y., Colot H.V., Jacquier A.C., Yu Q., Hall J.C., Baltimore D., and Rosbash M.A A family of unusually spliced biologically active transcripts encoded by a Drosophila clock gene. Nature 326: 42. Curtin K.D., Huang Z.J., and Rosbash M Temporally regulated nuclear entry of the Drosophila period protein contributes to the circadian clock. Neuron 14: 365. Darlington T.K., Wager-Smith K., Ceriani M.F., Staknis D., Gekakis N., Steeves T.D., Weitz C.J., Takahashi J.S., and Kay S.A Closing the circadian loop: CLOCK-induced transcription of its own inhibitors per and tim. Science 280: Forster T Intermolecular energy migration and fluorescence. Ann. Physics 2: 55. Gekakis N., Saez L., Delahaye-Brown A.M., Myers M.P., Sehgal A., Young M.W., and Weitz C.J Isolation of timeless by PER protein interaction: Defective interaction between timeless protein and long-period mutant PERL. Science 270: 732. Hardin P.E Essential and expendable features of the circadian timekeeping mechanism. Curr. Opin. Neurobiol. 16: 686. Hardin P.E., Hall J.C., and Rosbash M Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature 343: 536. Harms E., Young M.W., and Saez L CK1 and GSK3 in the Drosophila and mammalian circadian clock. Novartis Found. Symp. 253: 267. Hunter-Ensor M., Ousley A., and Sehgal A Regulation of the Drosophila protein timeless suggests a mechanism for resetting the circadian clock by light. Cell 84: 677. Jackson F.R., Bargiello T.A., Yun S.H., and Young M.W Product of per locus of Drosophila shares homology with proteoglycans. Nature 320: 185. Kim E.Y., Ko H.W., Yu W., Hardin P.E., and Edery I A DOUBLETIME kinase binding domain on the Drosophila PERIOD protein is essential for its hyperphosphorylation, transcriptional repression, and circadian clock function. Mol. Cell. Biol. 27: Kloss B., Rothenfluh A., Young M.W., and Saez L Phosphorylation of period is influenced by cycling physical associations of double-time, period, and timeless in the Drosophila clock. Neuron 30: 699. Kloss B., Price J.L., Saez L., Blau J., Rothenfluh A., Wesley C.S., and Young M.W The Drosophila clock gene double-time encodes a protein closely related to human casein kinase Iepsilon. Cell 94: 97. Ko H.W. and Edery I Analyzing the degradation of PERIOD protein by the ubiquitin-proteasome pathway in cultured Drosophila cells. Methods Enzymol. 393: 394. Ko H.W., Jiang J., and Edery I Role for Slimb in the degradation of Drosophila Period protein phosphorylated by Doubletime. Nature 420: 673. Konopka R.J. and Benzer S Clock mutants of Drosophila melanogaster. Proc. Natl. Acad. Sci. 68: Konopka R., Wells S., and Lee T Mosaic analysis of a Drosophila clock mutant. Mol. Gen. Genet. 190: 284. Lim C., Lee J., Choi C., Doh E., and Choe J Functional role of CREB-binding protein in the circadian clock system of Drosophila. Mol. Cell. Biol. 27: McDonald M.J. and Rosbash M Microarray analysis and organization of circadian gene expression in Drosophila. Cell 107: 567. Meyer P. and Young M.W The 2006 Pittendrigh/Aschoff Lecture: New roles for old proteins in the Drosophila circadian clock. J. Biol. Rhythms 22: 283. Meyer P., Saez L., and Young M.W PER-TIM interactions in living Drosophila cells: An interval timer for the circadian clock. Science 311: 226. Myers M.P., Wager-Smith K., Rothenfluh-Hilfiker A., and Young M.W Light-induced degradation of TIMELESS and entrainment of the Drosophila circadian clock. Science 271: Naidoo N., Song W., Hunter-Ensor M., and Sehgal A A role for the proteasome in the light response of the timeless clock protein. Science 285: Nawathean P. and Rosbash M The doubletime and CKII kinases collaborate to potentiate Drosophila PER transcriptional repressor activity. Mol. Cell 13: 213. Nawathean P., Menet J.S., and Rosbash M Assaying the Drosophila negative feedback loop with RNA interference in S2 cells. Methods Enzymol. 393: 610. Nawathean P., Stoleru D., and Rosbash M A small conserved domain of Drosophila PERIOD is important for circadian phosphorylation, nuclear localization, and transcriptional repressor activity. Mol. Cell. Biol. 27: Price J.L., Dembinska M.E., Young M.W., and Rosbash M Suppression of PERIOD protein abundance and circadian cycling by the Drosophila clock mutation timeless. EMBO J. 14: Price J.L., Blau J., Rothenfluh A., Abodeely M., Kloss B., and Young M.W double-time is a novel Drosophila clock gene that regulates PERIOD protein accumulation. Cell 94: 83. Reddy P., Zehring W. A., Wheeler D. A., Pirrotta V., Hadfield C., Hall J. C., and Rosbash M Molecular analysis of the period locus in Drosophila melanogaster and identification of a transcript involved in biological rhythms. Cell 38: 701. Rothenfluh A., Saez L., and Young M. W A TIMELESSindependent function for PERIOD proteins in the Drosophila clock. Neuron 26: 505. Saez L. and Young M.W In situ localization of the per clock protein during development of Drosophila melanogaster. Mol. Cell. Biol. 8: Regulation of nuclear entry of the Drosophila clock proteins period and timeless. Neuron 17: 911. Sathyanarayanan S., Zheng X., Xiao R., and Sehgal A Posttranslational regulation of Drosophila PERIOD protein by protein phosphatase 2A. Cell 116: 603. Schneider I Cell lines derived from late embryo stages of Drosophila melanogaster. J. Embryol. Exp. Morphol. 27: 353. Sehgal A., Price J.L., Man B., and Young M.W Loss of circadian behavioral rhythms and per RNA oscillations in the Drosophila mutant timeless. Science 263: Sehgal A., Rothenfluh-Hilfiker A., Hunter-Ensor M., Chen Y., Myers M.P., and Young M.W Rhythmic expression of timeless: A basis for promoting circadian cycles in period gene autoregulation. Science 270: 808. Shafer O.T., Rosbash M., and Truman J.W Sequential nuclear accumulation of the clock proteins period and timeless in the pacemaker neurons of Drosophila melanogaster. J. Neurosci. 22: Shafer O.T., Levine J.D., Truman J.W., and Hall J.C Flies by night: Effects of changing day length on Drosophila s circadian clock. Curr. Biol. 14: 424. Siwicki K.K., Eastman C., Petersen G., Rosbash M., and Hall J.C Antibodies to the period gene product of Drosophila reveal diverse tissue distribution and rhythmic changes in the visual system. Neuron 1: 141. Vosshall L.B., Price J.L., Sehgal A., Saez L., and Young M.W Block in nuclear localization of period protein by a second clock mutation, timeless. Science 263: Young M.W. and Kay S.A Time zones: A comparative genetics of circadian clocks. Nat. Rev. Genet. 2: 702.
The Nobel Assembly at Karolinska Institutet has today decided to award. the 2017 Nobel Prize in Physiology or Medicine. jointly to
 The Nobel Assembly at Karolinska Institutet has today decided to award the 2017 Nobel Prize in Physiology or Medicine jointly to Jeffrey C. Hall, Michael Rosbash and Michael W. Young for their discoveries
The Nobel Assembly at Karolinska Institutet has today decided to award the 2017 Nobel Prize in Physiology or Medicine jointly to Jeffrey C. Hall, Michael Rosbash and Michael W. Young for their discoveries
Altered Entrainment and Feedback Loop Function Effected by a Mutant Period Protein
 The Journal of Neuroscience, February 1, 2000, 20(3):958 968 Altered Entrainment and Feedback Loop Function Effected by a Mutant Period Protein Peter Schotland, Melissa Hunter-Ensor, Todd Lawrence, and
The Journal of Neuroscience, February 1, 2000, 20(3):958 968 Altered Entrainment and Feedback Loop Function Effected by a Mutant Period Protein Peter Schotland, Melissa Hunter-Ensor, Todd Lawrence, and
Scientific Background Discoveries of Molecular Mechanisms Controlling the Circadian Rhythm
 Scientific Background Discoveries of Molecular Mechanisms Controlling the Circadian Rhythm The 2017 Nobel Prize in Physiology or Medicine is awarded to Jeffrey C. Hall, Michael Rosbash and Michael W. Young
Scientific Background Discoveries of Molecular Mechanisms Controlling the Circadian Rhythm The 2017 Nobel Prize in Physiology or Medicine is awarded to Jeffrey C. Hall, Michael Rosbash and Michael W. Young
Clicker Question. The Need to Decompose. Mechanism and Reduction: Decomposing Circadian Clocks
 Mechanism and Reduction: Decomposing Circadian Clocks Clicker Question On the Deductive-Nomological (DN) model of reduction, which of the following does not figure in providing the explanation (i.e., is
Mechanism and Reduction: Decomposing Circadian Clocks Clicker Question On the Deductive-Nomological (DN) model of reduction, which of the following does not figure in providing the explanation (i.e., is
Transcriptional Feedback Loop Regulation, Function, and Ontogeny in Drosophila
 Transcriptional Feedback Loop Regulation, Function, and Ontogeny in Drosophila J. BENITO, H. ZHENG, F.S. NG, AND P.E. HARDIN Center for Research on Biological Clocks, Department of Biology, Texas A&M University,
Transcriptional Feedback Loop Regulation, Function, and Ontogeny in Drosophila J. BENITO, H. ZHENG, F.S. NG, AND P.E. HARDIN Center for Research on Biological Clocks, Department of Biology, Texas A&M University,
The tim SL Mutant Affects a Restricted Portion of the Drosophila melanogaster Circadian Cycle
 JO RutURNAL ila et al. OF / tim BIO SL LOGICAL MUTANT RHYTHMS / October 1998 The tim SL Mutant Affects a Restricted Portion of the Drosophila melanogaster Circadian Cycle Joan E. Rutila, * Olga Maltseva,
JO RutURNAL ila et al. OF / tim BIO SL LOGICAL MUTANT RHYTHMS / October 1998 The tim SL Mutant Affects a Restricted Portion of the Drosophila melanogaster Circadian Cycle Joan E. Rutila, * Olga Maltseva,
Biological Clocks. Lu Chen, Ph.D. MCB, UC Berkeley. Why Does Melatonin Now Outsell Vitamin C??
 Biological Clocks Lu Chen, Ph.D. MCB, UC Berkeley 1 Why Does Melatonin Now Outsell Vitamin C?? Wake / sleep complaints are extremely prevalent. Much melatonin is consumed in an attempt to overcome the
Biological Clocks Lu Chen, Ph.D. MCB, UC Berkeley 1 Why Does Melatonin Now Outsell Vitamin C?? Wake / sleep complaints are extremely prevalent. Much melatonin is consumed in an attempt to overcome the
Biological Clocks. Lu Chen, Ph.D. MCB, UC Berkeley. What is biological clock?
 Biological Clocks Lu Chen, Ph.D. MCB, UC Berkeley 1 What is biological clock? All eukaryotes and some prokaryotes display changes in gene activity, biochemistry, physiology, and behavior that wax and wane
Biological Clocks Lu Chen, Ph.D. MCB, UC Berkeley 1 What is biological clock? All eukaryotes and some prokaryotes display changes in gene activity, biochemistry, physiology, and behavior that wax and wane
What Is There Left to Learn about the Drosophila Clock?
 What Is There Left to Learn about the Drosophila Clock? J. BLAU, F. BLANCHARD, B. COLLINS, D. DAHDAL, A. KNOWLES, D. MIZRAK, AND M. RUBEN Department of Biology, New York University, New York, New York
What Is There Left to Learn about the Drosophila Clock? J. BLAU, F. BLANCHARD, B. COLLINS, D. DAHDAL, A. KNOWLES, D. MIZRAK, AND M. RUBEN Department of Biology, New York University, New York, New York
The Drosophila melanogaster life cycle
 The Drosophila melanogaster life cycle Eclosion The first phenotype described in Drosophila as an endogenous rhythm (1954): Number of eclosed flies Hamblen et al., Gene3cs 1998 rhythm with a periodicity
The Drosophila melanogaster life cycle Eclosion The first phenotype described in Drosophila as an endogenous rhythm (1954): Number of eclosed flies Hamblen et al., Gene3cs 1998 rhythm with a periodicity
Report. NEMO Kinase Contributes to Core Period Determination by Slowing the Pace of the Drosophila Circadian Oscillator
 Current Biology 21, 756 761, May 10, 2011 ª2011 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2011.02.037 NEMO Kinase Contributes to Core Period Determination by Slowing the Pace of the Drosophila
Current Biology 21, 756 761, May 10, 2011 ª2011 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2011.02.037 NEMO Kinase Contributes to Core Period Determination by Slowing the Pace of the Drosophila
TIMELESS-dependent positive and negative autoregulation in the Drosophila circadian clock
 The EMBO Journal Vol.18 No.3 pp.675 686, 1999 TIMELESS-dependent positive and negative autoregulation in the Drosophila circadian clock Vipin Suri, Anne Lanjuin and Michael Rosbash 1 Departments of Biochemistry
The EMBO Journal Vol.18 No.3 pp.675 686, 1999 TIMELESS-dependent positive and negative autoregulation in the Drosophila circadian clock Vipin Suri, Anne Lanjuin and Michael Rosbash 1 Departments of Biochemistry
Molecular Cogs of the Insect Circadian Clock
 ZOOLOGICAL SCIENCE 20: 947 955 (2003) 2003 Zoological Society of Japan [REVIEW] Molecular Cogs of the Insect Circadian Clock Naoto Shirasu 1, Yasuyuki Shimohigashi 1, Yoshiya Tominaga 2, and Miki Shimohigashi
ZOOLOGICAL SCIENCE 20: 947 955 (2003) 2003 Zoological Society of Japan [REVIEW] Molecular Cogs of the Insect Circadian Clock Naoto Shirasu 1, Yasuyuki Shimohigashi 1, Yoshiya Tominaga 2, and Miki Shimohigashi
The CREB-binding protein affects the circadian regulation of behaviour
 The CREB-binding protein affects the circadian regulation of behaviour Christian Maurer, Tobias Winter, Siwei Chen, Hsiu-Cheng Hung and Frank Weber Biochemistry Center Heidelberg, University of Heidelberg,
The CREB-binding protein affects the circadian regulation of behaviour Christian Maurer, Tobias Winter, Siwei Chen, Hsiu-Cheng Hung and Frank Weber Biochemistry Center Heidelberg, University of Heidelberg,
Stochastic simulations
 Stochastic simulations Application to circadian clocks Didier Gonze Circadian rhythms Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Circadian rhythms
Stochastic simulations Application to circadian clocks Didier Gonze Circadian rhythms Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Circadian rhythms
Simulation of Drosophila Circadian Oscillations, Mutations, and Light Responses by a Model with VRI, PDP-1, and CLK
 Title: Simulation of Drosophila Circadian Oscillations, Mutations, and Light Responses by a Model with VRI, PDP-1, and CLK Authors: Paul Smolen, Paul E. Hardin *, Brian S. Lo, Douglas A. Baxter, and John
Title: Simulation of Drosophila Circadian Oscillations, Mutations, and Light Responses by a Model with VRI, PDP-1, and CLK Authors: Paul Smolen, Paul E. Hardin *, Brian S. Lo, Douglas A. Baxter, and John
Advance in circadian rhythm genetics in mammals
 16 2 2004 4 Chinese Bulletin of Life Sciences Vol. 16, No. 2 Apr., 2004 1004-0374 (2004) 02-0104-05 1 100101 2 434025 9 24, Q41 A Advance in circadian rhythm genetics in mammals XU Zu-Yuan 1,2 (1 Beijing
16 2 2004 4 Chinese Bulletin of Life Sciences Vol. 16, No. 2 Apr., 2004 1004-0374 (2004) 02-0104-05 1 100101 2 434025 9 24, Q41 A Advance in circadian rhythm genetics in mammals XU Zu-Yuan 1,2 (1 Beijing
Stochastic simulations
 Circadian rhythms Stochastic simulations Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Application to circadian clocks Didier Gonze Circadian rhythms
Circadian rhythms Stochastic simulations Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Application to circadian clocks Didier Gonze Circadian rhythms
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Simulation of Drosophila Circadian Oscillations, Mutations, and Light Responses by a Model with VRI, PDP-1, and CLK
 2786 Biophysical Journal Volume 86 May 2004 2786 2802 Simulation of Drosophila Circadian Oscillations, Mutations, and Light Responses by a Model with VRI, PDP-1, and CLK Paul Smolen, Paul E. Hardin,* Brian
2786 Biophysical Journal Volume 86 May 2004 2786 2802 Simulation of Drosophila Circadian Oscillations, Mutations, and Light Responses by a Model with VRI, PDP-1, and CLK Paul Smolen, Paul E. Hardin,* Brian
Post-translational timing mechanisms of the Drosophila circadian clock
 FEBS Letters 585 (2011) 1443 1449 journal homepage: www.febsletters.org Review Post-translational timing mechanisms of the Drosophila circadian clock Frank Weber, Daniela Zorn, Christoph Rademacher, Hsiu-Cheng
FEBS Letters 585 (2011) 1443 1449 journal homepage: www.febsletters.org Review Post-translational timing mechanisms of the Drosophila circadian clock Frank Weber, Daniela Zorn, Christoph Rademacher, Hsiu-Cheng
Molecular Analysis of Sleep:Wake Cycles in Drosophila
 Molecular Analysis of Sleep:Wake Cycles in Drosophila A. SEHGAL,* W. JOINER,* A. CROCKER,* K. KOH,* S. SATHYANARAYANAN, Y. FANG,* M. WU,* J.A. WILLIAMS, AND X. ZHENG* *Howard Hughes Medical Institute,
Molecular Analysis of Sleep:Wake Cycles in Drosophila A. SEHGAL,* W. JOINER,* A. CROCKER,* K. KOH,* S. SATHYANARAYANAN, Y. FANG,* M. WU,* J.A. WILLIAMS, AND X. ZHENG* *Howard Hughes Medical Institute,
Drosophila Photoreceptors Contain an Autonomous Circadian Oscillator That Can Function without period mrna Cycling
 The Journal of Neuroscience, January 15, 1998, 18(2):741 750 Drosophila Photoreceptors Contain an Autonomous Circadian Oscillator That Can Function without period mrna Cycling Yuzhong Cheng 1 and Paul
The Journal of Neuroscience, January 15, 1998, 18(2):741 750 Drosophila Photoreceptors Contain an Autonomous Circadian Oscillator That Can Function without period mrna Cycling Yuzhong Cheng 1 and Paul
Supplementary Figure 1. Expression of the inducible tper2 is proportional to Dox/Tet concentration in Rosa-DTG/Per2 Per2-luc/wt MEFs.
 Supplementary Figure 1. Expression of the inducible tper2 is proportional to Dox/Tet concentration in Rosa-DTG/Per2 Per2-luc/wt MEFs. (a) Dose-responsive expression of tper2 by Dox. Note that there are
Supplementary Figure 1. Expression of the inducible tper2 is proportional to Dox/Tet concentration in Rosa-DTG/Per2 Per2-luc/wt MEFs. (a) Dose-responsive expression of tper2 by Dox. Note that there are
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras
 Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Circadian Transcription Contributes to Core Period Determination in Drosophila
 Circadian Transcription Contributes to Core Period Determination in Drosophila Sebastian Kadener 1,3, Jerome S. Menet 2,3, Rebecca Schoer 1,3, Michael Rosbash 1,2,3* PLoS BIOLOGY 1 Department of Biology,
Circadian Transcription Contributes to Core Period Determination in Drosophila Sebastian Kadener 1,3, Jerome S. Menet 2,3, Rebecca Schoer 1,3, Michael Rosbash 1,2,3* PLoS BIOLOGY 1 Department of Biology,
Welcome to Bi !
 Welcome to Bi156 2012! Professors: Paul Patterson (php@caltech.edu) Kai Zinn (zinnk@caltech.edu) TAs: Janna Nawroth Yanan Sui ysui@caltech.edu Student presentations Select a topic
Welcome to Bi156 2012! Professors: Paul Patterson (php@caltech.edu) Kai Zinn (zinnk@caltech.edu) TAs: Janna Nawroth Yanan Sui ysui@caltech.edu Student presentations Select a topic
Even a stopped clock tells the right time twice a day: circadian timekeeping in Drosophila
 Pflugers Arch - Eur J Physiol (2007) 454:857 867 DOI 10.1007/s00424-006-0188-9 INVITED REVIEW Even a stopped clock tells the right time twice a day: circadian timekeeping in Drosophila Ben Collins & Justin
Pflugers Arch - Eur J Physiol (2007) 454:857 867 DOI 10.1007/s00424-006-0188-9 INVITED REVIEW Even a stopped clock tells the right time twice a day: circadian timekeeping in Drosophila Ben Collins & Justin
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
Oscillation and Light Induction of timeless mrna in the Mammalian Circadian Clock
 The Journal of Neuroscience, 1999, Vol. 19 RC15 1of6 Oscillation and Light Induction of timeless mrna in the Mammalian Circadian Clock Shelley A. Tischkau, 1 Jeffrey A. Barnes, 1 Fang-Ju Lin, 2 Edith M.
The Journal of Neuroscience, 1999, Vol. 19 RC15 1of6 Oscillation and Light Induction of timeless mrna in the Mammalian Circadian Clock Shelley A. Tischkau, 1 Jeffrey A. Barnes, 1 Fang-Ju Lin, 2 Edith M.
Behavioral and molecular analyses suggest that circadian output is disrupted by disconnected mutants
 The EMBO Journal vol. 1 1 no. 1 pp. 1-6, 1992 Behavioral and molecular analyses suggest that circadian output is disrupted by disconnected mutants in D.melanogaster Paul E.Hardin1,2,3, Jeffrey C.Hall2
The EMBO Journal vol. 1 1 no. 1 pp. 1-6, 1992 Behavioral and molecular analyses suggest that circadian output is disrupted by disconnected mutants in D.melanogaster Paul E.Hardin1,2,3, Jeffrey C.Hall2
Post-translational modifications in circadian rhythms
 Review Post-translational modifications in circadian rhythms Arun Mehra 1, Christopher L. Baker 1, Jennifer J. Loros 1,2 and Jay C. Dunlap 1,2 1 Department of Genetics, Dartmouth Medical School, Hanover,
Review Post-translational modifications in circadian rhythms Arun Mehra 1, Christopher L. Baker 1, Jennifer J. Loros 1,2 and Jay C. Dunlap 1,2 1 Department of Genetics, Dartmouth Medical School, Hanover,
The period (per) gene in Drosophila melanogaster plays a central
 Seasonal behavior in Drosophila melanogaster requires the photoreceptors, the circadian clock, and phospholipase C B. H. Collins*, E. Rosato, and C. P. Kyriacou* Departments of *Genetics and Biology, University
Seasonal behavior in Drosophila melanogaster requires the photoreceptors, the circadian clock, and phospholipase C B. H. Collins*, E. Rosato, and C. P. Kyriacou* Departments of *Genetics and Biology, University
Posttranslational Control of the Neurospora Circadian Clock
 Posttranslational Control of the Neurospora Circadian Clock J. CHA, G.HUANG, J.GUO, AND Y. LIU Department of Physiology, University of Texas Southwestern Medical Center, Dallas, Texas 75390-9040 The eukaryotic
Posttranslational Control of the Neurospora Circadian Clock J. CHA, G.HUANG, J.GUO, AND Y. LIU Department of Physiology, University of Texas Southwestern Medical Center, Dallas, Texas 75390-9040 The eukaryotic
Circadian Rhythms in Physiology and Behavior. The Persistence of Memory, Salvador Dali, 1931
 Circadian Rhythms in Physiology and Behavior The Persistence of Memory, Salvador Dali, 1931 Homeostasis and Rhythms? Homeostasis (Bernard, 1878): All the vital mechanisms, however varied they may be, have
Circadian Rhythms in Physiology and Behavior The Persistence of Memory, Salvador Dali, 1931 Homeostasis and Rhythms? Homeostasis (Bernard, 1878): All the vital mechanisms, however varied they may be, have
Bio 401 Sp2014. Multiple Choice (Circle the letter corresponding to the correct answer)
 MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
Problem Set 5 KEY
 2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
Fig. S1. Subcellular localization of overexpressed LPP3wt-GFP in COS-7 and HeLa cells. Cos7 (top) and HeLa (bottom) cells expressing for 24 h human
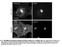 Fig. S1. Subcellular localization of overexpressed LPP3wt-GFP in COS-7 and HeLa cells. Cos7 (top) and HeLa (bottom) cells expressing for 24 h human LPP3wt-GFP, fixed and stained for GM130 (A) or Golgi97
Fig. S1. Subcellular localization of overexpressed LPP3wt-GFP in COS-7 and HeLa cells. Cos7 (top) and HeLa (bottom) cells expressing for 24 h human LPP3wt-GFP, fixed and stained for GM130 (A) or Golgi97
2013 John Wiley & Sons, Inc. All rights reserved. PROTEIN SORTING. Lecture 10 BIOL 266/ Biology Department Concordia University. Dr. S.
 PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
Significance. 1 To whom correspondence should be addressed.
 Pacemaker-neuron dependent disturbance of the molecular clockwork by a Drosophila CLOCK mutant homologous to the mouse Clock mutation Euna Lee a,b, Eunjoo Cho c, Doo Hyun Kang a,b, Eun Hee Jeong b, Zheng
Pacemaker-neuron dependent disturbance of the molecular clockwork by a Drosophila CLOCK mutant homologous to the mouse Clock mutation Euna Lee a,b, Eunjoo Cho c, Doo Hyun Kang a,b, Eun Hee Jeong b, Zheng
Diagrams as Vehicles for Scientific Reasoning. Adele Abrahamsen Center for Research in Language University of California, San Diego
 Diagrams as Vehicles for Scientific Reasoning William Bechtel Department of Philosophy, Center for Chronobiology, and Interdisciplinary Program in Cognitive Science University of California, San Diego
Diagrams as Vehicles for Scientific Reasoning William Bechtel Department of Philosophy, Center for Chronobiology, and Interdisciplinary Program in Cognitive Science University of California, San Diego
Circadian Phase Entrainment via Nonlinear Model Predictive Control
 Paper No. 573b Circadian Phase Entrainment via Nonlinear Model Predictive Control Neda Bagheri, Jörg Stelling, and Francis J. Doyle III 1 Department of Electrical and Computer Engineering Department of
Paper No. 573b Circadian Phase Entrainment via Nonlinear Model Predictive Control Neda Bagheri, Jörg Stelling, and Francis J. Doyle III 1 Department of Electrical and Computer Engineering Department of
Diagrams as Tools for Scientific Reasoning. Adele Abrahamsen and William Bechtel University of California, San Diego
 Diagrams as Tools for Scientific Reasoning Adele Abrahamsen and William Bechtel University of California, San Diego Abstract We contend that diagrams are tools not only for communication but also for supporting
Diagrams as Tools for Scientific Reasoning Adele Abrahamsen and William Bechtel University of California, San Diego Abstract We contend that diagrams are tools not only for communication but also for supporting
High-amplitude circadian rhythms in Drosophila driven by. calcineurin-mediated post-translational control of sarah
 Genetics: Early Online, published on May 3, 218 as 1.1534/genetics.118.388 High-amplitude circadian rhythms in Drosophila driven by calcineurin-mediated post-translational control of sarah Sin Ho Kweon
Genetics: Early Online, published on May 3, 218 as 1.1534/genetics.118.388 High-amplitude circadian rhythms in Drosophila driven by calcineurin-mediated post-translational control of sarah Sin Ho Kweon
Analysis of Period mrna Cycling in Drosophila Head and Body Tissues Indicates that Body Oscillators Behave Differently from Head Oscillators
 MOLECULAR AND CELLULAR OLOGY, Nov. 1994, p. 7211-7218 0270-7306/94/$04.00+0 Copyright ( 1994, American Society for Microbiology Vol. 14, No. 11 Analysis of Period mrna Cycling in Drosophila Head and ody
MOLECULAR AND CELLULAR OLOGY, Nov. 1994, p. 7211-7218 0270-7306/94/$04.00+0 Copyright ( 1994, American Society for Microbiology Vol. 14, No. 11 Analysis of Period mrna Cycling in Drosophila Head and ody
MOLECULAR BASES OF CIRCADIAN RHYTHMS
 Annu. Rev. Cell Dev. Biol. 2001. 17:215 53 Copyright c 2001 by Annual Reviews. All rights reserved MOLECULAR BASES OF CIRCADIAN RHYTHMS Stacey L. Harmer, Satchidananda Panda, and Steve A. Kay Department
Annu. Rev. Cell Dev. Biol. 2001. 17:215 53 Copyright c 2001 by Annual Reviews. All rights reserved MOLECULAR BASES OF CIRCADIAN RHYTHMS Stacey L. Harmer, Satchidananda Panda, and Steve A. Kay Department
Neurons and Hormones 1. How do animals perform the right behaviors at the right time? In the right context?
 Neurons and Hormones 1 How do animals perform the right behaviors at the right time? In the right context? Active at night only What if conflicting signals? Magnetic cues are always present But migrate
Neurons and Hormones 1 How do animals perform the right behaviors at the right time? In the right context? Active at night only What if conflicting signals? Magnetic cues are always present But migrate
Organization of the Drosophila Circadian Control Circuit
 Current Biology 18, R84 R93, January 22, 2008 ª2008 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2007.11.061 Organization of the Drosophila Circadian Control Circuit Review Michael N. Nitabach 1
Current Biology 18, R84 R93, January 22, 2008 ª2008 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2007.11.061 Organization of the Drosophila Circadian Control Circuit Review Michael N. Nitabach 1
BioNSi: A Discrete Biological Network Simulator Tool
 BioNSi: A Discrete Biological Network Simulator Tool Amir Rubinstein 1, Noga Bracha 1, Liat Rudner 1, Noga Zucker 1, Hadas E. Sloin 2, and Benny Chor 1 1 Blavatnik School of Computer Science, Tel Aviv
BioNSi: A Discrete Biological Network Simulator Tool Amir Rubinstein 1, Noga Bracha 1, Liat Rudner 1, Noga Zucker 1, Hadas E. Sloin 2, and Benny Chor 1 1 Blavatnik School of Computer Science, Tel Aviv
Circadian Gene Expression in the Suprachiasmatic Nucleus
 Circadian Gene Expression in the Suprachiasmatic Nucleus 901 Circadian Gene Expression in the Suprachiasmatic Nucleus M U Gillette and S-H Tyan, University of Illinois at Urbana-Champaign, Urbana, IL,
Circadian Gene Expression in the Suprachiasmatic Nucleus 901 Circadian Gene Expression in the Suprachiasmatic Nucleus M U Gillette and S-H Tyan, University of Illinois at Urbana-Champaign, Urbana, IL,
Light-Mediated TIM Degradation within Drosophila Pacemaker Neurons (s-lnvs) Is Neither Necessary nor Sufficient for Delay Zone Phase Shifts
 Report Light-Mediated TIM egradation within rosophila Pacemaker Neurons (s) Is Neither Necessary nor Sufficient for elay Zone Phase Shifts hih-hang nthony Tang, Erica Hinteregger,, Yuhua Shang, and Michael
Report Light-Mediated TIM egradation within rosophila Pacemaker Neurons (s) Is Neither Necessary nor Sufficient for elay Zone Phase Shifts hih-hang nthony Tang, Erica Hinteregger,, Yuhua Shang, and Michael
Molecular genetics of the fruit-fly circadian clock
 (2006) 14, 729 738 & 2006 Nature Publishing Group All rights reserved 1018-4813/06 $30.00 www.nature.com/ejhg REVIEW Ezio Rosato 1, Eran Tauber 1 and Charalambos P Kyriacou*,1 1 Department of Genetics,
(2006) 14, 729 738 & 2006 Nature Publishing Group All rights reserved 1018-4813/06 $30.00 www.nature.com/ejhg REVIEW Ezio Rosato 1, Eran Tauber 1 and Charalambos P Kyriacou*,1 1 Department of Genetics,
SUPPLEMENTARY INFORMATION
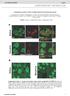 doi: 10.1038/nature06994 A phosphatase cascade by which rewarding stimuli control nucleosomal response A. Stipanovich*, E. Valjent*, M. Matamales*, A. Nishi, J.H. Ahn, M. Maroteaux, J. Bertran-Gonzalez,
doi: 10.1038/nature06994 A phosphatase cascade by which rewarding stimuli control nucleosomal response A. Stipanovich*, E. Valjent*, M. Matamales*, A. Nishi, J.H. Ahn, M. Maroteaux, J. Bertran-Gonzalez,
Interactions between circadian neurons control temperature synchronization of Drosophila behavior
 University of Massachusetts Medical School escholarship@umms GSBS Student Publications Graduate School of Biomedical Sciences 2007-10-05 Interactions between circadian neurons control temperature synchronization
University of Massachusetts Medical School escholarship@umms GSBS Student Publications Graduate School of Biomedical Sciences 2007-10-05 Interactions between circadian neurons control temperature synchronization
Drosophila Free-Running Rhythms Require Intercellular Communication
 Drosophila Free-Running Rhythms Require Intercellular Communication Ying Peng 1,2, Dan Stoleru 1,2, Joel D. Levine 1, Jeffrey C. Hall 1, Michael Rosbash 1,2* PLoS BIOLOGY 1 Department of Biology, Brandeis
Drosophila Free-Running Rhythms Require Intercellular Communication Ying Peng 1,2, Dan Stoleru 1,2, Joel D. Levine 1, Jeffrey C. Hall 1, Michael Rosbash 1,2* PLoS BIOLOGY 1 Department of Biology, Brandeis
Evidence that the TIM Light Response Is Relevant to Light-Induced Phase Shifts in Drosophila melanogaster
 Neuron, Vol. 21, 225 234, July, 1998, Copyright 1998 by Cell Press Evidence that the TIM Light Response Is Relevant to Light-Induced Phase Shifts in Drosophila melanogaster Vipin Suri,* Zuwei Qian,* Jeffrey
Neuron, Vol. 21, 225 234, July, 1998, Copyright 1998 by Cell Press Evidence that the TIM Light Response Is Relevant to Light-Induced Phase Shifts in Drosophila melanogaster Vipin Suri,* Zuwei Qian,* Jeffrey
Diagrams as Tools for Scientific Reasoning
 Rev.Phil.Psych. (2015) 6:117 131 DOI 10.1007/s13164-014-0215-2 Diagrams as Tools for Scientific Reasoning Adele Abrahamsen & William Bechtel Published online: 12 November 2014 # Springer Science+Business
Rev.Phil.Psych. (2015) 6:117 131 DOI 10.1007/s13164-014-0215-2 Diagrams as Tools for Scientific Reasoning Adele Abrahamsen & William Bechtel Published online: 12 November 2014 # Springer Science+Business
Transcriptional and Translational Mechanisms Controlling Circadian Rhythms in Drosophila: A Dissertation
 University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 6-14-2013 Transcriptional and Translational Mechanisms Controlling Circadian
University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 6-14-2013 Transcriptional and Translational Mechanisms Controlling Circadian
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
CIRCADIAN SIGNALING NETWORKS
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P. Matthias and RG Clerc Roger G. Clerc 07.05.2014 CIRCADIAN SIGNALING NETWORKS Master pacemaker SCN «Slave clocks»
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P. Matthias and RG Clerc Roger G. Clerc 07.05.2014 CIRCADIAN SIGNALING NETWORKS Master pacemaker SCN «Slave clocks»
Setting the clock by nature: Circadian rhythm in the fruitfly Drosophila melanogaster
 FEBS Letters 585 (2011) 1435 1442 journal homepage: www.febsletters.org Review Setting the clock by nature: Circadian rhythm in the fruitfly Drosophila melanogaster Nicolai Peschel, Charlotte Helfrich-Förster
FEBS Letters 585 (2011) 1435 1442 journal homepage: www.febsletters.org Review Setting the clock by nature: Circadian rhythm in the fruitfly Drosophila melanogaster Nicolai Peschel, Charlotte Helfrich-Förster
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
PER-TIM Interactions with the Photoreceptor Cryptochrome Mediate Circadian Temperature Responses in Drosophila
 PLoS BIOLOGY PER-TIM Interactions with the Photoreceptor Cryptochrome Mediate Circadian Temperature Responses in Drosophila Rachna Kaushik 1,2,3, Pipat Nawathean 1,2,3, Ania Busza 4,5,6, Alejandro Murad
PLoS BIOLOGY PER-TIM Interactions with the Photoreceptor Cryptochrome Mediate Circadian Temperature Responses in Drosophila Rachna Kaushik 1,2,3, Pipat Nawathean 1,2,3, Ania Busza 4,5,6, Alejandro Murad
Representing Time in Scientific Diagrams
 Representing Time in Scientific Diagrams William Bechtel (bechtel@ucsd.edu) Daniel Burnston (dburnston@ucsd.edu) Benjamin Sheredos (bsheredos@ucsd.edu) Department of Philosophy and Center for Circadian
Representing Time in Scientific Diagrams William Bechtel (bechtel@ucsd.edu) Daniel Burnston (dburnston@ucsd.edu) Benjamin Sheredos (bsheredos@ucsd.edu) Department of Philosophy and Center for Circadian
Emerging roles for microrna in the regulation of Drosophila circadian clock
 https://doi.org/10.1186/s12868-018-0401-8 BMC Neuroscience REVIEW Open Access Emerging roles for microrna in the regulation of Drosophila circadian clock Yongbo Xue and Yong Zhang * Abstract Background:
https://doi.org/10.1186/s12868-018-0401-8 BMC Neuroscience REVIEW Open Access Emerging roles for microrna in the regulation of Drosophila circadian clock Yongbo Xue and Yong Zhang * Abstract Background:
Supplementary Discussion 1: The mechanistic model of NIK1-mediated antiviral
 doi:10.1038/nature14171 SUPPLEMENTARY DISCUSSION Supplementary Discussion 1: The mechanistic model of NIK1-mediated antiviral signaling 1. Stress-induced oligomerization of the extracellular domain of
doi:10.1038/nature14171 SUPPLEMENTARY DISCUSSION Supplementary Discussion 1: The mechanistic model of NIK1-mediated antiviral signaling 1. Stress-induced oligomerization of the extracellular domain of
Circadian rhythms new functions for old clock genes?
 Clock genes Perspective Circadian rhythms new functions for old clock genes? The mechanisms of circadian clocks, which time daily events, are being investigated by characterizing clock genes that affect
Clock genes Perspective Circadian rhythms new functions for old clock genes? The mechanisms of circadian clocks, which time daily events, are being investigated by characterizing clock genes that affect
Theoretical models for circadian rhythms in Neurospora and Drosophila
 C.R. Acad. Sci. Paris, Sciences de la vie / Life Sciences 323 (2000) 57 67 2000 Académie des sciences/éditions scientifiques et médicales Elsevier SAS. Tous droits réservés S0764446900001116/FLA Theoretical
C.R. Acad. Sci. Paris, Sciences de la vie / Life Sciences 323 (2000) 57 67 2000 Académie des sciences/éditions scientifiques et médicales Elsevier SAS. Tous droits réservés S0764446900001116/FLA Theoretical
The Implications of Multiple Circadian Clock Origins Michael Rosbash
 Essay The Implications of Multiple Circadian Clock Origins Michael Rosbash In the Beginning Genetics has had an awesome impact on our understanding of basic processes like circadian rhythms, which were
Essay The Implications of Multiple Circadian Clock Origins Michael Rosbash In the Beginning Genetics has had an awesome impact on our understanding of basic processes like circadian rhythms, which were
Nature Genetics: doi: /ng.3731
 Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
The ins and outs of circadian timekeeping Steven A Brown* and Ueli Schibler
 588 The ins and outs of circadian timekeeping Steven A Brown* and Ueli Schibler Recent research in Drosophila and in mammals has generated fascinating new models for how circadian clocks in these organisms
588 The ins and outs of circadian timekeeping Steven A Brown* and Ueli Schibler Recent research in Drosophila and in mammals has generated fascinating new models for how circadian clocks in these organisms
8 Suppression Analysis
 Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 8 Suppression Analysis OVERVIEW Suppression
Genetic Techniques for Biological Research Corinne A. Michels Copyright q 2002 John Wiley & Sons, Ltd ISBNs: 0-471-89921-6 (Hardback); 0-470-84662-3 (Electronic) 8 Suppression Analysis OVERVIEW Suppression
supplementary information
 DOI: 10.1038/ncb2157 Figure S1 Immobilization of histone pre-mrna to chromatin leads to formation of histone locus body with associated Cajal body. Endogenous histone H2b(e) pre-mrna is processed with
DOI: 10.1038/ncb2157 Figure S1 Immobilization of histone pre-mrna to chromatin leads to formation of histone locus body with associated Cajal body. Endogenous histone H2b(e) pre-mrna is processed with
Signaling components that drive circadian rhythms Garrick K Wang* and Amita Sehgal
 331 Signaling components that drive circadian rhythms Garrick K Wang* and Amita Sehgal In the past year, knowledge of how information is relayed in the regulation of circadian rhythms has advanced considerably.
331 Signaling components that drive circadian rhythms Garrick K Wang* and Amita Sehgal In the past year, knowledge of how information is relayed in the regulation of circadian rhythms has advanced considerably.
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
REGULATORY MECHANISMS OF TRANSCRIPTION FACTOR FUNCTION
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) RG Clerc 04.04.2012 REGULATORY MECHANISMS OF TRANSCRIPTION FACTOR FUNCTION Protein synthesized Protein phosphorylated
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) RG Clerc 04.04.2012 REGULATORY MECHANISMS OF TRANSCRIPTION FACTOR FUNCTION Protein synthesized Protein phosphorylated
A Genetic Program for Embryonic Development
 Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Concept 18.4: A program of differential gene expression leads to the different cell types in a multicellular organism During embryonic development, a fertilized egg gives rise to many different cell types
Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists
 Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists of: (i) the YPet domain (an enhanced YFP); (ii) the
Supplementary Figure 1. Procedures for p38 activity imaging in living cells. (a) Schematic model of the p38 activity reporter. The reporter consists of: (i) the YPet domain (an enhanced YFP); (ii) the
The Success of Decomposition
 11/21/11 Mechanism and Levels of Organization: Recomposing and Situating Circadian Clocks The Success of Decomposition Moving beyond per, researchers in the 1990s and early 2000s identified many clock
11/21/11 Mechanism and Levels of Organization: Recomposing and Situating Circadian Clocks The Success of Decomposition Moving beyond per, researchers in the 1990s and early 2000s identified many clock
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Molecular and Neuronal Analysis of Circadian Photoresponses in Drosophila: A Dissertation
 University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 2007-10-25 Molecular and Neuronal Analysis of Circadian Photoresponses
University of Massachusetts Medical School escholarship@umms GSBS Dissertations and Theses Graduate School of Biomedical Sciences 2007-10-25 Molecular and Neuronal Analysis of Circadian Photoresponses
s u p p l e m e n ta ry i n f o r m at i o n
 Figure S1 Characterization of tet-off inducible cell lines expressing GFPprogerin and GFP-wt lamin A. a, Western blot analysis of GFP-progerin- or GFP-wt lamin A- expressing cells before induction (0d)
Figure S1 Characterization of tet-off inducible cell lines expressing GFPprogerin and GFP-wt lamin A. a, Western blot analysis of GFP-progerin- or GFP-wt lamin A- expressing cells before induction (0d)
Materiale per uso didattico 1
 BCII - Perroteau - shuttling citoplasma nucleus 1 ERK regulation by scaffolds and anchors 2 Materiale per uso didattico 1 ERK Pathway regulation by feedback loops and phosphatases 3 ERK has more
BCII - Perroteau - shuttling citoplasma nucleus 1 ERK regulation by scaffolds and anchors 2 Materiale per uso didattico 1 ERK Pathway regulation by feedback loops and phosphatases 3 ERK has more
mcry1 and mcry2 Are Essential Components of the Negative Limb of the Circadian Clock Feedback Loop
 Cell, Vol. 98, 193 205, July 23, 1999, Copyright 1999 by Cell Press mcry1 and mcry2 Are Essential Components of the Negative Limb of the Circadian Clock Feedback Loop Kazuhiko Kume,* # Mark J. Zylka,*
Cell, Vol. 98, 193 205, July 23, 1999, Copyright 1999 by Cell Press mcry1 and mcry2 Are Essential Components of the Negative Limb of the Circadian Clock Feedback Loop Kazuhiko Kume,* # Mark J. Zylka,*
[2010] Kwang Huei Low ALL RIGHTS RESERVED
![[2010] Kwang Huei Low ALL RIGHTS RESERVED [2010] Kwang Huei Low ALL RIGHTS RESERVED](/thumbs/94/118553955.jpg) [2010] Kwang Huei Low ALL RIGHTS RESERVED THERMOSENSITIVE SPLICING OF A CLOCK GENE AND SEASONAL ADAPTATION by KWANG HUEI LOW A dissertation submitted to the Graduate School-New Brunswick Rutgers, The State
[2010] Kwang Huei Low ALL RIGHTS RESERVED THERMOSENSITIVE SPLICING OF A CLOCK GENE AND SEASONAL ADAPTATION by KWANG HUEI LOW A dissertation submitted to the Graduate School-New Brunswick Rutgers, The State
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
SUPPLEMENTARY INFORMATION In format provided by JAATTELA (NOVEMBER 2005)
 Box S1: Methods for studying lysosomal function and integrity Volume and distribution of the acidic compartment. Acridine orange is a metachromatic fluorochrome and a weak base that accumulates in the
Box S1: Methods for studying lysosomal function and integrity Volume and distribution of the acidic compartment. Acridine orange is a metachromatic fluorochrome and a weak base that accumulates in the
Modeling Rhythms on Differents Levels: Cells, Tissues, and Organisms
 Modeling Rhythms on Differents Levels: Cells, Tissues, and Organisms Hanspeter Herzel Institute for Theoretical Biology (ITB) Charité and Humboldt University Berlin Molecular Chronobiology SCN-neuron nucleus
Modeling Rhythms on Differents Levels: Cells, Tissues, and Organisms Hanspeter Herzel Institute for Theoretical Biology (ITB) Charité and Humboldt University Berlin Molecular Chronobiology SCN-neuron nucleus
Supplementary Figure 1
 Supplementary Figure 1 Kif1a RNAi effect on basal progenitor differentiation Related to Figure 2. Representative confocal images of the VZ and SVZ of rat cortices transfected at E16 with scrambled or Kif1a
Supplementary Figure 1 Kif1a RNAi effect on basal progenitor differentiation Related to Figure 2. Representative confocal images of the VZ and SVZ of rat cortices transfected at E16 with scrambled or Kif1a
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Problem Set #5 4/3/ Spring 02
 Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Rapid blue-light mediated induction of protein interactions in living cells
 Nature Methods Rapid blue-light mediated induction of protein interactions in living cells Matthew J Kennedy, Robert M Hughes, Leslie A Peteya, Joel W Schwartz, Michael D Ehlers & Chandra L Tucker Supplementary
Nature Methods Rapid blue-light mediated induction of protein interactions in living cells Matthew J Kennedy, Robert M Hughes, Leslie A Peteya, Joel W Schwartz, Michael D Ehlers & Chandra L Tucker Supplementary
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/305/ra106/dc1 Supplementary Materials for Controlling Long-Term Signaling: Receptor Dynamics Determine Attenuation and Refractory Behavior of the TGF-β Pathway
www.sciencesignaling.org/cgi/content/full/6/305/ra106/dc1 Supplementary Materials for Controlling Long-Term Signaling: Receptor Dynamics Determine Attenuation and Refractory Behavior of the TGF-β Pathway
N.T.Kumar*,Thiago R,Sandro B,Vinicius R,Lisane V,Vanderlei P Versor Inovação,Santo André SP,Brazil. id* :
 General Theoretical Aspects of Cell Mechanics from Circadian Oscillator Point of View Based on Higher Order Logic(HOL) A Short Communication to Develop Informatics Framework Involving Bio-Chemical Concepts
General Theoretical Aspects of Cell Mechanics from Circadian Oscillator Point of View Based on Higher Order Logic(HOL) A Short Communication to Develop Informatics Framework Involving Bio-Chemical Concepts
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
SUPPLEMENTARY INFORMATION
 Figure S1 Treatment with both Sema6D and Plexin-A1 sirnas induces the phenotype essentially identical to that induced by treatment with Sema6D sirna alone or Plexin-A1 sirna alone. (a,b) The cardiac tube
Figure S1 Treatment with both Sema6D and Plexin-A1 sirnas induces the phenotype essentially identical to that induced by treatment with Sema6D sirna alone or Plexin-A1 sirna alone. (a,b) The cardiac tube
