Accepted Article Preview: Published ahead of advance online publication
|
|
|
- Owen Page
- 5 years ago
- Views:
Transcription
1 Accepted Article Preview: Published ahead of advance online publication Laboratory recommendations for scoring deep molecular responses following treatment for chronic myeloid leukemia OPEN N C P Cross, H White, D Colomer, H Ehrencrona, L Foroni, E Gottardi, T Lange, T Lion, K M Polakova, S Dulucq, G Martinelli, E O Leibundgut, N Pallisgaard, G Barbany, T Sacha, R Talmaci, B Izzo, G Saglio, F Pane, M C Mu ller, A Hochhaus Cite this article as: N C P Cross, H White, D Colomer, H Ehrencrona, L Foroni, E Gottardi, T Lange, T Lion, K M Polakova, S Dulucq, G Martinelli, E O Leibundgut, N Pallisgaard, G Barbany, T Sacha, R Talmaci, B Izzo, G Saglio, F Pane, M C Mu ller, A Hochhaus, Laboratory recommendations for scoring deep molecular responses following treatment for chronic myeloid leukemia, Leukemia accepted article preview 5 February 2015; doi: /leu This is a PDF file of an unedited peer-reviewed manuscript that has been accepted for publication. NPG are providing this early version of the manuscript as a service to our customers. The manuscript will undergo copyediting, typesetting and a proof review before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers apply. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit creativecommons.org/licenses/by/4.0/ Received 16 January 2015; accepted 21 January 2015; Accepted article preview online 5 February 2015
2 Laboratory recommendations for scoring deep molecular responses following treatment for chronic myeloid leukemia Nicholas C.P. Cross 1,2, Helen White 1,2, Dolors Colomer 3, Hans Ehrencrona 4, Letizia Foroni 5, Enrico Gottardi 6, Thoralf Lange 7, Thomas Lion 8, Katerina Machova Polakova 9, Stéphanie Dulucq 10, Giovanni Martinelli 11, Elisabeth Oppliger Leibundgut 12, Niels Pallisgaard 13, Gisela Barbany 14, Tomasz Sacha 15, Rodica Talmaci 16, Barbara Izzo 17, Giuseppe Saglio 6, Fabrizio Pane 17,18, Martin C. Müller 19, Andreas Hochhaus 20 1 Faculty of Medicine, University of Southampton, Southampton, UK 2 National Genetics Reference Laboratory (Wessex), Salisbury, UK 3 Hematopathology Unit, Hospital Clinic, IDIBAPS, Barcelona, Spain 4 Department of Clinical Genetics, Laboratory Medicine, Lund University, Lund, Sweden 5 Imperial Molecular Pathology Laboratory, Centre for Haematology, Imperial College London, London, UK 6 Department of Clinical and Biological Science, University of Turin, Turin, Italy 7 Abteilung für Hämatologie und internistische Onkologie, Universität Leipzig, Leipzig, Germany 8 Children s Cancer Research Institute/LabDia Labordiagnostik and Medical University, Vienna, Austria 9 Institute of Hematology and Blood Transfusion, Prague, Czech Republic 10 Laboratoire Hematologie, CHU Bordeaux, Hematopoiese Leucemique et Cibles Therapeutiques, INSERM U1035, Université Bordeaux, France 11 Department of Experimental, Diagnostic and Specialty Medicine, University of Bologna, Italy 12 Molecular Diagnostics Laboratory, Department of Hematology, University Hospital Bern, Bern, Switzerland 13 Klinisk Biokemi, Vejle Sygehus, Vejle, Denmark 14 Department of Molecular Medicine and Surgery, Clinical Genetics, Karolinska Institutet, Stockholm, Sweden 1
3 15 Hematology Department, Jagiellonian University, Krakow, Poland 16 Hematology Department - Fundeni Clinical Institute, University of Medicine and Pharmacy "Carol Davila", Bucharest, Romania 17 Department of Clinical Medicine and Surgery, University "Federico II" of Naples, Naples, Italy 18 CEINGE Biotecnologie Avanzate, Naples, Italy 19 III. Medizinische Klinik, Universitätsmedizin Mannheim, Mannheim, Germany 20 Abteilung Hämatologie/Onkologie, Klinik für Innere Medizin II, Universitätsklinikum Jena, Jena, Germany Correspondence to: Professor Nicholas C.P. Cross Wessex Regional Genetics Laboratory Salisbury District Hospital Salisbury SP2 8BJ, UK Tel Fax ncpc@soton.ac.uk 2
4 Abstract Treatment of chronic myeloid leukemia (CML) with tyrosine kinase inhibitors has advanced to a stage where many patients achieve very low or undetectable levels of disease. Remarkably, some of these patients remain in sustained remission when treatment is withdrawn, suggesting that they may be at least operationally cured of their disease. Accurate definition of deep molecular responses (MR) is therefore increasingly important for optimal patient management and comparison of independent data sets. We previously published proposals for broad standardized definitions of MR at different levels of sensitivity. Here we present detailed laboratory recommendations, developed as part of the European Treatment and Outcome Study (EUTOS) for CML, to enable testing laboratories to score MR in a reproducible manner for CML patients expressing the most common BCR-ABL1 variants. Introduction Molecular monitoring provides important prognostic information for individual chronic myeloid leukemia (CML) patients undergoing therapy, and international treatment recommendations incorporate specific time-dependent molecular milestones to help determine whether a patient is responding optimally or not. 1, 2 Molecular measurements are made by reverse transcriptase quantitative polymerase chain reaction (RT-qPCR) to estimate the amount of BCR-ABL1 mrna relative to an internal reference gene, most commonly ABL1, GUSB, or BCR. 3, 4 The results are expressed on an International Scale (IS) as a percentage, with 100% BCR-ABL IS corresponding to the International Randomized Study of Interferon and STI571 (IRIS) study standardized baseline 3
5 and 0.1% BCR-ABL IS being defined as a major molecular response (MMR or MR 3 ; 3 log reduction from the standardized baseline). 3 Expression of results on the IS depends on each testing laboratory either having obtained a laboratory-specific conversion factor (CF) by sample exchange with an established reference laboratory, or by using kits and reagents that have been calibrated to the World Health Organisation International Genetic Reference Panel for quantitation of BCR-ABL1 mrna. 4-9 Efforts to standardize molecular monitoring to the IS focused initially on detectable residual disease and in particular whether a patient had or had not achieved particular milestones, for example 10% BCR-ABL IS or 0.1% BCR-ABL IS at various time points. However with longer follow up, it became apparent that many patients treated with imatinib achieved deeper levels of response, with BCR-ABL1 becoming undetectable in a minority of cases. 10 This, along with the fact that second generation TKIs produce faster and deeper responses compared to imatinib, 11, 12 prompted the need for robust, standardized definitions of deep molecular response (MR). Such definitions are particularly important in the context of studies that are enrolling patients 13, 14 with sustained deep responses into treatment-free protocols. We previously published proposals for broad standardized definitions of MR at different levels of sensitivity (MR 4, MR 4.5 etc; collectively referred to as deep MR ) which were endorsed by the European LeukemiaNet in their most recent recommendations for treatment of CML patients. 1, 15 These broad definitions, however, and clinical studies that have been published to date, do 4
6 not provide the technical details and interpretation to enable laboratories to categorize patients in a standardized fashion. As part of the European Treatment and Outcome Study (EUTOS) we have developed laboratory proposals, as detailed below, to enable testing laboratories to define MR in a reproducible manner. These proposals were developed by consensus over several meetings and are described in detail in this paper, along with several examples. The terminology employed is based on the recommendations of the Minimum Information for Publication of Quantitative Real-Time PCR Experiments (MIQE) guidelines 16 and the proposal focuses on qpcr assays for the most common BCR-ABL1 variants (e13a2 and/or e14a2; 97% of CML patients) that use an external plasmid calibrator to estimate numbers of target molecules. Reference genes other than ABL1 The published definitions of MR focus on the use of ABL1 as a reference gene since this is used by the majority of laboratories worldwide. 15 Of the principal alternative reference genes, 3 GUSB is used by a significant minority of European laboratories whereas BCR is used primarily in Australasia and some US laboratories. We have focused here on extending the MR definitions when BCR-ABL1 is undetectable to include GUSB; further work will be required to extend these definitions to include BCR. To determine the correspondence between ABL1 and GUSB, we collected data from three centres that routinely analysed the expression of both genes in parallel. We focused on CML 5
7 samples that were <10% BCR-ABL IS and had >10,000 ABL1 copies. Of 1567 samples, the median ratio of GUSB/ABL1 was 2.4 in the same volume of cdna and therefore we consider that, for the purpose of defining deep MR, 10,000 ABL1 transcripts are equivalent to 24,000 GUSB transcripts. The previously published 15 definitions of MR can therefore be expanded as follows: MR 4 ( 4-log reduction from IRIS baseline) = either (i) detectable disease 0.01% BCR- ABL IS or (ii) undetectable disease in cdna with 10,000-31,999 ABL1 transcripts or 24,000-76,999 GUSB transcripts MR 4.5 ( 4.5-log reduction from IRIS baseline) = either (i) detectable disease % BCR-ABL IS or (ii) undetectable disease in cdna with 32,000-99,999 ABL1 transcripts or 77, ,999 GUSB transcripts MR 5 ( 5-log reduction from IRIS baseline) = either (i) detectable disease 0.001% BCR- ABL IS or (ii) undetectable disease in cdna with 100,000 ABL1 transcripts 240,000 GUSB transcripts Although GUSB laboratories may use these definitions, we suggest that they should ideally derive their own correspondence between ABL1 and GUSB (or other reference gene) using at least remission (<10% BCR-ABL IS ) samples to derive their own cut offs for different MR levels. Prior to making this comparison, the amplification conditions should be optimized and in particular the amplification efficiency for the two genes should be the same. This can be achieved easily for ABL1, GUSB and BCR (and BCR-ABL1) using the ERM-AD623 plasmid. 17 For 6
8 laboratory developed tests we further recommend that ERM-AD623 is used directly as a qpcr calibrator for routine analysis or indirectly as a calibrator for in house plasmid dilutions. Defining detectable and undetectable disease There are several ways in which testing laboratories differ in how they define disease as detectable or undetectable. For individual amplification reactions and runs, we recommend that the established Europe Against Cancer (EAC) criteria are used 18. In particular: The cut-off for positivity should correspond to a quantification cycle (Cq) of intercept + 1 (which should generally lead to cut-offs of Cq). In other words, samples with a Cq higher than intercept + 1 should be considered as undetectable. The no template control wells and reagent blanks should ideally not cross the threshold at any point but should certainly be at least 2 Cq above the intercept Cq for that run. If this is not the case then the run must be considered as failed. A major variable between centres is the number of replicate assays that are performed for each sample and the way that those replicates are considered to yield the final result. Typically, both BCR-ABL1 and the reference gene are tested in duplicate, although some centres perform triplicate assays and some only perform single assays. If replicate assays are performed for BCR- ABL1 (as recommended from RNA 19, 20 or cdna 21 to help improve the accuracy of results) and any of the individual replicates are positive according to the criteria above, we recommend that 7
9 the final result is considered as positive, i.e. detectable disease. Even when testing in triplicate and two replicates are scored as undetectable and one is scored as detectable, the overall result should be scored as detectable or positive. The EAC defines assay sensitivity by using normalized copy number and ΔΔCt methods, both of which relate the level of MRD to pretreatment levels for individual patients. 22 This is not compatible with the IS in CML, which relates MRD levels to the IRIS standardized baseline, and therefore an alternative approach is required. Scoring MR when disease is detectable In general, measurable residual disease 23 should be assigned a value on the IS and scored as MR 4 if 0.01% BCR-ABL IS, MR 4.5 if % BCR-ABL IS etc, provided that the sample fulfils minimum quality criteria, i.e ABL1 10,000 or GUSB 24,000 in each replicate. 21 If replicate analyses are performed and the values between replicates are comparable 21 then the number of BCR-ABL1 and reference gene transcripts should be the total value across replicates and the final result expressed on the IS, i.e. [(sum of BCR-ABL1 copies)/(sum of reference gene copies)] x conversion factor x 100 (see examples 1-3). Since the reference gene in this context is used to estimate the amount of cdna tested for BCR-ABL1, any difference in the number of replicates performed for BCR-ABL1 and the reference gene will need to be taken into account, (see example 4). In addition, we recommend that for scoring MR 4.5, the total reference gene number should be 32,000-99,999 ABL1 transcripts or 77, ,999 GUSB transcripts regardless of 8
10 whether the disease is detectable or undetectable. For scoring MR 5 the total reference gene number should be ABL1 100,000 or GUSB 240,000 (Table 1; see example 5). Many centers score positive samples with a Cq higher than that of the lowest plasmid standard as low level positive, positive outside the quantifiable range (POQR), <10 BCR-ABL1 (if the lowest standard is 10), <4 BCR-ABL1 (if the lowest standard is 4, etc). Indeed some guidelines specifically recommend that values should not be estimated if they require extrapolation beyond the span of the standard plasmid calibration curve. 21 This presents a problem for scoring low levels of disease and means, for example, that a laboratory using 10 as the lowest standard and a CF = 1 would need to achieve ABL1 reference gene values of 100,000 or greater to be able to score a sample with low level detectable disease as MR 4 and a value of 320,000 to score a similar sample as MR 4.5 (<10 BCR-ABL1/320,000 ABL1 = % BCR-ABL1). Despite the significant errors in quantifying small numbers of target molecules, we suggest that all level positive replicates should be assigned a specific number of BCR-ABL1 transcripts by extrapolating below the lowest plasmid standard. Testing laboratories have generally not rigorously determined their in house limit of detection (LoD; defined as the lowest concentration of target that can be detected with 95% confidence) for BCR-ABL1 transcripts. One reason for this is that standardized reagents have not been available to perform LoD analysis in a reproducible manner. Now, with the availability of the ERM-AD623 plasmid 17 and other calibration reagents 8, we recommend that laboratories 9
11 specifically measure 24 and optimize their BCR-ABL1 LoD. Clearly, the accuracy and precision by which MR can be scored critically depends on the BCR-ABL1 LoD being maximized. A laboratory with a poor LoD may score samples as undetectable and deep MR, whereas a laboratory with an optimized LoD may detect BCR-ABL1 in the same sample and score it as not deep MR. Several studies have indicated that qpcr can be routinely optimized to detect single target molecules Assuming that a single BCR-ABL1 cdna target can be detected and that there is no background signal (Limit of Blank = 0), the LoD is given by the Poisson distribution as 3 BCR- ABL1 targets, since for a sample with an average of 3 targets/unit volume, there is a 95% chance that any unit volume will contain at least one target (Figure 1). Thus, we recommend that any replicate scored as positive should be assigned a value of 3 BCR-ABL1 copies, i.e. positive replicates with estimated copy numbers of <3 should be scored as 3 (see examples 6-8). Alternative technologies, e.g. digital PCR, are likely to be more accurate than qpcr for estimating small numbers of target molecules and may well become the method of choice for 28, 29 more accurate definition of low level positive disease. Scoring MR when disease is undetectable Analysis of multiple replicates can increase the sensitivity of detection simply by increasing the amount of sample that is tested. This approach has been used to design very sensitive assays to detect BCR-ABL1 by qrt-pcr in healthy individuals, 30 for genomic DNA based tests for BCR- ABL1 in CML 31-33, for detection of minimal residual disease in lymphoid disorders 25 and for other applications such as non-invasive prenatal testing 34. When BCR-ABL1 is undetectable in all 10
12 replicates from the same sample we recommend that the final result is given as [undetectable BCR-ABL1]/[sum of reference gene in all the replicates]. We suggest that for routine analysis a maximum of two or three replicates are performed (examples 9-11) although for specific studies it may be desirable to perform more replicates. Stringent quality criteria are essential, specifically replicates with <10,000 ABL1 or <24,000 GUSB transcripts should be considered as inevaluable for determining deep MR (examples 12 and 13), and laboratories should maximize their LoD for BCR-ABL1 to avoid false negative results. As above, any difference in the number of replicates performed for BCR-ABL1 and the reference gene should be taken into account (example 14). Examples (i) BCR-ABL1 detected in at least one replicate Example 1 (Lab conversion factor = 0.8): - BCR-ABL1 replicate 1: detectable in 2μl cdna, estimated 7 copies - BCR-ABL1 replicate 2: detectable in 2μl cdna, estimated 3 copies - ABL1 replicate 1: 24,000 copies in 2μl cdna - ABL1 replicate 2: 28,000 copies in 2μl cdna Result = (sum BCR-ABL1 = 10)/(sum ABL1 = 52,000) x0.8 x100 = 11
13 0.015% = MMR but not MR 4 Example 2 (Lab conversion factor = 1.8): - BCR-ABL1 replicate 1: undetectable in 5μl cdna - BCR-ABL1 replicate 2: detectable in 5μl cdna, estimated 3 copies - GUSB replicate 1: 43,000 copies in 5μl cdna - GUSB replicate 2: 49,000 copies in 5μl cdna Result = (sum BCR-ABL1 = 3)/(sum GUSB = 92,000) x1.8 x100 = % = MR 4 Comment: Testing laboratories use different amounts of RNA to make cdna, make different volumes of cdna and use different volumes of cdna for individual qpcr assays. The number of reference gene transcripts should be estimated in the same volume of cdna used to test for BCR-ABL1. The use of other reference genes, e.g. BCR, is possible but the numbers of transcripts required to define different levels of MR remain to be determined. Example 3 (Lab conversion factor = 0.5): - BCR-ABL1 replicate 1: undetectable in 5μl cdna - BCR-ABL1 replicate 2: detectable in 5μl cdna, estimated 3 copies - ABL1 replicate 1: 9,000 copies in 5μl cdna 12
14 - ABL1 replicate 2: 8,000 copies in 5μl cdna Result = inevaluable for MR Comment: Although the [(sum of BCR-ABL1)/(sum of reference gene)] x conversion factor x 100 is less than 0.01%, the sample should be considered as inevaluable for assessment of MR as the ABL1 copy number in each replicate is <10,000. Example 4 (Lab conversion factor = 0.8): - BCR-ABL1 replicate 1: undetectable in 5μl cdna - BCR-ABL1 replicate 2: undetectable in 5μl cdna - BCR-ABL1 replicate 3: detectable in 5μl cdna, estimated 4 copies - ABL1 replicate 1: 14,000 copies in 5μl cdna - ABL1 replicate 2: 15,000 copies in 5μl cdna Result = (sum BCR-ABL1 = 4)/(sum ABL1 = 29,000 x1.5) x0.8 x100 = % = MR 4 Comment: The sum of the reference gene copy number is multiplied by 1.5 (equivalent to multiplying the mean copy number x 3) because only two replicates were performed for the 13
15 reference gene whereas three replicates were performed for BCR-ABL1. In general we consider that it is better to perform the same number of replicates for BCR-ABL1 and the reference gene. Example 5 (Lab conversion factor = 0.25): - BCR-ABL1 replicate 1: undetectable in 2μl cdna - BCR-ABL1 replicate 2: detectable in 2μl cdna, estimated 3 copies - ABL1 replicate 1: 12,000 copies in 2μl cdna - ABL1 replicate 2: 14,000 copies in 2μl cdna Result = (sum BCR-ABL1 = 3)/(sum ABL1 = 26,000) x0.25 x100 = %; sum of ABL1 <32,000 = MR 4 Comment: Although the [(sum of BCR-ABL1)/(sum of reference gene)] x conversion factor x 100 is <0.0032%, the total ABL1 value is less than 32,000 and should thus be considered as MR 4. Considering the extreme examples of 31,999 ABL1 transcripts and either 0 or 3 BCR-ABL1 transcripts, it is apparent that this discrepancy only arises if the CF is <0.35 if using ABL1 as a reference gene (or <0.82 if using GUSB). Example 6 (Lab conversion factor = 0.8): - BCR-ABL1 replicate 1: undetectable in 5μl cdna - BCR-ABL1 replicate 2: detectable in 5μl cdna, estimated 2 copies 14
16 - ABL1 replicate 1: 18,000 copies in 5μl cdna - ABL1 replicate 2: 16,500 copies in 5μl cdna Result = (sum BCR-ABL1 = 3)/(sum ABL1 = 34,500) x0.8 x100 = 0.007% = MR 4 Comment: Each positive replicate should be assigned a value of 3 copies and therefore the second BCR-ABL1 replicate is scored as 3 copies. Example 7 (Lab conversion factor = 0.8): - BCR-ABL1 replicate 1: detectable in 5μl cdna, estimated 2 copies - BCR-ABL1 replicate 2: detectable in 5μl cdna, estimated 1 copy - ABL1 replicate 1: 18,000 copies in 5μl cdna - ABL1 replicate 2: 16,500 copies in 5μl cdna Result = (sum BCR-ABL1 = 6)/(sum ABL1 = 34,500) x0.8 x100 = 0.014% = MMR but not MR 4 Comment: Each positive replicate should be assigned a value of 3 copies and therefore each BCR-ABL1 replicate is scored as 3 copies. 15
17 Example 8 (Lab conversion factor = 0.8): - BCR-ABL1 replicate 1: detectable in 5μl cdna, estimated 2 copies - BCR-ABL1 replicate 2: detectable in 5μl cdna, estimated 5 copies - BCR-ABL1 replicate 3: detectable in 5μl cdna, estimated 7 copies - ABL1 replicate 1: 34,000 copies in 5μl cdna - ABL1 replicate 2: 38,500 copies in 5μl cdna - ABL1 replicate 3: 32,500 copies in 5μl cdna Result = (sum BCR-ABL1 = 15)/(sum ABL1 = 105,000) x0.8 x100 = 0.011% = MMR but not MR 4 Comment: Each positive replicate should be assigned a value of 3 copies. (ii) BCR-ABL1 undetected in all replicates Example 9: - BCR-ABL1 replicate 1: undetectable in 5μl cdna - BCR-ABL1 replicate 2: undetectable in 5μl cdna - ABL1 replicate 1: 16,500 copies in 5μl cdna - ABL1 replicate 2: 18,000 copies in 5μl cdna 16
18 Result = undetectable BCR-ABL1 in 34,500 ABL1 = MR 4.5 Example 10: - BCR-ABL1 single analysis: undetectable in 5μl cdna - ABL1 single analysis: 45,000 copies in 5μl cdna Result = undetectable BCR-ABL1 in 45,000 ABL1 = MR 4.5 Comment: although single analyses are performed by some centres, replicate assays from RNA or cdna improves the accuracy of results Example 11: - BCR-ABL1 replicate 1: undetectable in 2μl cdna - BCR-ABL1 replicate 2: undetectable in 2μl cdna - BCR-ABL1 replicate 3: undetectable in 2μl cdna - ABL1 replicate 1: 24,000 copies in 2μl cdna - ABL1 replicate 2: 22,500 copies in 2μl cdna - ABL1 replicate 3: 24,000 copies in 2μl cdna Result = undetectable BCR-ABL1 in 70,500 ABL1 = MR
19 Example 12: - BCR-ABL1 replicate 1: undetectable in 5μl cdna - BCR-ABL1 replicate 2: undetectable in 5μl cdna - ABL1 replicate 1: 7,000 copies in 5μl cdna - ABL1 replicate 2: 8,000 copies in 5μl cdna Result = inevaluable for MR as ABL1 <10,000 in each replicate Example 13: - BCR-ABL1 replicate 1: undetectable in 5μl cdna - BCR-ABL1 replicate 2: undetectable in 5μl cdna - ABL1 replicate 1: 6,000 copies in 5μl cdna - ABL1 replicate 2: 14,000 copies in 5μl cdna Result = inevaluable for MR Comment: One replicate is <10,000 ABL1 and hence the sample should be considered as inevaluable for MR. Since the two ABL1 replicates are discordant the reference gene qpcr could be repeated. Example 14: 18
20 - BCR-ABL1 replicate 1: undetectable in 2μl cdna - BCR-ABL1 replicate 2: undetectable in 2μl cdna - BCR-ABL1 replicate 3: undetectable in 2μl cdna - ABL1 replicate 1: 16,500 copies in 2μl cdna - ABL1 replicate 2: 18,000 copies in 2μl cdna Result = undetectable BCR-ABL1 in (34,500 x1.5 = 51,750 ABL1) = MR 4.5 Comment: The sum of the reference gene copy number is multiplied by 1.5 because only two replicates were performed for the reference gene whereas three replicates were performed for BCR-ABL1. Concluding remarks The remarkable progress in the treatment of CML has demanded definitions of deep MR that are stretching the technology of molecular monitoring to its limit. The recommendations described here are an attempt to develop standardized laboratory approaches that strike a reasonable balance between scientific accuracy and clinical reality. It should be recognized that there is considerable inherent uncertainty in defining very low levels of disease and that it will be important to continue to look at trends over time in order to recognize sustained MR. Furthermore, the reproducible application of the recommendations depends critically on the ability of testing laboratories to measure absolute numbers of reference gene transcripts in a 19
21 comparable manner, as well as their ability to maximize the LoD for BCR-ABL1 and minimize inter-assay variability. It is obvious that future methodological improvements that increase the amount of sample tested (as determined by the number of reference gene transcripts) will increase the precision and accuracy of scoring MR 4 or MR 4.5, as well as enabling even deeper levels of MR to be determined. We recognize that these recommendations may need to be adapted to local requirements and changing technologies. We also recognize that laboratory recommendations in isolation are meaningless and that the critical question is the clinical significance of achieving deep levels of MR. We anticipate that the standardized definitions described here will help to progress clinical studies that aim ultimately to cure CML as well as providing a common framework for reporting routine results. CONFLICT OF INTEREST None of the authors have a relevant conflict of interest. This work was supported by the European LeukemiaNet via the European Treatment and Outcome Study (EUTOS) for CML. EUTOS is funded by Novartis but the recommendations described in this paper were developed, finalised and written entirely by the authors. 20
22 ACKNOWLEDGEMENT We would like to thank all the EUTOS laboratory members and other individuals who have contributed to the discussions that have led to the development of these recommendations. GM was supported by NGS-PTL FP7 European Project, and ER regional project. FIGURE LEGEND LoD of BCR-ABL1 detection. The graph shows the Poisson distribution with a mean of 3 BCR- ABL1 targets per well. The percentage of wells with 0 10 targets per well is indicated (20,000 computer generated random datapoints; Minitab version 16, Coventry, UK) and shows that 95% of wells has at least one BCR-ABL1 target. Since the LoD is defined as the lowest concentration of target that can be detected with 95% confidence, the maximal theoretical LOD is 3 copies. 21
23 References 1. Baccarani M, Deininger MW, Rosti G, Hochhaus A, Soverini S, Apperley JF, et al. European LeukemiaNet recommendations for the management of chronic myeloid leukemia: Blood 2013; 122: O'Brien S, Radich JP, Abboud CN, Akhtari M, Altman JK, Berman E, et al. Chronic myelogenous leukemia, version J Natl Compr Canc Netw 2014; 12: Hughes T, Deininger M, Hochhaus A, Branford S, Radich J, Kaeda J, et al. Monitoring CML patients responding to treatment with tyrosine kinase inhibitors: review and recommendations for harmonizing current methodology for detecting BCR-ABL transcripts and kinase domain mutations and for expressing results. Blood 2006; 108: Cross NC. Standardisation of molecular monitoring for chronic myeloid leukaemia. Best Pract Res Clin Haematol 2009; 22: Branford S, Cross NC, Hochhaus A, Radich J, Saglio G, Kaeda J, et al. Rationale for the recommendations for harmonizing current methodology for detecting BCR-ABL transcripts in patients with chronic myeloid leukaemia. Leukemia 2006; 20:
24 6. Branford S, Fletcher L, Cross NC, Muller MC, Hochhaus A, Kim DW, et al. Desirable performance characteristics for BCR-ABL measurement on an international reporting scale to allow consistent interpretation of individual patient response and comparison of response rates between clinical trials. Blood 2008; 112: White HE, Matejtschuk P, Rigsby P, Gabert J, Lin F, Lynn Wang Y, et al. Establishment of the first World Health Organization International Genetic Reference Panel for quantitation of BCR-ABL mrna. Blood 2010; 116: e White HE, Hedges J, Bendit I, Branford S, Colomer D, Hochhaus A, et al. Establishment and validation of analytical reference panels for the standardization of quantitative BCR- ABL1 measurements on the international scale. Clin Chem 2013; 59: Maute C, Nibourel O, Rea D, Coiteux V, Grardel N, Preudhomme C, et al. Calibration of BCR-ABL1 mrna quantification methods using genetic reference materials is a valid strategy to report results on the international scale. Clin Biochem 2014; 47: Hehlmann R, Grimwade D, Simonsson B, Apperley J, Baccarani M, Barbui T, et al. The European LeukemiaNet: achievements and perspectives. Haematologica 2011; 96:
25 11. Saglio G, Kim DW, Issaragrisil S, le Coutre P, Etienne G, Lobo C, et al. Nilotinib versus imatinib for newly diagnosed chronic myeloid leukemia. N Engl J Med 2010; 362: Kantarjian H, Shah NP, Hochhaus A, Cortes J, Shah S, Ayala M, et al. Dasatinib versus imatinib in newly diagnosed chronic-phase chronic myeloid leukemia. N Engl J Med 2010; 362: Mahon FX, Rea D, Guilhot J, Guilhot F, Huguet F, Nicolini F, et al. Discontinuation of imatinib in patients with chronic myeloid leukaemia who have maintained complete molecular remission for at least 2 years: the prospective, multicentre Stop Imatinib (STIM) trial. Lancet Oncol 2010; 11: Rousselot P, Charbonnier A, Cony-Makhoul P, Agape P, Nicolini FE, Varet B, et al. Loss of major molecular response as a trigger for restarting tyrosine kinase inhibitor therapy in patients with chronic-phase chronic myelogenous leukemia who have stopped imatinib after durable undetectable disease. J Clin Oncol 2014; 32: Cross NC, White HE, Muller MC, Saglio G, Hochhaus A. Standardized definitions of molecular response in chronic myeloid leukemia. Leukemia 2012; 26:
26 16. Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 2009; 55: White H, Deprez L, Corbisier P, Hall V, Lin F, al. e. A certified plasmid reference material for the standardisation of BCR-ABL1 mrna quantification by eal time quantitative PCR Leukemia 2014; in press. 18. Gabert J, Beillard E, van der Velden VH, Bi W, Grimwade D, Pallisgaard N, et al. Standardization and quality control studies of 'real-time' quantitative reverse transcriptase polymerase chain reaction of fusion gene transcripts for residual disease detection in leukemia - a Europe Against Cancer program. Leukemia 2003; 17: Branford S, Hughes T. Diagnosis and monitoring of chronic myeloid leukemia by qualitative and quantitative RT-PCR. Methods Mol Med 2006; 125: Jennings LJ, Smith FA, Halling KC, Persons DL, Kamel-Reid S, Molecular Oncology Resource Committee of the College of American P. Design and analytic validation of 25
27 BCR-ABL1 quantitative reverse transcription polymerase chain reaction assay for monitoring minimal residual disease. Arch Pathol Lab Med 2012; 136: Foroni L, Wilson G, Gerrard G, Mason J, Grimwade D, White HE, et al. Guidelines for the measurement of BCR-ABL1 transcripts in chronic myeloid leukaemia. Br J Haematol 2011; 153: Beillard E, Pallisgaard N, van der Velden VH, Bi W, Dee R, van der Schoot E, et al. Evaluation of candidate control genes for diagnosis and residual disease detection in leukemic patients using 'real-time' quantitative reverse-transcriptase polymerase chain reaction (RQ-PCR) - a Europe against cancer program. Leukemia 2003; 17: Goldman JM, Gale RP. What does MRD in leukemia really mean? Leukemia 2014; 28: CLSI. Evaluation of Detection Capability for Clinical Laboratory Measurement Procedures; Approved Guideline - 2nd Edition (EP17-A2). Wayne, PA (USA): Clinical and Laboratory Standards Institute.;
28 25. Morley AA, Latham S, Brisco MJ, Sykes PJ, Sutton R, Hughes E, et al. Sensitive and specific measurement of minimal residual disease in acute lymphoblastic leukemia. J Mol Diagn 2009; 11: Ladetto M, Lobetti-Bodoni C, Mantoan B, Ceccarelli M, Boccomini C, Genuardi E, et al. Persistence of minimal residual disease in bone marrow predicts outcome in follicular lymphomas treated with a rituximab-intensive program. Blood 2013; 122: Morley AA. Digital PCR: A brief history. Biomolecular Detection and Quantification 2014; 1: Goh HG, Lin M, Fukushima T, Saglio G, Kim D, Choi SY, et al. Sensitive quantitation of minimal residual disease in chronic myeloid leukemia using nanofluidic digital polymerase chain reaction assay. Leuk Lymphoma 2011; 52: Jennings LJ, George D, Czech J, Yu M, Joseph L. Detection and quantification of BCR-ABL1 fusion transcripts by droplet digital PCR. J Mol Diagn 2014; 16: Bose S, Deininger M, Gora-Tybor J, Goldman JM, Melo JV. The presence of typical and atypical BCR-ABL fusion genes in leukocytes of normal individuals: biologic significance 27
29 and implications for the assessment of minimal residual disease. Blood 1998; 92: Sobrinho-Simoes M, Wilczek V, Score J, Cross NC, Apperley JF, Melo JV. In search of the original leukemic clone in chronic myeloid leukemia patients in complete molecular remission after stem cell transplantation or imatinib. Blood 2010; 116: Ross DM, Branford S, Seymour JF, Schwarer AP, Arthur C, Bartley PA, et al. Patients with chronic myeloid leukemia who maintain a complete molecular response after stopping imatinib treatment have evidence of persistent leukemia by DNA PCR. Leukemia 2010; 24: Bartley PA, Ross DM, Latham S, Martin-Harris MH, Budgen B, Wilczek V, et al. Sensitive detection and quantification of minimal residual disease in chronic myeloid leukaemia using nested quantitative PCR for BCR-ABL DNA. Int J Lab Hematol 2010; 32: e Hromadnikova I, Houbova B, Hridelova D, Voslarova S, Kofer J, Komrska V, et al. Replicate real-time PCR testing of DNA in maternal plasma increases the sensitivity of non-invasive fetal sex determination. Prenat Diagn 2003; 23:
30 Table: Summary of reference gene numbers required for scoring deep molecular response MR 4 MR 4.5 MR 5 Minimum sum of reference gene transcripts irrespective of whether BCR-ABL1 is detected or not a 10,000 ABL1 24,000 GUSB 32,000 ABL1 77,000 GUSB 100,000 ABL1 240,000 GUSB BCR-ABL IS level for positive samples b 0.01% % 0.001% a numbers of reference gene transcripts in same volume of cdna that is tested for BCR-ABL1. The minimum number in any individual replicate should be 10,000 ABL1 or 24,000 GUSB. b provided that the minimum reference gene copy numbers in the row above are fulfilled
31
Impact of Age on Efficacy and Toxicity of Nilotinib in Patients With Chronic Myeloid Leukemia in Chronic Phase (CML-CP): ENEST1st Sub-Analysis
 Impact of Age on Efficacy and Toxicity of Nilotinib in Patients With Chronic Myeloid Leukemia in Chronic Phase (CML-CP): ENEST1st Sub-Analysis Abstract 479 Giles FJ, Rea D, Baccarani M, Cross NCP, Steegmann
Impact of Age on Efficacy and Toxicity of Nilotinib in Patients With Chronic Myeloid Leukemia in Chronic Phase (CML-CP): ENEST1st Sub-Analysis Abstract 479 Giles FJ, Rea D, Baccarani M, Cross NCP, Steegmann
Milestones and Monitoring
 Curr Hematol Malig Rep (2015) 10:167 172 DOI 10.1007/s11899-015-0258-1 CHRONIC MYELOID LEUKEMIAS (E JABBOUR, SECTION EDITOR) Milestones and Monitoring Alessandro Morotti 1 & Carmen Fava 1 & Giuseppe Saglio
Curr Hematol Malig Rep (2015) 10:167 172 DOI 10.1007/s11899-015-0258-1 CHRONIC MYELOID LEUKEMIAS (E JABBOUR, SECTION EDITOR) Milestones and Monitoring Alessandro Morotti 1 & Carmen Fava 1 & Giuseppe Saglio
MRD in CML (BCR-ABL1)
 MRD in CML (BCR-ABL1) Moleculaire Biologie en Cytometrie cursus Barbara Denys LAbo Hematologie UZ Gent 6 mei 2011 2008 Universitair Ziekenhuis Gent 1 Myeloproliferative Neoplasms o WHO classification 2008:
MRD in CML (BCR-ABL1) Moleculaire Biologie en Cytometrie cursus Barbara Denys LAbo Hematologie UZ Gent 6 mei 2011 2008 Universitair Ziekenhuis Gent 1 Myeloproliferative Neoplasms o WHO classification 2008:
Molecular Detection of BCR/ABL1 for the Diagnosis and Monitoring of CML
 Molecular Detection of BCR/ABL1 for the Diagnosis and Monitoring of CML Imran Mirza, MD, MS, FRCPC Pathology & Laboratory Medicine Institute Sheikh Khalifa Medical City, Abu Dhabi, UAE. imirza@skmc.ae
Molecular Detection of BCR/ABL1 for the Diagnosis and Monitoring of CML Imran Mirza, MD, MS, FRCPC Pathology & Laboratory Medicine Institute Sheikh Khalifa Medical City, Abu Dhabi, UAE. imirza@skmc.ae
Chronic Myeloid Leukaemia
 Chronic Myeloid Leukaemia Molecular Response: What is really important? Jeff Szer The Royal Melbourne Hospital PROBABILITY, % PROBABILITY OF SURVIVAL AFTER MYELOABLATIVE TRANSPLANTS FOR CML IN CHRONIC
Chronic Myeloid Leukaemia Molecular Response: What is really important? Jeff Szer The Royal Melbourne Hospital PROBABILITY, % PROBABILITY OF SURVIVAL AFTER MYELOABLATIVE TRANSPLANTS FOR CML IN CHRONIC
Molecular monitoring in chronic myeloid leukemia how low can you go?
 CHRONIC MYELOID LEUKEMIA Molecular monitoring in chronic myeloid leukemia how low can you go? Susan Branford Department of Genetics and Molecular Pathology, Centre for Cancer Biology, SA Pathology, Adelaide,
CHRONIC MYELOID LEUKEMIA Molecular monitoring in chronic myeloid leukemia how low can you go? Susan Branford Department of Genetics and Molecular Pathology, Centre for Cancer Biology, SA Pathology, Adelaide,
The concept of TFR (Treatment Free Remission) in CML
 The concept of TFR (Treatment Free Remission) in CML Giuseppe Saglio University of Turin, Italy What can we expect today on long-term therapy with TKIs in CML? German CML study IV Relative and overall
The concept of TFR (Treatment Free Remission) in CML Giuseppe Saglio University of Turin, Italy What can we expect today on long-term therapy with TKIs in CML? German CML study IV Relative and overall
The BCR-ABL1 fusion. Epidemiology. At the center of advances in hematology and molecular medicine
 At the center of advances in hematology and molecular medicine Philadelphia chromosome-positive chronic myeloid leukemia Robert E. Richard MD PhD rrichard@uw.edu robert.richard@va.gov Philadelphia chromosome
At the center of advances in hematology and molecular medicine Philadelphia chromosome-positive chronic myeloid leukemia Robert E. Richard MD PhD rrichard@uw.edu robert.richard@va.gov Philadelphia chromosome
When to change therapy? Andreas Hochhaus Universitätsklinikum Jena, Germany
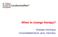 When to change therapy? Andreas Hochhaus Universitätsklinikum Jena, Germany Chromosome 22 Chromosome 9 e1 1b m-bcr M-bcr e1 e2 b1 b5 5 3 BCR ABL 5 3 1a a2 a3 μ -bcr e19 a11 e1a2 b2a2 b3a2 e19a2 p190 bcr-abl
When to change therapy? Andreas Hochhaus Universitätsklinikum Jena, Germany Chromosome 22 Chromosome 9 e1 1b m-bcr M-bcr e1 e2 b1 b5 5 3 BCR ABL 5 3 1a a2 a3 μ -bcr e19 a11 e1a2 b2a2 b3a2 e19a2 p190 bcr-abl
External quality assessment of p210 BCR-ABL1 transcript quantification by RT- qpcr: Findings and recommendations
 Received: 20 February 2018 Revised: 3 July 2018 Accepted: 17 July 2018 DOI: 10.1111/ijlh.12919 ORIGINAL ARTICLE External quality assessment of p210 BCR-ABL1 transcript quantification by RT- qpcr: Findings
Received: 20 February 2018 Revised: 3 July 2018 Accepted: 17 July 2018 DOI: 10.1111/ijlh.12919 ORIGINAL ARTICLE External quality assessment of p210 BCR-ABL1 transcript quantification by RT- qpcr: Findings
Oxford Style Debate on STOPPING Treatment.
 Oxford Style Debate on STOPPING Treatment. This house believes that there are good reasons NOT to stop CML treatment. It should be done within clinical trials, OR only in expert centers where frequent,
Oxford Style Debate on STOPPING Treatment. This house believes that there are good reasons NOT to stop CML treatment. It should be done within clinical trials, OR only in expert centers where frequent,
Deep molecular responses for treatment- free remission in chronic myeloid leukemia
 Cancer Medicine REVIEW Open Access Deep molecular responses for treatment- free remission in chronic myeloid leukemia Stéphanie Dulucq 1 & Francois-Xavier Mahon 2 1 Laboratoire d Hématologie, Centre Hospitalier
Cancer Medicine REVIEW Open Access Deep molecular responses for treatment- free remission in chronic myeloid leukemia Stéphanie Dulucq 1 & Francois-Xavier Mahon 2 1 Laboratoire d Hématologie, Centre Hospitalier
Molecular monitoring of CML patients
 EHA, Education Session, CML Stockholm, 14 June 2013 Molecular monitoring of CML patients Martin C. Müller Medical Faculty Mannheim Ruprecht-Karls-University Heidelberg Mannheim, Germany Disclosures Research
EHA, Education Session, CML Stockholm, 14 June 2013 Molecular monitoring of CML patients Martin C. Müller Medical Faculty Mannheim Ruprecht-Karls-University Heidelberg Mannheim, Germany Disclosures Research
Stopping Treatment in CML and dose reduction in clinical practice: Can we do it safely? YES WE CAN!
 Stopping Treatment in CML and dose reduction in clinical practice: Can we do it safely? YES WE CAN! Dragana Milojković The Hammersmith Hospital, London, UK Leukemic burden Current Aim of TKI therapy Molecular
Stopping Treatment in CML and dose reduction in clinical practice: Can we do it safely? YES WE CAN! Dragana Milojković The Hammersmith Hospital, London, UK Leukemic burden Current Aim of TKI therapy Molecular
Recent advances in the path toward the cure for chronic myeloid leukemia
 VOLUME 46 ㆍ NUMBER 3 ㆍ September 2011 THE KOREAN JOURNAL OF HEMATOLOGY REVIEW ARTICLE Recent advances in the path toward the cure for chronic myeloid leukemia Dong-Wook Kim Department of Hematology, Seoul
VOLUME 46 ㆍ NUMBER 3 ㆍ September 2011 THE KOREAN JOURNAL OF HEMATOLOGY REVIEW ARTICLE Recent advances in the path toward the cure for chronic myeloid leukemia Dong-Wook Kim Department of Hematology, Seoul
Molecular monitoring in CML and the prospects for treatment-free remissions
 MANAGING TYPICAL AND ATYPICAL CHRONIC MYELOID LEUKEMIA Molecular monitoring in CML and the prospects for treatment-free remissions Michael W. Deininger 1,2 1 Huntsman Cancer Institute, The University of
MANAGING TYPICAL AND ATYPICAL CHRONIC MYELOID LEUKEMIA Molecular monitoring in CML and the prospects for treatment-free remissions Michael W. Deininger 1,2 1 Huntsman Cancer Institute, The University of
EUROPEAN LEUKEMIANET RECOMMENDATIONS FOR CHRONIC MYELOID LEUKEMIA
 EUROPEAN LEUKEMIANET RECOMMENDATIONS FOR CHRONIC MYELOID LEUKEMIA SAN DIEGO, 11 DECEMBER 2011 AMSTERDAM, 14 JUNE 2012 BALTIMORE, 20 SEPTEMBER 2012 ATLANTA, 6 DECEMBER 2012 ELN, CML Panel Jane Apperley
EUROPEAN LEUKEMIANET RECOMMENDATIONS FOR CHRONIC MYELOID LEUKEMIA SAN DIEGO, 11 DECEMBER 2011 AMSTERDAM, 14 JUNE 2012 BALTIMORE, 20 SEPTEMBER 2012 ATLANTA, 6 DECEMBER 2012 ELN, CML Panel Jane Apperley
Treatment free remission in CML: from the concept to practice. François-Xavier Mahon. Cancer Center Bordeaux Université Bordeaux, France
 Treatment free remission in CML: from the concept to practice François-Xavier Mahon Cancer Center Bordeaux Université Bordeaux, France CML is a model 1960 1970 1980 1990 2000 2010 Philadelphia chromosome
Treatment free remission in CML: from the concept to practice François-Xavier Mahon Cancer Center Bordeaux Université Bordeaux, France CML is a model 1960 1970 1980 1990 2000 2010 Philadelphia chromosome
FEP Medical Policy Manual
 FEP Medical Policy Manual Effective Date: January 15, 2018 Related Policies: None BCR-ABL1 Testing in Chronic Myelogenous Leukemia and Description In the treatment of Philadelphia chromosome positive leukemias,
FEP Medical Policy Manual Effective Date: January 15, 2018 Related Policies: None BCR-ABL1 Testing in Chronic Myelogenous Leukemia and Description In the treatment of Philadelphia chromosome positive leukemias,
Stopping treatment how much we understand about mechanisms to stop successfully today, and where are the limits? Andreas Hochhaus
 Stopping treatment how much we understand about mechanisms to stop successfully today, and where are the limits? Andreas Hochhaus Frankfurt I 27.5.2017 Aims of CML Therapy Leukemia cells >10 12 CHR 10
Stopping treatment how much we understand about mechanisms to stop successfully today, and where are the limits? Andreas Hochhaus Frankfurt I 27.5.2017 Aims of CML Therapy Leukemia cells >10 12 CHR 10
Talpaz M. et al. Dasatinib in Imatinib-resistant Philadelphia chromosomepositive leukemias. N Engl J Med (2006) 354;24:
 References Sprycel Talpaz M. et al. Dasatinib in Imatinib-resistant Philadelphia chromosomepositive leukemias. N Engl J Med (2006) 354;24:2531-2541. National Comprehensive Cancer Network. Clinical Practice
References Sprycel Talpaz M. et al. Dasatinib in Imatinib-resistant Philadelphia chromosomepositive leukemias. N Engl J Med (2006) 354;24:2531-2541. National Comprehensive Cancer Network. Clinical Practice
Patient With Chronic Myeloid Leukemia in Complete Cytogenetic Response: What Does It Mean, and What Does One Do Next?
 VOLUME 32 NUMBER 5 FEBRUARY 10 2014 JOURNAL OF CLINICAL ONCOLOGY ONCOLOGY GRAND ROUNDS Patient With Chronic Myeloid Leukemia in Complete Cytogenetic Response: What Does It Mean, and What Does One Do Next?
VOLUME 32 NUMBER 5 FEBRUARY 10 2014 JOURNAL OF CLINICAL ONCOLOGY ONCOLOGY GRAND ROUNDS Patient With Chronic Myeloid Leukemia in Complete Cytogenetic Response: What Does It Mean, and What Does One Do Next?
Chronic Myeloid Leukemia A Disease of Young at Heart but Not of Body
 Chronic Myeloid Leukemia A Disease of Young at Heart but Not of Body Jeffrey H Lipton, PhD MD FRCPC Staff Physician, Princess Margaret Cancer Centre Professor of Medicine University of Toronto POGO November,
Chronic Myeloid Leukemia A Disease of Young at Heart but Not of Body Jeffrey H Lipton, PhD MD FRCPC Staff Physician, Princess Margaret Cancer Centre Professor of Medicine University of Toronto POGO November,
The significance of major and stable molecular responses in chronic myeloid leukemia in the tyrosine kinase inhibitor era
 Review Article The significance of major and stable molecular responses in chronic myeloid leukemia in the tyrosine kinase inhibitor era Ilana Zalcberg Renault 1 Vanesa Scholl 1 Rocio Hassan 1 Paola Capelleti
Review Article The significance of major and stable molecular responses in chronic myeloid leukemia in the tyrosine kinase inhibitor era Ilana Zalcberg Renault 1 Vanesa Scholl 1 Rocio Hassan 1 Paola Capelleti
La terapia della LMC: è possibile guarire senza trapianto? Fabrizio Pane
 La terapia della LMC: è possibile guarire senza trapianto? Fabrizio Pane What could be the concept of Cure in CML? Sustained DMR with or without TKI therapy And 100% CML-related survival And QoL comparable
La terapia della LMC: è possibile guarire senza trapianto? Fabrizio Pane What could be the concept of Cure in CML? Sustained DMR with or without TKI therapy And 100% CML-related survival And QoL comparable
FEP Medical Policy Manual
 FEP Medical Policy Manual Effective Date: January 15, 2019 Related Policies: None BCR-ABL1 Testing in Chronic Myelogenous Leukemia and Acute Description In the treatment of Philadelphia chromosome positive
FEP Medical Policy Manual Effective Date: January 15, 2019 Related Policies: None BCR-ABL1 Testing in Chronic Myelogenous Leukemia and Acute Description In the treatment of Philadelphia chromosome positive
CML: Role of combination treatments, Interferon and immunotherapy in CML
 CML: 2017 Role of combination treatments, Interferon and immunotherapy in CML Andreas Burchert Philipps Universität Marburg Universitätsklinikum Gießen und Marburg GmbH Key Developments that will Change
CML: 2017 Role of combination treatments, Interferon and immunotherapy in CML Andreas Burchert Philipps Universität Marburg Universitätsklinikum Gießen und Marburg GmbH Key Developments that will Change
Background. Linhartova et al. Molecular Cancer (2015) 14:89 DOI /s
 Linhartova et al. Molecular Cancer (2015) 14:89 DOI 10.1186/s12943-015-0363-8 LETTER TO THE EDITOR Open Access Characterization of 46 patient-specific BCR-ABL1 fusions and detection of SNPs upstream and
Linhartova et al. Molecular Cancer (2015) 14:89 DOI 10.1186/s12943-015-0363-8 LETTER TO THE EDITOR Open Access Characterization of 46 patient-specific BCR-ABL1 fusions and detection of SNPs upstream and
Development and evaluation of a secondary reference panel for BCR-ABL1 quantification on the International Scale
 Leukemia (2016) 30, 1844 1852 2016 Macmillan Publishers Limited, part of Springer Nature. All rights reserved 0887-6924/16 www.nature.com/leu OPEN ORIGINAL ARTICLE Development and evaluation of a secondary
Leukemia (2016) 30, 1844 1852 2016 Macmillan Publishers Limited, part of Springer Nature. All rights reserved 0887-6924/16 www.nature.com/leu OPEN ORIGINAL ARTICLE Development and evaluation of a secondary
Treatment free remission 2016
 Treatment free remission 2016 Pr Ph Rousselot Université de Versailles Saint-Quentin-en-Yvelines Hôpital André Mignot, Hôpitaux de Versailles, France How many patients still on imatinib? Versailles cohort
Treatment free remission 2016 Pr Ph Rousselot Université de Versailles Saint-Quentin-en-Yvelines Hôpital André Mignot, Hôpitaux de Versailles, France How many patients still on imatinib? Versailles cohort
Establishment and Validation of Analytical Reference Panels for the Standardization of Quantitative BCR-ABL1 Measurements on the International Scale
 Papers in Press. Published March 7, 23 as doi:.373/clinchem.22.96477 The latest version is at http://hwmaint.clinchem.org/cgi/doi/.373/clinchem.22.96477 Clinical Chemistry 59:6 (23) Molecular Diagnostics
Papers in Press. Published March 7, 23 as doi:.373/clinchem.22.96477 The latest version is at http://hwmaint.clinchem.org/cgi/doi/.373/clinchem.22.96477 Clinical Chemistry 59:6 (23) Molecular Diagnostics
Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+)
 Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+) Version: CMLBiomarkers 1.0.0.2 Protocol Posting Date: June 2017 This biomarker template is
Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+) Version: CMLBiomarkers 1.0.0.2 Protocol Posting Date: June 2017 This biomarker template is
CML CML CML. tyrosine kinase inhibitor CML. 22 t(9;22)(q34;q11) chronic myeloid leukemia CML ABL. BCR-ABL c- imatinib mesylate CML CML BCR-ABL
 1 Key Wordschronic myeloid leukemiaimatinib mesylate tyrosine kinase inhibitor chronic myeloid leukemia CML imatinib mesylate CML CML CML CML Ph 10 1 30 50 3 5 CML α IFNα Ph Ph cytogenetic response CRmajor
1 Key Wordschronic myeloid leukemiaimatinib mesylate tyrosine kinase inhibitor chronic myeloid leukemia CML imatinib mesylate CML CML CML CML Ph 10 1 30 50 3 5 CML α IFNα Ph Ph cytogenetic response CRmajor
Overview of CML related sessions at 21 th EHA Meeting in Copenhagen Preliminary Program
 Overview of CML related sessions at 21 th EHA Meeting in Copenhagen Preliminary Program Time slots Sessions Location June 9 th (Thursday) 8.00 10.00 Satellite Symposium: Improving outcomes: individualising
Overview of CML related sessions at 21 th EHA Meeting in Copenhagen Preliminary Program Time slots Sessions Location June 9 th (Thursday) 8.00 10.00 Satellite Symposium: Improving outcomes: individualising
BCR-ABL1 Kinase Domain Mutation Status (including educational clinical scenario)
 BCR-ABL1 Kinase Domain Mutation Status 171802 (including educational clinical scenario) Issue date: 23rd February 2018 Closing date: 23rd March 2018 Results for the BCR-ABL1 Kinase Domain Mutation Status
BCR-ABL1 Kinase Domain Mutation Status 171802 (including educational clinical scenario) Issue date: 23rd February 2018 Closing date: 23rd March 2018 Results for the BCR-ABL1 Kinase Domain Mutation Status
Blast Phase Chronic Myelogenous Leukemia
 Blast Phase Chronic Myelogenous Leukemia Benjamin Powers, MD; and Suman Kambhampati, MD The dramatic improvement in survival with tyrosine kinase inhibitors has not been demonstrated in the advanced blast
Blast Phase Chronic Myelogenous Leukemia Benjamin Powers, MD; and Suman Kambhampati, MD The dramatic improvement in survival with tyrosine kinase inhibitors has not been demonstrated in the advanced blast
RESEARCH ARTICLE. Introduction. Methods Wiley Periodicals, Inc.
 BCR/ABL level at 6 months identifies good risk CML subgroup after failing early molecular response at 3 months following imatinib therapy for CML in chronic phase AJH Dennis (Dong Hwan) Kim, 1 * Nada Hamad,
BCR/ABL level at 6 months identifies good risk CML subgroup after failing early molecular response at 3 months following imatinib therapy for CML in chronic phase AJH Dennis (Dong Hwan) Kim, 1 * Nada Hamad,
Research Article The Hasford Score May Predict Molecular Response in Chronic Myeloid Leukemia Patients: A Single Institution Experience
 Disease Markers Volume 216, Article ID 7531472, 5 pages http://dx.doi.org/1.1155/216/7531472 Research Article The Hasford Score May Predict Molecular Response in Chronic Myeloid Leukemia Patients: A Single
Disease Markers Volume 216, Article ID 7531472, 5 pages http://dx.doi.org/1.1155/216/7531472 Research Article The Hasford Score May Predict Molecular Response in Chronic Myeloid Leukemia Patients: A Single
"Molecular Monitoring Strategies to Track Response to BCR-ABL Kinase Inhibitors in CML"
 Association of Molecular Pathology USCAP Companion Meeting Sunday, February 12, 2006 7:00 PM Dan Jones, MD, PhD Associate Professor Medical Director, Molecular Diagnostic Laboratory Division of Pathology
Association of Molecular Pathology USCAP Companion Meeting Sunday, February 12, 2006 7:00 PM Dan Jones, MD, PhD Associate Professor Medical Director, Molecular Diagnostic Laboratory Division of Pathology
Radowan Elnair 1 and Ahmed Galal 2*
 Elnair and Galal BMC Cancer (2018) 18:1097 https://doi.org/10.1186/s12885-018-5004-3 CASE REPORT Open Access Finding the right BCR-ABL1 tyrosine kinase inhibitor: a case report of successful treatment
Elnair and Galal BMC Cancer (2018) 18:1097 https://doi.org/10.1186/s12885-018-5004-3 CASE REPORT Open Access Finding the right BCR-ABL1 tyrosine kinase inhibitor: a case report of successful treatment
I nuovi guariti? La malattia minima residua nella leucemia mieloide cronica. Fabrizio Pane
 I nuovi guariti? La malattia minima residua nella leucemia mieloide cronica Fabrizio Pane What could be the concept of Cure in CML? Sustained DMR with or without TKI therapy And 100% CML-related survival
I nuovi guariti? La malattia minima residua nella leucemia mieloide cronica Fabrizio Pane What could be the concept of Cure in CML? Sustained DMR with or without TKI therapy And 100% CML-related survival
Outlook CML 2016: What is being done on the way to cure
 New Horizons 2011 Outlook CML 2016: What is being done on the way to cure Gianantonio Rosti Dept. Of Hematology and Oncology St. Orsola-Malpighi University Hospital Bologna (Italy) GIMEMA CML Working Party
New Horizons 2011 Outlook CML 2016: What is being done on the way to cure Gianantonio Rosti Dept. Of Hematology and Oncology St. Orsola-Malpighi University Hospital Bologna (Italy) GIMEMA CML Working Party
NEW DRUGS IN HEMATOLOGY
 NEW DRUGS IN HEMATOLOGY BOLOGNA, 15-17 April 2013 TYROSINE KINASE INHIBITORS IN CHRONIC MYELOID LEUKEMIA MICHELE BACCARANI michele.baccarani@unibo.it Historic Development of CML Therapy Palliative Therapy
NEW DRUGS IN HEMATOLOGY BOLOGNA, 15-17 April 2013 TYROSINE KINASE INHIBITORS IN CHRONIC MYELOID LEUKEMIA MICHELE BACCARANI michele.baccarani@unibo.it Historic Development of CML Therapy Palliative Therapy
Achieving deeper molecular response is associated with a better clinical outcome in chronic myeloid leukemia patients on imatinib frontline therapy
 Published Ahead of Print on December 20, 2013, as doi:10.3324/haematol.2013.095158. Copyright 2013 Ferrata Storti Foundation. Achieving deeper molecular response is associated with a better clinical outcome
Published Ahead of Print on December 20, 2013, as doi:10.3324/haematol.2013.095158. Copyright 2013 Ferrata Storti Foundation. Achieving deeper molecular response is associated with a better clinical outcome
10 YEARS EXPERIENCE OF TYROSINE KINASE INHIBITOR THERAPY FOR CML IN OXFORD
 10 YEARS EXPERIENCE OF TYROSINE KINASE INHIBITOR THERAPY FOR CML IN OXFORD Dalia Khan 1, Noemi Roy 1, Vasha Bari 1, Grant Vallance 1, Helene Dreau 1, Timothy Littlewood 1, Andrew Peniket 1, Paresh Vyas
10 YEARS EXPERIENCE OF TYROSINE KINASE INHIBITOR THERAPY FOR CML IN OXFORD Dalia Khan 1, Noemi Roy 1, Vasha Bari 1, Grant Vallance 1, Helene Dreau 1, Timothy Littlewood 1, Andrew Peniket 1, Paresh Vyas
HOW I TREAT CML. 4. KONGRES HEMATOLOGOV IN TRANSFUZIOLOGOV SLOVENIJE Z MEDNARODNO UDELEŽBO Terme Olimia, Podčetrtek,
 HOW I TREAT CML 4. KONGRES HEMATOLOGOV IN TRANSFUZIOLOGOV SLOVENIJE Z MEDNARODNO UDELEŽBO Terme Olimia, Podčetrtek, 12. - 14. april, 2012 Gianantonio Rosti Dpt of Hematology and Oncological Sciences S.
HOW I TREAT CML 4. KONGRES HEMATOLOGOV IN TRANSFUZIOLOGOV SLOVENIJE Z MEDNARODNO UDELEŽBO Terme Olimia, Podčetrtek, 12. - 14. april, 2012 Gianantonio Rosti Dpt of Hematology and Oncological Sciences S.
ELN Recommendations on treatment choice and response. Gianantonio Rosti, MD, Department of Hematology, University of Bologna, Italy
 ELN Recommendations on treatment choice and response Gianantonio Rosti, MD, Department of Hematology, University of Bologna, Italy ELN 2013 Response to Front-line Treatment Baseline 3 months 6 months OPTIMAL
ELN Recommendations on treatment choice and response Gianantonio Rosti, MD, Department of Hematology, University of Bologna, Italy ELN 2013 Response to Front-line Treatment Baseline 3 months 6 months OPTIMAL
Molecular testing for the BCR-ABL1 fusion gene by
 Conversion, Correction, and International Scale Standardization Results From a Multicenter External Quality Assessment Study for BCR-ABL1 Testing Michael Griffiths, BSc; Simon J. Patton, PhD; Alberto Grossi,
Conversion, Correction, and International Scale Standardization Results From a Multicenter External Quality Assessment Study for BCR-ABL1 Testing Michael Griffiths, BSc; Simon J. Patton, PhD; Alberto Grossi,
Molecular Monitoring of Residual Disease in Chronic Myeloid Leukemia by Genomic DNA Compared with Conventional mrna Analysis
 Journal of Molecular Diagnostics, Vol. 11, No. 5, September 2009 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology DOI: 10.2353/jmoldx.2009.080150 Molecular
Journal of Molecular Diagnostics, Vol. 11, No. 5, September 2009 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology DOI: 10.2353/jmoldx.2009.080150 Molecular
RT-qPCR and RT-Digital PCR: A Comparison of Different Platforms for the Evaluation of Residual Disease in Chronic Myeloid Leukemia
 Papers in Press. Published December 15, 2016 as doi:10.1373/clinchem.2016.262824 The latest version is at http://hwmaint.clinchem.aaccjnls.org/cgi/doi/10.1373/clinchem.2016.262824 Clinical Chemistry 63:2
Papers in Press. Published December 15, 2016 as doi:10.1373/clinchem.2016.262824 The latest version is at http://hwmaint.clinchem.aaccjnls.org/cgi/doi/10.1373/clinchem.2016.262824 Clinical Chemistry 63:2
Greater Manchester and Cheshire Cancer Network Chronic Myeloid Leukaemia v3 2012
 Greater Manchester and Cheshire Cancer Network Chronic Myeloid Leukaemia v3 2012 Dr Simon Watt Dr Shiva Natarajan 1.0 Introduction The landscape in chronic myeloid leukaemia (CML) has changed dramatically
Greater Manchester and Cheshire Cancer Network Chronic Myeloid Leukaemia v3 2012 Dr Simon Watt Dr Shiva Natarajan 1.0 Introduction The landscape in chronic myeloid leukaemia (CML) has changed dramatically
MP BCR-ABL1 Testing in Chronic Myelogenous Leukemia and Acute Lymphoblastic Leukemia
 Medical Policy BCBSA Ref. Policy: 2.04.85 Last Review: 10/18/2018 Effective Date: 10/18/2018 Section: Medicine Related Policies 8.01.30 Hematopoietic Cell Transplantation for Chronic Myelogenous Leukemia
Medical Policy BCBSA Ref. Policy: 2.04.85 Last Review: 10/18/2018 Effective Date: 10/18/2018 Section: Medicine Related Policies 8.01.30 Hematopoietic Cell Transplantation for Chronic Myelogenous Leukemia
Study Design and Endpoints
 Complete Molecular Response (CMR) Rate With Nilotinib in Patients With CML-CP Without CMR After 2 Years on Imatinib: Preliminary Results From the Randomized ENESTcmr Trial Timothy P. Hughes, Jeffrey H.
Complete Molecular Response (CMR) Rate With Nilotinib in Patients With CML-CP Without CMR After 2 Years on Imatinib: Preliminary Results From the Randomized ENESTcmr Trial Timothy P. Hughes, Jeffrey H.
How I monitor residual disease in chronic myeloid leukemia
 CLINICAL TRIALS AND OBSERVATIONS How I treat How I monitor residual disease in chronic myeloid leukemia Jerald P. Radich 1 1 Clinical Research Division, Fred Hutchinson Cancer Research Center, Seattle,
CLINICAL TRIALS AND OBSERVATIONS How I treat How I monitor residual disease in chronic myeloid leukemia Jerald P. Radich 1 1 Clinical Research Division, Fred Hutchinson Cancer Research Center, Seattle,
Analytical and economic evaluation of the fully automated Xpert BCR-ABL assay from Cepheid
 Analytical and economic evaluation of the fully automated Xpert BCR-ABL assay from Cepheid Jean-Michel Cayuela Saint-Louis hospital Paris July 5, 2012 Talk outline Chronic myeloid leukemia > Genetic bases,
Analytical and economic evaluation of the fully automated Xpert BCR-ABL assay from Cepheid Jean-Michel Cayuela Saint-Louis hospital Paris July 5, 2012 Talk outline Chronic myeloid leukemia > Genetic bases,
Goals for chronic myeloid leukemia TK inhibitor treatment: how little disease is too much?
 MINIMAL RESIDUAL DISEASE:CHRONIC MYELOID LEUKEMIA AND ACUTE LYMPHOCYTIC LEUKEMIA Goals for chronic myeloid leukemia TK inhibitor treatment: how little disease is too much? Michael J. Mauro 1 1 Memorial
MINIMAL RESIDUAL DISEASE:CHRONIC MYELOID LEUKEMIA AND ACUTE LYMPHOCYTIC LEUKEMIA Goals for chronic myeloid leukemia TK inhibitor treatment: how little disease is too much? Michael J. Mauro 1 1 Memorial
Original Study. Abstract
 Original Study The Effectiveness of Tyrosine Kinase Inhibitors and Molecular Monitoring Patterns in Newly Diagnosed Patients With Chronic Myeloid Leukemia in the Community Setting Nicholas J. Di Bella,
Original Study The Effectiveness of Tyrosine Kinase Inhibitors and Molecular Monitoring Patterns in Newly Diagnosed Patients With Chronic Myeloid Leukemia in the Community Setting Nicholas J. Di Bella,
Imatinib dose intensification, combination therapies. Andreas Hochhaus Universitätsklinikum Jena, Germany
 Imatinib dose intensification, combination therapies Andreas Hochhaus Universitätsklinikum Jena, Germany Apperley JF. Lancet Oncol. 2007 High OCT-1 activity is associated with faster MMR in imatinib treated
Imatinib dose intensification, combination therapies Andreas Hochhaus Universitätsklinikum Jena, Germany Apperley JF. Lancet Oncol. 2007 High OCT-1 activity is associated with faster MMR in imatinib treated
How I treat high risck CML
 Torino, September 14, 2018 How I treat high risck CML Patrizia Pregno Hematology Dept. Citta della Salute e della Scienza Torino Disclosures Advisory Board: Novartis, Pfizer, Incyte Speaker Honoraria:
Torino, September 14, 2018 How I treat high risck CML Patrizia Pregno Hematology Dept. Citta della Salute e della Scienza Torino Disclosures Advisory Board: Novartis, Pfizer, Incyte Speaker Honoraria:
Dati sulla sospensione della terapia
 Leucemia Mieloide Cronica Dati sulla sospensione della terapia Gianantonio Rosti, Bologna BCR-ABL Loading in CML Patients 100% 10% 1% MMR MR 4 MR 4.5 STIM study design N=100 Start Imatinib CMR Sustained
Leucemia Mieloide Cronica Dati sulla sospensione della terapia Gianantonio Rosti, Bologna BCR-ABL Loading in CML Patients 100% 10% 1% MMR MR 4 MR 4.5 STIM study design N=100 Start Imatinib CMR Sustained
Chronic myeloid leukemia (CML)
 Bioscientia Int. Support Services Dept. Konrad-Adenauer-Strasse 17 55218 Ingelheim Germany Phone 49(0)6132-781-203 49(0)6132-781-224 49(0)6132-781-165 Fax 49(0)6132-781-236 int.support@bioscientia.com
Bioscientia Int. Support Services Dept. Konrad-Adenauer-Strasse 17 55218 Ingelheim Germany Phone 49(0)6132-781-203 49(0)6132-781-224 49(0)6132-781-165 Fax 49(0)6132-781-236 int.support@bioscientia.com
Kinetics of BCR-ABL Transcripts in Imatinib Mesylate treated Chronic Phase CML (CPCML), A Predictor of Response and Progression Free Survival
 International journal of Biomedical science ORIGINAL ARTICLE Kinetics of BCR-ABL Transcripts in Imatinib Mesylate treated Chronic Phase CML (CPCML), A Predictor of Response and Progression Free Survival
International journal of Biomedical science ORIGINAL ARTICLE Kinetics of BCR-ABL Transcripts in Imatinib Mesylate treated Chronic Phase CML (CPCML), A Predictor of Response and Progression Free Survival
Overview of CML related sessions at 23 rd EHA Meeting in Stockholm
 Overview of CML related sessions at 23 rd EHA Meeting in Stockholm Time slots Sessions Location June 14th (Thursday) 8.00 10.00 Satellite Symposium: Targeting the individual: Personalized treatment in
Overview of CML related sessions at 23 rd EHA Meeting in Stockholm Time slots Sessions Location June 14th (Thursday) 8.00 10.00 Satellite Symposium: Targeting the individual: Personalized treatment in
Chronic Myeloid Leukemia
 Chronic Myeloid Leukemia Session Chair: Armand Keating, MD Speakers: Timothy Hughes, MD, MBBS; Michael J. Mauro, MD; and Jane F. Apperley, MBChB ABL Kinase Inhibitor Therapy for CML: Baseline Assessments
Chronic Myeloid Leukemia Session Chair: Armand Keating, MD Speakers: Timothy Hughes, MD, MBBS; Michael J. Mauro, MD; and Jane F. Apperley, MBChB ABL Kinase Inhibitor Therapy for CML: Baseline Assessments
Implementation of Management Guidelines
 Implementation of Management Guidelines For Chronic Myeloid Leukemia Perspectives in the United States David Rizzieri, MD; and Joseph O. Moore, MD ABSTRACT Clinical practice guidelines are developed to
Implementation of Management Guidelines For Chronic Myeloid Leukemia Perspectives in the United States David Rizzieri, MD; and Joseph O. Moore, MD ABSTRACT Clinical practice guidelines are developed to
SESSION III: Chronic myeloid leukemia PONATINIB. Gianantonio Rosti, MD, Department of Hematology, University of Bologna, Italy
 SESSION III: Chronic myeloid leukemia PONATINIB Gianantonio Rosti, MD, Department of Hematology, University of Bologna, Italy Ponatinib A Pan-BCR-ABL Inhibitor Rationally designed inhibitor of BCR- ABL
SESSION III: Chronic myeloid leukemia PONATINIB Gianantonio Rosti, MD, Department of Hematology, University of Bologna, Italy Ponatinib A Pan-BCR-ABL Inhibitor Rationally designed inhibitor of BCR- ABL
CML and Future Perspective. Hani Al-Hashmi, MD
 CML and Future Perspective Hani Al-Hashmi, MD Objectives Learning from CML history Outcome of interest to clinician Patient and community interest!! Learning from CML history Survival Probability (All
CML and Future Perspective Hani Al-Hashmi, MD Objectives Learning from CML history Outcome of interest to clinician Patient and community interest!! Learning from CML history Survival Probability (All
Chronic myelogenous leukemia (CML) is a slowprogressing
 At a Glance Practical Implications p e148 Author Information p e151 Full text and PDF Web exclusive Patterns of Specific Testing for Patients With Chronic Myelogenous Leukemia Original Research Allison
At a Glance Practical Implications p e148 Author Information p e151 Full text and PDF Web exclusive Patterns of Specific Testing for Patients With Chronic Myelogenous Leukemia Original Research Allison
Guidelines and real World: Management of CML in chronic and advanced phases. Carolina Pavlovsky. FUNDALEU May 2017 Frankfurt
 Guidelines and real World: Management of CML in chronic and advanced phases Carolina Pavlovsky. FUNDALEU 26-28 May 217 Frankfurt Some Issues in CML 217 First Line treatment: Imatinib vs 2nd generation
Guidelines and real World: Management of CML in chronic and advanced phases Carolina Pavlovsky. FUNDALEU 26-28 May 217 Frankfurt Some Issues in CML 217 First Line treatment: Imatinib vs 2nd generation
Laboratory Tools for Diagnosis and Monitoring Response in Patients with Chronic Myeloid Leukemia
 in chromosome Laboratory Tools for Diagnosis and Monitoring Response in Patients with Chronic Myeloid Leukemia Tali Tohami PhD 1, Arnon Nagler MD 2,3 and Ninette Amariglio PhD 1 1 Hematology Laboratory
in chromosome Laboratory Tools for Diagnosis and Monitoring Response in Patients with Chronic Myeloid Leukemia Tali Tohami PhD 1, Arnon Nagler MD 2,3 and Ninette Amariglio PhD 1 1 Hematology Laboratory
Journal of Molecular Biology Research Vol. 1, No. 1; December 2011
 Evaluation of Molecular Response to Imatinib in Iraqi Chronic Myeloid Leukemia Patients Using Real Time Reveres Transcriptase-Polymerase Chain Reaction (RT-RT-PCR) Taqman Assay Maysaa Abdul Razzaq Dhahii
Evaluation of Molecular Response to Imatinib in Iraqi Chronic Myeloid Leukemia Patients Using Real Time Reveres Transcriptase-Polymerase Chain Reaction (RT-RT-PCR) Taqman Assay Maysaa Abdul Razzaq Dhahii
Low doses of tyrosine kinase inhibitors in CML
 CML Horizons Conference Warsaw 4-6 May 2018 Low doses of tyrosine kinase inhibitors in CML Delphine Rea, MD, PhD Pôle Hématologie Oncologie Radiothérapie INSERM UMR-1160 Centre Hospitalo-Universitaire
CML Horizons Conference Warsaw 4-6 May 2018 Low doses of tyrosine kinase inhibitors in CML Delphine Rea, MD, PhD Pôle Hématologie Oncologie Radiothérapie INSERM UMR-1160 Centre Hospitalo-Universitaire
Sequencing Treatment in Chronic Myeloid Leukemia: The First Choice May Be the Hardest
 Sequencing Treatment in Chronic Myeloid Leukemia: The First Choice May Be the Hardest Mark L. Heaney, MD, PhD Dr Heaney is an associate clinical professor of medicine at Columbia University Medical Center
Sequencing Treatment in Chronic Myeloid Leukemia: The First Choice May Be the Hardest Mark L. Heaney, MD, PhD Dr Heaney is an associate clinical professor of medicine at Columbia University Medical Center
Introduction NEOPLASIA
 NEOPLASIA Desirable performance characteristics for BCR-ABL measurement on an international reporting scale to allow consistent interpretation of individual patient response and comparison of response
NEOPLASIA Desirable performance characteristics for BCR-ABL measurement on an international reporting scale to allow consistent interpretation of individual patient response and comparison of response
1 Educational session salient points. Tim Hughes, Vivian Oehler and Rick Van Etten. Tim - Imatinib is a less toxic drug than what we are seeing with
 1 Educational session salient points. Tim Hughes, Vivian Oehler and Rick Van Etten. Tim - Imatinib is a less toxic drug than what we are seeing with both Nilotinib and Dasatinib. Nilotinib for the cardio
1 Educational session salient points. Tim Hughes, Vivian Oehler and Rick Van Etten. Tim - Imatinib is a less toxic drug than what we are seeing with both Nilotinib and Dasatinib. Nilotinib for the cardio
Stopping TKI s in CML- Are we There Yet? Joseph O. Moore, MD Duke Cancer Institute
 Stopping TKI s in CML- Are we There Yet? Joseph O. Moore, MD Duke Cancer Institute Natural History of CML Accumulation of immature myeloid cells New cytogenetic changes Chronic Phase Accelerated Phase
Stopping TKI s in CML- Are we There Yet? Joseph O. Moore, MD Duke Cancer Institute Natural History of CML Accumulation of immature myeloid cells New cytogenetic changes Chronic Phase Accelerated Phase
Clinical Utility of Droplet ddpcr, moving to diagnostics. Koen De Gelas, PhD, CRIG ddpcr mini symposium, 15/05/2018
 1 Clinical Utility of Droplet ddpcr, moving to diagnostics 2 Koen De Gelas, PhD, CRIG ddpcr mini symposium, 15/05/2018 Disclaimer Disclaimer: all consumables, instruments, applications and software covered
1 Clinical Utility of Droplet ddpcr, moving to diagnostics 2 Koen De Gelas, PhD, CRIG ddpcr mini symposium, 15/05/2018 Disclaimer Disclaimer: all consumables, instruments, applications and software covered
Starting & stopping therapy in Chronic Myeloid Leukemia: What more is needed? Richard A. Larson, MD University of Chicago March 2019
 Starting & stopping therapy in Chronic Myeloid Leukemia: What more is needed? Richard A. Larson, MD University of Chicago March 2019 Disclosures Richard A. Larson, MD Research funding to the University
Starting & stopping therapy in Chronic Myeloid Leukemia: What more is needed? Richard A. Larson, MD University of Chicago March 2019 Disclosures Richard A. Larson, MD Research funding to the University
Overview of CML related sessions at 22 nd EHA Meeting in Madrid
 Overview of CML related sessions at 22 nd EHA Meeting in Madrid Time slots Sessions Location June 22 nd (Thursday) 13.30 15.30 Satellite Symposium: Emerging trends in chronic myeloid leukemia and acute
Overview of CML related sessions at 22 nd EHA Meeting in Madrid Time slots Sessions Location June 22 nd (Thursday) 13.30 15.30 Satellite Symposium: Emerging trends in chronic myeloid leukemia and acute
-Glucuronidase Is an Optimal Normalization Control Gene for Molecular Monitoring of Chronic Myelogenous Leukemia
 JMD CME Program Journal of Molecular Diagnostics, Vol. 8, No. 3, July 2006 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology DOI: 10.2353/jmoldx.2006.050150
JMD CME Program Journal of Molecular Diagnostics, Vol. 8, No. 3, July 2006 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology DOI: 10.2353/jmoldx.2006.050150
journal of medicine The new england Long-Term Outcomes of Imatinib Treatment for Chronic Myeloid Leukemia abstract
 The new england journal of medicine established in 1812 March 9, 2017 vol. 376 no. 10 Long-Term Outcomes of Imatinib Treatment for Chronic Myeloid Leukemia Andreas Hochhaus, M.D., Richard A. Larson, M.D.,
The new england journal of medicine established in 1812 March 9, 2017 vol. 376 no. 10 Long-Term Outcomes of Imatinib Treatment for Chronic Myeloid Leukemia Andreas Hochhaus, M.D., Richard A. Larson, M.D.,
Ponatinib Withdrawal Update
 Hello. This is Dr. Stuart Goldberg from the Leukemia Division at the John Theurer Cancer Center at Hackensack University Medical Center in Hackensack, New Jersey. I am speaking on behalf of ManagingCML.com
Hello. This is Dr. Stuart Goldberg from the Leukemia Division at the John Theurer Cancer Center at Hackensack University Medical Center in Hackensack, New Jersey. I am speaking on behalf of ManagingCML.com
Treatment-free remission following frontline nilotinib in patients with chronic myeloid leukemia in chronic phase: results from the ENESTfreedom study
 OPEN Leukemia (2017) 31, 1525 1531 www.nature.com/leu ORIGINAL ARTICLE Treatment-free remission following frontline nilotinib in patients with chronic myeloid leukemia in chronic phase: results from the
OPEN Leukemia (2017) 31, 1525 1531 www.nature.com/leu ORIGINAL ARTICLE Treatment-free remission following frontline nilotinib in patients with chronic myeloid leukemia in chronic phase: results from the
Chronic myeloid leukemia: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up
 clinical practice guidelines Annals of Oncology 23 (Supplement 7): vii72 vii77, 2012 doi:10.1093/annonc/mds228 Chronic myeloid leukemia: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up
clinical practice guidelines Annals of Oncology 23 (Supplement 7): vii72 vii77, 2012 doi:10.1093/annonc/mds228 Chronic myeloid leukemia: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up
In this issue of the journal two articles deal with different
 EDITORIALS & PERSPECTIVES Monitoring treatment of chronic myeloid leukemia Michele Baccarani, 1 Fabrizio Pane, 2 and Giuseppe Saglio 3 1 Department of Hematology-Oncology "L. and A. Seràgnoli" University
EDITORIALS & PERSPECTIVES Monitoring treatment of chronic myeloid leukemia Michele Baccarani, 1 Fabrizio Pane, 2 and Giuseppe Saglio 3 1 Department of Hematology-Oncology "L. and A. Seràgnoli" University
EVI-1 oncogene expression predicts survival in chronic phase CML patients resistant to imatinib
 EVI-1 oncogene expression predicts survival in chronic phase CML patients resistant to imatinib Mustafa Daghistani Department of Haematology, Imperial College London at Hammersmith Hospital, Du Cane Road,
EVI-1 oncogene expression predicts survival in chronic phase CML patients resistant to imatinib Mustafa Daghistani Department of Haematology, Imperial College London at Hammersmith Hospital, Du Cane Road,
Tyrosine kinase inhibitors (TKIs) act by preventing disease
 SUBJECT REVIEW Emergence of BCR-ABL Kinase Domain Associated with Newly Diagnosed Chronic Myeloid Leukemia: A Meta-Analysis of Clinical Trials of Tyrosine Kinase Inhibitors Iulia D. Ursan, PharmD; Ruixuan
SUBJECT REVIEW Emergence of BCR-ABL Kinase Domain Associated with Newly Diagnosed Chronic Myeloid Leukemia: A Meta-Analysis of Clinical Trials of Tyrosine Kinase Inhibitors Iulia D. Ursan, PharmD; Ruixuan
CML: definition. CML epidemiology. CML diagnosis. CML: peripheralbloodsmear. Cytogenetic abnormality of CML
 MolecularDiagnostic.be Third Scientific Meeting Molecular Diagnostics.be t(9;22) CML: definition Management of CML patients treated with TKI: the place of molecular monitoring Antwerp, December 13 th 11
MolecularDiagnostic.be Third Scientific Meeting Molecular Diagnostics.be t(9;22) CML: definition Management of CML patients treated with TKI: the place of molecular monitoring Antwerp, December 13 th 11
Discontinuation of Imatinib in Patients with Chronic Myeloid Leukemia Who Have Maintained Complete Molecular Response: Update Results of the STIM
 Discontinuation of Imatinib in Patients with Chronic Myeloid Leukemia Who Have Maintained Complete Molecular Response: Update Results of the STIM François-Xavier Mahon, Delphine Réa, Joëlle Guilhot, François
Discontinuation of Imatinib in Patients with Chronic Myeloid Leukemia Who Have Maintained Complete Molecular Response: Update Results of the STIM François-Xavier Mahon, Delphine Réa, Joëlle Guilhot, François
Is there a best TKI for chronic phase CML?
 MANAGING TYPICAL AND ATYPICAL CHRONIC MYELOID LEUKEMIA Is there a best TKI for chronic phase CML? Richard A. Larson 1 1 Section of Hematology/Oncology, Department of Medicine, and Comprehensive Cancer
MANAGING TYPICAL AND ATYPICAL CHRONIC MYELOID LEUKEMIA Is there a best TKI for chronic phase CML? Richard A. Larson 1 1 Section of Hematology/Oncology, Department of Medicine, and Comprehensive Cancer
Defining therapy goals for major molecular remission in chronic myeloid leukemia: results of the randomized CML Study IV
 Leukemia (2018) 32:1222 1228 https://doi.org/10.1038/s41375-018-0055-7 ARTICLE Chronic myelogenous leukemia Defining therapy goals for major molecular remission in chronic myeloid leukemia: results of
Leukemia (2018) 32:1222 1228 https://doi.org/10.1038/s41375-018-0055-7 ARTICLE Chronic myelogenous leukemia Defining therapy goals for major molecular remission in chronic myeloid leukemia: results of
Philadelphia Positive (Ph+) Chronic Myeloid Leukaemia
 Advocacy toolkit In this toolkit, we take a look into the treatments for Philadelphia-positive Chronic Myeloid Leukaemia (CML) that have led to treatment free remission (TFR) being one of the biggest topics
Advocacy toolkit In this toolkit, we take a look into the treatments for Philadelphia-positive Chronic Myeloid Leukaemia (CML) that have led to treatment free remission (TFR) being one of the biggest topics
Update on Tyrosine Kinase Inhibitor Therapy for Chronic Myelogenous Leukemia
 Update on Tyrosine Kinase Inhibitor Therapy for Chronic Myelogenous Leukemia Chronic myelogenous leukemia (CML), a hematologic malignancy associated with a chromosomal mutation commonly known as the Philadelphia
Update on Tyrosine Kinase Inhibitor Therapy for Chronic Myelogenous Leukemia Chronic myelogenous leukemia (CML), a hematologic malignancy associated with a chromosomal mutation commonly known as the Philadelphia
Updated review of nilotinib as frontline treatment for newly diagnosed Philadelphia chromosome-positive chronic myeloid leukemia
 For reprint orders, please contact: reprints@futuremedicine.com Updated review of nilotinib as frontline treatment for newly diagnosed Philadelphia chromosome-positive chronic myeloid leukemia Nilotinib,
For reprint orders, please contact: reprints@futuremedicine.com Updated review of nilotinib as frontline treatment for newly diagnosed Philadelphia chromosome-positive chronic myeloid leukemia Nilotinib,
ipsogen BCR-ABL1 Mbcr IS-MMR Kit Handbook
 July 2016 ipsogen BCR-ABL1 Mbcr IS-MMR Kit Handbook 24 Version 1 Quantitative in vitro diagnostics For use with Rotor-Gene Q, Applied Biosystems, ABI PRISM, and LightCycler instruments 670723 QIAGEN GmbH,
July 2016 ipsogen BCR-ABL1 Mbcr IS-MMR Kit Handbook 24 Version 1 Quantitative in vitro diagnostics For use with Rotor-Gene Q, Applied Biosystems, ABI PRISM, and LightCycler instruments 670723 QIAGEN GmbH,
Dynamics of chronic myeloid leukemia response to dasatinib, nilotinib, and high-dose imatinib
 Chronic Myeloid Leukemia Dynamics of chronic myeloid leukemia response to dasatinib, nilotinib, and high-dose imatinib ARTICLES Adam Olshen, * Min Tang, * Jorge Cortes, Mithat Gonen, Timothy Hughes, Susan
Chronic Myeloid Leukemia Dynamics of chronic myeloid leukemia response to dasatinib, nilotinib, and high-dose imatinib ARTICLES Adam Olshen, * Min Tang, * Jorge Cortes, Mithat Gonen, Timothy Hughes, Susan
Second Tyrosine Kinase Inhibitor Discontinuation Attempt in Patients With Chronic Myeloid Leukemia
 Second Tyrosine Kinase Inhibitor Discontinuation Attempt in Patients With Chronic Myeloid Leukemia Laurence Legros, MD, PhD 1,2 ; Franck E. Nicolini, MD, PhD 3,4 ; Gabriel Etienne, MD, PhD 5 ; Philippe
Second Tyrosine Kinase Inhibitor Discontinuation Attempt in Patients With Chronic Myeloid Leukemia Laurence Legros, MD, PhD 1,2 ; Franck E. Nicolini, MD, PhD 3,4 ; Gabriel Etienne, MD, PhD 5 ; Philippe
Adecade ago imatinib mesylate, the first tyrosine
 CHRONIC MYELOID LEUKEMIA: AFTER A DECADE OF IMATIINIB Monitoring disease response to tyrosine kinase inhibitor therapy in CML Timothy P. Hughes 1 and Susan Branford 1 1 Centre for Cancer Biology, Departments
CHRONIC MYELOID LEUKEMIA: AFTER A DECADE OF IMATIINIB Monitoring disease response to tyrosine kinase inhibitor therapy in CML Timothy P. Hughes 1 and Susan Branford 1 1 Centre for Cancer Biology, Departments
Published Ahead of Print on August 31, 2017, as doi: /haematol Copyright 2017 Ferrata Storti Foundation.
 Published Ahead of Print on August 31, 2017, as doi:10.3324/haematol.2017.175265. Copyright 2017 Ferrata Storti Foundation. Impact of hospital experience on the quality of tyrosine kinase inhibitor response
Published Ahead of Print on August 31, 2017, as doi:10.3324/haematol.2017.175265. Copyright 2017 Ferrata Storti Foundation. Impact of hospital experience on the quality of tyrosine kinase inhibitor response
Tomonaga, Masao; Miyazaki, Yasushi. available at
 NAOSITE: Nagasaki University's Ac Title Author(s) Citation Successful treatment of a chronic-p myelogenous leukemia patient with a interferon-alfa Itonaga, Hidehiro; Tsushima, Hideki Imanishi, Daisuke;
NAOSITE: Nagasaki University's Ac Title Author(s) Citation Successful treatment of a chronic-p myelogenous leukemia patient with a interferon-alfa Itonaga, Hidehiro; Tsushima, Hideki Imanishi, Daisuke;
