SUPPLEMENTAY FIGURES AND TABLES
|
|
|
- Shavonne Harvey
- 6 years ago
- Views:
Transcription
1 SUPPLEMENTAY FIGURES AND TABLES Supplementary Figure S1: Validation of mir expression by quantitative real-time PCR and time course studies on mir- 29a-3p and mir-210-3p. A. The graphs illustrate the expression profiles of 8 selected mirs which were obtained with the use of mir microarrays (Array) and quantitative real-time PCR (qpcr). The numbers in red indicate the r 2 correlation values obtained after comparison of the microarray and qpcr results. Each qpcr value is the mean of 3 biological replicates ± SD. B. The panel shows the time-course of mir-29a-3p up-regulation and mir-210-3p down-regulation by ATRA and/or lapatinib. The results were obtained by qpcr analysis. Each result is the mean of 3 replicates ± SD. *Significantly different (p < 0.05, Student s t-test) **Significantly different (p < 0.01, Student s t-test).
2 Supplementary Figure S2: Flow chart of mir and target-mrna integrated analysis. Two active datasets from SKBR3 cells were used for the analysis: 1) mirs selected for significant changes in expression by one of the factors in the described microarray experiments (ATRA, Lapatinib, or interaction, 36 h treatment; two-way ANOVA, p < 0.01); 2) mrnas selected in the same way as in 1) from previously obtained microarray data (E-MEXP-3192; (48 hours; p < 0.001). The MAGIA (MiRNA- And-Genes-Integrated-Analysis) web-tool ( was used to predict mir/target mrna interactions in these two datasets. The steps for the analysis were: 1) identification of putative mir/target-mrna pairs by any one of three prediction algorithms (PITA, miranda, TargetScan; filters set as default); 2) for each of the putative mir/target-mrna pairs, selection of those showing a significant negative correlation between mir and mrna-expression (Spearman correlation < 0.9, q-value <0.05).
3 Supplementary Figure S3: Expression of Module-1, Module-2 and Module-3 mir mimics in transfected SKBR3 cells. RNA extracted from SKBR3 cells transfected with 30 nm of the indicated mir mimics or scrambled negative control (Neg) were analyzed after 48 hours by qpcr. Specific Taqman assays were used to quantify the levels of the indicated mirs in both Neg and mir-mimictransfected cells. Values represent the mean ± SD of 3 transfections. A. The panel shows the results for selected Module-1 (mir-188-5p) and Module-3 (mir-575 and mir p) mirs. B. The panel shows the results for selected Module-2 mirs.
4 Supplementary Table S1. mir profiles in SKBR3 cells following treatment with ATRA and/or lapatinib. The table lists 174 mirs whose expression is significantly altered by treatment of SKBR3 cells with 100 nm ATRA, 100 nm Lapatinib or 100 nm of both ATRA and lapatinib for 36 hours. Columns 3 6 indicate the mir expression values measured in the 4 experimental conditions. The data are expressed in linear intensity values and they correspond to the mean of 5 biological replicates. Columns 7 10 report the p-values obtained after 2-way ANOVA as indicated. Supplementary Table S2. Predicted mir/target-mrna interactions. The table lists the mir/targetmrna interactions predicted by the MAGIA algorithm with a Spearman correlation value < 0.9 and a q-value <0.05. The number in the first column indicates the module (Module-1 to -4) mirs belong to. The 0 symbol is used for mirs that are not organized in any module. Supplementary Table S3. mir and target-mrna nodes in Module1-4. The table lists all the mirs and target-mrnas belonging to the four most interconnected modules (Module-1 to -4). Node = mir or target-mrna; Degree = degree of connectivity.
5 Supplementary Table S4. Breast cancer cell lines characteristics. The Table lists the characteristics of the indicated cell lines. The IC 50 and IC 25 values (concentrations causing a 50% and 25% reduction in the number of cells) are indicated for ATRA, Lapatinib (Lap) and doxorubicin (Doxo). While the data for ATRA and Doxo were determined experimentally, the Lap data are extracted from the following article: Konecny, G.E., M.D. Pegram, N. Venkatesan, R. Finn, G. Yang, M. Rahmeh, M. Untch, D.W. Rusnak, G. Spehar, R.J. Mullin, B.R. Keith, T.M. Gilmer, M. Berger, et al. Activity of the dual kinase inhibitor lapatinib (GW572016) against HER-2-overexpressing and trastuzumab-treated breast cancer cells. Cancer Res, 2006; 66: p The breast cancer cell lines were used to evaluate the effects exerted by the three compounds on the expression of mir-125a-5p, mir-193a-5p and mir-210-3p belonging to Module-2. The same cell lines were also used to determine the effects of the three mirs on cell growth, apoptosis and motility. TN = triple-negative; ER = Estrogen receptor. Cell line Phenotype ER HER2 ATRA Lap IC 50 (nm) Doxo IC 25 (nm) IC 50 (nm) MDA-MB231 Basal/TN 3, , MDA-MB157 Basal/TN 37 6, MDA-MB-453 Luminal + > 10, 000 3, MCF-7 Luminal , SKBR3 Luminal
6 Supplementary Table S5. Literature review for the mirs present in Module 1 4. The table lists the references reporting on the biological action of the mirs present in Module-1 to -4. Column 3 lists the mir target-mrnas involved.
7 Supplementary Table S6. Amplimers and probes. The table contains the list of the amplimers and probes used for the Taqman assays along with the list of the mir mimics used throughout the study. mir qpcr assays Source Reference Code hsa-mir-29a-3p Exiqon hsa-mir-125a-5p Life Technologies hsa-mir-141-3p Exiqon hsa-mir-188-5p Life Technologies hsa-mir-193a-5p Life Technologies hsa-mir-203 Life Technologies hsa-mir-210-3p Life Technologies has mir-425-3p Exiqon hsa-mir-575 Life Technologies hsa-mir-720 Exiqon Z-30 Life Technologies Gene expression qpcr assays HIPK2 Life Technologies Hs _m1 PLCXD1 Life Technologies Hs _m1 RPLP0 Life Technologies Hs _m1 Actin Life Technologies Hs _m1 MiR mimics hsa-mir-29a-3p Life Technologies PM12499 hsa-mir-125a-5p Life Technologies PM12561 hsa-mir-193a-5p Life Technologies PM11786 hsa-mir-188-5p Life Technologies PM12963 hsa-mir-210-3p Life Technologies PM10516 hsa-mir-575 Life Technologies PM11506 hsa-mir-874-3p Life Technologies PM12355 hsa-mir p Life Technologies PM13447 Negative Control #1 Life Technologies AM17110
(A) Dose response curves of HMLE_shGFP (blue circle), HMLE_shEcad (red square),
 Supplementary Figures and Tables Figure S1. Validation of EMT-selective small molecules (A) Dose response curves of HMLE_shGFP (blue circle), HMLE_shEcad (red square), and HMLE_Twist (black diamond) cells
Supplementary Figures and Tables Figure S1. Validation of EMT-selective small molecules (A) Dose response curves of HMLE_shGFP (blue circle), HMLE_shEcad (red square), and HMLE_Twist (black diamond) cells
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figures Supplementary Figure 1 DOT1L regulates the expression of epithelial and mesenchymal markers. (a) The expression levels and cellular localizations of EMT markers were confirmed by
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
TEB. Id4 p63 DAPI Merge. Id4 CK8 DAPI Merge
 a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
a Duct TEB b Id4 p63 DAPI Merge Id4 CK8 DAPI Merge c d e Supplementary Figure 1. Identification of Id4-positive MECs and characterization of the Comma-D model. (a) IHC analysis of ID4 expression in the
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
microrna-200b and microrna-200c promote colorectal cancer cell proliferation via
 Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
Supplementary Materials microrna-200b and microrna-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs Supplementary Table 1.
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
www.sciencesignaling.org/cgi/content/full/7/310/ra11/dc1 Supplementary Materials for STAT3 Induction of mir-146b Forms a Feedback Loop to Inhibit the NF-κB to IL-6 Signaling Axis and STAT3-Driven Cancer
RNA preparation from extracted paraffin cores:
 Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Supplementary Information
 Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplementary Information Supplementary Figure 1. Effect of mir mimics and anti-mirs on DTPs a, Representative fluorescence microscopy images of GFP vector control or mir mimicexpressing parental and DTP
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/406/ra126/dc1 Supplementary Materials for The microrna mir-485 targets host and influenza virus transcripts to regulate antiviral immunity and restrict viral
www.sciencesignaling.org/cgi/content/full/8/406/ra126/dc1 Supplementary Materials for The microrna mir-485 targets host and influenza virus transcripts to regulate antiviral immunity and restrict viral
Supplementary materials and methods.
 Supplementary materials and methods. Microarray printing and data analysis. The LNA-modified oligonucleotide probe set for all annotated mirnas from mouse (Mus musculus) and human (Homo sapiens) in the
Supplementary materials and methods. Microarray printing and data analysis. The LNA-modified oligonucleotide probe set for all annotated mirnas from mouse (Mus musculus) and human (Homo sapiens) in the
SUPPLEMENTARY INFORMATION. Supp. Fig. 1. Autoimmunity. Tolerance APC APC. T cell. T cell. doi: /nature06253 ICOS ICOS TCR CD28 TCR CD28
 Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
Supp. Fig. 1 a APC b APC ICOS ICOS TCR CD28 mir P TCR CD28 P T cell Tolerance Roquin WT SG Icos mrna T cell Autoimmunity Roquin M199R SG Icos mrna www.nature.com/nature 1 Supp. Fig. 2 CD4 + CD44 low CD4
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplemental Data. TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim.
 Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
Supplemental Data TGF-β-mediated mir-181a expression promotes breast cancer metastasis by targeting Bim. Molly A. Taylor 1, Khalid Sossey-Alaoui 2, Cheryl L. Thompson 3, David Danielpour 4, and William
The clinical relevance of circulating, cell-free and exosomal micrornas as biomarkers for gynecological tumors
 Department of Tumor Biology The clinical relevance of circulating, cell-free and exosomal micrornas as biomarkers for gynecological tumors cfdna Copenhagen April 6-7, 2017 Heidi Schwarzenbach, PhD Tumor
Department of Tumor Biology The clinical relevance of circulating, cell-free and exosomal micrornas as biomarkers for gynecological tumors cfdna Copenhagen April 6-7, 2017 Heidi Schwarzenbach, PhD Tumor
SUPPLEMENTARY FIGURES AND TABLES
 SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
SUPPLEMENTARY FIGURES AND TABLES Supplementary Figure S1: CaSR expression in neuroblastoma models. A. Proteins were isolated from three neuroblastoma cell lines and from the liver metastasis of a MYCN-non
RmiR package vignette
 RmiR package vignette Francesco Favero * October 17, 2016 Contents 1 Introduction 1 2 Coupling mirna and gene expression data 2 2.1 Default parameters.............................. 3 2.2 Change the at.least
RmiR package vignette Francesco Favero * October 17, 2016 Contents 1 Introduction 1 2 Coupling mirna and gene expression data 2 2.1 Default parameters.............................. 3 2.2 Change the at.least
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Discovery Dataset. PD Liver Luminal B/ Her-2+ Letrozole. PD Supraclavicular Lymph node. PD Supraclavicular Lymph node Luminal B.
 Discovery Dataset 11T pt1cpn2am1(liver) 2009 2010 Liver / Her-2+ 2011 Death Letrozole CHT pt1cpn0(sn)m0 Supraclavicular Lymph node Death 12T 2003 2006 2006 2008 Anastrozole Local RT+Examestane Fulvestrant
Discovery Dataset 11T pt1cpn2am1(liver) 2009 2010 Liver / Her-2+ 2011 Death Letrozole CHT pt1cpn0(sn)m0 Supraclavicular Lymph node Death 12T 2003 2006 2006 2008 Anastrozole Local RT+Examestane Fulvestrant
Supplementary webappendix
 Supplementary webappendix This webappendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Ueda T, Volinia S, Okumura H, et al.
Supplementary webappendix This webappendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Ueda T, Volinia S, Okumura H, et al.
Inhibition of TGFβ enhances chemotherapy action against triple negative breast cancer by abrogation of
 SUPPLEMENTAL DATA Inhibition of TGFβ enhances chemotherapy action against triple negative breast cancer by abrogation of cancer stem cells and interleukin-8 Neil E. Bhola 1, Justin M. Balko 1, Teresa C.
SUPPLEMENTAL DATA Inhibition of TGFβ enhances chemotherapy action against triple negative breast cancer by abrogation of cancer stem cells and interleukin-8 Neil E. Bhola 1, Justin M. Balko 1, Teresa C.
HIF-inducible mir-191 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment.
 HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
HIF-inducible mir-9 promotes migration in breast cancer through complex regulation of TGFβ-signaling in hypoxic microenvironment. Neha Nagpal, Hafiz M Ahmad, Shibu Chameettachal3, Durai Sundar, Sourabh
developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success
 micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients.
 Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Supplementary Materials Supplementary Figures Figure S1. Reduction in glomerular mir-146a levels correlate with progression to higher albuminuria in diabetic patients. Figure S2. Expression level of podocyte
Yong Wu, Ph.D. Division of Cancer Research and Training (DCRT) Charles R. Drew University of Medicine & Science
 Yong Wu, Ph.D. Division of Cancer Research and Training (DCRT) Charles R. Drew University of Medicine & Science Jay Vadgama, Ph.D Chief, Division of Cancer Research and Training Background One in 8 women
Yong Wu, Ph.D. Division of Cancer Research and Training (DCRT) Charles R. Drew University of Medicine & Science Jay Vadgama, Ph.D Chief, Division of Cancer Research and Training Background One in 8 women
Synthetic microrna Reference Standards Genomics Research Group ABRF 2015
 Synthetic microrna Reference Standards Genomics Research Group ABRF 2015 Don A. Baldwin, Ph.D. support@signalbiology.com Reference samples for Platform evaluation Protocol development Assay service improvement
Synthetic microrna Reference Standards Genomics Research Group ABRF 2015 Don A. Baldwin, Ph.D. support@signalbiology.com Reference samples for Platform evaluation Protocol development Assay service improvement
Profiles of gene expression & diagnosis/prognosis of cancer. MCs in Advanced Genetics Ainoa Planas Riverola
 Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Supplementary Figure 1
 Supplementary Figure 1 14 12 SEM4C PLXN2 8 SEM4C C 3 Cancer Cell Non Cancer Cell Expression 1 8 6 6 4 log2 ratio Expression 2 1 4 2 2 p value.1 D Supplementary Figure 1. Expression of Sema4C and Plexin2
Supplementary Figure 1 14 12 SEM4C PLXN2 8 SEM4C C 3 Cancer Cell Non Cancer Cell Expression 1 8 6 6 4 log2 ratio Expression 2 1 4 2 2 p value.1 D Supplementary Figure 1. Expression of Sema4C and Plexin2
Supplementary Figure 1
 Supplementary Figure 1 Constitutive EGFR signaling does not activate canonical EGFR signals (a) U251EGFRInd cells with or without tetracycline exposure (24h, 1µg/ml) were treated with EGF for 15 minutes
Supplementary Figure 1 Constitutive EGFR signaling does not activate canonical EGFR signals (a) U251EGFRInd cells with or without tetracycline exposure (24h, 1µg/ml) were treated with EGF for 15 minutes
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/364/ra18/dc1 Supplementary Materials for The tyrosine phosphatase (Pez) inhibits metastasis by altering protein trafficking Leila Belle, Naveid Ali, Ana Lonic,
www.sciencesignaling.org/cgi/content/full/8/364/ra18/dc1 Supplementary Materials for The tyrosine phosphatase (Pez) inhibits metastasis by altering protein trafficking Leila Belle, Naveid Ali, Ana Lonic,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities
 Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
UNIVERSITA DEGLI STUDI DI TORINO. Department of Oncological Sciences. Ph.D. in Complex Systems applied to Post-Genomic Biology.
 UNIVERSITA DEGLI STUDI DI TORINO Department of Oncological Sciences Ph.D. in Complex Systems applied to Post-Genomic Biology XXIII Cycle mir-148b is a major coordinator in a Relapse-associated mir signature
UNIVERSITA DEGLI STUDI DI TORINO Department of Oncological Sciences Ph.D. in Complex Systems applied to Post-Genomic Biology XXIII Cycle mir-148b is a major coordinator in a Relapse-associated mir signature
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the
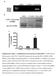 Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was
 Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
Supplementary Figure 1 IMQ-Induced Mouse Model of Psoriasis. IMQ cream was painted on the shaved back skin of CBL/J and BALB/c mice for consecutive days. (a, b) Phenotypic presentation of mouse back skin
ANALYTISCHE STRATEGIE Tissue Imaging. Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich
 ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
mir-630 targets IGF1R to regulate response to HER-targeting drugs and overall cancer cell progression in HER2 overexpressing
 mir-630 targets IGF1R to regulate response to HER-targeting drugs and overall cancer cell progression in HER2 overexpressing breast cancer The Harvard community has made this article openly available.
mir-630 targets IGF1R to regulate response to HER-targeting drugs and overall cancer cell progression in HER2 overexpressing breast cancer The Harvard community has made this article openly available.
Benchmark study: Exiqon mircury LNA microrna arrays vs. Supplier A s DNA based capture probes. Includes comparison of:
 Benchmark study: Exiqon mircury LNA microrna arrays vs. Supplier A s DNA based capture probes Includes comparison of: I. II. III. Signal levels Relative quantification False differences For complete experimental
Benchmark study: Exiqon mircury LNA microrna arrays vs. Supplier A s DNA based capture probes Includes comparison of: I. II. III. Signal levels Relative quantification False differences For complete experimental
Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2
 Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2 activity. Upper panel: Representative histograms for FACS
Supplementary Fig. 1: ATM is phosphorylated in HER2 breast cancer cell lines. (A) ATM is phosphorylated in SKBR3 cells depending on ATM and HER2 activity. Upper panel: Representative histograms for FACS
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
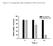 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
doi:10.1038/nature12215 Supplementary Figure 1. The effects of full and dissociated GR agonists in supporting BFU-E self-renewal divisions. BFU-Es were cultured in self-renewal medium with indicated GR
Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the. mutated sequence.
 Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the mutated sequence. 1 Supplementary Figure 2. Expression of mir-182 and SMAD7 in various cell lines. (A) Basal levels of mir-182 expression
Supplementary Figure 1. The mir-182 binding site of SMAD7 3 UTR and the mutated sequence. 1 Supplementary Figure 2. Expression of mir-182 and SMAD7 in various cell lines. (A) Basal levels of mir-182 expression
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting
 mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
mir-509-5p and mir-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer Hidekazu Hiramoto, M.D. 1,3, Tomoki Muramatsu, Ph.D. 1, Daisuke Ichikawa,
Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.
 Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.5, E18.5, P4, and P8. Values shown are means from four technical
Supplemental Figure S1. PLAG1 kidneys contain fewer glomeruli (A) Quantitative PCR for Igf2 and PLAG1 in whole kidneys taken from mice at E15.5, E18.5, P4, and P8. Values shown are means from four technical
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
www.sciencesignaling.org/cgi/content/full/2/1/ra81/dc1 Supplementary Materials for Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12- Dependent Vascular Protection Alma Zernecke,* Kiril Bidzhekov,
Supplementary Figure 1. Characterization of ALDH-positive cell population in MCF-7 cells. (a) Expression level of stem cell markers in MCF-7 cells or
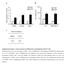 Supplementary Figure 1. Characterization of ALDH-positive cell population in MCF-7 cells. (a) Expression level of stem cell markers in MCF-7 cells or ALDH-positive cell population by qpcr. Data represent
Supplementary Figure 1. Characterization of ALDH-positive cell population in MCF-7 cells. (a) Expression level of stem cell markers in MCF-7 cells or ALDH-positive cell population by qpcr. Data represent
Supplementary Figure 1
 Supplementary Figure 1 6 HE-50 HE-116 E-1 HE-108 Supplementary Figure 1. Targeted drug response curves of endometrial cancer cells. Endometrial cancer cell lines were incubated with serial dilutions of
Supplementary Figure 1 6 HE-50 HE-116 E-1 HE-108 Supplementary Figure 1. Targeted drug response curves of endometrial cancer cells. Endometrial cancer cell lines were incubated with serial dilutions of
Differential altered expression of let-7a and mir-205 tumor-suppressor mirnas in different subtypes of breast cancer under treatment with Taxol
 Original papers Differential altered expression of let-7a and mir-205 tumor-suppressor mirnas in different subtypes of breast cancer under treatment with Taxol Faezeh Asghari 1,2,A D, Navideh Haghnavaz
Original papers Differential altered expression of let-7a and mir-205 tumor-suppressor mirnas in different subtypes of breast cancer under treatment with Taxol Faezeh Asghari 1,2,A D, Navideh Haghnavaz
Package TargetScoreData
 Title TargetScoreData Version 1.14.0 Author Yue Li Package TargetScoreData Maintainer Yue Li April 12, 2018 Precompiled and processed mirna-overexpression fold-changes from 84 Gene
Title TargetScoreData Version 1.14.0 Author Yue Li Package TargetScoreData Maintainer Yue Li April 12, 2018 Precompiled and processed mirna-overexpression fold-changes from 84 Gene
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Correspondence: mirna regulation of Sdf1 chemokine signaling provides genetic robustness to germ cell migration
 Correspondence: mirna regulation of Sdf1 chemokine signaling provides genetic robustness to germ cell migration Alison A. Staton, Holger Knaut, and Antonio J. Giraldez Supplementary Note Materials and
Correspondence: mirna regulation of Sdf1 chemokine signaling provides genetic robustness to germ cell migration Alison A. Staton, Holger Knaut, and Antonio J. Giraldez Supplementary Note Materials and
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Supplementary Figure 1 Supplementary Figure 1 Schematic depiction of the tandem Fc GDF15. Supplementary Figure 2 Supplementary Figure 2 Gfral mrna levels in the brains of both wild-type and knockout Gfral
Systematic exploration of autonomous modules in noisy microrna-target networks for testing the generality of the cerna hypothesis
 RESEARCH Systematic exploration of autonomous modules in noisy microrna-target networks for testing the generality of the cerna hypothesis Danny Kit-Sang Yip 1, Iris K. Pang 2 and Kevin Y. Yip 1,3,4* *
RESEARCH Systematic exploration of autonomous modules in noisy microrna-target networks for testing the generality of the cerna hypothesis Danny Kit-Sang Yip 1, Iris K. Pang 2 and Kevin Y. Yip 1,3,4* *
Cellular and molecular determinants of all-trans retinoic acid sensitivity in breast cancer: Luminal phenotype and RARa expression
 Research Article Cellular and molecular determinants of all-trans retinoic acid sensitivity in breast cancer: Luminal phenotype and RARa expression Floriana Centritto 1,, Gabriela Paroni 1,, Marco Bolis
Research Article Cellular and molecular determinants of all-trans retinoic acid sensitivity in breast cancer: Luminal phenotype and RARa expression Floriana Centritto 1,, Gabriela Paroni 1,, Marco Bolis
Supplementary appendix
 Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Cortazar P, Zhang L, Untch M, et al. Pathological
Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Cortazar P, Zhang L, Untch M, et al. Pathological
Supplementary Materials for
 www.advances.sciencemag.org/cgi/content/full/1/3/e1400244/dc1 Supplementary Materials for PlGF-induced VEGFR1-dependent vascular remodeling determines opposing antitumor effects and drug resistance to
www.advances.sciencemag.org/cgi/content/full/1/3/e1400244/dc1 Supplementary Materials for PlGF-induced VEGFR1-dependent vascular remodeling determines opposing antitumor effects and drug resistance to
CONTRACTING ORGANIZATION: Baylor College of Medicine Houston, TX 77030
 AD Award Number: W81XWH-05-1-0428 TITLE: MicroRNA and Breast Cancer Progression PRINCIPAL INVESTIGATOR: Konstantin Galaktionov, Ph.D. CONTRACTING ORGANIZATION: Baylor College of Medicine Houston, TX 77030
AD Award Number: W81XWH-05-1-0428 TITLE: MicroRNA and Breast Cancer Progression PRINCIPAL INVESTIGATOR: Konstantin Galaktionov, Ph.D. CONTRACTING ORGANIZATION: Baylor College of Medicine Houston, TX 77030
Title: Dysregulated mir-183 Inhibits Migration in Breast Cancer
 Author's response to reviews Title: Dysregulated mir-183 Inhibits Migration in Breast Cancer Authors: Aoife J Lowery (aoife.lowery@gmail.com) Nicola Miller (nicola.miller@nuigalway.ie) Roisin M Dwyer (roisin.dwyer@nuigalway.ie)
Author's response to reviews Title: Dysregulated mir-183 Inhibits Migration in Breast Cancer Authors: Aoife J Lowery (aoife.lowery@gmail.com) Nicola Miller (nicola.miller@nuigalway.ie) Roisin M Dwyer (roisin.dwyer@nuigalway.ie)
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplementary Figure 1. MLN8237 treatments show an unusual camel-back response pattern.
 Supplementary Figure 1. MLN8237 treatments show an unusual camel-ack response pattern. NCI-H1819 cells were treated with serial concentrations of MLN8237 and four days after treatment, cell viaility was
Supplementary Figure 1. MLN8237 treatments show an unusual camel-ack response pattern. NCI-H1819 cells were treated with serial concentrations of MLN8237 and four days after treatment, cell viaility was
Part-4. Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death
 Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
SubLasso:a feature selection and classification R package with a. fixed feature subset
 SubLasso:a feature selection and classification R package with a fixed feature subset Youxi Luo,3,*, Qinghan Meng,2,*, Ruiquan Ge,2, Guoqin Mai, Jikui Liu, Fengfeng Zhou,#. Shenzhen Institutes of Advanced
SubLasso:a feature selection and classification R package with a fixed feature subset Youxi Luo,3,*, Qinghan Meng,2,*, Ruiquan Ge,2, Guoqin Mai, Jikui Liu, Fengfeng Zhou,#. Shenzhen Institutes of Advanced
The mir-199a/brm/egr1 axis is a determinant of anchorage-independent growth in epithelial tumor cell lines
 Supplementary information Supplementary Figure -9 Supplementary Table -4 The mir-99a/brm/egr axis is a determinant of anchorage-independent growth in epithelial tumor cell lines Kazuyoshi Kobayashi, Kouhei
Supplementary information Supplementary Figure -9 Supplementary Table -4 The mir-99a/brm/egr axis is a determinant of anchorage-independent growth in epithelial tumor cell lines Kazuyoshi Kobayashi, Kouhei
Micro-RNA web tools. Introduction. UBio Training Courses. mirnas, target prediction, biology. Gonzalo
 Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3021 Supplementary figure 1 Characterisation of TIMPless fibroblasts. a) Relative gene expression of TIMPs1-4 by real time quantitative PCR (RT-qPCR) in WT or ΔTimp fibroblasts (mean ±
DOI: 10.1038/ncb3021 Supplementary figure 1 Characterisation of TIMPless fibroblasts. a) Relative gene expression of TIMPs1-4 by real time quantitative PCR (RT-qPCR) in WT or ΔTimp fibroblasts (mean ±
micrornas (mirna) and Biomarkers
 micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier
 Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Supplementary Figure S1 Expression of mir-181b in EOC (A) Kaplan-Meier curves for progression-free survival (PFS) and overall survival (OS) in a cohort of patients (N=52) with stage III primary ovarian
Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3
 Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Nature Genetics: doi: /ng.3731
 Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Supplementary Figure 1 Circadian profiles of Adarb1 transcript and ADARB1 protein in mouse tissues. (a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver
Bmi-1 regulates stem cell-like properties of gastric cancer cells via modulating mirnas
 Wang et al. Journal of Hematology & Oncology (2016) 9:90 DOI 10.1186/s13045-016-0323-9 RESEARCH Bmi-1 regulates stem cell-like properties of gastric cancer cells via modulating mirnas Open Access Xiaofeng
Wang et al. Journal of Hematology & Oncology (2016) 9:90 DOI 10.1186/s13045-016-0323-9 RESEARCH Bmi-1 regulates stem cell-like properties of gastric cancer cells via modulating mirnas Open Access Xiaofeng
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland X Approved for public release; distribution unlimited
 AD (Leave blank) Award Number: W81XWH-08-1-0458 TITLE: Altered MicroRNA Activity Promotes Resistance to Endocrine Therapy PRINCIPAL INVESTIGATOR: Diana M. Cittelly CONTRACTING ORGANIZATION: University
AD (Leave blank) Award Number: W81XWH-08-1-0458 TITLE: Altered MicroRNA Activity Promotes Resistance to Endocrine Therapy PRINCIPAL INVESTIGATOR: Diana M. Cittelly CONTRACTING ORGANIZATION: University
Bhatnagar et al, 2010 Cell Death and Disease Manuscript # CDDIS T
 Bhatnagar et al, Cell Death and Disease Manuscript # CDDIS--98-T Supplemental Materials. Supplemental Figure Legends Supplemental Figure (A) WPE-NA and WPE-NB6 cells were treated with 4 nm of Docetaxel
Bhatnagar et al, Cell Death and Disease Manuscript # CDDIS--98-T Supplemental Materials. Supplemental Figure Legends Supplemental Figure (A) WPE-NA and WPE-NB6 cells were treated with 4 nm of Docetaxel
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Amelio et al., http://www.jcb.org/cgi/content/full/jcb.201203134/dc1 Figure S1. mir-24 regulates proliferation and by itself induces
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Amelio et al., http://www.jcb.org/cgi/content/full/jcb.201203134/dc1 Figure S1. mir-24 regulates proliferation and by itself induces
Research progress on the use of estrogen receptor agonist for treatment of spinal cord injury
 Research progress on the use of estrogen receptor agonist for treatment of spinal cord injury Swapan K. Ray, PhD Professor, Department of Pathology, Microbiology, and Immunology USC School of Medicine,
Research progress on the use of estrogen receptor agonist for treatment of spinal cord injury Swapan K. Ray, PhD Professor, Department of Pathology, Microbiology, and Immunology USC School of Medicine,
Understanding PI3K/mTOR/MEK signalling in breast cancer. Euphemia Leung Auckland Cancer Society Research Centre University of Auckland
 Understanding PI3K/mTOR/MEK signalling in breast cancer Euphemia Leung Auckland Cancer Society Research Centre University of Auckland Receptor status of human breast cancer On a popula>on basis ER+/PR+
Understanding PI3K/mTOR/MEK signalling in breast cancer Euphemia Leung Auckland Cancer Society Research Centre University of Auckland Receptor status of human breast cancer On a popula>on basis ER+/PR+
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained
 Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
Supplementary Figure 1. Confocal immunofluorescence showing mitochondrial translocation of Drp1. Cardiomyocytes treated with H 2 O 2 were prestained with MitoTracker (red), then were immunostained with
Supplementary information
 Supplementary information Exosomes mediate the cell-to-cell transmission of interferon alpha-induced antiviral activity Jianhua Li, Kuancheng Liu, Yang Liu, Yan Xu, Fei Zhang, Huijuan Yang, Jiangxia Liu,
Supplementary information Exosomes mediate the cell-to-cell transmission of interferon alpha-induced antiviral activity Jianhua Li, Kuancheng Liu, Yang Liu, Yan Xu, Fei Zhang, Huijuan Yang, Jiangxia Liu,
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
SUPPLEMENTAL FILE. mir-22 and mir-29a are members of the androgen receptor cistrome modulating. LAMC1 and Mcl-1 in prostate cancer
 1 SUPPLEMENTAL FILE 2 3 mir-22 and mir-29a are members of the androgen receptor cistrome modulating LAMC1 and Mcl-1 in prostate cancer 4 5 6 Lorenza Pasqualini 1, Huajie Bu 1,2, Martin Puhr 1, Narisu Narisu
1 SUPPLEMENTAL FILE 2 3 mir-22 and mir-29a are members of the androgen receptor cistrome modulating LAMC1 and Mcl-1 in prostate cancer 4 5 6 Lorenza Pasqualini 1, Huajie Bu 1,2, Martin Puhr 1, Narisu Narisu
Title page. Title: MicroRNA-155 Controls Exosome Synthesis and Promotes Gemcitabine Resistance in
 Title page Title: MicroRNA- Controls Synthesis and Promotes Gemcitabine Resistance in Pancreatic Ductal Adenocarcinoma Authors Manabu Mikamori, Daisaku Yamada, Hidetoshi Eguchi, Shinichiro Hasegawa, Tomoya
Title page Title: MicroRNA- Controls Synthesis and Promotes Gemcitabine Resistance in Pancreatic Ductal Adenocarcinoma Authors Manabu Mikamori, Daisaku Yamada, Hidetoshi Eguchi, Shinichiro Hasegawa, Tomoya
A gene expression profile indicative of early stage HER2 targeted therapy response
 O Neill et al. Molecular Cancer 213, 12:69 RESEARCH Open Access A gene expression profile indicative of early stage HER2 targeted therapy response Fiona O Neill 1,2, Stephen F Madden 1, Martin Clynes 1,
O Neill et al. Molecular Cancer 213, 12:69 RESEARCH Open Access A gene expression profile indicative of early stage HER2 targeted therapy response Fiona O Neill 1,2, Stephen F Madden 1, Martin Clynes 1,
Inferring condition-specific mirna activity from matched mirna and mrna expression data
 Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL-
 Supplementary Material for mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL- XL Inhibition by Suppressing MCL-1 Anthony C. Faber 1,2 *, Erin M. Coffee
Supplementary Material for mtor Inhibition Specifically Sensitizes Colorectal Cancers with KRAS or BRAF Mutations to BCL-2/BCL- XL Inhibition by Suppressing MCL-1 Anthony C. Faber 1,2 *, Erin M. Coffee
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
microrna PCR System (Exiqon), following the manufacturer s instructions. In brief, 10ng of
 SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
ONCOLOGY LETTERS 9: , 2015
 ONCOLOGY LETTERS 9: 1351-1358, 2015 Lapatinib combined with neoadjuvant paclitaxel trastuzumab based chemotherapy in patients with human epidermal growth factor receptor 2 positive breast cancer: A meta
ONCOLOGY LETTERS 9: 1351-1358, 2015 Lapatinib combined with neoadjuvant paclitaxel trastuzumab based chemotherapy in patients with human epidermal growth factor receptor 2 positive breast cancer: A meta
From reference genes to global mean normalization
 From reference genes to global mean normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle qpcr Symposium USA November 9, 2009 Millbrae, CA outline what is normalization
From reference genes to global mean normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle qpcr Symposium USA November 9, 2009 Millbrae, CA outline what is normalization
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Comparative study of multicellular tumor spheroid formation methods and implications for drug screening
 Supporting Information Comparative study of multicellular tumor spheroid formation methods and implications for drug screening Maria F. Gencoglu, Lauren E. Barney, Christopher L. Hall, Elizabeth A. Brooks,
Supporting Information Comparative study of multicellular tumor spheroid formation methods and implications for drug screening Maria F. Gencoglu, Lauren E. Barney, Christopher L. Hall, Elizabeth A. Brooks,
A novel and universal method for microrna RT-qPCR data normalization
 A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
To determine the effect of over-expression and/or ligand activation of. PPAR / on cell cycle, cell lines were cultured as described above until ~80%
 Supplementary Materials and Methods Cell cycle analysis To determine the effect of over-expression and/or ligand activation of PPAR / on cell cycle, cell lines were cultured as described above until ~80%
Supplementary Materials and Methods Cell cycle analysis To determine the effect of over-expression and/or ligand activation of PPAR / on cell cycle, cell lines were cultured as described above until ~80%
Products for cfdna and mirna isolation. Subhead Circulating Cover nucleic acids from plasma
 MACHEREY-NAGEL Products for cfdna and mirna isolation Bioanalysis Subhead Circulating Cover nucleic acids from plasma n Flexible solutions for small and large blood plasma volumes n Highly efficient recovery
MACHEREY-NAGEL Products for cfdna and mirna isolation Bioanalysis Subhead Circulating Cover nucleic acids from plasma n Flexible solutions for small and large blood plasma volumes n Highly efficient recovery
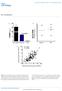 DOI: 10.1038/ncb2210 b. ICAM1 ng ml -1 P = 0.0001 Small RNA (15-30nts) ng ml -1 Cell Lysate Exosome HDL Plasma HDL Normal Human HDL mirnas R = 0.45 P < 0.0001 Normal Human Exosome mirnas Figure S1. Characterization
DOI: 10.1038/ncb2210 b. ICAM1 ng ml -1 P = 0.0001 Small RNA (15-30nts) ng ml -1 Cell Lysate Exosome HDL Plasma HDL Normal Human HDL mirnas R = 0.45 P < 0.0001 Normal Human Exosome mirnas Figure S1. Characterization
R2 Training Courses. Release The R2 support team
 R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
R2 Training Courses Release 2.0.2 The R2 support team Nov 08, 2018 Students Course 1 Student Course: Investigating Intra-tumor Heterogeneity 3 1.1 Introduction.............................................
SUPPLEMENTARY FIGURE LEGENDS
 SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
SUPPLEMENTARY FIGURE LEGENDS Supplementary Figure 1 Negative correlation between mir-375 and its predicted target genes, as demonstrated by gene set enrichment analysis (GSEA). 1 The correlation between
