Handling Immunogenetic Data Managing and Validating HLA Data
|
|
|
- Tracey Burns
- 6 years ago
- Views:
Transcription
1 Handling Immunogenetic Data Managing and Validating HLA Data Steven J. Mack PhD Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012
2 Overview 1. Master Analytical Dataset 2. The Perils of Using MS Excel 3. HLA Ambiguity 4. Internal Validation and Standardization of Datasets 5. Analytical Validation of HLA Population Data
3 Create a Master Dataset For Analysis/Sharing Create a single master data-file for research datasets Integrate demographic, phenotypic, and genotypic information Each analytical element (sample) is a row Each variable (genotype, phenotype, measurement) is a column Where possible, encode variable data numerically avoids errors, ambiguity of meaning, and saves space A data dictionary accompanies and explains the master data-file Describe the meaning and significance of each variable Define the numerical codes for each variable 1 = affected, 0 = control Define the code(s) for missing values -9 = unknown
4 Should I Store My Data in Excel? Microsoft Excel doesn t speak HLA IMGT/HLA db version Allele Name Excel Error 1.*.* *.* *.* 01:01:01 1:01:01 AM 3.*.* 01: Opening a text file by right-clicking and selecting Excel will result in Excel errors like these. Use Excel at your own risk; store data using a text format; consider using database software.
5 Consider Short-Term and Long-Term Needs for HLA Data Short-Term Need Interim Reporting, Submission Deadline, Data Analysis May often require an abstracted best guess to summarize the data. But that best guess may only be useful at the time it is made. Long-Term Need Archiving, Storage, Deposition into Public Domain Registry, Meta-Analysis, Multi-cycle projects Need to be able to make full use of these data in the future; The goal is to maximize available information, which often requires that primary or raw data be maintained. Should be storing both cleaned and ambiguous data.
6 HLA Data Ambiguity Allele ambiguity results when the polymorphisms that distinguish alleles fall outside of the regions assessed by the genotyping system. A*02:03:01/A*02:253/A*02:264 identical exon 2&3 sequence Genotype ambiguity results from an inability to establish chromosomal phase between identified polymorphisms. DRB1*04:01:01+DRB1*13:01:01 or DRB1*04:01:01+ DRB1*13:117 or DRB1*04:13+DRB1*13:02:01 or DRB1*04:14+DRB1*14:21 or DRB1*04:35+DRB1*13:40 or DRB1*04:38+DRB1*13:20 identical heterozygous exon 2 sequence An HLA genotype can include both allele and genotype ambiguity A*02:03:01/A*02:253/A*02:264+A*03:01:02 or A*02:171:01+A*03:50 or A*02:171:02+A*03:66
7 NMDP Allele Codes Coding Ambiguous HLA Data DRB1*04:02:01+DRB1*11:20 or DRB1*04:14+DRB1*11:16 Can be represented as DRB1*04:BK+DRB1*11:YR DRB1*04:BK represents DRB1*04:02/DRB1*04:14 DRB1*11:YR represents DRB1*11:20/DRB1*11:16 But these codes also represent excluded genotypes DRB1*04:02:01+DRB1*11:16 and DRB1*04:14+DRB1*11:20 Allele Codes increase ambiguity and omit information in the 3 rd & 4 th fields
8 Coding Ambiguous HLA Data Allele Groups G groups: Alleles with identical nucleotide sequence in the exons encoding the peptide binding domain (exon 2 for class II alleles, and exons 2 & 3 for class I alleles) A*02:03:01G = A*02:03:01/A*02:253/A*02:264 A*02:07:01G = A*02:07:01/A*02:07:02/A*02:15N/A*02:265 P groups: Alleles with identical peptide sequence in the peptide binding domain, with the exclusion of null alleles. A*02:03P = A*02:03:01/A*02:03:02/A*02:03:03/A*02:03:04/A*02:253/A*02:264 A*02:07P = A*02:07:01/A*02:07:02/A*02:265
9 Recording Ambiguous HLA Data Genotype Strings Genotype List String (GL String) Uses specific operators to describe the relationships between alleles, allowing genotype data to be recorded in a single line. Order of Precedence Data Delimiter Operator Description 1 ^ Gene/Locus 2 Genotype list 3 + Genotype 4 ~ Haplotype 5 / Alleles
10 GL String Representation of Ambiguous HLA Data Allelic ambiguity delimiter (forward slash) Defines an allele list / A*23:26/A*23:39 allele allele Possible alleles at locus A
11 GL String Representation of Ambiguous HLA Data ~ Haplotype delimiter (tilde) Applied in cis to identify a haplotype DRB5*01:02~DRB1*15:04 allele at locus DRB5 allele at locus DRB1 same chromosome
12 GL String Representation of Ambiguous HLA Data Genotype delimiter (plus sign) Identifies alleles on different chromosomes, genotype (trans) Delimits haplotype May also indicate gene duplication (ambiguous cis) + A*02:302+A*23:26/A*23:39 allele allele allele HLA-A allelic ambiguity genotype
13 GL String Representation of Ambiguous HLA Data Genotype list delimiter (pipe) Distinguishes ambiguous genotypes A*02:69+A*23:30 A*02:302+A*23:26/A*23:39 allele allele allele ambiguous allele list genotype genotype
14 GL String Representation of Ambiguous HLA Data Locus delimiter (carat) Distinguishes loci A*02:69+A*23:30 A*02:302+A*23:26/A*23:39^B*44:02:13+B*49:08 ^ allele allele allele ambiguous allele list allele allele possible genotype for A possible genotype for A Ambiguous genotype list for HLA-A HLA-B genotype
15 Alternative Format for Recording Ambiguous HLA Data UNIFORMAT Also allows ambiguous genotype data to be recorded in a single line, using different operators (colons, commas, spaces, tabs) than in GL String. identifier {tab mark} allele,allele [{space} allele,allele...][{tab mark} allele, allele...][{tab mark}#comments] sample identifier ambiguous alleles at one locus ambiguous alleles at additional loci comments
16 Know Your Nomenclature Version Identify the IMGT/HLA database release number applicable to your data The release number identifies which alleles should and should not be in your data IMGT/HLA db rel Allele 1.13 A* A* B* B* DPB1* DPB1*104:01 This information allows you to check your data for naming/recording errors
17 Dataset Validation The Allele Name Translation Tool (ANTT) can be used to validate/update the allele names in column-formatted datasets against/to any IMGT/HLA db release. Parses forward-slash (/) allele ambiguity delimiters; the next version will parse any delimiters (e.g. all GL String delimiters). Documents right-truncated allele names and unrecognized allele names. Identifies the id and row-column position of errors. DRB1*08:02:00 could not be found in the HLA-DRB1.upd translation file.[id = 003][Row = 4 Column = 2] DRB1*04:08 appears to be a truncated version of the DRB1*04:08:01 allele, and was translated to DRB1*0408. [id = 005][Row = 6 Column = 2]
18 Internal Standardization of Data Data Consistency Record data consistently across individuals (and datasets). For individuals typed Record homozygotes as diploid A*02:03:01/A*02:253/A*02:264+A*02:03:01/A*02:253/A*02:264 Use a code to identify missing data ****, -9, missing, etc. Use a code to identify absent loci when recording structural variants DRB1*01:01:01~DRB3*BLANK~DRB4*BLANK~DRB5*BLANK+DRB1*15:01:01~DRB3*BLANK~DRB4*BLANK~DRB5*01:01:01
19 Internal Standardization of Data Data Modification Document any post-typing modifications made to data. How did A*02:69+A*23:30 A*02:302+A*23:26/A*23:39 become A*02:302+A*23:39? For analysis across individuals and datasets Analyze allele names at the same level of polymorphism Avoid A*01:01:01:01, A*01:01:01, A*01:01, and A*01 in the same analysis You will have to throw out some data/information. Analyze alleles at the same sequence level For exons 2/2& 3 testing, don t analyze alleles in the same G group separately. Analyze DRB1*14:01:01 and DRB1*14:54 as DRB1*14:01:01G. Analyze allele names in the same nomenclature context Don t analyze A*2416 and A*3108; update to a single nomenclature.
20 Analytical Validation of HLA Population Data The Hardy-Weinberg (HW) model can give you insights into data-quality Hardy-Weinberg Equilibrium The frequency of the alleles should predict the frequency of the genotypes. If it does not (HW deviation), you may have problems with your data. Multi-locus HW deviation Sampling error Related individuals Populations mixed together (admixture) Solution: Review inclusion criteria and remove individuals Critical Typing problem (uncommon) Single-locus HW deviation Typing error Excess of homozygotes due to missed alleles Excess of heterozygotes due to poor assignment Solution: Review and redo typings Selection (unexpected in control-populations)
21 Example of HW Data Validation 13 DQB1 alleles in a population of n=109 Genotype Observed Count Expected Count p-value DQB1*03:03:02+DQB1*02:01: DQB1*03:03:02+DQB1*03:03: Chen s test of individual genotypes in PyPop ( HW deviation due to poor detection of DQB1*02:01:01 in the presence of DQB1*03:03:02, resulting from a SNP in DQB1*02:01:01 under a PCR primer. For population/control datasets, the first analysis done should be a HW test.
22 Much Thanks To Pierre-Antoine Gourraud Standard Methods for the Management Jill A. Hollenbach of Immunogenetic Data. Frank T. Christiansen Thomas Barnetche and Brian D. Tait (eds.), Immunogenetics: Richard Single Methods and Applications in Clinical Practice, Steven J. Mack Methods in Molecular Biology, vol pages doi: / _12 Children s Hospital Oakland Henry A. Erlich Janelle Noble Elizabeth Trachtenberg Immunogenetics Colleagues Glenys Thomson Alicia Sanchez-Mazas Owen D. Solberg Martin Maiers Carolyn Hurley Marcel Tilanus Christien Voorter Immunogenetics Community
23 Handling Immunogenetic Data Managing Highly Polymorphic Data for Disease Association Studies Jill A. Hollenbach, PhD, MPH Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012
24 Immunogenetic data require special handling in disease association studies
25 Immunogenetic data require special handling in disease association studies 1 Highly polymorphic loci
26 Immunogenetic data require special handling in disease association studies 1 Highly polymorphic loci Many rare alleles >>> sparse cells
27 Immunogenetic data require special handling in disease association studies 1 Highly polymorphic loci Many rare alleles >>> sparse cells Need to identify all disease associated alleles
28 Immunogenetic data require special handling in disease association studies 1 Highly polymorphic loci Many rare alleles >>> sparse cells Need to identify all disease associated alleles 2 Strong linkage disequilibrium
29 Immunogenetic data require special handling in disease association studies 1 Highly polymorphic loci Many rare alleles >>> sparse cells Need to identify all disease associated alleles 2 Strong linkage disequilibrium Identify which loci are primary
30 Immunogenetic data require special handling in disease association studies 1 Highly polymorphic loci Many rare alleles >>> sparse cells Need to identify all disease associated alleles 2 Strong linkage disequilibrium Identify which loci are primary 3Immunogenetic loci show strong population structure
31 Case-Control Study
32 Case-Control Study Statistical tests
33 Case-Control Study Statistical tests First step: Population analyses
34 Case-Control Study Statistical tests First step: Population analyses Tests for fit to HWEP
35 Case-Control Study Statistical tests First step: Population analyses Tests for fit to HWEP Calculation of allele and haplotype frequencies
36 Case-Control Study Statistical tests First step: Population analyses Tests for fit to HWEP Calculation of allele and haplotype frequencies Association tests
37 Case-Control Study Statistical tests First step: Population analyses Tests for fit to HWEP Calculation of allele and haplotype frequencies Association tests Contingency tables /chi-squared test
38 Case-Control Study Statistical tests First step: Population analyses Tests for fit to HWEP Calculation of allele and haplotype frequencies Association tests Contingency tables /chi-squared test Logistic regression
39 Case-Control Study Statistical tests First step: Population analyses Tests for fit to HWEP Calculation of allele and haplotype frequencies Association tests Contingency tables /chi-squared test Logistic regression Other tests/special cases (Cochran-Armitage test for trend; survival analysis; etc)
40 Case-Control Study Statistical tests First step: Population analyses Tests for fit to HWEP Calculation of allele and haplotype frequencies Association tests Contingency tables /chi-squared test Logistic regression Other tests/special cases (Cochran-Armitage test for trend; survival analysis; etc)
41 Case-Control Study Contingency tables
42 Case-Control Study Contingency tables Test difference (independence) of frequency distributions for categorical variables between groups - 2 test
43 Case-Control Study Contingency tables Test difference (independence) of frequency distributions for categorical variables between groups - 2 test Always constructed with raw counts, not frequency data
44 Case-Control Study Contingency tables Test difference (independence) of frequency distributions for categorical variables between groups - 2 test Always constructed with raw counts, not frequency data Analyses can be performed at the allele, genotype, haplotype, amino acid or other levels
45 Sparse cells in contingency tables
46 Chi-squared Test Statistic: (O - E) 2 c 2 = E å all cells O is the observed cell counts E is the expected cell counts, where E = Sparse cells in contingency tables (row total column total) 2N
47 Chi-squared Test Statistic: (O - E) 2 c 2 = E å all cells O is the observed cell counts E is the expected cell counts, where E = Sparse cells in contingency tables (row total column total) 2N c 2 test is inappropriate if any expected count is less than 1 or if the expected count is less than five in more than 20% of all cells in a contingency table *** aka sparse cells
48 Sparse cells in contingency tables DRB1 case control n
49 Sparse cells in contingency tables DRB1 case control n
50 Sparse cells in contingency tables DRB1 case control binned n
51 Sparse cells in contingency tables DRB1 case control p-value binned
52 Sparse cells in contingency tables DRB1 case control p-value binned
53 Identifying all disease associated alleles
54 Identifying all disease associated alleles Relative predispositional effects method (RPE; Payami et al 1989)
55 Identifying all disease associated alleles Relative predispositional effects method (RPE; Payami et al 1989) Method to identify all heterogeneity in disease risk at locus of interest
56 Identifying all disease associated alleles Relative predispositional effects method (RPE; Payami et al 1989) Method to identify all heterogeneity in disease risk at locus of interest Contingency table testing reveals overall difference in allele frequency distributions at a locus
57 Identifying all disease associated alleles Relative predispositional effects method (RPE; Payami et al 1989) Method to identify all heterogeneity in disease risk at locus of interest Contingency table testing reveals overall difference in allele frequency distributions at a locus But we want to identify all alleles that contribute significantly
58 Identifying all disease associated alleles Relative predispositional effects method (RPE; Payami et al 1989) Method to identify all heterogeneity in disease risk at locus of interest Contingency table testing reveals overall difference in allele frequency distributions at a locus But we want to identify all alleles that contribute significantly Alleles with the strongest predisposing or protective effects sequentially removed from analysis until no further heterogeneity in risk effects is seen
59 Identifying all disease associated alleles DRB1 case control p-value binned
60 Identifying all disease associated alleles DRB1 case control p-value binned
61 Identifying all disease associated alleles
62 Identifying all disease associated alleles
63 Identifying the primary locus
64 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls
65 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03:
66 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved.
67 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03)
68 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) (0.10)/(0.04)=
69 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) (0.10)/(0.04)=2.5
70 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.10)/(0.04)=2.5
71 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.10)/(0.04)=2.5 (0.05)/(0.02)=
72 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.10)/(0.04)=2.5 (0.05)/(0.02)=2.5
73 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.10)/(0.04)=2.5 (0.05)/(0.02)=2.5
74 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved.
75 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03)
76 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) (0.12)/(0.02)=
77 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) (0.12)/(0.02)=6
78 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.12)/(0.02)=6
79 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.12)/(0.02)=6 (0.05)/(0.02)=
80 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.12)/(0.02)=6 (0.05)/(0.02)=2.5
81 Identifying the primary locus The frequencies of two haplotypes of an allele at the predisposing locus may differ between patients and controls DRB1~DQB1 Haplotype Case Control 07:01~02: :01~03: HOWEVER: the relative frequency of their ratios will be the same if the second locus is not involved. f case (07:01~02:01)/f case (07:01~03:03) f cont (07:01~02:01)/f cont (07:01~03:03) (0.12)/(0.02)=6 (0.05)/(0.02)=2.5
82 Population substructure in disease association studies
83 Population substructure in disease association studies (f) PopX (f) LocusA PopY
84 Population substructure in disease association studies (f) PopX (f) LocusA PopY
85 Population substructure in disease association studies (f) PopX cases controls (f) LocusA PopY cases controls
86 Population substructure in disease association studies (f) PopX cases controls LocusA PopY No association (f) cases controls
87 Population substructure in disease association studies (f) PopX cases controls LocusA PopY No association (f) cases controls
88 Population substructure in disease association studies (f) PopX cases controls (f) LocusA PopY No association PopXY cases controls
89 Population substructure in disease association studies (f) PopX cases controls (f) LocusA PopY No association PopXY cases controls cases controls
90 Population substructure in disease association studies (f) PopX cases controls (f) LocusA PopY No association PopXY cases controls cases controls
91 Population substructure in disease association studies (f) PopX cases controls (f) LocusA PopY No association PopXY cases controls p<.05 cases controls
92 Population substructure in disease association studies cases controls p<.05
93 Population substructure in disease association studies (f) PopX cases controls (f) LocusA PopY No association PopXY cases controls p<.05 cases controls
94 Population substructure in disease association studies (f) PopX cases controls (f) LocusA PopY No association PopXY cases controls p<.05 cases controls
95 Population substructure in disease association studies cases controls p<.05
96 Immunogenetic data require special handling in disease association studies
97 Immunogenetic data require special handling in disease association studies Highly polymorphic loci
98 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles
99 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles Binning
100 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles Binning Need to identify all associated alleles
101 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles Binning Need to identify all associated alleles Relative predispositional effects
102 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles Binning Need to identify all associated alleles Relative predispositional effects Strong linkage disequilibrium
103 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles Binning Need to identify all associated alleles Relative predispositional effects Strong linkage disequilibrium Identify which loci are primary
104 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles Binning Need to identify all associated alleles Relative predispositional effects Strong linkage disequilibrium Identify which loci are primary Condition within haplotypes
105 Immunogenetic data require special handling in disease association studies Highly polymorphic loci Combine low frequency alleles Binning Need to identify all associated alleles Relative predispositional effects Strong linkage disequilibrium Identify which loci are primary Condition within haplotypes Control for population substructure
106 For further discussion see: Hollenbach JA, Mack SJ, Thomson G, Gourraud PA. Analytical methods for disease association studies with immunogenetic data. Methods Mol Biol. 2012;882: DOI: / _14
107 Thank you!
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS
 DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
Identifying Biologically Relevant Amino Acids in Immunogenetic Studies
 Identifying Biologically Relevant Amino Acids in Immunogenetic Studies Richard M. Single Department of Mathematics and Statistics University of Vermont Outline HLA background and nomenclature Asymmetric
Identifying Biologically Relevant Amino Acids in Immunogenetic Studies Richard M. Single Department of Mathematics and Statistics University of Vermont Outline HLA background and nomenclature Asymmetric
HLA Mismatches. Professor Steven GE Marsh. Anthony Nolan Research Institute EBMT Anthony Nolan Research Institute
 HLA Mismatches Professor Steven GE Marsh HLA Mismatches HLA Genes, Structure, Polymorphism HLA Nomenclature HLA Mismatches in HSCT Defining a mismatch HLA Mismatches HLA Genes, Structure, Polymorphism
HLA Mismatches Professor Steven GE Marsh HLA Mismatches HLA Genes, Structure, Polymorphism HLA Nomenclature HLA Mismatches in HSCT Defining a mismatch HLA Mismatches HLA Genes, Structure, Polymorphism
HLA-A * L
 What is Nomenclature? HLA Nomenclature 3.0 Signifies DNA Subtype Differences outside the coding region (introns) HLA-A * 4 0 01 0 L Steve Spellman Sr. Manager, NMDP Asst. Scientific Director, CIBMTR Locus
What is Nomenclature? HLA Nomenclature 3.0 Signifies DNA Subtype Differences outside the coding region (introns) HLA-A * 4 0 01 0 L Steve Spellman Sr. Manager, NMDP Asst. Scientific Director, CIBMTR Locus
The Human Major Histocompatibility Complex
 The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
An association analysis of the HLA gene region in latent autoimmune diabetes in adults
 Diabetologia (2007) 50:68 73 DOI 10.1007/s00125-006-0513-z SHORT COMMUNICATION An association analysis of the HLA gene region in latent autoimmune diabetes in adults M. Desai & E. Zeggini & V. A. Horton
Diabetologia (2007) 50:68 73 DOI 10.1007/s00125-006-0513-z SHORT COMMUNICATION An association analysis of the HLA gene region in latent autoimmune diabetes in adults M. Desai & E. Zeggini & V. A. Horton
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES. Irina Durbală
 ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
Human Leukocyte Antigens and donor selection
 Human Leukocyte Antigens and donor selection Duangtawan Thammanichanond, MD. PhD. Histocompatibility and Immunogenetics Laboratory, Faculty of Medicine Ramathibodi Hospital, Mahidol University Outline
Human Leukocyte Antigens and donor selection Duangtawan Thammanichanond, MD. PhD. Histocompatibility and Immunogenetics Laboratory, Faculty of Medicine Ramathibodi Hospital, Mahidol University Outline
HLA Disease Associations Methods Manual Version (July 25, 2011)
 HLA Disease Associations Methods Manual Version 0.1.0 (July 25, 2011) HLA Disease Associations: Detecting Primary and Secondary Disease Predisposing Genes Available online at: The Immunology Database and
HLA Disease Associations Methods Manual Version 0.1.0 (July 25, 2011) HLA Disease Associations: Detecting Primary and Secondary Disease Predisposing Genes Available online at: The Immunology Database and
Research: Genetics HLA class II gene associations in African American Type 1 diabetes reveal a protective HLA-DRB1*03 haplotype
 Research: Genetics HLA class II gene associations in African American Type 1 diabetes reveal a protective HLA-DRB1*03 haplotype J. M. M. Howson 1,2, M. S. Roy 3, L. Zeitels 1, H. Stevens 1 and J. A. Todd
Research: Genetics HLA class II gene associations in African American Type 1 diabetes reveal a protective HLA-DRB1*03 haplotype J. M. M. Howson 1,2, M. S. Roy 3, L. Zeitels 1, H. Stevens 1 and J. A. Todd
Documentation of Changes to EFI Standards: v 5.6.1
 Modified Standard B - PERSONNEL QUALIFICATIONS B1.000 The laboratory must employ one or more individuals who meet the qualifications and fulfil the responsibilities of the Director/Co-Director, Technical
Modified Standard B - PERSONNEL QUALIFICATIONS B1.000 The laboratory must employ one or more individuals who meet the qualifications and fulfil the responsibilities of the Director/Co-Director, Technical
Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97
 88 Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97 ARTICLE HLA gene and haplotype frequency in renal transplant recipients and donors of Uttar Pradesh (North India) S Agrawal, AK Singh, RK
88 Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97 ARTICLE HLA gene and haplotype frequency in renal transplant recipients and donors of Uttar Pradesh (North India) S Agrawal, AK Singh, RK
ASHI Proficiency Testing Program Summary Report. Survey 2013-HT1 / HLA Typing
 ASHI Proficiency Testing Program Summary Report Survey 2013-HT1 / HLA Typing Shipping Date: February 26,2013 / Results Due Date:April 5,2013 / Report Date: May 17,2013 The ASHI HT proficiency testing survey
ASHI Proficiency Testing Program Summary Report Survey 2013-HT1 / HLA Typing Shipping Date: February 26,2013 / Results Due Date:April 5,2013 / Report Date: May 17,2013 The ASHI HT proficiency testing survey
Decomposition of the Genotypic Value
 Decomposition of the Genotypic Value 1 / 17 Partitioning of Phenotypic Values We introduced the general model of Y = G + E in the first lecture, where Y is the phenotypic value, G is the genotypic value,
Decomposition of the Genotypic Value 1 / 17 Partitioning of Phenotypic Values We introduced the general model of Y = G + E in the first lecture, where Y is the phenotypic value, G is the genotypic value,
Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005)
 Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005) Stephen Spellman Research Manager NMDP Scientific Services Maria Brown Scientific Services Specialist Data Management Conference 2007 1
Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005) Stephen Spellman Research Manager NMDP Scientific Services Maria Brown Scientific Services Specialist Data Management Conference 2007 1
Diversity and Frequencies of HLA Class I and Class II Genes of an East African Population
 Open Journal of Genetics, 2014, 4, 99-124 Published Online April 2014 in SciRes. http://www.scirp.org/journal/ojgen http://dx.doi.org/10.4236/ojgen.2014.42013 Diversity and Frequencies of HLA Class I and
Open Journal of Genetics, 2014, 4, 99-124 Published Online April 2014 in SciRes. http://www.scirp.org/journal/ojgen http://dx.doi.org/10.4236/ojgen.2014.42013 Diversity and Frequencies of HLA Class I and
BST227 Introduction to Statistical Genetics. Lecture 4: Introduction to linkage and association analysis
 BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
Whole-genome detection of disease-associated deletions or excess homozygosity in a case control study of rheumatoid arthritis
 HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
HLA Amino Acid Polymorphisms and Kidney Allograft Survival. Supplemental Digital Content
 HLA Amino Acid Polymorphisms and Kidney Allograft Survival Supplemental Digital Content Malek Kamoun, MD, PhD a, Keith P. McCullough, MS b, Martin Maiers, MS c, Marcelo A. Fernandez Vina, MD, PhD d, Hongzhe
HLA Amino Acid Polymorphisms and Kidney Allograft Survival Supplemental Digital Content Malek Kamoun, MD, PhD a, Keith P. McCullough, MS b, Martin Maiers, MS c, Marcelo A. Fernandez Vina, MD, PhD d, Hongzhe
Effects of age-at-diagnosis and duration of diabetes on GADA and IA-2A positivity
 Effects of age-at-diagnosis and duration of diabetes on GADA and IA-2A positivity Duration of diabetes was inversely correlated with age-at-diagnosis (ρ=-0.13). However, as backward stepwise regression
Effects of age-at-diagnosis and duration of diabetes on GADA and IA-2A positivity Duration of diabetes was inversely correlated with age-at-diagnosis (ρ=-0.13). However, as backward stepwise regression
Statistical Tests for X Chromosome Association Study. with Simulations. Jian Wang July 10, 2012
 Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
Evaluation of MIA FORA NGS HLA test and software. Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group
 Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Association between the -77T>C polymorphism in the DNA repair gene XRCC1 and lung cancer risk
 Association between the -77T>C polymorphism in the DNA repair gene XRCC1 and lung cancer risk B.B. Sun, J.Z. Wu, Y.G. Li and L.J. Ma Department of Respiratory Medicine, People s Hospital Affiliated to
Association between the -77T>C polymorphism in the DNA repair gene XRCC1 and lung cancer risk B.B. Sun, J.Z. Wu, Y.G. Li and L.J. Ma Department of Respiratory Medicine, People s Hospital Affiliated to
2/10/2016. Evaluation of MIA FORA NGS HLA test and software. Disclosure. NGS-HLA typing requirements for the Stanford Blood Center
 Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Bio 312, Spring 2017 Exam 3 ( 1 ) Name:
 Bio 312, Spring 2017 Exam 3 ( 1 ) Name: Please write the first letter of your last name in the box; 5 points will be deducted if your name is hard to read or the box does not contain the correct letter.
Bio 312, Spring 2017 Exam 3 ( 1 ) Name: Please write the first letter of your last name in the box; 5 points will be deducted if your name is hard to read or the box does not contain the correct letter.
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/20898 holds various files of this Leiden University dissertation. Author: Jöris, Monique Maria Title: Challenges in unrelated hematopoietic stem cell transplantation.
Cover Page The handle http://hdl.handle.net/1887/20898 holds various files of this Leiden University dissertation. Author: Jöris, Monique Maria Title: Challenges in unrelated hematopoietic stem cell transplantation.
Significance of the MHC
 CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
SNPrints: Defining SNP signatures for prediction of onset in complex diseases
 SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
Minimal Requirements for Histocompatibility & Immunogenetics Laboratory
 Minimal Requirements for Histocompatibility & Immunogenetics Laboratory The 4 th WBMT Congress and Workshop Riyadh, KSA - January 15-17, 2017 HLA Discovery, 1958 The Nobel Prize in Physiology or Medicine
Minimal Requirements for Histocompatibility & Immunogenetics Laboratory The 4 th WBMT Congress and Workshop Riyadh, KSA - January 15-17, 2017 HLA Discovery, 1958 The Nobel Prize in Physiology or Medicine
For more information about how to cite these materials visit
 Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Analysis of single gene effects 1. Quantitative analysis of single gene effects. Gregory Carey, Barbara J. Bowers, Jeanne M.
 Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
Supplementary Figure 1 Dosage correlation between imputed and genotyped alleles Imputed dosages (0 to 2) of 2-digit alleles (red) and 4-digit alleles
 Supplementary Figure 1 Dosage correlation between imputed and genotyped alleles Imputed dosages (0 to 2) of 2-digit alleles (red) and 4-digit alleles (green) of (A) HLA-A, HLA-B, (C) HLA-C, (D) HLA-DQA1,
Supplementary Figure 1 Dosage correlation between imputed and genotyped alleles Imputed dosages (0 to 2) of 2-digit alleles (red) and 4-digit alleles (green) of (A) HLA-A, HLA-B, (C) HLA-C, (D) HLA-DQA1,
Association mapping (qualitative) Association scan, quantitative. Office hours Wednesday 3-4pm 304A Stanley Hall. Association scan, qualitative
 Association mapping (qualitative) Office hours Wednesday 3-4pm 304A Stanley Hall Fig. 11.26 Association scan, qualitative Association scan, quantitative osteoarthritis controls χ 2 test C s G s 141 47
Association mapping (qualitative) Office hours Wednesday 3-4pm 304A Stanley Hall Fig. 11.26 Association scan, qualitative Association scan, quantitative osteoarthritis controls χ 2 test C s G s 141 47
Association between the CYP11B2 gene 344T>C polymorphism and coronary artery disease: a meta-analysis
 Association between the CYP11B2 gene 344T>C polymorphism and coronary artery disease: a meta-analysis Y. Liu, H.L. Liu, W. Han, S.J. Yu and J. Zhang Department of Cardiology, The General Hospital of the
Association between the CYP11B2 gene 344T>C polymorphism and coronary artery disease: a meta-analysis Y. Liu, H.L. Liu, W. Han, S.J. Yu and J. Zhang Department of Cardiology, The General Hospital of the
Example HLA-B and abacavir. Roujeau 2014
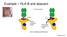 Example HLA-B and abacavir Roujeau 2014 FDA requires testing for abacavir Treatment with abacavir is generally well tolerated, but 5% of the patients experience hypersensitivity reactions that can be life
Example HLA-B and abacavir Roujeau 2014 FDA requires testing for abacavir Treatment with abacavir is generally well tolerated, but 5% of the patients experience hypersensitivity reactions that can be life
HLA and new technologies. Vicky Van Sandt
 HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
FONS Nové sekvenační technologie vklinickédiagnostice?
 FONS 2010 Nové sekvenační technologie vklinickédiagnostice? Sekvenování amplikonů Sequence capture Celogenomové sekvenování FONS 2010 Sekvenování amplikonů Amplicon sequencing - amplicon sequencing enables
FONS 2010 Nové sekvenační technologie vklinickédiagnostice? Sekvenování amplikonů Sequence capture Celogenomové sekvenování FONS 2010 Sekvenování amplikonů Amplicon sequencing - amplicon sequencing enables
Validation of the MIA FORA NGS FLEX Assay Using Buccal Swabs as the Sample Source
 S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. Introduction
S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. Introduction
Systems of Mating: Systems of Mating:
 8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
2) Cases and controls were genotyped on different platforms. The comparability of the platforms should be discussed.
 Reviewers' Comments: Reviewer #1 (Remarks to the Author) The manuscript titled 'Association of variations in HLA-class II and other loci with susceptibility to lung adenocarcinoma with EGFR mutation' evaluated
Reviewers' Comments: Reviewer #1 (Remarks to the Author) The manuscript titled 'Association of variations in HLA-class II and other loci with susceptibility to lung adenocarcinoma with EGFR mutation' evaluated
Histocompatibility Evaluations for HSCT at JHMI. M. Sue Leffell, PhD. Professor of Medicine Laboratory Director
 Histocompatibility Evaluations for HSCT at JHMI M. Sue Leffell, PhD Professor of Medicine Laboratory Director JHMI Patient Population Adults Peds NMDP data >20,000 HSCT JHMI HSCT Protocols Bone marrow
Histocompatibility Evaluations for HSCT at JHMI M. Sue Leffell, PhD Professor of Medicine Laboratory Director JHMI Patient Population Adults Peds NMDP data >20,000 HSCT JHMI HSCT Protocols Bone marrow
5/2/18. After this class students should be able to: Stephanie Moon, Ph.D. - GWAS. How do we distinguish Mendelian from non-mendelian traits?
 corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
Association between interleukin-17a polymorphism and coronary artery disease susceptibility in the Chinese Han population
 Association between interleukin-17a polymorphism and coronary artery disease susceptibility in the Chinese Han population G.B. Su, X.L. Guo, X.C. Liu, Q.T. Cui and C.Y. Zhou Department of Cardiothoracic
Association between interleukin-17a polymorphism and coronary artery disease susceptibility in the Chinese Han population G.B. Su, X.L. Guo, X.C. Liu, Q.T. Cui and C.Y. Zhou Department of Cardiothoracic
DEFINITIONS: POPULATION: a localized group of individuals belonging to the same species
 DEFINITIONS: POPULATION: a localized group of individuals belonging to the same species SPECIES: a group of populations whose individuals have the potential to interbreed and produce fertile offspring
DEFINITIONS: POPULATION: a localized group of individuals belonging to the same species SPECIES: a group of populations whose individuals have the potential to interbreed and produce fertile offspring
Title:The effect of CD14 and TLR4 gene polimorphisms on asthma phenotypes in adult Turkish asthma patients: a genetic study
 Author's response to reviews Title:The effect of CD14 and TLR4 gene polimorphisms on asthma phenotypes in adult Turkish asthma patients: a genetic study Authors: Fusun Sahin (fusunsahin19700@hotmail.com)
Author's response to reviews Title:The effect of CD14 and TLR4 gene polimorphisms on asthma phenotypes in adult Turkish asthma patients: a genetic study Authors: Fusun Sahin (fusunsahin19700@hotmail.com)
Type 1 diabetes is an autoimmune disease characterized
 ORIGINAL ARTICLE HLA Class I and Genetic Susceptibility to Type Diabetes Results From the Type Diabetes Genetics Consortium Janelle A. Noble, Ana Maria Valdes, Michael D. Varney, 3 Joyce A. Carlson, 4
ORIGINAL ARTICLE HLA Class I and Genetic Susceptibility to Type Diabetes Results From the Type Diabetes Genetics Consortium Janelle A. Noble, Ana Maria Valdes, Michael D. Varney, 3 Joyce A. Carlson, 4
IMMUNOLOGY. Elementary Knowledge of Major Histocompatibility Complex and HLA Typing
 IMMUNOLOGY Elementary Knowledge of Major Histocompatibility Complex and HLA Typing Tapasya Srivastava and Subrata Sinha Department of Biochemistry All India Institute of Medical Sciences New Delhi - 110029
IMMUNOLOGY Elementary Knowledge of Major Histocompatibility Complex and HLA Typing Tapasya Srivastava and Subrata Sinha Department of Biochemistry All India Institute of Medical Sciences New Delhi - 110029
An Introduction to Quantitative Genetics I. Heather A Lawson Advanced Genetics Spring2018
 An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
Section 8.1 Studying inheritance
 Section 8.1 Studying inheritance Genotype and phenotype Genotype is the genetic constitution of an organism that describes all the alleles that an organism contains The genotype sets the limits to which
Section 8.1 Studying inheritance Genotype and phenotype Genotype is the genetic constitution of an organism that describes all the alleles that an organism contains The genotype sets the limits to which
White Paper Estimating Genotype-Specific Incidence for One or Several Loci
 White Paper 23-01 Estimating Genotype-Specific Incidence for One or Several Loci Authors: Mike Macpherson Brian Naughton Andro Hsu Joanna Mountain Created: September 5, 2007 Last Edited: November 18, 2007
White Paper 23-01 Estimating Genotype-Specific Incidence for One or Several Loci Authors: Mike Macpherson Brian Naughton Andro Hsu Joanna Mountain Created: September 5, 2007 Last Edited: November 18, 2007
PopGen4: Assortative mating
 opgen4: Assortative mating Introduction Although random mating is the most important system of mating in many natural populations, non-random mating can also be an important mating system in some populations.
opgen4: Assortative mating Introduction Although random mating is the most important system of mating in many natural populations, non-random mating can also be an important mating system in some populations.
Tutorial on Genome-Wide Association Studies
 Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Patient Consult Report
 Patient Consult Report Clinical History Currently 36 years old History of firstborn son now about 5 years old followed by secondary recurrent pregnancy loss with 9 losses (all between 6 and 11 weeks) over
Patient Consult Report Clinical History Currently 36 years old History of firstborn son now about 5 years old followed by secondary recurrent pregnancy loss with 9 losses (all between 6 and 11 weeks) over
Transmission Disequilibrium Test in GWAS
 Department of Computer Science Brown University, Providence sorin@cs.brown.edu November 10, 2010 Outline 1 Outline 2 3 4 The transmission/disequilibrium test (TDT) was intro- duced several years ago by
Department of Computer Science Brown University, Providence sorin@cs.brown.edu November 10, 2010 Outline 1 Outline 2 3 4 The transmission/disequilibrium test (TDT) was intro- duced several years ago by
Genome-wide association studies for human narcolepsy and other complex diseases
 ETHZ-JST Joint WS Sept. 15-16, 2008 Genome-wide association studies for human narcolepsy and other complex diseases Katsushi Tokunaga Department of Human Genetics Graduate School of Medicine The University
ETHZ-JST Joint WS Sept. 15-16, 2008 Genome-wide association studies for human narcolepsy and other complex diseases Katsushi Tokunaga Department of Human Genetics Graduate School of Medicine The University
HLA Complex Genetics & Biology
 HLA Complex Genetics & Biology M. Tevfik DORAK Environmental & Occupational Health College of Public Health Florida International University Miami, USA http://www.dorak.info Mount Sinai Medical Center
HLA Complex Genetics & Biology M. Tevfik DORAK Environmental & Occupational Health College of Public Health Florida International University Miami, USA http://www.dorak.info Mount Sinai Medical Center
Bio 1M: Evolutionary processes
 Bio 1M: Evolutionary processes Evolution by natural selection Is something missing from the story I told last chapter? Heritable variation in traits Selection (i.e., differential reproductive success)
Bio 1M: Evolutionary processes Evolution by natural selection Is something missing from the story I told last chapter? Heritable variation in traits Selection (i.e., differential reproductive success)
Review Article Association between HLA-DQ Gene Polymorphisms and HBV-Related Hepatocellular Carcinoma
 Hindawi Gastroenterology Research and Practice Volume 2017, Article ID 7150386, 11 pages https://doiorg/101155/2017/7150386 Review Article Association between HLA-DQ Gene Polymorphisms and HBV-Related
Hindawi Gastroenterology Research and Practice Volume 2017, Article ID 7150386, 11 pages https://doiorg/101155/2017/7150386 Review Article Association between HLA-DQ Gene Polymorphisms and HBV-Related
Introduction to LOH and Allele Specific Copy Number User Forum
 Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Introduction to LOH and Allele Specific Copy Number User Forum Jonathan Gerstenhaber Introduction to LOH and ASCN User Forum Contents 1. Loss of heterozygosity Analysis procedure Types of baselines 2.
Allelic and Haplotype Frequencies of the p53 Polymorphisms in Brain Tumor Patients
 Physiol. Res. 51: 59-64, 2002 Allelic and Haplotype Frequencies of the p53 Polymorphisms in Brain Tumor Patients E. BIROŠ, I. KALINA, A. KOHÚT 1, E. BOGYIOVÁ 2, J. ŠALAGOVIČ, I. ŠULLA 3 Department of Medical
Physiol. Res. 51: 59-64, 2002 Allelic and Haplotype Frequencies of the p53 Polymorphisms in Brain Tumor Patients E. BIROŠ, I. KALINA, A. KOHÚT 1, E. BOGYIOVÁ 2, J. ŠALAGOVIČ, I. ŠULLA 3 Department of Medical
The MHC and Transplantation Brendan Clark. Transplant Immunology, St James s University Hospital, Leeds, UK
 The MHC and Transplantation Brendan Clark Transplant Immunology, St James s University Hospital, Leeds, UK Blood Groups Immunofluorescent staining has revealed blood group substance in the cell membranes
The MHC and Transplantation Brendan Clark Transplant Immunology, St James s University Hospital, Leeds, UK Blood Groups Immunofluorescent staining has revealed blood group substance in the cell membranes
appstats26.notebook April 17, 2015
 Chapter 26 Comparing Counts Objective: Students will interpret chi square as a test of goodness of fit, homogeneity, and independence. Goodness of Fit A test of whether the distribution of counts in one
Chapter 26 Comparing Counts Objective: Students will interpret chi square as a test of goodness of fit, homogeneity, and independence. Goodness of Fit A test of whether the distribution of counts in one
Exploring the Importance of Single Nucleotide Polymorphisms of HSPA9 in DNA of Sarcoma Patients
 University of New Hampshire University of New Hampshire Scholars' Repository Honors Theses and Capstones Student Scholarship Summer 2013 Exploring the Importance of Single Nucleotide Polymorphisms of HSPA9
University of New Hampshire University of New Hampshire Scholars' Repository Honors Theses and Capstones Student Scholarship Summer 2013 Exploring the Importance of Single Nucleotide Polymorphisms of HSPA9
Retrospective Genetic Analysis of Efficacy and Adverse Events in a Rheumatoid Arthritis Population Treated with Methotrexate and Anti-TNF-α
 Retrospective Genetic Analysis of Efficacy and Adverse Events in a Rheumatoid Arthritis Population Treated with Methotrexate and Anti-TNF-α Foti A 1, Lichter D 1, Shadick NA 2, Maher NE 2, Ginsburg GS
Retrospective Genetic Analysis of Efficacy and Adverse Events in a Rheumatoid Arthritis Population Treated with Methotrexate and Anti-TNF-α Foti A 1, Lichter D 1, Shadick NA 2, Maher NE 2, Ginsburg GS
Genetics and Genomics in Medicine Chapter 8 Questions
 Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
FTO Polymorphisms Are Associated with Obesity But Not with Diabetes in East Asian Populations: A Meta analysis
 BIOMEDICAL AND ENVIRONMENTAL SCIENCES 22, 449 457 (2009) www.besjournal.com FTO Polymorphisms Are Associated with Obesity But Not with Diabetes in East Asian Populations: A Meta analysis BO XI #, + AND
BIOMEDICAL AND ENVIRONMENTAL SCIENCES 22, 449 457 (2009) www.besjournal.com FTO Polymorphisms Are Associated with Obesity But Not with Diabetes in East Asian Populations: A Meta analysis BO XI #, + AND
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Mendel Short IGES 2003 Data Preparation. Eric Sobel. Department of of Human Genetics UCLA School of of Medicine
 Mendel Short Course @ IGES 2003 Data Preparation Eric Sobel Department of of Human Genetics UCLA School of of Medicine 02 November 2003 Mendel Short Course @ IGES Slide 1 Web Sites Mendel5: www.genetics.ucla.edu/software
Mendel Short Course @ IGES 2003 Data Preparation Eric Sobel Department of of Human Genetics UCLA School of of Medicine 02 November 2003 Mendel Short Course @ IGES Slide 1 Web Sites Mendel5: www.genetics.ucla.edu/software
MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH
 MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH TEJASWINI M. DHAWALE, M.D. HEME FELLOWS CONFERENCE NOVEMBER 08, 2013 CASE PRESENTATION 51 yo M with history of MDS (unilinear
MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH TEJASWINI M. DHAWALE, M.D. HEME FELLOWS CONFERENCE NOVEMBER 08, 2013 CASE PRESENTATION 51 yo M with history of MDS (unilinear
Roadmap. Inbreeding How inbred is a population? What are the consequences of inbreeding?
 1 Roadmap Quantitative traits What kinds of variation can selection work on? How much will a population respond to selection? Heritability How can response be restored? Inbreeding How inbred is a population?
1 Roadmap Quantitative traits What kinds of variation can selection work on? How much will a population respond to selection? Heritability How can response be restored? Inbreeding How inbred is a population?
Myoglobin A79G polymorphism association with exercise-induced skeletal muscle damage
 Myoglobin A79G polymorphism association with exercise-induced skeletal muscle damage T. Cui and M.S. Jiang College of Physical Education, Shandong University of Finance and Economics, Ji nan, Shandong,
Myoglobin A79G polymorphism association with exercise-induced skeletal muscle damage T. Cui and M.S. Jiang College of Physical Education, Shandong University of Finance and Economics, Ji nan, Shandong,
RNA based high-resolution HLA sequencing based typing
 Chapter 2 RNA based high-resolution HLA sequencing based typing Judith Reinders, Anna Houben, Alain van Mill, Erik Rozemuller, Jan van den Tweel and Marcel Tilanus Department of Pathology, University Medical
Chapter 2 RNA based high-resolution HLA sequencing based typing Judith Reinders, Anna Houben, Alain van Mill, Erik Rozemuller, Jan van den Tweel and Marcel Tilanus Department of Pathology, University Medical
How to Find an Unrelated Donor Theory & Technology
 How to Find an Unrelated Donor Theory & Technology Carlheinz R. Müller Zentrales Knochenmarkspender-Register für die Bundesrepublik Deutschland (ZKRD) Ulm, Germany How to Find an Unrelated Donor HLA-Basics
How to Find an Unrelated Donor Theory & Technology Carlheinz R. Müller Zentrales Knochenmarkspender-Register für die Bundesrepublik Deutschland (ZKRD) Ulm, Germany How to Find an Unrelated Donor HLA-Basics
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
New trends in donor selection in Europe: "best match" versus haploidentical. Prof Jakob R Passweg
 New trends in donor selection in Europe: "best match" versus haploidentical Prof Jakob R Passweg HSCT change in donor type: 1990-2015 9000 H S C T 8000 7000 6000 5000 4000 HLA identical sibling/twin Haplo-identical
New trends in donor selection in Europe: "best match" versus haploidentical Prof Jakob R Passweg HSCT change in donor type: 1990-2015 9000 H S C T 8000 7000 6000 5000 4000 HLA identical sibling/twin Haplo-identical
HLA-DRB1*1101: A Significant Risk Factor for Sarcoidosis in Blacks and Whites
 Am. J. Hum. Genet. 73:720 735, 2003 HLA-DRB1*1101: A Significant Risk Factor for Sarcoidosis in Blacks and Whites Milton D. Rossman, 1 Bruce Thompson, 3 Margaret Frederick, 3 Mary Maliarik, 4 Michael C.
Am. J. Hum. Genet. 73:720 735, 2003 HLA-DRB1*1101: A Significant Risk Factor for Sarcoidosis in Blacks and Whites Milton D. Rossman, 1 Bruce Thompson, 3 Margaret Frederick, 3 Mary Maliarik, 4 Michael C.
6/19/2012. Who is in the room today? What is your level of understanding of Donor Antigens and Candidate Unacceptables in KPD?
 6/19/212 KPD Webinar Series: Part 3 Demystifying the OPTN Kidney Paired Donation Pilot Program Unacceptable HLA antigens: The key to finding a compatible donor for your patient M. Sue Leffell, Ph.D. J.
6/19/212 KPD Webinar Series: Part 3 Demystifying the OPTN Kidney Paired Donation Pilot Program Unacceptable HLA antigens: The key to finding a compatible donor for your patient M. Sue Leffell, Ph.D. J.
Research Article Computational Approaches to Facilitate Epitope-Based HLA Matching in Solid Organ Transplantation
 Hindawi Journal of Immunology Research Volume 2017, Article ID 9130879, 9 pages https://doi.org/10.1155/2017/9130879 Research Article Computational Approaches to Facilitate Epitope-Based HLA Matching in
Hindawi Journal of Immunology Research Volume 2017, Article ID 9130879, 9 pages https://doi.org/10.1155/2017/9130879 Research Article Computational Approaches to Facilitate Epitope-Based HLA Matching in
Selection at one locus with many alleles, fertility selection, and sexual selection
 Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/49010 holds various files of this Leiden University dissertation. Author: Heide, A. van der Title: Unravelling narcolepsy : from pathophysiology to measuring
Cover Page The handle http://hdl.handle.net/1887/49010 holds various files of this Leiden University dissertation. Author: Heide, A. van der Title: Unravelling narcolepsy : from pathophysiology to measuring
HOST-PARASITE INTERPLAY
 HOST-PARASITE INTERPLAY Adriano Casulli EURLP, ISS (Rome, Italy) HOST-PARASITE INTERPLAY WP3 (parasite virulence vs human immunity) (Parasite) Task 3.1: Genotypic characterization Task 3.6: Transcriptome
HOST-PARASITE INTERPLAY Adriano Casulli EURLP, ISS (Rome, Italy) HOST-PARASITE INTERPLAY WP3 (parasite virulence vs human immunity) (Parasite) Task 3.1: Genotypic characterization Task 3.6: Transcriptome
Supplementary Figures
 Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
National Disease Research Interchange Annual Progress Report: 2010 Formula Grant
 National Disease Research Interchange Annual Progress Report: 2010 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Disease Research Interchange received $62,393
National Disease Research Interchange Annual Progress Report: 2010 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Disease Research Interchange received $62,393
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
Supplementary Figure 1. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations.
 Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
(b) What is the allele frequency of the b allele in the new merged population on the island?
 2005 7.03 Problem Set 6 KEY Due before 5 PM on WEDNESDAY, November 23, 2005. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Two populations (Population One
2005 7.03 Problem Set 6 KEY Due before 5 PM on WEDNESDAY, November 23, 2005. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Two populations (Population One
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Complex HLA-DR and -DQ Interactions Confer Risk of Narcolepsy- Cataplexy in Three Ethnic Groups
 Am. J. Hum. Genet. 68:686 699, 001 Complex HLA-DR and -DQ Interactions Confer Risk of Narcolepsy- Cataplexy in Three Ethnic Groups Emmanuel Mignot, 1 Ling Lin, 1 William Rogers, 1 Yutaka Honda 4 Xiaohong
Am. J. Hum. Genet. 68:686 699, 001 Complex HLA-DR and -DQ Interactions Confer Risk of Narcolepsy- Cataplexy in Three Ethnic Groups Emmanuel Mignot, 1 Ling Lin, 1 William Rogers, 1 Yutaka Honda 4 Xiaohong
Additive and interaction effects at three amino acid positions in HLA-DQ and HLA-DR molecules drive type 1 diabetes risk
 Additive and interaction effects at three amino acid positions in HLA-DQ and HLA-DR molecules drive type 1 diabetes risk Xinli Hu 1-6 *, Aaron J. Deutsch 1-5 *, Tobias L. Lenz 2,7, Suna Onengut-Gumuscu
Additive and interaction effects at three amino acid positions in HLA-DQ and HLA-DR molecules drive type 1 diabetes risk Xinli Hu 1-6 *, Aaron J. Deutsch 1-5 *, Tobias L. Lenz 2,7, Suna Onengut-Gumuscu
Performing. linkage analysis using MERLIN
 Performing linkage analysis using MERLIN David Duffy Queensland Institute of Medical Research Brisbane, Australia Overview MERLIN and associated programs Error checking Parametric linkage analysis Nonparametric
Performing linkage analysis using MERLIN David Duffy Queensland Institute of Medical Research Brisbane, Australia Overview MERLIN and associated programs Error checking Parametric linkage analysis Nonparametric
Association between ERCC1 and ERCC2 gene polymorphisms and susceptibility to pancreatic cancer
 Association between ERCC1 and ERCC2 gene polymorphisms and susceptibility to pancreatic cancer M.G. He, K. Zheng, D. Tan and Z.X. Wang Department of Hepatobiliary Surgery, Nuclear Industry 215 Hospital
Association between ERCC1 and ERCC2 gene polymorphisms and susceptibility to pancreatic cancer M.G. He, K. Zheng, D. Tan and Z.X. Wang Department of Hepatobiliary Surgery, Nuclear Industry 215 Hospital
Mendelian Genetics using Fast Plants Report due Sept. 15/16. Readings: Mendelian genetics: Hartwell Chapter 2 pp , Chapter 5 pp
 1 Biology 423L Sept. 1/2 Mendelian Genetics using Fast Plants Report due Sept. 15/16. Readings: Mendelian genetics: Hartwell Chapter 2 pp. 13-27, Chapter 5 pp. 127-130. FASTPLANTS: Williams et al. (1986)
1 Biology 423L Sept. 1/2 Mendelian Genetics using Fast Plants Report due Sept. 15/16. Readings: Mendelian genetics: Hartwell Chapter 2 pp. 13-27, Chapter 5 pp. 127-130. FASTPLANTS: Williams et al. (1986)
University of Groningen. Metabolic risk in people with psychotic disorders Bruins, Jojanneke
 University of Groningen Metabolic risk in people with psychotic disorders Bruins, Jojanneke IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from
University of Groningen Metabolic risk in people with psychotic disorders Bruins, Jojanneke IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.
 Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Outline. How archaics shaped the modern immune system. The immune system. Innate immune system. Adaptive immune system
 Outline How archaics shaped the modern immune system Alan R. Rogers February 14, 2018 Why the immune system is sensitive to archaic introgression. Archaic MHC alleles The OAS1 innate immunity locus 1 /
Outline How archaics shaped the modern immune system Alan R. Rogers February 14, 2018 Why the immune system is sensitive to archaic introgression. Archaic MHC alleles The OAS1 innate immunity locus 1 /
Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population
 Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population J. Zhu 1 *, F. He 2 *, D.D. Zhang 2 *, J.Y. Yang 2, J. Cheng 1, R. Wu 1, B. Gong 2, X.Q. Liu
Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population J. Zhu 1 *, F. He 2 *, D.D. Zhang 2 *, J.Y. Yang 2, J. Cheng 1, R. Wu 1, B. Gong 2, X.Q. Liu
Available online at , 2(1):27-37
 Available online at www.apjbg.com Asia-Pacific Journal of Blood Types and Genes 2018, 2(1):27-37 APJBG Allele and haplotype frequencies of human leukocyte antigen-a, -B, -C, -DRB1, and -DQB1 in Chinese
Available online at www.apjbg.com Asia-Pacific Journal of Blood Types and Genes 2018, 2(1):27-37 APJBG Allele and haplotype frequencies of human leukocyte antigen-a, -B, -C, -DRB1, and -DQB1 in Chinese
