Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors
|
|
|
- Nickolas Preston
- 5 years ago
- Views:
Transcription
1 Hypothesis Volume 9(11) Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors Monika 1 *, Janmeet Kour 1 & Kulwinder Singh 2 1Department of Biotechnology, Mata Gujri College, Fatehgarh Sahib , Punjab, India; 2 Department of Biotechnology, Punjabi University, Patiala , Punjab, India; Monika monika187@rediffmail.com; Phone: ; *Corresponding author Received June 03, 2013; Accepted June 04, 2013; Published June 29, 2013 Abstract: The leukotrienes constitute a group of arachidonic acid-derived compounds with biologic activities suggesting important roles in inflammation and immediate hypersensitivity. Epidermis-type lipoxygenase-3 (ALOXE3), a distinct subclass within the multigene family of mammalian lipoxygenases, is a novel isoenzyme involved in the metabolism of leukotrienes and plays a very important role in skin barrier functions. Lipoxygenase selective inhibitors such as azelastine and zileuton are currently used to reduce inflammatory response. Nausea, pharyngolaryngeal pain, headache, nasal burning and somnolence are the most frequently reported adverse effects of these drugs. Therefore, there is still a need to develop more potent lipoxygenase inhibitors. In this paper, we report the screening of various compounds from the ZINC database (contains over 21 million compounds) using the Molegro Virtual Docker software against the ALOXE3 protein. Screening was performed using molecular constraints tool to filter compounds with physico-chemical properties similar to the 1N8Q bound ligand protocatechuic acid. The analysis resulted in 4319 Lipinski compliant hits which are docked and scored to identify structurally novel ligands that make similar interactions to those of known ligands or may have different interactions with other parts of the binding site. Our screening approach identified four molecules ZINC ; ZINC ; ZINC & ZINC with MolDock score of , , & kcal/mol, respectively. Their energy scores were better than the 1N8Q bound co-crystallized ligand protocatechuic acid (with MolDock score of kcal/mol). All the ligands were docked within the binding pocket forming interactions with amino acid residues. Keywords: lipoxygenase, ZINC database, virtual screening, Molegro Virtual Docker. Background: Lipoxygenases are a class of enzymes that convert arachidonic, linoleic, and other polyunsaturated fatty acid into biologically active metabolites involved in the inflammatory and immune responses [1]. There are five active LOXs found in human beings: 5-LOX, 12S-LOX, 12R-LOX, 15-LOX-1, and 15-LOX-2. A sixth gene family member, epidermal LOX type 3 (elox3, gene symbol ALOXE3) was described first in the mouse [2], and in humans in 2001 [3]. Widely expressed in animals and plants, and also in some bacteria and fungi, LOX enzymes participate in diverse physiological and pathological processes. Mammalian 12R-LOX and epidermal lipoxygenase-3 (elox3), for example, play an indispensable role in skin barrier formation [4, 5]. A distinctive characteristic of LOX enzymes in general is that the resting enzyme requires activation before catalytic reactions with fatty acids and molecular oxygen can begin [6]. elox3 is an enzyme that in humans is encoded by the ALOXE3 gene. Unlike other lipoxygenases, the elox3 enzyme does not act directly on fatty acids. Instead, it processes the product of another lipoxygenase reaction, a fatty acid hydroperoxide. The substance produced is later converted to a signaling molecule that is involved in the growth and division (proliferation) and specialization (differentiation) of skin cells. The elox3 enzyme is thought to play a role in the Bioinformation 9(11): (2013) Biomedical Informatics
2 formation and maintenance of the fat (lipid) membrane of the cells that make up the outermost layer of the skin (the epidermis). The epidermis helps prevent water loss, regulates body temperature, and protects against infection. At least 10 mutations in the ALOXE3 gene have been found to cause nonbullous congenital ichthyosiform erythroderma (NBCIE). Most of these mutations change single protein building blocks (amino acids) in the elox3 enzyme. Many ALOXE3 gene mutations lead to the production of a nonfunctional elox3 enzyme, which impairs the formation of the lipid membrane of cells within the epidermis. Problems with this protective barrier underlie the skin abnormalities and other features of NBCIE [7]. approaches can be highly effective in reducing the dataset to be docked to the order of 10 3 to 10 4 compounds [14]. Several classes of compounds having selective lipoxygenase inhibitory activity have been reported in the literature. Many 5-LOX inhibitors have subsequently been developed, and several have displayed efficacy in asthma models, such as allergen-induced bronchoconstriction. For example, 1, 5- diarylpyrazoles, indolizines and indoles [8]. However, evidence suggests that adverse reactions such as neuropsychiatric events, including sleep disorders, behavioral changes, headache, nasal burning and somnolence are associated with prolonged use of lipoxygenase selective inhibitors like zileuton and azelastine [9, 10]. Thus, there is still a need for novel, selective, and potent inhibitors of active LOXs with an improved profile compared to current lipoxygenase inhibitors. Traditional synthesis of a series of new compounds utilizing combinatorial chemistry and high-throughput screening can be carried out at high cost and also are time consuming; whereas on the other hand, screening small molecule databases for novel compounds represents an alternative process. Docking various ligands to the protein of interest followed by scoring to determine the affinity of binding and to reveal the strength of interaction has become increasingly important in the context of drug discovery. Screening large databases of compounds can provide a feasible, alternative technique against highthroughput screening, but depends on the fast and accuracy of the docking algorithm. Hence, in this paper we report screening a library of compounds from ZINC database [11] against elox3 protein, 1N8Q (PDB ID) with bound ligand protocatechuic acid extracted from protein data bank, by utilizing a fast, exhaustive docking software Molegro Virtual Docker [12]. In recent years, virtual screening has reached a status of a dynamic and lucrative technology in probing for novel drug like compounds or so called hits in the pharmaceutical industry [13]. The technique employed is based on the prediction of binding modes and binding affinities of each compound in the dataset by means of docking to an X-ray crystallographic structure. Virtual screening utilizes docking and scoring of each compound from a dataset. It is important to visualize the docked poses of high-scoring compounds because many ligands are docked in different orientations and may often miss interactions that are known to be important for the target receptor. This sort of study becomes more difficult as the size of the dataset increases. Therefore, an alternative approach is to eliminate unpromising compounds before docking by restricting the dataset to drug like compounds, by filtering the dataset based on appropriate property and substructural features and by performing diversity analysis. These Figure 1: Diagram illustrating the interactions between 1N8Q protein and protocatechuic acid. Legend: black dashed lines - hydrogen bonds [15]. Methodology: Receptor X-ray structure The 3D coordinates of the crystal structure of elox3 in complex with protocatechuic acid (PDB code: 1N8Q) was selected as the receptor model in virtual screening program. We used the chemical compound library, ZINC database and the docking program Molegro Virtual Docker for the study. Figure 1, illustrating the hydrogen bond interactions between protocatechuic acid and elox3 protein (PDB code: 1N8Q). Ligand ZINC database ZINC, an acronym for ZINC is not commercial, a free database for virtual screening contains over 21 million compounds in ready-to-dock, 3D formats, available at the URL Molecules in ZINC are annotated by molecular property that include molecular weight, number of rotatable bonds, calculated log P, number of hydrogen-bond donors, hydrogen-bond acceptors, chiral centers, chiral double bonds (E/Z isomerism), polar and apolar desolvation energy (in kcal/mol), net charge and rigid fragments. The database contains lead-like molecules, having molecular weight in the range 150 to 350 with calculated log P < 4, number of hydrogen bond donors 3 three, and number of hydrogen-bond acceptors 6. ZINC provides several search criteria such as molecular property constraint, ZINC codes, vendor based, and molecular substructure search [11]. Molegro Virtual Docker Molegro is a Danish company, developing novel high-quality drug discovery and data mining software, founded in 2005 by Bioinformation 9(11): (2013) Biomedical Informatics
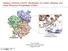
3 Rene Thomsen and Mikael Hvidtfeldt Christensen [12]. We used the template docking available in Molegro virtual docker and evaluated MolDock, rerank and protein-ligand interaction scores from MolDock and MolDock [GRID] options. Template docking is based on extracting the chemical properties like the pharmacophore elements of a ligand bound in the active site and using that information for docking structurally similar analogs. We used the default settings, including a grid resolution of 0.30 Å for grid generation and a 15 Å radius from the template as the binding site. We used the MolDock optimizer as a search algorithm, and the number of runs was set to 10. A population size of 50, maximum iteration of 2000, scaling factor of 0.50, crossover rate of 0.90 and a variationbased termination scheme for parameter settings were used. The maximum number of poses was set to a default value of 5. We docked these 4319 compounds using Molegro virtual docker and evaluated binding compatibility with the receptor based on docked energy (in kcal/mol). The docking tool generated 50 conformers for each docked molecule in about 1-2 minutes of CPU time. The virtual screening technique employed in this study identified diverse, yet specific ligands that bind in a comparable manner similar to protocatechuic acid binding for elox3. This approach identified four compounds ZINC ([5-(5,7-dimethyl-1H-indol-2-yl)- 1H-pyrrol-3-yl] methanamine); ZINC (4-(4-methyl- 1H-indol-2-yl)-1H-pyrrol-3-amine); ZINC ([5-(1Hindol-2-yl)-1H-pyrrol-3-yl]methanamine) & ZINC (4- [(E)-2-(2- thienyl)vinyl]benzene-1,2-diamine) from the ZINC database for compatible binding with elox3 Table 1 (see supplementary material). Screening Before screening ZINC database, the docking protocol was validated. 1N8Q protein bound ligand protocatechuic acid was docked into the binding pocket to obtain the docked pose and the RMSD (Root Mean Square Deviation) of all atoms between these two conformations indicating that the parameters for docking simulation were good in reproducing the X-ray crystal structure. Therefore, the ZINC database was screened for compounds similar to protocatechuic acid structural features (structure based search) and by providing molecular constraints (property based search). ZINC database was also screened for compounds having properties similar to the known inhibitors of LOXs. The physicochemical properties such as log P value, H-bond donors, Hbond acceptors, molecular weight and rotational bonds, for protocatechuic acid ligand and known inhibitors of LOXs were calculated using the molinspiration server [16]. Discussion: Lipinski's "Rule of 5" states, that most "drug-like" molecules have log P < or = 5, number of hydrogen bond acceptors < or = 10, number of hydrogen bond donors < or = 5 and molecular weight < or = 500 g/mol. Molecules violating more than one of these rules may have problems with bioavailability. The rule is called "Rule of 5", because the border values are 5, 500, 2*5, and 5 [17]. Number of rotatable bonds is a simple topological parameter which is a measure of molecular flexibility. It has been shown to be a very good descriptor of oral bioavailability of drugs [18]. Rotatable bond is defined as any single non-ring bond, bounded to nonterminal heavy (i.e., non-hydrogen) atom. Amide C-N bonds are not considered because of their high rotational energy barrier. The physico-chemical properties (log P value, H-bond acceptors, H-bond donors, molecular weight and rotational bonds) for protocatechuic acid ligand and known inhibitors of LOXs (aromadedrin, azelastin, robinetin, zileuton, caffeic and eriodictyol) were calculated using the molinspiration server. For these compounds range of calculated log P values was from 0.88 to 4.222, number of H- bond acceptors ranged from 4 to 7, number of H-bond donors ranged from 1 to 5, molecular weight ranged from g/mol to g/mol and number of rotational bonds ranged from 1 to 3. We searched the ZINC database based on range values of properties of protocatechuic acid and known inhibitors of LOXs and identified 4319 hits that were Lipinski compliants. Figure 2: Schematic diagram illustrating the docking of ZINC (pink) (ligand with best MolDock score) to elox3 protein (pdb id 1N8Q) to form a protein-ligand complex. elox3 co-crystallized protocatechuic acid ligand resulted in MolDock score of kcal/mol. Therefore, any molecule from the dataset which shows a score lower than kcal/mol would be regarded as ligands with higher binding affinity. In other words, these four zinc ligands would act as inhibitors against elox3 protein and such screening analysis forms the basis when millions of ligands are available in compound libraries such as the ZINC database. The docked energies of the four zinc ligands are given in (Table 1). Active site of elox3 offers many different binding modes for these compounds as they are strongly dependent on the attached substituent. All the ligands were docked deeply within the binding pocket region forming interactions with Thr337, Ser326, Lys332 and Glu333. In (Table 1) the binding affinities and the possible number of interactions based on chemical property identifiers are reported. Our screening approach identified four molecules ZINC (MolDock score of kcal/mol); ZINC (MolDock score of kcal/mol); ZINC (MolDock score of kcal/mol) and ZINC (MolDock score of kcal/mol) from the ZINC database. Their energy scores were better than the 1N8Q bound co-crystallized ligand Bioinformation 9(11): (2013) Biomedical Informatics
4 protocatechuic acid with MolDock score of kcal/mol. Figure 2, illustrating the protein-ligand complex between ZINC and elox3 protein (PDB code: 1N8Q). Conclusion: Finding novel compounds as starting points for lead optimization is a major challenge in drug discovery. Virtual screening is a powerful technique for identifying hit molecules as starting points for medicinal chemistry. The number of methods and software which use the ligand and target-based virtual screening approaches is increasing at a rapid pace. It has been clearly demonstrated that the approach utilized in this study is successful in finding four novel elox3 inhibitors from the ZINC database. ZINC , in particular, showed high binding affinity with MolDock score of kcal/mol against 1N8Q. The ligand was docked deeply within the binding pocket region forming hydrogen bond interaction with Thr337. Therefore, this study states the importance of small molecule libraries and their use to enhance drug discovery process prior synthesis. This approach to screen novel compounds as elox3 inhibitors from ZINC database depends on various parameters such as Lipinski s rule of 5, pharmacophoric groups attached on the ligand, size of the dataset and compound libraries among others. Further, work can be extended to study the receptor ligand interactions experimentally and evaluation of their biological activity would help in designing compounds based on virtual screening techniques. Acknowledgment: We would like to extend our heartfelt thanks to Molegro ApS for giving us a fully functional trial version for a period of 30 days during which all the in-silico docking work was carried out. References: [1] Catalano A & Procopio A, Histol Histopathol : 969 [PMID: ] [2] Kinzig A et al. Genomics : 158 [PMID: ] [3] Krieg P et al. Genomics : 323 [PMID: ] [4] Brash AR et al. FEBS J : 3494 [PMID: ] [5] Furstenberger G et al. Prostaglandins Other Lipid Mediat : 128 [PMID: ] [6] Ford-Hutchinson AW et al. Annu Rev Biochem : 383 [PMID: ] [7] [8] Hofmann B & Steinhilber D, Expert Opin Ther Pat [PMID: ] [9] [10] Corren J et al. Clin Ther : 543 [PMID: ] [11] Irwin JJ & Shoichet BK, J Chem Inf Model : 177 [PMID: ] [12] Thomsen R & Christensen MH, J Med Chem : 3315 [PMID: ] [13] Shoichet BK, Nature : 862 [PMID: ] [14] Warren GL et al. J Med Chem : 5912 [PMID: ] [15] d=1n8q [16] [17] Lipinski CA et al. Adv Drug Deliv Rev : 3 [PMID: ] [18] Veber DF et al. J Med Chem : 2615 [PMID: ] Edited by P Kangueane Citation: Monika et al. Bioinformation 9(11): (2013) License statement: This is an open-access article, which permits unrestricted use, distribution, and reproduction in any medium, for non-commercial purposes, provided the original author and source are credite Bioinformation 9(11): (2013) Biomedical Informatics
5 Supplementary material: Table 1: Interaction parameters of elox3 with the predicted ZINC ligands and protocatechuic acid. S. No. Compound Structure MolDock Score (kcal/mol) 1 protocatechuic acid Rerank Score HBond Interacting residues His523, Gln514 2 ZINC Thr337 3 ZINC Ser326, Thr337 4 ZINC Lys332, Glu333 5 ZINC Glu333 Bioinformation 9(11): (2013) Biomedical Informatics
PHARMACOPHORE MODELING
 CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
A Combinatorial approach: To design inhibitory molecules on Hemagglutinin protein of H1N1 virus (Swine Flu)
 www.bioinformation.net Hypothesis Volume 9(11) A Combinatorial approach: To design inhibitory molecules on Hemagglutinin protein of H1N1 virus (Swine Flu) Chekkara Venkata Satya Siva Prasad*, Kamal Kumar
www.bioinformation.net Hypothesis Volume 9(11) A Combinatorial approach: To design inhibitory molecules on Hemagglutinin protein of H1N1 virus (Swine Flu) Chekkara Venkata Satya Siva Prasad*, Kamal Kumar
Asian Journal of Biomedical and Pharmaceutical Sciences 1 (6) 2012, 01-05
 Asian Journal of Biomedical and Pharmaceutical Sciences 1 (6) 2012, 01-05 RESEARCH ARTICLE ISSN 2249-622X Virtual Screening of Xanthones in Combating Malaria Targeting Plasmodium Falciparum Erythrocyte
Asian Journal of Biomedical and Pharmaceutical Sciences 1 (6) 2012, 01-05 RESEARCH ARTICLE ISSN 2249-622X Virtual Screening of Xanthones in Combating Malaria Targeting Plasmodium Falciparum Erythrocyte
Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma (PPARγ)
 3rd International Conference on Computation for Science and Technology (ICCST-3) Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma
3rd International Conference on Computation for Science and Technology (ICCST-3) Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma
Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids
 www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
Trans-10,cis-12 Conjugated Linoleic Acid a Novel Inhibitor of Inflammatory Mediators
 American Journal of Bioinformatics Research 2016, 6(2): 27-31 DOI: 10.5923/j.bioinformatics.20160602.01 Trans-10,cis-12 Conjugated Linoleic Acid a Novel Inhibitor of Inflammatory Mediators Rashmi H. K.,
American Journal of Bioinformatics Research 2016, 6(2): 27-31 DOI: 10.5923/j.bioinformatics.20160602.01 Trans-10,cis-12 Conjugated Linoleic Acid a Novel Inhibitor of Inflammatory Mediators Rashmi H. K.,
Molecular Docking Studies of Shc1 Protein: A Drug Target to Treat Obesity
 IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 9, Issue 6 Ver. III (Nov -Dec. 2014), PP 63-68 Molecular Docking Studies of Shc1 Protein: A Drug
IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 9, Issue 6 Ver. III (Nov -Dec. 2014), PP 63-68 Molecular Docking Studies of Shc1 Protein: A Drug
Synopsis of the thesis on
 Synopsis of the thesis on COMPUTATIONAL INVESTIGATION OF PROTEIN-LIGAND INTERACTIONS IN ANTI-DIABETIC AGENTS A dissertation submitted in the partial fulfillment for the award of the degree of DOCTOR OF
Synopsis of the thesis on COMPUTATIONAL INVESTIGATION OF PROTEIN-LIGAND INTERACTIONS IN ANTI-DIABETIC AGENTS A dissertation submitted in the partial fulfillment for the award of the degree of DOCTOR OF
Computer Aided Docking Studies on Anticancer Drugs for Breast Cancer
 International Journal of Information & Computation Technology. ISSN 0974-2239 Volume 2, Number 1 (2012), pp. 49-55 International Research Publications House http://www. ripublication.com Computer Aided
International Journal of Information & Computation Technology. ISSN 0974-2239 Volume 2, Number 1 (2012), pp. 49-55 International Research Publications House http://www. ripublication.com Computer Aided
INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER
 ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
open access Hypothesis Volume 10(7) Rajesh Kumar Pathak 1, 2, Mamta Baunthiyal 1 *, Gohar Taj 2 & Anil Kumar 2
 www.bioinformation.net Hypothesis Volume 10(7) Virtual screening of natural inhibitors to the predicted HBx protein structure of Hepatitis B Virus using molecular docking for identification of potential
www.bioinformation.net Hypothesis Volume 10(7) Virtual screening of natural inhibitors to the predicted HBx protein structure of Hepatitis B Virus using molecular docking for identification of potential
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Virtual screening of 2,3-disubstituted-4(3H)- quinazolinones possessing benzenesulfonamide moiety for COX-2 inhibitor
 www.bioinformation.net Hypothesis Volume 7(5) Virtual screening of 2,3-disubstituted-4(3H)- quinazolinones possessing benzenesulfonamide moiety for COX-2 inhibitor Hayun 1 *, Arry Yanuar 1, Muhammad Hanafi
www.bioinformation.net Hypothesis Volume 7(5) Virtual screening of 2,3-disubstituted-4(3H)- quinazolinones possessing benzenesulfonamide moiety for COX-2 inhibitor Hayun 1 *, Arry Yanuar 1, Muhammad Hanafi
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
A HLA-DRB supertype chart with potential overlapping peptide binding function
 A HLA-DRB supertype chart with potential overlapping peptide binding function Arumugam Mohanapriya 1,2, Satish Nandagond 1, Paul Shapshak 3, Uma Kangueane 1, Pandjassarame Kangueane 1, 2 * 1 Biomedical
A HLA-DRB supertype chart with potential overlapping peptide binding function Arumugam Mohanapriya 1,2, Satish Nandagond 1, Paul Shapshak 3, Uma Kangueane 1, Pandjassarame Kangueane 1, 2 * 1 Biomedical
Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II inhibitors
 International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol. 3, No.3,pp 1622-1627, July-Sept 2011 Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II
International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol. 3, No.3,pp 1622-1627, July-Sept 2011 Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II
NIH Public Access Author Manuscript J Am Chem Soc. Author manuscript; available in PMC 2008 September 29.
 NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
Clinical Study on Serum Nitric oxide levels in Osteoarthritis and Structure-Based Drug Design of New Inducible Nitric Oxide Synthase inhibitors
 World Journal of Pharmaceutical Sciences ISS (Print): 2321-3310; ISS (Online): 2321-3086 Published by Atom and Cell Publishers All Rights Reserved Available online at: http://www.wjpsonline.com/ Research
World Journal of Pharmaceutical Sciences ISS (Print): 2321-3310; ISS (Online): 2321-3086 Published by Atom and Cell Publishers All Rights Reserved Available online at: http://www.wjpsonline.com/ Research
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor!
 Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Molecular docking approaches in identification of High affinity inhibitors of Human SMO receptor
 www.bioinformation.net Hypothesis Volume 10(12) Molecular docking approaches in identification of High affinity inhibitors of Human SMO receptor Uday Raj Akare 1, Srinivas Bandaru 2, Uzma Shaheen 2, Pramod
www.bioinformation.net Hypothesis Volume 10(12) Molecular docking approaches in identification of High affinity inhibitors of Human SMO receptor Uday Raj Akare 1, Srinivas Bandaru 2, Uzma Shaheen 2, Pramod
Structure-based analysis of curcumin and conventional drugs targeting tumor-inducing protein PHF20
 www.bioinformation.net Volume 14(9) Hypothesis Structure-based analysis of curcumin and conventional drugs targeting tumor-inducing protein PHF20 Vibha Agrawal 1, Aradhya Mishra 1, Shivani Tiwari 1, Akhileshwar
www.bioinformation.net Volume 14(9) Hypothesis Structure-based analysis of curcumin and conventional drugs targeting tumor-inducing protein PHF20 Vibha Agrawal 1, Aradhya Mishra 1, Shivani Tiwari 1, Akhileshwar
COMPARATIVE VEGF RECEPTOR TYROSINE KINASE MODELING FOR THE DEVELOPMENT OF HIGHLY SPECIFIC INHIBITORS OF TUMOR ANGIOGENESIS
 243 COMPARATIVE VEGF RECEPTOR TYROSINE KINASE MODELING FOR THE DEVELOPMENT OF HIGHLY SPECIFIC INHIBITORS OF TUMOR ANGIOGENESIS ULRIKE SCHMIDT 1 JESSICA AHMED 1 ulrike.schmidt@charite.de jessica.ahmed@charite.de
243 COMPARATIVE VEGF RECEPTOR TYROSINE KINASE MODELING FOR THE DEVELOPMENT OF HIGHLY SPECIFIC INHIBITORS OF TUMOR ANGIOGENESIS ULRIKE SCHMIDT 1 JESSICA AHMED 1 ulrike.schmidt@charite.de jessica.ahmed@charite.de
DESIGNING, DOCKING AND TOXICITY STUDIES OF NOVEL HIV-1 PROTEASE INHIBITORS
 IMPACT: International Journal of Research in Applied atural and Social Sciences (IMPACT: IJRASS) ISS(E): 2321-8851; ISS(P): 2347-4580 Vol. 3, Issue 9, Sep 2015, 139-146 Impact Journals DESIGIG, DCKIG AD
IMPACT: International Journal of Research in Applied atural and Social Sciences (IMPACT: IJRASS) ISS(E): 2321-8851; ISS(P): 2347-4580 Vol. 3, Issue 9, Sep 2015, 139-146 Impact Journals DESIGIG, DCKIG AD
Protein Refinement in Discovery Studio 1.7. Presented by: Francisco Hernandez-Guzman, PhD
 Protein Refinement in Discovery Studio 1.7 Presented by: Francisco Hernandez-Guzman, PhD February 2007 Agenda Part I: Protein Refinement in Discovery Studio 1.7 (20mins) The problem Novel Algorithms at
Protein Refinement in Discovery Studio 1.7 Presented by: Francisco Hernandez-Guzman, PhD February 2007 Agenda Part I: Protein Refinement in Discovery Studio 1.7 (20mins) The problem Novel Algorithms at
EGFRIndb: Epidermal Growth Factor Receptor Inhibitor database
 EGFRIndb: Epidermal Growth Factor Receptor Inhibitor database Inderjit S Yadav 1,3, Harinder Singh 2, Mohd. Imran Khan 1, Ashok Chaudhury 3, G.P.S. Raghava 2 and Subhash M. Agarwal 1 * 1 Bioinformatics
EGFRIndb: Epidermal Growth Factor Receptor Inhibitor database Inderjit S Yadav 1,3, Harinder Singh 2, Mohd. Imran Khan 1, Ashok Chaudhury 3, G.P.S. Raghava 2 and Subhash M. Agarwal 1 * 1 Bioinformatics
Research Article. Insilico predictions of inhibitors of novel statin structural analogues with HMG-CoA reductase
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(3):942-950 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico predictions of inhibitors of novel statin
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(3):942-950 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico predictions of inhibitors of novel statin
In Silico docking studies of lipoxygenase inhibitory activity of commercially available flavonoids
 Available online at www.scholarsresearchlibrary.com J. Comput. Method. Mol. Design, 2011, 1 (4):65-72 (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA In Silico docking
Available online at www.scholarsresearchlibrary.com J. Comput. Method. Mol. Design, 2011, 1 (4):65-72 (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA In Silico docking
Time Minimizing in anticancer drug discovery. Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia
 Time Minimizing in anticancer drug discovery Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia Abstract Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia
Time Minimizing in anticancer drug discovery Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia Abstract Farida Suhud University of Surabaya, Raya Kalirungkut, Surabaya, Indonesia
OBJECTIVE. Lipids are largely hydrocarbon derivatives and thus represent
 Paper 4. Biomolecules and their interactions Module 20: Saturated and unsaturated fatty acids, Nomenclature of fatty acids and Essential and non-essential fatty acids OBJECTIVE The main aim of this module
Paper 4. Biomolecules and their interactions Module 20: Saturated and unsaturated fatty acids, Nomenclature of fatty acids and Essential and non-essential fatty acids OBJECTIVE The main aim of this module
The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation
 www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
IN-SILICO APPROACH FOR THE ASSESSMENT OF ORAL CANCER PROPERTY ON LIMONIA ACIDISSIMA
 IJPSR (2016), Vol. 7, Issue 3 (Research Article) Received on 16 September, 2015; received in revised form, 29 October, 2015; accepted, 12 December, 2015; published 01 March, 2016 IN-SILICO APPROACH FOR
IJPSR (2016), Vol. 7, Issue 3 (Research Article) Received on 16 September, 2015; received in revised form, 29 October, 2015; accepted, 12 December, 2015; published 01 March, 2016 IN-SILICO APPROACH FOR
CHAPTER 1 INTRODUCTION
 CHAPTER 1 INTRODUCTION Understanding the mechanism of sweetness has been a challenging problem. It is necessary to understand this in order to design safer sweet molecules, for in many diseases (like diabetes,
CHAPTER 1 INTRODUCTION Understanding the mechanism of sweetness has been a challenging problem. It is necessary to understand this in order to design safer sweet molecules, for in many diseases (like diabetes,
Available Online through
 ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
Bioinformation Volume 5
 Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
IDENTIFICATION OF POTENTIAL INHIBITORS FOR LOWERING CHOLESTEROL LEVEL BY INHIBITING PROPROTEIN CONVERTASE SUBTILISIN KEXIN 9
 Vol 9, Issue 6, 2016 Online - 2455-3891 Print - 0974-2441 Research Article IDENTIFICATION OF POTENTIAL INHIBITORS FOR LOWERING CHOLESTEROL LEVEL BY INHIBITING PROPROTEIN CONVERTASE SUBTILISIN KEXIN 9 ABSTRACT
Vol 9, Issue 6, 2016 Online - 2455-3891 Print - 0974-2441 Research Article IDENTIFICATION OF POTENTIAL INHIBITORS FOR LOWERING CHOLESTEROL LEVEL BY INHIBITING PROPROTEIN CONVERTASE SUBTILISIN KEXIN 9 ABSTRACT
Qualitative and Quantitative analysis of 3D predicted arachidonate 15-lipoxygenase-B (15-LOX- 2) from Homo sapiens
 www.bioinformation.net Hypothesis Volume 8(12) Qualitative and Quantitative analysis of 3D predicted arachidonate 15-lipoxygenase-B (15-LOX- 2) from Homo sapiens Neha Arora 1, Vinay Kumar Singh 2, Kavita
www.bioinformation.net Hypothesis Volume 8(12) Qualitative and Quantitative analysis of 3D predicted arachidonate 15-lipoxygenase-B (15-LOX- 2) from Homo sapiens Neha Arora 1, Vinay Kumar Singh 2, Kavita
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Molecular modelling and insilico drug docking studies on breast cancer target protein (TNRC9) using cheminformatics software and tools
 Available online at www.ijntps.org ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES Molecular modelling and insilico drug docking studies on breast cancer target protein
Available online at www.ijntps.org ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES Molecular modelling and insilico drug docking studies on breast cancer target protein
Exploring in silico affinity of flavonoids and tannins to human fibroblast growth factorinducible14 (Fn14), a member of TNF receptor super family
 www.bioinformation.net Hypothesis Volume 9(12) Exploring in silico affinity of flavonoids and tannins to human fibroblast growth factorinducible14 (Fn14), a member of TNF receptor super family CVS Siva
www.bioinformation.net Hypothesis Volume 9(12) Exploring in silico affinity of flavonoids and tannins to human fibroblast growth factorinducible14 (Fn14), a member of TNF receptor super family CVS Siva
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
CHAPTER 21: Amino Acids, Proteins, & Enzymes. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Binding property of HIV p24 and Reverse transcriptase by chalcones from Pongamia pinnata seeds
 www.bioinformation.net Volume 14(6) Hypothesis Binding property of HIV p24 and Reverse transcriptase by chalcones from Pongamia pinnata seeds Manikannan Mathaiyan 1, Arumugam Suresh 1*, Rangasamy Balamurugan
www.bioinformation.net Volume 14(6) Hypothesis Binding property of HIV p24 and Reverse transcriptase by chalcones from Pongamia pinnata seeds Manikannan Mathaiyan 1, Arumugam Suresh 1*, Rangasamy Balamurugan
More powerful virus inhibitors from structure-based analysis of
 More powerful virus inhibitors from structure-based analysis of HEV71 capsid-binding molecules Luigi De Colibus, Xiangxi Wang, John A. B. Spyrou, James Kelly, Jingshan Ren, Jonathan Grimes, Gerhard Puerstinger,
More powerful virus inhibitors from structure-based analysis of HEV71 capsid-binding molecules Luigi De Colibus, Xiangxi Wang, John A. B. Spyrou, James Kelly, Jingshan Ren, Jonathan Grimes, Gerhard Puerstinger,
Docking Analysis of Darunavir as HIV Protease Inhibitors
 Journal of Computational Methods in Molecular Design, 2012, 2 (1):39-43 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Docking Analysis
Journal of Computational Methods in Molecular Design, 2012, 2 (1):39-43 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Docking Analysis
GPCR Structure-Based Virtual Screening Approach for CB2 Antagonist Search
 Article GPCR Structure-Based Virtual Screening Approach for CB2 Antagonist Search Jian-Zhong Chen, Junmei Wang, and Xiang-Qun Xie J. Chem. Inf. Model., 2007, 47 (4), 1626-1637 DOI: 10.1021/ci7000814 Publication
Article GPCR Structure-Based Virtual Screening Approach for CB2 Antagonist Search Jian-Zhong Chen, Junmei Wang, and Xiang-Qun Xie J. Chem. Inf. Model., 2007, 47 (4), 1626-1637 DOI: 10.1021/ci7000814 Publication
News. Volume 8: November 2, 2009
 FightAIDS@Home News Volume 8: November 2, 2009 The FightAIDS@Home Project uses the volunteered computing power of the World Community Grid to test candidate compounds against the variations (or mutants
FightAIDS@Home News Volume 8: November 2, 2009 The FightAIDS@Home Project uses the volunteered computing power of the World Community Grid to test candidate compounds against the variations (or mutants
Creating a high-throughput screening database to propose ligands capable of modulating the HIV-1 protease receptor
 Vol. 5(7), pp. 224-234, July, 2013 DOI 10.5897/JAHR12.056 ISSN 2141-2359 2013 Academic Journals http://www.academicjournals.org/jahr Journal of AIDS and HIV Research Full Length Research Paper Creating
Vol. 5(7), pp. 224-234, July, 2013 DOI 10.5897/JAHR12.056 ISSN 2141-2359 2013 Academic Journals http://www.academicjournals.org/jahr Journal of AIDS and HIV Research Full Length Research Paper Creating
Prediction of temperature factors from protein sequence
 www.bioinformation.net Hypothesis Volume 9(3) Prediction of temperature factors from protein sequence Shrihari Sonavane 1 *, Ashok A Jaybhaye 2 & Ajaykumar G Jadhav 1 1Department of Microbiology, Institute
www.bioinformation.net Hypothesis Volume 9(3) Prediction of temperature factors from protein sequence Shrihari Sonavane 1 *, Ashok A Jaybhaye 2 & Ajaykumar G Jadhav 1 1Department of Microbiology, Institute
Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose
 www.bioinformation.net Hypothesis Volume 9(16) Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose Nurul Bahiyah Ahmad Khairudin* & Nur Shima Fadhilah Mazlan Malaysia
www.bioinformation.net Hypothesis Volume 9(16) Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose Nurul Bahiyah Ahmad Khairudin* & Nur Shima Fadhilah Mazlan Malaysia
Trilateral Project WM4
 ANNEX 2: Comments of the JPO Trilateral Project WM4 Comparative studies in new technologies Theme: Comparative study on protein 3-dimensional (3-D) structure related claims 1. Introduction As more 3-D
ANNEX 2: Comments of the JPO Trilateral Project WM4 Comparative studies in new technologies Theme: Comparative study on protein 3-dimensional (3-D) structure related claims 1. Introduction As more 3-D
In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents
 Research Article In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents C. N. Hemalatha 1, Elancheziyan K 1, D. Pavithra 2, M. Vijey Aanandhi 3 * ABSTRACT
Research Article In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents C. N. Hemalatha 1, Elancheziyan K 1, D. Pavithra 2, M. Vijey Aanandhi 3 * ABSTRACT
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Molecular modeling of Ruellia tuberosa compounds as -amylase inhibitor: an in silico comparation between human and rat enzyme model
 www.bioinformation.net Volume 10(4) Hypothesis Molecular modeling of Ruellia tuberosa L compounds as amylase inhibitor: an in silico comparation between human and rat enzyme model Dyah Ratna Wulan 1, 2,
www.bioinformation.net Volume 10(4) Hypothesis Molecular modeling of Ruellia tuberosa L compounds as amylase inhibitor: an in silico comparation between human and rat enzyme model Dyah Ratna Wulan 1, 2,
Pelagia Research Library. Docking studies on PPARγ of novel α-phenoxy phenyl propionic acid derivatives as antidiabetic agents
 Available online at www.pelagiaresearchlibrary.com Der Pharmacia Sinica, 2011, 2 (2): 327-332 ISSN: 0976-8688 CDEN (USA): PSHIBD Docking studies on PPARγ of novel α-phenoxy phenyl propionic acid derivatives
Available online at www.pelagiaresearchlibrary.com Der Pharmacia Sinica, 2011, 2 (2): 327-332 ISSN: 0976-8688 CDEN (USA): PSHIBD Docking studies on PPARγ of novel α-phenoxy phenyl propionic acid derivatives
Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase in the treatment of dermatophytoses by molecular docking
 International Journal of Advanced Research in Medical Sciences 2017 Volume 1 Issue 1 Pages: 1-5 Original Research Paper Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase
International Journal of Advanced Research in Medical Sciences 2017 Volume 1 Issue 1 Pages: 1-5 Original Research Paper Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase
Investigating the Inhibition of 5-LO Enzyme by Main Cannabinoids Contained in Cannabis sativa and Seized Resin Cannabis by Using Molecular Modeling
 Research Article imedpub Journals www.imedpub.com DOI: 10.21767/2573-4466.100010 Investigating the Inhibition of 5-LO Enzyme by Main Cannabinoids Contained in Cannabis sativa and Seized Resin Cannabis
Research Article imedpub Journals www.imedpub.com DOI: 10.21767/2573-4466.100010 Investigating the Inhibition of 5-LO Enzyme by Main Cannabinoids Contained in Cannabis sativa and Seized Resin Cannabis
Bio 100 Serine Proteases 9/26/11
 Assigned Reading: 4th ed. 6.4.1 The Chymotrypsin Mechanism Involves Acylation And Deacylation Of A Ser Residue p. 213 BOX 20-1 Penicillin and β-lactamase p. 779 6.5.7 Some Enzymes Are Regulated By Proteolytic
Assigned Reading: 4th ed. 6.4.1 The Chymotrypsin Mechanism Involves Acylation And Deacylation Of A Ser Residue p. 213 BOX 20-1 Penicillin and β-lactamase p. 779 6.5.7 Some Enzymes Are Regulated By Proteolytic
LIPOPREDICT: Bacterial lipoprotein prediction server
 www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Combining molecular modelling with experiments: Sulfonylureas and glinides as new PPARγ agonists
 Combining molecular modelling with experiments: Sulfonylureas and glinides as new PPARγ agonists Marco Scarsi Biozentrum - Swiss Institute of Bioinformatics Drug discovery and development 12-15 years of
Combining molecular modelling with experiments: Sulfonylureas and glinides as new PPARγ agonists Marco Scarsi Biozentrum - Swiss Institute of Bioinformatics Drug discovery and development 12-15 years of
LIGAND BASED PHARMACOPHORE DEVELOPMENT FOR COLORECTAL CANCER DRUGS
 The Professional Medical Journal www.theprofesional.com ORIGINAL PROF-2566 LIGAND BASED PHARMACOPHORE DEVELOPMENT FOR COLORECTAL CANCER DRUGS 1. PhD Scholar (Bioinformatics) M.A.J.U. Islamabad. 2. Postgraduate
The Professional Medical Journal www.theprofesional.com ORIGINAL PROF-2566 LIGAND BASED PHARMACOPHORE DEVELOPMENT FOR COLORECTAL CANCER DRUGS 1. PhD Scholar (Bioinformatics) M.A.J.U. Islamabad. 2. Postgraduate
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
Structural characterization of a flavonoid glycosyltransferase from Withania somnifera
 www.bioinformation.net Hypothesis Volume 8(19) Structural characterization of a flavonoid glycosyltransferase from Withania somnifera Santosh Kumar Ramachandra Jadhav 1, Krunal Arvind Patel 1, Bhushan
www.bioinformation.net Hypothesis Volume 8(19) Structural characterization of a flavonoid glycosyltransferase from Withania somnifera Santosh Kumar Ramachandra Jadhav 1, Krunal Arvind Patel 1, Bhushan
The Binding Mode of by Electron Crystallography
 The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
Chapter 3: Amino Acids and Peptides
 Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation
 6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
Insight into the mechanism of lipids binding and uptake by CD36 receptor
 www.bioinformation.net Hypothesis Volume 11(6) Insight into the mechanism of lipids binding and uptake by CD36 receptor Zineb Tarhda*, Azeddine Ibrahimi Biotechnology lab (MedBiotech), Faculté de Médecine
www.bioinformation.net Hypothesis Volume 11(6) Insight into the mechanism of lipids binding and uptake by CD36 receptor Zineb Tarhda*, Azeddine Ibrahimi Biotechnology lab (MedBiotech), Faculté de Médecine
Austroinulin from Stevia rebaudiana a Potential Agonist of the Peroxisome Proliferator-activated Receptor (PPARγ): an in-silico Study.
 Research Article http://www.alliedacademies.org/systems-biology-proteome-research/ Austroinulin from Stevia rebaudiana a Potential Agonist of the Peroxisome Proliferator-activated Receptor (PPARγ): an
Research Article http://www.alliedacademies.org/systems-biology-proteome-research/ Austroinulin from Stevia rebaudiana a Potential Agonist of the Peroxisome Proliferator-activated Receptor (PPARγ): an
Drug designing and validation studies on human asthma disease gene NPSR1 using Cheminformatics tools
 Available online at wwwijntpsorg ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES Drug designing and validation studies on human asthma disease gene NPSR1 using Cheminformatics
Available online at wwwijntpsorg ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES Drug designing and validation studies on human asthma disease gene NPSR1 using Cheminformatics
VIRTUAL SCREENS WITH MULTIPLE RECEPTOR CONFORMATIONS. Rob Swift, UCI Wednesday, August 3, 2011
 VIRTUAL SCREENS WITH MULTIPLE RECEPTR CNFRMATINS Rob Swift, UCI Wednesday, August 3, 2011 utline verview of multi receptor virtual screening in CADD Case studies 1. Trypanosoma brucei RNA editing ligase
VIRTUAL SCREENS WITH MULTIPLE RECEPTR CNFRMATINS Rob Swift, UCI Wednesday, August 3, 2011 utline verview of multi receptor virtual screening in CADD Case studies 1. Trypanosoma brucei RNA editing ligase
DESIGN DOCKING STUDIES ON ANTIAPOPTOTIC PROTEIN INHIBITORS AS A NOVEL TARGET ON BREAST CANCER
 DESIGN DOCKING STUDIES ON ANTIAPOPTOTIC PROTEIN INHIBITORS AS A NOVEL TARGET ON BREAST CANCER Wagh Jyoti. Gorakha a,b,c *, Abhilasha Mithal a a Jayoti Vidyapeeth Women s University, Jaipur - Ajmer Express
DESIGN DOCKING STUDIES ON ANTIAPOPTOTIC PROTEIN INHIBITORS AS A NOVEL TARGET ON BREAST CANCER Wagh Jyoti. Gorakha a,b,c *, Abhilasha Mithal a a Jayoti Vidyapeeth Women s University, Jaipur - Ajmer Express
Chemical Nature of the Amino Acids. Table of a-amino Acids Found in Proteins
 Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Targeting BACE 1(Beta secretase) through Polyphenolic compounds -A computational insilico approach with emphasis on binding site analysis
 Journal of Computational Methods in Molecular Design, 2014, 4 (1):14-24 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Targeting BACE 1(Beta
Journal of Computational Methods in Molecular Design, 2014, 4 (1):14-24 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Targeting BACE 1(Beta
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
In-Silico Anticancer activity of cow urine extract of Kappaphycus alvarezii
 Available online at www.ijntps.org ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES In-Silico Anticancer activity of cow urine extract of Kappaphycus alvarezii RESEARCH
Available online at www.ijntps.org ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES In-Silico Anticancer activity of cow urine extract of Kappaphycus alvarezii RESEARCH
Reactions and amino acids structure & properties
 Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Break Desired Compound into 3 Smaller Ones
 Break Desired Compound into 3 Smaller nes A B C Indole ydrazine Phenol When creating a covalent link between model compounds move the charge on the deleted into the carbon to maintain integer charge (i.e.
Break Desired Compound into 3 Smaller nes A B C Indole ydrazine Phenol When creating a covalent link between model compounds move the charge on the deleted into the carbon to maintain integer charge (i.e.
Visualizing Biopolymers and Their Building Blocks
 Visualizing Biopolymers and Their Building Blocks By Sharlene Denos (UIUC) & Kathryn Hafner (Danville High) Living things are primarily composed of carbon-based (organic) polymers. These are made up many
Visualizing Biopolymers and Their Building Blocks By Sharlene Denos (UIUC) & Kathryn Hafner (Danville High) Living things are primarily composed of carbon-based (organic) polymers. These are made up many
Predicting Inactive Conformations of Protein Kinases Using Active Structures: Conformational Selection of Type-II Inhibitors
 Predicting Inactive Conformations of Protein Kinases Using Active Structures: Conformational Selection of Type-II Inhibitors Min Xu, Lu Yu, Bo Wan, Long Yu, Qiang Huang* State Key Laboratory of Genetic
Predicting Inactive Conformations of Protein Kinases Using Active Structures: Conformational Selection of Type-II Inhibitors Min Xu, Lu Yu, Bo Wan, Long Yu, Qiang Huang* State Key Laboratory of Genetic
Chemical Surface Transformation 1
 Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
Molecular Docking Studies on Pyrazolopyrimidine and their Derivatives as Human Phosphoinositide 3-Kinase Inhibitors
 Cloud Publications International Journal of Advanced Bioinformatics and Computational Biology 2012, Volume 1, Issue 1, pp. 1-11, Article ID Med-30 Research Article Open Access Molecular Docking Studies
Cloud Publications International Journal of Advanced Bioinformatics and Computational Biology 2012, Volume 1, Issue 1, pp. 1-11, Article ID Med-30 Research Article Open Access Molecular Docking Studies
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Molecular Dynamics Simulation Study Explaining Inhibitor Selectivity in Different Class of Histone Deacetylases
 Journal of Biomolecular Structure & Dynamics, ISSN 0739-1102 Volume 29, Issue Number 4, (2012) Adenine Press (2012) Abstract Molecular Dynamics Simulation Study Explaining Inhibitor Selectivity in Different
Journal of Biomolecular Structure & Dynamics, ISSN 0739-1102 Volume 29, Issue Number 4, (2012) Adenine Press (2012) Abstract Molecular Dynamics Simulation Study Explaining Inhibitor Selectivity in Different
Workshop on Analysis and prediction of contacts in proteins
 Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Homology modelling of CB1 receptor and selection of potential inhibitor against Obesity
 www.bioinformation.net Hypothesis Volume 8(11) Homology modelling of CB1 receptor and selection of potential inhibitor against Obesity Mahesh Shrinivasan 1 *, Sinosh Skariyachan 2, Vaka Aparna 1 & Vinod
www.bioinformation.net Hypothesis Volume 8(11) Homology modelling of CB1 receptor and selection of potential inhibitor against Obesity Mahesh Shrinivasan 1 *, Sinosh Skariyachan 2, Vaka Aparna 1 & Vinod
CH 3. Lipids CHAPTER SUMMARY
 H 3 C H 3 C 15 H 3 C H Views of Cholesterol APTER SUMMARY 15.1 The Nature of can best be defined as biomolecules which are soluble to a great extent in solvents. In contrast to carbohydrates, proteins
H 3 C H 3 C 15 H 3 C H Views of Cholesterol APTER SUMMARY 15.1 The Nature of can best be defined as biomolecules which are soluble to a great extent in solvents. In contrast to carbohydrates, proteins
Journal of Chemical and Pharmaceutical Research, 2016, 8(9): Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2016, 8(9):73-80 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 QSAR and Docking Studies of Pubchem Derivatives as
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2016, 8(9):73-80 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 QSAR and Docking Studies of Pubchem Derivatives as
A. There are about 100 elements; 25 of them are necessary for life. B. Carbon atoms can form long chains, leading to a huge number of possible
 Ch. 2 How Cells Function 2.1 Chemical reactions take place inside cells. 1. All cells are made of the same elements. A. There are about 100 elements; 25 of them are necessary for life. B. The smallest
Ch. 2 How Cells Function 2.1 Chemical reactions take place inside cells. 1. All cells are made of the same elements. A. There are about 100 elements; 25 of them are necessary for life. B. The smallest
You must answer FOUR of the SIX questions. Use a SEPARATE answer book for EACH question.
 UNIVERSITY OF EAST ANGLIA School of Pharmacy Main Series UG Examination 2016-17 DRUG DESIGN AND MECHANISMS OF DRUG ACTION PHA-5001Y Time allowed: 2 hours You must answer FOUR of the SIX questions. Use
UNIVERSITY OF EAST ANGLIA School of Pharmacy Main Series UG Examination 2016-17 DRUG DESIGN AND MECHANISMS OF DRUG ACTION PHA-5001Y Time allowed: 2 hours You must answer FOUR of the SIX questions. Use
Targeting PPIs in Oncology using a Fragment-Based Drug Discovery Approach Justin F. Bower CRUK Beatson Institute
 Targeting PPIs in Oncology using a Fragment-Based Drug Discovery Approach Justin F. Bower CRUK Beatson Institute Cancer Research UK Beatson Institute Establish an outstanding basic research programme into
Targeting PPIs in Oncology using a Fragment-Based Drug Discovery Approach Justin F. Bower CRUK Beatson Institute Cancer Research UK Beatson Institute Establish an outstanding basic research programme into
International Journal of Research in Pharmacy and Science
 Research article Available online www.ijrpsonline.com I: 2249 3522 International Journal of Research in Pharmacy and cience Pharmacophore Mapping and Docking on Triazepane Derivatives as DPP-IV Inhibitors
Research article Available online www.ijrpsonline.com I: 2249 3522 International Journal of Research in Pharmacy and cience Pharmacophore Mapping and Docking on Triazepane Derivatives as DPP-IV Inhibitors
Module Contact: Dr Paul McDermott, PHA Copyright of the University of East Anglia Version 1
 UIVERSITY F EAST AGLIA School of Pharmacy Main Series UG Examination 2012-13 LIFE SCIECES CHEMISTRY PHA-1ECY Time allowed: 2 hours You must answer FUR (4) of the SIX (6) questions. Use a SEPARATE answer
UIVERSITY F EAST AGLIA School of Pharmacy Main Series UG Examination 2012-13 LIFE SCIECES CHEMISTRY PHA-1ECY Time allowed: 2 hours You must answer FUR (4) of the SIX (6) questions. Use a SEPARATE answer
