Papers. Clinical application of the Quantiplex HCV RNA 2.0 and Amplicor HCV Monitor assays for quantifying serum hepatitis C virus RNA
|
|
|
- Lesley Clark
- 6 years ago
- Views:
Transcription
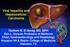
1 J Clin Pathol 1999;52: Papers Hepatobiliary Division, Department of Internal Medicine, Kaohsiung Medical College Hospital, No 100, Shih-Chuan 1st Rd, Kaohsiung, Taiwan, Republic of China M-L Yu W-L Chuang S-C Chen Z-Y Lin M-Y Hsieh L-Y Wang W-Y Chang Correspondence to: Dr Chang Accepted for publication 21 July 1999 Clinical application of the Quantiplex HCV RNA 2.0 and HCV Monitor assays for quantifying serum hepatitis C virus RNA Ming-Lung Yu, Wan-Long Chuang, Shinn-Cherng Chen, Zu-Yau Lin, Ming-Yuh Hsieh, Liang-Yen Wang, Wen-Yu Chang Abstract Aim To compare the performance characteristics and clinical application of two diverent technologies for quantifying serum hepatitis C virus (HCV) RNA levels. Methods HCV RNA was quantified by HCV Monitor assay () and Quantiplex HCV RNA 2.0 assay () in 119 sera from 107 HCV infected patients. Results Both assays had similar sensitivity (79.4% for ; 86.0% for bdna- 2), acceptable coeycients of variation (5.3% in ; 2.6% in ), and good linearity (r 2 > 0.98). There was a positive correlation between quantification values of both methods (r = 0.683, p < 0.001). The values were on an average 1.76 log lower than results. Male subjects and HCV genotype 1b were significantly associated with higher viral load determined by, but not with viral load measured by. In 70 chronic HCV infected patients treated with interferon alfa, mean (SD) pretreatment viral load in 27 complete responders (3.47 (0.84) logs for, 5.63 (0.58) for ) was significantly lower than in non-responders (4.43 (1.01) logs for, 6.10 (0.67) logs for ; p < 0.001). Cut ov points of 3.9 logs for and 5.8 logs for were determined to be the best for predicting response to interferon alfa, giving acceptable sensitivity (70.4%, 74.1%), specificity (72.1%, 65.1%), and accuracy (71.4%, 68.6%), respectively. Conclusions Both the and assays are clinically useful methods for HCV RNA quantification and are reliable for predicting the outcome of treatment, despite diverences in absolute quantification values and in the correlation between HCV genotypes and viral load. (J Clin Pathol 1999;52: ) Keywords: hepatitis C virus; virus quantification assay; interferon Hepatitis C virus (HCV), a highly heterogeneous, single stranded RNA virus, is the major aetiological agent in parenterally transmitted non-a, non-b hepatitis and often causes persistent infection leading to chronic liver disease and primary hepatocellular carcinoma. 1 3 Serum HCV RNA (viral load) may correlate with the clinical stage of disease 4 and HCV genotype. 5 A low HCV viral load has also been observed to be one of the important predictive markers of successful treatment with interferon alfa. 6 8 Assays for serum HCV RNA quantification should therefore be accurate and reproducible to permit careful clinical assessment of diverent HCV patient populations. Competitive reverse transcription polymerase chain reaction (RT-PCR) is useful for evaluating the state of viral replication. 4 5 However, the method is costly, time consuming, and technically challenging, as reflected by large variations in sensitivity among diverent laboratories. Two commercial assays for HCV RNA quantification have become available and are widely used: the branched DNA assay (Chiron Corporation, Emeryville, California, USA) and the HCV Monitor assay (Roche, Nutley, New Jersey, USA) There are relatively few studies comparing the current version of bdna assay 11 (Quantiplex HCV RNA 2.0; ) and the original version of the assay, particularly in assessing Log HCV RNA concentration (copies or eq/ml) r 2 = 0.98 r 2 = Log (threefold dilutions of serum) Figure 1 Analysis of HCV RNA in serial dilutions of high titre serum by two diverent quantitative assays, the Quantiplex branched-dna 2.0 () assay and the Roche HCV Monitor assay.
2 808 Yu, Chuang, Chen, et al Table 1 Correlation between HCV genotypes and quantification values by the and assays Genotype No HCV RNA concentration* Spearman s rank correlation coeycient p Value All types (1.09) 5.93 (0.69) < b (0.99) 5.99 (0.73) < a (0.87) 6.06 (0.63) < b (1.57) 5.88 (1.05) <0.05 3a Mixed (1.06) 5.78 (0.57) Unclassified (1.07) 5.57 (0.53) * Expressed as mean (SD) of logarithmic transformation (log copy or equivalent/ml). Table 2 Background of HCV infected patients and their serum HCV RNA levels measured by the and assays Characteristics No * * All patients (1.10) 5.93 (0.65) Age (years) Below (1.09) 5.88 (0.65) 40 to (1.07) 5.97 (0.64) Above (1.18) 5.92 (0 72) Sex Male (1.08) 6.03 (0.61) Female (1.07) 5.82 (0.68) Mode of transmission Transfusion (1.13) 5.88 (0.60) Percutaneous (1.07) 5.93 (0 62) procedure Unknown (1.09) 6.01 (0 67) Alanine aminotransferase Normal (< 25 IU/l) (0.84) 5.46 (0.55) Abnormal (> 25 IU/l) (1.10) 5.96 (0.65) Histology Acute hepatitis (1.24) 5.80 (0.81) Chronic persistent (0.89) 6.11 (0.76) hepatitis Chronic active (1.08) 5.94 (0.59) hepatitis Liver cirrhosis (1.16) 5.91 (0.58) Hepatocellular (1.34) 5.74 (0 86) carcinoma HCV subtype 1b (0.99) 5.99 (0.73) Non-1b (0.98) 5.90 (0.60) 2a (0.86) 6.05 (0.58) 2b (1.58) 5.92 (0.91) 3a Unclassified (1.17) 5.54 (0.50) Mixed infection (1.06) 5.78 (0.57) * Mean (SD) deviation of logarithmic transformation of log copies or equivalents/ml. Probability of diverence in sex and viral load by the assay (p < 0.05). Probability of diverences in HCV genotypes and viral load by the assay (p < 0.001). their ability to predict response to interferon alfa treatment in patients with chronic hepatitis C. Objectives of the current study were: (1) to compare the clinical sensitivity, reproducibility, linearity, and correlation of the and assays in patients of diverent HCV genotypes and stages of infection; and (2) to Table 3 Stepwise multiple linear regression analysis of factors associated with serum HCV RNA levels Assay variables CoeYcients Standard error of coeycient Adjusted multiple r 2 p Value assay / Intercept (log copies/ml) 4.38 / Sex (male = 1, female = 2 ) <0.05 / HCV genotype (genotype 1 = 1, <0.01 non-genotype 1b = 0) BDNA-2 assay /Intercept (log eq/ml) 5.93 Factors such as age, sex, mode of transmission, aminotransferase, histology, and HCV genotype were analysed by stepwise multiple linear regression. Males and patients infected with genotype 1b virus had significantly higher HCV RNA levels, as determined with the assay. No factors were significantly associated with viral load measured by the assay. determine the clinical value of these two commercial HCV viral load assays for predicting response to interferon alfa treatment. Methods PATIENTS One hundred and seven Taiwanese patients were enrolled in the study. Seventy patients with chronic hepatitis C were treated with 6 MU of interferon alfa given intramuscularly three times a week for 24 weeks. Complete responders were defined as patients showing normal alanine aminotransferase (ALT) levels and clearance of HCV RNA by nested RT-PCR at the end of the treatment and for six months after the cessation of treatment. All other patients were classified as nonresponders. All serum samples were positive for HCV RNA, including 107 samples collected at the time of liver biopsy or aspiration cytology for measurement of baseline viral load, and an additional 12 samples from 12 patients at the third month of interferon alfa treatment. DETECTION/QUANTIFICATION OF SERUM HCV RNA AND GENOTYPING HCV RNA was detected qualitatively by nested RT-PCR using 5' non-coding region (5'NCR) specific primers. 12 The detection limit of the nested RT-PCR is 50 copies/ml. HCV genotyping was determined by method described by Okamoto et al. 13 Levels of serum HCV RNA were measured with the and assays according to the manufacturer s instructions, which were followed exactly. HCV RNA was detected directly in the assay by a series of probe hybridisations to boost the signal coming from each HCV RNA. The relative intensity of the signal was compared against an external standard curve, giving a quantification of million equivalents (Meq)/ml. The assay is based on reverse transcription and amplification of HCV RNA with primers targeting an xbp region within the 5' NCR of the HCV genome. An internal quantitation standard (an RNA molecule derived from genotype 1 sequence of the HCV) was coamplified with the HCV RNA target. Both the amplified products of the internal standard and sample RNA were serially diluted and detected by hybridization in a colorimetric assay. Results were calculated manually according to the manufacturer s instructions, giving a quantification range of copies/ ml. STATISTICAL ANALYSIS Data are expressed as mean (SD) after logarithmic transformation of original values. The χ 2 test, Student s t test, analysis of variance, Spearman s rank correlation coefficient, simple linear regression, and stepwise multiple linear regression were used. We compared the area under the curve with the receiver operating characteristics (ROC) analysis and made an attempt to derive a suitable clinical cut ov for each assay that would best predict the response to interferon alfa treatment. For the purpose of analysing the
3 Quantification of hepatitis C virus RNA levels 809 Log serum HCV RNA concentration p < p < Detection limit 2 CR NR CR NR Figure 2 Pretreatment levels of serum hepatitis C virus RNA in responders (CR) and non-responders (NR) to interferon alfa treatment at 6 MU thrice weekly for 24 weeks assessed by the Roche HCV Monitor assay () and Quantiplex branched-dna 2.0 assay (). Of 70 patients, 27 were responders. The bars represent mean pretreatment level of serum hepatitis C virus RNA in each group. data by suitable statistical methods, we assigned a nominal value of 0.1 Meq/ml to samples that were negative by but positive for HCV RNA by nested RT-PCR, and a nominal value of 500 copies/ml to samples that were negative but positive by nested RT-PCR. Results PERFORMANCE CHARACTERISTICS OF THE Detection limit AMPLICOR AND ASSAYS Of 107 samples for baseline viral load, HCV RNA was quantifiable in 85 patients (79.4%) by and in 92 (86.0%) by. Eighty (74.8%) samples were quantifiable by both assays, while 10 (9.4%) were under the detection limit of both assays. For 12 PCR positive samples collected at the third month of interferon alfa treatment, the percentage of quantifiable samples by and was similar (41.7% and 58.3%, respectively). To investigate assay reproducibility, each assay was tested on two occasions using diverent lots of reagents with separate aliquots of a Table 4 Correlation between pretreatment levels of serum HCV RNA and response to interferon alfa treatment Pretreatment serum HCV RNA level CR (n=27) NR (n=43) p Value (log copies/ml) Low (< 3) 13 (68.4%) 6 (31.6%) <0.05* Median (3 4.5) 9 (39.1%) 14 (60.9%) High (> 4.5) 5 (17.9%) 23 (82.1%) (log eq/ml) Low (< 5.3) 8 (66.7%) 4 (33.3%) <0.05* Median ( ) 15 (38.5%) 24 (61.5%) High (> 6.5) 4 (21.1%) 15 (78.9%) CR, complete responder; eq, equivalent; NR, non-responder. *χ 2 test with linear correlation. Table 5 Prediction of complete response to interferon alfa treatment based on the calculated clinical cut ov values for serum HCV RNA levels Assays Cut ov values Sensitivity (n/n) Specificity (n/n) Accuracy (n/n) (copies/ml) (3.9 logs) 70.4% (19/27) 72.1% (31/43) 71.4% (50/70) (eq/ml) (5.8 logs) 74.1% (20/27) 65.1% (28/43) 68.6% (48/70) Seventy patients with chronic hepatitis C treated with interferon alfa, 6 MU thrice weekly for 24 weeks, were studied. The complete response rate was 38.6 % (27/70). panel of 16 samples. The mean per cent coeycient of variation based upon repeat testing was 5.3% (range 0.8% to 10.4%) for and 2.6% (range 1.3% to 7.1%) for. Linearity of quantification was examined with a dilution panel made by six threefold serial dilutions of a high titre serum sample into normal human sera. As shown in fig 1, both and had good linearity within the linear range (r 2 = 0.99 and 0.98, respectively). HCV RNA quantification values measured by the and assays were positively correlated (Spearman s rank correlation coeycient, r = 0.683, p < 0.001). The values were on average 1.76 log lower than the results. Non-parametric tests of correlation gave a high correlation coeycient upon pairwise comparison of samples with diverent genotypes (table 1). However, the mean diverence in quantification values between these two assays was 1.1 log for genotype 1 and 2.1 logs for non-genotype 1 (p < 0.05). HCV VIRAL LOAD AND CLINICAL MANIFESTATIONS Patient age, sex, mode of transmission, liver biochemistry, liver histology, and viral genotype were analysed to evaluate the relation between HCV viral load and clinical manifestation of HCV infection. By univariate analysis, viral load determined with did not correlate with patient s age, mode of transmission, and severity of liver disease (table 2). HCV viral load was significantly diverent between male and female patients (p < 0.05), and among diverent HCV genotypes (p < 0.001). In further analysis using stepwise multiple linear regression, sex and HCV genotype remained as significant factors avecting serum HCV RNA levels measured by (table 3). For the quantification results, none of the factors analysed was found to be correlated with viral load by either univariate or multivariate analyses. PRETREATMENT HCV RNA LEVELS AND RESPONSE TO INTERFERON ALFA TREATMENT In 70 interferon alfa treated patients, pretreatment viral load in 27 complete responders (38.6%) (3.47 (0.84) logs for, 5.63 (0.58) for ) was significantly lower than that in non-responders (4.43 (1.01) logs for, 6.10 (0.67) logs for ; p < 0.001, fig 2). There was no association between sex, age, pretreatment ALT levels, liver histology, HCV genotype, kind of interferon, or history of blood transfusion with the interferon response. After dividing patients into three groups with low, medium, and high viral load, the percentage of complete responders was observed to decrease with higher pretreatment serum HCV RNA concentrations (p < 0.05, χ 2 test with linear correlation) (table 4). In an attempt to accurately predict interferon alfa response by the pretreatment viral load, the most appropriate cut ov in pretreatment HCV RNA was calculated by using the ROC analysis. In this analysis, patients with a pretreatment viral load lower than the cut ov
4 810 Yu, Chuang, Chen, et al value would be considered as probable complete responders, while those with a viral load higher than the cut ov value would be regarded as probable non-responders. Using a cut ov value of 3.9 logs for and 5.8 logs for, both the and assays had acceptable sensitivity (70.5% and 74.1%), specificity (72.1% and 65.1%), and accuracy (71.4% and 68.6%) in predicting complete response to interferon alfa given at 6 MU thrice weekly for 24 weeks (table 5). Discussion Serum HCV RNA is an important predictive factor of response to interferon treatment in patients with chronic hepatitis C. 6 8 Easy, reliable, standardised quantitative HCV RNA tests with good accuracy and reproducibility are therefore needed to popularise the routine use of viral load in the clinical management of HCV patients. We undertook this study to compare the commercially available and assays for clinical sensitivity, reproducibility, linearity of quantitation, and ability to quantify all HCV genotypes with equal sensitivity. Both assays showed good reproducibility, based on repetitive testing of a panel of 16 clinical samples in our study. The between run coeycients of variation were 5.3% and 2.6% in the and assays, respectively. This contrasts with the poor reproducibility in the assay reported by Hawkins et al. 16 Possible explanations for this discrepancy might be the small number of samples tested in the present study and in that by Hawkins, or diverences in technical expertise in performing the test at various laboratories. We observed that the and assays had similar clinical sensitivity, despite the vastly diverent quantification limits assigned by the manufacturers. Although quantification results by these two tests were positively correlated, the values were 1.76 log lower than the values obtained with the assay. This diverence could result from factors such as quantification standards of diverent lengths, sequences, and chemical properties. An important observation in our study was that there were discrepancies in the relation between genotypes and HCV viral load depending on the quantification method used. HCV genotype 1b samples were shown to have significantly higher viral load than other genotypes in the assay. No genotype dependent diverences in viral load was observed for the assay, consistent with previous reports, 16 as well as with larger surveys of blood donors and patients when values obtained with the first generation bdna assay were corrected for lower sensitivities for genotypes 2 and 3. Furthermore, the mean diverence in quantitation values between these two assays was 1.1 log for genotype 1 and 2.1 logs for non-genotype 1, even though the and results correlated well with each other for each of the HCV genotypes. DiVerences in viral load among HCV genotypes may have been caused by the absolute eyciency of HCV RNA detection rather than being a real biological phenomenon. HCV exists as a heterogeneous group of viruses sharing approximately 70% homology, and shows significant variations even in the most conserved regions of the HCV genome. The assay uses the most conserved regions of genotype 1 HCV as target sequences for amplification and probe hybridisation. However, minor sequence variations between the genotype 1 sequences used in PCR primers and target sequences of non-genotype 1 viruses, 20 and genotype specific diverences in the strength of base pairings and in the secondary structure of the 5'NCR, may interfere with primer annealing and thus lead to underestimation of viral load for non-genotype 1 samples. 21 An unexpected finding in our study was that male sex was associated with significantly higher levels of HCV RNA by the results. Similar results had been observed in dialysis patients by using competitive PCR 22 and in blood donors using the corrected bdna-1 results, 18 but not in other studies. 4 9 The lower viral load in female subjects might be caused by sex specific diverences in viral replication that could not be detected by the assay in the present study. On the other hand, the observation might be an opportunistically statistical bias: confounding factors could not be controlled for with multivariate analysis owing to the fact that the assay might have genotype specific diverences in detection eyciency, and that a diverence in genotype distribution among male and female subjects (42.4% male, 31.3% female for genotype 1 patients) contributed to sex specific diverences in viral load. Our study, as well as most other, 6 8 documented that pretreatment HCV RNA level is one of the factors influencing the outcome of interferon alfa treatment, and may in fact be the single most important determinant. 6 As it would be very useful for clinicians to be able to use a certain pretreatment viral load for gauging responsiveness to interferon therapy, we made an attempt to derive a suitable clinical cut ov for each assay that would best predict the response to interferon. By using the ROC analysis, patients with an HCV viral load of less than 3.9 logs in the assay or 5.8 logs in the assay were considered likely to be complete responders to interferon given a dose of 6 MU thrice weekly for 24 weeks. When these cut ov points were used to analyse the interferon response in our treated patients retrospectively, the predictive rate was 70% with either of the two assays, suggesting that either assay could be used for routine viral load quantification to tailor treatment regimens. In patients with a high viral load who are expected to respond poorly to conventional interferon treatment, a higher dose, a longer duration of interferon treatment, or a combination of ribavirin plus interferon may be appropriate. 23 CONCLUSIONS Both the and assays are clinically useful methods for HCV RNA quan-
5 Quantification of hepatitis C virus RNA levels 811 tification, despite the existence of method related discrepancies in quantification values and in the correlation between HCV genotypes and viral load. Both assays are reliable for monitoring virological changes throughout the course of HCV infection and in selecting patients for treatment, and also may provide decision making information for future therapeutic strategies. However, the two assays cannot be used interchangeably owing to inherent diverences in technology and performance characteristics. Clinicians should weigh the advantages and weaknesses of each assay in selecting one particular assay for clinical studies and patient management. 1 Choo QL, Kuo G, Weiner AJ, et al. Isolation of a cdna clone derived from a blood-borne non-a, non-b viral hepatitis genome. Science 1989;244: Kuo G, Choo QL, Alter HJ, et al. An assay for circulating antibodies to a major etiologic virus of human non-a, non-b hepatitis. Science 1989;244: Alter MJ, Margolis HS, Krawczynski K, et al. The natural history of community-acquired hepatitis C in the United States. N Engl J Med 1992;327: Kato N, Yokosuka O, Hosoda K, et al. Quantification of hepatitis C virus by competitive reverse transcriptionpolymerase chain reaction: increase of the virus in advanced liver disease. Hepatology 1993;18: Hayashi J, Kishihara Y, Yoshimura E, et al. Relationship of genotype to level of hepatitis C viremia determined by competitive polymerase chain reaction. J Infect 1995;30: Hagiwara H, Hayashi N, Mita E, et al. Quantitative analysis of hepatitis C virus RNA in serum during interferon alfa therapy. Gastroenterology 1993;104: Toyoda H, Nakano S, Kumada T, et al. Comparison of serum hepatitis C virus RNA concentration by branched DNA probe assay with competitive reverse transcription polymerase chain reaction as a predictor of response to interferon-α therapy in chronic hepatitis C patients. JMed Virol 1996;48: Trabaud MA, Bailly F, Si-Ahmed SN, et al. Comparison of HCV RNA assays for the detection and quantification of hepatitis C virus RNA levels in serum of patients with chronic hepatitis C treated with interferon. J Med Virol 1997;52: Lau JYN, Davis GL, KniVen J, et al. Significance of serum hepatitis C virus RNA levels in chronic hepatitis C. Lancet 1993;341: Colucci G, Gutekunst K. Development of a quantitative PCR assay for monitoring HCV viremia levels in patients with chronic hepatitis C. J Viral Hepatitis 1997;4: Detmer J, Lagier R, Flynn J, et al. Accurate quantification of hepatitis C virus (HCV) RNA from all HCV genotype by using branched-dna technology. J Clin Microbiol 1996;34: Yu ML, Chuang WL, Lu SN, et al. The genotypes of hepatitis C virus in patients with chronic hepatitis C virus infection in Southern Taiwan. Kaohsiung J Med Sci 1996;12: Okamoto H, Tokita H, Sakamoto M, et al. Characterization of the genomic sequence of type V (or 3a) hepatitis C virus isolates and PCR primers for specific detection. J Gen Virol 1993;74: Hanley JA, McNeil BJ. A method of comparing the areas under receiver operating characteristic curves derived from the same cases. Radiology 1983;148: Zweig MH, Campbell G. Receiver-operating characteristic (ROC) plots: a fundamental evaluation tool in clinical medicine. Clin Chem 1993;39: Hawkins A, Davidson F, Simmonds P. Comparison of plasma virus loads among individuals infected with hepatitis C virus (HCV) genotypes 1, 2, and 3 by Quantiplex HCV RNA assay versions 1 and 2, Roche Monitor assay, and an in-house limiting dilution method. J Clin Microbiol 1997;35: Collins ML, Zayati C, Detmer JJ, et al. Preparation and characterization of RNA standards for use in quantitative branched DNA hybridization assays. Anal Biochem 1995; 226: Smith DB, Davidson F, Yap PL, et al. Levels of hepatitis C virus in blood donors infected with diverent viral genotypes. J Infect Dis 1996;173: Lau JYN, Davis GL, Prescott LE, et al. Distribution of hepatitis C virus genotypes determined by line probe assay in patients with chronic hepatitis C seen at tertiary referral centers in the United States. Ann Intern Med 1996;124: Lau JYN, Simmonds P, Urdea MS. Implications of variations of conserved regions of hepatitis C virus genome. Lancet 1995;346: Smith DB, Mellor J, Jarvis LM, et al. Variation of hepatitis C virus 5' non-coding region: implications for secondary structure, virus detection and typing. J Gen Virol 1995;6: Dubois DB, Gretch D, Rosa CD, et al. Quantitation of hepatitis C viral RNA in sera of hemodialysis patients: gender-related diverences in viral load. Am J Kidney Dis 1994;24: Reichard O, Norkrans G, Fryden A, et al. Randomised, double-blind, placebo-controlled trial of interferon α-2b with and without ribavirin for chronic hepatitis C. Lancet 1998;351:83 7. J Clin Pathol: first published as /jcp on 1 November Downloaded from on 14 May 2018 by guest. Protected by copyright.
W ith an estimated million people worldwide
 141 ORIGINAL ARTICLE Clinical evaluation of the COBAS Amplicor HBV monitor test for measuring serum HBV DNA and comparison with the Quantiplex branched DNA signal amplification assay in Taiwan C-Y Dai,
141 ORIGINAL ARTICLE Clinical evaluation of the COBAS Amplicor HBV monitor test for measuring serum HBV DNA and comparison with the Quantiplex branched DNA signal amplification assay in Taiwan C-Y Dai,
Performance Characteristics of the COBAS AMPLICOR Hepatitis C Virus MONITOR Test, Version 2.0
 Clinical Chemistry / EVALUATION OF QUANTITATIVE HEPATITIS C VIRUS ASSAY Performance Characteristics of the COBAS AMPLICOR Hepatitis C Virus MONITOR Test, Version 2.0 Maria Erali, MS, 1 Edward R. Ashwood,
Clinical Chemistry / EVALUATION OF QUANTITATIVE HEPATITIS C VIRUS ASSAY Performance Characteristics of the COBAS AMPLICOR Hepatitis C Virus MONITOR Test, Version 2.0 Maria Erali, MS, 1 Edward R. Ashwood,
Comparison of Quantitative HCV RNA Assays in Chronic Hepatitis C
 MICROBIOLOGY AND INFECTIOUS DISEASE Original Article Comparison of Quantitative HCV RNA Assays in Chronic Hepatitis C SHERAJ JACOB, MD, DEBORAH BAUDY, ELIZABETH JONES, LIZHE XU, PhD, ANDREW MASON, MBBS,
MICROBIOLOGY AND INFECTIOUS DISEASE Original Article Comparison of Quantitative HCV RNA Assays in Chronic Hepatitis C SHERAJ JACOB, MD, DEBORAH BAUDY, ELIZABETH JONES, LIZHE XU, PhD, ANDREW MASON, MBBS,
Genotype Dependence of Hepatitis C Virus Load Measurement in Commercially Available Quantitative Assays
 JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1999, p. 2525 2532 Vol. 37, No. 8 0095-1137/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Genotype Dependence of Hepatitis C
JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1999, p. 2525 2532 Vol. 37, No. 8 0095-1137/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Genotype Dependence of Hepatitis C
Performance of an automated system for quantification of hepatitis C virus RNA
 Journal of Virological Methods 86 (2000) 55 60 www.elsevier.com/locate/jviromet Performance of an automated system for quantification of hepatitis C virus RNA Anne Marie Roque Afonso a, *, Josiane Didier
Journal of Virological Methods 86 (2000) 55 60 www.elsevier.com/locate/jviromet Performance of an automated system for quantification of hepatitis C virus RNA Anne Marie Roque Afonso a, *, Josiane Didier
Antiviral Therapy 11: Hepatobiliary Division, Department of Internal Medicine, Kaohsiung Medical University Hospital, Kaohsiung, Taiwan 2
 Antiviral Therapy 11:985 994 A sustained virological response to interferon or interferon/ribavirin reduces hepatocellular carcinoma and improves survival in chronic hepatitis C: a nationwide, multicentre
Antiviral Therapy 11:985 994 A sustained virological response to interferon or interferon/ribavirin reduces hepatocellular carcinoma and improves survival in chronic hepatitis C: a nationwide, multicentre
CALIBRATION OF ANALYTICAL STANDARDS FOR HBV-DNA, HCV-RNA AND HIV-1 RNA IN GENOME COPIES BY A REFERENCE METHOD
 CALIBRATION OF ANALYTICAL STANDARDS FOR HBV-DNA, HCV-RNA AND HIV-1 RNA IN GENOME COPIES BY A REFERENCE METHOD AAJ van Drimmelen, E.R. Bax and W.G.V. Quint, BioQControl (BQC), Delft Diagnostic Laboratories
CALIBRATION OF ANALYTICAL STANDARDS FOR HBV-DNA, HCV-RNA AND HIV-1 RNA IN GENOME COPIES BY A REFERENCE METHOD AAJ van Drimmelen, E.R. Bax and W.G.V. Quint, BioQControl (BQC), Delft Diagnostic Laboratories
HEPATITIS C VIRUS GENOTYPING IN CHRONIC HEPATITIS C PATIENTS
 HEPATITIS C VIRUS GENOTYPING IN CHRONIC HEPATITIS C PATIENTS I. Qattan Centres for Hepatology, Royal Free & University College Medical School, London V. Emery Department of Virology, Royal Free & University
HEPATITIS C VIRUS GENOTYPING IN CHRONIC HEPATITIS C PATIENTS I. Qattan Centres for Hepatology, Royal Free & University College Medical School, London V. Emery Department of Virology, Royal Free & University
Interferon and ribavirin therapy for chronic hepatitis C virus genotype 6: A comparison with genotype 1
 Title Interferon and ribavirin therapy for chronic hepatitis C virus genotype 6: A comparison with genotype 1 Author(s) Hui, CK; Yuen, MF; Sablon, E; Chan, AOO; Wong, BCY; Lai, CL Citation Journal Of Infectious
Title Interferon and ribavirin therapy for chronic hepatitis C virus genotype 6: A comparison with genotype 1 Author(s) Hui, CK; Yuen, MF; Sablon, E; Chan, AOO; Wong, BCY; Lai, CL Citation Journal Of Infectious
HCV HIV +*+ Human immunodeficiency virus HIV hepatitis C virus HCV HIV HCV HCV HIV HIV
 ,**- The Japanese Society for AIDS Research The Journal of AIDS Research +0 HIV HCV + + + + + + + + +, +, :HCV HIV HAART / : +*+ +*0,**- Human immunodeficiency virus HIV hepatitis C virus HCV HIV,* HCV
,**- The Japanese Society for AIDS Research The Journal of AIDS Research +0 HIV HCV + + + + + + + + +, +, :HCV HIV HAART / : +*+ +*0,**- Human immunodeficiency virus HIV hepatitis C virus HCV HIV,* HCV
SYNOPSIS. Clinical Study Report AI Addendum #1. Open-label Dosing Phase
 Name of Sponsor/Company: Bristol-Myers Squibb Name of Finished Product: Individual Study Table Referring to the Dossier (For National Authority Use Only) Name of Active Ingredient: Entecavir SYNOPSIS Clinical
Name of Sponsor/Company: Bristol-Myers Squibb Name of Finished Product: Individual Study Table Referring to the Dossier (For National Authority Use Only) Name of Active Ingredient: Entecavir SYNOPSIS Clinical
Serum and liver HCV RNA levels in patients with chronic hepatitis C: correlation with clinical and histological features
 856 Department of Clinical and Experimental Medicine, University of Padova, Italy L De Moliner P Pontisso G L De Salvo L Cavalletto L Chemello A Alberti Correspondence to: Dr P Pontisso, Clinica Medica
856 Department of Clinical and Experimental Medicine, University of Padova, Italy L De Moliner P Pontisso G L De Salvo L Cavalletto L Chemello A Alberti Correspondence to: Dr P Pontisso, Clinica Medica
ORIGINAL INVESTIGATION. Age-Related Response to Interferon Alfa Treatment in Women vs Men With Chronic Hepatitis C Virus Infection
 ORIGINAL INVESTIGATION Age-Related to Interferon Alfa Treatment in Women vs Men With Chronic Hepatitis C Virus Infection Jun Hayashi, MD, PhD; Yasuhiro Kishihara, MD; Kumiko Ueno, MD; Kouzaburo Yamaji,
ORIGINAL INVESTIGATION Age-Related to Interferon Alfa Treatment in Women vs Men With Chronic Hepatitis C Virus Infection Jun Hayashi, MD, PhD; Yasuhiro Kishihara, MD; Kumiko Ueno, MD; Kouzaburo Yamaji,
Evaluation of Interferon Treatment in Cirrhotic Patients with Hepatitis C
 Evaluation of Interferon Treatment in Cirrhotic Patients with Hepatitis C Tatsuya IDE, Michio SATA, Hiroshi SUZUKI, Shiroh MURASHIMA, Ichiroh MIYAJIMA, Miki SHIRACHI and Kyuichi TANIKAWA The Second Department
Evaluation of Interferon Treatment in Cirrhotic Patients with Hepatitis C Tatsuya IDE, Michio SATA, Hiroshi SUZUKI, Shiroh MURASHIMA, Ichiroh MIYAJIMA, Miki SHIRACHI and Kyuichi TANIKAWA The Second Department
P0141 HBV 1000 copies/ml genotype reference panel
 P0141 HBV 1000 copies/ml genotype reference panel P0141 The kit insert contains a detailed protocol and should be read carefully before testing the run control to ensure optimal performance Table of contents
P0141 HBV 1000 copies/ml genotype reference panel P0141 The kit insert contains a detailed protocol and should be read carefully before testing the run control to ensure optimal performance Table of contents
Anti-hepatitis C virus core IgM antibodies correlate with hepatitis C recurrence and its severity in liver transplant patients
 698 Hepatology Unit, Hôtel-Dieu, Lyon 6988, France S N Si Ahmed Department of Liver Transplantation, Croix-Rousse, Lyon 697, France M Adham C Ducerf J Baulieux Laboratoire d Hygiène, Faculté Rockefeller,
698 Hepatology Unit, Hôtel-Dieu, Lyon 6988, France S N Si Ahmed Department of Liver Transplantation, Croix-Rousse, Lyon 697, France M Adham C Ducerf J Baulieux Laboratoire d Hygiène, Faculté Rockefeller,
Pegylated Interferon Alfa-2b (Peg-Intron) Plus Ribavirin (Rebetol)in the Treatment of Chronic Hepatitis C: A Local Experience
 Pegylated Interferon Alfa-2b (Peg-Intron) Plus Ribavirin (Rebetol)in the Treatment of Chronic Hepatitis C: A Local Experience E L Seow, PH Robert Ding Island Hospital, Penang, Malaysia. Introduction Hepatitis
Pegylated Interferon Alfa-2b (Peg-Intron) Plus Ribavirin (Rebetol)in the Treatment of Chronic Hepatitis C: A Local Experience E L Seow, PH Robert Ding Island Hospital, Penang, Malaysia. Introduction Hepatitis
Viral Hepatitis Diagnosis and Management
 Viral Hepatitis Diagnosis and Management CLINICAL BACKGROUND Viral hepatitis is a relatively common disease (25 per 100,000 individuals in the United States) caused by a diverse group of hepatotropic agents
Viral Hepatitis Diagnosis and Management CLINICAL BACKGROUND Viral hepatitis is a relatively common disease (25 per 100,000 individuals in the United States) caused by a diverse group of hepatotropic agents
Supplementary materials: Predictors of response to pegylated interferon in chronic hepatitis B: a
 Supplementary materials: Predictors of response to pegylated interferon in chronic hepatitis B: a real-world hospital-based analysis Yin-Chen Wang 1, Sien-Sing Yang 2*, Chien-Wei Su 1, Yuan-Jen Wang 3,
Supplementary materials: Predictors of response to pegylated interferon in chronic hepatitis B: a real-world hospital-based analysis Yin-Chen Wang 1, Sien-Sing Yang 2*, Chien-Wei Su 1, Yuan-Jen Wang 3,
Hepatitis C. Core slides
 Hepatitis C Core slides This material was prepared by the Viral Hepatitis Prevention Board The slides (or subsets) can be reproduced for educational use only, with reference to the original source and
Hepatitis C Core slides This material was prepared by the Viral Hepatitis Prevention Board The slides (or subsets) can be reproduced for educational use only, with reference to the original source and
Quantification of HBV, HCV genotype and HIV subtype panels
 Quantification of HBV, HCV genotype and HIV subtype panels Harry van Drimmelen 1,2, Wim Quint 2, Nico Lelie 3 and the international NAT study group 1. Biologicals Quality Control, 2. DDL Diagnostic Laboratory,
Quantification of HBV, HCV genotype and HIV subtype panels Harry van Drimmelen 1,2, Wim Quint 2, Nico Lelie 3 and the international NAT study group 1. Biologicals Quality Control, 2. DDL Diagnostic Laboratory,
HBV Core and Core-Related Antigen Quantitation in Chinese Patients with. Chronic Hepatitis B Genotype B and C Virus Infection
 Title page HBV Core and Core-Related Antigen Quantitation in Chinese Patients with Chronic Hepatitis B Genotype B and C Virus Infection Short Title: Quantitation of HBc and HBcrAg in Chinese patients Akinori
Title page HBV Core and Core-Related Antigen Quantitation in Chinese Patients with Chronic Hepatitis B Genotype B and C Virus Infection Short Title: Quantitation of HBc and HBcrAg in Chinese patients Akinori
Monitoring Hepatitis C
 Monitoring Hepatitis C Section Six Monitoring Hepatitis C Screening for hepatitis C is not routinely done, so you may have to request a test from your medical provider. This usually involves an antibody
Monitoring Hepatitis C Section Six Monitoring Hepatitis C Screening for hepatitis C is not routinely done, so you may have to request a test from your medical provider. This usually involves an antibody
Sequential Serum Hepatitis C Viral RNA Levels Longitudinally Assessed by Branched DNA Signal Amplification
 Sequential Serum Hepatitis C Viral RNA Levels Longitudinally Assessed by Branched DNA Signal Amplification STUART C. GORDON, 1,2 PETER J. DAILEY, 3 ANN L. SILVERMAN, 1,2 BILAL A. KHAN, 1 VALLISTRAM P.
Sequential Serum Hepatitis C Viral RNA Levels Longitudinally Assessed by Branched DNA Signal Amplification STUART C. GORDON, 1,2 PETER J. DAILEY, 3 ANN L. SILVERMAN, 1,2 BILAL A. KHAN, 1 VALLISTRAM P.
TRANSFUSION-ASSOCIATED HEPATITIS G VIRUS INFECTION AND ITS RELATION TO LIVER DISEASE
 TRANSFUSION-ASSOCIATED HEPATITIS G VIRUS INFECTION AND ITS RELATION TO LIVER DISEASE THE INCIDENCE OF TRANSFUSION-ASSOCIATED HEPATITIS G VIRUS INFECTION AND ITS RELATION TO LIVER DISEASE HARVEY J. ALTER,
TRANSFUSION-ASSOCIATED HEPATITIS G VIRUS INFECTION AND ITS RELATION TO LIVER DISEASE THE INCIDENCE OF TRANSFUSION-ASSOCIATED HEPATITIS G VIRUS INFECTION AND ITS RELATION TO LIVER DISEASE HARVEY J. ALTER,
on October 4, 2018 by guest
 JCM Accepts, published online ahead of print on 3 July 2012 J. Clin. Microbiol. doi:10.1128/jcm.01249-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 Title: Performance of
JCM Accepts, published online ahead of print on 3 July 2012 J. Clin. Microbiol. doi:10.1128/jcm.01249-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 Title: Performance of
Technical Bulletin No. 162
 CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 162 cobas 6800 HCV Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 162 cobas 6800 HCV Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
Viral Hepatitis The Preventive Potential of Antiviral Therapy. Thomas Berg
 Viral Hepatitis The Preventive Potential of Antiviral Therapy Thomas Berg Therapeutic and preventive strategies in patients with hepatitis virus infection Treatment of acute infection Treatment of chronic
Viral Hepatitis The Preventive Potential of Antiviral Therapy Thomas Berg Therapeutic and preventive strategies in patients with hepatitis virus infection Treatment of acute infection Treatment of chronic
Proposed 1 st IS for Hepatitis E Virus RNA WHO/BS/
 www.pei.de Proposed 1 st IS for Hepatitis E Virus RNA WHO/BS/09.2126 S. Baylis, Division of Virology, Paul-Ehrlich-Institut, Langen, Germany SoGAT XXII 14 th -15 th April 2011, Rome, Italy Presented at:
www.pei.de Proposed 1 st IS for Hepatitis E Virus RNA WHO/BS/09.2126 S. Baylis, Division of Virology, Paul-Ehrlich-Institut, Langen, Germany SoGAT XXII 14 th -15 th April 2011, Rome, Italy Presented at:
Strengths and Limitations of Commercial Tests for Hepatitis C Virus RNA Quantification
 JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 2004, p. 421 425 Vol. 42, No. 1 0095-1137/04/$08.00 0 DOI: 10.1128/JCM.42.1.421 425.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Strengths
JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 2004, p. 421 425 Vol. 42, No. 1 0095-1137/04/$08.00 0 DOI: 10.1128/JCM.42.1.421 425.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Strengths
Protection against persistence of hepatitis C
 Mechanisms of disease Protection against persistence of hepatitis C Shruti H Mehta, Andrea Cox, Donald R Hoover, Xiao-Hong Wang, Qing Mao, Stuart Ray, Steffanie A Strathdee, David Vlahov, David L Thomas
Mechanisms of disease Protection against persistence of hepatitis C Shruti H Mehta, Andrea Cox, Donald R Hoover, Xiao-Hong Wang, Qing Mao, Stuart Ray, Steffanie A Strathdee, David Vlahov, David L Thomas
Virologic factors related to interferon-a-induced thyroid dysfunction in patients with chronic hepatitis C
 European Journal of Endocrinology (2000) 142 431 437 ISSN 0804-4643 CLINICAL STUDY Virologic factors related to interferon-a-induced thyroid dysfunction in patients with chronic hepatitis C Ming-Chia Hsieh,
European Journal of Endocrinology (2000) 142 431 437 ISSN 0804-4643 CLINICAL STUDY Virologic factors related to interferon-a-induced thyroid dysfunction in patients with chronic hepatitis C Ming-Chia Hsieh,
Laboratory and Clinical Diagnosis of HCV Infection
 Laboratory and Clinical Diagnosis of HCV Infection Jean-Michel Pawlotsky,, MD, PhD Department of Virology (EA 3489) Henri Mondor Hospital University of Paris XII Créteil,, France I Nonspecific Liver Tests
Laboratory and Clinical Diagnosis of HCV Infection Jean-Michel Pawlotsky,, MD, PhD Department of Virology (EA 3489) Henri Mondor Hospital University of Paris XII Créteil,, France I Nonspecific Liver Tests
HIV-1 Viral Load Real Time (RG)
 -1 Viral Load Real Time (RG) Real Time RT-PCR type 1 RNA quantification assay MSP Reg. pending Valdense 3616. 11700. Montevideo. Uruguay. phone (598) 2 336 83 01. Fax (598) 2 336 71 60. Info@atgen.com.uy
-1 Viral Load Real Time (RG) Real Time RT-PCR type 1 RNA quantification assay MSP Reg. pending Valdense 3616. 11700. Montevideo. Uruguay. phone (598) 2 336 83 01. Fax (598) 2 336 71 60. Info@atgen.com.uy
Peginterferon alpha-2b plus ribavirin for chronic hepatitis C virus mixed genotype infection
 ORIGINAL ARTICLE July-August, Vol. 13 No. 4, 2014: 350-355 Peginterferon alpha-2b plus ribavirin for chronic hepatitis C virus mixed genotype infection Ching-Chung Lin,*,, Chia-Hsien Wu, Huan-Lin Chen,
ORIGINAL ARTICLE July-August, Vol. 13 No. 4, 2014: 350-355 Peginterferon alpha-2b plus ribavirin for chronic hepatitis C virus mixed genotype infection Ching-Chung Lin,*,, Chia-Hsien Wu, Huan-Lin Chen,
Virological Tools and Monitoring in the DAA Era
 Virological Tools and Monitoring in the DAA Era Prof. Jean-Michel Pawlotsky, MD, PhD National Reference Center for Viral Hepatitis B, C and delta Department of Virology & INSERM U955 Henri Mondor Hospital
Virological Tools and Monitoring in the DAA Era Prof. Jean-Michel Pawlotsky, MD, PhD National Reference Center for Viral Hepatitis B, C and delta Department of Virology & INSERM U955 Henri Mondor Hospital
Molecular Diagnosis Future Directions
 Molecular Diagnosis Future Directions Philip Cunningham NSW State Reference Laboratory for HIV/AIDS & Molecular Diagnostic Medicine Laboratory, SydPath St Vincent s Hospital Sydney Update on Molecular
Molecular Diagnosis Future Directions Philip Cunningham NSW State Reference Laboratory for HIV/AIDS & Molecular Diagnostic Medicine Laboratory, SydPath St Vincent s Hospital Sydney Update on Molecular
29th Viral Hepatitis Prevention Board Meeting
 29th Viral Hepatitis Prevention Board Meeting Madrid, November 2006 Treatment of chronic hepatitis B José M. Sánchez-Tapias Liver Unit Hospital Clínic University of Barcelona Spain CHRONIC HBV INFECTION
29th Viral Hepatitis Prevention Board Meeting Madrid, November 2006 Treatment of chronic hepatitis B José M. Sánchez-Tapias Liver Unit Hospital Clínic University of Barcelona Spain CHRONIC HBV INFECTION
Standardization of Hepatitis C Virus RNA Quantification
 Standardization of Hepatitis C Virus RNA Quantification JEAN-MICHEL PAWLOTSKY, 1,2 MAGALI BOUVIER-ALIAS, 1 CHRISTOPHE HEZODE, 3 FRANCOISE DARTHUY, 1 JOCELYNE REMIRE, 1 AND DANIEL DHUMEAUX 2,3 It was recently
Standardization of Hepatitis C Virus RNA Quantification JEAN-MICHEL PAWLOTSKY, 1,2 MAGALI BOUVIER-ALIAS, 1 CHRISTOPHE HEZODE, 3 FRANCOISE DARTHUY, 1 JOCELYNE REMIRE, 1 AND DANIEL DHUMEAUX 2,3 It was recently
Laboratory for Clinical and Biological Studies, University of Miami Miller School of Medicine, Miami, FL, USA.
 000000 00000 0000 000 00 0 bdna () 00000 0000 000 00 0 Nuclisens () 000 00 0 000000 00000 0000 000 00 0 Amplicor () Comparison of Amplicor HIV- monitor Test, NucliSens HIV- QT and bdna Versant HIV RNA
000000 00000 0000 000 00 0 bdna () 00000 0000 000 00 0 Nuclisens () 000 00 0 000000 00000 0000 000 00 0 Amplicor () Comparison of Amplicor HIV- monitor Test, NucliSens HIV- QT and bdna Versant HIV RNA
Diagnostic Methods of HBV infection. Zohreh Sharifi,ph.D of Virology Research center, Iranian Blood Transfusion Organization (IBTO)
 Diagnostic Methods of HBV infection Zohreh Sharifi,ph.D of Virology Research center, Iranian Blood Transfusion Organization (IBTO) Hepatitis B-laboratory diagnosis Detection of HBV infection involves
Diagnostic Methods of HBV infection Zohreh Sharifi,ph.D of Virology Research center, Iranian Blood Transfusion Organization (IBTO) Hepatitis B-laboratory diagnosis Detection of HBV infection involves
Should Elderly CHC Patients (>70 years old) be Treated?
 Should Elderly CHC Patients (>70 years old) be Treated? Deepak Amarapurkar Consultant Gastroenterologist & Hepatologist Bombay Hospital & Medical Research Center, Mumbai & Jagjivanram Western Railway Hospital,
Should Elderly CHC Patients (>70 years old) be Treated? Deepak Amarapurkar Consultant Gastroenterologist & Hepatologist Bombay Hospital & Medical Research Center, Mumbai & Jagjivanram Western Railway Hospital,
Prise en charge actuelle de l'hépatite C et nouvelles approches thérapeutiques
 Prise en charge actuelle de l'hépatite C et nouvelles approches thérapeutiques Future Complications of Darius Moradpour Service de Gastro-entérologie et d'hépatologie Centre Hospitalier Universitaire Vaudois
Prise en charge actuelle de l'hépatite C et nouvelles approches thérapeutiques Future Complications of Darius Moradpour Service de Gastro-entérologie et d'hépatologie Centre Hospitalier Universitaire Vaudois
HBV PUBLIC HEALTH IMPLICATIONS
 جزايری دکتر سيد محمد آزمايشگاه ھپاتيت B -دانشکده بھداشت ويروس شناسی- گروه دانشگاه علوم پزشکی تھران کنگره ارتقا کيفيت- ١٣٩٢ HBV PUBLIC HEALTH IMPLICATIONS 2 billion people have been infected by HBV worldwide.
جزايری دکتر سيد محمد آزمايشگاه ھپاتيت B -دانشکده بھداشت ويروس شناسی- گروه دانشگاه علوم پزشکی تھران کنگره ارتقا کيفيت- ١٣٩٢ HBV PUBLIC HEALTH IMPLICATIONS 2 billion people have been infected by HBV worldwide.
Trends in molecular diagnostics
 Trends in molecular diagnostics Detection of target genes of interest Quantification Infectious diseases HIV Hepatitis C & B TB / MAC Cytomegalovirus Herpes simplex Varicella zoster CT/GC HPV Profiling
Trends in molecular diagnostics Detection of target genes of interest Quantification Infectious diseases HIV Hepatitis C & B TB / MAC Cytomegalovirus Herpes simplex Varicella zoster CT/GC HPV Profiling
Prospective Multicenter Clinical Evaluation of AMPLICOR and COBAS AMPLICOR Hepatitis C Virus Tests
 JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2001, p. 4005 4012 Vol. 39, 11 0095-1137/01/$04.00 0 DOI: 10.1128/JCM.39.11.4005 4012.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2001, p. 4005 4012 Vol. 39, 11 0095-1137/01/$04.00 0 DOI: 10.1128/JCM.39.11.4005 4012.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
Prediction of HBsAg Loss by Quantitative HBsAg Kinetics during Long-Term 2015
 THAI J 16 GASTROENTEROL Treatment with Nucleos(t)ide Original Analogues Article Prediction of HBsAg Loss by Quantitative HBsAg Kinetics during Long-Term Treatment with Nucleos(t)ide Analogues Sombutsook
THAI J 16 GASTROENTEROL Treatment with Nucleos(t)ide Original Analogues Article Prediction of HBsAg Loss by Quantitative HBsAg Kinetics during Long-Term Treatment with Nucleos(t)ide Analogues Sombutsook
TO Approved for public release, unlimited
 UNCLASSIFIED AD NUMBER ADB232218 NEW LIMITATION CHANGE TO Approved for public release, unlimited distribution FROM Distribution authorized to U.S. Gov't. agencies only; Proprietary Information; Jul 96.
UNCLASSIFIED AD NUMBER ADB232218 NEW LIMITATION CHANGE TO Approved for public release, unlimited distribution FROM Distribution authorized to U.S. Gov't. agencies only; Proprietary Information; Jul 96.
Hepatitis C: Management of Previous Non-responders with First Line Protease Inhibitors
 Hepatitis C: Management of Previous Non-responders with First Line Protease Inhibitors Fred Poordad, MD The Texas Liver Institute Clinical Professor of Medicine University of Texas Health Science Center
Hepatitis C: Management of Previous Non-responders with First Line Protease Inhibitors Fred Poordad, MD The Texas Liver Institute Clinical Professor of Medicine University of Texas Health Science Center
COMPARISON OF HCV TREATMENT; HCV MONO-INFECTED AND HCV/HBV CO-INFECTED PATIENTS.
 The Professional Medical Journal DOI: 10.17957/TPMJ/16.3242 ORIGINAL PROF-3242 1. (MBBS, M.Phil), Department of Biochemistry, Foundation University Medical College, Islamabad. 2. (MBBS, FCPS), Department
The Professional Medical Journal DOI: 10.17957/TPMJ/16.3242 ORIGINAL PROF-3242 1. (MBBS, M.Phil), Department of Biochemistry, Foundation University Medical College, Islamabad. 2. (MBBS, FCPS), Department
entecavir, 0.5mg and 1mg film-coated tablets and 0.05 mg/ml oral solution, Baraclude SMC No. (747/11) Bristol-Myers Squibb Pharmaceuticals Ltd
 entecavir, 0.5mg and 1mg film-coated tablets and 0.05 mg/ml oral solution, Baraclude SMC No. (747/11) Bristol-Myers Squibb Pharmaceuticals Ltd 09 December 2011 The Scottish Medicines Consortium (SMC) has
entecavir, 0.5mg and 1mg film-coated tablets and 0.05 mg/ml oral solution, Baraclude SMC No. (747/11) Bristol-Myers Squibb Pharmaceuticals Ltd 09 December 2011 The Scottish Medicines Consortium (SMC) has
Appendix This appendix was part of the submitted manuscript and has been peer reviewed. It is posted as supplied by the authors.
 Appendix This appendix was part of the submitted manuscript and has been peer reviewed. It is posted as supplied by the authors. Appendix to: Thompson AJV; Expert panel representing the Gastroenterological
Appendix This appendix was part of the submitted manuscript and has been peer reviewed. It is posted as supplied by the authors. Appendix to: Thompson AJV; Expert panel representing the Gastroenterological
Viral Hepatitis. Dr Melissa Haines Gastroenterologist Waikato Hospital
 Viral Hepatitis Dr Melissa Haines Gastroenterologist Waikato Hospital Viral Hepatitis HAV HBV HCV HDV HEV Other viral: CMV, EBV, HSV Unknown Hepatitis A Hepatitis A Transmitted via the faecal-oral route
Viral Hepatitis Dr Melissa Haines Gastroenterologist Waikato Hospital Viral Hepatitis HAV HBV HCV HDV HEV Other viral: CMV, EBV, HSV Unknown Hepatitis A Hepatitis A Transmitted via the faecal-oral route
L\\\\\\1 V\\ High dose interferon treatment in chronic hepatitis C. Group A. Group B. Dosage. Treatment period
 SI 14 Gut 1993; supplement: SI 14-S118 Institute of Medical Science, School ofmedicine, St Marianna University, Japan S Iino Correspondence to: Dr S Iino, Institute of Medical Science, School of Medicine,
SI 14 Gut 1993; supplement: SI 14-S118 Institute of Medical Science, School ofmedicine, St Marianna University, Japan S Iino Correspondence to: Dr S Iino, Institute of Medical Science, School of Medicine,
-HCV genome is about 9400 nucleotides long, it is ssrna and positive sense -the 10 viral proteins are first made as a large polyprotein -individual
 2013: HCV Genome -HCV genome is about 9400 nucleotides long, it is ssrna and positive sense -the 10 viral proteins are first made as a large polyprotein -individual proteins are released from polyprotein
2013: HCV Genome -HCV genome is about 9400 nucleotides long, it is ssrna and positive sense -the 10 viral proteins are first made as a large polyprotein -individual proteins are released from polyprotein
Hepatocellular Carcinoma: Can We Slow the Rising Incidence?
 Hepatocellular Carcinoma: Can We Slow the Rising Incidence? K.Rajender Reddy M.D. Professor of Medicine Director of Hepatology Medical Director of Liver Transplantation University of Pennsylvania Outline
Hepatocellular Carcinoma: Can We Slow the Rising Incidence? K.Rajender Reddy M.D. Professor of Medicine Director of Hepatology Medical Director of Liver Transplantation University of Pennsylvania Outline
ARTICLE IN PRESS. A Treatment Algorithm for the Management of Chronic Hepatitis B Virus Infection in the United States: An Update
 CLINICAL GASTROENTEROLOGY AND HEPATOLOGY 2006;4:xxx REVIEW A Treatment Algorithm for the Management of Chronic Hepatitis B Virus Infection in the United States: An Update EMMET B. KEEFFE,* DOUGLAS T. DIETERICH,
CLINICAL GASTROENTEROLOGY AND HEPATOLOGY 2006;4:xxx REVIEW A Treatment Algorithm for the Management of Chronic Hepatitis B Virus Infection in the United States: An Update EMMET B. KEEFFE,* DOUGLAS T. DIETERICH,
Role of Hepatitis B Virus Genotypes in Chronic Hepatitis B Exacerbation
 BRIEF REPORT Role of Hepatitis B Virus Genotypes in Chronic Hepatitis B Exacerbation Man-Fung Yuen, 1 Erwin Sablon, 2 Danny Ka-Ho Wong, 1 He-Jun Yuan, 1 Benjamin Chun-Yu Wong, 1 Annie On-On Chan, 1 and
BRIEF REPORT Role of Hepatitis B Virus Genotypes in Chronic Hepatitis B Exacerbation Man-Fung Yuen, 1 Erwin Sablon, 2 Danny Ka-Ho Wong, 1 He-Jun Yuan, 1 Benjamin Chun-Yu Wong, 1 Annie On-On Chan, 1 and
Hepatitis C: A Hepatologist s Approach to an Infectious Disease
 CLINICAL PRACTICE Ellie J. C. Goldstein, Section Editor INVITED ARTICLE Hepatitis C: A Hepatologist s Approach to an Infectious Disease Peter M. Rosenberg Department of Medicine, St. John s Health Center,
CLINICAL PRACTICE Ellie J. C. Goldstein, Section Editor INVITED ARTICLE Hepatitis C: A Hepatologist s Approach to an Infectious Disease Peter M. Rosenberg Department of Medicine, St. John s Health Center,
Hepatitis C Virus (HCV)
 Clinical Practice Guidelines Hepatitis C Virus (HCV) OBJECTIVE The purpose is to guide the appropriate diagnosis and management of Hepatitis C Virus (HCV). GUIDELINE These are only guidelines, and are
Clinical Practice Guidelines Hepatitis C Virus (HCV) OBJECTIVE The purpose is to guide the appropriate diagnosis and management of Hepatitis C Virus (HCV). GUIDELINE These are only guidelines, and are
Rama Nada. - Malik
 - 2 - Rama Nada - - Malik 1 P a g e We talked about HAV in the previous lecture, now we ll continue the remaining types.. Hepatitis E It s similar to virus that infect swine, so its most likely infect
- 2 - Rama Nada - - Malik 1 P a g e We talked about HAV in the previous lecture, now we ll continue the remaining types.. Hepatitis E It s similar to virus that infect swine, so its most likely infect
Viral interaction and responses in chronic hepatitis C and B coinfected patients with interferon-alpha plus ribavirin combination therapy
 Antiviral Therapy 10:125 133 Viral interaction and responses in chronic hepatitis C and B coinfected patients with interferon-alpha plus ribavirin combination therapy Wan-Long Chuang, Chia-Yen Dai, Wen-Yu
Antiviral Therapy 10:125 133 Viral interaction and responses in chronic hepatitis C and B coinfected patients with interferon-alpha plus ribavirin combination therapy Wan-Long Chuang, Chia-Yen Dai, Wen-Yu
Hepatitis B Virus therapy. Maria Buti Hospital Universitario Valle Hebron Barcelona Spain
 Hepatitis B Virus therapy Maria Buti Hospital Universitario Valle Hebron Barcelona Spain Disclosures Advisor: AbbVie, Boehringer Ingelheim, Bristol-Myers Squibb, Gilead Sciences, Janssen, Merck Sharp &
Hepatitis B Virus therapy Maria Buti Hospital Universitario Valle Hebron Barcelona Spain Disclosures Advisor: AbbVie, Boehringer Ingelheim, Bristol-Myers Squibb, Gilead Sciences, Janssen, Merck Sharp &
Polymerase Chain Reaction RNA Assays To Establish
 JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1992, p. 2145-2149 0095-1137/92/082145-05$02.00/0 Copyright 1992, American Society for Microbiology Vol. 30, No. 8 Use of Aminotransferase, Hepatitis C Antibody,
JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 1992, p. 2145-2149 0095-1137/92/082145-05$02.00/0 Copyright 1992, American Society for Microbiology Vol. 30, No. 8 Use of Aminotransferase, Hepatitis C Antibody,
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Viral hepatitis and Hepatocellular Carcinoma
 Viral hepatitis and Hepatocellular Carcinoma Hashem B. El-Serag, MD, MPH Dan L. Duncan Professor of Medicine Chief, Gastroenterology and Hepatology Houston VA & Baylor College of Medicine Houston, TX Outline
Viral hepatitis and Hepatocellular Carcinoma Hashem B. El-Serag, MD, MPH Dan L. Duncan Professor of Medicine Chief, Gastroenterology and Hepatology Houston VA & Baylor College of Medicine Houston, TX Outline
An Update HBV Treatment
 An Update HBV Treatment Epidemiology Natural history Treatment Daryl T.-Y. Lau, MD, MPH Associate Professor of Medicine Director of Translational Liver Research Division of Gastroenterology BIDMC, Harvard
An Update HBV Treatment Epidemiology Natural history Treatment Daryl T.-Y. Lau, MD, MPH Associate Professor of Medicine Director of Translational Liver Research Division of Gastroenterology BIDMC, Harvard
HEPATITIS C VIRUS (HCV) GENOTYPE TESTING
 CLINICAL GUIDELINES For use with the UnitedHealthcare Laboratory Benefit Management Program, administered by BeaconLBS HEPATITIS C VIRUS (HCV) GENOTYPE TESTING Policy Number: PDS - 027 Effective Date:
CLINICAL GUIDELINES For use with the UnitedHealthcare Laboratory Benefit Management Program, administered by BeaconLBS HEPATITIS C VIRUS (HCV) GENOTYPE TESTING Policy Number: PDS - 027 Effective Date:
Hepatitis C Virus RNA Profiles in Chronically Infected Individuals: Do They Relate to Disease Activity?
 Hepatitis C Virus RNA Profiles in Chronically Infected Individuals: Do They Relate to Disease Activity? PATRIZIA PONTISSO, 1 GIORGIO BELLATI, 2 MAURIZIA BRUNETTO, 2 LILIANA EMELLO, 1 GUIDO COLLOREDO, 2
Hepatitis C Virus RNA Profiles in Chronically Infected Individuals: Do They Relate to Disease Activity? PATRIZIA PONTISSO, 1 GIORGIO BELLATI, 2 MAURIZIA BRUNETTO, 2 LILIANA EMELLO, 1 GUIDO COLLOREDO, 2
Monitoring of HCV RNA. Hepatitis C Requirements of Antiviral Therapy Monitoring
 Monitoring of HCV RNA Hepatitis C Requirements of Antiviral Therapy Monitoring Hepatitis C Requirements of Antiviral Therapy Monitoring Author: Ute Hofmann,, Konrad-Zuse-Str. 1, 07745 Jena, Germany Beatrix
Monitoring of HCV RNA Hepatitis C Requirements of Antiviral Therapy Monitoring Hepatitis C Requirements of Antiviral Therapy Monitoring Author: Ute Hofmann,, Konrad-Zuse-Str. 1, 07745 Jena, Germany Beatrix
Natural History of HBV Infection
 Natural History of HBV Infection Joseph JY Sung MD PhD Institute of Digestive Disease Department of Medicine & Therapeutics Prince of Wales Hospital The Chinese University of Hong Kong HBV Infection 2
Natural History of HBV Infection Joseph JY Sung MD PhD Institute of Digestive Disease Department of Medicine & Therapeutics Prince of Wales Hospital The Chinese University of Hong Kong HBV Infection 2
Clinical Outcome of Hepatitis C as a Function of Mode of Transmission
 Clinical Outcome of Hepatitis C as a Function of Mode of Transmission STUART C. GORDON, 1,2 NASSER BAYATI, 1 AND ANN L. SILVERMAN 1,2 Abbreviations: HCV, hepatitis C virus, ALT, alanine aminotransferase.
Clinical Outcome of Hepatitis C as a Function of Mode of Transmission STUART C. GORDON, 1,2 NASSER BAYATI, 1 AND ANN L. SILVERMAN 1,2 Abbreviations: HCV, hepatitis C virus, ALT, alanine aminotransferase.
Technical Bulletin No. 161
 CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 161 cobas 6800 HIV-1 Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 161 cobas 6800 HIV-1 Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
Introduction to the Impact of Resistance in Hepatitis C
 Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Hepatitis C Virus. https://www.labcorp.com/wps/wcm/connect/labcorp+content/labcorp/education+and+re...
 Page 1 of 16 Hepatitis C Virus Data reflected in this report are based solely on the collection of samples submitted to LabCorp for testing. Refer to the limitations section of this report for additional
Page 1 of 16 Hepatitis C Virus Data reflected in this report are based solely on the collection of samples submitted to LabCorp for testing. Refer to the limitations section of this report for additional
Hepatitis C Management and Treatment
 Hepatitis C Management and Treatment Kaya Süer Near East University Faculty of Medicine Infectious Diseases and Clinical Microbiology 1 Discovery of Hepatitis C Key facts Hepatitis C: the virus can cause
Hepatitis C Management and Treatment Kaya Süer Near East University Faculty of Medicine Infectious Diseases and Clinical Microbiology 1 Discovery of Hepatitis C Key facts Hepatitis C: the virus can cause
Pharmacological management of viruses in obese patients
 Cubist Pharmaceuticals The Shape of Cures to Come Pharmacological management of viruses in obese patients Dr. Dimitar Tonev, Medical Director UKINORD 1 Disclosures } The author is a pharmaceutical physician
Cubist Pharmaceuticals The Shape of Cures to Come Pharmacological management of viruses in obese patients Dr. Dimitar Tonev, Medical Director UKINORD 1 Disclosures } The author is a pharmaceutical physician
Feature. Downloaded from by guest on 06 November 2018
 Hepatitis C Virus: Epidemiology, Diagnosis, and Patient Management Peter P. Chou, PhD, DABCC, FACB (Quest Diagnostics Nichols Institute, Chantilly, VA) DOI: 10.1309/BNK8PH8FEPJ0VCH2 Laboratory photo courtesy
Hepatitis C Virus: Epidemiology, Diagnosis, and Patient Management Peter P. Chou, PhD, DABCC, FACB (Quest Diagnostics Nichols Institute, Chantilly, VA) DOI: 10.1309/BNK8PH8FEPJ0VCH2 Laboratory photo courtesy
Performance of Version 2.0 of the Cobas AmpliPrep/Cobas TaqMan Real-Time PCR Assay for Hepatitis B Virus DNA Quantification
 JOURNAL OF CLINICAL MICROBIOLOGY, Oct. 2010, p. 3641 3647 Vol. 48, No. 10 0095-1137/10/$12.00 doi:10.1128/jcm.01306-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. Performance
JOURNAL OF CLINICAL MICROBIOLOGY, Oct. 2010, p. 3641 3647 Vol. 48, No. 10 0095-1137/10/$12.00 doi:10.1128/jcm.01306-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. Performance
The treatment of choice for chronic hepatitis C is
 Early Identification of HCV Genotype 1 Patients Responding to 24 Weeks Peginterferon -2a (40 kd)/ribavirin Therapy Donald M. Jensen, 1 Timothy R. Morgan, 2 Patrick Marcellin, 3 Paul J. Pockros, 4 K. Rajender
Early Identification of HCV Genotype 1 Patients Responding to 24 Weeks Peginterferon -2a (40 kd)/ribavirin Therapy Donald M. Jensen, 1 Timothy R. Morgan, 2 Patrick Marcellin, 3 Paul J. Pockros, 4 K. Rajender
The ABCs of Viral Hepatitis Diagnosis. Ila Singh, M.D., Ph.D. P & S Viral Hepatitis. Hepatitis A, B, C, D, E and G viruses
 The ABCs of Viral Hepatitis Diagnosis Ila Singh, M.D., Ph.D. P & S 14-453 is132@columbia.edu Viral Hepatitis Hepatotropic viruses Hepatitis A, B, C, D, E and G viruses Generalized infection plus infection
The ABCs of Viral Hepatitis Diagnosis Ila Singh, M.D., Ph.D. P & S 14-453 is132@columbia.edu Viral Hepatitis Hepatotropic viruses Hepatitis A, B, C, D, E and G viruses Generalized infection plus infection
Estimation of Seroprevalence of Hepatitis B Virus and Hepatitis C Virus in Taiwan from a Large-scale Survey of Free Hepatitis Screening Participants
 ORIGINAL ARTICLE Estimation of Seroprevalence of Hepatitis B Virus and Hepatitis C Virus in Taiwan from a Large-scale Survey of Free Hepatitis Screening Participants Chien-Hung Chen, 1 Pei-Ming Yang, 1
ORIGINAL ARTICLE Estimation of Seroprevalence of Hepatitis B Virus and Hepatitis C Virus in Taiwan from a Large-scale Survey of Free Hepatitis Screening Participants Chien-Hung Chen, 1 Pei-Ming Yang, 1
Pelagia Research Library. European Journal of Experimental Biology, 2015, 5(10):1-5
 Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 2015, 5(10):1-5 ISSN: 2248 9215 CODEN (USA): EJEBAU Molecular diagnosis of human immuno deficiency virus (HIV)
Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 2015, 5(10):1-5 ISSN: 2248 9215 CODEN (USA): EJEBAU Molecular diagnosis of human immuno deficiency virus (HIV)
MedInform. HBV DNA loss in Bulgarian patients on NUC therapy. Speed related factors. (NUC related speed of HBV DNA loss in Bulgaria) Original Article
 DOI: 10.18044/Medinform.201852.897 ISSUE 3, 2018 HBV DNA loss in Bulgarian patients on NUC therapy. Speed related factors. (NUC related speed of HBV DNA loss in Bulgaria) Donika Krasteva, Radosveta Tomova,
DOI: 10.18044/Medinform.201852.897 ISSUE 3, 2018 HBV DNA loss in Bulgarian patients on NUC therapy. Speed related factors. (NUC related speed of HBV DNA loss in Bulgaria) Donika Krasteva, Radosveta Tomova,
29th Viral Hepatitis Prevention Board Meeting
 29th Viral Hepatitis Prevention Board Meeting Madrid, November 2006 Treatment of chronic hepatitis C José M. Sánchez-Tapias Liver Unit Hospital Clínic University of Barcelona Spain CHRONIC HEPATITIS C
29th Viral Hepatitis Prevention Board Meeting Madrid, November 2006 Treatment of chronic hepatitis C José M. Sánchez-Tapias Liver Unit Hospital Clínic University of Barcelona Spain CHRONIC HEPATITIS C
Experience with Standardisation of Blood Virology NAT. Clare Morris Division of Retrovirology National Institute for Biological Standards and Control
 Experience with Standardisation of Blood Virology NAT Clare Morris Division of Retrovirology National Institute for Biological Standards and Control Background of Blood Virology Standardisation In the
Experience with Standardisation of Blood Virology NAT Clare Morris Division of Retrovirology National Institute for Biological Standards and Control Background of Blood Virology Standardisation In the
Long-term tracking of hepatitis B viral load and the relationship with risk for hepatocellular carcinoma in men
 Carcinogenesis vol.29 no.1 pp.106 112, 2008 doi:10.1093/carcin/bgm252 Advance Access publication November 13, 2007 Long-term tracking of hepatitis B viral load and the relationship with risk for hepatocellular
Carcinogenesis vol.29 no.1 pp.106 112, 2008 doi:10.1093/carcin/bgm252 Advance Access publication November 13, 2007 Long-term tracking of hepatitis B viral load and the relationship with risk for hepatocellular
Assays to Address Emerging Threats to Blood Safety
 Assays to Address Emerging Threats to Blood Safety Jeffrey M. Linnen, Ph.D. Director, Product Development Gen-Probe Incorporated San Diego, CA The IPFA/PEI 17th Workshop on Surveillance and Screening of
Assays to Address Emerging Threats to Blood Safety Jeffrey M. Linnen, Ph.D. Director, Product Development Gen-Probe Incorporated San Diego, CA The IPFA/PEI 17th Workshop on Surveillance and Screening of
Current therapy for hepatitis C: pegylated interferon and ribavirin
 Clin Liver Dis 7 (2003) 149 161 Current therapy for hepatitis C: pegylated interferon and ribavirin John G. McHutchison, MD a, Michael W. Fried, MD b, * a Duke Clinical Research Institute, Duke University
Clin Liver Dis 7 (2003) 149 161 Current therapy for hepatitis C: pegylated interferon and ribavirin John G. McHutchison, MD a, Michael W. Fried, MD b, * a Duke Clinical Research Institute, Duke University
Fulminant hepatic failure in acute hepatitis C: increased risk in chronic carriers of hepatitis B virus
 Gut 1999;45:613 617 613 Liver Research Unit, Chang Gung Memorial Hospital and Medical College, Taipei, Taiwan C M Chu CTYeh Y F Liaw Correspondence to: Dr C M Chu, Liver Research Unit, Chang Gung Memorial
Gut 1999;45:613 617 613 Liver Research Unit, Chang Gung Memorial Hospital and Medical College, Taipei, Taiwan C M Chu CTYeh Y F Liaw Correspondence to: Dr C M Chu, Liver Research Unit, Chang Gung Memorial
Automated Quantitative Analysis of Hepatitis B Virus DNA by Using the Cobas Amplicor HBV Monitor Test
 JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1999, p. 2793 2797 Vol. 37, No. 9 0095-1137/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Automated Quantitative Analysis of
JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1999, p. 2793 2797 Vol. 37, No. 9 0095-1137/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Automated Quantitative Analysis of
A "State-of-the-Art" Conference Hepatitis C: A Meeting Ground for the Generalist and the Specialist
 A "State-of-the-Art" Conference Hepatitis C: A Meeting Ground for the Generalist and the Specialist Information regarding pathogenesis and appropriate management of chronic hepatitis C continues to evolve.
A "State-of-the-Art" Conference Hepatitis C: A Meeting Ground for the Generalist and the Specialist Information regarding pathogenesis and appropriate management of chronic hepatitis C continues to evolve.
Hepatitis B Virus therapy. Maria Buti Hospital Universitario Valle Hebron Barcelona Spain
 Hepatitis B Virus therapy Maria Buti Hospital Universitario Valle Hebron Barcelona Spain Disclosures Advisor: AbbVie, Boehringer Ingelheim, Bristol-Myers Squibb, Gilead Sciences, Janssen, Merck Sharp &
Hepatitis B Virus therapy Maria Buti Hospital Universitario Valle Hebron Barcelona Spain Disclosures Advisor: AbbVie, Boehringer Ingelheim, Bristol-Myers Squibb, Gilead Sciences, Janssen, Merck Sharp &
QUANTITATIVE HEPATITIS C VIRUS (HCV) RNA
 CLINICAL GUIDELINES For use with the UnitedHealthcare Laboratory Benefit Management Program, administered by BeaconLBS QUANTITATIVE HEPATITIS C VIRUS (HCV) RNA Policy Number: PDS - 028 Effective Date:
CLINICAL GUIDELINES For use with the UnitedHealthcare Laboratory Benefit Management Program, administered by BeaconLBS QUANTITATIVE HEPATITIS C VIRUS (HCV) RNA Policy Number: PDS - 028 Effective Date:
PROPOSAL FOR THE DEVELOPMENT OF AN INTERNATIONAL REFERENCE PREPARATION FOR HEPATITIS D VIRUS RNA
 PROPOSAL FOR THE DEVELOPMENT OF AN INTERNATIONAL REFERENCE PREPARATION FOR HEPATITIS D VIRUS RNA SoGAT Clinical Diagnostics II 30 September / 1 October 2009, Istanbul Michael Chudy Julia Kreß C. Micha
PROPOSAL FOR THE DEVELOPMENT OF AN INTERNATIONAL REFERENCE PREPARATION FOR HEPATITIS D VIRUS RNA SoGAT Clinical Diagnostics II 30 September / 1 October 2009, Istanbul Michael Chudy Julia Kreß C. Micha
L iver steatosis is a frequent histological finding in patients
 420 LIVER Effect of antiviral treatment on evolution of liver steatosis in patients with chronic hepatitis C: indirect evidence of a role of hepatitis C virus genotype 3 in steatosis L Castéra, C Hézode,
420 LIVER Effect of antiviral treatment on evolution of liver steatosis in patients with chronic hepatitis C: indirect evidence of a role of hepatitis C virus genotype 3 in steatosis L Castéra, C Hézode,
Hepatitis Delta Virus and GBV-C Infection in Two Neighboring Hepatitis B Virus and Hepatitis C Virus Endemic Villages in Taiwan
 Original Article 137 Hepatitis Delta Virus and GBV-C Infection in Two Neighboring Hepatitis B Virus and Hepatitis C Virus Endemic Villages in Taiwan Chang-Jung Chang 1, MD; Jui-Chin Chiang 1,2,6, MD; Sheng-Nan
Original Article 137 Hepatitis Delta Virus and GBV-C Infection in Two Neighboring Hepatitis B Virus and Hepatitis C Virus Endemic Villages in Taiwan Chang-Jung Chang 1, MD; Jui-Chin Chiang 1,2,6, MD; Sheng-Nan
Hepatitis delta: often forgotten?
 15 th Annual Resistance and Antiviral Therapy Meeting Dr Sarah Hughes King s College Hospital, London Thursday 29 September 2011, Royal College of Physicians, London Hepatitis delta: often forgotten? Dr
15 th Annual Resistance and Antiviral Therapy Meeting Dr Sarah Hughes King s College Hospital, London Thursday 29 September 2011, Royal College of Physicians, London Hepatitis delta: often forgotten? Dr
Conventional Interferon Alfa-2b and Ribavirin for 12 Versus 24 Weeks in HCV Genotype 2 or 3
 ORIGINAL ARTICLE Conventional Interferon Alfa-2b and Ribavirin for 12 Versus 24 Weeks in HCV Genotype 2 or 3 Javed Iqbal Farooqi and Rukhsana Javed Farooqi* ABSTRACT Objective: To determine the efficacy
ORIGINAL ARTICLE Conventional Interferon Alfa-2b and Ribavirin for 12 Versus 24 Weeks in HCV Genotype 2 or 3 Javed Iqbal Farooqi and Rukhsana Javed Farooqi* ABSTRACT Objective: To determine the efficacy
The Alphabet Soup of Viral Hepatitis Testing
 The Alphabet Soup of Viral Hepatitis Testing August 18, 2011 Patricia Slev, PhD, DABCC Medical Director, Serologic Hepatitis and Retrovirus Laboratory, ARUP Laboratories Assistant Professor of Pathology,
The Alphabet Soup of Viral Hepatitis Testing August 18, 2011 Patricia Slev, PhD, DABCC Medical Director, Serologic Hepatitis and Retrovirus Laboratory, ARUP Laboratories Assistant Professor of Pathology,
