Compensatory mutations at the HIV cleavage sites p7/p1 and p1/p6 gag in therapy-naive and therapy-experienced patients
|
|
|
- Candace Stafford
- 6 years ago
- Views:
Transcription
1 Antiviral Therapy 11: Compensatory mutations at the HIV cleavage sites p7/p1 and p1/p6 gag in therapy-naive and therapy-experienced patients Jens Verheyen 1 *, Elena Litau 1, Tobias Sing 2, Martin Däumer 1, Melanie Balduin 1, Mark Oette 3, Gerd Fätkenheuer 4, Jürgen K Rockstroh 5, Ulrike Schuldenzucker 6, Daniel Hoffmann 7, Herbert Pfister 1 and Rolf Kaiser 1 1 Institute of Virology, University of Cologne, Cologne, Germany 2 Max Planck Institute for Informatics, Saarbrücken, Germany 3 Clinic for Gastroenterology, Hepatology and infectious Diseases, University Clinic Düsseldorf, Germany 4 Department of Internal Medicine 1, University of Cologne, Cologne, Germany 5 Department of Internal Medicine 1, University of Bonn, Bonn, Germany 6 Caesar, Bonn, Germany 7 Center for Medical Biotechnology, University of Duisburg-Essen, Essen, Germany *Corresponding author: Tel: ; Fax: ; jens.verheyen@uk-koeln.de Background: Mutations in the genome of HIV conferring drug resistance are a major reason for the failure of antiretroviral therapy, but they often compromise viral fitness. Protease (PR) cleavage site (CS) mutations could compensate for impaired replication capacity of drug-resistant viruses. Patients and methods: We analysed the cleavage sites p1/p7 and p1/p6 gag of 500 HIV-1 subtype B infected patients. The collective consists of 275 therapy-naive and 225 therapy-experienced patients with at least one primary PR mutation, from whom eight underwent therapy-interruption in different clinical settings. Results: Multiple mutations within the CS p7/p1 and p1/p6 gag accumulated in therapy-experienced isolates (p7/p1: A431V K436R I437V and p1/p6 gag: L449F/V P452S P453L/A). Further rare CS mutations were totally absent in therapy-naive viruses. Sixty percent of all therapy-experienced viruses exhibited at least one therapy-associated CS mutation, but so did 10% of therapy-naive viruses. The analysis of CS and PR mutations in therapy-experienced viruses revealed several positive correlations A431V with L24I M46I/L I54V V82A; I437V with I54V V82F/T/S; L449V with I54M/L/S/T/A; and L449F/R452S/P453L: with D30N I84V whereas P453L and V82A were negatively correlated. Mutagenetic trees constructed form this cross-sectional data showed an ordered accumulation of the most prominent CS mutations along two pathways L90M I84V P453L and I54 V82 A431V followed by either M46L or L24I. Furthermore, eight viruses with at least one therapyassociated mutation at each CS displayed an outstanding maintenance of major PR mutations during therapy interruption. Conclusions: These findings emphasize the relevance of CS mutations in the evolution of HIV resistance to PR inhibitors. Therefore, therapy-associated CS mutations should be considered in HIV resistance tests to estimate viral fitness in different clinical settings. Introduction Highly active antiretroviral therapy (HAART) effectively inhibits the replication of the HIV in most patients. Apart from compliance problems, the emergence of resistance-associated mutations within the target proteins protease (PR) and reverse transcriptase (RT) is one of the major reasons for therapy failure. Some PR mutations conferring drug resistance also influence the enzymatic activity of the PR, leading to decreased viral fitness compared with wild type. Compensatory changes were observed at secondary sites within the PR after the occurrence of primary PR mutations that had affected enzymatic activity [1]. The processing of HIV precursor proteins into functional viral proteins by the impaired mutant PR could be improved through modification of its substrate peptides [2]. The 2006 International Medical Press
2 J Verheyen et al. cleavage sites (CS) within these precursor proteins exhibit different cleavage rates, demonstrating the gradual activation of viral enzymes and liberation of structural proteins as a crucial step in virus replication. The initial cleavage takes place at CS p2/p7, followed by cleavage of secondary CS p1/p6 gag and p17/p24 and finally of the less adapted tertiary CS p7/p1 and p24/p7 [3,4]. With the introduction of protease inhibitors (PIs) into the antiretroviral therapy, the occurrence of CS mutations has been reported from cell culture experiments and in vivo studies [5,6]. Mainly mutations at the two CS p7/p1 and p1/p6 gag have been discussed as compensatory mutations restoring the compromised enzymatic activity of mutated PRs. In vitro experiments proved the compensatory character of certain CS mutations (A431V/L449F/P453L) for specific PR mutations [7 10]. A great variety of CS mutations have been associated with therapy exposure and with certain PR mutations. However, the patient cohorts and analysed mutations varied largely and a comprehensive approach for analysing these CS mutations with regard to resistance profiles is still missing. The knowledge of relations between resistance to different PI and compensatory pathways could provide deeper insights into the evolution of drug resistance. Moreover, viral fitness gains increasing attention in different clinical settings, especially during therapy-interruption or maintenance of a failing therapy regimen. Great efforts are therefore being made to establish fitness assays that could measure compensatory effects of mutations and verify results from observations of cell culture or patient cohorts. We analysed gag mutations in numerous HIVsubtype B samples between therapy-naive (TN) patients along with a comparable number of patients who had failed previous therapies, with primary PR mutations. The purpose of this strategy was to distinguish natural polymorphisms from therapy associated mutations. Patients and methods Patients Blood samples were collected in the years 2001 to 2004 from two groups: from therapy-naive TN patients from the RESINA study [13 15], all of whom were infected with HIV-1 subtype B without primary protease mutations; and from therapy-experienced (TE) patients from the university clinics of Cologne, Düsseldorf and Bonn, all of whom were also infected with HIV-1 subtype B. The TN viruses were analysed before the initiation of HAART after a variable time of HIV infection. Most TE patients were heavily pretreated and obtained a genotypic HIV resistance test after therapy failure, revealing at least one of known primary PR mutations: L24I, D30N, V32I, M46I/L, I50L/V, I54V/M/S/L/T/A, V82A/F/T/S, I84V, N88S/T, L90M. Eight patients with highly resistant viruses underwent therapy interruption for different clinical reasons, and HIV genotypes were compared before and during therapy interruption. Genotypic resistance tests HIV from all patients were genotyped and the genotypic resistance test was performed as described previously [14]. Briefly, a OneStep RT-PCR (Qiagen, Hilden, Germany) with primers RES1, 5 -GAAGAAATGATGACAGCATGTCAGGG-3 (nucleotides [nt] 1,819 1,844) and RES2, 5 -TAATT- TATTACTTGTTCATTTCCTCCAAT-3 (nt 4,173 4,202) was carried out, followed by a nested PCR with HotStartTaq (Qiagen, Hilden, Germany) and primers Res3, 5 -AGACAGGCTAATTTTT- TAGGGA-3 (nt 2,074 2,095) and Res4, 5 -ATGGYTCTTGATAAATTTGATATGTCC-3 (nt 3,559 3,585). The population based sequencing of HIV-1 pol region was performed with the ViroSeq HIV-1 Genotyping System sequencing module (Applied Biosystems, Foster City, CA, USA). HIV-1 pol sequences were assembled and edited with the ViroSeq HIV-1 Genotyping software v2.5 (Applied Biosystems, Foster City, CA, USA). Analysis of the C-terminal HIV gag gene Viral RNA was isolated from patient s plasma using QIAamp viral RNA mini kit (Qiagen, Hilden, Germany) according to manufacturer s protocol. Reverse Transcription and PCR were carried out using OneStep RT-PCR kit (Qiagen, Hilden, Germany) and primers C-gag1, 5 -CCAATTCCCC- CTTCATTTTTGG-3 (nt 2,404 2,382) and C-gag2, 5 -GGCTGTTGGAAATGTGGAAAGGA-3 (nt 2,023 2,045). The 0.4 kilobase pair PCR product was purified with the QIAquick spin PCR purification kit (Qiagen, Hilden, Germany). The sequence reaction was performed with the same separated primers and ABI reaction mix. Extension products were purified using MultiScreen purification plates (Millipore, Bedford, MA, USA) and Sephadex G-50 superfine (Amersham Biosciences, Uppsala, Sweden) and were run on an ABI Prism 310 capillary sequencer. The obtained sequences were assembled and edited by using the Seqman software and translated thereafter into protein sequences according to the gag and pol reading frame. Each sequence was aligned to the Hxb2 reference sequence and amino acid substitutions were documented (gag mutations in italic, PR mutations in non-italic). Mixtures of International Medical Press
3 Compensatory cleavage site mutations in HIV mutant and wild-type amino acids at resistanceassociated PR positions and CS positions were counted as mutant. Statistical analysis Categorical variables were analysed using either the χ 2 test of association or Fisher s exact test, according to the size of the analysed group. P<0.05 was considered as significant. The mtreemix software package is described by Beerenwinkel et al. [16,17]. Briefly, assuming that the appearance of mutations is permanent, a local maximum likelihood mutagenetic tree indicating their accumulation order can be estimated from cross-sectional data using a combination of a graph-theoretical method with an Expectation- Maximization approach. Results CS p7/p1 and p1/p6 gag mutations in TN and TE viruses Five amino acids in both directions from each cleavage site were compared (Table 1). The two carboxy-terminal gag cleavage sites (p7/p1 and p1/p6 gag) proved significant differences between the two groups. The leading therapy-associated mutations within the p7/p1 CS were A431V (P<0.0001) and I437V (P<0.0001), which were found only rarely in TN viruses. The mutation K436R tended to be more frequent in TE viruses but barely reached significance (P=0.029), whereas I437L occurred in similar number in both groups. The variety of CS mutations in comparison to HxB2 at the CS p1/p6 gag was much higher than at CS p7/p1. Five p1/p6 gag mutations were significantly more frequent in TE viruses (L449V (P<0.005), L449F (P<0.005), R452S (P<0.05), P453A (P<0.01) and P453L (P<0.0001)). At both CS some rare mutations were present almost exclusively among TE viruses: K436N (n=1), K436E (n=1), I437M (n=1), L449H (n=2), R451E (n=1), R451SS (n=1), R452K (n=1), R452Q (n=1), P453I (TE [n=4] vs TN [n=1]) P453S (n=3) and P453V (n=2), but each individual mutation failed significance due to their rare occurrence. By contrast, L449P occurred in similar number in both groups and was usually accompanied by P453 mutations, mainly P453L, which was much more frequent in viruses from TE patients when L449P was missing (TN [n=5], TE [n=46]) was further on named P453L *. However, P453T without additional L449P still lacked significance (P=0.176). No single S451 mutation exhibited was associated with exposure to antiretroviral therapy, however S451T failed only barely significance (TN [n=3] vs TE [n=8]; P=0.072; Table 1). Table 1. CS mutations in viruses from TN and TE patients TN viruses TE viruses CS mutations (n=275) (n=225) P-value CS p7/p1, n (%) A431 3 (1.1) 61 (27.1) V 1 (0.4) 60 (26.7) P< D 2 (0.7) 0 (0.0) I 0 (0.0) 1 (0.4) K (5.5) 23 (10.2) R 12 (4.3) 21 (9.3) P<0.05 G 2 (0.7) 0 (0.0) T 1 (0.4) 0 (0.0) N/E 0 (0.0) 2 (0.0) I (6.2) 42 (18.7) L 8 (2.9) 6 (2.7) V 9 (3.3) 35 (15.6) P< M 0 (0.0) 1 (0.4) CS p1/p6 gag, n (%) L (10.5) 41 (18.2) F 4 (1.5) 17 (7.6) P<0.01 V 2 (0.7) 12 (5.3) P<0.01 P 21 (7.6) 10 (4.4) I 1 (0.4) 0 (0.0) H 0 (0.0) 2 (0.9) S (21.5) 36 (16.0) N 41 (14.9) 21 (9.3) SS 0 (0.0) 1 (0.4) E 0 (0.0) 1 (0.4) G/R/T 16 (5.8) 12 (5.3) R452 2 (0.7) 8 (3.6) G 1 (0.4) 2 (0.9) L 1 (0.4) 0 (0.0) K/Q 0 (0.0) 2 (0.9) S 0 (0.0) 4 (1.8) P<0.05 P (10.5) 76 (33.8) L 19 (6.9) 53 (23.6) P<0.001 I 1 (0.4) 4 (1.8) T 9 (3.3) 10 (4.4) A 0 (0.0) 6 (2.7) P<0.01 S/V 0 (0.0) 5 (2.2) P<0.05 L449P+P453 L 14 (5.1) 7 (3.1) I/V/T 4 (0.0) 3 (0.9) P 4 (1.5) 0 (0.0) Mutations at the cleavage sites p7/p1 and p1/p6 gag in comparison to HxB2 are indicated in absolute number and percent for TN and TE viruses. Cleavage site (CS) mutations A431V, K436R, I437V, L449F/V, R452S and P453L/A accumulate significantly in therapy-experienced (TE) viruses. S451SS indicates an insertion at this position. TN, therapy niave. Correlation of CS and PR mutations TE viruses with therapy-associated CS mutations (A431V, K436R, I437V, L449F/V, R452S and P453A/L*) were analysed with regard to their PR Antiviral Therapy 11:7 881
4 J Verheyen et al. mutations. A431V was significantly associated with the presence of PR mutations L24I (P<0,001), M46I/L (P<0.0001), I54V (P<0.001) and V82A (P<0.001). I437V was also linked with I54V (P<0.05) and mutations at position V82 (without V82A) (P<0.001). Different associations could be distinguished at gag position L449. Whereas L449F was significantly associated with PR mutations D30N (P<0.05), I84V (P<0.01) and N88D/S/T P<0.05), L449V was correlated with PR mutations at position I54 (P<0.01). In the case of L449P mutations, which were detected in similar numbers in TN and TE viruses, even an association with a less severe resistance profile could be notified. Viruses with P453L without an additional L449P were positively correlated with the occurrence of PR mutations D30N (P=0.069) and I84V (P<0.0001). They were also significantly less often detected in viruses harbouring the PR mutation V82A (P<0.05). No significant differences or tendencies could be found for viruses with K436R and P453A mutations, especially for the latter due to the low number of occurrence (Table 2). Moreover notable was that viruses with R452S mutations (n=4) simultaneously had PR mutation D30N (n=2) or PR mutations I54V, I84V and L90M (n=2). Correlation of CS and PR mutation patterns Sixty percent of all TE viruses had at least one therapy-associated CS mutation compared with 10% of TN viruses. If also the rare CS mutations (A431I, K436N/E, I437M, L449H, S451E, S451SS R452S/K/Q, P453I/T/S/V) were considered, 65,3% TE vs 12,4% TN viruses were positive for at least one CS mutation. Moreover TE viruses without any of these CS mutations (n=78) had a relatively low average sum of primary PR mutations (2.18 ±1.45); mainly TE viruses with only one single PR mutation had no CS mutations (37/78). Vice versa, therapyassociated CS mutations were quite frequent in TE viruses with more than one primary PR mutation. The percentage of TE viruses with one, two or more primary PR mutations, which were positive for at least one CS mutation, raised from 31.1% to 46.7% and 74.1%, respectively. More than one therapy-associated CS mutation at one cleavage site was only found in 10.2% of all TE viruses (p7/p1: 7.1%, p1/p6 gag: 3.1%). Particularly, the few viruses with two therapy-associated mutations at CS p1/p6 gag (n=7) had a surprisingly high amount of primary PR mutations (3.7 ±1.25). Two frequent but rarely combined (n=4) pairs of PR mutations were I54A V82A (n=51) and Table 2. CS and PR mutations in TE viruses All TE viruses A431V K436R I437V L449F L449V L449P P453A P453L* (n=225), (n=60), (n=21), (n=35), (n=17), (n=12), (n=10), (n=6), n=46 Mutation n (%) n (%) n (%) n (%) n (%) n (%) n (%) n (%) n (%) L24I 23 (10.1) 14 (23.3) 1 (4.8) 3 (8.6) 3 (17.6) 1 (8.3%) 0 (0.0) 0 (0.0) 2 (4.3) D30N 18 (7.9) 0 (0.0) 2 (9.5) 1 (2.9) 4 * (23.5) 0 (0.0%) 0 (0.0) 0 (0.0) 7 (15.2) V32I 21 (9.3) 2 (3.3) 4 (19.0) 1 (2.9) 2 (11.8) 1 (8.3) 0 (0.0) 1 (16.7) 1 (2.2) L33F 42 (18.7) 12 (20.0) 6 (28.6) 8 (22.9) 5 (29.4) 2 (16.7) 1 (10.0) 3 (50.0) 7 (15.2) M46I 72 (31.7) 27* (45.0) 6 (28.6) 13 (37.1) 7 (41.2) 3 (25.0) 2 (20.0) 4 (66.7) 20 (43.5) M46L 31 (13.7) 17 (28.3) 2 (9.5) 3 (8.6) 3 (17.6) 2 (16.7) 0 (0.0) 1 (16.7) 3 (6.5) I47V 14 (6.2) 3 (5.0) 3 (14.3) 1 (2.9) 2 (11.8) 1 (8.3) 0 (0.0) 1 (16.7) 2 (4.3) G48V 13 (5.8) 3 (5.0) 2 (9.5) 1 (2.9) 1 (5.9) 1 (8.3) 1 (10.0) 0 (0.0) 1 (2.2) I50L/V 6 (2.7) 1 (1.7) 1 (4.8) 2 (5.7) 0 (0.0) 0 (0.0) 0 (0.0) 0 (0.0) 3 (6.5) I54V 78 (34.7) 31 (51.7) 8 (38.1) 19* (54.3) 6 (35.3) 5 (41.7) 2 (20.0) 2 (33.3) 17 (37.0) I54L/M/S/T/A 30 (13.9) 7 (11.7) 3 (14.3) 6 (17.1) 4 (17.6) 5* (41.7) 1 (0.0) 3 (50.0) 6 (13.0) V82A 83 (36.9) 34 (56.7) 10 (47.6) 14 (40.0) 5 (29.4) 5 (41.7) 2 (20.0) 2 (33.3) 9 (19.6) V82F/T/S 22 (9.8) 7 (11.7) 1 (0.0) 10 (28.6) 0 (0.0) 0 (0.0) 2 (20.0) 0 (0.0) 4 (8.7) I84V 49 (21.7) 17 (28.3) 2 (9.5) 12 (34.3) 8* (47.1) 3 (25.0) 2 (20.0) 3 (50.0) 26 (56.5) N88S/T 5 (2.2) 2 (3.3) 2 (9.5) 1 (2.9) 1 (5.9) 0 (0.0) 0 (0.0) 0 (0.0) 1 (2.2) L90M 147 (65.3) 35 (58.3) 12 (57.1) 23 (65.7) 12 (70.6) 8 (66.7) 4 (40.0) 6 (100.0) 35 (76.1) Average 2.83 ± ± ± ± ± ± ± ± ±1.52 primary PR mutations *P<0.05. P<0.01. CS, cleavage site; PR, protease; TE, therapy experienced International Medical Press
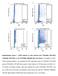
5 Compensatory cleavage site mutations in HIV Table 3. Correlation of CS and PR mutation patterns All TE V82A+ L90M+ viruses I54V I84V D30N (n=225) (n=51) (n=42) (n=18) Mutation n (%) n (%) n (%) n (%) A431V 60 (26.7) 23 (45.1) 13 (31.0) 0 (0.0) K436R 21 (9.3) 6 (11.8) 2 (4.8) 2 (11.1) I437V 35 (15.6) 11 (21.6) 10 (23.8) 1 (5.6) At least 100 (44.4) 33 (64.7) 21 (50.0) 3 (16.7) one p7/p1 CS mutation L449V 12 (5.3) 4 (7.8) 3 (7.1) 0 (0.0) L449F 17 (7.6) 4 (7.8) 8 (19.0) 4* (22.2) R452S 4 (1.8) 0 (0.0) 2 (4.8) 2 (11.1) P453L 46 (20.4) 7 (13.7) 24 (57.1) 7 (38.9) P453A 6 (2.7) 1 (2.0) 3 (7.1) 0 (0.0) At least one 78 (34.7) 15 (29.4) 35 (83.3) 13 (72.2) p1/p6 gag CS mutation At least one 135 (60.0) 35 (68.6) 37 (88.1) 13 (72.2) CS mutation at each CS *P<0.05. P<0.01. CS, cleavage site; PR, protease. Figure 1. Resistance pathways for cleavage site mutations A431V and 453L* and protease mutations at position 54 and 82 as well as L24I, M46L, I84V and L90M Arrows indicate order of appearance. Simultaneous evolution along different pathways is possible, but a mutation can only occur in a sample, if all its predecessors (as seen from the root) were also present. Numerical edge labels indicate conditional probability of appearance of a mutation, given that its predecessor is present. Fifty seven percent of samples could be explained by an ordered accumulation along two pathways (right), whereas 43% followed an unordered appearance or perhaps other pathways. L90M I84V (n=41) both involved in different resistance pathways. Interestingly, viruses with PR mutations I54A V82A had more often therapyassociated mutations at the CS p7/p1 (P<0.005) and viruses with the PR mutations L90M I84V at the CS p1/p6 gag (P<0.0001). CS mutations at this site were also independently associated with the detection of the PR mutation D30N (P<0.0001) (Table 3). To identify the association of CS mutations with known evolutionary pathways we analysed our crosssectional with the mtreemix package. Two major pathways were revealed; one consisting of PR mutations at positions I54 and V82, the other consisting of PR mutations L90M and I84V. P453L* was found to be part of the L90M I84V pathway, whereas mutation A431V was part of the I54 V82 pathway. Remarkably, our analysis showed that mutation A431V typically occurred before protease mutations L24I or M46L (Figure 1). Fifty-seven percent of our samples could be explained with this model, whereas the remaining 43% were attributed to spontaneous, unordered appearance of mutations, or other not yet considered resistance pathways in this model (D30N, I54V independently of V82 mutations). Combinations of p7/p1 and p1/p6 gag CS mutations Forty-three (19.1%) of TE viruses had at least one therapy-associated CS mutation at each site (p7/p1 and p1/p6 gag). Almost no TE viruses with one primary PR mutation (2/45) and only a few with two primary PR mutations (6/45) belonged to this group, but 13 of 36 TE viruses with 5 or more primary PR mutations had therapy-associated CS mutations at both CS. These viruses were associated with PR mutations L24I, M46I/L, 54 and 82. PR mutations I84V/L90M and D30N were only barely detected (n=3) in contrast to viruses with CS p1/p6 gag mutations alone (n=10). A follow-up of eight patients who underwent therapy interruption for different clinical reasons, demonstrated an outstanding maintenance of primary PR mutations during therapy-interruption. HIV genotypic resistance tests were performed before therapy interruption and compared with results obtained during therapy interruption. The median time in therapy interruption, before the second HIV genotype was determined, was 41 days ( days). Five of eight viruses showed no change in primary PR mutations within therapy-interruption Antiviral Therapy 11:7 883
6 J Verheyen et al. Table 4. Analysis of 8 patients before and within therapy interruption Patient G1-PR G1-CS Time in TI G2-PR G2-CS 1 10F, 41K, 46L, 54V, 71V, 72T, 431V, 451N, 453L 80 41K, 63P 431V, 451N, 453L 82A, 90M 2 10I, 32I, 33F, 46I, 47V, 54M, 63P, 431V, 449F/V, 453L 70 54M, 63P, 77I, 84V, 90M 431V, 449V, 453L 71L, 77I, 84V, 90M 3 10I, 33I, 54V, 63P, 71V, 84V, 90M 431V, 437V, 449F, 453A 41 10I, 33I, 54V, 63P, 71V, 431V, 449F, 453A 84V, 90M 4 10F, 20M, 33I, 43T, 47V, 53L, 54V, 437V, 453L 41 10F, 20M, 33I, 43T, 47V, 53L, 54V, 437V, 453L 63P, 71V, 73S, 74S, 82A, 90M 63P, 71V, 73S, 74S, 82A, 90M 5 10I, 20I, 36I, 46I, 54V, 63P, 71V, 431V, 453L 67 10I, 20I, 46I, 54V, 63P, 71V, 431V, 453L 84V, 90M 84V, 90M 6 10F, 46I, 63P, 84V, 90M 431V, 449F 43 10F, 46I, 63P, 84V, 90M 431V, 449F 7 63P, 71V*, 73S*, 84V, 90M 431V, 453L 30 63P, 77I, 84V, 90M 431V, 453L 8 10I, 33I, 36I, 46I, 54V, 63P, 71V, 431V, 449F, 453A I, 33I, 36I, 46I, 54V, 63P, 71V, 431V, 449F, 453A 84V, 90M 84V, 90M All patients were heavily pre-treated and harbouring multiple primary protease (PR) mutations (G1-PR). Additionally all viruses revealed mutations at both cleavage site (CS; p7/p1 and p1/p6 gag; G1-CS) simultaneously. The time in therapy interruption (TI) before the next genotype (G2-PR/CS) was in median 43d. In patient 7 71V* and 73* were simultaneously detected with a wt virus at these two positions before TI. after median 43 days ( days). Notably, the HIV of patient 5 had no CS mutations and only two primary PR mutations 2,8 years before time point 1. In patient 7 wild-type and mutant viruses were detected at positions 71 and 73 before therapy interruption, so that in therapy interruption no PR mutations were lost but one mutant virus ruled out the other mutant virus. In patient 1, wild-type HIV was detected after 80 days of therapy interruption without any primary PR mutation, whereas after 42 days of therapy interruption the mutant virus was still detected. Interestingly, mutations at both CS were still detectable at day 80 without PR mutations. A follow-up after the initiation of a new, PIcontaining, therapy was also available in this patient and revealed the reoccurrence of the former mutant virus (saquinavir and lopinavir). HIV of patient 2 lost three PR mutations M46I, I47V and L33F, similar to Patient 1, the CS mutations remained almost unchanged and with the selective pressure of a new, also PI containing, therapy regime (lopinavir and amprenavir), these three PR mutations re-occurred (Table 4). Discussion From the introduction of PI into the antiretroviral therapy, CS mutations associated with the exposure to these drugs have been reported from in vitro and in vivo studies [5,6,18,19]. In recent years the restoration of the otherwise compromized replication capacity of PR mutant viruses could be demonstrated for some CS mutations such as A431V, L449F and P453L [7 9,20]. Other CS mutations varied from study to study, mainly due to a great variation of analysed viruses and incomparable clinical settings, so that their impact on the evolution of PI resistance still remained uncertain. Some CS mutations associated with therapy exposure in our study (A431V, I437V, L449F/V and P453L) were also found in similar quantities in previous studies. K436R occurred significantly more often in TE viruses similar to Bally et al. [22], but was not associated with any PR mutation, meaning that the impact as a compensatory mutation remains to be determined in further studies. Some new mutations at CS p1/p6 gag (K452S, P453A) could complete the spectrum of therapy-associated CS mutations. The role and impact of the rare CS mutations (A431I, K436N/E, I437M, L449H, S451E, S451SS R452S/K/Q, P453I/T/S/V) has to be verified due to the low number of positive viruses in further studies and follow-ups. Notably, viruses with K436E and I437T CS mutations have been found in in vitro selection experiments without additional PR mutations and displayed 5 8-fold resistance to all clinical by approved PI, due to an increased p7/p1 processing [23]. An increased resistance to amprenavir has furthermore been described in viruses with CS mutations L449F and P453L compared with viruses with International Medical Press
7 Compensatory cleavage site mutations in HIV the same PR gene without CS mutations previously [24]. CS mutations can therefore not only compensate for the impaired fitness of viruses with primary PR mutations, but can also directly contribute to PI resistance. Three natural polymorphisms (I437L, R451N and L449P) that were not associated with PR mutations or therapy exposure were identified in our study. L449P has been discussed controversially as a natural polymorphism or as compensatory CS mutation selecting certain PR mutations. L449P as natural polymorphism is further supported by the fact that TE viruses with this mutation showed fewer primary PR mutations than the other TE viruses. As L449P is almost always accompanied by P453 mutations, these have to be separated from single P453 mutations and explain former descriptions of P453L as a natural polymorphism selecting for certain PR mutations [22]. The meaning of compensatory mutations in TN viruses remains unclear. In comparison to 2% of TN patients with HIV already harbouring primary PR mutations [14], the number of viruses with putative compensatory mutations is much higher (12.1%). Whether a resistant virus is transmitted in these patients but failed to form the dominant strain or is still resting in a cellular reservoir could not be verified yet. The follow-up of these patients is furthermore important as it could be speculated that these viruses have a lower genetic barrier for the acquisition of certain PR mutations. Bally et al. [22] still found A431V and L449F exclusively in TE viruses, which is in contrast to the 0.4% and 1.4% of TN viruses in our study. Results from Gallego et al. [25] were difficult to compare since natural polymorphisms and compensatory mutations were not distinguished, but they seemed to reach similar quantities. We have shown that therapy-associated CS p7/p1 and p1/p6 gag are strongly associated with two different selected PR mutation profiles. On the one hand A431V and I437V could be grouped together according to selected PR mutations at positions 54 and 82, and on the other hand L449F, R452S and P453L were associated with PR mutations D30N and I84V, although L449F has been mentioned before in the context of different PR mutations [10,12]. The rare L449V correlated with PR mutations at position I54 (with and without additional mutations at position 82) could not be clearly assigned to one group. Some associations of CS mutations and PR mutations seen in our study have already been reported: A431V-V82/I54V/M46I/L [21,26], I437V V82 [26] and L449F/P453L I84V [10,12,22]. Newly revealed correlations such as A431V L24I, I437V I54V, L449V I54 and L449F/P453L D30N further support the interesting linkage of CS mutations to particular PR mutant profiles. Most CS p7/p1 and p1/p6 gag mutations were not first-line mutations during the evolution of PI resistance, since they were predominantly observed in viruses with two or more primary PR mutations. The PR mutation combinations and 84V 90M are associated with divergent resistance pathway. Whereas viruses with PR mutations at positions 54 and 82 have decreased susceptibility for lopinavir [27,28], the resistance factors against saquinavir are not altered at all [29]; the opposite is described for the combination of PR mutations L90M I84V, which contribute to intermediate resistance against saquinavir without influencing the susceptibility to lopinavir [29,30]. Interestingly, the two combinations were associated with different CS mutations (54 82: p7/p1 CS mutations and 84V 90M: p1/p6 gag CS mutations). Moreover, the cross-sectional data analysis integrated the two most prominent CS mutations (A431V and P453L*) in the different resistance pathways (90M 84V 453L* and V). For A431V, a further evolution to either M46L or L24I was estimated by the mutagenetic tree analysis, underlining the importance of A431V for the lopinavir resistance profile [27]. This is supported by the fact that A431V occurred in cell culture experiments after exposure to lopinavir [31]. Assuming that existing mutations in CS p1/p6 gag lower the genetic barrier to saquinavir resistance, one it should be realized that D30N, conferring only resistance to nelfinavir [29,30], was associated with p1/p6 gag mutations. This might imply that pre-treatment with nelfinavir reduces the genetic barrier to saquinavir resistance not only by selection for L90M, but also for D30N. Two important studies analysed the waning of PR mutations during therapy-interruption in HIV-infected patients. Deeks et al. [32] reported that the waning of PR mutations began in median after 6 weeks (interquartile range: 4 7 weeks) and was complete within 2 weeks (median). The time observed by Miller et al. [33] was even shorter. They found a complete wild-type PR in all patients after 30 days of therapy interruption. A431V as single CS mutation disappeared in these patients together with primary PR mutations. In comparison, our group of highly resistant viruses with at least one compensatory mutation at both CS revealed an outstanding ability to maintain PR mutations during therapy interruption. Although the limited number of patients in our study precludes general conclusions on viral evolution during therapy-interruptions, it is tempting to speculate that mutations at p7/p1 and p1/p6 gag CS favoured the maintenance of resistant viruses. Antiviral Therapy 11:7 885
8 J Verheyen et al. Altogether, our results underline the importance of therapy-associated mutations at CS p7/p1 and p1/p6 gag in the evolution of resistance to single or multiple PI, restoring the otherwise compromised viral fitness. It will be interesting to complete the knowledge about viral fitness by analysis of other CS (p17/p24, p24/p7 p2/p7) as well as other HIV proteins (p6). Acknowledgments This study was supported by the Koeln Fortune Program and a grant of the German Ministry of Health and Social Security (AZ /3). We thank Zebulon Tolman for the critical reading of our manuscript. References 1. Berkhout B. HIV-1 evolution under pressure of protease inhibitors: climbing the stairs of viral fitness. J Biomed Sci 1999; 6: Prabu-Jeyabalan M, Nalivaika EA, King NM, Schiffer CA. Structural basis for coevolution of a human immunodeficiency virus type 1 nucleocapsid-p1 cleavage site with a V82A drug-resistant mutation in viral protease. J Virol 2004; 78: Pettit SC, Henderson GJ, Schiffer CA, Swanstrom R. Replacement of the P1 amino acid of human immunodeficiency virus type 1 Gag processing sites can inhibit or enhance the rate of cleavage by the viral protease. J Virol 2002; 76: Shehu-Xhilaga M, Kraeusslich HG, Pettit S, et al. Proteolytic processing of the p2/nucleocapsid cleavage site is critical for human immunodeficiency virus type 1 RNA dimer maturation. J Virol 2001; 75: Doyon L, Croteau G, Thibeault D, Poulin F, Pilote L, Lamarre D. Second locus involved in human immunodeficiency virus type 1 resistance to protease inhibitors. J Virol 1996; 70: Zhang YM, Imamichi H, Imamichi T, et al. Drug resistance during indinavir therapy is caused by mutations in the protease gene and in its Gag substrate cleavage sites. J Virol 1997; 71: Bleiber G, Munoz M, Ciuffi A, Meylan P, Telenti A. Individual contributions of mutant protease and reverse transcriptase to viral infectivity, replication, and protein maturation of antiretroviral drug-resistant human immunodeficiency virus type 1. J Virol 2001; 75: Myint L, Matsuda M, Matsuda Z, et al. Gag noncleavage site mutations contribute to full recovery of viral fitness in protease inhibitor-resistant human immunodeficiency virus type 1. Antimicrob Agents Chemother 2004; 48: Robinson LH, Myers RE, Snowden BW, Tisdale M, Blair ED. HIV type 1 protease cleavage site mutations and viral fitness: implications for drug susceptibility phenotyping assays. AIDS Res Hum Retroviruses 2000; 16: Feher A, Weber IT, Bagossi P, et al. Effect of sequence polymorphism and drug resistance on two HIV-1 Gag processing sites. Eur J Biochem 2002; 269: Cote HC, Brumme ZL, Harrigan PR. Human immunodeficiency virus type 1 protease cleavage site mutations associated with protease inhibitor cross-resistance selected by indinavir, ritonavir, and/or saquinavir. J Virol 2001; 75: Prado JG, Wrin T, Beauchaine J, Ruiz L, Petropoulos CJ, Frost SD, Clotet B, D Aquila RT, Martinez-Picado J. Amprenavir-resistant HIV-1 exhibits lopinavir cross-resistance and reduced replication capacity. AIDS 2002; 16: Oette M, Haussinger D. [Drug resistance in antiretroviral therapy of HIV infection]. Med Klin (Munich) 2003; 98: Oette M, Kaiser R, Daumer M, et al. Primary drug-resistance in HIV-positive patients on initiation of first-line antiretroviral therapy in Germany. Eur J Med Res 2004; 9: Oette M, Kaiser R, Daumer M, et al. Primary HIV drug resistance and efficacy of first-line antiretroviral therapy guided by resistance testing. J Acquir Immune Defic Syndr 2006; 41: Beerenwinkel N, Daumer M, Sing T, Rahnenfuhrer J, Lengauer T, Selbig J, Hoffmann D, Kaiser R. Estimating HIV evolutionary pathways and the genetic barrier to drug resistance. J Infect Dis 2005; 191: Beerenwinkel N, Rahnenfuhrer J, Kaiser R, Hoffmann D, Selbig J, Lengauer T. Mtreemix: a software package for learning and using mixture models of mutagenetic trees. Bioinformatics 2005; 21: Mammano F, Petit C, Clavel F. Resistance-associated loss of viral fitness in human immunodeficiency virus type 1: phenotypic analysis of protease and gag coevolution in protease inhibitor-treated patients. J Virol 1998; 72: Zennou V, Mammano F, Paulous S, Mathez D, Clavel F. Loss of viral fitness associated with multiple Gag and Gag-Pol processing defects in human immunodeficiency virus type 1 variants selected for resistance to protease inhibitors in vivo. J Virol 1998; 72: Maguire MF, Guinea R, Griffin P, Macmanus et al. Changes in human immunodeficiency virus type 1 Gag at positions L449 and P453 are linked to I50V protease mutants in vivo and cause reduction of sensitivity to amprenavir and improved viral fitness in vitro. J Virol 2002; 76: Lastere S, Dalban C, Collin G, et al. Impact of insertions in the HIV-1 p6 PTAPP region on the virological response to amprenavir. Antivir Ther 2004; 9: Bally F, Martinez R, Peters S, Sudre P, Telenti A. Polymorphism of HIV type 1 gag p7/p1 and p1/p6 cleavage sites: clinical significance and implications for resistance to protease inhibitors. AIDS Res Hum Retroviruses 2000; 16: Nijhuis M, van Maarseveen N, Schipper P, et al. Novel HIV gag based protease drug resistance mechanism caused by an increased processing of the NC/p1 cleavage site. Antiviral Therapy 2005; 10:S Maguire M, Shortino D, Klein A, et al. Emergence of resistance to protease inhibitor amprenavir in human immunodeficiency virus type 1-infected patients: selection of four alternative viral protease genotypes and influence of viral susceptibility to coadministered reverse transcriptase nucleoside inhibitors. Antimicrob Agents Chemother 2002; 46: Gallego O, de Mendoza C, Corral A, Soriano V. Changes in the human immunodeficiency virus p7-p1-p6 gag gene in drug-naive and pretreated patients. J Clin Microbiol 2003; 41: Ueda TM ML, Shiino T, Nishizawa M, Matsuda M, Sugiura, W. Analysis of interference and co evolution between protease inhibitor resistant mutations and gag mutations. Antivir Ther 2005; 10:S Parkin NT, Chappey C, Petropoulos CJ. Improving lopinavir genotype algorithm through phenotype correlations: novel mutation patterns and amprenavir cross-resistance. AIDS 2003; 17: Mo H, King MS, King K, Molla A, Brun S, Kempf DJ. Selection of resistance in protease inhibitor-experienced, human immunodeficiency virus type 1-infected subjects International Medical Press
9 Compensatory cleavage site mutations in HIV failing lopinavir- and ritonavir-based therapy: mutation patterns and baseline correlates. J Virol 2005; 79: Beerenwinkel N, Schmidt B, Walter H, et al. Diversity and complexity of HIV-1 drug resistance: a bioinformatics approach to predicting phenotype from genotype. Proc Natl Acad Sci USA 2002; 99: Kempf DJ, Isaacson JD, King MS, et al. Identification of genotypic changes in human immunodeficiency virus protease that correlate with reduced susceptibility to the protease inhibitor lopinavir among viral isolates from protease inhibitor-experienced patients. J Virol 2001; 75: Accepted for publication 22 June Carrillo A, Stewart KD, Sham HL, et al. In vitro selection and characterization of human immunodeficiency virus type 1 variants with increased resistance to ABT- 378, a novel protease inhibitor. J Virol 1998; 72: Deeks SG, Wrin T, Liegler T, et al. Virologic and immunologic consequences of discontinuing combination antiretroviral-drug therapy in HIV-infected patients with detectable viremia. N Engl J Med 2001; 344: Miller V, Sabin C, Hertogs K, et al. Virological and immunological effects of treatment interruptions in HIV-1 infected patients with treatment failure. Aids 2000; 14: Antiviral Therapy 11:7 887
10
ORIGINAL ARTICLE /j x. Brescia, Italy
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
Title. HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in
 Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
viruses ISSN
 Viruses 2010, 2, 1411-1426; doi:10.3390/v2071411 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Review Role of Gag in HIV Resistance to Protease Inhibitors François Clavel 1,2,3, * and
Viruses 2010, 2, 1411-1426; doi:10.3390/v2071411 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Review Role of Gag in HIV Resistance to Protease Inhibitors François Clavel 1,2,3, * and
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication
 Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
X/01/$ DOI: /JVI Copyright 2001, American Society for Microbiology. All Rights Reserved.
 JOURNAL OF VIROLOGY, Jan. 2001, p. 589 594 Vol. 75, No. 2 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.2.589 594.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
JOURNAL OF VIROLOGY, Jan. 2001, p. 589 594 Vol. 75, No. 2 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.2.589 594.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
HIV-1 Protease Correlated Cleavage Site Mutations. Enhance Inhibitor Resistance
 JVI Accepts, published online ahead of print on 12 August 2009 J. Virol. doi:10.1128/jvi.00628-09 Copyright 2009, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 12 August 2009 J. Virol. doi:10.1128/jvi.00628-09 Copyright 2009, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Int.J.Curr.Microbiol.App.Sci (2017) 6(5):
 International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 6 Number 5 (2017) pp. 1322-1330 Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2017.605.143
International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 6 Number 5 (2017) pp. 1322-1330 Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2017.605.143
Improving PI drug resistance scores. Jens Verheyen, MD Institute of Virology University Duisburg-Essen
 Improving PI drug resistance scores Jens Verheyen, MD Institute of Virology University Duisburg-Essen Overview Why can all PI drug resistance scores be improved? Do we still need to improve PI drug resistance
Improving PI drug resistance scores Jens Verheyen, MD Institute of Virology University Duisburg-Essen Overview Why can all PI drug resistance scores be improved? Do we still need to improve PI drug resistance
Case Study. Dr Sarah Sasson Immunopathology Registrar. HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital.
 Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
The Danish HIV Cohort Study, Copenhagen University Hospital, Rigshospitalet, Copenhagen, Denmark 4
 Antiviral Therapy 2009 14: 995 1000 (doi: 10.3851/IMP1412) Original article The incidence rate of HIV type-1 drug resistance in patients on antiretroviral therapy: a nationwide population-based Danish
Antiviral Therapy 2009 14: 995 1000 (doi: 10.3851/IMP1412) Original article The incidence rate of HIV type-1 drug resistance in patients on antiretroviral therapy: a nationwide population-based Danish
Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review
 pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
The E138A substitution in HIV-1 reverse transcriptase decreases in vitro. susceptibility to emtricitabine as indicated by competitive fitness assays
 AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
Resistance Workshop. 3rd European HIV Drug
 3rd European HIV Drug Resistance Workshop March 30-April 1 st, 2005 Christine Hughes, PharmD Clinical Associate Professor Faculty of Pharmacy & Pharmaceutical Sciences University of Alberta Tenofovir resistance
3rd European HIV Drug Resistance Workshop March 30-April 1 st, 2005 Christine Hughes, PharmD Clinical Associate Professor Faculty of Pharmacy & Pharmaceutical Sciences University of Alberta Tenofovir resistance
Because accurate and reproducible phenotypic susceptibility
 BRIEF REPORT: CLINICAL SCIENCE Comparison of the Precision and Sensitivity of the Antivirogram and PhenoSense HIV Drug Susceptibility Assays Jie Zhang, MS,* Soo-Yon Rhee, MS,* Jonathan Taylor, PhD, and
BRIEF REPORT: CLINICAL SCIENCE Comparison of the Precision and Sensitivity of the Antivirogram and PhenoSense HIV Drug Susceptibility Assays Jie Zhang, MS,* Soo-Yon Rhee, MS,* Jonathan Taylor, PhD, and
Clinical utility of NGS for the detection of HIV and HCV resistance
 18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
Scottish Medicines Consortium
 Scottish Medicines Consortium darunavir 300mg tablets (Prezista ) No. (378/07) Tibotec (a division of Janssen-Cilag Ltd) 4 May 2007 The Scottish Medicines Consortium has completed its assessment of the
Scottish Medicines Consortium darunavir 300mg tablets (Prezista ) No. (378/07) Tibotec (a division of Janssen-Cilag Ltd) 4 May 2007 The Scottish Medicines Consortium has completed its assessment of the
Improved prediction of response to antiretroviral combination therapy using the genetic barrier to drug resistance
 Antiviral Therapy 12:169 178 Improved prediction of response to antiretroviral combination therapy using the genetic barrier to drug resistance André Altmann 1 *, Niko Beerenwinkel 2, Tobias Sing 1, Igor
Antiviral Therapy 12:169 178 Improved prediction of response to antiretroviral combination therapy using the genetic barrier to drug resistance André Altmann 1 *, Niko Beerenwinkel 2, Tobias Sing 1, Igor
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
HIV-1 Subtypes: An Overview. Anna Maria Geretti Royal Free Hospital
 HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
Risk of HIV-1 low level viremia to treatment. Germany. Nadine Lübke Düsseldorf
 Risk of HIV-1 low level viremia to treatment failure in the AREVIR-RESINA cohort in Germany Nadine Lübke Düsseldorf FDA Approval of antiretroviral drugs 1980-84 1985-89 1990-94 1995-99 2000-04 2005-09
Risk of HIV-1 low level viremia to treatment failure in the AREVIR-RESINA cohort in Germany Nadine Lübke Düsseldorf FDA Approval of antiretroviral drugs 1980-84 1985-89 1990-94 1995-99 2000-04 2005-09
Antiviral Therapy 2011; 16: (doi: /IMP1812)
 Antiviral Therapy 2011; 16:725 732 (doi: 10.3851/IMP1812) Original article Single genome sequencing of HIV-1 gag and protease resistance mutations at virologic failure during the OK04 trial of simplified
Antiviral Therapy 2011; 16:725 732 (doi: 10.3851/IMP1812) Original article Single genome sequencing of HIV-1 gag and protease resistance mutations at virologic failure during the OK04 trial of simplified
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation
 Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
High Failure Rate of the ViroSeq HIV-1 Genotyping System for Drug Resistance Testing in Cameroon, a Country with Broad HIV-1 Genetic Diversity
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
History (August 2010) Therapy for Experienced Patients. History (September 2010) History (November 2010) 12/2/11
 (August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
(August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
Gag Non-Cleavage Site Mutations Contribute to Full Recovery of Viral Fitness in Protease Inhibitor-Resistant Human Immunodeficiency Virus Type 1
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 2004, p. 444 452 Vol. 48, No. 2 0066-4804/04/$08.00 0 DOI: 10.1128/AAC.48.2.444 452.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 2004, p. 444 452 Vol. 48, No. 2 0066-4804/04/$08.00 0 DOI: 10.1128/AAC.48.2.444 452.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
Milan, Italy. Received 15 March 2002; returned 22 July 2002; revised 12 September 2002; accepted 27 September 2002
 Journal of Antimicrobial Chemotherapy (2003) 51, 135 139 DOI: 10.1093/jac/dkg016 Comparison of levels of HIV-1 resistance to protease inhibitors by recombinant versus conventional virus phenotypic assay
Journal of Antimicrobial Chemotherapy (2003) 51, 135 139 DOI: 10.1093/jac/dkg016 Comparison of levels of HIV-1 resistance to protease inhibitors by recombinant versus conventional virus phenotypic assay
Interactive selective pressures of HLA-restricted immune responses and antiretroviral drugs on HIV-1
 Antiviral Therapy 10:551 555 Interactive selective pressures of HLA-restricted immune responses and antiretroviral drugs on HIV-1 Mina John, Corey B Moore, Ian R James and Simon A Mallal* Centre for Clinical
Antiviral Therapy 10:551 555 Interactive selective pressures of HLA-restricted immune responses and antiretroviral drugs on HIV-1 Mina John, Corey B Moore, Ian R James and Simon A Mallal* Centre for Clinical
ORIGINAL ARTICLE. Ó 2004 Copyright by the European Society of Clinical Microbiology and Infectious Diseases
 ORIGINAL ARTICLE Effect of polymorphisms on the replicative capacity of protease inhibitor-resistant HIV-1 variants under drug pressure C. Suñé 1, L. Brennan 1, D. R. Stover 2 and T. Klimkait 1,3 1 Basel
ORIGINAL ARTICLE Effect of polymorphisms on the replicative capacity of protease inhibitor-resistant HIV-1 variants under drug pressure C. Suñé 1, L. Brennan 1, D. R. Stover 2 and T. Klimkait 1,3 1 Basel
Introduction to the Impact of Resistance in Hepatitis C
 Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Monitoring for Drug Resistance by Genotyping. Urvi M Parikh, PhD MTN Virology Core Lab
 Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
Immune pressure analysis of protease and reverse transcriptase genes of primary HIV-1 subtype C isolates from South Africa
 African Journal of Biotechnology Vol. 10(24), pp. 4784-4793, 6 June, 2011 Available online at http://www.academicjournals.org/ajb DOI: 10.5897/AJB10.560 ISSN 1684 5315 2011 Academic Journals Full Length
African Journal of Biotechnology Vol. 10(24), pp. 4784-4793, 6 June, 2011 Available online at http://www.academicjournals.org/ajb DOI: 10.5897/AJB10.560 ISSN 1684 5315 2011 Academic Journals Full Length
Hepatitis C Resistance Associated Variants (RAVs)
 Hepatitis C Resistance Associated Variants (RAVs) Atif Zaman, MD MPH Oregon Health & Science University Professor of Medicine Division of Gastroenterology and Hepatology Nothing to disclose Disclosure
Hepatitis C Resistance Associated Variants (RAVs) Atif Zaman, MD MPH Oregon Health & Science University Professor of Medicine Division of Gastroenterology and Hepatology Nothing to disclose Disclosure
Deep-Sequencing of HIV-1
 Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Low genetic barrier to large increases in HIV-1 cross-resistance to protease inhibitors during salvage therapy
 Antiviral Therapy 11:143 154 Low genetic barrier to large increases in HIV-1 cross-resistance to protease inhibitors during salvage therapy Laurence Morand-Joubert 1,2, Charlotte Charpentier 3,4, Gwendoline
Antiviral Therapy 11:143 154 Low genetic barrier to large increases in HIV-1 cross-resistance to protease inhibitors during salvage therapy Laurence Morand-Joubert 1,2, Charlotte Charpentier 3,4, Gwendoline
Clinical Implications of Mutations at Reverse Transcriptase Codon 135 on Response to NNRTI-Based Therapy
 8 The Open Virology Journal, 2007, 1, 8-13 Clinical Implications of Mutations at Reverse Transcriptase Codon 135 on Response to NNRTI-Based Therapy Harout K. Tossonian 1, Jesse D. Raffa 2, Jason Grebely
8 The Open Virology Journal, 2007, 1, 8-13 Clinical Implications of Mutations at Reverse Transcriptase Codon 135 on Response to NNRTI-Based Therapy Harout K. Tossonian 1, Jesse D. Raffa 2, Jason Grebely
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E,
 Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
It takes more than just a single target
 It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
Micropathology Ltd. University of Warwick Science Park, Venture Centre, Sir William Lyons Road, Coventry CV4 7EZ
 www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
Introduction to HIV Drug Resistance. Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School
 Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY. 16th International WAVLD symposium, 10th OIE Seminar
 NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY S. Van Borm, I. Monne, D. King and T. Rosseel 16th International WAVLD symposium, 10th OIE Seminar 07.06.2013 Viral livestock
NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY S. Van Borm, I. Monne, D. King and T. Rosseel 16th International WAVLD symposium, 10th OIE Seminar 07.06.2013 Viral livestock
Special Contribution Questions to and Answers from the International AIDS Society USA Resistance Testing Guidelines Panel
 Special Contribution Questions to and Answers from the International AIDS Society USA Resistance Testing Guidelines Panel In 1996 the International AIDS Society USA convened an international panel of experts
Special Contribution Questions to and Answers from the International AIDS Society USA Resistance Testing Guidelines Panel In 1996 the International AIDS Society USA convened an international panel of experts
NIH Public Access Author Manuscript J Acquir Immune Defic Syndr. Author manuscript; available in PMC 2013 September 01.
 NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
JCM (Revised version, June 23 th 2011)
 JCM Accepts, published online ahead of print on 6 July 2011 J. Clin. Microbiol. doi:10.1128/jcm.00908-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
JCM Accepts, published online ahead of print on 6 July 2011 J. Clin. Microbiol. doi:10.1128/jcm.00908-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Microbiology Review Division of Antiviral Drug Products (HFD-530)
 Microbiology Review Division of Antiviral Drug Products (HFD-530) NDA#: 21-785 Serial #: 000 Reviewer: N. Battula Date submitted: June 17,2004 Date received: June 18,2004 Date assigned: July 01,2004 Date
Microbiology Review Division of Antiviral Drug Products (HFD-530) NDA#: 21-785 Serial #: 000 Reviewer: N. Battula Date submitted: June 17,2004 Date received: June 18,2004 Date assigned: July 01,2004 Date
Genotypic Testing for HIV-1 Drug Resistance ( ) Robert W. Shafer, M.D.
 1 Genotypic Testing for HIV-1 Drug Resistance (11-30-03) by Robert W. Shafer, M.D. Division of Infectious Diseases and Geographic Medicine, Stanford University, Stanford, CA, U.S. A. Correspondence: Robert
1 Genotypic Testing for HIV-1 Drug Resistance (11-30-03) by Robert W. Shafer, M.D. Division of Infectious Diseases and Geographic Medicine, Stanford University, Stanford, CA, U.S. A. Correspondence: Robert
Resistance to Integrase Strand Transfer Inhibitors
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
DIVISION OF ANTIVIRAL DRUG PRODUCTS (HFD-530) MICROBIOLOGY REVIEW NDA:
 Atazanavir (BMS-232632, ATV) Molecular Formula: C 38 H 52 N 6 O 7 H 2 SO 4 Molecular Weight: 704.9 Dosage Form(s): 100/150/200 mg capsule Route(s) of Administration: Oral Pharmacological Category: Indication(s):
Atazanavir (BMS-232632, ATV) Molecular Formula: C 38 H 52 N 6 O 7 H 2 SO 4 Molecular Weight: 704.9 Dosage Form(s): 100/150/200 mg capsule Route(s) of Administration: Oral Pharmacological Category: Indication(s):
MedChem 401~ Retroviridae. Retroviridae
 MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
Received 23 September 2004/Accepted 16 January 2005
 JOURNAL OF VIROLOGY, May 2005, p. 5907 5913 Vol. 79, No. 10 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.10.5907 5913.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Phenotypic
JOURNAL OF VIROLOGY, May 2005, p. 5907 5913 Vol. 79, No. 10 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.10.5907 5913.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Phenotypic
Global Pharmaceutical Research and Development, Abbott Laboratories, Abbott Park, Illinois 60064
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Jan. 2003, p. 350 359 Vol. 47, No. 1 0066-4804/03/$08.00 0 DOI: 10.1128/AAC.47.1.350 359.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Jan. 2003, p. 350 359 Vol. 47, No. 1 0066-4804/03/$08.00 0 DOI: 10.1128/AAC.47.1.350 359.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System
 Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System Joel O. Wertheim 1,2, Alexandra M. Oster 3, Jeffrey A. Johnson 3, William M. Switzer 3, Neeraja Saduvala
Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System Joel O. Wertheim 1,2, Alexandra M. Oster 3, Jeffrey A. Johnson 3, William M. Switzer 3, Neeraja Saduvala
The New England Journal of Medicine
 IROLOGIC ND IMMUNOLOGIC CONSEQUENCES OF DISCONTINUING COMBINTION NTIRETROIRL-DRUG THERPY IN HI-INFECTED PTIENTS WITH DETECTBLE IREMI STEEN G. DEEKS, M.D., TERRI WRIN, B.S., TERI LIEGLER, PH.D., REBECC
IROLOGIC ND IMMUNOLOGIC CONSEQUENCES OF DISCONTINUING COMBINTION NTIRETROIRL-DRUG THERPY IN HI-INFECTED PTIENTS WITH DETECTBLE IREMI STEEN G. DEEKS, M.D., TERRI WRIN, B.S., TERI LIEGLER, PH.D., REBECC
Clinical Applications of Resistance Stuart C. Ray, MD
 Clinical Applications of Resistance Stuart C. Ray, MD Professor of Medicine and Oncology Director, Infectious Diseases Fellowship Training Program Johns Hopkins University School of Medicine Disclosures
Clinical Applications of Resistance Stuart C. Ray, MD Professor of Medicine and Oncology Director, Infectious Diseases Fellowship Training Program Johns Hopkins University School of Medicine Disclosures
Scottish Medicines Consortium
 Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
I m B m. 1 f ub I B. D m B. f u. 1 f ua 1 D I A. I A m. D m A. f a. 1 f u. D m B ) D m A )(I m B. 1 f ua. 1 (I m A. log (I A. log f.
 Supplementary Material Appendix 1 Here we show that independent inhibition by a single drug of two distinct steps (A and ) in the viral life cycle results in a non-linear median effect dose-response curve
Supplementary Material Appendix 1 Here we show that independent inhibition by a single drug of two distinct steps (A and ) in the viral life cycle results in a non-linear median effect dose-response curve
Originally published as:
 Originally published as: Ratsch, B.A., Bock, C.-T. Viral evolution in chronic hepatitis B: A branched way to HBeAg seroconversion and disease progression? (2013) Gut, 62 (9), pp. 1242-1243. DOI: 10.1136/gutjnl-2012-303681
Originally published as: Ratsch, B.A., Bock, C.-T. Viral evolution in chronic hepatitis B: A branched way to HBeAg seroconversion and disease progression? (2013) Gut, 62 (9), pp. 1242-1243. DOI: 10.1136/gutjnl-2012-303681
Selection of High-Level Resistance to Human Immunodeficiency Virus Type 1 Protease Inhibitors
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 2003, p. 759 769 Vol. 47, No. 2 0066-4804/03/$08.00 0 DOI: 10.1128/AAC.47.2.759 769.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Feb. 2003, p. 759 769 Vol. 47, No. 2 0066-4804/03/$08.00 0 DOI: 10.1128/AAC.47.2.759 769.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
Characterization of Novel HIV Drug Resistance Mutations Using Clustering, Multidimensional Scaling and SVM-Based Feature Ranking
 Characterization of Novel HIV Drug Resistance Mutations Using Clustering, Multidimensional Scaling and SVM-Based Feature Ranking Tobias Sing 1, Valentina Svicher 2, Niko Beerenwinkel 3, Francesca Ceccherini-Silberstein
Characterization of Novel HIV Drug Resistance Mutations Using Clustering, Multidimensional Scaling and SVM-Based Feature Ranking Tobias Sing 1, Valentina Svicher 2, Niko Beerenwinkel 3, Francesca Ceccherini-Silberstein
HCV Resistance Associated variants: impact on chronic hepatitis C treatment
 HCV Resistance Associated variants: impact on chronic hepatitis C treatment Dr. Stéphane Chevaliez Associate Professor of Medicine at the University of Paris-Est. History of Resistance in HCV Concern Only
HCV Resistance Associated variants: impact on chronic hepatitis C treatment Dr. Stéphane Chevaliez Associate Professor of Medicine at the University of Paris-Est. History of Resistance in HCV Concern Only
Evolutionary pathways of transmitted drug-resistant HIV-1
 J Antimicrob Chemother 2011; 66: 1467 1480 doi:10.1093/jac/dkr157 Advance Access publication 18 April 2011 Evolutionary pathways of transmitted drug-resistant HIV-1 Marieke Pingen 1,2, Monique Nijhuis
J Antimicrob Chemother 2011; 66: 1467 1480 doi:10.1093/jac/dkr157 Advance Access publication 18 April 2011 Evolutionary pathways of transmitted drug-resistant HIV-1 Marieke Pingen 1,2, Monique Nijhuis
Second-Line Therapy NORTHWEST AIDS EDUCATION AND TRAINING CENTER
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
Changes in Human Immunodeficiency Virus Type 1 Populations after Treatment Interruption in Patients Failing Antiretroviral Therapy
 JOURNAL OF VIROLOGY, July 2001, p. 6410 6417 Vol. 75, No. 14 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.14.6410 6417.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Changes
JOURNAL OF VIROLOGY, July 2001, p. 6410 6417 Vol. 75, No. 14 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.14.6410 6417.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Changes
Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal Introduction
 Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
14 TH EUROPEAN HIV & HEPATITIS MEETING Abst#_O_06
 14 TH EUROPEAN HIV & HEPATITIS MEETING 2016 Abst#_O_06 Patients with pre-existent NRTI- and NNRTI-resistance have a higher risk to lose virological suppression under tenofovir/emtricitabine/rilpivirine
14 TH EUROPEAN HIV & HEPATITIS MEETING 2016 Abst#_O_06 Patients with pre-existent NRTI- and NNRTI-resistance have a higher risk to lose virological suppression under tenofovir/emtricitabine/rilpivirine
HIV/AIDS CID 2003:37 (1 July) 113
 MAJOR ARTICLE HIV/AIDS Antiretroviral Drug Resistance Testing in Adults Infected with Human Immunodeficiency Virus Type 1: 2003 Recommendations of an International AIDS Society USA Panel Martin S. Hirsch,
MAJOR ARTICLE HIV/AIDS Antiretroviral Drug Resistance Testing in Adults Infected with Human Immunodeficiency Virus Type 1: 2003 Recommendations of an International AIDS Society USA Panel Martin S. Hirsch,
Evaluation and Management of Virologic Failure
 National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
Drug-Selected Resistance Mutations and Non-B Subtypes in Antiretroviral-Naive Adults with Established Human Immunodeficiency Virus Infection
 MAJOR ARTICLE Drug-Selected Resistance Mutations and Non-B Subtypes in Antiretroviral-Naive Adults with Established Human Immunodeficiency Virus Infection George J. Hanna, 1,a Henri U. Balaguera, 2,a Kenneth
MAJOR ARTICLE Drug-Selected Resistance Mutations and Non-B Subtypes in Antiretroviral-Naive Adults with Established Human Immunodeficiency Virus Infection George J. Hanna, 1,a Henri U. Balaguera, 2,a Kenneth
Management of NRTI Resistance
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Management of NRTI Resistance David Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Management of NRTI Resistance David Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington
Diagnostic Methods of HBV and HDV infections
 Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
modified dye uptake assay including formazan test EC 90 not tested plaque reduction assay
 Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762)
 Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
From Mosquitos to Humans: Genetic evolution of Zika Virus
 Article: From Mosquitos to Humans: Genetic evolution of Zika Virus Renata Pellegrino, PhD Director, Sequencing lab Center for Applied Genomics The Children s Hospital of Philadelphia Journal Club Clinical
Article: From Mosquitos to Humans: Genetic evolution of Zika Virus Renata Pellegrino, PhD Director, Sequencing lab Center for Applied Genomics The Children s Hospital of Philadelphia Journal Club Clinical
HIV Drug Resistance: An Overview
 Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
Running title: Gag genetic determinants and virological response
 AAC Accepts, published online ahead of print on 3 May 2010 Antimicrob. Agents Chemother. doi:10.1128/aac.00194-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions.
AAC Accepts, published online ahead of print on 3 May 2010 Antimicrob. Agents Chemother. doi:10.1128/aac.00194-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions.
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
Genetic evolution of HIV in patients remaining on a stable HAART regimen despite insufficient viral suppression
 Scandinavian Journal of Infectious Diseases, 2005; 37: 890 /901 ORIGINAL ARTICLE Genetic evolution of HIV in patients remaining on a stable HAART regimen despite insufficient viral suppression THOMAS B.
Scandinavian Journal of Infectious Diseases, 2005; 37: 890 /901 ORIGINAL ARTICLE Genetic evolution of HIV in patients remaining on a stable HAART regimen despite insufficient viral suppression THOMAS B.
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
Antiviral Therapy 2011; 16: (doi: /IMP1851)
 Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
Mapping Evolutionary Pathways of HIV-1 Drug Resistance. Christopher Lee, UCLA Dept. of Chemistry & Biochemistry
 Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Lamei Chen, Alla Perlina and Christopher J. Lee /JVI
 Positive Selection Detection in 40,000 Human Immunodeficiency Virus (HIV) Type 1 Sequences Automatically Identifies Drug Resistance and Positive Fitness Mutations in HIV Protease and Reverse Transcriptase
Positive Selection Detection in 40,000 Human Immunodeficiency Virus (HIV) Type 1 Sequences Automatically Identifies Drug Resistance and Positive Fitness Mutations in HIV Protease and Reverse Transcriptase
Received 4 August 2005/Accepted 7 December 2005
 JOURNAL OF VIROLOGY, Mar. 2006, p. 2472 2482 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2472 2482.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Extensive Recombination
JOURNAL OF VIROLOGY, Mar. 2006, p. 2472 2482 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2472 2482.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Extensive Recombination
Evaluation of Dried Blood Spots (DBS) for Human Immunodeficiency Virus (HIV-1) Drug Resistance Testing
 Evaluation of Dried Blood Spots (DBS) for Human Immunodeficiency Virus (HIV-1) Drug Resistance Testing Dawit Assefa 2, Woldaregay E.Abegaz 3, Teferi Gedif 2, Belete Tegbaru 1, Dereje Teshome 1, Tesfaye
Evaluation of Dried Blood Spots (DBS) for Human Immunodeficiency Virus (HIV-1) Drug Resistance Testing Dawit Assefa 2, Woldaregay E.Abegaz 3, Teferi Gedif 2, Belete Tegbaru 1, Dereje Teshome 1, Tesfaye
Update on HIV Drug Resistance. Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
DATA SHEET. Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter calf thymus DNA.
 Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Rapid, phenotypic HIV-1 drug sensitivity assay for protease and reverse transcriptase inhibitors
 Journal of Clinical Virology 13 (1999) 71 80 Rapid, phenotypic HIV-1 drug sensitivity assay for protease and reverse transcriptase inhibitors Hauke Walter a, Barbara Schmidt a, Klaus Korn a, Anne-Mieke
Journal of Clinical Virology 13 (1999) 71 80 Rapid, phenotypic HIV-1 drug sensitivity assay for protease and reverse transcriptase inhibitors Hauke Walter a, Barbara Schmidt a, Klaus Korn a, Anne-Mieke
Antiretroviral Prophylaxis and HIV Drug Resistance. John Mellors University of Pittsburgh
 Antiretroviral Prophylaxis and HIV Drug Resistance John Mellors University of Pittsburgh MTN Annual 2008 Outline Two minutes on terminology Origins of HIV drug resistance Lessons learned from ART Do these
Antiretroviral Prophylaxis and HIV Drug Resistance John Mellors University of Pittsburgh MTN Annual 2008 Outline Two minutes on terminology Origins of HIV drug resistance Lessons learned from ART Do these
Hepatitis C Virus resistance screening and phenotypic NS3-PI-Resistance characterization. Arevir Meeting, 08./09.05.
 Hepatitis C Virus resistance screening and phenotypic NS3-PI-Resistance characterization Arevir Meeting, 08./09.05.2014 Daniel Rupp HCV prevalence 130-170 million patients world wide Ca. 75% of infections
Hepatitis C Virus resistance screening and phenotypic NS3-PI-Resistance characterization Arevir Meeting, 08./09.05.2014 Daniel Rupp HCV prevalence 130-170 million patients world wide Ca. 75% of infections
HIV replication and selection of resistance: basic principles
 HIV replication and selection of resistance: basic principles 26th International HIV Drug Resistance and Treatment Strategies Workshop Douglas Richman 6 November 2017 CLINICAL DATA DURING SIXTEEN WEEKS
HIV replication and selection of resistance: basic principles 26th International HIV Drug Resistance and Treatment Strategies Workshop Douglas Richman 6 November 2017 CLINICAL DATA DURING SIXTEEN WEEKS
The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide polymorphisms at thymidine analogue mutation sites
 Journal of Antimicrobial Chemotherapy Advance Access published June 7, 2013 J Antimicrob Chemother doi:10.1093/jac/dkt204 The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide
Journal of Antimicrobial Chemotherapy Advance Access published June 7, 2013 J Antimicrob Chemother doi:10.1093/jac/dkt204 The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide
BASIC AND TRANSLATIONAL SCIENCE
 BASIC AND TRANSLATIONAL SCIENCE Prediction of Virological Response and Assessment of Resistance Emergence to the HIV-1 Attachment Inhibitor BMS-626529 During 8-Day Monotherapy With Its Prodrug BMS-663068
BASIC AND TRANSLATIONAL SCIENCE Prediction of Virological Response and Assessment of Resistance Emergence to the HIV-1 Attachment Inhibitor BMS-626529 During 8-Day Monotherapy With Its Prodrug BMS-663068
The frequency of HIV-I drug resistance mutations among treatment-naïve individuals at a tertiary care centre in south India
 ORIGINAL RESEARCH ARTICLE The frequency of HIV-I drug resistance mutations among treatment-naïve individuals at a tertiary care centre in south India A J Kandathil MSc*, R Kannangai MD PhD*, O C Abraham
ORIGINAL RESEARCH ARTICLE The frequency of HIV-I drug resistance mutations among treatment-naïve individuals at a tertiary care centre in south India A J Kandathil MSc*, R Kannangai MD PhD*, O C Abraham
Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study
 Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study Prabhav Kalaghatgi Max Planck Institute for Informatics March 20th 2013 HIV epidemic (2009) Prabhav Kalaghatgi 2/18
Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study Prabhav Kalaghatgi Max Planck Institute for Informatics March 20th 2013 HIV epidemic (2009) Prabhav Kalaghatgi 2/18
Recognition of HIV-1 subtypes and antiretroviral drug resistance using weightless neural networks
 Recognition of HIV-1 subtypes and antiretroviral drug resistance using weightless neural networks Caio R. Souza 1, Flavio F. Nobre 1, Priscila V.M. Lima 2, Robson M. Silva 2, Rodrigo M. Brindeiro 3, Felipe
Recognition of HIV-1 subtypes and antiretroviral drug resistance using weightless neural networks Caio R. Souza 1, Flavio F. Nobre 1, Priscila V.M. Lima 2, Robson M. Silva 2, Rodrigo M. Brindeiro 3, Felipe
Update on HIV-1 Drug Resistance and Tropism Testing
 Update on HIV-1 Drug Resistance and Tropism Testing Daniel R. Kuritzkes, MD Section of Retroviral Therapeutics Brigham and Women s Hospital Harvard Medical School ACTHIV 2011: A State-of-the-Science Conference
Update on HIV-1 Drug Resistance and Tropism Testing Daniel R. Kuritzkes, MD Section of Retroviral Therapeutics Brigham and Women s Hospital Harvard Medical School ACTHIV 2011: A State-of-the-Science Conference
2 nd Line Treatment and Resistance. Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012
 2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
A Novel Duplex Real-Time Reverse-Transcription PCR Assay for the Detection of Influenza A and the Novel Influenza A(H1N1) Strain
 Viruses 2009, 1, 1204-1208; doi:10.3390/v1031204 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Article A Novel Duplex Real-Time Reverse-Transcription PCR Assay for the Detection of Influenza
Viruses 2009, 1, 1204-1208; doi:10.3390/v1031204 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Article A Novel Duplex Real-Time Reverse-Transcription PCR Assay for the Detection of Influenza
The legally binding text is the original French version TRANSPARENCY COMMITTEE OPINION. 14 December 2011
 The legally binding text is the original French version TRANSPARENCY COMMITTEE OPINION 14 December 2011 PREZISTA 400 mg, film-coated tablet B/60 (CIP code: 393 138-3) Applicant: JANSSEN-CILAG darunavir
The legally binding text is the original French version TRANSPARENCY COMMITTEE OPINION 14 December 2011 PREZISTA 400 mg, film-coated tablet B/60 (CIP code: 393 138-3) Applicant: JANSSEN-CILAG darunavir
Somnuek Sungkanuparph, M.D.
 HIV Drug Resistance Somnuek Sungkanuparph, M.D. Associate Professor Division of Infectious Diseases Department of Medicine Faculty of Medicine Ramathibodi Hospital Mahidol University Adjunct Professor
HIV Drug Resistance Somnuek Sungkanuparph, M.D. Associate Professor Division of Infectious Diseases Department of Medicine Faculty of Medicine Ramathibodi Hospital Mahidol University Adjunct Professor
Geno2pheno: estimating phenotypic drug resistance from HIV-1 genotypes
 3850 3855 Nucleic Acids Research, 2003, Vol. 31, No. 13 DOI: 10.1093/nar/gkg575 Geno2pheno: estimating phenotypic drug resistance from HIV-1 genotypes Niko Beerenwinkel*, Martin Däumer 1, Mark Oette 2,
3850 3855 Nucleic Acids Research, 2003, Vol. 31, No. 13 DOI: 10.1093/nar/gkg575 Geno2pheno: estimating phenotypic drug resistance from HIV-1 genotypes Niko Beerenwinkel*, Martin Däumer 1, Mark Oette 2,
