The Journal of Infectious Diseases BRIEF REPORT
|
|
|
- Eugene Nicholson
- 6 years ago
- Views:
Transcription
1 The Journal of Infectious Diseases BRIEF REPORT Prevalence of Drug-Resistant Minority Variants in Untreated HIV-1 Infected Individuals With and Those Without Transmitted Drug Resistance Detected by Sanger Sequencing Dana S. Clutter, 1,a Shuntai Zhou, 7,a Vici Varghese, 1 Soo-Yon Rhee, 1 Benjamin A. Pinsky, 1,2 W. Jeffrey Fessel, 4 Daniel B. Klein, 5 Ean Spielvogel, 7 Susan P. Holmes, 3 Leo B. Hurley, 6 Michael J. Silverberg, 6 Ronald Swanstrom, 7 and Robert W. Shafer 1 1 Division of Infectious Diseases and Geographic Medicine and 2 Department of Pathology, Stanford University School of Medicine, and 3 Department of Statistics, Stanford University, Stanford, 4 Department of Internal Medicine, San Francisco Medical Center, Kaiser Permanente Northern California, San Francisco, 5 Department of Infectious Diseases, San Leandro Medical Center, Kaiser Permanente Northern California, San Leandro, and 6 Division of Research, Kaiser Permanente Northern California, Oakland, California; 7 Department of Biochemistry and Biophysics, University of North Carolina Chapel Hill Minority variant human immunodeficiency virus type 1 (HIV- 1) nonnucleoside reverse transcriptase inhibitor (NNRTI) resistance mutations are associated with an increased risk of virological failure during treatment with NNRTI-containing regimens. To determine whether individuals to whom variants with isolated NNRTI-associated drug resistance were transmitted are at increased risk of virological failure during treatment with a non-nnrti containing regimen, we identified minority variant resistance mutations in 33 individuals with isolated NNRTI-associated transmitted drug resistance and 49 matched controls. We found similar proportions of overall and nucleoside reverse transcriptase inhibitor associated minority variant resistance mutations in both groups, suggesting that isolated NNRTI-associated transmitted drug resistance may not be a risk factor for virological failure during treatment with a non-nnrti containing regimen. Keywords. HIV-1; drug resistance; minority variant; next-generation sequencing; antiretroviral therapy. Standard genotypic resistance testing using Sanger sequencing is recommended upon diagnosis of human immunodeficiency virus type 1 (HIV-1) infection to identify individuals to whom drug-resistant HIV-1 was transmitted. Sanger sequencing typically detects drug resistance mutations (DRMs) present in at least 20% 30% of circulating plasma viruses [1]. However, Received 17 May 2017; editorial decision 13 July 2017; accepted 17 July 2017; published online July 25, Presented in part: IDWeek 2016, New Orleans, Louisiana, 29 October a D. S. C. and S. Z. contributed equally to this report. Correspondence: D. S. Clutter, MD, MS, Division of Infectious Diseases and Geographic Medicine, Stanford University School of Medicine, 300 Pasteur Dr, Rm L-134, Stanford, CA (dclutter@stanford.edu). The Journal of Infectious Diseases 2017;216: The Author Published by Oxford University Press for the Infectious Diseases Society of America. All rights reserved. For permissions, journals.permissions@oup.com. DOI: /infdis/jix338 DRMs present below this threshold, commonly referred to as minority variant DRMs, have been shown to increase the risk of virological failure during an initial nonnucleoside reverse transcriptase inhibitor (NNRTI) containing regimen [2]. The prevalence of transmitted drug resistance (TDR) in the United States ranges from 10% to 20%, with NNRTIs being the most commonly affected drug class [3, 4]. Nucleoside reverse transcriptase inhibitor (NRTI) associated and NNRTIassociated DRMs emerge commonly in patients with virological failure during an NNRTI-containing regimen. However, in the absence of selective antiretroviral pressure, transmitted NRTIassociated DRMs decay more rapidly than transmitted NNRTIassociated DRMs [5]. Therefore, we investigated whether individuals with NNRTI-associated TDR detected by Sanger sequencing are more likely than those without TDR detected by Sanger sequencing to harbor minority variant DRMs that would increase their risk of virological failure even during a non-nnrti containing regimen. To determine whether TDR is associated with an increased prevalence of minority variant DRMs, we performed Illumina next-generation sequencing of a portion of the reverse transcriptase coding region from plasma samples collected before antiretroviral therapy from individuals with isolated NNRTIassociated TDR and individuals without TDR. For accurate identification and quantification of minority variants, we used the Primer ID method, which corrects for polymerase chain reaction (PCR) bias and for PCR and sequencing errors and reveals the sampling depth of the viral population [6]. METHODS Study Population The study population included HIV-1 infected, antiretroviral-naive individuals in the Kaiser Permanente Medical Care Program Northern California (KPNC) undergoing Sanger sequencing at Stanford Health Care Diagnostic Virology Laboratory between April 2004 and September The TDR prevalence in this population during the study period was 12.5%. The TDR group comprised 33 individuals with isolated NNRTI-associated TDR, defined as having variants with 1 NNRTI-associated surveillance DRM but no NRTI-associated or protease inhibitor associated surveillance DRMs detected by Sanger sequencing [7]. The control group comprised 49 antiretroviral-naive individuals without any TDR by Sanger sequencing who were matched to the TDR group for HIV-1 RNA level, CD4 + T-cell count, sampling year, and antiretroviral regimen where possible. Additional inclusion criteria were a viral load of > copies/ml and an available cryopreserved sample for analysis by next-generation sequencing. The Stanford University and KPNC institutional review boards approved this study. BRIEF REPORT JID 2017:216 (1 August) 387
2 Sanger Sequencing RNA extracted from plasma was reverse transcribed to complementary DNA, using SuperScript One-Step reverse transcription PCR with Platinum Taq (ThermoFisher Scientific, Waltham MA). AmpliTaq DNA polymerase was used for nested PCR analysis. Bidirectional sequencing encompassing protease and the first 250 codons of reverse transcriptase was performed using Big Dye Terminator Cycle Sequencing with electrophoretic resolution of products on an ABI 3730 sequencer (ThermoFisher Scientific). The GenBank accession numbers for the Sanger sequences are in the Supplementary Materials. Primer ID Illumina Next-Generation Sequencing Primer ID Illumina/MiSeq sequencing was performed using a previously published protocol [6]. RNA was extracted from plasma, using the QIAamp viral RNA mini kit (Qiagen, Germantown, MD). Barcoded primers (Primer IDs) and Superscript III First Strand Synthesis System (Invitrogen, Carlsbad, CA) were used to generate complementary DNA (primer sequences are in the Supplementary Materials). The complementary DNA was purified to remove the unincorporated Primer ID oligos and then used as a template in 2 rounds of PCR, with MiSeq-indexed primers incorporated in the second round of PCR. Sequencing libraries were purified, quantified, and pooled for sequencing by Illumina MiSeq 300-base paired-end platform. The Illumina bcl2fastq pipeline (v.1.8.4) was used to for initial processing of sequence data. The Primer ID consensus pipeline was then used to create template consensus sequences for each starting viral RNA genome by pooling reads with identical Primer IDs, with the number of unique Primer IDs in a run defining the sampling depth of the viral RNA population [6] (available at: github.com/swanstromlab/pid). The HXB2 numbering for the sequenced regions was (reverse transcriptase codons ). To distinguish mutations present in the viral RNA from the low rate of residual error, a Poisson threshold set at 10 times the frequency of the residual error was determined for each sample. Additionally, all template consensus sequences containing stop codons were excluded [6]. Statistical Analysis We defined minority variant mutations as amino acid differences from the consensus B sequence that were detected by next-generation sequencing but not Sanger sequencing. We compared the prevalence and frequencies of minority variant DRMs between the TDR and control groups, using the Fisher exact test and Wilcoxon rank sum test, respectively. We defined potential APOBEC-associated DRMs as DRMs that could result from G A substitutions within the canonical APOBEC3F (GA AA) or APOBEC3G (GG AG) dinucleotide contexts: D67N, E138K, M184I, G190S/E, and M230I [8]. Human APOBEC3F/G enzymes are retroviral restriction factors that generate de novo viral mutations in a characteristic sequence context to impair viral fitness. We performed 2 analyses of clinical outcomes: (1) we compared the proportions of individuals with virological failure between the TDR and control groups, and (2) we compared the proportions of individuals with virological failure between individuals with and those without minority variant DRMs conferring reduced susceptibility to any antiretroviral in the regimen they received. Reduced susceptibility to an antiretroviral was defined as intermediate or high-level resistance according to the Stanford HIV Drug Resistance Database interpretation system (version 8.2) [9]. Virological failure was defined as a confirmed HIV RNA level of >75 copies/ml at week 48. The Fisher exact test was used for both comparisons. RESULTS Characteristics of the TDR and Control Groups Overall, 90% of study individuals were male, 55% were white, and the median age was 42 years. The median CD4 + T-cell count was 339 cells/mm 3, and the median viral load was 4.8 log copies/ ml. Fifty percent and 35% of patients initiated a regimen containing raltegravir or boosted atazanavir, respectively. Eightyeight percent of regimens included the NRTIs emtricitabine and tenofovir. The median year of pretherapy Sanger sequencing was Demographic and clinical characteristics were similar between the TDR and control groups, with the exception that more control individuals initiated a regimen not containing raltegravir or boosted atazanavir (Supplementary Table 1). Primer ID Illumina Sampling Depth The median number of template consensus sequences was 619 overall (interquartile range [IQR], ). It was higher in the control group (1045; IQR, ) than in the TDR group (445; IQR, ; P =.03, by the Wilcoxon rank sum test). The median lower limit of detection of a minority variant with 95% confidence for a sample was 0.5% overall (IQR,.2% 1.2%), 0.7% in the TDR group (IQR,.3% 1.8%), and 0.3% in the control group (IQR,.2%.8%; P =.03, by the Wilcoxon rank sum test). The lowest limit of detection was 0.04% in the TDR group and 0.03% in the control group. Distribution of Drug Resistance Mutations Overall, 110 DRMs were detected in 66 of 82 study individuals (80%). Of the 110 DRMs, 35 were NNRTI-associated DRMs detected by Sanger sequencing among the 33 individuals in the TDR group. Each of these mutations was also detected by next-generation sequencing. The additional 75 DRMs were minority variants, as they were detected only by next-generation sequencing. Overall, minority variant DRMs were detected in 49 study individuals (60%). The most-common minority variant DRMs were G190E (in 16 individuals), K219R (in 11), and K101E, M184I, and K219E (in 5 each; Table 1). Figure 1 shows that there were 2 main distributions of DRMs: those occurring at low frequencies detectable only by next-generation sequencing and those occurring at high frequencies that were detected 388 JID 2017:216 (1 August) BRIEF REPORT
3 Table 1. Minority Variant Drug Resistance Mutations (DRMs) in the Control Group and the Group With Transmitted Drug Resistance (TDR) 40 Variable Control Group (n = 49) TDR Group (n = 33) P a Minority variant DRM pattern, patients, no. (%) Any 30 (61) 19 (58).82 NRTI only 22 (45) 10 (30).25 NNRTI only 19 (39) 11 (33).65 NNRTI and NRTI 11 (22) 2 (6).06 Minority variant DRMs, median frequency, % (number of occurrences) Any 0.2 (48) 0.2 (27).82 NRTI Overall 0.2 (25) 0.2 (16).29 M41L (1) K65R (1) D67N b 0.2 (3) 0 K70E (1) K70R (1) V75M 0.1 (1) 0 M184I b 0.4 (5) 0 M184V 0.1 (1) 0.6 (1) L210W 11.4 (1) 0 T215I 0.2 (2) 0.5 (2) T215S 0.2 (3) 0 K219E 1.1 (1) 0.1 (4) K219N 0.7 (1) 0 K219Q 0.8 (1) 0 K219R 0.2 (6) 0.4 (5) NNRTI Overall 0.2 (23) 0.5 (11).10 K101E 0.2 (3) 5.7 (2) K103N 4.4 (1) 0.3 (1) K103S (4) Y181C 0.2 (3) 0.1 (1) Y181V 0.2 (1) 0 Y188H 0.1 (1) 0 G190E b 0.2 (13) 1.2 (3) P225H 1.5 (1) 0 Frequency is calculated as the percentage of an individual s virus population that bears a specific DRM. Abbreviations: NNRTI, nonnucleoside reverse transcriptase inhibitor; NRTI, nucleoside reverse transcriptase inhibitor. a Comparisons of the proportions of patients with a particular minority variant DRM pattern between the control and TDR groups were made using the Fisher exact test. Comparisons of the DRM frequencies among those individuals with a particular DRM in the control and TDR groups were made using the Wilcoxon rank sum test. b Mutations that could potentially arise from APOBEC-mediated G A editing [8]. by both sequencing methods. Among the 75 minority variant DRMs, 48 occurred at frequencies of 0.1% 0.3%, whereas 27 occurred at frequencies of 0.4% 14%. Comparison of Minority Variants in the TDR and Control Groups There was no difference in the proportion of individuals in the TDR (19 of 33 [58%]) and control (30 of 49 [61%]) groups with minority variant DRMs (P =.82; Table 1). NRTI-associated minority variant DRMs were present in 39% of individuals overall, including 10 (30%) in the TDR group and 22 (45%) in DRMs detected, no Frequency Sequencing Assay NGS Sanger + NGS Figure 1. Histogram of drug resistance mutation (DRM) frequency, by method of sequencing. The histogram plots the number of DRMs detected at each frequency (percentage of the total virus population). DRMs in red were detected only by next-generation sequencing (NGS), and those in blue were detected by both Sanger sequencing and NGS. the control group (P =.25). NNRTI-associated minority variant DRMs were present in 37% of individuals overall, including 11 (33%) in the TDR group and 19 (39%) in the control group (P =.65). Combined NRTI-associated plus NNRTI-associated minority variant DRMs were present in 16% of individuals overall, including 2 (6%) in the TDR group and 11 (22%) in the control group (P =.06). There was no difference in the median frequency of minority variant DRMs observed in the TDR and control groups (0.2% in both groups; P =.82). Of the 75 minority variant DRMs, 24 (32%) were potential APOBEC-associated DRMs, including G190E in 16 individuals, M184I in 5 individuals, and D67N in 3 individuals. There was no difference in the proportions of potential APOBEC-associated minority variant DRMs between the TDR and control groups. Virological Outcome There was no difference in the proportions of individuals with virological failure between the TDR and control groups (2 of 33 vs 5 of 49; P =.70). Poor medication adherence was documented in 4 of 7 individuals with virological failure. Two individuals in the control group and none in the TDR group underwent repeat Sanger sequencing. The DRM M184V was detected during virological failure in one of the control individuals who lacked baseline minority variant DRMs. Two of 33 individuals in the TDR group and 8 of 49 in the control group had minority variant DRMs associated with intermediate or high-level resistance to 1 antiretroviral in their treatment regimen (P =.30). One of the 8 patients in BRIEF REPORT JID 2017:216 (1 August) 389
4 the control group with a pretherapy minority variant DRM (M184I at a frequency of 0.1%) developed virological failure during treatment with a boosted atazanavir-based regimen. Repeat resistance testing was not performed, and the individual achieved resuppression following substitution of lopinavir for atazanavir. DISCUSSION We used Primer ID Illumina next-generation sequencing to quantify the number of minority variant DRMs in 33 antiretroviral-naive individuals with isolated NNRTI-associated TDR detected by Sanger sequencing and 49 matched controls without TDR. The overall proportion of individuals with minority variant DRMs was 60%. This proportion is higher than that reported in previous next-generation sequencing studies, which have reported detecting minority variant DRMs in 20% 35% of antiretroviral-naive individuals [10]. Previous next-generation sequencing studies, however, generally ignored minority variant DRMs detected at levels of <1.0% because of the high probability that, in the absence of a method such as Primer ID, variants detected at a lower frequency include errors introduced during PCR amplification and sequencing [11]. The minority variant DRMs we observed likely represent a combination of transmitted DRMs that faded to levels no longer detectable by Sanger sequencing and de novo mutations resulting from HIV-1 reverse transcriptase misincorporation or APOBEC-mediated editing. Indeed, most of the DRMs in this study had a frequency either close to 100% or close to 0%. This bimodal distribution is consistent with 2 sources of DRMs. The finding that nearly one third of the minority variant DRMs were potential APOBEC-associated DRMs suggests that many of these were de novo mutations. In particular, G190E, which was present in 16 individuals, is rarely detected by Sanger sequencing and is extremely unfit [12]. Previous noncomparative studies have shown that individuals with TDR detected by Sanger sequencing may have additional minority variant DRMs [13, 14]. In this study, however, we found that individuals with TDR were not more likely than those without TDR to have additional minority variant DRMs. A potential explanation for the similar prevalence of minority variant DRMs among those with and those without NNRTI-associated TDR is that minority variant DRMs resulting from de novo mutation are likely to be evenly distributed among individuals with and those without TDR, potentially obscuring differences arising from the fading of transmitted DRMs. The absence of an excess of minority variant DRMs among individuals with isolated NNRTI-associated TDR may help explain previous reports that antiretroviral-naive individuals with isolated NNRTI-associated TDR who receive a non-nnrti-containing regimen achieve similar rates of virological suppression as compared to individuals without TDR [15]. However, our study was underpowered to assess the clinical significance of minority variant DRMs because only 10 individuals had minority variant DRMs associated with reduced susceptibility to any of the antiretrovirals in their regimen. Supplementary Data Supplementary materials are available at The Journal of Infectious Diseases online. Consisting of data provided by the authors to benefit the reader, the posted materials are not copyedited and are the sole responsibility of the authors, so questions or comments should be addressed to the corresponding author. Notes Financial support. This work was supported by the National Institutes of Health (grants KL2 TR and UL1 TR [to D. S. C.] and R01 AI [to S.-Y. R., S. P. H., V. V., and R. W. S.], and R56 AI44667 [to R. S.]), the UNC Center for AIDS Research of the National Institutes of Health (grant AI50410), and the UNC Lineberger Comprehensive Cancer Center of the National Institutes of Health (grant P30 CA 16068). Potential conflicts of interest. D. S. C. has received research funding through the Bristol-Myers Squibb Virology Fellows Research Program. R. S. is a coinventor of Primer ID and will receive royalties from Cellular Research related to the patent/licensing of the technology. R. W. S. is a consultant for Abbott Diagnostics, ViiV Healthcare, and Qiagen Molecular Diagnostics and has received research funding from Gilead Sciences, Bristol-Myers Squibb, Merck, and Vela Diagnostics. W. J. F. and D. B. K. are employed by The Permanente Medical Group. M. J. S. has received research grants from Pfizer and Merck. All other authors report no potential conflicts of interest. All authors have submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest. Conflicts that the editors consider relevant to the content of the manuscript have been disclosed. References 1. Leitner T, Halapi E, Scarlatti G, et al. Analysis of heterogeneous viral populations by direct DNA sequencing. Biotechniques 1993; 15: Li JZ, Paredes R, Ribaudo HJ, et al. Low-frequency HIV-1 drug resistance mutations and risk of NNRTI-based antiretroviral treatment failure: a systematic review and pooled analysis. JAMA 2011; 305: Rhee SY, Blanco JL, Jordan MR, et al. Geographic and temporal trends in the molecular epidemiology and genetic mechanisms of transmitted HIV-1 drug resistance: an individual-patient- and sequence-level meta-analysis. PLoS Med 2015; 12:e JID 2017:216 (1 August) BRIEF REPORT
5 4. Buchacz K, Young B, Palella FJ Jr, et al. Trends in use of genotypic resistance testing and frequency of major drug resistance among antiretroviral-naive persons in the HIV Outpatient Study, J Antimicrob Chemother 2015; 70: Jain V, Sucupira MC, Bacchetti P, et al. Differential persistence of transmitted HIV-1 drug resistance mutation classes. J Infect Dis 2011; 203: Zhou S, Jones C, Mieczkowski P, Swanstrom R. Primer ID validates template sampling depth and greatly reduces the error rate of next-generation sequencing of HIV-1 genomic RNA populations. J Virol 2015; 89: Bennett DE, Camacho RJ, Otelea D, et al. Drug resistance mutations for surveillance of transmitted HIV-1 drug-resistance: 2009 update. PLoS One 2009; 4:e Rhee SY, Sankaran K, Varghese V, et al. HIV-1 Protease, Reverse Transcriptase, and Integrase Variation. J Virol 2016; 90: Tang MW, Liu TF, Shafer RW. The HIVdb system for HIV-1 genotypic resistance interpretation. Intervirology 2012; 55: Simen BB, Simons JF, Hullsiek KH, et al.; Terry Beirn Community Programs for Clinical Research on AIDS. Low-abundance drug-resistant viral variants in chronically HIV-infected, antiretroviral treatment-naive patients significantly impact treatment outcomes. J Infect Dis 2009; 199: Thys K, Verhasselt P, Reumers J, Verbist BM, Maes B, Aerssens J. Performance assessment of the Illumina massively parallel sequencing platform for deep sequencing analysis of viral minority variants. J Virol Methods 2015; 221: Huang W, Gamarnik A, Limoli K, Petropoulos CJ, Whitcomb JM. Amino acid substitutions at position 190 of human immunodeficiency virus type 1 reverse transcriptase increase susceptibility to delavirdine and impair virus replication. J Virol 2003; 77: Dauwe K, Staelens D, Vancoillie L, Mortier V, Verhofstede C. Deep sequencing of HIV-1 RNA and DNA in newly diagnosed patients with baseline drug resistance showed no indications for hidden resistance and is biased by strong interference of hypermutation. J Clin Microbiol 2016; 54: Toni TA, Asahchop EL, Moisi D, et al. Detection of human immunodeficiency virus (HIV) type 1 M184V and K103N minority variants in patients with primary HIV infection. Antimicrob Agents Chemother 2009; 53: Clutter DS, Fessel WJ, Rhee SY, et al. Response to therapy in antiretroviral therapy-naive patients with isolated nonnucleoside reverse transcriptase inhibitor-associated transmitted drug resistance. J Acquir Immune Defic Syndr (1999) 2016; 72: BRIEF REPORT JID 2017:216 (1 August) 391
Title: Response to Therapy in Antiretroviral Therapy-Naïve Patients with Isolated ACCEPTED
 JAIDS Journal of Acquired Immune Deficiency Syndromes Publish Ahead of Print DOI: 10.1097/QAI.0000000000000942 Title: Response to Therapy in Antiretroviral Therapy-Naïve Patients with Isolated Nonnucleoside
JAIDS Journal of Acquired Immune Deficiency Syndromes Publish Ahead of Print DOI: 10.1097/QAI.0000000000000942 Title: Response to Therapy in Antiretroviral Therapy-Naïve Patients with Isolated Nonnucleoside
Antiviral Therapy 2011; 16: (doi: /IMP1851)
 Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
NNRTI Resistance NORTHWEST AIDS EDUCATION AND TRAINING CENTER
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER NNRTI Resistance David H. Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington Last Updated:
NORTHWEST AIDS EDUCATION AND TRAINING CENTER NNRTI Resistance David H. Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington Last Updated:
Deep-Sequencing of HIV-1
 Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Management of NRTI Resistance
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Management of NRTI Resistance David Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Management of NRTI Resistance David Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington
Title. HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in
 Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
Clinical utility of NGS for the detection of HIV and HCV resistance
 18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
ICAAC/IDSA DC, Oct. 26, 2008
 Tenofovir (TDF)- or Abacavir (ABC)-selected Minority Subpopulations in Viremic Subjects Detected by Ultra-deep Sequencing R. T. D Aquila 1, E. Rouse 2, J. Horton 2, A. Kheshti 1,3, S. Raffanti 1,3, K.
Tenofovir (TDF)- or Abacavir (ABC)-selected Minority Subpopulations in Viremic Subjects Detected by Ultra-deep Sequencing R. T. D Aquila 1, E. Rouse 2, J. Horton 2, A. Kheshti 1,3, S. Raffanti 1,3, K.
Introduction to HIV Drug Resistance. Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School
 Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review
 pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
Low-frequency HIV-1 drug resistance mutations can be clinically significant but must be interpreted with caution
 J Antimicrob Chemother 2010; 65: 1322 1326 doi:10.1093/jac/dkq139 Advance Access publication 13 May 2010 Low-frequency HIV-1 drug resistance mutations can be clinically significant but must be interpreted
J Antimicrob Chemother 2010; 65: 1322 1326 doi:10.1093/jac/dkq139 Advance Access publication 13 May 2010 Low-frequency HIV-1 drug resistance mutations can be clinically significant but must be interpreted
Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosure The speaker serves as a consultant to, and has received
Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosure The speaker serves as a consultant to, and has received
Evaluation and Management of Virologic Failure
 National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
Resistance to Integrase Strand Transfer Inhibitors
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
The E138A substitution in HIV-1 reverse transcriptase decreases in vitro. susceptibility to emtricitabine as indicated by competitive fitness assays
 AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation
 Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
The new epidemic of drug resistant HIV-1
 The new epidemic of drug resistant HIV-1 Gillian Hunt Centre for HIV and STI National Institute for Communicable Diseases ICREID March 2018 Status of the global HIV epidemic (2016) WHO Global Summary on
The new epidemic of drug resistant HIV-1 Gillian Hunt Centre for HIV and STI National Institute for Communicable Diseases ICREID March 2018 Status of the global HIV epidemic (2016) WHO Global Summary on
HIV Drug Resistance among Adolescents and Young Adults Failing HIV Therapy in Zimbabwe
 HIV Drug Resistance among Adolescents and Young Adults Failing HIV Therapy in Zimbabwe V Kouamou 1, J Manasa 1, D Katzenstein 1, A McGregor 1, CE Ndhlovu 1 & AT Makadzange 1. 1 University of Zimbabwe Introduction
HIV Drug Resistance among Adolescents and Young Adults Failing HIV Therapy in Zimbabwe V Kouamou 1, J Manasa 1, D Katzenstein 1, A McGregor 1, CE Ndhlovu 1 & AT Makadzange 1. 1 University of Zimbabwe Introduction
Antiviral Therapy 2016; 21: (doi: /IMP2987)
 Antiviral Therapy 2016; 21:175 180 (doi: 10.3851/IMP2987) Case report Virological failure in two patients with HIV-1 RNA viral loads >1,000,000 copies/ml initiated on elvitegravir/cobicistat/emtricitabine/tenofovir
Antiviral Therapy 2016; 21:175 180 (doi: 10.3851/IMP2987) Case report Virological failure in two patients with HIV-1 RNA viral loads >1,000,000 copies/ml initiated on elvitegravir/cobicistat/emtricitabine/tenofovir
HIV Drug Resistance. Together, we can change the course of the HIV epidemic one woman at a time.
 HIV Drug Resistance Together, we can change the course of the HIV epidemic one woman at a time. #onewomanatatime #thewellproject What Is Resistance? HIV drugs are designed to keep the amount of HIV virus
HIV Drug Resistance Together, we can change the course of the HIV epidemic one woman at a time. #onewomanatatime #thewellproject What Is Resistance? HIV drugs are designed to keep the amount of HIV virus
Second-Line Therapy NORTHWEST AIDS EDUCATION AND TRAINING CENTER
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
Update on HIV-1 Drug Resistance and Tropism Testing
 Update on HIV-1 Drug Resistance and Tropism Testing Daniel R. Kuritzkes, MD Section of Retroviral Therapeutics Brigham and Women s Hospital Harvard Medical School ACTHIV 2011: A State-of-the-Science Conference
Update on HIV-1 Drug Resistance and Tropism Testing Daniel R. Kuritzkes, MD Section of Retroviral Therapeutics Brigham and Women s Hospital Harvard Medical School ACTHIV 2011: A State-of-the-Science Conference
New technologies reaching the clinic
 New technologies reaching the clinic Martin Däumer May 31, 2018 Deep-sequencing Standard Sanger-sequencing...PQIYMDDHTRE... Ultra-deep-sequencing...PQIYMDDHTRE......PQIYMDDHTRE......PQIYVDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE...
New technologies reaching the clinic Martin Däumer May 31, 2018 Deep-sequencing Standard Sanger-sequencing...PQIYMDDHTRE... Ultra-deep-sequencing...PQIYMDDHTRE......PQIYMDDHTRE......PQIYVDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE...
History (August 2010) Therapy for Experienced Patients. History (September 2010) History (November 2010) 12/2/11
 (August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
(August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E,
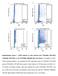 Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Introduction to the Impact of Resistance in Hepatitis C
 Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
14 TH EUROPEAN HIV & HEPATITIS MEETING Abst#_O_06
 14 TH EUROPEAN HIV & HEPATITIS MEETING 2016 Abst#_O_06 Patients with pre-existent NRTI- and NNRTI-resistance have a higher risk to lose virological suppression under tenofovir/emtricitabine/rilpivirine
14 TH EUROPEAN HIV & HEPATITIS MEETING 2016 Abst#_O_06 Patients with pre-existent NRTI- and NNRTI-resistance have a higher risk to lose virological suppression under tenofovir/emtricitabine/rilpivirine
Because accurate and reproducible phenotypic susceptibility
 BRIEF REPORT: CLINICAL SCIENCE Comparison of the Precision and Sensitivity of the Antivirogram and PhenoSense HIV Drug Susceptibility Assays Jie Zhang, MS,* Soo-Yon Rhee, MS,* Jonathan Taylor, PhD, and
BRIEF REPORT: CLINICAL SCIENCE Comparison of the Precision and Sensitivity of the Antivirogram and PhenoSense HIV Drug Susceptibility Assays Jie Zhang, MS,* Soo-Yon Rhee, MS,* Jonathan Taylor, PhD, and
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
Whole genome deep sequencing of HIV reveals extensive multi-class drug resistance in Nigerian patients failing first-line antiretroviral therapy
 Whole genome deep sequencing of HIV reveals extensive multi-class drug resistance in Nigerian patients failing first-line antiretroviral therapy K El Bouzidi 1,, RP Datir 1, V Kwaghe 3, S Roy 1, D Frampton
Whole genome deep sequencing of HIV reveals extensive multi-class drug resistance in Nigerian patients failing first-line antiretroviral therapy K El Bouzidi 1,, RP Datir 1, V Kwaghe 3, S Roy 1, D Frampton
Resistance Workshop. 3rd European HIV Drug
 3rd European HIV Drug Resistance Workshop March 30-April 1 st, 2005 Christine Hughes, PharmD Clinical Associate Professor Faculty of Pharmacy & Pharmaceutical Sciences University of Alberta Tenofovir resistance
3rd European HIV Drug Resistance Workshop March 30-April 1 st, 2005 Christine Hughes, PharmD Clinical Associate Professor Faculty of Pharmacy & Pharmaceutical Sciences University of Alberta Tenofovir resistance
Dr Carole Wallis, PhD Medical Director, BARC-SA Head of the Specialty Molecular Division, Lancet Laboratories, South Africa
 Dr Carole Wallis, PhD Medical Director, BARC-SA Head of the Specialty Molecular Division, Lancet Laboratories, South Africa Transmitted drug resistance Resistance patterns in first-line failures in adults
Dr Carole Wallis, PhD Medical Director, BARC-SA Head of the Specialty Molecular Division, Lancet Laboratories, South Africa Transmitted drug resistance Resistance patterns in first-line failures in adults
Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation
 Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation By: Blake M. Hauser Senior Honors Thesis Department of Biology College of Arts and Sciences The University of North Carolina
Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation By: Blake M. Hauser Senior Honors Thesis Department of Biology College of Arts and Sciences The University of North Carolina
The Danish HIV Cohort Study, Copenhagen University Hospital, Rigshospitalet, Copenhagen, Denmark 4
 Antiviral Therapy 2009 14: 995 1000 (doi: 10.3851/IMP1412) Original article The incidence rate of HIV type-1 drug resistance in patients on antiretroviral therapy: a nationwide population-based Danish
Antiviral Therapy 2009 14: 995 1000 (doi: 10.3851/IMP1412) Original article The incidence rate of HIV type-1 drug resistance in patients on antiretroviral therapy: a nationwide population-based Danish
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication
 Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
2 nd Line Treatment and Resistance. Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012
 2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING
 INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING Dr. Danni Kirwan ID/Microbiology SpR St. George s Hospital, London ARV initiation in treatment-naïve patients BHIVA,
INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING Dr. Danni Kirwan ID/Microbiology SpR St. George s Hospital, London ARV initiation in treatment-naïve patients BHIVA,
HIV DNA Genotyping by UDS compared with cumulative HIV RNA Genotypes in Pretreated Patients
 HIV DNA Genotyping by UDS compared with cumulative HIV RNA Genotypes in Pretreated Patients Mathieu BLOT, Yannis DUFFOURD, Hélène GIRAUDON, Emilie TISSERAND, Arnaud SALMON-ROUSSEAU, Sophie MAHY, Marielle
HIV DNA Genotyping by UDS compared with cumulative HIV RNA Genotypes in Pretreated Patients Mathieu BLOT, Yannis DUFFOURD, Hélène GIRAUDON, Emilie TISSERAND, Arnaud SALMON-ROUSSEAU, Sophie MAHY, Marielle
It takes more than just a single target
 It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
See Important Reminder at the end of this policy for important regulatory and legal information.
 Clinical Policy: Enfuvirtide (Fuzeon) Reference Number: CP.PHAR.41 Effective Date: 10.01.18 Last Review Date: 07.13.18 Line of Business: Oregon Health Plan Revision Log See Important Reminder at the end
Clinical Policy: Enfuvirtide (Fuzeon) Reference Number: CP.PHAR.41 Effective Date: 10.01.18 Last Review Date: 07.13.18 Line of Business: Oregon Health Plan Revision Log See Important Reminder at the end
Optimizing 2 nd and 3 rd Line Antiretroviral Therapy in Children and Adolescents
 Optimizing 2 nd and 3 rd Line Antiretroviral Therapy in Children and Adolescents Victor Musiime, MBChB, MMED, PhD Senior Lecturer, Makerere University Investigator, Joint Clinical Research Centre (JCRC)
Optimizing 2 nd and 3 rd Line Antiretroviral Therapy in Children and Adolescents Victor Musiime, MBChB, MMED, PhD Senior Lecturer, Makerere University Investigator, Joint Clinical Research Centre (JCRC)
Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal Introduction
 Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
HIV Drug Resistance: An Overview
 Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
HIV-1 resistance testing from proviral DNA
 HIV-1 resistance testing from proviral DNA Alexander Thielen AREVIR 2018 Resistance testing from proviral DNA? amount of samples with low viral loads increasing desire to switch under successful therapy
HIV-1 resistance testing from proviral DNA Alexander Thielen AREVIR 2018 Resistance testing from proviral DNA? amount of samples with low viral loads increasing desire to switch under successful therapy
DATA SHEET. Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter calf thymus DNA.
 Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Antiretroviral Treatment Strategies: Clinical Case Presentation
 Antiretroviral Treatment Strategies: Clinical Case Presentation Department of Internal Medicine, Far Eastern Memorial Hospital, New Taipei City, Taiwan Chia-Jui, Yang M.D Disclosure No conflicts of interests.
Antiretroviral Treatment Strategies: Clinical Case Presentation Department of Internal Medicine, Far Eastern Memorial Hospital, New Taipei City, Taiwan Chia-Jui, Yang M.D Disclosure No conflicts of interests.
Journal of Infectious Diseases Advance Access published April 18, Using Ultradeep Pyrosequencing to Study HIV 1 Co receptor Usage in Primary and
 Journal of Infectious Diseases Advance Access published April 18, 2013 1 Using Ultradeep Pyrosequencing to Study HIV 1 Co receptor Usage in Primary and Dual Infection Gabriel A. Wagner 1, Mary E. Pacold
Journal of Infectious Diseases Advance Access published April 18, 2013 1 Using Ultradeep Pyrosequencing to Study HIV 1 Co receptor Usage in Primary and Dual Infection Gabriel A. Wagner 1, Mary E. Pacold
Virological suppression and PIs. Diego Ripamonti Malattie Infettive - Bergamo
 Virological suppression and PIs Diego Ripamonti Malattie Infettive - Bergamo Ritonavir-boosted PIs Boosted PIs: 3 drugs in one The intrinsic antiretroviral activity Viral suppression and high baseline
Virological suppression and PIs Diego Ripamonti Malattie Infettive - Bergamo Ritonavir-boosted PIs Boosted PIs: 3 drugs in one The intrinsic antiretroviral activity Viral suppression and high baseline
Case Study. Dr Sarah Sasson Immunopathology Registrar. HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital.
 Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
Pornpimon Nimitsantiwong 1, Chorthip Wathitphan 1, Sukanta Kaveepatharanon 1, Kanoknun Thanomphakorn 1, Wasun Chantratita 1,2 and Ekawat Pasomsub 1
 Southeast Asian J Trop Med Public Health COMPARISON OF SENTOSA SQ DEEP SEQUENCING-BASED HIV-1 GENOTYPING COUPLED TO INTEGRATED WORKFLOW WITH SANGER SEQUENCING METHOD FOR DETECTION OF DRUG RESISTANCE MUTATIONS
Southeast Asian J Trop Med Public Health COMPARISON OF SENTOSA SQ DEEP SEQUENCING-BASED HIV-1 GENOTYPING COUPLED TO INTEGRATED WORKFLOW WITH SANGER SEQUENCING METHOD FOR DETECTION OF DRUG RESISTANCE MUTATIONS
HIV DRUG RESISTANCE IN AFRICA
 HIV DRUG RESISTANCE IN AFRICA Francis Ssali Joint Clinical Research Centre, Kampala Interest Meeting Mombasa May 10 th 2012 Scope 1. HIV-DR testing in Africa 2. The Epidemiology of HIV-DR in Africa 3.
HIV DRUG RESISTANCE IN AFRICA Francis Ssali Joint Clinical Research Centre, Kampala Interest Meeting Mombasa May 10 th 2012 Scope 1. HIV-DR testing in Africa 2. The Epidemiology of HIV-DR in Africa 3.
Antiviral Therapy 2014; 19: (doi: /IMP2748)
 Antiviral Therapy 214; 19:435 441 (doi: 1.3851/IMP2748) Short communication A decade of HIV-1 drug resistance in the United States: trends and characteristics in a large protease/reverse transcriptase
Antiviral Therapy 214; 19:435 441 (doi: 1.3851/IMP2748) Short communication A decade of HIV-1 drug resistance in the United States: trends and characteristics in a large protease/reverse transcriptase
DNA Genotyping in HIV Infection
 Frontier AIDS Education and Training Center DNA Genotyping in HIV Infection Steven C. Johnson M.D. Director, University of Colorado HIV/AIDS Clinical Program; Professor of Medicine, Division of Infectious
Frontier AIDS Education and Training Center DNA Genotyping in HIV Infection Steven C. Johnson M.D. Director, University of Colorado HIV/AIDS Clinical Program; Professor of Medicine, Division of Infectious
HIV and drug resistance Simon Collins UK-CAB 1 May 2009
 HIV and drug resistance Simon Collins UK-CAB 1 May 2009 slides: thanks to Prof Clive Loveday, Intl. Clinical Virology Centre www.icvc.org.uk Tip of the iceberg = HIV result, CD4, VL Introduction: resistance
HIV and drug resistance Simon Collins UK-CAB 1 May 2009 slides: thanks to Prof Clive Loveday, Intl. Clinical Virology Centre www.icvc.org.uk Tip of the iceberg = HIV result, CD4, VL Introduction: resistance
Anumber of clinical trials have demonstrated
 IMPROVING THE UTILITY OF PHENOTYPE RESISTANCE ASSAYS: NEW CUT-POINTS AND INTERPRETATION * Richard Haubrich, MD ABSTRACT The interpretation of a phenotype assay is determined by the cut-point, which defines
IMPROVING THE UTILITY OF PHENOTYPE RESISTANCE ASSAYS: NEW CUT-POINTS AND INTERPRETATION * Richard Haubrich, MD ABSTRACT The interpretation of a phenotype assay is determined by the cut-point, which defines
Dr Marta Boffito Chelsea and Westminster Hospital, London
 Dr Marta Boffito Chelsea and Westminster Hospital, London Speaker Name Statement Dr Marta Boffito has received travel and research grants from and has been an advisor for Janssen, Roche, Pfizer, ViiV,
Dr Marta Boffito Chelsea and Westminster Hospital, London Speaker Name Statement Dr Marta Boffito has received travel and research grants from and has been an advisor for Janssen, Roche, Pfizer, ViiV,
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2007 Paper 221 Biomarker Discovery Using Targeted Maximum Likelihood Estimation: Application to the
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2007 Paper 221 Biomarker Discovery Using Targeted Maximum Likelihood Estimation: Application to the
Dynamics of Compartmentalized HIV-1 Populations in the Central Nervous System Patrick R. Harrington
 Dynamics of Compartmentalized HIV-1 Populations in the Central Nervous System Patrick R. Harrington Postdoctoral Fellow Laboratory of Ronald Swanstrom Lineberger Comprehensive Cancer Center University
Dynamics of Compartmentalized HIV-1 Populations in the Central Nervous System Patrick R. Harrington Postdoctoral Fellow Laboratory of Ronald Swanstrom Lineberger Comprehensive Cancer Center University
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Determinants of residual viraemia during combination HIV treatment: Impacts of baseline HIV RNA levels and treatment choice
 DOI: 10.1111/hiv.12323 ORIGINAL RESEARCH Determinants of residual viraemia during combination HIV treatment: Impacts of baseline HIV RNA levels and treatment choice E McKinnon, 1, * A Castley, 2,3, * L
DOI: 10.1111/hiv.12323 ORIGINAL RESEARCH Determinants of residual viraemia during combination HIV treatment: Impacts of baseline HIV RNA levels and treatment choice E McKinnon, 1, * A Castley, 2,3, * L
Integrase Strand Transfer Inhibitors on the Horizon
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Integrase Strand Transfer Inhibitors on the Horizon David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, University of Washington Presentation
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Integrase Strand Transfer Inhibitors on the Horizon David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, University of Washington Presentation
HIV-1 Subtypes: An Overview. Anna Maria Geretti Royal Free Hospital
 HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES
 PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES Mark A. Wainberg McGill University AIDS Centre Jewish General Hospital Montreal, Quebec,
PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES Mark A. Wainberg McGill University AIDS Centre Jewish General Hospital Montreal, Quebec,
Management of patients with antiretroviral treatment failure: guidelines comparison
 The editorial staff Management of patients with antiretroviral treatment failure: guidelines comparison A change of therapy should be considered for patients if they experience sustained rebound in viral
The editorial staff Management of patients with antiretroviral treatment failure: guidelines comparison A change of therapy should be considered for patients if they experience sustained rebound in viral
ORIGINAL ARTICLE /j x. Brescia, Italy
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
Tools to Monitor HIV Infection in 2013 and Beyond.
 Tools to Monitor HIV Infection in 2013 and Beyond. Federico García, fegarcia@ugr.es Servicio de Microbiología Univ. Hospital San Cecilio Granada, Spain Outline Address clinical questions: Ultra sensitive
Tools to Monitor HIV Infection in 2013 and Beyond. Federico García, fegarcia@ugr.es Servicio de Microbiología Univ. Hospital San Cecilio Granada, Spain Outline Address clinical questions: Ultra sensitive
NIH Public Access Author Manuscript J Acquir Immune Defic Syndr. Author manuscript; available in PMC 2013 September 01.
 NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762)
 Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System
 Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System Joel O. Wertheim 1,2, Alexandra M. Oster 3, Jeffrey A. Johnson 3, William M. Switzer 3, Neeraja Saduvala
Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System Joel O. Wertheim 1,2, Alexandra M. Oster 3, Jeffrey A. Johnson 3, William M. Switzer 3, Neeraja Saduvala
Time taken to reach undetectable viral loads in therapy-naïve HIV patients commencing ART
 Time taken to reach undetectable viral loads in therapy-naïve HIV patients commencing ART Authors M A Ismail, E Okpo, S Baguley, A Butt, D Brawley, I Tonna 4 University of Aberdeen, MBChB Office, School
Time taken to reach undetectable viral loads in therapy-naïve HIV patients commencing ART Authors M A Ismail, E Okpo, S Baguley, A Butt, D Brawley, I Tonna 4 University of Aberdeen, MBChB Office, School
Clinical use of HIV-DNA quantity and resistance testing
 Clinical use of HIV-DNA quantity and resistance testing Personal fees and travel grants Abbott Molecular Gilead Sciences Janssen-Cilag Merck Sharp and Dohme ViiV Healthcare Research and educational grants
Clinical use of HIV-DNA quantity and resistance testing Personal fees and travel grants Abbott Molecular Gilead Sciences Janssen-Cilag Merck Sharp and Dohme ViiV Healthcare Research and educational grants
THE HIV LIFE CYCLE. Understanding How Antiretroviral Medications Work
 THE HIV LIFE CYCLE Understanding How Antiretroviral Medications Work DEFINITIONS Host: The animal or cell that another organism lives in. In HIV human CD4 T-cells are the host for HIV. Nucleus: The core
THE HIV LIFE CYCLE Understanding How Antiretroviral Medications Work DEFINITIONS Host: The animal or cell that another organism lives in. In HIV human CD4 T-cells are the host for HIV. Nucleus: The core
Dolutegravir-Rilpivirine (Juluca)
 Dolutegravir-Rilpivirine (Juluca) David H. Spach, MD Clinical Director, MW AETC Professor of Medicine Division of Infectious Diseases University of Washington Last Updated: November 30, 2017 ANTIRETROVIRAL
Dolutegravir-Rilpivirine (Juluca) David H. Spach, MD Clinical Director, MW AETC Professor of Medicine Division of Infectious Diseases University of Washington Last Updated: November 30, 2017 ANTIRETROVIRAL
Transmitted and Acquired HIV Drug Resistance in Latin America. Dr. Luis Enrique Soto Ramírez MEXICO
 Transmitted and Acquired HIV Drug Resistance in Latin America Dr. Luis Enrique Soto Ramírez MEXICO Disclosure Advisory boards and speaker for: Abbvie GSK/ViiV MSD Roche Diagnostics OVERVIEW The development
Transmitted and Acquired HIV Drug Resistance in Latin America Dr. Luis Enrique Soto Ramírez MEXICO Disclosure Advisory boards and speaker for: Abbvie GSK/ViiV MSD Roche Diagnostics OVERVIEW The development
Resistance Characteristics of Integrase Inhibitors
 Resistance Characteristics of Integrase Inhibitors Madrid, November 2016 Jonathan M Schapiro, MD National Hemophilia Center, Israel Stanford University School of Medicine, USA Disclaimer Presentation includes
Resistance Characteristics of Integrase Inhibitors Madrid, November 2016 Jonathan M Schapiro, MD National Hemophilia Center, Israel Stanford University School of Medicine, USA Disclaimer Presentation includes
Clinical skills building - HIV drug resistance
 Clinical skills building - HIV drug resistance Richard Lessells Clinical case 44-year old HIV-positive male HIV diagnosis 2010 Pre-treatment CD4+ count not known Initiated first-line ART (TDF/FTC/EFV)
Clinical skills building - HIV drug resistance Richard Lessells Clinical case 44-year old HIV-positive male HIV diagnosis 2010 Pre-treatment CD4+ count not known Initiated first-line ART (TDF/FTC/EFV)
What s New in Acute HIV Infection?
 3 4 Disclosure I have received research grants awarded to my institution from Gilead Sciences, Inc. ntiretroviral medications have been provided by Gilead Sciences, Inc. Susan Little, M.D. Professor of
3 4 Disclosure I have received research grants awarded to my institution from Gilead Sciences, Inc. ntiretroviral medications have been provided by Gilead Sciences, Inc. Susan Little, M.D. Professor of
Innovative diagnostics for HIV, HBV and HCV
 Innovative diagnostics for HIV, HBV and HCV Dan Otelea National Institute for Infectious Diseases Bucharest, Romania Disclaimer No conflicts of interest Innovative diagnostics for HIV, HBV and HCV - is
Innovative diagnostics for HIV, HBV and HCV Dan Otelea National Institute for Infectious Diseases Bucharest, Romania Disclaimer No conflicts of interest Innovative diagnostics for HIV, HBV and HCV - is
Antiviral Activity of Tenofovir Alafenamide against HIV-1 with Thymidine Analog Mutation(s) and M184V
 Antiviral Activity of Tenofovir Alafenamide against HIV-1 with Thymidine Analog Mutation(s) and M184V Christian Callebaut, PhD Gilead Sciences, Foster City, CA, USA HIV DART AND EMERGING VIRUSES 12/08/2016
Antiviral Activity of Tenofovir Alafenamide against HIV-1 with Thymidine Analog Mutation(s) and M184V Christian Callebaut, PhD Gilead Sciences, Foster City, CA, USA HIV DART AND EMERGING VIRUSES 12/08/2016
Somnuek Sungkanuparph, M.D.
 HIV Drug Resistance Somnuek Sungkanuparph, M.D. Associate Professor Division of Infectious Diseases Department of Medicine Faculty of Medicine Ramathibodi Hospital Mahidol University Adjunct Professor
HIV Drug Resistance Somnuek Sungkanuparph, M.D. Associate Professor Division of Infectious Diseases Department of Medicine Faculty of Medicine Ramathibodi Hospital Mahidol University Adjunct Professor
Scottish Medicines Consortium
 Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
HIV medications HIV medication and schedule plan
 Living with HIV (human immunodeficiency virus) It may be scary to find out that you re HIV-positive or have AIDS. Coping with this news may be difficult. Although HIV is a serious infection, people with
Living with HIV (human immunodeficiency virus) It may be scary to find out that you re HIV-positive or have AIDS. Coping with this news may be difficult. Although HIV is a serious infection, people with
Transmission of integrase resistance HIV
 Transmission of integrase resistance HIV Charles Boucher, MD, PhD Clinical Virology, Dept. Viroscience, Erasmus Medical Center, Erasmus Universiy, The Netherlands Major resistance mutations (Stanford)
Transmission of integrase resistance HIV Charles Boucher, MD, PhD Clinical Virology, Dept. Viroscience, Erasmus Medical Center, Erasmus Universiy, The Netherlands Major resistance mutations (Stanford)
HIV replication and selection of resistance: basic principles
 HIV replication and selection of resistance: basic principles 26th International HIV Drug Resistance and Treatment Strategies Workshop Douglas Richman 6 November 2017 CLINICAL DATA DURING SIXTEEN WEEKS
HIV replication and selection of resistance: basic principles 26th International HIV Drug Resistance and Treatment Strategies Workshop Douglas Richman 6 November 2017 CLINICAL DATA DURING SIXTEEN WEEKS
Clinical Implications of Mutations at Reverse Transcriptase Codon 135 on Response to NNRTI-Based Therapy
 8 The Open Virology Journal, 2007, 1, 8-13 Clinical Implications of Mutations at Reverse Transcriptase Codon 135 on Response to NNRTI-Based Therapy Harout K. Tossonian 1, Jesse D. Raffa 2, Jason Grebely
8 The Open Virology Journal, 2007, 1, 8-13 Clinical Implications of Mutations at Reverse Transcriptase Codon 135 on Response to NNRTI-Based Therapy Harout K. Tossonian 1, Jesse D. Raffa 2, Jason Grebely
MDR HIV and Total Therapeutic Failure. Douglas G. Fish, MD Albany Medical College Albany, New York Cali, Colombia March 30, 2007
 MDR HIV and Total Therapeutic Failure Douglas G. Fish, MD Albany Medical College Albany, New York Cali, Colombia March 30, 2007 Objectives Case study Definitions Fitness Pathogenesis of resistant virus
MDR HIV and Total Therapeutic Failure Douglas G. Fish, MD Albany Medical College Albany, New York Cali, Colombia March 30, 2007 Objectives Case study Definitions Fitness Pathogenesis of resistant virus
Clinical Significance of Human Immunodeficiency Virus Type 1 Replication Fitness
 CLINICAL MICROBIOLOGY REVIEWS, Oct. 2007, p. 550 578 Vol. 20, No. 4 0893-8512/07/$08.00 0 doi:10.1128/cmr.00017-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Clinical Significance
CLINICAL MICROBIOLOGY REVIEWS, Oct. 2007, p. 550 578 Vol. 20, No. 4 0893-8512/07/$08.00 0 doi:10.1128/cmr.00017-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Clinical Significance
I m B m. 1 f ub I B. D m B. f u. 1 f ua 1 D I A. I A m. D m A. f a. 1 f u. D m B ) D m A )(I m B. 1 f ua. 1 (I m A. log (I A. log f.
 Supplementary Material Appendix 1 Here we show that independent inhibition by a single drug of two distinct steps (A and ) in the viral life cycle results in a non-linear median effect dose-response curve
Supplementary Material Appendix 1 Here we show that independent inhibition by a single drug of two distinct steps (A and ) in the viral life cycle results in a non-linear median effect dose-response curve
High Failure Rate of the ViroSeq HIV-1 Genotyping System for Drug Resistance Testing in Cameroon, a Country with Broad HIV-1 Genetic Diversity
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
Purpose Methods Demographics of patients in the study Outcome. Efficacy Adverse Event. Limitation
 ANDREW LEE Purpose Methods Demographics of patients in the study Outcome Efficacy Adverse Event Limitation Dolutegravir Integrase inhibitor Plasma half life 14hours Tivicay FDA (US)- 13 August 2013 50mg
ANDREW LEE Purpose Methods Demographics of patients in the study Outcome Efficacy Adverse Event Limitation Dolutegravir Integrase inhibitor Plasma half life 14hours Tivicay FDA (US)- 13 August 2013 50mg
SINGLE. Efficacy and safety of dolutegravir (DTG) in treatment-naïve subjects
 SINGLE Efficacy and safety of dolutegravir (DTG) in treatment-naïve subjects SE/HIV/0023/14 January 2014 PHASE III DTG TRIALS IN TREATMENT-NAÏVE ADULT SUBJECTS WITH HIV SINGLE 1 N=833 Phase III non-inferiority,
SINGLE Efficacy and safety of dolutegravir (DTG) in treatment-naïve subjects SE/HIV/0023/14 January 2014 PHASE III DTG TRIALS IN TREATMENT-NAÏVE ADULT SUBJECTS WITH HIV SINGLE 1 N=833 Phase III non-inferiority,
Prevalence of Transmitted Drug Resistance Mutations among Naive HIV-infected patients ( ) in Northwest Spain
 Prevalence of Transmitted Drug Resistance Mutations among Naive HIV-infected patients (2014-2016) in Northwest Spain Berta Pernas 1, Andrés Tabernilla 1, Marta Grandal 1, Angelina Cañizares 2, Sofía Ortún
Prevalence of Transmitted Drug Resistance Mutations among Naive HIV-infected patients (2014-2016) in Northwest Spain Berta Pernas 1, Andrés Tabernilla 1, Marta Grandal 1, Angelina Cañizares 2, Sofía Ortún
Involvement of Novel Human Immunodeficiency Virus Type 1 Reverse Transcriptase Mutations in the Regulation of Resistance to Nucleoside Inhibitors
 JOURNAL OF VIROLOGY, July 2006, p. 7186 7198 Vol. 80, No. 14 0022-538X/06/$08.00 0 doi:10.1128/jvi.02084-05 Copyright 2006, American Society for Microbiology. All Rights Reserved. Involvement of Novel
JOURNAL OF VIROLOGY, July 2006, p. 7186 7198 Vol. 80, No. 14 0022-538X/06/$08.00 0 doi:10.1128/jvi.02084-05 Copyright 2006, American Society for Microbiology. All Rights Reserved. Involvement of Novel
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
HIV drug resistance and tracing outcomes among antiretroviral therapy defaulters in Malawi
 HIV drug resistance and tracing outcomes among antiretroviral therapy defaulters in Malawi Bello G 1, ParkinN 2, KagoliM 1, ChipetaS 1, CzaickiN 3, Pry J 3, OdenyT 3, NyasuluI 4, LapointeH 5, Doherty M
HIV drug resistance and tracing outcomes among antiretroviral therapy defaulters in Malawi Bello G 1, ParkinN 2, KagoliM 1, ChipetaS 1, CzaickiN 3, Pry J 3, OdenyT 3, NyasuluI 4, LapointeH 5, Doherty M
DOI: /hiv British HIV Association HIV Medicine (2014), 15, SHORT COMMUNICATION
 DOI: 10.1111/hiv.12071 SHORT COMMUNICATION Week 96 analysis of rilpivirine or efavirenz in HIV-1-infected patients with baseline viral load 100 000 copies/ml in the pooled ECHO and THRIVE phase 3, randomized,
DOI: 10.1111/hiv.12071 SHORT COMMUNICATION Week 96 analysis of rilpivirine or efavirenz in HIV-1-infected patients with baseline viral load 100 000 copies/ml in the pooled ECHO and THRIVE phase 3, randomized,
Update on HIV Drug Resistance. Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
The Journal of Infectious Diseases
 The Journal of Infectious Diseases BRIEF REPORT Selection of Rilpivirine-Resistant HIV- 1 in a Seroconverter From the SSAT 040 Trial Who Received the 300-mg Dose of Long-Acting Rilpivirine (TMC278LA) Kerri
The Journal of Infectious Diseases BRIEF REPORT Selection of Rilpivirine-Resistant HIV- 1 in a Seroconverter From the SSAT 040 Trial Who Received the 300-mg Dose of Long-Acting Rilpivirine (TMC278LA) Kerri
Ongoing HIV Replication During ART Reconsidered
 Open Forum Infectious Diseases PERSPECTIVES Ongoing HIV Replication During ART Reconsidered Mary F. Kearney, 1 Ann Wiegand, 1 Wei Shao, 2 William R. McManus, 1 Michael J. Bale, 1 Brian Luke, 2 Frank Maldarelli,
Open Forum Infectious Diseases PERSPECTIVES Ongoing HIV Replication During ART Reconsidered Mary F. Kearney, 1 Ann Wiegand, 1 Wei Shao, 2 William R. McManus, 1 Michael J. Bale, 1 Brian Luke, 2 Frank Maldarelli,
