Determination of HLA-A, -C, -B, -DRB1 allele and haplotype frequency in Japanese population based on family study
|
|
|
- Blaze Ray
- 5 years ago
- Views:
Transcription
1 Tissue Antigens ISSN Determination of HLA-A, -C, -B, -DRB1 allele and haplotype frequency in Japanese population based on family study N. Ikeda 1,H.Kojima 1,M.Nishikawa 1, K. Hayashi 1, T. Futagami 1, T. Tsujino 1, Y. Kusunoki 1, N. Fujii 1, S. Suegami 1, Y. Miyazaki 1, D. Middleton 2, H. Tanaka 1 &H.Saji 1 1 HLA Foundation Laboratory, Kyoto, Japan 2 Transplant Immunology, Royal Liverpool University Hospital Trust and University of Liverpool, Liverpool, UK Key words allele frequency; family study; haplotype frequency; human leucocyte antigen; Japanese population Correspondence Hiroh Saji HLA Foundation Laboratory 2F#1 Kyoto Research Park 134, Chudoji Minami-machi Kyoto Japan Tel: Fax: hla@hla.or.jp Received 18 August 2014; revised 14 January 2015; accepted 3 February 2015 Abstract The present study investigates the human leucocyte antigen (HLA) allele and haplotype frequencies in Japanese population. We carried out the frequency analysis in 5824 families living across Japanese archipelago. The studied population has mainly been typed for the purpose of transplant, especially the hematopoietic stem cell transplantation (HSCT). We determined HLA class I (A, B, and C) and HLA class II (DRB1) using Luminex technology. The haplotypes were directly counted by segregation. A total of 44 HLA-A, 29 HLA-C, 75 HLA-B, and 42 HLA-DRB1 alleles were identified. In the HLA haplotypes of A-C-B-DRB1 and C-B, the pattern of linkage disequilibrium peculiar to Japanese population has been confirmed. Moreover, the haplotype frequencies based on family study was compared with the frequencies estimated by maximum likelihood estimation (MLE), and the equivalent results were obtained. The allele and haplotype frequencies obtained in this study could be useful for anthropology, transplantation therapy, and disease association studies. doi: /tan Introduction The human leukocyte antigen (HLA) gene family is characterized by extreme degree of genetic polymorphism and linkage disequilibrium (LD). The varieties in polymorphism and LD patterns of HLA gene family show a tendency to be unique in each ethnic group (1, 2). HLA antigens have been known to play an important role in immune responses. In hematopoietic stem cell transplantation (HSCT), HLA matching between donors and recipients lowers the risk of graft rejection and graft-versus-host disease (GVHD) (3, 4). Morishima et al. suggested that the genetic difference derived from HLA haplotype is associated with acute GVHD in allogeneic HSCT (5). Therefore, HLA haplotype cannot be excluded from consideration during donor selection because of potential contribution from proteins encoded by non-hla genes inherited with HLA genes. Several studies reported analysis of HLA allele and haplotype frequency data in the Japanese population (6 9). However, these studies failed to present accurate and detailed information due to the haplotypes estimation using software or small sample size, or both. This necessitates developing a method which can produce accurate and detailed gene distribution. The present study aims to obtain a more exact and detailed HLA haplotype distribution from 18,604 members of 5824 Japanese families, whose HLA haplotypes were determined by descent. Our study also attempts to determine the frequency of specific haplotypes, C-B, A-B-DRB1, and A-C-B-DRB1, used in donor search. In addition, it was ascertained whether the haplotype frequencies estimated by maximum likelihood estimation (MLE) would be equivalent to the frequencies found in the present family study. Materials and method Subjects A total of 18,604 members (including patients and normal subjects) from 5824 families, distributed in all parts of Japan, were enrolled for this study. Among these families, there were patients, considered for transplantation, especially HSCT. The families were divided into three groups (Table 1): (i) families with both parents with one or more children, (ii) families with one parent with one or more children, and (iii) families with no parents but having two or more children. The families with more than two generations were counted as separate families. Informed consent was obtained from all the participants of this study by the clinicians who ordered HLA typing. For comparing the haplotype frequencies obtained by family study and using MLE software, unrelated 4500 people were John Wiley & Sons A/S. Published by John Wiley & Sons Ltd
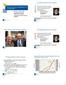
2 N. Ikeda et al. HLA allele and haplotype frequency in Japanese population Table 1 Breakdown of the family structure Number of children Number of families a A B Parents One parent No parent Total 5824 a The families with more than two generations are counted as separate families. a/b C c/d D E 2 b/d a/c e/f chosen at random from the total subjects of present study (18,604 members). They were genetically unrelated, because one person was chosen from each of 4500 random chosen families. The overlaps of blood relationship with three or more generation were avoided in these families. The allele and haplotype frequencies calculated from these 4500 people were very similar to the frequencies from the total subjects [allele frequencies (AF) data not shown]. Samples DNA samples were obtained from peripheral lymphocytes or buccal cells using a JetQuick Blood & Cell Culture Kits (GENOMED, Löhne, Germany) or QuickGene DNA Tissue Kit (KURABO, Osaka, Japan) according to the manufacturer s protocols. HLA allele typing HLA (-A, -C, -B, and -DRB1) four-digit allele typing was performed using Luminex 200 system (Luminex, Austin, TX) and WAKFlow HLA Typing kit (Wakunaga, Hiroshima, Japan) (10 12). HLA alleles were assigned automatically using WAKFLOW Typing software (Wakunaga, Hiroshima, Japan). The primer sequences of WAKFLOW Typing kit are specifically designed to make allele determination easier in Japanese population, and by default the analysis with WAKFLOW Typing software is based on the AF of the donors registered with Japan Marrow Donor Program (JMDP) which are available on the website, (9). Therefore, this method can determine alleles with frequencies of 0.1% and greater in the Japanese population. A few alleles which could not be determined by this method as rare alleles were considered as secondary; these alleles were determined using Luminex 200 system and LAB Type SSO kit (One Lambda, Los Angeles, CA) assigned using the HLA Fusion software (One Lambda). In brief, exon 2 for HLA-DRB1; exons 2 and 3 for HLA-A, -B, and -C were amplified in these methods. Haplotype determination The haplotypes were determined by segregation. This study was designed with the aim to assess genetic linkage with high certainty. Nevertheless, the results also included some partial haplotypes because of the possibility of one or more recombination F A, mother 1; B, father 1; C, child 1-1; D, child 1-2 D, mother 2; E, father 2; F, child 2-1; G, child 2-2; H, child 2-3 G H a/f a/e c/f 1, family 1 2, family 2 1+2, family 1+2 ex). Haplotype identification results are assumed as below. a. A * 24:02 B * 52:01 DRB1 * 15:02 A has a/b. D has a/c. b. A * 33:03 B * 44:03 DRB1 * 13:02 B has c/d. E has e/f. c. A * 24:02 B * 07:02 DRB1 * 01:01 C has b/d. F has a/f. d. A * 24:02 B * 54:01 DRB1 * 04:05 D has a/c. G has a/e. 08:03 H has c/f. e. A * 02:07 B * 46:01 DRB1 * f. A * 11:01 B * 15:01 DRB1 * 04:06 Figure 1 Avoiding haplotype duplication. The extended haplotypes of a and c are redundant in families 1 and 2. In this case, six haplotypes were counted as family in a family. The haplotype frequencies were calculated by using haplotypes without taking into account the recombination. Thus, we counted the haplotypes of parents and not of children because fathers and mothers are genetically unrelated, while the haplotypes created by recombination were those of children only. Some haplotypes of parents whose children had recombinant haplotypes could not been determined because of two patterns of their combinations. These haplotypes were determined and counted as not less frequent but frequent haplotype phase. Specifically, assuming that four haplotypes are Hp1, Hp2, Hp3, and Hp4, those frequencies are HF1, HF2, HF3, and HF4, and two estimable phases are Hp1, Hp2 and Hp3, Hp4, HF1 was multiplied by HF2, HF3 was multiplied by HF4, and the haplotypes of the estimable phase with larger product were counted. For the comparison to the result by MLE, the haplo.em program, which was evaluated by HAPLO.STATS (version 1.6.0) software operated in the R language, was used (13 16). For genetic markers measured on unrelated subjects with linkage phase unknown, this program computes the maximum likelihood estimates of haplotype probabilities using the progressive insertion algorithm that progressively inserts batches of loci into haplotypes of growing lengths John Wiley & Sons A/S. Published by John Wiley & Sons Ltd 253
3 HLA allele and haplotype frequency in Japanese population N. Ikeda et al. Table 2 HLA-A, -C, -B, -DRB1 allele frequencies in Japan a HLA-A serological HLA-A HLA-C serological HLA-C HLA-B serological HLA-B HLA-DRB1 serological HLADRB1 A1 A*01: Cw1 C*01: B7 B*07: DR1 DRB1*01: A2 A*02: C*01: B*07: DRB1*01: A*02: Cw2 C*02: B*07: DR4 DRB1*04: A*02: Cw4 C*04: B8 B*08: DRB1*04: A*02: Cw5 C*05: B13 B*13: DRB1*04: A*02: Cw6 C*06: B*13: DRB1*04: A*02: Cw7 C*07: B18 B*18: DRB1*04: A*02: C*07: B22 B*56: DRB1*04: A203 A*02: C*07: B*55: DRB1*04: A210 A*02: Cw8 C*08: B27 B*27: DRB1*04: A3 A*03: C*08: B*27: DRB1*04: A*03: C*08: B*27: DR6 DRB1*14: A11 A*11: Cw9 C*03: B35 B*35: DR7 DRB1*07: A*11: Cw10 C*03: B*35: DR8 DRB1*08: A*11: C*03: B*35: DRB1*08: A24 A*24: Cw12 C*12: B*35: DRB1*08: A*24: C*12: B37 B*37: DR9 DRB1*09: A*24: Cw14 C*14: B38 B*38: DR10 DRB1*10: A*24: C*14: B*38: DR11 DRB1*11: A*24: Cw15 C*15: B39 B*39: DRB1*11: A*24: C*15: B*39: DRB1*11: A*24: C*15: B*39: DR12 DRB1*12: A2403 A*24: Cw17 C*17: B3901 B*39: DRB1*12: A26 A*26: undefined C*03: B3902 B*39: DR13 DRB1*13: A*26: C*03: B41 B*41: DRB1*13: A*26: C*07: B44 B*44: DRB1*13: A*26: C*03: B*44: DR14 DRB1*14: A*26: C*04: B45 B*45: DRB1*14: A29 A*29: C*16: B46 B*46: DRB1*14: A30 A*30: B*46: DRB1*14: A*30: B48 B*48: DRB1*14: A31 A*31: B50 B*50: DRB1*14: A32 A*32: B51 B*51: DRB1*14: A33 A*33: B5102 B*51: DR1403 DRB1*14: A*33: B5103 B*51: DR1404 DRB1*14: A34 A*34: B52 B*52: DR15 DRB1*15: A66 A*66: B53 B*53: DRB1*15: A68 A*68: B54 B*54: DRB1*15: Null A*02:53N B55 B*55: DR16 DRB1*16: Undefined A*24: B*55: DR17 DRB1*03: A*11: B56 B*56: undefined DRB1*04: A*24: B57 B*57: DRB1*08: A*26: B58 B*58: A*31: B59 B*59: B60 B*40: B*40: B*40: B61 B*40: B*40: B*40: B*40: B*40: B*40: John Wiley & Sons A/S. Published by John Wiley & Sons Ltd
4 N. Ikeda et al. HLA allele and haplotype frequency in Japanese population Table 2 Continued HLA-A serological HLA-A HLA-C serological HLA-C HLA-B serological HLA-B HLA-DRB1 serological HLADRB1 B62 B*15: B*15: B*15: B*15: B*15: B*15: B*15: B64 B*14: B65 B*14: B67 B*67: B71 B*15: B75 B*15: B*15: B77 B*15: B78 B*78: B81 B*81: Unknown b B*54:21 b Null B*15:26N undefined B*35: B*51: B*52: B*52: GF, gene frequencies; HLA, human leucocyte antigen. a GF are calculated by using 19,183 haplotypes counted manually. Nomenclature of serological specificities, refer to Tissue Antigens 2010: 75: (15). b The allele is not in the paper (15). Statistical analysis The haplotypes were counted manually using Microsoft Excel spreadsheets. The haplotypes that extended three or more generations were counted once (Figure 1). The allele and haplotype frequencies were calculated by using 19,183 haplotypes, counted as mentioned above. Relative LD values (RD) were computed for each haplotypes (17, 18). The exact test for deviation from Hardy Weinberg Equilibrium were evaluated by GENEPOP software, version 4.2 (19, 20), which uses a Markov Chain (MC) algorithm (dememorization = 10,000, batches = 10,000, and iterations per batch = 10,000) to estimate the P-value. The expected prevalence (P) of the allele or the haplotype under Hardy Weinberg proportions were calculated from AF by using the following equation: P = 1 (1 AF) 2. in Table 2 need specific attention for HLA allele matching in unrelated HSCT between the Japanese, as they are present at high frequencies within a serotype (allele family) match. Haplotype segregation analysis based on family study Haplotype frequencies of HLA-A-C-B-DRB1 Table 3 lists 60 haplotypes with frequencies higher than 0.2% in the population. Approximately, 38% of the entire Japanese population is expected to carry one or two of the five most common haplotypes. These data have also been submitted to Allele Frequency Net Database (AFND) (22). The four-loci haplotypes with frequencies equal to or more than 0.01% and the AF can be found at the AFND website, (22). Results HLA-A, -B, -C, and -DRB1 AF Table 2 presents the list of the AF of HLA-A, -B, -C, and -DRB1 loci (21). We identified 44 HLA-A, 75 HLA-B, 29 HLA-C, and 42 HLA-DRB1 alleles and found A*24:02 to be 36.48%, the highest in Japanese population; thus, it is distributed in approximately 60% of the population. The alleles underlined HLA-A-B-DRB1 haplotype sets with the same serotypes Table 4 lists the sets of three-loci haplotypes at frequencies of 0.2% or greater, which would have the same serotype. Haplotype frequencies of HLA-C-B Table 5 lists 64 haplotypes with frequencies higher than 0.1 % in the population. Half of these haplotypes have RD values 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd 255
5 HLA allele and haplotype frequency in Japanese population N. Ikeda et al. Table 3 HLA A C B DRB1 haplotype frequencies in Japanese a A C B DRB1 HF (%) RD A C B DRB1 HF (%) RD A*24:02 C*12:02 B*52:01 DRB1*15: A*02:07 C*01:02 B*46:01 DRB1*09: A*33:03 C*14:03 B*44:03 DRB1*13: A*33:03 C*03:02 B*58:01 DRB1*13: A*24:02 C*07:02 B*07:02 DRB1*01: A*26:01 C*03:03 B*35:01 DRB1*04: A*24:02 C*01:02 B*54:01 DRB1*04: A*02:01 C*03:03 B*15:11 DRB1*09: A*02:07 C*01:02 B*46:01 DRB1*08: A*24:02 C*04:01 B*15:01 DRB1*04: A*11:01 C*04:01 B*15:01 DRB1*04: A*24:02 C*14:02 B*51:01 DRB1*14: A*24:02 C*01:02 B*59:01 DRB1*04: A*24:02 C*03:04 B*40:01 DRB1*04: A*11:01 C*01:02 B*54:01 DRB1*04: A*26:02 C*03:03 B*15:01 DRB1*14: A*26:01 C*03:04 B*40:02 DRB1*09: A*24:02 C*03:04 B*40:02 DRB1*04: A*24:02 C*08:01 B*40:06 DRB1*09: A*02:01 C*07:02 B*07:02 DRB1*01: A*24:02 C*14:02 B*51:01 DRB1*09: A*02:06 C*07:02 B*07:02 DRB1*01: A*31:01 C*14:02 B*51:01 DRB1*08: A*11:01 C*01:02 B*54:01 DRB1*08: A*33:03 C*14:03 B*44:03 DRB1*08: A*11:01 C*01:02 B*55:02 DRB1*04: A*26:02 C*08:01 B*40:06 DRB1*09: A*31:01 C*14:02 B*51:01 DRB1*04: A*02:01 C*03:04 B*13:01 DRB1*12: A*02:01 C*15:02 B*51:01 DRB1*15: A*24:02 C*01:02 B*46:01 DRB1*08: A*03:01 C*05:01 B*44:02 DRB1*13: A*02:06 C*08:01 B*40:06 DRB1*09: A*11:01 C*07:02 B*67:01 DRB1*15: A*11:01 C*07:02 B*39:01 DRB1*08: A*24:02 C*14:03 B*44:03 DRB1*13: A*26:01 C*03:04 B*40:02 DRB1*08: A*01:01 C*06:02 B*37:01 DRB1*10: A*02:06 C*03:03 B*35:01 DRB1*15: A*24:02 C*03:03 B*35:01 DRB1*15: A*24:02 C*12:02 B*52:01 DRB1*09: A*24:02 C*03:04 B*40:01 DRB1*11: A*31:01 C*14:02 B*51:01 DRB1*14: A*31:01 C*14:02 B*51:01 DRB1*09: A*02:06 C*07:02 B*39:01 DRB1*15: A*26:03 C*03:03 B*15:01 DRB1*09: A*24:02 C*03:04 B*40:02 DRB1*09: A*02:06 C*01:02 B*54:01 DRB1*04: A*02:01 C*01:02 B*54:01 DRB1*04: A*02:06 C*03:03 B*35:01 DRB1*09: A*26:03 C*03:03 B*15:01 DRB1*15: A*02:01 C*01:02 B*46:01 DRB1*08: A*11:01 C*07:02 B*67:01 DRB1*16: A*02:01 C*08:01 B*40:06 DRB1*09: A*02:06 C*01:02 B*59:01 DRB1*04: A*02:06 C*08:01 B*48:01 DRB1*04: A*24:02 C*03:03 B*15:07 DRB1*04: A*31:01 C*04:01 B*56:01 DRB1*09: A*24:02 C*07:04 B*15:18 DRB1*04: A*31:01 C*07:02 B*07:02 DRB1*01: HF, haplotype frequencies; HLA, human leucocyte antigen; RD, relative linkage disequilibrium value. a Four-locus haplotypes with HF >0.2% are listed. Table 4 Haplotype sets assigned same serotype of HLA A B DRB1 a Set no. Haplotype HF (%) 1 A*02:01-B*07:02-DRB1*01: A*02:06-B*07:02-DRB1*01: A*02:01-B*46:01-DRB1*08: A*02:07-B*46:01-DRB1*08: A*02:01-B*54:01-DRB1*04: A*02:06-B*54:01-DRB1*04: A*02:01-B*40:06-DRB1*09: A*02:06-B*40:06-DRB1*09: A*24:02-B*40:02-DRB1*09: A*24:02-B*40:06-DRB1*09: A*24:02-B*15:01-DRB1*04: A*24:02-B*15:07-DRB1*04: A*24:02-B*35:01-DRB1*04: A*24:02-B*35:01-DRB1*04: A*26:01-B*40:02-DRB1*09: A*26:01-B*40:06-DRB1*09: A*26:02-B*40:06-DRB1*09: HF, haplotype frequency; HLA, human leucocyte antigen. a The analysis objects are HF with more than 0.2%. more than 0.7, suggesting conservation of HLA-C-B linkage. B*40:02 and B*40:06 which correspond to the same serotype (B61) have high frequencies alleles of B61 and need attention for matching in HSCT. Focusing on these, Table 5 shows the frequency of C*03:04-B*40:02 as 6.26%, while the frequency of B*40:02 is 7.95% (Table 2); therefore, the frequency of C*03:04-B*40:02 linkage would account for 79% of B*40:02 alleles. Similarly, linkage of B*40:06 with C*08:01 would account for 81% of the presence of B*40:06. Thus, it is important to differentiate serotypes such as B61 when the constituting alleles are in linkage with different HLA-C alleles. Observed recombination A total observation number of recombination events were 136 in 134 families. These were divided into two groups: (i) the haplotypes of 103 parents (75.7%) could be determined, (ii) the haplotypes of 33 parents (24.3%) could not be determined and thus were inferred. Table 6 summarizes the observation number of recombination events in informative families which contain the parents and three or more children. The genotypes John Wiley & Sons A/S. Published by John Wiley & Sons Ltd
6 N. Ikeda et al. HLA allele and haplotype frequency in Japanese population Table 5 HLA C B haplotype frequencies in Japanese a C B HF (%) RD C B HF (%) RD C*12:02 B*52: C*08:01 B*15: C*01:02 B*54: C*03:04 B*51: C*14:02 B*51: C*04:01 B*35: C*14:03 B*44: C*04:01 B*40: C*03:04 B*40: C*03:04 B*40: C*03:03 B*35: C*05:01 B*44: C*07:02 B*07: C*03:04 B*35: C*01:02 B*46: C*03:04 B*40: C*08:01 B*40: C*15:02 B*40: C*07:02 B*39: C*01:03 B*46: C*03:03 B*15: C*07:02 B*39: C*03:04 B*40: C*15:02 B*40: C*04:01 B*15: C*06:02 B*13: C*01:02 B*55: C*07:02 B*38: C*01:02 B*59: C*03:04 B*15: C*08:01 B*48: C*03:03 B*55: C*15:02 B*51: C*03:03 B*48: C*07:02 B*67: C*01:02 B*56: C*03:04 B*13: C*03:03 B*40: C*08:01 B*35: C*08:03 B*54: C*03:03 B*40: C*01:02 B*51: C*07:02 B*40: C*07:02 B*39: C*08:03 B*48: C*01:02 B*56: C*07:04 B*15: C*15:02 B*51: C*03:03 B*15: C*01:02 B*40: C*15:02 B*40: C*12:02 B*27: C*01:02 B*15: C*15:02 B*15: C*03:03 B*15: C*08:03 B*15: C*03:02 B*58: C*03:03 B*55: C*04:01 B*56: C*07:02 B*40: C*07:02 B*15: C*04:01 B*15: C*06:02 B*37: C*04:01 B*48: HF, haplotype frequencies; HLA, human leucocyte antigen; RD, relative linkage disequilibrium value. a C-B haplotypes with HF >0.1% are listed. Table 6 Number of HLA A/C, C/B, B/DRB1 recombination evens by each family structure Number of children Number of families A/C a n (%R/T) B/DRB1 a n (%R/T) A/DRB1 a n (%R/T) Parents 5 5 b b b (0.89) 1 (0.45) 3 (1.34) (0.50) 8 (0.57) 15 (1.07) Total (0.54) 9 (0.54) 18 (1.08) HLA, human leucocyte antigen. a X/Y, number of recombination between X and Y; %R/T, % of recombination per transmission. b No observation of recombination. of these parents are heterozygote at all loci of HLA-A, -B, -C, and -DRB1. Group (ii) is not included in these informative families. The transmission of recombination (%R/T) shows the HLA-A-DRB1 recombination probability as 1.08% per child. Furthermore, Table 6 also indicates the recombination probabilities of HLA-A-C and B-DRB1 are 0.54%. Comparison to the result by MLE Table 7 shows the haplotype frequencies of HLA-A-C-B-DRB1 based on family study (result-fs) and based on MLE (result-mle) with frequencies more than 0.5%. In the frequent haplotypes with frequencies not less than 0.12%, result-mle tends to be higher than result-fs. In the low-frequent haplotypes with less than 0.12%, result-mle tends to be lower than result-fs. In addition, result-mle could not be detected in 585 haplotypes of the 2099 haplotypes with frequencies less than 0.12% in result-fs. On the allele data used for this comparison, four loci showed Hardy Weinberg Equilibrium: the P-values of the exact test at HLA-A, -B, -C, and DRB1 loci were , , , and , respectively. Discussion The AF presented in Table 2 show the high frequency of A*24:02 at 36.48%; therefore, around 60% of the population could be administered peptide vaccines such as WT1 peptide 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd 257
7 HLA allele and haplotype frequency in Japanese population N. Ikeda et al. Table 7 Comparison to maximum likelihood estimation (n = 4500) a A C B DRB1 FS (%) MLE (%) A*24:02-C*12:02-B*52:01-DRB1*15: % 8.298% A*33:03-C*14:03-B*44:03-DRB1*13: % 4.478% A*24:02-C*07:02-B*07:02-DRB1*01: % 3.857% A*24:02-C*01:02-B*54:01-DRB1*04: % 2.490% A*02:07-C*01:02-B*46:01-DRB1*08: % 2.053% A*11:01-C*04:01-B*15:01-DRB1*04: % 1.326% A*24:02-C*01:02-B*59:01-DRB1*04: % 1.167% A*11:01-C*01:02-B*54:01-DRB1*04: % 0.815% A*26:01-C*03:04-B*40:02-DRB1*09: % 0.892% A*24:02-C*08:01-B*40:06-DRB1*09: % 0.830% A*24:02-C*14:02-B*51:01-DRB1*09: % 0.803% A*31:01-C*14:02-B*51:01-DRB1*08: % 0.619% A*33:03-C*14:03-B*44:03-DRB1*08: % 0.589% A*26:02-C*08:01-B*40:06-DRB1*09: % 0.526% A*24:02-C*01:02-B*46:01-DRB1*08: % 0.504% A*02:01-C*03:04-B*13:01-DRB1*12: % 0.502% FS, haplotype frequencies based on family study; MLE, haplotype frequencies estimated by maximum likelihood estimation. a Four-locus haplotypes with frequencies >0.5% in result-fs are listed. vaccine presented by HLA molecules coded A*24:02 (23, 24). Moreover, around 80% of the population could be administered the vaccines (25, 26) presented by HLA molecules coded by one of three alleles as A*24:02 (36.5%), A*02:01 (11.6%), and A*02:06 (9.1%). The haplotype frequencies are also characteristic of Japanese population. The haplotypes of high frequencies are well conserved. Approximately one third (38%) of Japanese population carry the five most common haplotypes. We believe that this is strongly influenced by the founder effect: the ancestors having the common haplotypes migrated to Japan, thus, their haplotypes have been concentrated. While searching in the AFND (22), we found that the common haplotypes are homologous to those residing in the neighboring countries, especially in South Korea (11, 12, 27 31). In Korean population, the frequent haplotypes are similar to the ones in Japanese population. The three major Japanese haplotypes A*24:02-C*12:02-B*52:01-DRB1*15:02, A*33:03- C*14:03-B*44:03-DRB1*13:02, and A*24:02-C*07:02-B*07: 02-DRB1*01:01 have also high frequencies in Korea (1.9%, 4.2%, 2.9%, respectively) (27), suggesting the migration of some ancestors though the Korean Peninsula. The haplotype analysis not only helps in understanding the history of human migration but also in matching for unrelated donor searches for HSCT. In JMDP, the HLA compatibility with a donor is evaluated by both serotype and genotype. Furthermore, allele matching is a better evaluation of compatibility compared to the serotype matching; the rejection and GVHD risk of bone marrow transplant (BMT) have been found to be lower with allele-level matching compared to the serotype-level matching (3, 32). Accordingly, focusing on the alleles in every serotype, A2, A26, etc. increases the possibility of allele mismatch in spite of serotype match (Table 2). As shown in Table 4, HLA-A2, B61, B15, and DR4 especially increase the allele mismatch risk. However, compared with South Korean haplotype analysis, South Korea has 15 sets of haplotypes to pay attention to in HSCT matching (27) but Japan has eight sets (Table 4). It also shows that for most HLA haplotypes at the serotype-level, HLA haplotype matching is almost allele-level matching in HSCT between the Japanese. Although HLA-C locus is not indispensable for registering information in HSCT, HLA-C allele matching is important (32, 33). Even without information of donor HLA-C allele, the C-B haplotypes can predict HLA-C allele, although not always, because of well-conserved C-B linkage (Table 5); the conservation is possible due to short genetic linkage distance. In other words, HLA-B allele matching increases the possibility of HLA-C allele matching. Accordingly, analysis in the HLA distribution in Japanese population may contribute in planning the strategies of HLA matching for HSCT. Result-MLE was similar to result-fs (Table 7). Table 7 indicates the haplotype frequencies estimated by the software are very similar to real family-derived haplotypes. If the detection of the low frequency haplotypes is needed, determination of haplotypes by descent or a large sample size appears to be necessary. In conclusion, this study of determination of HLA allele and haplotype frequencies using family samples not only serves as a tool for elucidating linkage of each HLA locus but also acts as a tool in detecting HLA gene mutations in human germ cells such as recombination. The data obtained in this study will be useful in various fields such as anthropology, transplantation therapy, and disease association studies. Acknowledgments We would like to thank Jun Ohashi from Department of Biological Sciences, Graduate School of Science, The University of Tokyo, for valuable comments and suggestions about choosing unrelated samples and the exact test for deviation from Hardy Weinberg equilibrium. Conflicts of interest The authors have no conflicts of interest. References 1. Cao K, Hollenbach J, Shi X et al. Analysis of the frequencies of HLA-A, B, and C alleles and haplotypes in the five major ethnic groups of the United States reveals high levels of diversity in these loci and contrasting distribution patterns in these populations. Hum Immunol 2001: 62: Imanishi T, Wakisaka A, Gojobori T. Genetic relationships among various human populations indicated by MHC polymorphisms. In: Tsuji K, Aizawa M, Sasazuki T, eds. HLA Oxford: Oxford University Press, 1992, John Wiley & Sons A/S. Published by John Wiley & Sons Ltd
8 N. Ikeda et al. HLA allele and haplotype frequency in Japanese population 3. Lee SJ, Klein J, Haagenson M et al. High-resolution donor-recipient HLA matching contributes to the success of unrelated donor marrow transplantation. Blood 2007a: 110: Horan J, Wang T, Haagenson M et al. Evaluation of HLA matching in unrelated hematopoietic stem cell transplantation for nonmalignant disorders. Blood 2012: 120: Morishima S, Ogawa S, Matsubara A et al. Impact of highly conserved HLA haplotype on acute graft-versus-host disease. Blood 2010: 115: Nakaoka H, Mitsunaga S, Hosomichi K et al. Detection of ancestry informative HLA alleles confirms the admixed origins of Japanese population. PLoS One 2013: 8: e Tokunaga K, Ishikawa Y, Ogawa A et al. Sequence-based association analysis of HLA class I and II alleles in Japanese supports conservation of common haplotypes. Immunogenetics 1997: 46: Saito S, Ota S, Yamada E, Inoko H, Ota M. Allele frequencies and haplotypic associations defined by allelic DNA typing at HLA class I and class II loci in the Japanese population. Tissue Antigens 2000: 56: Moriyama Y, Kato K, Mura T, Juji T. Analysis of HLA gene frequencies and HLA haplotype frequencies for bone marrow donors in Japan. MHC 2006: 12: Ogata S, Shi L, Matsushita M et al. Polymorphisms of human leucocyte antigen genes in Maonan people in China. Tissue Antigens 2007: 69: Yao Y, Shi L, Shi L et al. Distribution of HLA-A, -B, -Cw, and -DRB1 alleles and haplotypes in an isolated Han population in Southwest China. Tissue Antigens 2009: 73: Shi L, Yao YF, Shi L et al. HLA alleles and haplotypes distribution in Dai population in Yunnan province, Southwest China. Tissue Antigens 2010: 75: Shichi D, Kikkawa EF, Ota M et al. The haplotype block, NFKBIL1-ATP6V1G2-BAT1-MICB-MICA, within the class III - class I boundary region of the human major histocompatibility complex may control susceptibility to hepatitis C virus-associated dilated cardiomyopathy. Tissue Antigens 2005: 66: Kamatani Y, Wattanapokayakit S, Ochi H et al. A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat Genet 2009: 41: Zhao H, Qi Y, Wang Y et al. Interactive contribution of serine/threonine kinase 39 gene multiple polymorphisms to hypertension among northeastern Han Chinese. Sci Rep 2014: 4: Wang ZT, Hu JJ, Fan R, Zhou J, Zhong J. RAGE gene three polymorphisms with Crohn s disease susceptibility in Chinese Han population. World J Gastroenterol 2014: 20: Lewontin RC. The Interaction of Selection and Linkage. I. General Considerations; Heterotic Models. Genetics 1964: 49: Hedrick PW. Gametic disequilibrium measures: proceed with caution. Genetics 1987: 117: Raymond M, Rousset F. GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Heredity 1995: 86: Rousset F. GENEPOP 007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol 2007: 8: Marsh SGE, Albert ED, Bodmer WF et al. Nomenclature for factors of the HLA system, Tissue Antigens 2010: 75: Gonzalez-Galarza FF, Christmas S, Middleton D, Jones AR. Allele frequency net: A database and online repository for immune gene frequencies in worldwide populations. Nucleic Acids Res 2011: 39: D Tsuboi A, Oka Y, Udaka K et al. Enhanced induction of human WT1-specific cytotoxic T lymphocytes with a 9-mer WT1 peptide modified at HLA-A*2402-binding residues. Cancer Immunol Immunother 2002: 51: Oka Y, Tsuboi A, Taguchi T et al. Induction of WT1 (Wilms tumor gene)-specific cytotoxic T lymphocytes by WT1 peptide vaccine and the resultant cancer regression. Proc Natl Acad Sci USA2004: 101: Kaida M, Morita-Hoshi Y, Soeda A et al. Phase 1 trial of Wilms tumor 1 (WT1) peptide vaccine and gemcitabine combination therapy in patients with advanced pancreatic or biliary tract cancer. J Immunother 2011: 34: Ohno S, Okuyama R, Aruga A, Sugiyama H, Yamamoto M. Phase I trial of Wilms Tumor 1 (WT1) peptide vaccine with GM-CSF or CpG in patients with solid malignancy. Anticancer Res 2012: 32: Lee KW, Oh DH, Lee C, Yang SY. Allelic and haplotypic diversity of HLA-A, -B, -C, -DRB1, and -DQB1 genes in the Korean population. Tissue Antigens 2005: 65: Yang G, Deng YJ, Hu SN et al. HLA-A, -B, and -DRB1 polymorphism defined by sequence based typing of the Han population in Northern China. Tissue Antigens 2006: 67: Chen S, Hu Q, Xie Y et al. Origin of Tibeto-Burman speakers: evidence from HLA allele distribution in Lisu and Nu inhabiting Yunnan of China. Hum Immunol 2007: 68: Lai MJ, Wen SH, Lin YH et al. Distributions of human leukocyte antigen-a, -B, and -DRB1 alleles and haplotypes based on 46,915 Taiwanese donors. Hum Immunol 2010: 71: Wen SH, Lai MJ, Yang KL. Human leukocyte antigen-a, -B, and -DRB1 haplotypes of cord blood units in the Tzu Chi Taiwan Cord Blood Bank. Hum Immunol 2008: 69: Flomenberg N, Baxter-Lowe LA, Confer D et al. Impact of HLA class I and class II high-resolution matching on outcomes of unrelated donor bone marrow transplantation: HLA-C mismatching is associated with a strong adverse effect on transplantation outcome. Blood 2004: 104: Kanda Y, Kanda J, Atsuta Y et al. Impact of a single human leucocyte antigen (HLA) allele mismatch on the outcome of unrelated bone marrow transplantation over two time periods. A retrospective analysis of 3003 patients from the HLA Working Group of the Japan Society for Blood and Marrow Transplantation. Br J Haematol 2013: 161: John Wiley & Sons A/S. Published by John Wiley & Sons Ltd 259
Allele and Haplotype Frequencies of Human Leukocyte Antigen-A, -B, -C, -DRB1, and -DQB1 From Sequence- Based DNA Typing Data in Koreans
 Original Article Diagnostic Immunology Ann Lab Med 2015;35:429-435 http://dx.doi.org/10.3343/alm.2015.35.4.429 ISSN 2234-3806 eissn 2234-3814 Allele and Haplotype Frequencies of Human Leukocyte Antigen-A,
Original Article Diagnostic Immunology Ann Lab Med 2015;35:429-435 http://dx.doi.org/10.3343/alm.2015.35.4.429 ISSN 2234-3806 eissn 2234-3814 Allele and Haplotype Frequencies of Human Leukocyte Antigen-A,
Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005)
 Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005) Stephen Spellman Research Manager NMDP Scientific Services Maria Brown Scientific Services Specialist Data Management Conference 2007 1
Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005) Stephen Spellman Research Manager NMDP Scientific Services Maria Brown Scientific Services Specialist Data Management Conference 2007 1
The Human Major Histocompatibility Complex
 The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
Documentation of Changes to EFI Standards: v 5.6.1
 Modified Standard B - PERSONNEL QUALIFICATIONS B1.000 The laboratory must employ one or more individuals who meet the qualifications and fulfil the responsibilities of the Director/Co-Director, Technical
Modified Standard B - PERSONNEL QUALIFICATIONS B1.000 The laboratory must employ one or more individuals who meet the qualifications and fulfil the responsibilities of the Director/Co-Director, Technical
Human Leukocyte Antigens and donor selection
 Human Leukocyte Antigens and donor selection Duangtawan Thammanichanond, MD. PhD. Histocompatibility and Immunogenetics Laboratory, Faculty of Medicine Ramathibodi Hospital, Mahidol University Outline
Human Leukocyte Antigens and donor selection Duangtawan Thammanichanond, MD. PhD. Histocompatibility and Immunogenetics Laboratory, Faculty of Medicine Ramathibodi Hospital, Mahidol University Outline
Histocompatibility Evaluations for HSCT at JHMI. M. Sue Leffell, PhD. Professor of Medicine Laboratory Director
 Histocompatibility Evaluations for HSCT at JHMI M. Sue Leffell, PhD Professor of Medicine Laboratory Director JHMI Patient Population Adults Peds NMDP data >20,000 HSCT JHMI HSCT Protocols Bone marrow
Histocompatibility Evaluations for HSCT at JHMI M. Sue Leffell, PhD Professor of Medicine Laboratory Director JHMI Patient Population Adults Peds NMDP data >20,000 HSCT JHMI HSCT Protocols Bone marrow
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS
 DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97
 88 Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97 ARTICLE HLA gene and haplotype frequency in renal transplant recipients and donors of Uttar Pradesh (North India) S Agrawal, AK Singh, RK
88 Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97 ARTICLE HLA gene and haplotype frequency in renal transplant recipients and donors of Uttar Pradesh (North India) S Agrawal, AK Singh, RK
Role of NMDP Repository in the Evolution of HLA Matching and Typing for Unrelated Donor HCT
 Role of NMDP Repository in the Evolution of HLA Matching and Typing for Unrelated Donor HCT Stephen Spellman, MBS Director, Immunobiology and Observational Research Assistant Scientific Director CIBMTR,
Role of NMDP Repository in the Evolution of HLA Matching and Typing for Unrelated Donor HCT Stephen Spellman, MBS Director, Immunobiology and Observational Research Assistant Scientific Director CIBMTR,
HLA-A*26 and Susceptibility of Iranian Patients with Non-Hodgkin Lymphoma
 HLA-A*26 and Susceptibility of Iranian Patients with Non-Hodgkin Lymphoma Arezou Sayad 1, Mohammad Taghi Akbari 2**, Mahshid Mehdizadeh 3,4, Mohammad Taheri 1, Abbas Hajifathali 3* 1 Department of Medical
HLA-A*26 and Susceptibility of Iranian Patients with Non-Hodgkin Lymphoma Arezou Sayad 1, Mohammad Taghi Akbari 2**, Mahshid Mehdizadeh 3,4, Mohammad Taheri 1, Abbas Hajifathali 3* 1 Department of Medical
Basel - 6 September J.-M. Tiercy National Reference Laboratory for Histocompatibility (LNRH) University Hospital Geneva
 Basel - 6 eptember 2012 J.-M. Tiercy National Reference Laboratory for Histocompatibility (LNRH) University Hospital Geneva Outline the HLA system is (a) complex anti-hla immunisation and alloreactivity
Basel - 6 eptember 2012 J.-M. Tiercy National Reference Laboratory for Histocompatibility (LNRH) University Hospital Geneva Outline the HLA system is (a) complex anti-hla immunisation and alloreactivity
The role of HLA in Allogeneic Hematopoietic Stem Cell Transplantation and Platelet Refractoriness.
 The role of HLA in Allogeneic Hematopoietic Stem Cell Transplantation and Platelet Refractoriness. Robert Liwski, MD, PhD, FRCPC Medical Director HLA Typing Laboratory Department of Pathology Dalhousie
The role of HLA in Allogeneic Hematopoietic Stem Cell Transplantation and Platelet Refractoriness. Robert Liwski, MD, PhD, FRCPC Medical Director HLA Typing Laboratory Department of Pathology Dalhousie
EBMT2008_1_21:EBMT :06 Pagina 46 * CHAPTER 3. Immunogenetics of allogeneic HSCT * 3.1. The role of HLA in HSCT. J.M.
 EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 46 * CHAPTER 3 Immunogenetics of allogeneic HSCT * 3.1 The role of HLA in HSCT J.M. Tiercy EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 47 CHAPTER 3.1 HLA and
EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 46 * CHAPTER 3 Immunogenetics of allogeneic HSCT * 3.1 The role of HLA in HSCT J.M. Tiercy EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 47 CHAPTER 3.1 HLA and
Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis
 Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis H.-Y. Zou, W.-Z. Yu, Z. Wang, J. He and M. Jiao Institute of Clinical Medicine, Urumqi General Hospital, Lanzhou
Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis H.-Y. Zou, W.-Z. Yu, Z. Wang, J. He and M. Jiao Institute of Clinical Medicine, Urumqi General Hospital, Lanzhou
25/10/2017. Clinical Relevance of the HLA System in Blood Transfusion. Outline of talk. Major Histocompatibility Complex
 Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
HLA Mismatches. Professor Steven GE Marsh. Anthony Nolan Research Institute EBMT Anthony Nolan Research Institute
 HLA Mismatches Professor Steven GE Marsh HLA Mismatches HLA Genes, Structure, Polymorphism HLA Nomenclature HLA Mismatches in HSCT Defining a mismatch HLA Mismatches HLA Genes, Structure, Polymorphism
HLA Mismatches Professor Steven GE Marsh HLA Mismatches HLA Genes, Structure, Polymorphism HLA Nomenclature HLA Mismatches in HSCT Defining a mismatch HLA Mismatches HLA Genes, Structure, Polymorphism
Profiling HLA motifs by large scale peptide sequencing Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009
 Profiling HLA motifs by large scale peptide sequencing 2009 Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009 HLA Background The human leukocyte antigen system (HLA) is the
Profiling HLA motifs by large scale peptide sequencing 2009 Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009 HLA Background The human leukocyte antigen system (HLA) is the
21/05/2018. Continuing Education. Presentation Recording. learn.immucor.com
 Transplant Webinar Series: Ep. 6 Donor Selection for Haematopoietic Stem Cell Transplantation Future Webinars The Role of NGS in the Transplant Setting Featuring Dr Sujatha Krishnakumar Sirona Genomics,
Transplant Webinar Series: Ep. 6 Donor Selection for Haematopoietic Stem Cell Transplantation Future Webinars The Role of NGS in the Transplant Setting Featuring Dr Sujatha Krishnakumar Sirona Genomics,
New trends in donor selection in Europe: "best match" versus haploidentical. Prof Jakob R Passweg
 New trends in donor selection in Europe: "best match" versus haploidentical Prof Jakob R Passweg HSCT change in donor type: 1990-2015 9000 H S C T 8000 7000 6000 5000 4000 HLA identical sibling/twin Haplo-identical
New trends in donor selection in Europe: "best match" versus haploidentical Prof Jakob R Passweg HSCT change in donor type: 1990-2015 9000 H S C T 8000 7000 6000 5000 4000 HLA identical sibling/twin Haplo-identical
MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH
 MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH TEJASWINI M. DHAWALE, M.D. HEME FELLOWS CONFERENCE NOVEMBER 08, 2013 CASE PRESENTATION 51 yo M with history of MDS (unilinear
MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH TEJASWINI M. DHAWALE, M.D. HEME FELLOWS CONFERENCE NOVEMBER 08, 2013 CASE PRESENTATION 51 yo M with history of MDS (unilinear
The MHC and Transplantation Brendan Clark. Transplant Immunology, St James s University Hospital, Leeds, UK
 The MHC and Transplantation Brendan Clark Transplant Immunology, St James s University Hospital, Leeds, UK Blood Groups Immunofluorescent staining has revealed blood group substance in the cell membranes
The MHC and Transplantation Brendan Clark Transplant Immunology, St James s University Hospital, Leeds, UK Blood Groups Immunofluorescent staining has revealed blood group substance in the cell membranes
How to Find an Unrelated Donor Theory & Technology
 How to Find an Unrelated Donor Theory & Technology Carlheinz R. Müller Zentrales Knochenmarkspender-Register für die Bundesrepublik Deutschland (ZKRD) Ulm, Germany How to Find an Unrelated Donor HLA-Basics
How to Find an Unrelated Donor Theory & Technology Carlheinz R. Müller Zentrales Knochenmarkspender-Register für die Bundesrepublik Deutschland (ZKRD) Ulm, Germany How to Find an Unrelated Donor HLA-Basics
CHAPTER 3 LABORATORY PROCEDURES
 CHAPTER 3 LABORATORY PROCEDURES CHAPTER 3 LABORATORY PROCEDURES 3.1 HLA TYPING Molecular HLA typing will be performed for all donor cord blood units and patients in the three reference laboratories identified
CHAPTER 3 LABORATORY PROCEDURES CHAPTER 3 LABORATORY PROCEDURES 3.1 HLA TYPING Molecular HLA typing will be performed for all donor cord blood units and patients in the three reference laboratories identified
Minimal Requirements for Histocompatibility & Immunogenetics Laboratory
 Minimal Requirements for Histocompatibility & Immunogenetics Laboratory The 4 th WBMT Congress and Workshop Riyadh, KSA - January 15-17, 2017 HLA Discovery, 1958 The Nobel Prize in Physiology or Medicine
Minimal Requirements for Histocompatibility & Immunogenetics Laboratory The 4 th WBMT Congress and Workshop Riyadh, KSA - January 15-17, 2017 HLA Discovery, 1958 The Nobel Prize in Physiology or Medicine
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/20898 holds various files of this Leiden University dissertation. Author: Jöris, Monique Maria Title: Challenges in unrelated hematopoietic stem cell transplantation.
Cover Page The handle http://hdl.handle.net/1887/20898 holds various files of this Leiden University dissertation. Author: Jöris, Monique Maria Title: Challenges in unrelated hematopoietic stem cell transplantation.
National Marrow Donor Program HLA-Matching Guidelines for Unrelated Marrow Transplants
 Biology of Blood and Marrow Transplantation 9:610-615 (2003) 2003 American Society for Blood and Marrow Transplantation 1083-8791/03/0910-0003$30.00/0 doi:10.1016/s1083-8791(03)00329-x National Marrow
Biology of Blood and Marrow Transplantation 9:610-615 (2003) 2003 American Society for Blood and Marrow Transplantation 1083-8791/03/0910-0003$30.00/0 doi:10.1016/s1083-8791(03)00329-x National Marrow
Clinical Relevance of the HLA System in Blood Transfusion. Dr Colin J Brown PhD FRCPath. October 2017
 Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
Application of Calculated Panel Reactive Antibody Using HLA Frequencies in Koreans
 Original Article Diagnostic Immunology Jang J-Y, et al. Ann Lab Med 2012;32:66-72 ISSN 2234-3806 eissn 2234-3814 Application of Calculated Panel Reactive Antibody Using HLA Frequencies in Koreans Ji-Young
Original Article Diagnostic Immunology Jang J-Y, et al. Ann Lab Med 2012;32:66-72 ISSN 2234-3806 eissn 2234-3814 Application of Calculated Panel Reactive Antibody Using HLA Frequencies in Koreans Ji-Young
HLA haplotype A33-B58-Cw10 may modulate radiographic development of bamboo spine in Taiwanese patients with primary ankylosing spondylitis
 Disease Markers 26 (2009) 93 96 93 DOI 10.3233/DMA-2009-0619 IOS Press HLA haplotype A33-B58-Cw10 may modulate radiographic development of bamboo spine in Taiwanese patients with primary ankylosing spondylitis
Disease Markers 26 (2009) 93 96 93 DOI 10.3233/DMA-2009-0619 IOS Press HLA haplotype A33-B58-Cw10 may modulate radiographic development of bamboo spine in Taiwanese patients with primary ankylosing spondylitis
Transplantation. Immunology Unit College of Medicine King Saud University
 Transplantation Immunology Unit College of Medicine King Saud University Objectives To understand the diversity among human leukocyte antigens (HLA) or major histocompatibility complex (MHC) To know the
Transplantation Immunology Unit College of Medicine King Saud University Objectives To understand the diversity among human leukocyte antigens (HLA) or major histocompatibility complex (MHC) To know the
Available online at , 2(1):27-37
 Available online at www.apjbg.com Asia-Pacific Journal of Blood Types and Genes 2018, 2(1):27-37 APJBG Allele and haplotype frequencies of human leukocyte antigen-a, -B, -C, -DRB1, and -DQB1 in Chinese
Available online at www.apjbg.com Asia-Pacific Journal of Blood Types and Genes 2018, 2(1):27-37 APJBG Allele and haplotype frequencies of human leukocyte antigen-a, -B, -C, -DRB1, and -DQB1 in Chinese
HLA and new technologies. Vicky Van Sandt
 HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
HLA and antigen presentation. Department of Immunology Charles University, 2nd Medical School University Hospital Motol
 HLA and antigen presentation Department of Immunology Charles University, 2nd Medical School University Hospital Motol MHC in adaptive immunity Characteristics Specificity Innate For structures shared
HLA and antigen presentation Department of Immunology Charles University, 2nd Medical School University Hospital Motol MHC in adaptive immunity Characteristics Specificity Innate For structures shared
XIV. HLA AND TRANSPLANTATION MEDICINE
 XIV. HLA AND TRANSPLANTATION MEDICINE A. Introduction 1. The HLA system includes a complex array of genes and their molecular products that are involved in immune regulation and cellular differentiation.
XIV. HLA AND TRANSPLANTATION MEDICINE A. Introduction 1. The HLA system includes a complex array of genes and their molecular products that are involved in immune regulation and cellular differentiation.
Haplo vs Cord vs URD Debate
 3rd Annual ASBMT Regional Conference for NPs, PAs and Fellows Haplo vs Cord vs URD Debate Claudio G. Brunstein Associate Professor University of Minnesota Medical School Take home message Finding a donor
3rd Annual ASBMT Regional Conference for NPs, PAs and Fellows Haplo vs Cord vs URD Debate Claudio G. Brunstein Associate Professor University of Minnesota Medical School Take home message Finding a donor
Significance of the MHC
 CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
Validation of the MIA FORA NGS FLEX Assay Using Buccal Swabs as the Sample Source
 S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. Introduction
S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. S. Krishnakumar, M. Li, C. Wang, M, Osada, R. Kuehn, M. Fukushima and Y. Thorstenson Immucor, Inc. Introduction
Association of HLA-DRB alleles and pulmonary tuberculosis in North Chinese patients
 Association of HLA-DRB alleles and pulmonary tuberculosis in North Chinese patients G.L. Shi, X.L. Hu, L. Yang, C.L. Rong, Y.L. Guo and C.X. Song Department of Clinical Immunology Laboratory, Beijing Tuberculosis
Association of HLA-DRB alleles and pulmonary tuberculosis in North Chinese patients G.L. Shi, X.L. Hu, L. Yang, C.L. Rong, Y.L. Guo and C.X. Song Department of Clinical Immunology Laboratory, Beijing Tuberculosis
Sylwia Mizia, 1 Dorota Dera-Joachimiak, 1 Malgorzata Polak, 1 Katarzyna Koscinska, 1 Mariola Sedzimirska, 1 and Andrzej Lange 1, 2. 1.
 Bone Marrow Research Volume 2012, Article ID 873695, 5 pages doi:10.1155/2012/873695 Clinical Study Both Optimal Matching and Procedure Duration Influence Survival of Patients after Unrelated Donor Hematopoietic
Bone Marrow Research Volume 2012, Article ID 873695, 5 pages doi:10.1155/2012/873695 Clinical Study Both Optimal Matching and Procedure Duration Influence Survival of Patients after Unrelated Donor Hematopoietic
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Gragert L, Eapen M, Williams E, et al. HLA match likelihoods
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Gragert L, Eapen M, Williams E, et al. HLA match likelihoods
HLA Amino Acid Polymorphisms and Kidney Allograft Survival. Supplemental Digital Content
 HLA Amino Acid Polymorphisms and Kidney Allograft Survival Supplemental Digital Content Malek Kamoun, MD, PhD a, Keith P. McCullough, MS b, Martin Maiers, MS c, Marcelo A. Fernandez Vina, MD, PhD d, Hongzhe
HLA Amino Acid Polymorphisms and Kidney Allograft Survival Supplemental Digital Content Malek Kamoun, MD, PhD a, Keith P. McCullough, MS b, Martin Maiers, MS c, Marcelo A. Fernandez Vina, MD, PhD d, Hongzhe
Trends in Hematopoietic Cell Transplantation. AAMAC Patient Education Day Oct 2014
 Trends in Hematopoietic Cell Transplantation AAMAC Patient Education Day Oct 2014 Objectives Review the principles behind allogeneic stem cell transplantation Outline the process of transplant, some of
Trends in Hematopoietic Cell Transplantation AAMAC Patient Education Day Oct 2014 Objectives Review the principles behind allogeneic stem cell transplantation Outline the process of transplant, some of
OneMatch Stem Cell and Marrow Network. Training Guide
 OneMatch Stem Cell and Marrow Network Training Guide What is OneMatch all about? OneMatch is a Canadian Program that matches and coordinates the collection of stem cells from potential donors to help save
OneMatch Stem Cell and Marrow Network Training Guide What is OneMatch all about? OneMatch is a Canadian Program that matches and coordinates the collection of stem cells from potential donors to help save
Rob Wynn RMCH & University of Manchester, UK. HCT in Children
 Rob Wynn RMCH & University of Manchester, UK HCT in Children Summary Indications for HCT in children Donor selection for Paediatric HCT Using cords Achieving engraftment in HCT Conditioning Immune action
Rob Wynn RMCH & University of Manchester, UK HCT in Children Summary Indications for HCT in children Donor selection for Paediatric HCT Using cords Achieving engraftment in HCT Conditioning Immune action
HLA Match Likelihoods for Hematopoietic Stem-Cell Grafts in the U.S. Registry
 The new england journal of medicine special article HLA Likelihoods for Hematopoietic Stem-Cell Grafts in the U.S. Registry Loren Gragert, B.S., B.A., Mary Eapen, M.B., B.S., Eric Williams, Ph.D., John
The new england journal of medicine special article HLA Likelihoods for Hematopoietic Stem-Cell Grafts in the U.S. Registry Loren Gragert, B.S., B.A., Mary Eapen, M.B., B.S., Eric Williams, Ph.D., John
OPTN/UNOS Policy Notice Review of HLA Tables (2016)
 Review of HLA Tables (2016) Sponsoring Committee: Policy/Bylaws Affected: Histocompatibility Policy 4.10 (Reference Tables of HLA Antigen Values and Split Equivalences) Public Comment: July 31, 2017 October
Review of HLA Tables (2016) Sponsoring Committee: Policy/Bylaws Affected: Histocompatibility Policy 4.10 (Reference Tables of HLA Antigen Values and Split Equivalences) Public Comment: July 31, 2017 October
Handling Immunogenetic Data Managing and Validating HLA Data
 Handling Immunogenetic Data Managing and Validating HLA Data Steven J. Mack PhD Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012 Overview 1. Master Analytical
Handling Immunogenetic Data Managing and Validating HLA Data Steven J. Mack PhD Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012 Overview 1. Master Analytical
Transplantation Immunology Booklet C
 Booklet C Pretest Correct Answers 3. (C) is correct. One of the major barriers to successful hematopoietic stem cell transplantation is the antigens coded for by the major histocompatibility gene complex.
Booklet C Pretest Correct Answers 3. (C) is correct. One of the major barriers to successful hematopoietic stem cell transplantation is the antigens coded for by the major histocompatibility gene complex.
General Terms: Appendix B. National Marrow Donor Program and The Medical College of Wisconsin
 Glossary of Terms This appendix is divided into two sections. The first section, General Terms, defines terms used throughout the CIBMTR data collection forms. The second section, FormsNet TM 2 Terms,
Glossary of Terms This appendix is divided into two sections. The first section, General Terms, defines terms used throughout the CIBMTR data collection forms. The second section, FormsNet TM 2 Terms,
The Use of HLA /HPA Selected Platelets
 The Use of HLA /HPA Selected Platelets Dr Colin Brown PhD FRCPath. Consultant Clinical Scientist Histocompatibility and Immunogenetics HLA and Transfusion Human Leucocyte Antigens HLA Human Platelet Antigens
The Use of HLA /HPA Selected Platelets Dr Colin Brown PhD FRCPath. Consultant Clinical Scientist Histocompatibility and Immunogenetics HLA and Transfusion Human Leucocyte Antigens HLA Human Platelet Antigens
Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany
 HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
Biol Blood Marrow Transplant 17: (2011) Ó 2011 American Society for Blood and Marrow Transplantation
 Outcomes of Patients with Myeloid Malignancies Treated with Allogeneic Hematopoietic Stem Cell Transplantation from Matched Unrelated Donors Compared with One Human Leukocyte Antigen Mismatched Related
Outcomes of Patients with Myeloid Malignancies Treated with Allogeneic Hematopoietic Stem Cell Transplantation from Matched Unrelated Donors Compared with One Human Leukocyte Antigen Mismatched Related
HLA and antigen presentation. Department of Immunology Charles University, 2nd Medical School University Hospital Motol
 HLA and antigen presentation Department of Immunology Charles University, 2nd Medical School University Hospital Motol MHC in adaptive immunity Characteristics Specificity Innate For structures shared
HLA and antigen presentation Department of Immunology Charles University, 2nd Medical School University Hospital Motol MHC in adaptive immunity Characteristics Specificity Innate For structures shared
The impact of HLA matching on unrelated donor hematopoietic stem cell transplantation in Korean children
 VOLUME 46 ㆍ NUMBER ㆍ March 0 THE KOREAN JOURNAL OF HEMATOLOGY ORIGINAL ARTICLE The impact of HLA matching on unrelated donor hematopoietic stem cell transplantation in Korean children Meerim Park, Kyung
VOLUME 46 ㆍ NUMBER ㆍ March 0 THE KOREAN JOURNAL OF HEMATOLOGY ORIGINAL ARTICLE The impact of HLA matching on unrelated donor hematopoietic stem cell transplantation in Korean children Meerim Park, Kyung
The Relationship between Minor Histocompatibility Antigens and Graft Versus Host Disease in Unrelated Peripheral Blood Stem Cell Transplants
 mhags and GVHD Page 1 The Relationship between Minor Histocompatibility Antigens and Graft Versus Host Disease in Unrelated Peripheral Blood Stem Cell Transplants Running Title: mhags and GVHD Conflict
mhags and GVHD Page 1 The Relationship between Minor Histocompatibility Antigens and Graft Versus Host Disease in Unrelated Peripheral Blood Stem Cell Transplants Running Title: mhags and GVHD Conflict
Review Article Association between HLA-DQ Gene Polymorphisms and HBV-Related Hepatocellular Carcinoma
 Hindawi Gastroenterology Research and Practice Volume 2017, Article ID 7150386, 11 pages https://doiorg/101155/2017/7150386 Review Article Association between HLA-DQ Gene Polymorphisms and HBV-Related
Hindawi Gastroenterology Research and Practice Volume 2017, Article ID 7150386, 11 pages https://doiorg/101155/2017/7150386 Review Article Association between HLA-DQ Gene Polymorphisms and HBV-Related
Immunogenetics in SARS: a casecontrol
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES MHL Ng 吳香玲 SH Cheng 鄭淑恆 KM Lau 劉建盟 GM Leung 梁卓偉 US Khoo 邱瑋璇 BCW Zee 徐仲鍈 JJY Sung 沈祖堯 Key Messages 1. Human leukocyte antigen (HLA) genotypes from 102
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES MHL Ng 吳香玲 SH Cheng 鄭淑恆 KM Lau 劉建盟 GM Leung 梁卓偉 US Khoo 邱瑋璇 BCW Zee 徐仲鍈 JJY Sung 沈祖堯 Key Messages 1. Human leukocyte antigen (HLA) genotypes from 102
HLA-A * L
 What is Nomenclature? HLA Nomenclature 3.0 Signifies DNA Subtype Differences outside the coding region (introns) HLA-A * 4 0 01 0 L Steve Spellman Sr. Manager, NMDP Asst. Scientific Director, CIBMTR Locus
What is Nomenclature? HLA Nomenclature 3.0 Signifies DNA Subtype Differences outside the coding region (introns) HLA-A * 4 0 01 0 L Steve Spellman Sr. Manager, NMDP Asst. Scientific Director, CIBMTR Locus
Influence of interleukin-18 gene polymorphisms on acute pancreatitis susceptibility in a Chinese population
 Influence of interleukin-18 gene polymorphisms on acute pancreatitis susceptibility in a Chinese population H.B. Gui 1, X.G. Du 2, Z.H. Fu 3 and X.M. Chen 1 1 Department of Emergency, The First Affiliated
Influence of interleukin-18 gene polymorphisms on acute pancreatitis susceptibility in a Chinese population H.B. Gui 1, X.G. Du 2, Z.H. Fu 3 and X.M. Chen 1 1 Department of Emergency, The First Affiliated
D7.1 Report summarising results of survey of EU countries to identify volumes and trends in relation to the import and export of stem cells
 Disclaimer: The content of this Deliverable represents the views of the author only and is his/her sole responsibility; it cannot be considered to reflect the views of the European Commission and/or the
Disclaimer: The content of this Deliverable represents the views of the author only and is his/her sole responsibility; it cannot be considered to reflect the views of the European Commission and/or the
Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population
 Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population J. Zhu 1 *, F. He 2 *, D.D. Zhang 2 *, J.Y. Yang 2, J. Cheng 1, R. Wu 1, B. Gong 2, X.Q. Liu
Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population J. Zhu 1 *, F. He 2 *, D.D. Zhang 2 *, J.Y. Yang 2, J. Cheng 1, R. Wu 1, B. Gong 2, X.Q. Liu
D7.1 Report summarising results of survey of EU countries to identify volumes and trends in relation to the import and export of stem cells
 Disclaimer: The content of this Deliverable represents the views of the author only and is his/her sole responsibility; it cannot be considered to reflect the views of the European Commission and/or the
Disclaimer: The content of this Deliverable represents the views of the author only and is his/her sole responsibility; it cannot be considered to reflect the views of the European Commission and/or the
Diversity and Frequencies of HLA Class I and Class II Genes of an East African Population
 Open Journal of Genetics, 2014, 4, 99-124 Published Online April 2014 in SciRes. http://www.scirp.org/journal/ojgen http://dx.doi.org/10.4236/ojgen.2014.42013 Diversity and Frequencies of HLA Class I and
Open Journal of Genetics, 2014, 4, 99-124 Published Online April 2014 in SciRes. http://www.scirp.org/journal/ojgen http://dx.doi.org/10.4236/ojgen.2014.42013 Diversity and Frequencies of HLA Class I and
BST227 Introduction to Statistical Genetics. Lecture 4: Introduction to linkage and association analysis
 BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES. Irina Durbală
 ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
The National Marrow Donor Program. Graft Sources for Hematopoietic Cell Transplantation. Simon Bostic, URD Transplant Recipient
 1988 199 1992 1994 1996 1998 2 22 24 26 28 21 212 214 216 218 Adult Donors Cord Blood Units The National Donor Program Graft Sources for Hematopoietic Cell Transplantation Dennis L. Confer, MD Chief Medical
1988 199 1992 1994 1996 1998 2 22 24 26 28 21 212 214 216 218 Adult Donors Cord Blood Units The National Donor Program Graft Sources for Hematopoietic Cell Transplantation Dennis L. Confer, MD Chief Medical
IL10 rs polymorphism is associated with liver cirrhosis and chronic hepatitis B
 IL10 rs1800896 polymorphism is associated with liver cirrhosis and chronic hepatitis B L.N. Cao 1, S.L. Cheng 2 and W. Liu 3 1 Kidney Disease Department of Internal Medicine, Xianyang Central Hospital,
IL10 rs1800896 polymorphism is associated with liver cirrhosis and chronic hepatitis B L.N. Cao 1, S.L. Cheng 2 and W. Liu 3 1 Kidney Disease Department of Internal Medicine, Xianyang Central Hospital,
Introduction to Clinical Hematopoietic Cell Transplantation (HCT) George Chen, MD Thursday, May 03, 2018
 Introduction to Clinical Hematopoietic Cell Transplantation (HCT) George Chen, MD Thursday, May 03, 2018 The transfer of hematopoietic progenitor and stem cells for therapeutic purposes Hematopoietic Cell
Introduction to Clinical Hematopoietic Cell Transplantation (HCT) George Chen, MD Thursday, May 03, 2018 The transfer of hematopoietic progenitor and stem cells for therapeutic purposes Hematopoietic Cell
Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs.
 Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs. C.J. SAVOIE, N. KAMIKAWAJI, T. SASAZUKI Dept. of Genetics, Medical Institute of Bioregulation, Kyushu
Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs. C.J. SAVOIE, N. KAMIKAWAJI, T. SASAZUKI Dept. of Genetics, Medical Institute of Bioregulation, Kyushu
HLA and more. Ilias I.N. Doxiadis. Geneva 03/04/2012.
 www.ebmt.org HLA and more Ilias I.N. Doxiadis Geneva 03/04/2012 HLA and more HLA and more / Doxiadis 2 Topic of the day Compatibility testing is a type of testing used to ensure compatibility of the system/application/website
www.ebmt.org HLA and more Ilias I.N. Doxiadis Geneva 03/04/2012 HLA and more HLA and more / Doxiadis 2 Topic of the day Compatibility testing is a type of testing used to ensure compatibility of the system/application/website
IMMUNOLOGY. Elementary Knowledge of Major Histocompatibility Complex and HLA Typing
 IMMUNOLOGY Elementary Knowledge of Major Histocompatibility Complex and HLA Typing Tapasya Srivastava and Subrata Sinha Department of Biochemistry All India Institute of Medical Sciences New Delhi - 110029
IMMUNOLOGY Elementary Knowledge of Major Histocompatibility Complex and HLA Typing Tapasya Srivastava and Subrata Sinha Department of Biochemistry All India Institute of Medical Sciences New Delhi - 110029
Seventy percent of people who are candidates for allogeneic hematopoietic
 274 7th Biennial Symposium on Minorities, the Medically Underserved and Cancer Supplement to Cancer The National Marrow Donor Program Meeting the Needs of the Medically Underserved Dennis L. Confer, M.D.
274 7th Biennial Symposium on Minorities, the Medically Underserved and Cancer Supplement to Cancer The National Marrow Donor Program Meeting the Needs of the Medically Underserved Dennis L. Confer, M.D.
Historical definition of Antigen. An antigen is a foreign substance that elicits the production of antibodies that specifically binds to the antigen.
 Historical definition of Antigen An antigen is a foreign substance that elicits the production of antibodies that specifically binds to the antigen. Historical definition of Antigen An antigen is a foreign
Historical definition of Antigen An antigen is a foreign substance that elicits the production of antibodies that specifically binds to the antigen. Historical definition of Antigen An antigen is a foreign
Supplementary Figure 1 Dosage correlation between imputed and genotyped alleles Imputed dosages (0 to 2) of 2-digit alleles (red) and 4-digit alleles
 Supplementary Figure 1 Dosage correlation between imputed and genotyped alleles Imputed dosages (0 to 2) of 2-digit alleles (red) and 4-digit alleles (green) of (A) HLA-A, HLA-B, (C) HLA-C, (D) HLA-DQA1,
Supplementary Figure 1 Dosage correlation between imputed and genotyped alleles Imputed dosages (0 to 2) of 2-digit alleles (red) and 4-digit alleles (green) of (A) HLA-A, HLA-B, (C) HLA-C, (D) HLA-DQA1,
Transplant Booklet D Page 1
 Booklet D Pretest Correct Answers 4. (A) is correct. Technically, performing a hematopoietic stem cell transplant is one of the simplest transplantation procedures. The hematopoietic stem cells are infused
Booklet D Pretest Correct Answers 4. (A) is correct. Technically, performing a hematopoietic stem cell transplant is one of the simplest transplantation procedures. The hematopoietic stem cells are infused
Outcomes of pediatric bone marrow transplantation for leukemia and myelodysplasia using matched. unrelated donors
 Outcomes of pediatric bone marrow transplantation for leukemia and myelodysplasia using matched sibling, mismatched related or matched unrelated donors Immunobiology Working Committee PIs: Peter Shaw and
Outcomes of pediatric bone marrow transplantation for leukemia and myelodysplasia using matched sibling, mismatched related or matched unrelated donors Immunobiology Working Committee PIs: Peter Shaw and
Effect of HLA mismatch on acute graft-versus-host disease
 Int J Hematol (2013) 98:300 308 DOI 10.1007/s12185-013-1405-x PROGRESS IN HEMATOLOGY New clinical and basic aspects of graft-versus-host disease Effect of HLA mismatch on acute graft-versus-host disease
Int J Hematol (2013) 98:300 308 DOI 10.1007/s12185-013-1405-x PROGRESS IN HEMATOLOGY New clinical and basic aspects of graft-versus-host disease Effect of HLA mismatch on acute graft-versus-host disease
Genome-wide association studies for human narcolepsy and other complex diseases
 ETHZ-JST Joint WS Sept. 15-16, 2008 Genome-wide association studies for human narcolepsy and other complex diseases Katsushi Tokunaga Department of Human Genetics Graduate School of Medicine The University
ETHZ-JST Joint WS Sept. 15-16, 2008 Genome-wide association studies for human narcolepsy and other complex diseases Katsushi Tokunaga Department of Human Genetics Graduate School of Medicine The University
the HLA complex Hanna Mustaniemi,
 the HLA complex Hanna Mustaniemi, 28.11.2007 The Major Histocompatibility Complex Major histocompatibility complex (MHC) is a gene region found in nearly all vertebrates encodes proteins with important
the HLA complex Hanna Mustaniemi, 28.11.2007 The Major Histocompatibility Complex Major histocompatibility complex (MHC) is a gene region found in nearly all vertebrates encodes proteins with important
HLA Complex Genetics & Biology
 HLA Complex Genetics & Biology M. Tevfik DORAK Environmental & Occupational Health College of Public Health Florida International University Miami, USA http://www.dorak.info Mount Sinai Medical Center
HLA Complex Genetics & Biology M. Tevfik DORAK Environmental & Occupational Health College of Public Health Florida International University Miami, USA http://www.dorak.info Mount Sinai Medical Center
MHC class I MHC class II Structure of MHC antigens:
 MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
An Introduction to Bone Marrow Transplant
 Introduction to Blood Cancers An Introduction to Bone Marrow Transplant Rushang Patel, MD, PhD, FACP Florida Hospital Medical Group S My RBC Plt Gran Polycythemia Vera Essential Thrombocythemia AML, CML,
Introduction to Blood Cancers An Introduction to Bone Marrow Transplant Rushang Patel, MD, PhD, FACP Florida Hospital Medical Group S My RBC Plt Gran Polycythemia Vera Essential Thrombocythemia AML, CML,
The Major Histocompatibility Complex of Genes
 The Major Histocompatibility Complex of Genes Topic 4 The Major Histocompatibility Complex Outline of Lectures The immunological reasons for transplant rejection How the MHC was discovered using inbred
The Major Histocompatibility Complex of Genes Topic 4 The Major Histocompatibility Complex Outline of Lectures The immunological reasons for transplant rejection How the MHC was discovered using inbred
Factors Influencing Haematopoietic Progenitor cell transplant outcome Optimising donor selection
 Factors Influencing Haematopoietic Progenitor cell transplant outcome Optimising donor selection Alison Logan Transplantation Laboratory Manchester Royal Infirmary Haematopoietic progenitor cell transplants
Factors Influencing Haematopoietic Progenitor cell transplant outcome Optimising donor selection Alison Logan Transplantation Laboratory Manchester Royal Infirmary Haematopoietic progenitor cell transplants
Myoglobin A79G polymorphism association with exercise-induced skeletal muscle damage
 Myoglobin A79G polymorphism association with exercise-induced skeletal muscle damage T. Cui and M.S. Jiang College of Physical Education, Shandong University of Finance and Economics, Ji nan, Shandong,
Myoglobin A79G polymorphism association with exercise-induced skeletal muscle damage T. Cui and M.S. Jiang College of Physical Education, Shandong University of Finance and Economics, Ji nan, Shandong,
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
Definition of MHC supertypes through clustering of MHC peptide binding repertoires
 Definition of MHC supertypes through clustering of MHC peptide binding repertoires Pedro A. Reche and Ellis L. Reinherz Laboratory of Immunobiology and Department of Medical Oncology, Dana-Farber Cancer
Definition of MHC supertypes through clustering of MHC peptide binding repertoires Pedro A. Reche and Ellis L. Reinherz Laboratory of Immunobiology and Department of Medical Oncology, Dana-Farber Cancer
10/18/2012. A primer in HLA: The who, what, how and why. What?
 A primer in HLA: The who, what, how and why What? 1 First recognized in mice during 1930 s and 1940 s. Mouse (murine) experiments with tumors Independent observations were made in humans with leukoagglutinating
A primer in HLA: The who, what, how and why What? 1 First recognized in mice during 1930 s and 1940 s. Mouse (murine) experiments with tumors Independent observations were made in humans with leukoagglutinating
Diagnosis of CMV infection UPDATE ECIL
 UPDATE ECIL-4 2011 Recommendations for CMV and HHV-6 management in patients with hematological diseases Per Ljungman, Rafael de la Camara, Hermann Einsele, Dan Engelhard, Pierre Reusser, Jan Styczynski,
UPDATE ECIL-4 2011 Recommendations for CMV and HHV-6 management in patients with hematological diseases Per Ljungman, Rafael de la Camara, Hermann Einsele, Dan Engelhard, Pierre Reusser, Jan Styczynski,
BE THE MATCH. The Role of HLA in Finding a Match for Bone Marrow or Peripheral Blood Stem Cell Transplantation
 BE THE MATCH The Role of HLA in Finding a Match for Bone Marrow or Peripheral Blood Stem Cell Transplantation Leukemia Leukemia is a type of cancer of the blood or bone marrow. It is characterized by an
BE THE MATCH The Role of HLA in Finding a Match for Bone Marrow or Peripheral Blood Stem Cell Transplantation Leukemia Leukemia is a type of cancer of the blood or bone marrow. It is characterized by an
Virtual Crossmatch in Kidney Transplantation
 Virtual Crossmatch in Kidney Transplantation Shiva Samavat Associate Professor of Nephrology Labbafinejad Hospital SBMU 2018.11.21 All transplant candidates are screened to determine the degree of humoral
Virtual Crossmatch in Kidney Transplantation Shiva Samavat Associate Professor of Nephrology Labbafinejad Hospital SBMU 2018.11.21 All transplant candidates are screened to determine the degree of humoral
IMMUNOGENETICS AND TRANSPLANTATION
 IMMUNOGENETICS AND TRANSPLANTATION WHAT S THE MHC GOT TO DO WITH TRANSPLANTATION? We have learned that the molecules of Class I and II MHC are involved in presenting antigenic peptides to the receptors
IMMUNOGENETICS AND TRANSPLANTATION WHAT S THE MHC GOT TO DO WITH TRANSPLANTATION? We have learned that the molecules of Class I and II MHC are involved in presenting antigenic peptides to the receptors
Corporate Medical Policy
 Corporate Medical Policy Hematopoietic Stem-Cell Transplantation for Autoimmune Diseases File Name: Origination: Last CAP Review: Next CAP Review: Last Review: hematopoietic_stem-cell_transplantation_for_autoimmune_diseases
Corporate Medical Policy Hematopoietic Stem-Cell Transplantation for Autoimmune Diseases File Name: Origination: Last CAP Review: Next CAP Review: Last Review: hematopoietic_stem-cell_transplantation_for_autoimmune_diseases
Robert B. Colvin, M.D. Department of Pathology Massachusetts General Hospital Harvard Medical School
 Harvard-MIT Division of Health Sciences and Technology HST.035: Principle and Practice of Human Pathology Dr. Robert B. Colvin Transplantation: Friendly organs in a hostile environment Robert B. Colvin,
Harvard-MIT Division of Health Sciences and Technology HST.035: Principle and Practice of Human Pathology Dr. Robert B. Colvin Transplantation: Friendly organs in a hostile environment Robert B. Colvin,
Distribution of HLA Antigens Class I and II in Iraqi Arab population
 Distribution of HLA s Class I and II in Iraqi Arab population *Ahmed A.A. Al-Hassan, MSc, Immunology ; **Sana a Al-Naseri, MD, ph.d, immunopathology *** Batool H. Al-Ghurabi, ph.d, Immunology **** Mohammed
Distribution of HLA s Class I and II in Iraqi Arab population *Ahmed A.A. Al-Hassan, MSc, Immunology ; **Sana a Al-Naseri, MD, ph.d, immunopathology *** Batool H. Al-Ghurabi, ph.d, Immunology **** Mohammed
CHAPTER 2 PROTOCOL DESIGN
 CHAPTER 2 PROTOCOL DESIGN CHAPTER 2 PROTOCOL DESIGN 2.1 ELIGIBILITY CRITERIA Participants fulfilling the following criteria will be eligible for enrollment in the protocol: 1. Participant is diagnosed
CHAPTER 2 PROTOCOL DESIGN CHAPTER 2 PROTOCOL DESIGN 2.1 ELIGIBILITY CRITERIA Participants fulfilling the following criteria will be eligible for enrollment in the protocol: 1. Participant is diagnosed
Transplantation and Cancer
 Transplantation and Cancer Immunology Bởi: OpenStaxCollege The immune responses to transplanted organs and to cancer cells are both important medical issues. With the use of tissue typing and anti-rejection
Transplantation and Cancer Immunology Bởi: OpenStaxCollege The immune responses to transplanted organs and to cancer cells are both important medical issues. With the use of tissue typing and anti-rejection
HLA-DR-matched Parental Donors for Allogeneic Hematopoietic Stem Cell Transplantation in Patients with High-risk Acute Leukemia
 BRIEF COMMUNICATION HLA-DR-matched Parental Donors for Allogeneic Hematopoietic Stem Cell Transplantation in Patients with High-risk Acute Leukemia Shang-Ju Wu, Ming Yao,* Jih-Luh Tang, Bo-Sheng Ko, Hwei-Fang
BRIEF COMMUNICATION HLA-DR-matched Parental Donors for Allogeneic Hematopoietic Stem Cell Transplantation in Patients with High-risk Acute Leukemia Shang-Ju Wu, Ming Yao,* Jih-Luh Tang, Bo-Sheng Ko, Hwei-Fang
Dr. Yi-chi M. Kong August 8, 2001 Benjamini. Ch. 19, Pgs Page 1 of 10 TRANSPLANTATION
 Benjamini. Ch. 19, Pgs 379-399 Page 1 of 10 TRANSPLANTATION I. KINDS OF GRAFTS II. RELATIONSHIPS BETWEEN DONOR AND RECIPIENT Benjamini. Ch. 19, Pgs 379-399 Page 2 of 10 II.GRAFT REJECTION IS IMMUNOLOGIC
Benjamini. Ch. 19, Pgs 379-399 Page 1 of 10 TRANSPLANTATION I. KINDS OF GRAFTS II. RELATIONSHIPS BETWEEN DONOR AND RECIPIENT Benjamini. Ch. 19, Pgs 379-399 Page 2 of 10 II.GRAFT REJECTION IS IMMUNOLOGIC
Histocompatibility antigens
 Histocompatibility antigens Tuesday 09 November 2010 Telegraph UK Livers grown in the laboratory could eventually solve organ transplant shortage. Made-to-measure organs for transplantation are a step
Histocompatibility antigens Tuesday 09 November 2010 Telegraph UK Livers grown in the laboratory could eventually solve organ transplant shortage. Made-to-measure organs for transplantation are a step
