Completion of The Cancer Genome Atlas, what did we learn?
|
|
|
- Cornelius Lawrence
- 5 years ago
- Views:
Transcription
1 Charles M. Perou, Ph.D. Departments of Genetics Lineberger Comprehensive Cancer Center University of North Carolina at Chapel Hill Completion of The Cancer Genome Atlas, what did we learn?
2 TCGA Breast AWG Workshop University of North Carolina September 29-30, 2011
3 TCGA Breast Papers 1. Comprehensive molecular portraits of human breast tumours. TCGA et al., Nature, 2012 (PMID: ) 2. The molecular diversity of Luminal A breast tumors. Ciriello et al., BCRT, 2013 (PMID: ) 3. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Hoadley et al., Cell, 2014 (PMID: ) 4. Comprehensive molecular portraits of invasive lobular breast cancer. Ciriello et al., Cell (PMID: ) 5. DNA defects, epigenetics, and gene expression in cancer-adjacent breast: a study from The Cancer Genome Atlas. Troester et al., NPJ Breast Cancer, 2016 (PMID: ) 6. The molecular basis of breast cancer pathological phenotypes. Heng et al., J Pathology, 2017 (PMID: ) 7. Comparison of breast cancer molecular features and survival by African and European ancestry in The Cancer Genome Atlas. Huo et al., JAMA Oncol, 2017 (PMID: )
4 Comprehensive molecular portraits of human breast tumours. TCGA et al., Nature, 2012 (PMID: ) 466 Invasive Breast Carcinomas (348 common to all 6 platforms) 1. Whole Exome (all exons) capture and sequencing of tumor and normal (somatic coding sequence mutations and germline variation) WashU (n=507) 2. Gene expression analysis (Agilent microarrays and mrna-sequencing) - UNC 3. MicroRNA expression analysis (Illumina sequencing) - UBC 4. Tumor DNA copy number analysis (AFFY 6.0 microarrays and sequencing) - Broad 5. Tumor DNA methylation analysis (Illumina methylation arrays transitioning to sequencing)- USC 6. Reverse Phase Protein Arrays to detect proteins and phospho-proteins - MDACC 7. Integrated computational analyses (Genome Data Analysis Centers)- 6 Universities/sites 4
5 507 Breast Tumors Significantly Mutated Gene List by mrna intrinsic-subtype TCGA et al., Nature, 2012 (PMID: )
6 507 Breast Tumors Significantly Mutated Gene List # Mutated FDR TP E+00 PIK3CA E+00 TTN E-04 GATA E+00 MAP3K E+00 MLL E+00 CDH E+00 USH2A E-03 FLG E-03 RYR E-02 MAP2K E+00 FAT E-02 RUNX E+00 PTEN E+00 NCOR E-05 PIK3R E-11 NF E-03 SPTA E-02 TBX E-13 CTCF E-05 AFF E-03 SPEN E-02 CACNA1E E-02 AKT E-11 PTPRD E-02 MUC5B E-01 PREX E-02 ATM E-02 ARID1A E-01 SF3B E-04 RPGR E-03 PCDH E-02 FAM47C E-01 KIAA NA RB E-03 KIAA E-02 CBLB E-01 KCNT E-01 ATP1A E-01 MED23 9 NA CBFB E-08 TBL1XR E-06 FOXA E-02 NEK E-02 MYB E-02 SEMA5A E-02 ATN E-02 PIWIL E-02 UNC5D E-01 WNK3 8 NA ZFP36L E-04 PTPN E-03 WNT7A E-02 DCAF4L E-02 NLRC E-02 UBC E-01 TLR E-01 CPXM E-01 PPEF E-01 ZNF E-01 DSPP E-01 DGKG 7 NA PCDHGB6 7 NA ERBB2 7 NA GPS E-04 OR2G E-03 CHD1L E-01 BAI E-01 DALRD E-01 PRKG E-01 KCNB2 6 NA ITPKB 6 NA CTNND2 6 NA MST1P9 6 NA NPAS4 6 NA WSCD2 6 NA DENND1B 6 NA MLLT10 6 NA CDKN1B E-03 GPR E-02 LOC E-01 SRPR E-01 OR5I E-01 C2orf E-01 OR2T E-01 PIP4K2C E-01 PIWIL2 5 NA RGS7 5 NA GSPT1 5 NA PPP1R3A 5 NA CD5L 5 NA PRRX1 5 NA OR6A E-02 HIST1H2BC E-02 SEPT E-02 C12orf E-02 HIST1H3B E-02 OR2L E-02 HIST1H1C E-02 FAM166A E-01 HLF E-01 KRAS E-01 C8orf E-01 BID E-01 HLA-A E-01 GIMAP E-01 LYSMD E-01 ZNF266 4 NA GP1BA 4 NA HLA-DRB1 4 NA OR8H2 4 NA PIP5K1C 4 NA COPS3 4 NA C4orf40 4 NA TRIM6-TRIM34 4 NA TCGA et al., Nature, 2012 (PMID: )
7 Significantly Mutated Gene List according to mrna-subtype Basal-like (n=93) Mutated FDR TP TTN 24 NA USH2A E-02 PIK3CA E-07 FAT3 8 NA FLG 8 NA MLL3 6 NA MUC5B 5 NA KIAA E-01 SPTA1 4 NA RB E-02 AFF2 4 NA PREX2 4 NA RPGR E-01 BAI E-01 UNC5D 4 NA CACNA1E 3 NA PCDH19 3 NA CTNND2 3 NA PRKG2 3 NA OR5I E-01 TRIM6- TRIM E-02 ZNF841 3 NA HLF E-02 NPDC E-01 HER2-enriched (n=57) Mutated FDR TP PIK3CA E+00 TTN 7 NA FLG 5 NA ATP1A E-02 USH2A 4 NA MLL3 4 NA RYR2 4 NA PIK3R E-02 KIAA NA SRPR E-03 SPTA1 3 NA AFF2 3 NA PREX2 3 NA CACNA1E 3 NA ARID1A 3 NA CBLB 3 NA CDH1 3 NA PTPN22 3 NA ZNF E-01 KCNT2 2 NA CPXM1 2 NA C12orf36 2 NA C8orf31 2 NA PTPRD 2 NA FAM47C 2 NA OR2G3 2 NA SF3B1 2 NA ARMCX E-02 MAP3K1 2 NA RUNX1 2 NA ERBB2 2 NA Luminal A (n=225) Mutated FDR PIK3CA E+00 GATA E+00 MAP3K E+00 TP E+00 TTN 26 NA CDH E+00 MLL E-11 MAP2K E+00 RUNX E+00 NCOR E-09 PTEN E-11 CTCF E-06 AKT E-09 SPEN 8 NA SF3B E-05 USH2A 7 NA TBX E-05 NF1 6 NA RYR2 6 NA ATN E-03 MED E-02 CBFB E-05 TBL1XR E-03 FLG 5 NA FOXA E-02 NEK E-03 WNT7A E-03 MYB E-02 PCDH19 5 NA DGKG E-02 PIK3R E-01 GPS E-03 PTPRD 4 NA ATM 4 NA PPEF1 4 NA ARID1A 4 NA UNC5D 4 NA ITPKB E-01 GP1BA E-02 Luminal B (n=126) Mutated FDR PIK3CA E+00 TP E+00 GATA E+00 TTN 19 NA RYR E-01 FAT3 8 NA MLL E-01 MAP3K E-07 CDH E-03 PTEN E-07 TBX E-04 KCNB E-03 NF1 5 NA PTPRD E-02 FAM47C 5 NA WNK3 5 NA SEMA5A 5 NA MLLT E-02 USH2A 4 NA FLG 4 NA PIK3R1 4 NA ATM 4 NA PIWIL1 4 NA MUC5B 4 NA ZFP36L E-02 SPTA1 4 NA NLRC4 4 NA PCDHGB6 4 NA RB1 4 NA NPAS E-01 PIWIL E-01 GSPT E-02 MAP2K E-01 RUNX E-01 NCOR1 3 NA AKT1 3 NA ARID1A 3 NA AFF2 3 NA MST1P9 3 NA CBLB 3 NA KCNT2 3 NA PTPN22 3 NA DCAF4L2 3 NA PREX2 3 NA KIAA NA PRRX1 3 NA OR2G E-02 COPS E-02 CCDC E-01 CRTAP E-01 AGAP E-01 TCGA et al., Nature, 2012 (PMID: )
8 MAP3K1 (39) MAP2K4 (21) ~12% mutually exclusive TCGA et al., Nature, 2012 (PMID: )
9 PIK3CA Mutations TCGA et al., Nature, 2012 (PMID: )
10 Mutation of GATA3 in Human Breast Tumors Usary et al., Oncogene 46, (2004) (PMID: )
11 54 GATA3 mutant tumors TCGA et al., Nature, 2012 (PMID: )
12 A class of GATA3 mutation reprograms the breast cancer transcriptional network through gain and loss of function Mokoti Takaku et al. (Paul Wade Lab at NIEHS), Nature Com., In Press (2018) GATA3 mutations compiled from METABRIC, TCGA Nature 2012 & Nat Commun 2016
13 A class of GATA3 mutation reprograms the breast cancer transcriptional network through gain and loss of function Mokoti Takaku et al. (Paul Wade Lab at NIEHS), Nature Com., In Press (2018)
14 Comprehensive molecular portraits of human breast tumours. TCGA et al., Nature, 2012 (PMID: ) 466 Invasive Breast Carcinomas (348 common to all 6 platforms) 1. Whole Exome (all exons) capture and sequencing of tumor and normal (somatic coding sequence mutations and germline variation) WashU (n=507) 2. Gene expression analysis (Agilent microarrays and mrna-sequencing) - UNC 3. MicroRNA expression analysis (Illumina sequencing) - UBC 4. Tumor DNA copy number analysis (AFFY 6.0 microarrays and sequencing) - Broad 5. Tumor DNA methylation analysis (Illumina methylation arrays transitioning to sequencing)- USC 6. Reverse Phase Protein Arrays to detect proteins and phospho-proteins - MDACC 7. Integrated computational analyses (Genome Data Analysis Centers)- 6 Universities/sites 14
15 Unsupervised Hierarchical Cluster of DNA methylation data analyzed for 802 tumors using 574 methylation probes (5 subtypes) TCGA et al., Nature, 2012 (PMID: )
16 Comprehensive molecular portraits of human breast tumours. TCGA et al., Nature, 2012 (PMID: ) 466 Invasive Breast Carcinomas (348 common to all 6 platforms) 1. Whole Exome (all exons) capture and sequencing of tumor and normal (somatic coding sequence mutations and germline variation) WashU (n=507) 2. Gene expression analysis (Agilent microarrays and mrna-sequencing) - UNC 3. MicroRNA expression analysis (Illumina sequencing) - UBC 4. Tumor DNA copy number analysis (AFFY 6.0 microarrays and sequencing) - Broad 5. Tumor DNA methylation analysis (Illumina methylation arrays transitioning to sequencing)- USC 6. Reverse Phase Protein Arrays to detect proteins and phospho-proteins - MDACC 7. Integrated computational analyses (Genome Data Analysis Centers)- 6 Universities/sites 16
17 Hierarchical cluster analysis of 402 breast tumors using 171 proteins/phospho-proteins (7 subtypes) TCGA et al., Nature, 2012 (PMID: )
18 Comprehensive molecular portraits of human breast tumours. TCGA et al., Nature, 2012 (PMID: ) 466 Invasive Breast Carcinomas (348 common to all 6 platforms) 1. Whole Exome (all exons) capture and sequencing of tumor and normal (somatic coding sequence mutations and germline variation) WashU (n=507) 2. Gene expression analysis (Agilent microarrays and mrna-sequencing) - UNC 3. MicroRNA expression analysis (Illumina sequencing) - UBC 4. Tumor DNA copy number analysis (AFFY 6.0 microarrays and sequencing) - Broad 5. Tumor DNA methylation analysis (Illumina methylation arrays transitioning to sequencing)- USC 6. Reverse Phase Protein Arrays to detect proteins and phospho-proteins - MDACC 7. Integrated computational analyses (Genome Data Analysis Centers)- 6 Universities/sites 18
19 TCGA Breast Tumor RB Pathway Integrated Analysis Luminal A Luminal B HER2E Basal-like TCGA et al., Nature, 2012 (PMID: )
20 mrna microrna Protein/RPPA DNA Copy Number DNA Methylation Mutations/Exomes TCGA et al., Nature, 2012 (PMID: )
21 Cluster of Clusters (Consensus Clustering of 5 platform classification systems) TCGA et al., Nature, 2012 (PMID: )
22 Normal Breast Luminal B Luminal A Claudin-low HER2-enriched Basal-like MEMo Pathway Analyses of Basal-like Tumors. Giovanni Ciriello and others from MSKCC (Sanders Lab) if BRCA1 germline and somatic, and BRCA2 germline and somatic, deleterious variants are added up, then ~20% of Basal-like tumors have alterations in BRCA1/2
23 TCGA MEMo Comparison of Breast Luminal vs. Breast Basal-like vs. Serous Ovarian TCGA et al., Nature, 2012 (PMID: )
24 TCGA DNA Copy Number Comparison of Breast tumors vs. Serous Ovarian tumors TCGA et al., Nature, 2012 (PMID: )
25 TCGA Pan Cancer Study (12 tumor types representing 3527 specimens) Tumor Type # Samples GBM AML 173 AML Lung Adenocarcinoma Lung Squamous Ovarian Endometrial Katie Hoadley Head & Neck Breast Kidney Clear Cell Colon Rectum Bladder Bladder 122 Breast 845 Colon 190 Endometrial 370 GBM 168 Head & Neck 303 Kidney Clear Cell 480 Lung Adeno 355 Lung Squamous 259 Ovarian Serous 265 Rectum 72 Hoadley et al., Cell, (PMID: )
26 mrna microrna Protein/RPPA DNA Copy Number DNA Methylation Mutations/Exomes (-/+) tissue mut class Myeloid TP53-related PIK3-related Eclectic DNA-damage VHL-related Hoadley et al., Cell, (PMID: )
27 Consensus Clustering to define how many groups/subtypes are present within the 12 tumor types (3527 samples) when using all genomic platforms at once 11 Cluster of Cluster Assignment (COCA) subtypes are seen Hoadley et al., Cell, (PMID: )
28 Hoadley et al., Cell, (PMID: )
29 12 Tissue of Origin Sites Translate into 11 Genomic COCA Subtypes Bladder Head & Neck Rectum Lung Adeno Lung Squam Breast Renal Endometrial Colon Ovary GBM AML LUADenriched Squamous -like Breast Luminal (includes all HER2+ and all Non-basal TNBCs) Breast Basal-like Renal Endo Rectum & Colon 131/139 Basal-like tumors are in this group and nothing else Bladder Ovary GBM AML Hoadley et al., Cell, (PMID: )
30 TCGA Cluster Analysis of 10,000 tumors x 5000 genes 33 tumor types studied including breast (n=1100), bladder, colon, rectum, head & neck, gastric, lung squamous & adenocarcinoma, melanoma, renal clear cell & chromophobe, ovarian, glioblastoma, prostate, endometrial, thyroid, pancreas, testicular and others Luminal Basal-like
31 HOW MANY ETIOLOGICAL SUBTYPES OF BREAST CANCER: TWO, THREE, OR MORE? William F. Anderson, Philip S. Rosenberg, Aleix Prat, Charles M. Perou, and Mark E. Sherman. JNCI, (2014). PMID: Clemmesen s Hook (Br. J. Radiol, 1948) n=2000
32 Invasive Lobular Carcinoma (ILC) 10-15% Invasive Ductal Carcinoma (IDC) 70-75% Ciriello, Gatza et al., Cell PMID:
33 Comprehensive Molecular Portraits of Invasive Lobular Breast Cancer Ciriello, Gatza et al., Cell 2015 (PMID: ) 817 breast cancer samples with genomic data from: RNAseq, Exomes, DNA Copy Number, Methylation, micrornas, and RPPA (~200 proteins) Andy Beck (Harvard) and TCGA Breast Pathology Working Group Ciriello, Gatza et al., Cell PMID:
34 Lobular versus Ductal genomic alterations (within only Luminal A) Ductal Lum A (% altered samples) Lobular LumA (% altered samples) Ciriello, Gatza et al., Cell PMID:
35 Lobular versus Ductal genomic alterations (within only Luminal A) Ductal Lum A (% altered samples) ILC: 5% IDC: 20% ILC: 9% IDC: 2% Lobular LumA (% altered samples) Ciriello, Gatza et al., Cell PMID:
36 FOXA1 mutations in breast cancer All Lobular FOXA1 mutations fall within the Fork-head domain Ciriello, Gatza et al., Cell PMID:
37 FOXA1 mutations in breast cancer FOXA1 mutations correlates with increased FOXA1 mrna expression FOXA1 mutations PAM50 FOXA1 ESR1 ILC tumor only FOXA1 mrna expression correlates with decreased DNA methylation of FOXA1 binding sites FOXA1 binding sites Most variable probes Ciriello, Gatza et al., Cell PMID:
38 Identification of Molecular Subtypes of Invasive Lobular Cancers Reactive-like Immune-related Proliferative Ciriello, Gatza et al., Cell PMID:
39 Comparison of Breast Cancer Molecular Features and Survival by African and European Ancestry in The Cancer Genome Atlas. Huo, Hu, et al., JAMA Oncology, 2017 (PMID: )
40 Comparison of Breast Cancer Molecular Features and Survival by African and European Ancestry in The Cancer Genome Atlas. Huo, Hu, et al., JAMA Oncology, 2017 (PMID: ) subtype while most molecular differences were eliminated after adjusting for intrinsic subtype, the study found 16 DNA methylation probes, 4 DNA copy number segments, 1 protein, and 142 genes that were differentially expressed
41 TCGA Breast Cancer Summary 1. Comprehensive molecular portraits of human breast tumours. TCGA et al., Nature, 2012 (PMID: ); multi-platform analysis identifies 4 major subtypes of breast cancers 2. The molecular diversity of Luminal A breast tumors. Ciriello et al., BCRT, 2013 (PMID: ); identified 4 subtypes within Luminal A based on DNA copy number changes 3. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Hoadley et al., Cell, 2014 (PMID: ); showed that amongst 3500 tumors from 12 diverse tumor types, breast cancer is 2 diseases (basal and luminal) 4. Comprehensive Molecular Portraits of Invasive Lobular Breast Cancer. Ciriello et al., Cell (PMID: ); identified many unique genomic features of lobulars, and 3 subtypes 5. DNA defects, epigenetics, and gene expression in cancer-adjacent breast: a study from The Cancer Genome Atlas. Troester et al., NPJ Breast Cancer, 2016 (PMID: ); a molecular study of normal breast tissues adjacent to tumors identified occult tumor in many normals 6. The molecular basis of breast cancer pathological phenotypes. Heng et al., J Pathology, 2017 (PMID: ); genomic and genetic correlates to multiple pathology features including tubule formation and mitotic index 7. Comparison of Breast Cancer Molecular Features and Survival by African and European Ancestry in The Cancer Genome Atlas. Huo et al., JAMA Oncol, 2017 (PMID: ); molecular comparison of tumors from African Americans versus Whites shows little molecular differences once race associated subtype frequency % are accounted for 41
42 TCGA is over, what is next? 1. Its not over, we are working on the Special Histologies Paper, which includes the full set of 1101 tumors with ~10 different histological subtypes 2. Need to do similar multi-platform work on the metastatic setting. The AURORA project in the USA and EU is focused on this goal 3. Need to do similar multi-platform work on the DCIS and earlier lesion setting. In USA there is a call for new grants for the Pre-Cancer Atlas Project 4. Need to do more work with proteins. In USA we have the Clinical Proteomic Tumor Analysis Consortium (CPTAC), which for breast cancer is lead by Matthew Ellis 5. Need to focus on data integration and using multiple data types together as tools for discovery, and as the possible biomarker(s) 42
43 TCGA Breast Cancer Analysis Working Group ( ) 43 University of North Carolina Chuck Perou (co-chair) Katie Hoadley Joel Parker Michael Gatza University of California Santa Cruz Buck Institute Chris Benz Christina Yau Sam Ng Ted Goldstein Kyle Ellrott Charlie Vaske Josh Stuart University of Southern California Peter Laird Swapna Mahurkar Simeen Malik Dan Weisenberger Nationwide Children s Hospital Jay Bowen Julie Gastier-Foster The University of Texas MD Anderson Cancer Center Roel Verhaak Rehan Akbani Nancy Shih Gordon Mills Memorial Sloan-Kettering Cancer Center Niki Schultz Giovanni Ciriello Ethan Cerami Arthur Goldberg Caitlin Byrne Anders Jacobsen Tari King Chris Sander National Cancer Institute Chunhua Yan John Demchok Laura Dillon Peter Fielding Margi Sheth Peter Good Jacqueline Palchik Heidi Sofia Kenna Shaw Baylor College of Medicine Chad Creighton Xiaosong Wang British Columbia Cancer Agency Andy Chu Elizabeth Chun Andy Mungall Gordon Robertson Dominik Stoll Broad Institute Andrew Cherniack Harvard Medical School Terrence Wu Yonghong Xiao Institute for Systems Biology Sheila Reynolds Ilya Shmulevich Lawrence Berkeley National Laboratory Paul Spellman Mayo Clinic Jim Ingle Windber Research Institute Hai Hu Richard Mural The Genome Institute at Washington University Matthew Ellis (co-chair) Li Ding Lucinda Fulton Daniel Koboldt Elaine Mardis
The Cancer Genome Atlas Pan-cancer analysis Katherine A. Hoadley
 The Cancer Genome Atlas Pan-cancer analysis Katherine A. Hoadley Department of Genetics Lineberger Comprehensive Cancer Center The University of North Carolina at Chapel Hill What is TCGA? The Cancer Genome
The Cancer Genome Atlas Pan-cancer analysis Katherine A. Hoadley Department of Genetics Lineberger Comprehensive Cancer Center The University of North Carolina at Chapel Hill What is TCGA? The Cancer Genome
Genomic landscape of breast cancer
 Genomic landscape of breast cancer Aleix Prat MD PhD Head of the Translational Genomics Group Attending Physician at the Breast Cancer Unit VHIO, Barcelona, Spain Major research advance for breast cancer
Genomic landscape of breast cancer Aleix Prat MD PhD Head of the Translational Genomics Group Attending Physician at the Breast Cancer Unit VHIO, Barcelona, Spain Major research advance for breast cancer
The Cancer Genome Atlas
 The Cancer Genome Atlas July 14, 2011 Kenna M. Shaw, Ph.D. Deputy Director The Cancer Genome Atlas Program TCGA: Core Objectives Launched in 2006 as a pilot and expanded in 2009, the goals of TCGA are
The Cancer Genome Atlas July 14, 2011 Kenna M. Shaw, Ph.D. Deputy Director The Cancer Genome Atlas Program TCGA: Core Objectives Launched in 2006 as a pilot and expanded in 2009, the goals of TCGA are
Perspectivas moleculares en la disección del cáncer de mama esporádico y hereditario
 Perspectivas moleculares en la disección del cáncer de mama esporádico y hereditario Aleix Prat, MD PhD Medical Oncology, Hospital Clínic Barcelona, Spain mrna microrna Protein DNA Copy Number DNA Methylation
Perspectivas moleculares en la disección del cáncer de mama esporádico y hereditario Aleix Prat, MD PhD Medical Oncology, Hospital Clínic Barcelona, Spain mrna microrna Protein DNA Copy Number DNA Methylation
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser
 Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
Comprehensive molecular portraits of human breast tumours
 ARTICLE doi:10.1038/nature11412 Comprehensive molecular portraits of human breast tumours The Cancer Genome Atlas Network We analysed primary breast cancers by genomic DNA copy number arrays, DNA methylation,
ARTICLE doi:10.1038/nature11412 Comprehensive molecular portraits of human breast tumours The Cancer Genome Atlas Network We analysed primary breast cancers by genomic DNA copy number arrays, DNA methylation,
The Cancer Genome Atlas & International Cancer Genome Consortium
 The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Transcriptional Profiles from Paired Normal Samples Offer Complementary Information on Cancer Patient Survival -- Evidence from TCGA Pan-Cancer Data
 Transcriptional Profiles from Paired Normal Samples Offer Complementary Information on Cancer Patient Survival -- Evidence from TCGA Pan-Cancer Data Supplementary Materials Xiu Huang, David Stern, and
Transcriptional Profiles from Paired Normal Samples Offer Complementary Information on Cancer Patient Survival -- Evidence from TCGA Pan-Cancer Data Supplementary Materials Xiu Huang, David Stern, and
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
Molecular Classification of Triple-Negative Breast Cancer (TNBC) Aleix Prat, MD
 Molecular Classification of Triple-Negative Breast Cancer (TNBC) Emerging genomic and clinical data Aleix Prat, MD Postdoctoral Research Associate at Perou Lab Lineberger Comprehensive Cancer Center University
Molecular Classification of Triple-Negative Breast Cancer (TNBC) Emerging genomic and clinical data Aleix Prat, MD Postdoctoral Research Associate at Perou Lab Lineberger Comprehensive Cancer Center University
ncounter Assay Automated Process Immobilize and align reporter for image collecting and barcode counting ncounter Prep Station
 ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
ncounter Assay ncounter Prep Station Automated Process Hybridize Reporter to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes Immobilize and
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
ncounter Assay Automated Process Capture & Reporter Probes Bind reporter to surface Remove excess reporters Hybridize CodeSet to RNA
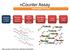 ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
ncounter Assay Automated Process Hybridize CodeSet to RNA Remove excess reporters Bind reporter to surface Immobilize and align reporter Image surface Count codes mrna Capture & Reporter Probes slides
Supplementary Figure 1. Copy Number Alterations TP53 Mutation Type. C-class TP53 WT. TP53 mut. Nature Genetics: doi: /ng.
 Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplemental Information. Integrated Genomic Analysis of the Ubiquitin. Pathway across Cancer Types
 Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Kinome Profiling: The Potential in ER-Negative Patients. Charles M. Perou, Ph.D. Departments of Genetics and Pathology
 Kinome Profiling: The Potential in ER-Negative Patients Charles M. Perou, Ph.D. Departments of Genetics and Pathology Lineberger Comprehensive Cancer Center University of North Carolina Chapel Hill, North
Kinome Profiling: The Potential in ER-Negative Patients Charles M. Perou, Ph.D. Departments of Genetics and Pathology Lineberger Comprehensive Cancer Center University of North Carolina Chapel Hill, North
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Biomarker development in the era of precision medicine. Bei Li, Interdisciplinary Technical Journal Club
 Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Next generation histopathological diagnosis for precision medicine in solid cancers
 Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making
 Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles
 On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles Ying-Wooi Wan 1,2,4, Claire M. Mach 2,3, Genevera I. Allen 1,7,8, Matthew L. Anderson 2,4,5 *, Zhandong Liu 1,5,6,7 * 1 Departments of Pediatrics
On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles Ying-Wooi Wan 1,2,4, Claire M. Mach 2,3, Genevera I. Allen 1,7,8, Matthew L. Anderson 2,4,5 *, Zhandong Liu 1,5,6,7 * 1 Departments of Pediatrics
Immune Characteriza/on of Basal- like Bladder Cancer
 Immune Characteriza/on of - like Bladder Cancer William Y. Kim, M.D. Departments of Medicine and Gene2cs Lineberger Comprehensive Cancer Center University of North Carolina Chapel Hill, NC 5 year survival
Immune Characteriza/on of - like Bladder Cancer William Y. Kim, M.D. Departments of Medicine and Gene2cs Lineberger Comprehensive Cancer Center University of North Carolina Chapel Hill, NC 5 year survival
Genomic tests to personalize therapy of metastatic breast cancers. Fabrice ANDRE Gustave Roussy Villejuif, France
 Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
Clinical Grade Genomic Profiling: The Time Has Come
 Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
List of Available TMAs in the PRN
 TMA RPCI_BrainCa01 RPCI_BrCa03 RPCI_BrCa04 RPCI_BrCa05 RPCI_BrCa0 RPCI_BrCa07 RPCI_BrCa08 RPCI_BrCa15 RPCI_BrCa1 RPCI_BrCa17 RPCI_BrCa18 RPCI_BrCa19 RPCI_BrCa20 RPCI_BrCa21 RPCI_BrCa24 RPCI_BrCa25 RPCI_BrCa2
TMA RPCI_BrainCa01 RPCI_BrCa03 RPCI_BrCa04 RPCI_BrCa05 RPCI_BrCa0 RPCI_BrCa07 RPCI_BrCa08 RPCI_BrCa15 RPCI_BrCa1 RPCI_BrCa17 RPCI_BrCa18 RPCI_BrCa19 RPCI_BrCa20 RPCI_BrCa21 RPCI_BrCa24 RPCI_BrCa25 RPCI_BrCa2
10/15/2012. Biologic Subtypes of TNBC. Topics. Topics. Histopathology Molecular pathology Clinical relevance
 Biologic Subtypes of TNBC Andrea L. Richardson M.D. Ph.D. Brigham and Women s Hospital Dana-Farber Cancer Institute Harvard Medical School Boston, MA Topics Histopathology Molecular pathology Clinical
Biologic Subtypes of TNBC Andrea L. Richardson M.D. Ph.D. Brigham and Women s Hospital Dana-Farber Cancer Institute Harvard Medical School Boston, MA Topics Histopathology Molecular pathology Clinical
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Analyses across Six Cancer Types Identify Basal-like Breast Cancer as a Unique Molecular Entity
 Genomic Analyses across Six Cancer Types Identify Basal-like Breast Cancer as a Unique Molecular Entity Aleix Prat, Barbara Adamo, Cheng Fan, Vicente Peg, Maria Vidal, Patricia Galván, Ana Vivancos, Paolo
Genomic Analyses across Six Cancer Types Identify Basal-like Breast Cancer as a Unique Molecular Entity Aleix Prat, Barbara Adamo, Cheng Fan, Vicente Peg, Maria Vidal, Patricia Galván, Ana Vivancos, Paolo
New breast cancer classification: traditional pathology and molecular subtypes. Dr. PN Mainwaring Centre for Personalised NanoMedicine
 New breast cancer classification: traditional pathology and molecular subtypes Dr. PN Mainwaring Centre for Personalised NanoMedicine AIBN@UQ Outline Traditional Pathology Haematoxylin & Eosin Descriptors;
New breast cancer classification: traditional pathology and molecular subtypes Dr. PN Mainwaring Centre for Personalised NanoMedicine AIBN@UQ Outline Traditional Pathology Haematoxylin & Eosin Descriptors;
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
New Drug development and Personalized Therapy in The Era of Molecular Medicine
 New Drug development and Personalized Therapy in The Era of Molecular Medicine Ramesh K. Ramanathan MD Virginia G. Piper Cancer Center Translational Genomics Research Institute Scottsdale, AZ Clinical
New Drug development and Personalized Therapy in The Era of Molecular Medicine Ramesh K. Ramanathan MD Virginia G. Piper Cancer Center Translational Genomics Research Institute Scottsdale, AZ Clinical
Genetically Engineered Murine Models of RCC William Y. Kim, M.D.
 Genetically Engineered Murine Models of RCC William Y. Kim, M.D. Co-director, Mouse Phase 1 Unit (MP1U) Departments of Medicine, Genetics, and Urology Lineberger Comprehensive Cancer Center University
Genetically Engineered Murine Models of RCC William Y. Kim, M.D. Co-director, Mouse Phase 1 Unit (MP1U) Departments of Medicine, Genetics, and Urology Lineberger Comprehensive Cancer Center University
Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change
 Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change Charles Matthew Quick, M.D. Associate Professor of Pathology Director of Gynecologic Pathology University of Arkansas for Medical Sciences
Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change Charles Matthew Quick, M.D. Associate Professor of Pathology Director of Gynecologic Pathology University of Arkansas for Medical Sciences
CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing.
 Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Contemporary Classification of Breast Cancer
 Contemporary Classification of Breast Cancer Laura C. Collins, M.D. Vice Chair of Anatomic Pathology Professor of Pathology Beth Israel Deaconess Medical Center and Harvard Medical School Boston, MA Outline
Contemporary Classification of Breast Cancer Laura C. Collins, M.D. Vice Chair of Anatomic Pathology Professor of Pathology Beth Israel Deaconess Medical Center and Harvard Medical School Boston, MA Outline
IPATIMUP Translational Research Unit Matching IPATIMUP s Expertise with Company s Research/Clinical Strategy
 IPATIMUP Translational Research Unit Matching IPATIMUP s Expertise with Company s Research/Clinical Strategy Innovative Projects Questions & Needs André Albergaria INDUSTRY Pharma R&D Clinical Research
IPATIMUP Translational Research Unit Matching IPATIMUP s Expertise with Company s Research/Clinical Strategy Innovative Projects Questions & Needs André Albergaria INDUSTRY Pharma R&D Clinical Research
Multigene Panel Testing for Hereditary Cancer Risk
 Multigene Panel Testing for Hereditary Cancer Risk Dana Zakalik, M.D. Director, Nancy and James Grosfeld Cancer Genetics Center Professor, OUWB Medical School MCC Annual Meeting November 4, 2015 Outline
Multigene Panel Testing for Hereditary Cancer Risk Dana Zakalik, M.D. Director, Nancy and James Grosfeld Cancer Genetics Center Professor, OUWB Medical School MCC Annual Meeting November 4, 2015 Outline
NeoTYPE Cancer Profiles
 NeoTYPE Cancer Profiles Multimethod Analysis of 25+ Hematologic Diseases and Solid Tumors Anatomic Pathology FISH Molecular The next generation of diagnostic, prognostic, and therapeutic assessment NeoTYPE
NeoTYPE Cancer Profiles Multimethod Analysis of 25+ Hematologic Diseases and Solid Tumors Anatomic Pathology FISH Molecular The next generation of diagnostic, prognostic, and therapeutic assessment NeoTYPE
NeoTYPE Cancer Profiles
 NeoTYPE Cancer Profiles 30+ Multimethod Assays for Hematologic Diseases and Solid Tumors Molecular FISH Anatomic Pathology The next generation of diagnostic, prognostic, and therapeutic assessment What
NeoTYPE Cancer Profiles 30+ Multimethod Assays for Hematologic Diseases and Solid Tumors Molecular FISH Anatomic Pathology The next generation of diagnostic, prognostic, and therapeutic assessment What
Breast cancer classification: beyond the intrinsic molecular subtypes
 Breast cancer classification: beyond the intrinsic molecular subtypes Britta Weigelt, PhD Signal Transduction Laboratory CRUK London Research Institute Summary Breast cancer heterogeneity Molecular classification
Breast cancer classification: beyond the intrinsic molecular subtypes Britta Weigelt, PhD Signal Transduction Laboratory CRUK London Research Institute Summary Breast cancer heterogeneity Molecular classification
Discovery Dataset. PD Liver Luminal B/ Her-2+ Letrozole. PD Supraclavicular Lymph node. PD Supraclavicular Lymph node Luminal B.
 Discovery Dataset 11T pt1cpn2am1(liver) 2009 2010 Liver / Her-2+ 2011 Death Letrozole CHT pt1cpn0(sn)m0 Supraclavicular Lymph node Death 12T 2003 2006 2006 2008 Anastrozole Local RT+Examestane Fulvestrant
Discovery Dataset 11T pt1cpn2am1(liver) 2009 2010 Liver / Her-2+ 2011 Death Letrozole CHT pt1cpn0(sn)m0 Supraclavicular Lymph node Death 12T 2003 2006 2006 2008 Anastrozole Local RT+Examestane Fulvestrant
Molecular classification of breast cancer implications for pathologists. Sarah E Pinder
 Molecular classification of breast cancer implications for pathologists Sarah E Pinder Courtesy of CW Elston Histological types Breast Cancer Special Types 17 morphological special types 25-30% of all
Molecular classification of breast cancer implications for pathologists Sarah E Pinder Courtesy of CW Elston Histological types Breast Cancer Special Types 17 morphological special types 25-30% of all
Advances in Brain Tumor Research: Leveraging BIG data for BIG discoveries
 Advances in Brain Tumor Research: Leveraging BIG data for BIG discoveries Jill Barnholtz-Sloan, PhD Associate Professor & Associate Director for Bioinformatics and Translational Informatics jsb42@case.edu
Advances in Brain Tumor Research: Leveraging BIG data for BIG discoveries Jill Barnholtz-Sloan, PhD Associate Professor & Associate Director for Bioinformatics and Translational Informatics jsb42@case.edu
The Role of Genetics in Ovarian Cancer Screening. Dawn DeLozier, Ph.D. Medical Geneticist, SAMC Associate Professor UCSF-Fresno
 The Role of Genetics in Ovarian Cancer Screening Dawn DeLozier, Ph.D. Medical Geneticist, SAMC Associate Professor UCSF-Fresno How Much Breast and Ovarian Cancer Is Hereditary? 15%-20% 5%-10% Breast Cancer
The Role of Genetics in Ovarian Cancer Screening Dawn DeLozier, Ph.D. Medical Geneticist, SAMC Associate Professor UCSF-Fresno How Much Breast and Ovarian Cancer Is Hereditary? 15%-20% 5%-10% Breast Cancer
USE OF CLUSTER ANALYSIS AS TRANSLATIONAL PHARMACOGENOMICS TOOL FOR BREAST CANCER GUIDED THERAPY. Ngozi Nwana
 USE OF CLUSTER ANALYSIS AS TRANSLATIONAL PHARMACOGENOMICS TOOL FOR BREAST CANCER GUIDED THERAPY By Ngozi Nwana A Dissertation Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor
USE OF CLUSTER ANALYSIS AS TRANSLATIONAL PHARMACOGENOMICS TOOL FOR BREAST CANCER GUIDED THERAPY By Ngozi Nwana A Dissertation Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor
The Center for PERSONALIZED DIAGNOSTICS
 The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
Insights from Sequencing the Melanoma Exome
 Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies
 Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
MolEcular Taxonomy of BReast cancer International Consortium (METABRIC)
 PERSPECTIVE 1 LARGE SCALE DATASET EXAMPLES MolEcular Taxonomy of BReast cancer International Consortium (METABRIC) BC Cancer Agency, Vancouver Samuel Aparicio, PhD FRCPath Nan and Lorraine Robertson Chair
PERSPECTIVE 1 LARGE SCALE DATASET EXAMPLES MolEcular Taxonomy of BReast cancer International Consortium (METABRIC) BC Cancer Agency, Vancouver Samuel Aparicio, PhD FRCPath Nan and Lorraine Robertson Chair
Breast cancer: IHC classification. Mogens Vyberg Professor of Clinical Pathology Director of NordiQC Aalborg University Hospital, Aalborg, Denmark
 Breast cancer: IHC classification Mogens Vyberg Professor of Clinical Pathology Director of NordiQC Aalborg University Hospital, Aalborg, Denmark http://upload.wikimedia.org/wikipedia/commons/1/1a/breast.svg
Breast cancer: IHC classification Mogens Vyberg Professor of Clinical Pathology Director of NordiQC Aalborg University Hospital, Aalborg, Denmark http://upload.wikimedia.org/wikipedia/commons/1/1a/breast.svg
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Examining Genetics and Genomics of Acute Myeloid Leukemia in 2017
 Examining Genetics and Genomics of Acute Myeloid Leukemia in 2017 Elli Papaemmanuil, PhD Memorial Sloan Kettering Cancer Center New York, New York, United States Today s Talk Cancer genome introduction
Examining Genetics and Genomics of Acute Myeloid Leukemia in 2017 Elli Papaemmanuil, PhD Memorial Sloan Kettering Cancer Center New York, New York, United States Today s Talk Cancer genome introduction
University of Pittsburgh Cancer Institute UPMC CancerCenter. Uma Chandran, MSIS, PhD /21/13
 University of Pittsburgh Cancer Institute UPMC CancerCenter Uma Chandran, MSIS, PhD chandran@pitt.edu 412-648-9326 2/21/13 University of Pittsburgh Cancer Institute Founded in 1985 Director Nancy Davidson,
University of Pittsburgh Cancer Institute UPMC CancerCenter Uma Chandran, MSIS, PhD chandran@pitt.edu 412-648-9326 2/21/13 University of Pittsburgh Cancer Institute Founded in 1985 Director Nancy Davidson,
Corporate Medical Policy
 Corporate Medical Policy Molecular Panel Testing of Cancers to Identify Targeted Therapies File Name: Origination: Last CAP Review: Next CAP Review: Last Review: molecular_panel_testing_of_cancers_to_identify_targeted_therapies
Corporate Medical Policy Molecular Panel Testing of Cancers to Identify Targeted Therapies File Name: Origination: Last CAP Review: Next CAP Review: Last Review: molecular_panel_testing_of_cancers_to_identify_targeted_therapies
Emerging Targets for Squamous Cell NSCLC. Ramaswamy Govindan M.D. Alvin J Siteman Cancer Center Washington University School of Medicine St Louis
 Emerging Targets for Squamous Cell NSCLC Ramaswamy Govindan M.D. Alvin J Siteman Cancer Center Washington University School of Medicine St Louis Structural variants Translocations Fusions Inversion Copy
Emerging Targets for Squamous Cell NSCLC Ramaswamy Govindan M.D. Alvin J Siteman Cancer Center Washington University School of Medicine St Louis Structural variants Translocations Fusions Inversion Copy
Sample Metrics. Allele Frequency (%) Read Depth Ploidy. Gene CDS Effect Protein Effect. LN Metastasis Tumor Purity Computational Pathology 80% 60%
 Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Comprehensive Analyses of Circulating Cell- Free Tumor DNA
 Comprehensive Analyses of Circulating Cell- Free Tumor DNA Boston, MA June 28th, 2016 Derek Murphy, Ph.D. Scientist, Research and Development Personal Genome Diagnostics Acquisition of Somatic Alterations
Comprehensive Analyses of Circulating Cell- Free Tumor DNA Boston, MA June 28th, 2016 Derek Murphy, Ph.D. Scientist, Research and Development Personal Genome Diagnostics Acquisition of Somatic Alterations
Clinical Grade Biomarkers in the Genomic Era Observations & Challenges
 Clinical Grade Biomarkers in the Genomic Era Observations & Challenges IOM Committee on Policy Issues in the Clinical Development & Use of Biomarkers for Molecularly Targeted Therapies March 31-April 1,
Clinical Grade Biomarkers in the Genomic Era Observations & Challenges IOM Committee on Policy Issues in the Clinical Development & Use of Biomarkers for Molecularly Targeted Therapies March 31-April 1,
Biomarkers in Imunotherapy: RNA Signatures as predictive biomarker
 Biomarkers in Imunotherapy: RNA Signatures as predictive biomarker Joan Carles, MD PhD Director GU, CNS and Sarcoma Program Department of Medical Oncology Vall d'hebron University Hospital Outline Introduction
Biomarkers in Imunotherapy: RNA Signatures as predictive biomarker Joan Carles, MD PhD Director GU, CNS and Sarcoma Program Department of Medical Oncology Vall d'hebron University Hospital Outline Introduction
Secuenciación masiva: papel en la toma de decisiones
 Secuenciación masiva: papel en la toma de decisiones Cancer is a Genetic Disease Development of cancer is driven by the acquisition of somatic genetic alterations: Nonsynonymous point mutations: missense.
Secuenciación masiva: papel en la toma de decisiones Cancer is a Genetic Disease Development of cancer is driven by the acquisition of somatic genetic alterations: Nonsynonymous point mutations: missense.
Predicting outcome in metastatic breast cancer
 Predicting outcome in metastatic breast cancer Aleix Prat, MD, PhD Medical Oncology Department Translational Genomics and Targeted Therapeutics in Solid Tumors Monday, 15 th January, Manchester, UK Disclosures
Predicting outcome in metastatic breast cancer Aleix Prat, MD, PhD Medical Oncology Department Translational Genomics and Targeted Therapeutics in Solid Tumors Monday, 15 th January, Manchester, UK Disclosures
Supplementary Figure 1: LUMP Leukocytes unmethylabon to infer tumor purity
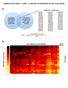 Supplementary Figure 1: LUMP Leukocytes unmethylabon to infer tumor purity A Consistently unmethylated sites (30%) in 21 cancer types 174,696
Supplementary Figure 1: LUMP Leukocytes unmethylabon to infer tumor purity A Consistently unmethylated sites (30%) in 21 cancer types 174,696
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
How will new biomarkers change bladder cancer management? John Kelly Professor of Uro-Oncology UCL
 How will new biomarkers change bladder cancer management? John Kelly Professor of Uro-Oncology UCL What do we mean by biomarker? a measurable indicator of the severity or presence of some disease state.
How will new biomarkers change bladder cancer management? John Kelly Professor of Uro-Oncology UCL What do we mean by biomarker? a measurable indicator of the severity or presence of some disease state.
Adenocarcinoma of the pancreas
 Adenocarcinoma of the pancreas SEMINARS IN DIAGNOSTIC PATHOLOGY 31 (2014) 443 451 Ralph H.Hruban, MD, David S. Klimstra, MD Paola Parente Anatomia Patologica Casa Sollievo della Sofferenza San Giovanni
Adenocarcinoma of the pancreas SEMINARS IN DIAGNOSTIC PATHOLOGY 31 (2014) 443 451 Ralph H.Hruban, MD, David S. Klimstra, MD Paola Parente Anatomia Patologica Casa Sollievo della Sofferenza San Giovanni
Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma
 Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma Nathaniel L Jones 1, Joanne Xiu 2, Sandeep K. Reddy 2, Ana I. Tergas 1, William
Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma Nathaniel L Jones 1, Joanne Xiu 2, Sandeep K. Reddy 2, Ana I. Tergas 1, William
Cancer develops as a result of the accumulation of somatic
 Cancer-mutation network and the number and specificity of driver mutations Jaime Iranzo a,1, Iñigo Martincorena b, and Eugene V. Koonin a,1 a National Center for Biotechnology Information, National Library
Cancer-mutation network and the number and specificity of driver mutations Jaime Iranzo a,1, Iñigo Martincorena b, and Eugene V. Koonin a,1 a National Center for Biotechnology Information, National Library
ARTICLE. Mutational landscape and significance across12majorcancertypes. OPEN doi: /nature12634
 ARTICLE OPEN doi:1.138/nature12634 Mutational landscape and significance across12majorcancertypes Cyriac Kandoth 1 *, Michael D. McLellan 1 *, Fabio Vandin 2, Kai Ye 1,3, Beifang Niu 1, Charles Lu 1, Mingchao
ARTICLE OPEN doi:1.138/nature12634 Mutational landscape and significance across12majorcancertypes Cyriac Kandoth 1 *, Michael D. McLellan 1 *, Fabio Vandin 2, Kai Ye 1,3, Beifang Niu 1, Charles Lu 1, Mingchao
Clasificación Molecular del Cáncer de Próstata. JM Piulats
 Clasificación Molecular del Cáncer de Próstata JM Piulats Introduction The Gleason score is the major method for prostate cancer tissue grading and the most important prognostic factor in this disease.
Clasificación Molecular del Cáncer de Próstata JM Piulats Introduction The Gleason score is the major method for prostate cancer tissue grading and the most important prognostic factor in this disease.
CHARLES M. PEROU. The University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA
 The Oncologist Molecular Stratification of Triple-Negative Breast Cancers CHARLES M. PEROU The University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA Key Words. Triple-negative breast
The Oncologist Molecular Stratification of Triple-Negative Breast Cancers CHARLES M. PEROU The University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA Key Words. Triple-negative breast
EGFR: fundamenteel en klinisch
 EGFR: fundamenteel en klinisch Guido Lammering MAASTRO Clinic Maastricht, NL What is EGFR?? The EGFR some facts 1186 amino acids 170 kda Expressed by all cells of epithelial origin Increased activation
EGFR: fundamenteel en klinisch Guido Lammering MAASTRO Clinic Maastricht, NL What is EGFR?? The EGFR some facts 1186 amino acids 170 kda Expressed by all cells of epithelial origin Increased activation
Simultaneous Identification of Multiple Driver Pathways in Cancer
 Simultaneous Identification of Multiple Driver Pathways in Cancer Mark D. M. Leiserson 1, Dima Blokh 2, Roded Sharan 2., Benjamin J. Raphael 1. * 1 Department of Computer Science and Center for Computational
Simultaneous Identification of Multiple Driver Pathways in Cancer Mark D. M. Leiserson 1, Dima Blokh 2, Roded Sharan 2., Benjamin J. Raphael 1. * 1 Department of Computer Science and Center for Computational
DATA STANDARDS AND QUALITY CONTROL MEMORANDUM DSQC #
 DATA STANDARDS AND QUALITY CONTROL MEMORANDUM DSQC #2006-01 CATEGORY: CLARIFICATION SUBJECT: RESCINDMENT - DSQC MEMORANDUM 2002-08 Coding Complex Morphologic Diagnoses (revised 8/02) EFFECTIVE: For Cases
DATA STANDARDS AND QUALITY CONTROL MEMORANDUM DSQC #2006-01 CATEGORY: CLARIFICATION SUBJECT: RESCINDMENT - DSQC MEMORANDUM 2002-08 Coding Complex Morphologic Diagnoses (revised 8/02) EFFECTIVE: For Cases
Package AIMS. June 29, 2018
 Type Package Package AIMS June 29, 2018 Title AIMS : Absolute Assignment of Breast Cancer Intrinsic Molecular Subtype Version 1.12.0 Date 2014-06-25 Description This package contains the AIMS implementation.
Type Package Package AIMS June 29, 2018 Title AIMS : Absolute Assignment of Breast Cancer Intrinsic Molecular Subtype Version 1.12.0 Date 2014-06-25 Description This package contains the AIMS implementation.
Histological Type. Morphological and Molecular Typing of breast Cancer. Nottingham Tenovus Primary Breast Cancer Study. Survival (%) Ian Ellis
 Morphological and Molecular Typing of breast Cancer Ian Ellis Molecular Medical Sciences, University of Nottingham Department of Histopathology, Nottingham University Hospitals NHS Trust Histological Type
Morphological and Molecular Typing of breast Cancer Ian Ellis Molecular Medical Sciences, University of Nottingham Department of Histopathology, Nottingham University Hospitals NHS Trust Histological Type
Cancer Somatic Mut. Cancer Germline Mut. Symbol Name GeneID Chr Chr Band
 The Cancer Gene Census is a catalogue of genes for which mutations have been causally implicated in cancer. Currently, more than 1% of all human genes are implicated via mutation in cancer. Approximately
The Cancer Gene Census is a catalogue of genes for which mutations have been causally implicated in cancer. Currently, more than 1% of all human genes are implicated via mutation in cancer. Approximately
Triple Negative Breast Cancer
 Triple Negative Breast Cancer Prof. Dr. Pornchai O-charoenrat Division of Head-Neck & Breast Surgery Department of Surgery Faculty of Medicine Siriraj Hospital Breast Cancer Classification Traditional
Triple Negative Breast Cancer Prof. Dr. Pornchai O-charoenrat Division of Head-Neck & Breast Surgery Department of Surgery Faculty of Medicine Siriraj Hospital Breast Cancer Classification Traditional
SUPPLEMENTARY INFORMATION
 Supplementary Information S1 Class I PI3K isoform alterations in cancer Alteration Type Cancer Type Frequency of Alteration Sample Size Range Class IA PIK3CA (p110α) Mutation Endometrial 10.3-53.0% 29-232
Supplementary Information S1 Class I PI3K isoform alterations in cancer Alteration Type Cancer Type Frequency of Alteration Sample Size Range Class IA PIK3CA (p110α) Mutation Endometrial 10.3-53.0% 29-232
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium
 Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Out-Patient Billing CPT Codes
 Out-Patient Billing CPT Codes Updated Date: August 3, 08 Client Billed Molecular Tests HPV DNA Tissue Testing 8764 No Medicare Billed - Molecular Tests NeoARRAY NeoARRAY SNP/Cytogenetic No 89 NeoLAB NeoLAB
Out-Patient Billing CPT Codes Updated Date: August 3, 08 Client Billed Molecular Tests HPV DNA Tissue Testing 8764 No Medicare Billed - Molecular Tests NeoARRAY NeoARRAY SNP/Cytogenetic No 89 NeoLAB NeoLAB
TCGA. The Cancer Genome Atlas
 TCGA The Cancer Genome Atlas TCGA: History and Goal History: Started in 2005 by the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) with $110 Million to catalogue
TCGA The Cancer Genome Atlas TCGA: History and Goal History: Started in 2005 by the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) with $110 Million to catalogue
Decipher Bladder Predicts Which Patients May Benefit from Neoadjuvant Chemotherapy Prior to Radical Cystectomy
 Decipher Bladder Predicts Which Patients May Benefit from Neoadjuvant Chemotherapy Prior to Cystectomy Contact the GenomeDx Customer Support Team 1.888.792.1601 (toll-free) customersupport@genomedx.com
Decipher Bladder Predicts Which Patients May Benefit from Neoadjuvant Chemotherapy Prior to Cystectomy Contact the GenomeDx Customer Support Team 1.888.792.1601 (toll-free) customersupport@genomedx.com
IntelliGENSM. Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community.
 IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
MEDICAL POLICY. SUBJECT: MOLECULAR PANEL TESTING OF CANCERS TO IDENTIFY TARGETED THERAPIES (Excluding NSCLC and CRC) EFFECTIVE DATE: 12/21/17
 MEDICAL POLICY SUBJECT: MOLECULAR PANEL TESTING OF PAGE: 1 OF: 5 If a product excludes coverage for a service, it is not covered, and medical policy criteria do not apply. If a commercial product, including
MEDICAL POLICY SUBJECT: MOLECULAR PANEL TESTING OF PAGE: 1 OF: 5 If a product excludes coverage for a service, it is not covered, and medical policy criteria do not apply. If a commercial product, including
Transcriptomic classification of genetically engineered mouse models of breast cancer identifies human subtype counterparts
 Pfefferle et al. Genome Biology 2013, 14:R125 RESEARCH Transcriptomic classification of genetically engineered mouse models of breast cancer identifies human subtype counterparts Open Access Adam D Pfefferle
Pfefferle et al. Genome Biology 2013, 14:R125 RESEARCH Transcriptomic classification of genetically engineered mouse models of breast cancer identifies human subtype counterparts Open Access Adam D Pfefferle
RNA preparation from extracted paraffin cores:
 Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester
 Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
Evolution of Pathology
 1 Traditional pathology Molecular pathology 2 Evolution of Pathology Gross Pathology Cellular Pathology Morphologic Pathology Molecular/Predictive Pathology Antonio Benivieni (1443-1502): First autopsy
1 Traditional pathology Molecular pathology 2 Evolution of Pathology Gross Pathology Cellular Pathology Morphologic Pathology Molecular/Predictive Pathology Antonio Benivieni (1443-1502): First autopsy
Supplementary Figures. Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated. for each lesion type.
 Supplementary Figures Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated for each lesion type. a b Supplementary Figure 2. Non-negative matrix factorization-derived
Supplementary Figures Supplementary Figure 1. Dinucleotide variant proportions. These are described and quantitated for each lesion type. a b Supplementary Figure 2. Non-negative matrix factorization-derived
Predictive biomarker profiling of > 1,900 sarcomas: Identification of potential novel treatment modalities
 Predictive biomarker profiling of > 1,900 sarcomas: Identification of potential novel treatment modalities Sujana Movva 1, Wenhsiang Wen 2, Wangjuh Chen 2, Sherri Z. Millis 2, Margaret von Mehren 1, Zoran
Predictive biomarker profiling of > 1,900 sarcomas: Identification of potential novel treatment modalities Sujana Movva 1, Wenhsiang Wen 2, Wangjuh Chen 2, Sherri Z. Millis 2, Margaret von Mehren 1, Zoran
Comparison of Triple Negative Breast Cancer between Asian and Western Data Sets
 2010 IEEE International Conference on Bioinformatics and Biomedicine Workshops Comparison of Triple Negative Breast Cancer between Asian and Western Data Sets Lee H. Chen Bioinformatics and Biostatistics
2010 IEEE International Conference on Bioinformatics and Biomedicine Workshops Comparison of Triple Negative Breast Cancer between Asian and Western Data Sets Lee H. Chen Bioinformatics and Biostatistics
Summary... 2 TRANSLATIONAL RESEARCH Tumour gene expression used to direct clinical decision-making for patients with advanced cancers...
 ESMO 2016 Congress 7-11 October, 2016 Copenhagen, Denmark Table of Contents Summary... 2 TRANSLATIONAL RESEARCH... 3 Tumour gene expression used to direct clinical decision-making for patients with advanced
ESMO 2016 Congress 7-11 October, 2016 Copenhagen, Denmark Table of Contents Summary... 2 TRANSLATIONAL RESEARCH... 3 Tumour gene expression used to direct clinical decision-making for patients with advanced
The use of diagnostic FFPE material in cancer epidemiology research
 The use of diagnostic FFPE material in cancer epidemiology research Neil O Callaghan Genetic Epidemiology Laboratory Department of Pathology The University of Melbourne www.pedigree.org.au Overview Who
The use of diagnostic FFPE material in cancer epidemiology research Neil O Callaghan Genetic Epidemiology Laboratory Department of Pathology The University of Melbourne www.pedigree.org.au Overview Who
Introduction to Systems Biology of Cancer Lecture 2
 Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers
 Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers Sung-Hou Kim University of California Berkeley, CA Global Bio Conference 2017 MFDS, Seoul, Korea June 28, 2017 Cancer
Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers Sung-Hou Kim University of California Berkeley, CA Global Bio Conference 2017 MFDS, Seoul, Korea June 28, 2017 Cancer
Relevancia práctica de la clasificación de subtipos intrínsecos en cáncer de mama Miguel Martín Instituto de Investigación Sanitaria Gregorio Marañón
 Relevancia práctica de la clasificación de subtipos intrínsecos en cáncer de mama Miguel Martín Instituto de Investigación Sanitaria Gregorio Marañón Universidad Complutense Madrid The new technologies
Relevancia práctica de la clasificación de subtipos intrínsecos en cáncer de mama Miguel Martín Instituto de Investigación Sanitaria Gregorio Marañón Universidad Complutense Madrid The new technologies
