Identification of Proteins Involved in Aldehyde Production for Bioluminescence
|
|
|
- Bruce Caldwell
- 5 years ago
- Views:
Transcription
1 JOURNAL OF BACTERIOLOGY, Aug. 1984, p /84/ $02.00/0 Copyright C) 1984, American Society for Microbiology Vol. 159, No. 2 In Vivo and In Vitro Acylation of Polypeptides in Vibrio harveyi: Identification of Proteins Involved in Aldehyde Production for Bioluminescence LEE A. WALL, DAVID M. BYERS, AND EDWARD A. MEIGHEN* McGill University, Department of Biochemistry, Montreal, Quebec H3G I Y6, Canada Received 14 February 1984/Accepted 21 May 1984 Incubation of soluble extracts from Vibrio harveyi with [3H]tetradecanoic acid (+ATP) resulted in the acylation of several polypeptides, including proteins with molecular masses near 20 kilodaltons (kda), and at least five polypeptides in the 30- to 60-kDa range. However, in growing cells pulse-labeled in vivo with [3HJtetradecanoic acid, only three of these polypeptides, with apparent molecular masses of 54, 42, and 32 kda, were specifically labeled. When extracts were acylated with [3H]tetradecanoyl coenzyme A, on the other hand, only the 32-kDa polypeptide was labeled. When luciferase-containing dark mutants of V. harveyi were investigated, acylated 32-kDa polypeptide was not detected in a fatty acid-stimulated mutant, whereas the 42- kda polypeptide appeared to be lacking in a mutant defective in aldehyde synthesis. Acylation of both of these polypeptides also increased specifically during induction of bioluminescence in V. harveyi. These results suggest that the role of the 32-kDa polypeptide is to supply free fatty acids, whereas the 42-kDa protein may be responsible for activation of fatty acids for their subsequent reduction to form the aldehyde substrates of the bioluminescent reaction. Light emission by bioluminescent marine bacteria is catalyzed by luciferase in a reaction which oxidizes reduced flavin mononucleotide and a long-chain aliphatic aldehyde to flavin mononucleotide and the corresponding fatty acid (2). Recently, we have reported a fatty acid reductase activity in extracts of the bioluminescent strain Photobacterilum phosphoreum which is capable of supplying the aldehyde substrates for the luminescent reaction of luciferase (6, 11). Purification of the P. phosphoreium fatty acid reductase has shown that it is a high-molecular-weight complex consisting of three different polypeptides (the three proteins of the P. phosphoreum fatty acid reductase complex have been designated 58,000 [58K], 50K, and 34K as previously described [14, 18], whereas proteins in Vibrio harveyi, to prevent confusion, have been designated by the apparent molecular masses on sodium dodecyl sulfate-polyacrylamide gel electrophoresis [SDS-PAGE] in kilodaltons [kda]): an acyl protein synthetase component (50K) which, in the presence of ATP, activates fatty acids to an acyl-sok intermediate, an acyl-coenzyme A (CoA) reductase subunit (58K) which catalyzes the reduction of acyl-50k or acyl-coa with NADPH, and a third polypeptide (34K) whose role in fatty acid utilization remains unknown (13, 14). In recent studies, it has also been shown that these polypeptides can be specifically identified in extracts of P. phosphoreum by acylation with [3H]tetradecanoic acid (+ATP), which labels the 50K and 34K polypeptides, or [3H]tetradecanoyl-CoA, which labels the 34K and 58K polypeptides (14, 18). In V. harveyi, the luminescent bacterium which has been studied in the greatest detail, the mechanism of the luciferase reaction has been well characterized, whereas the biosynthesis of the aldehyde substrate in this strain is only poorly understood. The existence of a luciferase-containing dark mutant of V. harveyi, which can be stimulated to produce light by the addition of exogenous fatty acids, has suggested that aldehyde biosynthesis may also proceed via fatty acid reduction (16). However, despite repeated attempts, fatty * Corresponding author. acid reductase activity has not yet been measured in vitro in lysates of V. harveyi, and therefore, the identity of the proteins involved remains unknown. In the present study, lysates of V. harveyi and dark mutants (whose luminescence is stimulated by fatty aidehyde) were acylated in vitro with [3H]tetradecanoic acid and [3H]tetradecanoyl-CoA in an attempt to establish which polypeptides may be involved in aldehyde biosynthesis. These techniques were supplemented by fatty acid labeling in vivo and resulted in the identification of polypeptides involved in supplying aldehyde for the bioluminescent system of V. harveyi. MATERIALS AND METHODS Materials. Acrylamide, N,N-methylenebisacrylamide, ATP, NADPH (type III), flavin mononucleotide, and,bmercaptoethanol were all obtained from Sigma Chemical Co. SDS-PAGE molecular mass standards (94 kda, phosphorylase b; 67 kda, bovine serum albumin; 43 kda, ovalbumin; 30 kda, carbonic anhydrase; 20 kda, soybean trypsin inhibitor; and 14 kda, a-lactalbumin) were obtained from Pharmacia Fine Chemicals. [3H]tetradecanoic acid (21 Ci/mmol) was prepared by New England Nuclear Corp. by reduction of cis-9-tetradecenoic acid with tritium gas in the presence of a 5% palladium-on-carbon catalyst. [3H]tetradecanoyl-CoA (1 Ci/mmol) was prepared from the radioactive fatty acid as previously described (13). Fatty acids and aldehydes were stored at -20 C as stock solutions in ethanol or isopropanol. Phosphate buffers were made by mixing the appropriate amounts of K2HPO4 and NaH2PO4. The bioluminescent bacterial strains used in these studies were P. phosphoreum NCMB 844 and V. harveyi B392. The dark mutants derived from V. harveyi by mutagenesis with nitrosoguanidine were a generous gift from J. W. Hastings (Harvard University). M17 is a dark mutant which can be stimulated to produce light by addition of exogenous aidehydes or fatty acids, as has been described previously (16). The acylation patterns of three different aldehyde synthesis mutants, which respond only to aldehyde and not fatty acids, 720
2 VOL. 159, 1984 were examined and found to be identical; hence, experiments on only one of these mutants (designated Aldl6) are presented in this report. Cell growth and lysis. P. phosphoreum was grown at 19 to 20 C in 3% NaCI complex media (11), whereas the V. harveyi strains were grown in 1% NaCI complex media at 26 to 27 C. Cultures (50 to 300 ml) were inoculated at an optical density of 660 nm (Aw) of 0.05 and grown in Erlenmeyer flasks in a reciprocatory shaker. The increase in A60 and in vivo luminescence of 1-ml samples was followed with growth. For M17 and Aldl6 cells, the maximum in vivo bioluminescence was measured after addition of 100,uM tetradecanoic acid or 100,M decanal, respectively. One light unit (LU) equals 6 x 109 quanta per s based on the standard of Hastings and Weber (4). For in vitro analyses, a constant amount of cells (A60 x volume [in milliliters] = 20) was isolated by centrifugation (15,000 x g, 15 min) and frozen at -20 C. P. phosphoreum cells were thawed and sonicated (20 s, three times) into 250 RI of 1 mm phosphate (ph 7.0) containing 1 mm EDTA and 1 mm 3-mercaptoethanol and then centrifuged to remove cellular debris. The supernatant was made 50 mm in phosphate (ph 7.0)-4-mercaptoethanol by addition of 1 M phosphate (ph 7.0) containing 1 M,B-mercaptoethanol. V. harveyi cells were extracted by sonication directly into 250 Rl of 50 mm phosphate (ph 7.0) containing 20 mm,-mercaptoethanol, followed by centrifugation. In vivo acylation. A constant amount of growing cells (A660 x volume [in milliliters] = 1.0) was removed and placed in a 1.5-mi Eppendorf tube with 2.5,uM [3H]tetradecanoic acid (21 Ci/mmol) added. The cells were incubated by shaking for 5 min at the proper growth temperature and then isolated immediately by centrifugation, washed twice with cold medium, and stored at -70 C. The labeled cells were extracted by sonication (20 s, three times) into 50 RI of 1 mm phosphate (ph 7.0) containing 1 mm f-mercaptoethanol. The cellular debris was removed by centrifugation, and after removal of a small aliquot for protein determination, the supernatant was diluted (1:1) with SDS-PAGE sample buffer containing 0.12 M Tris-hydrochloride (ph 6.8), 25% glycerol (vol/vol), 2.5% SDS, 0.35 M,B-mercaptoethanol, and 0.01% (wt/vol) bromophenol blue as tracking dye. The samples were then boiled for 5 min. In vitro acylation. For fatty acid acylation, 25 RI of the extracts was made up to a total volume of 100 RI containing 50 mm phosphate (ph 7.0), 5 mm ATP, 12.5,uM [3H]tetradecanoic acid (21 Ci/mmol), and 10 mm P-mercaptoethanol, with 10 mm MgSO4 for V. harveyi samples or without MgSO4 for P. phosphoreum samples. After 10 min, the reaction was stopped by mixing with an equal volume of SDS-PAGE sample buffer and boiling for 5 min. Proteins were acylated with acyl-coa by incubation with 10,uM [3H]tetradecanoyl-CoA (instead of tetradecanoic acid and ATP) for 30 s under the conditions described above. This short labeling period was used, as it resulted in the maximum incorporation into extracts from V. harveyi strains. Longer incubation times resulted in a decreased incorporation, apparently due to a loss of acyl-coa by high thioesterase activities in the extracts. Enzyme assays. The purification and the standard assay procedures for luciferase have been previously described (3, 7), with P. phosphoreum luciferase being measured in the presence of 0.01% dodecanal and V. harveyi luciferase being assayed with 0.002% dodecanal. Acyl-CoA reductase activity was measured by the luciferase-coupled assay in 50 mm phosphate (ph 7.0), containing ACYLATION OF POLYPEPTIDES IN V. HARVEYI mm,b-mercaptoethanol and 10 mm MgSO4 as previously described (13). Extracts were assayed in the presence of 10,uM NADPH and 5,uM tetradecanoyl-coa for 2 min, with 5,ug of P. phosphoreum luciferase added. Under these conditions, 1 LU corresponds to 4 pmol of aldehyde produced in the assay. SDS-PAGE. SDS-PAGE was performed by the system described by Laemmli (5) with 12% polyacrylamide resolving gels (8.5 cm) and 5% stacking gels. Gels were stained with Coomassie brilliant blue R (Aldrich Chemical Co.), destained, soaked in En3Hance (New England Nuclear Corp.), dried under vacuum, and fluorographed for 5 to 8 days with Kodak X-OMAT AR film as previously described (14). Protein assays. Protein concentrations were determined by the Bio-Rad dye binding assay (1), with bovine serum albumin used as a standard. RESULTS AND DISCUSSION We have previously demonstrated that the 50K (acyl protein synthetase) and the 34K subunits of the P. phosphoreum fatty acid reductase complex can be specifically labeled with [3H]tetradecanoic acid (+ATP) in cell extracts (14) with similar acylation patterns being observed in two other Photobacterium strains. In V. harveyi extracts, however, the relative intensity of acylated polypeptides and their molecular masses were found to be quite different from P. phosphoreum. The majority of the fatty acid label migrated at a position on SDS-PAGE (20 kda) which was close to that expected for the acyl-acyl carrier protein (acyl-acp), which migrates anomalously on this gel system (12). As polypeptides of higher molecular masses were only labeled to a low degree in extracts of V. harveyi, a clear relationship between the acylated polypeptides in V. harveyi and those of the fatty acid reductase complex in P. phosphoreum was not established. In the present investigation, experimental conditions were modified to enhance the labeling of V. harveyi proteins in vitro; these modifications included the omission of EDTA from the cell lysis buffer and the addition of Mg2+ during the incubation of the extracts with [3H]tetradecanoic acid (+ATP). The results are presented in Fig. la, lane 1. In contrast to the labeling pattern seen in P. phosphoreum (Fig. lb, lane 1), a number of acylated polypeptides are observed in V. harveyi extracts. In addition to the intensely labeled 20- kda bands, labeled polypeptides can be observed at 32 and 50 kda (two bands) as can lighter bands at 54 and 42 kda. The in vitro acylation with fatty acid of all these V. harveyi proteins was found to be ATP dependent, and in some experiments, the acylated 54-kDa band was found to be very low, or at an undetectable level, relative to the other labeled bands. The relatively large number of fatty acid-labeled polypeptides observed in extracts of V. harveyi could partially arise from nonspecific acylation under the in vitro conditions employed, thus raising the question of which of these acylated polypeptides, if any, are involved in aldehyde biosynthesis. Since V. harveyi cells are capable of taking up fatty acids from the growth medium (17),- growing cells were labeled with fatty acids to determine which of the acyl polypeptides are present in significant amounts in vivo. After pulse-labeling with [3H]tetradecanoic acid, SDS- PAGE of the soluble proteins, followed by fluorography, revealed essentially three labeled polypeptides (Fig. la, lane 2). These proteins corresponded in molecular mass to the 54-, 42-, and 32-kDa bands acylated in vitro with [3H]tetrade-
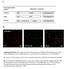
3 722 WALL, BYERS, AND MEIGHEN 54kDa- 5OkDa- 42 kda- - a m 32kDa- _oo.m 20kDa- Front- b -50K K FIG. 1. In vitro and in vivo acylation of V. harveyi and P. phosphoreiin. Cells were grown to near peak luminescence (A~(,,( 1.8, 400 LU/ml and A(,6) = 3.3, 3,000 LU/ml for V. Ihanievi and P. phosphorcien, respectively). In vivo and in vitro acylation followed by separation of the polypeptides by SDS-PAGE and fluorography were performed as described in the text. The fluorogram of the gel is presented. (a) V. hari'evi; each lane contains 50 p.g of protein. (b) P. phlosplhori-ei'n:1 each lane contains 25 p.g of protein. Lanes 1. in vitro acylation with V3H]tetradecanoic acid (-+ATP); lanes 2. in vivo acylation with 3Hltetradecanoic acid; lane 3. in vitro acylation with [3H ]tetradecanoyl-coa. canoic acid (+ATP) (Fig. la, lane 1), although the relative degree of labeling of the 32-kDa polypeptide was found to be much lower in vivo. Very minor in vivo-acylated bands near 50 kda were sometimes also detected; however, their level was always found to be low in comparison with the 54-, 42-, and 32-kDa acyl derivatives. The heavily acylated 20-kDa bands observed in vitro in V. harveyi extracts may be absent from samples labeled in vivo, but this conclusion must be viewed with caution since polypeptides in this region are only partially resolved from the large lipid band seen near the front of the gel in in vivo-labeled samples. An apparent lack of in vivo-labeled ACP derivatives in V. htiirvevi would be consistent with reports suggesting that Esclerichia coli does not activate exogenous fatty acids to acyl-acp in vivo (15), even though acyl-acp synthetase activity can be detected in E. coli extracts (10). Acylation of P. phosphoreum proteins was also investigated by labeling in vivo with [3H]tetradecanoic acid. The results (Fig. lb, lane 2) indicate that only 50K, the acylprotein synthetase subunit, is labeled in vivo with exogenous fatty acid in this bacteria. The absence of label associated with the 34K subunit suggests that the acylation of this polypeptide by [3H]tetradecanoic acid (+ATP) in vitro is J. BACTERIOL. different than its acylation in vivo under physiological conditions. Possible relationships between bioluminescence-specific proteins in P. pliospiworeiii and V. harieyi were further examined in vitro by acylation with [3H]tetradecanoyl-CoA. We have prevously shown that the 34K component and, to a lesser extent, the 58K (acyl-coa reductase) component of P. phosplioreiuml fatty acid reductase can be labeled with this substrate (18). Only the 32-kDa polypeptide was highly labeled when V. harlevi extracts were incubated with [3H]tetradecanoyl-CoA for short periods of time (Fig. la, lane 3). The strong, relatively specific labeling of the V. hanreyi 32-kDa polypeptide and the P. phosphoreiumw1 34K subunit by tetradecanoyl-coa (18) suggests that these two proteins may perform analogous roles in their respective bacterial strains. This conclusion is supported not only by their similarity in molecular weight, but also by the tendency of both proteins to be much more readily acylated in vitro than in vivo by fatty acid (Fig. 1). It is interesting in this regard that the most highly labeled polypeptides with tetradecanoyl-coa in a variety of other luminescent bacteria are also in the 30- to 35-kDa range (unpublished data). The bacterial bioluminescence system is growth dependent, as it is induced only at later stages of exponential growth (9). It is reasonable to expect that any enzyme activities associated with this system will be coinduced with in vivo luminescence, as has been demonstrated for luciferase (8) and for each of the fatty acid reductase components of P. phosphoreium (18). Thus, to establish a direct relationship between bioluminescence and acylated proteins in V. hari'evi, the in vivo acylation patterns from V. hanr'evi cells pulse-labeled before the onset of luminescence were compared with cells labeled near peak luminescence. In vivo acylation of the 54-, 42-, and 32-kDa polypeptides is significantly higher in induced cells (Fig. 2a). In vitro acylation with 3H-labeled fatty acid (Fig. 2b) also demonstrates induced acylation of the 42- and 32-kDa polypeptides, whereas labeling of the 20-kDa bands, as well as the 50-kDa polypeptides, appears to be growth independent. The increase in the 32-kDa polypeptide is also clearly seen by labeling extracts with [3H]tetradecanoyl-CoA (Fig. 2c). These results provide evidence that the three polypeptides which are acylated in vivo (54, 42, and 32 kda) are involved in V. harivevi bioluminescence and raise the question of what role these proteins play in this system. To investigate possible functions of the 54-, 42-, and 32- kda polypeptides, the acylation patterns of two different types of dark mutants (M17 and Aldl6) were investigated. Under normal growth conditions, the V. harvevi M17 mutant does not emit light, although it can be stimulated to luminesce by the addition of tetradecanoic acid or tetradecanal (16). Presumably, M17 possesses the ability to reduce exogenous fatty acids to the corresponding aldehydes, but it is incapable of aldehyde biosynthesis de novo. When cells of M17. which reached the same level as wild-type cells in terms of growth and extractable luciferase activity (Table 1), were incubated with [33H]tetradecanoic acid in vivo, both the 42- and 54-kDa polypeptides were acylated, although at a lower level than in wild-type cells (Fig. 3a). However, acylated 32-kDa polypeptide was not detected in the labeled M17 cells. Similarly, in vitro labeling of M17 extracts with [3H]tetradecanoic acid and ATP resulted in a very low level of acylation of the 32-kDa protein, whereas the labeling of all other polypeptides remained relatively constant with respect to wild type (Fig. 3b). The difference between wild-type and M17 strains is even more dramatically illustrated by labeling
4 VOL. 159, 1984 ACYLATION OF POLYPEPTIDES IN V. HARVEYI 723 a b C a b C 54kDa- 42kDa- 32 kda 54 kda- 42 k Da kda FIG. 2. Induction of acylation of polypeptides in V. harveyi. Cells of V. harveyi before induction of luminescence (Aw = 0.35, 0.5 LU/mI) or near peak luminescence (A6w = 1.5, 230 LU/ml) were labeled in vivo and in vitro. Protein from each sample (50,ug) was separated by SDS-PAGE, and the gel was fluorographed as described in the text. (a) In vivo or (b) in vitro labeling with [3H]tetradecanoic acid; (c) in vitro labeling with tetradecanoyl-coa. Lanes 1, preinduced cells; lanes 2, induced cells. with [3H]tetradecanoyl-CoA in vitro; the label associated with the 32-kDa polypeptide in wild-type extracts was absent in M17 (Fig. 3c). These results suggest that the 32-kDa polypeptide is either nonfunctional or absent in M17 cells and thus provide a clear link between this protein and bioluminescence in V. harveyi. Unlike M17, the Aldl6 dark mutant of V. harveyi is not capable of converting fatty acids to aldehydes and can therefore only be stimulated to luminesce if aldehyde is added exogenously. When Aldl6 cells (Table 1) were labeled with [3HJtetradecanoic acid in vivo, acylation of the 54- and 42-kDa polypeptides was extremely low, whereas the 32- kda protein was found to be acylated at a higher level than in wild-type cells (Fig. 3a). Fatty acid labeling of Aldl6 extracts in vitro revealed the absence of the acylated 42-kDa polypeptide only (Fig. 3b), whereas no significant difference in the [3HJtetradecanoyl-CoA labeling pattern was observed TABLE 1. Growth of V. hari'evi and dark mutants Inuvivos In vitro InvtoA reduc- Cell type A6w lumines- luciferase tasedumo cence (LU/mg) tase (pmol (U(LU/ m LUm) min-' mg-') Wild type M " Ald b "Maxirmum luminescence after addition of 100 FM tetradecanoic acid. b Maximum luminescence after addition of 100 FM decanal. w..._ ' FIG. 3. in vivo and in vitro acylation of dark mutants of V. harveyi. Wild-type, M17, and Aldl6 cells were all grown to equivalent levels as depicted in Table 1. The samples were labeled in vivo and in vitro, and 50,ug of protein was separated by SDS-PAGE as described in the text. The fluorogram of the SDS gel is presented. (a) In vivo or (b) in vitro acylation with [3H]tetradecanoic acid; (c) in vit,ro acylation with [3H]tetradecanoyl-CoA. Lanes 1, wild type; lanes 2, M17; lanes 3, Aldl6. between Aldl6 and wild-type extracts (Fig. 3c). Although it is not understood at present why the 54-kDa polypeptide in Aldi6 cells is labeled at wild-type levels in vitro, but not in vivo, the observations described above indicate that the 42- kda (and, perhaps, the 54-kDa) protein may be involved in the biosynthesis of long-chain aldehydes. Although the specific functions of these proteins cannot be directly assessed from the experiments described above, the phenotypic characteristics of the M17 and Aldl6 mutants do provide some clues regarding their roles in V. hars'evi bioluminescence. In M17 cells, the inability to provide the endogenous fatty acids required for aldehyde synthesis appears to be related to the lack of a functional 32-kDa polypeptide. A direct role of this protein in fatty acid synthesis can be ruled out since M17 cells grow as well as wild-type cells and both strains exhibit similar fatty acid synthetase activities in vitro (unpublished data). On the other hand, it is quite possible that this protein functions by supplying free fatty acids from an activated or storage form. This hypothesis could explain the increased acylation of the 32-kDa protein in the Aldl6 mutant in vivo (Pig. 3a), in which fatty acid conversion to aldehyde is blocked. Additional support for this idea comes from recent observations in our laboratory which indicate that the analogous 34K subunit purified from P. phosphoreum fatty acid reductase exhibits acyl esterase activity with a number of activated fatty acyl compounds (L. Carey, A. Rodriguez, and E. A. Meighen, J. Biol. Chem., in press). The function of the 42-kDa polypeptide in V. harv'eyi bioluminescence is probably more closely involved with the reduction of fatty acid to aldehyde, since both this activity and the acylated 42-kDa protein appear to be missing in Aldl6 cells. It is not likely that the 42-kDa protein corresponds to the acyl-coa reductase component of P. phosphoreum fatty acid reductase because we have recently
5 724 WALL, BYERS, AND MEIGHEN demonstrated that an aldehyde dehydrogenase, of higher molecular weight, is responsible for this activity in V. harveyi (D. M. Byers and E. A. Meighen, J. Biol. Chem., in press). In any case, it is illustrated in Table 1 that acyl- CoA reductase activity was similar in the wild-type and mutant strains. There is some evidence that the V. hars'eyi 42-kDa polypeptide may, in fact, be analogous to the acyl protein synthetase component (50K) of P. phosphoreium. Both proteins are labeled by [3HJtetradecanoic acid in vivo and in vitro, but not by [3H]tetradecanoyl-CoA (Fig. 1). Moreover, the increased in vivo acylation of the 42-kDa polypeptide in the wild type relative to M17 cells, in which the 32-kDa polypeptide is absent, could relate to the stimulation of P. phosphorelum 50K fatty acid acylation by the 34K subunit (14). The relationship between the aldehyde stimulatable mutants and the 32-, 42-, and 54-kDa acylated polypeptides is shown in the following pathway of aldehyde biosynthesis in V. harveyi: Mutant M17 Mutant Aldl6 Fatty acid precursor Fatty acid 32 kda 42 kda 54 kda? Aldehyde -. Fatty acid + light Luciferase Experiments are currently under way in our laboratory to further characterize the 42- and 32-kDa proteins, as well as the 54-kDa polypeptide, and their relationship to the fatty acid reductase reaction. ACKNOWLEDGMENTS This work was supported by a Medical Research Council Grant (MT-4314), a Medical Research Council Studentship (to L.A.W.), and a Medical Research Council Fellowship (to D.M.B.). We would like to thank J. W. Hastings (Harvard University) for supplying the V. harveyi mutants and Rose Szittner for technical assistance. LITERATURE CITED 1. Bradford, M. M A rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72: Dunn, D. K., G. A. Michalizyn, I. G. Bogacki, and E. A. Meighen Conversion of aldehyde to acid in the bioluminescent reaction. Biochemistry 12: J. BACTERIOL. 3. Gunsalus-Miguel, A., E. A. Meighen, M. Z. Nicoli, K. H. Nealson, and J. W. Hastings Purification and properties of bacterial luciferases. J. Biol. Chem. 247: Hastings, J. W., and G. Weber Total quantum flux of isotropic sources. J. Opt. Soc. Am. 53: Laemmli, U. K Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature (London) 227: Meighen, E. A Biosynthesis of aliphatic aldehydes for the bacterial bioluminescent reaction, stimulation by ATP and NADPH. Biochem. Biophys. Res. Commun. 87: Meighen, E. A., and i. Bartlet Complementation of subunits from different bacterial luciferases. J. Biol. Chem. 255: Michaliszyn, G. A., and E. A. Meighen Induced polypeptide synthesis during the development of bacterial bioluminescence. J. Biol. Chem. 251: Nealson, K. H., T. Platt, and J. W. Hastings Cellular control of the synthesis and activity of the bacterial luminescent system. J. Biol. Chem. 104: Ray, T. K., and J. E. Cronan, Jr Activation of long chain fatty acids with acyl carrier protein. Proc. Natl. Acad. Sci. U.S.A. 73: Riendeau, D., and E. Meighen Evidence for a fatty acid reductase catalyzing the synthesis of aldehydes for the bacterial bioluminescent reaction. J. Biol. Chem. 254: Rock, C. O., and J. E. Cronan, Jr Re-evaluation of the solution structure of acyl-carrier protein. J. Biol. Chem. 254: Rodriguez, A., D. Riendeau, and E. Meighen Purification of the acyl coenzyme A reductase component from a complex responsible for the reduction of fatty acids in bioluminescent bacteria. J. Biol. Chem. 258: Rodriguez, A., L. Wall, D. Riendeau, and E. Meighen Fatty acid acylation of proteins in bioluminescent bacteria. Biochemistry 22: Silbert, D. F., F. Ruch, and P. R. Vagelos Fatty acid replacement in a fatty acid auxotroph of Escherichia coli. J. Bacteriol. 95: Ulitzur, S., and J. W. Hastings Myristic acid stimulation of bacterial bioluminescence in 'aldehyde" mutants. Proc. Natl. Acad. Sci. U.S.A. 75: Ulitztar, S., and J. W. Hastings Reversible inhibition of bacterial bioluminescence by long chain fatty acids. Curr. Microbiol. 3: Wall, L., A. Rodriguez, and E. Meighen Differential acylation in vitro with tetradecanoyl-coa and tetradecanoic acid (+ATP) of three polypeptides shown to have induced synthesis in Photobacterium phosphoniirn. J. Biol. Chem. 259:
these authors to develop a bioassay for this compound.
 JOURNAL OF BACTERIOLOGY, JUIY 1989, p. 3866-3871 21-9193/89/73866-6$2./ Copyright ( 1989, American Society for Microbiology Vol. 171, No. 7 Inhibition of Vibrio harveyi Bioluminescence by Cerulenin: In
JOURNAL OF BACTERIOLOGY, JUIY 1989, p. 3866-3871 21-9193/89/73866-6$2./ Copyright ( 1989, American Society for Microbiology Vol. 171, No. 7 Inhibition of Vibrio harveyi Bioluminescence by Cerulenin: In
Control of Aldehyde Synthesis in the Luminous Bacterium Beneckea harveyi
 JOURNAL OF BACTERIOLOGY, Feb. 1979, p. 854-859 0021-9193/79/02-0854/06$02.00/0 Vol. 137, No. 2 Control of Aldehyde Synthesis in the Luminous Bacterium Beneckea harveyi S. ULITZUR' AND J. W. HASTINGS2*
JOURNAL OF BACTERIOLOGY, Feb. 1979, p. 854-859 0021-9193/79/02-0854/06$02.00/0 Vol. 137, No. 2 Control of Aldehyde Synthesis in the Luminous Bacterium Beneckea harveyi S. ULITZUR' AND J. W. HASTINGS2*
Elongation of Exogenous Fatty Acids by the Bioluminescent Bacterium Vibrio harveyi
 JOURNAL OF BACTERIOLOGY, Jan. 1989, p. 59-64 Vol. 171, No. 1 21-9193/89/159-6$2./ Copyright 1989, American Society for Microbiology Elongation of Exogenous Fatty Acids by the Bioluminescent Bacterium Vibrio
JOURNAL OF BACTERIOLOGY, Jan. 1989, p. 59-64 Vol. 171, No. 1 21-9193/89/159-6$2./ Copyright 1989, American Society for Microbiology Elongation of Exogenous Fatty Acids by the Bioluminescent Bacterium Vibrio
Activation To Form Acyl-Acyl Carrier Protein Intermediates and Lipid A in Vibrio harveyi
 JOURNAL OF BACrERIOLOGY, Jan. 1994, p. 77-83 0021-9193/94/$04.00+0 Copyright C) 1994, American Society for Microbiology Vol. 176, No. 1 Exogenous Myristic Acid Can Be Partially Degraded prior to Activation
JOURNAL OF BACrERIOLOGY, Jan. 1994, p. 77-83 0021-9193/94/$04.00+0 Copyright C) 1994, American Society for Microbiology Vol. 176, No. 1 Exogenous Myristic Acid Can Be Partially Degraded prior to Activation
Dental Research Institute, Faculty of Dentistry, University of Toronto, Toronto, Canada *For correspondence:
 Zymogram Assay for the Detection of Peptidoglycan Hydrolases in Streptococcus mutans Delphine Dufour and Céline M. Lévesque * Dental Research Institute, Faculty of Dentistry, University of Toronto, Toronto,
Zymogram Assay for the Detection of Peptidoglycan Hydrolases in Streptococcus mutans Delphine Dufour and Céline M. Lévesque * Dental Research Institute, Faculty of Dentistry, University of Toronto, Toronto,
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Purification and Characterization of Fatty Acyl-Acyl Carrier Protein Synthetase from Vibrio harveyi
 JOURNAL OF BACTERIOLOGY, Apr. 1993, p. 1865-187 21-9193/93/71865-6$2./ Copyright 1993, American Society for Microbiology Vol. 175, No. 7 Purification and Characterization of Fatty Acyl-Acyl Carrier Protein
JOURNAL OF BACTERIOLOGY, Apr. 1993, p. 1865-187 21-9193/93/71865-6$2./ Copyright 1993, American Society for Microbiology Vol. 175, No. 7 Purification and Characterization of Fatty Acyl-Acyl Carrier Protein
MW.SDS.70L and MW-SDS.200 Kits
 ~'A'.'.A'k'~ ~ ~ ':if';"7'~~'!11;~\ C HEM IC A I CQ P.O. ~X 14508,$T,LQV1S,MQ;, ~17,;U$A SDS MOLECULAR WEIGHT MARKERS IN A DISCONTINUOUS BUFFER July 1988 Technical Bulletin No. MWS-877L ORDER DIRECT: USA/Canada
~'A'.'.A'k'~ ~ ~ ':if';"7'~~'!11;~\ C HEM IC A I CQ P.O. ~X 14508,$T,LQV1S,MQ;, ~17,;U$A SDS MOLECULAR WEIGHT MARKERS IN A DISCONTINUOUS BUFFER July 1988 Technical Bulletin No. MWS-877L ORDER DIRECT: USA/Canada
Protein MultiColor Stable, Low Range
 Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
BIL 256 Cell and Molecular Biology Lab Spring, Tissue-Specific Isoenzymes
 BIL 256 Cell and Molecular Biology Lab Spring, 2007 Background Information Tissue-Specific Isoenzymes A. BIOCHEMISTRY The basic pattern of glucose oxidation is outlined in Figure 3-1. Glucose is split
BIL 256 Cell and Molecular Biology Lab Spring, 2007 Background Information Tissue-Specific Isoenzymes A. BIOCHEMISTRY The basic pattern of glucose oxidation is outlined in Figure 3-1. Glucose is split
Spore Formation Induced by Glycerol, Dimethyl Sulfoxide,
 JOURNAL OF BACTERIOLOGY, Dec. 1980, p. 1076-1082 0021-9193/80/12-1076/07$2.00/0 Vol. 144, No. 3 Patterns of Protein Production in Myxococcus xanthus During Spore Formation Induced by Glycerol, Dimethyl
JOURNAL OF BACTERIOLOGY, Dec. 1980, p. 1076-1082 0021-9193/80/12-1076/07$2.00/0 Vol. 144, No. 3 Patterns of Protein Production in Myxococcus xanthus During Spore Formation Induced by Glycerol, Dimethyl
Protocol for protein SDS PAGE and Transfer
 Protocol for protein SDS PAGE and Transfer According to Laemmli, (1970) Alaa El -Din Hamid Sayed, Alaa_h254@yahoo.com Serum Selection of a protein source cell cultures (bacteria, yeast, mammalian, etc.)
Protocol for protein SDS PAGE and Transfer According to Laemmli, (1970) Alaa El -Din Hamid Sayed, Alaa_h254@yahoo.com Serum Selection of a protein source cell cultures (bacteria, yeast, mammalian, etc.)
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
Acetyl-CoA or BC acyl-coa. Int1-ACP. FabH. FabF. Anti-FabF Platensimycin ACP. Fak
 Initiation Acetyl-CoA AccBCDA FabZ Int-ACP Elongation [N+] FabG Int1-ACP Acetyl-CoA or BC acyl-coa FabH FabD CoA Malonyl-ACP Malonyl CoA ACP Anti-FabI Triclosan AFN-15 FabI acyl-acp PlsX PlsY Pls C Lipoic
Initiation Acetyl-CoA AccBCDA FabZ Int-ACP Elongation [N+] FabG Int1-ACP Acetyl-CoA or BC acyl-coa FabH FabD CoA Malonyl-ACP Malonyl CoA ACP Anti-FabI Triclosan AFN-15 FabI acyl-acp PlsX PlsY Pls C Lipoic
SUPPLEMENTARY INFORMATION. Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was
 SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
Glycoprotein Synthesis by D-Glucosamine Hydrochloride
 JOURNAL OF VIROLOGY, Apr. 1974, p. 775-779 Copyright 0 1974 American Society for Microbiology Vol. 13, No. 4 Printed in U.S.A. Selective Inhibition of Newcastle Disease Virus-Induced Glycoprotein Synthesis
JOURNAL OF VIROLOGY, Apr. 1974, p. 775-779 Copyright 0 1974 American Society for Microbiology Vol. 13, No. 4 Printed in U.S.A. Selective Inhibition of Newcastle Disease Virus-Induced Glycoprotein Synthesis
Acetyl CoA Carboxylase: The Purified Transcarboxylase Component
 Proc. Nat. Acad. Sci. USA Vol. 68, No. 6, pp. 12591263, June 1971 Acetyl CoA Carboxylase: The Purified Transcarboxylase Component (acyl CoA binding/carboxylation/exchange reactions/biotin) ALFRED W. ALBERTS,
Proc. Nat. Acad. Sci. USA Vol. 68, No. 6, pp. 12591263, June 1971 Acetyl CoA Carboxylase: The Purified Transcarboxylase Component (acyl CoA binding/carboxylation/exchange reactions/biotin) ALFRED W. ALBERTS,
Williams Lab Recipes ANTIBIOTICS
 Williams Lab Recipes ANTIBIOTICS 1000x Ampicillin (sodium salt) 100mg/ml recipe 1. Measure out 1 g of Ampicillin tri hydrate 2. Add Milli-Q H2O to 10 ml 3. Add ~.1 g of NaOH pellets (half pellet or more
Williams Lab Recipes ANTIBIOTICS 1000x Ampicillin (sodium salt) 100mg/ml recipe 1. Measure out 1 g of Ampicillin tri hydrate 2. Add Milli-Q H2O to 10 ml 3. Add ~.1 g of NaOH pellets (half pellet or more
ELECTROPHORETIC STUDIES OF SONIC EXTRACTS OF PROTEUS VULGARIS
 ELECTROPHORETIC STUDIES OF SONIC EXTRACTS OF PROTEUS VULGARIS I. EFFECT OF GROWTH ENVIRONMENT ON ELECTROPHORETIC PATTERNS' SIDNEY D. RODENBERG Laboratory of Microbiology, Division of Biology, University
ELECTROPHORETIC STUDIES OF SONIC EXTRACTS OF PROTEUS VULGARIS I. EFFECT OF GROWTH ENVIRONMENT ON ELECTROPHORETIC PATTERNS' SIDNEY D. RODENBERG Laboratory of Microbiology, Division of Biology, University
Acetylene as a suicide substrate and active site probe for methane monooxygenase from Methylococcus capsulatus (Bath)
 FEMS Microbiology Letters 29 (1985) 105-109 105 Published by Elsevier FEM 02197 Acetylene as a suicide substrate and active site probe for methane monooxygenase from Methylococcus capsulatus (Bath) (Inhibitor
FEMS Microbiology Letters 29 (1985) 105-109 105 Published by Elsevier FEM 02197 Acetylene as a suicide substrate and active site probe for methane monooxygenase from Methylococcus capsulatus (Bath) (Inhibitor
Step 4. Step 4. Choose MW markers. Choose MW markers. Gel Electrophoresis of Proteins
 Gel Electrophoresis of Proteins Choose MW markers 16 Step 4 Gel Electrophoresis of Proteins A Practical Approach, Third Edition An overview of basic techniques to benefit any life scientist using electrophoretic
Gel Electrophoresis of Proteins Choose MW markers 16 Step 4 Gel Electrophoresis of Proteins A Practical Approach, Third Edition An overview of basic techniques to benefit any life scientist using electrophoretic
SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons
 LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
Student Number: To form the polar phase when adsorption chromatography was used.
 Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis.
 ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
Identification of NADPH-thioredoxin reductase system
 Proc. Nat. Acad. Sci. USA Vol. 72, No. 11, pp. 4233-4237, November 1975 Biochemistry Identification of NADPH-thioredoxin reductase system in Euglena gracilis* (ribonucleotide reduction) S. MUNAVALLIO,
Proc. Nat. Acad. Sci. USA Vol. 72, No. 11, pp. 4233-4237, November 1975 Biochemistry Identification of NADPH-thioredoxin reductase system in Euglena gracilis* (ribonucleotide reduction) S. MUNAVALLIO,
6. C-type cytochrome, soluble and membrane protein
 185 6. C-type cytochrome, soluble and membrane protein analysis of Rhodobacter sp SW2 and Rhodopseudomonas palustris TIE-1 ABSTRACT The ability to grown on Fe(II) is thought to be a primitive metabolism
185 6. C-type cytochrome, soluble and membrane protein analysis of Rhodobacter sp SW2 and Rhodopseudomonas palustris TIE-1 ABSTRACT The ability to grown on Fe(II) is thought to be a primitive metabolism
Kit for assay of thioredoxin
 FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Biochemistry: A Short Course
 Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples:
 Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Mengovirus Virions. growth (48-h cultures) were infected with a. cell at a density of 107 cells per ml of ABM42-
 JOURNAL OF VIROLOGY, Mar. 1977, p. 1256-1261 Copyright 1977 American Society for Microbiology Vol. 21, No. 3 Printed in U.S.A. Factors Affecting Composition and Thermostability of Mengovirus Virions CLIFFORD
JOURNAL OF VIROLOGY, Mar. 1977, p. 1256-1261 Copyright 1977 American Society for Microbiology Vol. 21, No. 3 Printed in U.S.A. Factors Affecting Composition and Thermostability of Mengovirus Virions CLIFFORD
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
HiPer Western Blotting Teaching Kit
 HiPer Western Blotting Teaching Kit Product Code: HTI009 Number of experiments that can be performed: 5/20 Duration of Experiment: ~ 2 days Day 1: 6-8 hours (SDS- PAGE and Electroblotting) Day 2: 3 hours
HiPer Western Blotting Teaching Kit Product Code: HTI009 Number of experiments that can be performed: 5/20 Duration of Experiment: ~ 2 days Day 1: 6-8 hours (SDS- PAGE and Electroblotting) Day 2: 3 hours
N-Glycosidase F Deglycosylation Kit
 For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
The Synthesis of Vitamin B, by some Mutant Strains of Escherichia coli
 597 MORRIS, J. G. (1959). J. gen. Mimobiol. 20, 5 974 The Synthesis of Vitamin B, by some Mutant Strains of Escherichia coli BY J. G. MORRIS Microbiology Unit, Department of Biochemistry, University of
597 MORRIS, J. G. (1959). J. gen. Mimobiol. 20, 5 974 The Synthesis of Vitamin B, by some Mutant Strains of Escherichia coli BY J. G. MORRIS Microbiology Unit, Department of Biochemistry, University of
Communication. Identification of Methionine N -Acetyltransferase from Saccharomyces cerevisiae
 Communication THE JOURNAL OP BIOLOGICAL CHEMISTRY Vol. 265, No. 7, Issue of March 5, pp. 3603-3606,lSSO 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U. S. A. Identification
Communication THE JOURNAL OP BIOLOGICAL CHEMISTRY Vol. 265, No. 7, Issue of March 5, pp. 3603-3606,lSSO 0 1990 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U. S. A. Identification
Isolation and Structural Characterization of Cap-Binding Proteins from Poliovirus-Infected HeLa Cells
 JOURNAL OF VIROLOGY, May 1985. p. 515-524 0022-538X/85/050515-10$02.00/0 Copyright C 1985, American Society for Microbiology Vol. 54, No. 2 Isolation and Structural Characterization of Cap-Binding Proteins
JOURNAL OF VIROLOGY, May 1985. p. 515-524 0022-538X/85/050515-10$02.00/0 Copyright C 1985, American Society for Microbiology Vol. 54, No. 2 Isolation and Structural Characterization of Cap-Binding Proteins
Luminescent platforms for monitoring changes in the solubility of amylin and huntingtin in living cells
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
MEK1 Assay Kit 1 Catalog # Lot # 16875
 MEK1 Assay Kit 1 Kit Components Assay Dilution Buffer (ADB), Catalog # 20-108. Three vials, each containing 1.0ml of assay dilution buffer (20mM MOPS, ph 7.2, 25mM ß-glycerol phosphate, 5mM EGTA, 1mM sodium
MEK1 Assay Kit 1 Kit Components Assay Dilution Buffer (ADB), Catalog # 20-108. Three vials, each containing 1.0ml of assay dilution buffer (20mM MOPS, ph 7.2, 25mM ß-glycerol phosphate, 5mM EGTA, 1mM sodium
Chap 3 Metabolism and Growth
 Chap 3 Metabolism and Growth I. Metabolism Definitions: Metabolism includes two parts: anabolism and catabolism Catabolism: Anabolism: Aerobic metabolism: catabolism anabolis m catabolis anabolis m Anaerobic
Chap 3 Metabolism and Growth I. Metabolism Definitions: Metabolism includes two parts: anabolism and catabolism Catabolism: Anabolism: Aerobic metabolism: catabolism anabolis m catabolis anabolis m Anaerobic
Lab Recipes. Destain 50% MeOH 40% milli-q water 10% Acetic Acid For 5 L total 2.5 liters Methanol 2 liters milli-q 0.5 liters Acetic Acid
 1% Blotto 1 liter TTBS (aka TBST) 50 g nonfat dry milk 10 g BSA (Albumin, Bovine) Spike with 2 of 5% Sodium Azide (in hood) Shake it up and put it in the fridge Coomassie Stain 50% MeOH 10% Acetic Acid
1% Blotto 1 liter TTBS (aka TBST) 50 g nonfat dry milk 10 g BSA (Albumin, Bovine) Spike with 2 of 5% Sodium Azide (in hood) Shake it up and put it in the fridge Coomassie Stain 50% MeOH 10% Acetic Acid
For the quantitative measurement of ATP Synthase Specific activity in samples from Human, Rat and Cow
 ab109716 ATP Synthase Specific Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of ATP Synthase Specific activity in samples from Human, Rat and Cow This product is for
ab109716 ATP Synthase Specific Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of ATP Synthase Specific activity in samples from Human, Rat and Cow This product is for
TECHNICAL BULLETIN. PhosDecor Fluorescent Phosphoprotein In-Gel Detection Kit. Catalog Number PDECOR Storage Temperature 2 8 C
 PhosDecor Fluorescent Phosphoprotein In-Gel Detection Kit Catalog Number PDECOR Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Phosphorylation is an important covalent posttranslational
PhosDecor Fluorescent Phosphoprotein In-Gel Detection Kit Catalog Number PDECOR Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Phosphorylation is an important covalent posttranslational
Lumino Firefly Luciferase Assay
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Lumino Firefly Luciferase Assay (Cat. # 786 1267, 786 1268) think proteins! think G-Biosciences
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Lumino Firefly Luciferase Assay (Cat. # 786 1267, 786 1268) think proteins! think G-Biosciences
For the rapid, sensitive and accurate quantification of Ras in various samples
 ab128504 Ras Assay Kit Instructions for Use For the rapid, sensitive and accurate quantification of Ras in various samples This product is for research use only and is not intended for diagnostic use.
ab128504 Ras Assay Kit Instructions for Use For the rapid, sensitive and accurate quantification of Ras in various samples This product is for research use only and is not intended for diagnostic use.
Keratin-like Proteins in Corneal and Conjunctival Epithelium are Different
 Keratin-like Proteins in Corneal and Conjunctival Epithelium are Different Shigeru Kinoshiro,* Judith Friend, Timothy C. Kiorpes, and Richard A. Thoft Using SDS polyacrylamide slab-gel electrophoresis,
Keratin-like Proteins in Corneal and Conjunctival Epithelium are Different Shigeru Kinoshiro,* Judith Friend, Timothy C. Kiorpes, and Richard A. Thoft Using SDS polyacrylamide slab-gel electrophoresis,
MORIMOTO LAB BUFFER AND SOLUTION RECIPES
 MORIMOTO LAB BUFFER AND SOLUTION RECIPES 30% Acrylamide 100ml 29g 2X Acyrlamide 1g N,N -methylenebisacrylamide Add ~300ml ddh 2 O. Heat to 37 C to dissolve chemicals. Adjust final volume to 500ml with
MORIMOTO LAB BUFFER AND SOLUTION RECIPES 30% Acrylamide 100ml 29g 2X Acyrlamide 1g N,N -methylenebisacrylamide Add ~300ml ddh 2 O. Heat to 37 C to dissolve chemicals. Adjust final volume to 500ml with
Cell Lysis Buffer. Catalog number: AR0103
 Cell Lysis Buffer Catalog number: AR0103 Boster s Cell Lysis Buffer is a ready-to-use Western blot related reagent solution used for efficient extraction of total soluble protein in nondenatured state
Cell Lysis Buffer Catalog number: AR0103 Boster s Cell Lysis Buffer is a ready-to-use Western blot related reagent solution used for efficient extraction of total soluble protein in nondenatured state
colorimetrically by the methylene blue method according to Fogo and manometrically. In the presence of excess sulfur the amount of oxygen taken up
 GLUTA THIONE AND SULFUR OXIDATION BY THIOBACILLUS THIOOXIDANS* BY ISAMU SUZUKI AND C. H. WERKMAN DEPARTMENT OF BACTERIOLOGY, IOWA STATE COLLEGE Communicated December 15, 1958 The ability of Thiobacillus
GLUTA THIONE AND SULFUR OXIDATION BY THIOBACILLUS THIOOXIDANS* BY ISAMU SUZUKI AND C. H. WERKMAN DEPARTMENT OF BACTERIOLOGY, IOWA STATE COLLEGE Communicated December 15, 1958 The ability of Thiobacillus
Expression constructs
 Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Total Phosphatidic Acid Assay Kit
 Product Manual Total Phosphatidic Acid Assay Kit Catalog Number MET- 5019 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Phosphatidic Acid (PA) is a critical precursor
Product Manual Total Phosphatidic Acid Assay Kit Catalog Number MET- 5019 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Phosphatidic Acid (PA) is a critical precursor
FUNCTION OF PYRIDOXAL PHOSPHATE: RESOLUTION AND PURIFICATION OF THE TRYPTOPHANASE ENZYME OF ESCHERICHIA COLI
 FUNCTION OF PYRIDOXAL PHOSPHATE: RESOLUTION AND PURIFICATION OF THE TRYPTOPHANASE ENZYME OF ESCHERICHIA COLI BY W. A. WOOD,* I. c. GUNSALUS, AND W. W. UMBREIT (From the Laboratory of Bacteriology, College
FUNCTION OF PYRIDOXAL PHOSPHATE: RESOLUTION AND PURIFICATION OF THE TRYPTOPHANASE ENZYME OF ESCHERICHIA COLI BY W. A. WOOD,* I. c. GUNSALUS, AND W. W. UMBREIT (From the Laboratory of Bacteriology, College
Recipes for Media and Solution Preparation SC-ura/Glucose Agar Dishes (20mL/dish, enough for 8 clones)
 Protocol: 300 ml Yeast culture preparation Equipment and Reagents needed: Autoclaved toothpicks Shaker Incubator set at 30 C Incubator set at 30 C 60 mm 2 sterile petri dishes Autoclaved glass test tubes
Protocol: 300 ml Yeast culture preparation Equipment and Reagents needed: Autoclaved toothpicks Shaker Incubator set at 30 C Incubator set at 30 C 60 mm 2 sterile petri dishes Autoclaved glass test tubes
Microbial Metabolism Quorum Sensing and Biofilms
 1 Microbial Metabolism Quorum Sensing and Biofilms Ching-Tsan Huang ( 黃慶璨 ) Office: Agronomy Hall, Room 111 Tel: (02) 33664454 E-mail: cthuang@ntu.edu.tw 2 Discovery of Cell-to-Cell Communication Stage
1 Microbial Metabolism Quorum Sensing and Biofilms Ching-Tsan Huang ( 黃慶璨 ) Office: Agronomy Hall, Room 111 Tel: (02) 33664454 E-mail: cthuang@ntu.edu.tw 2 Discovery of Cell-to-Cell Communication Stage
Very-Long Chain Fatty Acid Biosynthesis
 Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
Very-Long Chain Fatty Acid Biosynthesis Objectives: 1. Review information on the isolation of mutants deficient in VLCFA biosynthesis 2. Generate hypotheses to explain the absence of mutants with lesions
FEBS 1138 January Paul R. Buckland and Bernard Rees Smith
 Volume 166, number 1 FEBS 1138 January 1984 A structural comparison receptors by of guinea pig thyroid and fat TSH photoaffinity labelling Paul R. Buckland and Bernard Rees Smith Endocrine Immunology Unit,
Volume 166, number 1 FEBS 1138 January 1984 A structural comparison receptors by of guinea pig thyroid and fat TSH photoaffinity labelling Paul R. Buckland and Bernard Rees Smith Endocrine Immunology Unit,
Protocol for Western Blo
 Protocol for Western Blo ng SDS-PAGE separa on 1. Make appropriate percentage of separa on gel according to the MW of target proteins. Related recommenda ons and rou ne recipes of separa on/stacking gels
Protocol for Western Blo ng SDS-PAGE separa on 1. Make appropriate percentage of separa on gel according to the MW of target proteins. Related recommenda ons and rou ne recipes of separa on/stacking gels
Biosynthesis of Fatty Acids. By Dr.QUTAIBA A. QASIM
 Biosynthesis of Fatty Acids By Dr.QUTAIBA A. QASIM Fatty Acids Definition Fatty acids are comprised of hydrocarbon chains terminating with carboxylic acid groups. Fatty acids and their associated derivatives
Biosynthesis of Fatty Acids By Dr.QUTAIBA A. QASIM Fatty Acids Definition Fatty acids are comprised of hydrocarbon chains terminating with carboxylic acid groups. Fatty acids and their associated derivatives
Phospholipase D Activity of Gram-Negative Bacteria
 JOURNAL OF BACTERIOLOGY, Dec. 1975, p. 1148-1152 Copyright 1975 American Society for Microbiology Vol. 124, No. 3 Printed in U.S.A. Phospholipase D Activity of Gram-Negative Bacteria R. COLE AND P. PROULX*
JOURNAL OF BACTERIOLOGY, Dec. 1975, p. 1148-1152 Copyright 1975 American Society for Microbiology Vol. 124, No. 3 Printed in U.S.A. Phospholipase D Activity of Gram-Negative Bacteria R. COLE AND P. PROULX*
Protocol for purification of recombinant protein from 300 ml yeast culture
 Protocol for purification of recombinant protein from 300 ml yeast culture Equipment and reagents needed: Zirconia beads (0.5 mm diameter from BSP, Germany) Paint Shaker (at 4 C) Tube rotator for 15 ml
Protocol for purification of recombinant protein from 300 ml yeast culture Equipment and reagents needed: Zirconia beads (0.5 mm diameter from BSP, Germany) Paint Shaker (at 4 C) Tube rotator for 15 ml
Recombinant Trypsin, Animal Origin Free
 Recombinant Trypsin, Animal Origin Free PRODUCT INFORMATION: BioGenomics r-trypsin powder is ready to use, animal origin free optimized for cell culture applications. It is derived by r-dna technology.
Recombinant Trypsin, Animal Origin Free PRODUCT INFORMATION: BioGenomics r-trypsin powder is ready to use, animal origin free optimized for cell culture applications. It is derived by r-dna technology.
BCM 221 LECTURES OJEMEKELE O.
 BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
I mutants accumulate pyruvate when growing in the presence of isoleucine and
 THE iv-3 MUTANTS OF NEUROSPORA CRASSA 11. ACTIVITY OF ACETOHYDROXY ACID SYNTHETASE DINA F. CAROLINE, ROY W. HARDINGZ, HOMARE KUWANA3, T. SATYANARAYANA AND R.P. WAGNER4 Genetics Foundation, The University
THE iv-3 MUTANTS OF NEUROSPORA CRASSA 11. ACTIVITY OF ACETOHYDROXY ACID SYNTHETASE DINA F. CAROLINE, ROY W. HARDINGZ, HOMARE KUWANA3, T. SATYANARAYANA AND R.P. WAGNER4 Genetics Foundation, The University
ab Complex II Enzyme Activity Microplate Assay Kit
 ab109908 Complex II Enzyme Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of Complex II activity in samples from Human, Rat, Mouse and Cow This product is for research
ab109908 Complex II Enzyme Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of Complex II activity in samples from Human, Rat, Mouse and Cow This product is for research
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Conversion of green note aldehydes into alcohols by yeast alcohol dehydrogenase
 Conversion of green note aldehydes into alcohols by yeast alcohol dehydrogenase M.-L. Fauconnier 1, A. Mpambara 1, J. Delcarte 1, P. Jacques 2, P. Thonart 2 & M. Marlier 1 1 Unité de Chimie Générale et
Conversion of green note aldehydes into alcohols by yeast alcohol dehydrogenase M.-L. Fauconnier 1, A. Mpambara 1, J. Delcarte 1, P. Jacques 2, P. Thonart 2 & M. Marlier 1 1 Unité de Chimie Générale et
Annexure III SOLUTIONS AND REAGENTS
 Annexure III SOLUTIONS AND REAGENTS A. STOCK SOLUTIONS FOR DNA ISOLATION 0.5M Ethylene-diamine tetra acetic acid (EDTA) (ph=8.0) 1M Tris-Cl (ph=8.0) 5M NaCl solution Red cell lysis buffer (10X) White cell
Annexure III SOLUTIONS AND REAGENTS A. STOCK SOLUTIONS FOR DNA ISOLATION 0.5M Ethylene-diamine tetra acetic acid (EDTA) (ph=8.0) 1M Tris-Cl (ph=8.0) 5M NaCl solution Red cell lysis buffer (10X) White cell
Microbial Production of L-Threonine. Part III. Production by Methionine and Lysine Auxotrophs. Derived from ƒ -Amino-ƒÀ-hydroxyvaleric Acid Resistant
 I [Agr. Biol. Chem., Vol. 36, No. 7, p. 12091216, 1972] Microbial Production of L-Threonine Part III. Production by Methionine and Lysine Auxotrophs Derived from ƒ -Amino-ƒÀ-hydroxyvaleric Acid Resistant
I [Agr. Biol. Chem., Vol. 36, No. 7, p. 12091216, 1972] Microbial Production of L-Threonine Part III. Production by Methionine and Lysine Auxotrophs Derived from ƒ -Amino-ƒÀ-hydroxyvaleric Acid Resistant
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for
 Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
Chromatin IP (Isw2) Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles.
 Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
20S Proteasome Activity Assay Kit
 20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
Summary of fatty acid synthesis
 Lipid Metabolism, part 2 1 Summary of fatty acid synthesis 8 acetyl CoA + 14 NADPH + 14 H+ + 7 ATP palmitic acid (16:0) + 8 CoA + 14 NADP + + 7 ADP + 7 Pi + 7 H20 1. The major suppliers of NADPH for fatty
Lipid Metabolism, part 2 1 Summary of fatty acid synthesis 8 acetyl CoA + 14 NADPH + 14 H+ + 7 ATP palmitic acid (16:0) + 8 CoA + 14 NADP + + 7 ADP + 7 Pi + 7 H20 1. The major suppliers of NADPH for fatty
EFFECT OF CARBON SOURCES ON FORMATION OF a-amylase AND GLUCOAMYLASE BY
 J. Gen. App!. Microbiol,, 21, 51-59 (1975) EFFECT OF CARBON SOURCES ON FORMATION OF a-amylase AND GLUCOAMYLASE BY CLOSTRIDIUM ACETOBUTYLICUM BURT ENSLEY, JOHN J. McHUGH, AND LARRY L. BARTON Department
J. Gen. App!. Microbiol,, 21, 51-59 (1975) EFFECT OF CARBON SOURCES ON FORMATION OF a-amylase AND GLUCOAMYLASE BY CLOSTRIDIUM ACETOBUTYLICUM BURT ENSLEY, JOHN J. McHUGH, AND LARRY L. BARTON Department
Saccharomyces cerevisiae?
 JOURNAL OF BACTERIOLOGY, Aug. 1983, p. 623-627 21-9193/83/8623-5$2.O/ Copyright 1983, American Society for Microbiology Vol. 155, No. 2 What Is the Function of Nitrogen Catabolite Repression in Saccharomyces
JOURNAL OF BACTERIOLOGY, Aug. 1983, p. 623-627 21-9193/83/8623-5$2.O/ Copyright 1983, American Society for Microbiology Vol. 155, No. 2 What Is the Function of Nitrogen Catabolite Repression in Saccharomyces
hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide gel electrophoresis/genetics)
 Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
EFFECT OF DRYING ON PROTEIN PROFILE OF MURRAYA KOENIGII LEAVES S. Satheshkumar 1* and N. Punniamurthy 2
 International Journal of Science, Environment and Technology, Vol. 6, No 1, 2017, 861 865 ISSN 2278-3687 (O) 2277-663X (P) EFFECT OF DRYING ON PROTEIN PROFILE OF MURRAYA KOENIGII LEAVES S. Satheshkumar
International Journal of Science, Environment and Technology, Vol. 6, No 1, 2017, 861 865 ISSN 2278-3687 (O) 2277-663X (P) EFFECT OF DRYING ON PROTEIN PROFILE OF MURRAYA KOENIGII LEAVES S. Satheshkumar
Kit for assays of mammalian Trx
 FkTRX-04 Kit for assays of mammalian Trx The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
FkTRX-04 Kit for assays of mammalian Trx The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
(Mardeshev et al., 1948) and that the coenzyme of the decarboxylase has been
 STUDIES ON THE ASPARTIC ACID DECARBOXYLASE OF RHIZOBIUM TRIFOLII DANIEL BILLEN AND HERMAN C. LICHSTEIN Department of Bacteriology, University of Tennessee, Knoxville, Tennessee Received for publication
STUDIES ON THE ASPARTIC ACID DECARBOXYLASE OF RHIZOBIUM TRIFOLII DANIEL BILLEN AND HERMAN C. LICHSTEIN Department of Bacteriology, University of Tennessee, Knoxville, Tennessee Received for publication
MILK BIOSYNTHESIS PART 3: FAT
 MILK BIOSYNTHESIS PART 3: FAT KEY ENZYMES (FROM ALL BIOSYNTHESIS LECTURES) FDPase = fructose diphosphatase Citrate lyase Isocitrate dehydrogenase Fatty acid synthetase Acetyl CoA carboxylase Fatty acyl
MILK BIOSYNTHESIS PART 3: FAT KEY ENZYMES (FROM ALL BIOSYNTHESIS LECTURES) FDPase = fructose diphosphatase Citrate lyase Isocitrate dehydrogenase Fatty acid synthetase Acetyl CoA carboxylase Fatty acyl
Coupled, interconnecting reactions
 Metabolism: Basic concepts Hand-out for the CBT version November 2011 This module is based on 'Biochemistry' by Berg, Tymoczko and Stryer, seventh edition (2011), Chapter 15: Metabolism: Basic Concepts
Metabolism: Basic concepts Hand-out for the CBT version November 2011 This module is based on 'Biochemistry' by Berg, Tymoczko and Stryer, seventh edition (2011), Chapter 15: Metabolism: Basic Concepts
2-Deoxyglucose (2DG) Uptake Measurement kit
 Document#:K2DG13516E For research use only. Not for clinical diagnosis. Catalog No. CSR-OKP-PMG-K1E 2-Deoxyglucose (2DG) Uptake Measurement kit Introduction Measurement of 2-deoxyglucose (2DG) uptake in
Document#:K2DG13516E For research use only. Not for clinical diagnosis. Catalog No. CSR-OKP-PMG-K1E 2-Deoxyglucose (2DG) Uptake Measurement kit Introduction Measurement of 2-deoxyglucose (2DG) uptake in
BIOL 347L Laboratory Three
 Introduction BIOL 347L Laboratory Three Osmosis in potato and carrot samples Osmosis is the movement of water molecules through a selectively permeable membrane into a region of higher solute concentration,
Introduction BIOL 347L Laboratory Three Osmosis in potato and carrot samples Osmosis is the movement of water molecules through a selectively permeable membrane into a region of higher solute concentration,
Stabilization and enhanced reactivity of actinorhodin polyketide synthase minimal complex in polymer/nucleotide coacervate droplets
 Stabilization and enhanced reactivity of actinorhodin polyketide synthase minimal complex in polymer/nucleotide coacervate droplets John Crosby,* Tom Treadwell, Michelle Hammerton, Konstantinos Vasilakis,
Stabilization and enhanced reactivity of actinorhodin polyketide synthase minimal complex in polymer/nucleotide coacervate droplets John Crosby,* Tom Treadwell, Michelle Hammerton, Konstantinos Vasilakis,
Europium Labeling Kit
 Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin
 SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
Synthesis of Vitamin B6 by a Mutant of Escherichia coli K12 and the Action of 4 -Deoxypyridoxine
 Journal of General Microbiology (1979), 110, 285-289. Printed in Great Britain 28 5 Synthesis of Vitamin B6 by a Mutant of Escherichia coli K12 and the Action of 4 -Deoxypyridoxine By THOMAS A. SCOTT AND
Journal of General Microbiology (1979), 110, 285-289. Printed in Great Britain 28 5 Synthesis of Vitamin B6 by a Mutant of Escherichia coli K12 and the Action of 4 -Deoxypyridoxine By THOMAS A. SCOTT AND
Aperto Cell Lysis and Protein Solubilization Users Manual
 Aperto Cell Lysis and Protein Solubilization Users Manual Revision 2 THIS MANUAL APPLIES TO THE FOLLOWING PRODUCTS: 3A8600 Aperto, 5X Cell Lysis Buffer. 20mL 3A8610 Aperto, 5X Cell Lysis Buffer. 100mL
Aperto Cell Lysis and Protein Solubilization Users Manual Revision 2 THIS MANUAL APPLIES TO THE FOLLOWING PRODUCTS: 3A8600 Aperto, 5X Cell Lysis Buffer. 20mL 3A8610 Aperto, 5X Cell Lysis Buffer. 100mL
For the rapid, sensitive and accurate detection of long-chain Free Fatty Acids in various samples.
 ab65341 Free Fatty Acid Quantification Kit Instructions for Use For the rapid, sensitive and accurate detection of long-chain Free Fatty Acids in various samples. This product is for research use only
ab65341 Free Fatty Acid Quantification Kit Instructions for Use For the rapid, sensitive and accurate detection of long-chain Free Fatty Acids in various samples. This product is for research use only
unlike the wild-type strain, cannot incorporate exogenous fatty of a second long-chain acyl-coa synthetase which occurs in the
 Proc. Nati. Acad. Sci. USA Vol. 76, No. 9, pp. 4390-4394, September 1979 Biochemistry Involvement of long-chain acyl coenzyme A for lipid synthesis in repression of acetyl-coenzyme A carboxylase in Candida
Proc. Nati. Acad. Sci. USA Vol. 76, No. 9, pp. 4390-4394, September 1979 Biochemistry Involvement of long-chain acyl coenzyme A for lipid synthesis in repression of acetyl-coenzyme A carboxylase in Candida
Supporting Information for:
 Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Small-scale Triton X-114 Extraction of Hydrophobic Proteins Yuzuru Taguchi * and Hermann M. Schätzl
 Small-scale Triton X-114 Extraction of Hydrophobic Proteins Yuzuru Taguchi * and Hermann M. Schätzl Comparative Biology and Experimental Medicine, University of Calgary, Calgary, Canada *For correspondence:
Small-scale Triton X-114 Extraction of Hydrophobic Proteins Yuzuru Taguchi * and Hermann M. Schätzl Comparative Biology and Experimental Medicine, University of Calgary, Calgary, Canada *For correspondence:
(Anderson, 1946) containing sodium chloride, sodium-potassium phosphate. added to this basic medium in a concentration sufficient for maximum growth.
 THE EFFECTS OF A TRYPTOPHAN-HISTIDINE DEFICIENCY IN A MUTANT OF ESCHERICHIA COLI MARGOT K. SANDS AND RICHARD B. ROBERTS Carnegie Institution of Washington, Department of Terrestrial Magnetism, Washington,
THE EFFECTS OF A TRYPTOPHAN-HISTIDINE DEFICIENCY IN A MUTANT OF ESCHERICHIA COLI MARGOT K. SANDS AND RICHARD B. ROBERTS Carnegie Institution of Washington, Department of Terrestrial Magnetism, Washington,
ab ATP Synthase Enzyme Activity Microplate Assay Kit
 ab109714 ATP Synthase Enzyme Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of ATP Synthase activity in samples from Human, Rat and Cow This product is for research
ab109714 ATP Synthase Enzyme Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of ATP Synthase activity in samples from Human, Rat and Cow This product is for research
Free Fatty Acid Assay Kit
 Free Fatty Acid Assay Kit Catalog Number KA1667 100 assays Version: 05 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
Free Fatty Acid Assay Kit Catalog Number KA1667 100 assays Version: 05 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 General Information...
Glycine Synthesis and Metabolism in Escherichia coli
 JOURNAL OF BACTERIOLOGY, Apr., 1965 Copyright a 1965 American Society for Microbiology Vol. 89, No. 4 Printed in U.S.A. Glycine Synthesis and Metabolism in Escherichia coli LEWIS I. PIZER Departmiient
JOURNAL OF BACTERIOLOGY, Apr., 1965 Copyright a 1965 American Society for Microbiology Vol. 89, No. 4 Printed in U.S.A. Glycine Synthesis and Metabolism in Escherichia coli LEWIS I. PIZER Departmiient
Synthesis and Degradation of Liver Acetyl Coenzyme A Carboxylase
 Proc. Nat. Acad. Sci. USA Vol. 68, No. 9, pp. 2288-2292, September 1971 Synthesis and Degradation of Liver Acetyl Coenzyme A Carboxylase in Genetically Obese Mice (increased hepatic lipogenesis/immunochemical
Proc. Nat. Acad. Sci. USA Vol. 68, No. 9, pp. 2288-2292, September 1971 Synthesis and Degradation of Liver Acetyl Coenzyme A Carboxylase in Genetically Obese Mice (increased hepatic lipogenesis/immunochemical
