Efficient formation of influenza virus-like particles: dependence on the expression levels of viral proteins
|
|
|
- Linda Stafford
- 5 years ago
- Views:
Transcription
1 Journal of General Virology (1999), 80, Printed in Great Britain... Efficient formation of influenza virus-like particles: dependence on the expression levels of viral proteins Paulino Go mez-puertas, 1 Ignacio Mena, 1 Mar Castillo, 1 Amparo Vivo, 2 Esperanza Pe rez-pastrana 2 and Agustı n Portela 1 Centro Nacional de Biologı a Fundamental 1 and Centro Nacional de Microbiologı a 2, Instituto de Salud Carlos III, Majadahonda 28220, Madrid, Spain It has previously been demonstrated in this laboratory that an influenza virus-like chloramphenicol acetyltransferase (CAT) RNA could be expressed in COS-1 cells that synthesized all ten influenza A virus-encoded proteins from recombinant plasmids. It was also shown that supernatant fluids harvested from these cultures contained virus-like particles (VLPs) that could deliver an enclosed CAT RNA to MDCK cells. Here, it is shown that the levels of expression of the reporter gene in the COS-1 and/or MDCK cells can be altered drastically by modifying the concentrations of the recombinant plasmids transfected in the COS-1 cells. Thus, it was observed that overexpression of NS2 reduced CAT expression in COS-1 cells, whereas overexpression of M2 and NS1 proteins dramatically decreased transmission of the CAT RNA to the MDCK cultures. These results are discussed with reference to the roles of these proteins during virus replication. From these experiments, a ratio of transfected plasmids was found that increased the efficiency of the previously described system by fold. Under these optimized conditions, it was demonstrated that VLPs can be formed in the absence of neuraminidase expression and that these VLPs remained aggregated to each other and to cell membranes. Moreover, it is shown that CAT RNA transmission was dependent on specific interactions of the ribonucleoprotein complex with other viral structural polypeptides. These data demonstrate the usefulness of this encapsidation packaging system for the study of different aspects of the influenza virus life-cycle. Introduction Influenza A virions are pleomorphic, enveloped particles with a diameter of nm. The viral genome, which consists of eight negative-sense, single-stranded RNAs, has a coding capacity for ten polypeptides. The virion contains three integral membrane proteins, haemagglutinin (HA), neuraminidase (NA) and the M2 ion channel protein. Six other viral proteins are found within the virion membrane. Four of them [nucleoprotein (NP), PB1, PB2 and PA] are associated with the viral genome to form ribonucleoprotein (RNP) complexes and the other two polypeptides, M1 and NS2 [also called NEP; O Neill et al. (1998)], interact with each other and with the Author for correspondence: Agustı n Portela. Fax aportela isciii.es Present address: Department of Neuropharmacology, CVN-9, The Scripps Research Institute, North Torrey Pines Road, La Jolla, CA 92037, USA RNPs. The NS1 protein is the only non-structural component of the virus (Lamb, 1989; Lamb & Krug, 1996). The virus is internalized by receptor-mediated endocytosis and, after fusion of the viral and endosomal membranes, the infecting RNPs are transported from the cytosol to the cell nucleus (Martin & Helenius, 1991), where replication and transcription of the viral genome takes place (Herz et al., 1981). The newly synthesized RNPs are then exported from the cell nucleus to the cytoplasm (Martin & Helenius, 1991; Whittaker et al., 1996; O Neill et al., 1998) and should reach the proximity of the cellular membrane, where virus budding occurs (Compans & Dimmock, 1969). Although the interactions between the virus components that govern formation of virion particles are poorly understood, it is thought that contacts between the RNPs and other virus components are critically important. In fact, it has been shown that interactions between the RNPs and the M1 and NS2 proteins modulate the nuclear cytoplasmic transport of RNPs (Martin & Helenius, 1991; Whittaker et al., 1996; SGM BGDF
2 P. Go mez-puertas and others O Neill et al., 1998) and morphological (Murti et al., 1992) and biochemical (Zvonarjev & Ghendon, 1980; Zhirnov, 1992) observations suggest that M1 protein associates with the RNPs in the virion. Despite these reports, functional evidence demonstrating the importance of the interactions between RNPs and other virus factors for the formation of infectious particles is still lacking. Contacts between the cytoplasmic tails of the virus membrane proteins and the virion internal components are also important for formation of the budding particle. Thus, viruses lacking the cytoplasmic tail of HA or NA or both have reduced infectivity and show alterations in their morphology (Jin et al., 1994, 1997; Garcı a-sastre & Palese, 1995; Mitnaul et al., 1996). The role of NA in the formation of infectious virions has been the subject of a number of studies. It has been shown that NAdeficient viruses produce particles that form large aggregates on the cell surface (Palese et al., 1974; Liu et al., 1995) and that these particles, when released from the cell, can complete a round of replication in animals and cell culture (Liu et al., 1995). These results indicate, therefore, that NA activity is needed to prevent formation of virus aggregates and to allow the release of fully assembled virus particles from the cell surface. It should be mentioned that the NA-deficient mutants, which were selected in the presence of bacterial NA, contained a deleted NA segment that retains the capacity to code for an N-terminal NA peptide (Yang et al., 1997). It is, however, not known whether expression of this N-terminal sequence, which includes the membrane-anchoring region of NA, is required for formation of virions. Recently, we described a system in which a synthetic influenza A virus-like chloramphenicol acetyltransferase (CAT) RNA could be encapsidated, replicated and packaged into virus-like particles (VLPs) in cells expressing all virus-encoded polypeptides from plasmids (Mena et al., 1996). This system is analogous to those described for the negative-sense RNA viruses vesicular stomatitis virus (Pattnaik & Wertz, 1991), rabies virus (Conzelmann & Schnell, 1994), Bunyamwera virus (Bridgen & Elliott, 1996) and human respiratory syncytial virus (Teng & Collins, 1998). The influenza virus rescue system was, however, very inefficient with respect to transmission of the CAT RNA by the VLPs. Therefore, the system was not suitable for systematic studies of the roles played by the viral proteins during virus replication. Here, we describe the optimization of the rescue system by a factor of fold and show the usefulness of this system for the study of the role of NA in formation of VLPs and to demonstrate that interactions of RNP components with other virus factors are required for formation of functional VLPs. Methods Cell cultures and viruses. COS-1 and MDCK cells were maintained in Dulbecco s modified Eagle s medium (DMEM) containing 10% foetal calf serum. Recombinant vaccinia virus (vtf7-3), which expresses T7 RNA polymerase, was kindly provided by B. Moss (National Institutes of Health, Bethesda, MA, USA) (Fuerst et al., 1986). The influenza virus strains used were A Victoria 3 75 (H3N2), A Puerto Rico 8 34 (H1N1) and B Panama Plasmids and RNAs. Plasmids pgem-pb1, pgem-pb2, pgem- PA, pgem-np, pgem-ha, pgem-na, pgem-m1, pgem-m2, pgem- NS1 and pgem-ns2, encoding the influenza virus polypeptides PB1, PB2, PA, NP, HA, NA, M1, M2, NS1 and NS2, respectively, from the A Victoria 3 75 strain, have been described previously (Mena et al., 1994, 1996). In these plasmids, the virus genes are cloned downstream of the T7 promoter of plasmids pgem-3 or pgem-4 (Promega). Plasmid pgem-m3-1, which contains a cdna copy of mrna (derived from the M segment) (Lamb et al., 1981) cloned under the control of the T7 promoter of plasmid pgem-3, was derived by RT PCR from mrna isolated from cells infected with influenza virus A Victoria Plasmids pgb-pb1-89.1, pgb-pb2-2, pgb-pa-4 and pgb-np-7, encoding the influenza virus proteins PB1, PB2, PA and NP, respectively, from the B Panama strain, have been described previously (Jambrina et al., 1997). Plasmids pivacat1 S (Piccone et al., 1993) and pt7nsbcat (Barclay & Palese, 1995) were kindly provided by P. Palese and W. S. Barclay (Mount Sinai School of Medicine, New York, USA). These plasmids were used to generate influenza A and B virus-like model RNAs after transcription with T7 RNA polymerase. These RNAs contain the CAT gene in negative polarity flanked by the 5 and 3 untranslated sequences of the RNA segment encoding the NS proteins of the corresponding influenza A or B virus. Plasmid concentrations were estimated spectrophotometrically by assuming that an A of 1 corresponded to 50 µg ml DNA. Antibodies and immunoblotting. Monoclonal antibodies (MAbs) M 58 p51 G, HA1-50 and M F4, which recognize the A Victoria 3 75 NP and HA proteins, and a rabbit antiserum that recognizes the C terminus of NP have been described previously (Arrese & Portela, 1996; Lo pez et al., 1986; Sa nchez-fauquier et al., 1987). Goat antiserum against M2 protein was a gift from Alan Hay (National Institute for Medical Research, London, UK). A polyclonal antiserum against M1 protein was obtained by immunizing rabbits with a denatured, histidine-tagged M1 protein. Western blotting analysis with antisera to M2, M1 or NP and MAb HA1-50 was carried out as described previously (Arrese & Portela, 1996). Construction of plasmids pgem-m1, pgem-m2 and pgem-ns2. Plasmid pgem-m2 contains a cdna copy of the M2 mrna cloned in the polylinker of plasmid pgem-3 (Promega). In this plasmid, the nucleotide sequence following the T7 promoter is GGGAGACCGGAATTCGAGCTCGGTACCCTCTTCAagcaaacgcaggtagatatcgaaagatg, where the pgem-3 vector sequence is shown in capitals and the influenza virus-derived sequence is shown in lower case. The EcoRI site present in the vector is underlined and the ATG initiation codon of the M2 protein is indicated in italics. A new EcoRI site was introduced in this plasmid by converting the sequence ta into at. This mutagenesis step was carried with the Transformer site-directed mutagenesis kit (Clontech). The mutagenized plasmid was then digested with EcoRI and circularized to yield plasmid pgem-m2. This plasmid contains the sequence GGGAGACCGGAATTCgaaagatg following the T7 promoter and therefore lacks most of the virus sequences present upstream of the ATG codon in plasmid pgem-m2. An analogous strategy was used to generate plasmids pgem-m1 and pgem-ns2 from plasmids pgem-m1 and pgem-ns2, respectively. The only difference was that an XbaI site, instead of the EcoRI site, was introduced to prepare plasmid pgem-m1, since plasmid pgem- M1 was derived from the pgem-4 vector. The sequences downstream of the T7 promoter in these plasmids were GGGAGACCGGAAGCTT- BGDG
3 Influenza VLPs and viral protein expression Fig. 1. Optimization of input amounts of plasmids pgem-m2, pgem-m1 and pgem-ns2. (A) COS-1 cells were infected with vtf7-3 and transfected with plasmids encoding proteins PB1, PB2, PA, NP, HA, NA, M1 and NS2 (the amounts are indicated in the text) and the indicated amounts (µg) of plasmid pgem-m2. The cultures were transfected 5 h later with a synthetic CAT RNA. The supernatants from these cultures were harvested 72 h post-infection and added to MDCK cells that were superinfected with a helper influenza virus as detailed in Methods. Aliquots of the COS-1 and MDCK cell extracts were assayed for CAT activity and autoradiographs of the corresponding TLC plates are shown in the figure, with the positions of [ 14 C]chloramphenicol and 3-acetylated [ 14 C]chloramphenicol indicated by open and filled arrowheads, respectively. CAT activities in MDCK cells were expressed as a percentage of that obtained in the sample yielding the highest activity (as shown bellow each lane on the TLC). CAT activities in COS-1 cells were expressed as a percentage by considering the activity of the COS-1 cell sample corresponding to that yielding the highest CAT level in MDCK cells as 100%. The relative CAT activities are shown below each of the samples of the corresponding TLC plate. The lower panel corresponds to a Western blot of the COS-1 cell extracts probed with antibodies (α) against the proteins indicated. In sample M1, the plasmid pgem-m1 was not included in the transfection mixture. (B) The experiment was performed as described in (A) except that the amount of plasmid pgem-m2 included in the transfection mixture was 150 ng and the concentrations of pgem-m1 and pgem-ns2 were varied as indicated. GCATGCCTGCAGGTCGACTCTAGAaagatg (pgem-m1 ) and GGGAGACCGGAATTCgacataatg (pgem-ns2 ). Assay for detection of functional VLPs. The standard protocol was carried out basically as described by Mena et al. (1996). The major modifications were (i) that plasmids pgem-m1, pgem-m2 and pgem-ns2 were used instead of plasmids pgem-m1, pgem-m2 and pgem-ns2 and (ii) that the influenza virus strain A Puerto Rico 8 34 substituted for A Victoria 3 75 as helper virus, since the former grows to higher titres in cell culture. Briefly, the protocol was as follows. Cultures of COS-1 cells (10 cells) growing in 35 mm diameter dishes in the presence of DMEM-Ara-Ant (DMEM containing 40 µg ml cytosine β-d-arabinofuranoside, 100 U ml penicillin and 100 µg ml streptomycin) were infected with vtf7-3 (m.o.i. of 5). After virus adsorption, the cultures were incubated with 1 ml DMEM-Ara-Ant that was supplemented with a 100 µl mixture that contained cationic liposomes and the plasmids indicated in each case. After the optimization experiments, the optimal amounts of the plasmids in the transfection mixture were: pgem-pb1 (0 6 µg), pgem-pb2 (0 6 µg), pgem-pa (0 2 µg), pgem-np (2 µg), pgem-ha (0 6 µg), pgem-na (0 6 µg), pgem-m1 (0 15 µg), pgem-m2 (0 15 µg) and pgem-ns2 (0 1 µg) (see text for details). After 5 h incubation with the plasmids, the cells were transfected again with a mixture containing 0 5 µg synthetic CAT RNA. After incubation overnight, the medium was replaced with 1 ml DMEM-Ara-Ant and the cultures were incubated for an additional 48 h. Cell supernatants were then harvested and cell extracts were prepared. Aliquots of the cell extracts were used for Western blotting and for determination of CAT activity (see below). The supernatant collected from the COS-1 cells was clarified by centrifugation for 15 min in a microcentrifuge and subjected to three cycles of freezing and thawing. To test for the presence of VLPs, an aliquot of the supernatant (typically 400 µl) was incubated with trypsin (2 5 µg ml) for 15 min at 37 C and added to 10 MDCK cells, growing in 35 mm diameter dishes in the presence of DMEM-Ara-Ant. After 1 h incubation, the cells were superinfected with influenza virus A Puerto Rico 8 34 (or B Panama when indicated) at an m.o.i. of 5 and the cultures were incubated for 17 h. Cell extracts were then prepared for determination of CAT activity. Routinely, for these assays a sample corresponding to 5000 COS-1 cells and to MDCK cells was incubated for 1 h at 37 C with [ C]chloramphenicol as described previously (Mena et al., 1996). Quantification of CAT activity was performed by phosphorimaging of the acetylated spots detected on TLC plates by using a Fujix Bas 1000 phosphorimager and the software PCBAS v2.09. BGDH
4 P. Go mez-puertas and others Electron microscopy. Transfected cultures were pre-cooled for 15 min at 4 C and incubated for 30 min with the anti-ha MAb M F4 (diluted 1:5 in DMEM containing 5% sheep serum). After washing with DMEM 5% sheep serum, the samples were incubated for 30 min with a secondary antibody (10 nm gold-labelled, goat anti-mouse IgG; Auroprobe EM GAM IgG G10; Amersham) diluted 1: 20 in DMEM. After washing with Tris-buffered saline (25 mm Tris HCl, ph 7 4, 2 68 mm KCl, 137 mm NaCl), the cells were fixed with 2% glutaraldehyde in 0 1 M cacodylate buffer (ph 7 4) and post-fixed with 1% osmium tetroxide prepared in the same buffer for 1 h at 4 C. The fixed cells were then dehydrated and embedded in epoxy resin EPON 812. Sections prepared with an LKB Ultratome IV were post-stained in 1% aqueous uranyl acetate for 15 min and then 3 min in lead citrate and were visualized with a Philips 400T electron microscope at 80 kv. Results Optimization of the CAT rescue system As detailed previously (Mena et al., 1996), the protocol followed to generate and detect recombinant VLPs included two steps. In the first step, COS-1 cells were infected with vtf7-3 and transfected with the pgem-derived plasmids encoding all influenza virus polypeptides and with a synthetic, negative-sense CAT RNA. In the second step, the supernatant fluids collected from the transfected COS-1 cells were treated with trypsin and incubated with fresh MDCK cultures that were then superinfected with a wild-type influenza helper virus. Detection of CAT activity in COS-1 cells demonstrated expression of the model CAT RNA by the recombinant polymerase, whereas detection of CAT activity in the MDCK cell extracts indicated that the COS-1 cell supernatant contained functional VLPs, i.e. VLPs competent to deliver an enclosed CAT RNA into MDCK cells. It has been shown for a number of systems in which a model RNA is replicated and or packaged that obtaining the highest replication and or packaging levels depends on finding the optimal ratio of transfected plasmids (Pattnaik & Wertz, 1990; Dunn et al., 1995; Sanz-Ezquerro et al., 1995; Teng & Collins, 1998). It was therefore decided to test the concentration-dependent effects of the transfected plasmids on the formation of influenza VLPs by following the protocol outlined above. The experiments were initiated with only the nine plasmids that encode the viral structural components, since we had demonstrated previously that formation of VLPs does not require expression of NS1 protein (Mena et al., 1996). The concentrations of the plasmids encoding the virus core proteins were those found previously to be optimal for expression of a synthetic CAT RNA (Sanz-Ezquerro et al., 1995) and the starting concentrations of the remaining plasmids were those used in the previous report (1 µg of each plasmid). The first experiment was to optimize the concentration of plasmid pgem-m2 (Fig. 1 A), since preliminary assays indicated that the concentration of the M2 protein drastically affected the CAT activity detected in MDCK cells (not shown). It was observed that increasing the amount of transfected M2 plasmid resulted in greater accumulation of the M2 protein in COS-1 cells, as expected from the transient expression system used (Fuerst et al., 1986). To detect alterations in the expression levels of the other recombinant proteins, which could arise as a result of transfecting different doses of a particular plasmid, we routinely checked for the accumulation of NP and or HA in the different cell samples. As can be observed in Fig. 1(A), there were no significant changes in the accumulation of these two proteins in any of the samples analysed. Moreover, the reporter gene activity measured in COS-1 cells was not affected by the level of expression of M2, indicating that this protein does not modify the functionality of the core proteins to express the input CAT RNA. However, the concentration of the M2 plasmid affected formation of functional VLPs drastically. Transfection of very small amounts of the plasmid (5 ng) were sufficient to allow detection of functional VLPs and maximum detection was achieved after transfection of 150 ng plasmid. From that point on, CAT activity in MDCK cells decreased until it reached virtually background values with transfection of 0 9 µg plasmid. On the basis of these results, all subsequent transfection experiments were carried out with 150 ng pgem-m2. It was next decided to titrate the plasmids encoding the M1 and NS2 proteins. In preliminary tests, it was observed that transfecting more than 300 ng pgem-ns2 had an inhibitory effect on CAT expression in COS-1 cells (data not shown, see below), whereas transfection of small amounts of the M1 plasmid (50 ng) resulted in low levels of CAT expression in the MDCK cultures (data not shown). Since M1 and NS2 proteins appear to interact with each other (Yasuda et al., 1993; O Neill et al., 1998), it was decided to carry out co-transfection experiments with the plasmids encoding these two proteins. On the basis of preliminary experiments, the doses chosen for these analyses varied from 150 to 1300 ng for pgem-m1 and from 30 to 300 ng for pgem-ns2 (Fig. 1B). For each of the three doses of pgem-m1 tested, 100 ng of the NS2 plasmid always yielded the highest CAT activities in MDCK cells; the sample transfected with the smallest amount of M1 plasmid was the one which yielded the highest reporter gene activity in MDCK cells. It was again observed that there was a direct correlation between the amount of plasmid pgem- M1 transfected and the concentration of M1 protein in the COS-1 cultures. The accumulation of NS2 protein was not tested because of the lack of an appropriate immunological reagent. Once the optimal amounts of the M1 (150 ng) and NS2 (100 ng) plasmids were determined, the dose-dependent effects of HA and NA plasmids in the CAT rescue system were investigated. As shown in Fig. 2(A), the different amounts of plasmids tested did not have any significant effect on the CAT activity obtained in COS-1 cells and the doses of these two plasmids that yielded the highest level of CAT expression in MDCK cells were 600 ng. It was then decided to study the effect of NP concentration on the system and to confirm the inhibitory effect (mentioned BGDI
5 Influenza VLPs and viral protein expression Fig. 2. Optimization of input amounts of plasmids pgem-ha, pgem-na, pgem-np and pgem-ns2. (A) The experiment was performed as described in Fig. 1(B) except that the amounts of plasmids pgem-m1 and pgem-ns2 included in the transfection mixture were 150 and 100 ng, respectively. The concentrations of pgem-ha and pgem-na were varied as indicated. (B) The experiment was performed as described in (A) except that the amounts of pgem-ha and pgem-na included in the transfection mixture were 600 ng each. The concentrations of plasmids pgem-np and pgem-ns2 were varied as indicated. above) of NS2 on expression of the CAT RNA in COS-1 cells (Fig. 2B). It was observed that reducing the amount of NP plasmid led to a reduction in the level of CAT expression in COS-1 cells. A similar effect was observed by increasing the amount of plasmid pgem-ns2 in the transfection mixture. In both cases, the CAT activity detected in MDCK cells was roughly proportional to that detected in COS-1 cells. One interpretation of these results would be that either small amounts of NP or large amounts of NS2 reduced the production of viral RNPs (containing the CAT RNA) in transfected cells. Consequently, less viral RNP would be packaged into VLPs and lower CAT activity would be detected in the MDCK cell extracts. On the basis of the results obtained, the doses of plasmids encoding NP and NS2 were maintained at 2 µg and 100 ng, respectively. Once the system had been optimized in the absence of NS1 protein, we re-examined the effect of this protein on formation of VLPs. It was confirmed that NS1 protein was not required for efficient transmission of CAT RNA to MDCK cells (Fig. 3 A) (Mena et al., 1996). Strikingly, it was observed that increasing the amount of NS1 plasmid from 100 ng to 2 µg resulted in a 100-fold reduction in the CAT activity detected in MDCK cells, whereas within this dose range, there was only a 3-fold reduction in the CAT activity observed in the COS- 1 cell extracts. Finally, the possible effect of mrna on the rescue system was tested. mrna is a spliced product derived from RNA segment 7 and it contains an open reading frame for a 9 amino acid peptide that has never been found in infected cell cultures (Lamb et al., 1981). As shown in Fig. 3(B), expression of mrna did not have a significant effect on the reporter gene activities reached in COS-1 and MDCK cells. It should be pointed out that the M1 mrna (derived from plasmid pgem- M1 ) will not be spliced to generate mrna, since it lacks the 5 splice site. Moreover, it is worth mentioning that the T7- derived transcripts produced in the vaccinia T7 system are not expected to be spliced, since they are synthesized in the cytoplasm. From the above experiments, we determined the optimal concentration of the nine plasmids encoding the virus structural polypeptides that allowed efficient formation of functional VLPs. By considering the amounts of the MDCK cell extract tested in the CAT assays (5% as compared to 20 50% in the previous report) as well as the CAT activities observed, it was calculated that the optimized rescue system was fold more efficient than the previous one (Mena et al., 1996) for CAT transmission to the MDCK cells. CAT transmission to MDCK cells is mediated by VLPs Using the optimized conditions, a series of experiments identical to those described in the previous report (Mena et al., 1996) were carried out to characterize the particles that transmitted the CAT RNA. It was confirmed, with three different plasmid preparations, that expression of all viral BGDJ
6 P. Go mez-puertas and others Fig. 3. Optimization of input amounts of plasmids pgem-ns1 and pgem-m3. The experiment was performed as described in Fig. 2(B). The concentrations of plasmids pgem-ns1 (A) and pgem-m3-1 (B) were varied as indicated. N.A., Not analysed. In sample NS2, the plasmid pgem-ns2 was not included in the transfection mixture. Fig. 4. Viral proteins required for transmission of CAT RNA and characterization of the VLPs that transmit the CAT RNA to MDCK cells. (A) COS-1 cells were infected with vtf7-3 and transfected with the nine plasmids encoding the structural viral polypeptides (sample NS1) or with only eight of them (samples HA, NA, M1, M2 and NS2, where the plasmid encoding the indicated gene product was also omitted). The cultures were then transfected with a CAT RNA and the supernatants from these cultures were tested on MDCK cells. Aliquots of the COS-1 and MDCK cell extracts were assayed for CAT activity as detailed in Methods. (B) The supernatant obtained from COS-1 cells expressing the nine structural virus polypeptides and transfected with a CAT RNA was harvested. Aliquots of this supernatant were then incubated with trypsin, RNase A or MAbs specific for influenza virus HA (MAb 234/1/F4) or NP (MAb M58/p51/G) as indicated. The treated supernatants were next added to MDCK cells, which were then mock-infected or infected with the influenza virus strain A/Puerto Rico/8/34 as indicated (Inf. virus). Cell extracts were prepared and assayed for CAT activity as described above. structural proteins was required for detection of CAT activity in MDCK cells (Fig. 4A). Moreover, it was demonstrated that treatment of the COS-1 cell supernatant with trypsin and superinfection with an influenza helper virus were absolute requirements for detection of reporter gene activity in MDCK cells (Fig. 4B). In addition, it was shown that transmission of the CAT RNA to MDCK cells could be abolished by incubation of the COS-1 cell supernatant with a neutralizing anti-ha MAb but that it was unaffected by treatment of this supernatant with a MAb to NP, an antiserum to vaccinia virus or RNase A (Fig. 4B and data not shown). Taken together, these results allowed us to conclude that the CAT activity detected in the MDCK cell extracts was transmitted by VLPs enclosing the CAT RNA and that these VLPs should resemble authentic influenza virions, since expression of all virus structural proteins was required for CAT RNA transmission. Electron microscopy of VLPs In an attempt to visualize the recombinant VLPs, thin sections of COS-1 cells expressing all virus structural proteins were incubated with an anti-ha MAb and examined by BGEA
7 BGEB Fig. 5. Immunoelectron microscopy of transfected COS-1 cells. COS-1 cells were infected with vtf7-3 and transfected with the nine plasmids encoding all the influenza virus structural proteins ( NS1) or with a similar mixture that also lacked the NA plasmid ( NA). At 60 h post-infection, cells were incubated with an anti- HA MAb (M234/1/F4) and decorated with a 10 nm immunogold conjugate before fixation as described in Methods. Bars represent 200 nm. Influenza VLPs and viral protein expression
8 P. Go mez-puertas and others Fig. 6. Formation of heterotypic VLPs. Four COS-1 cell cultures (samples 1 4) were infected with vtf7-3 and then transfected with DNA mixtures containing the pgem-derived plasmids corresponding to the influenza A virus proteins HA, NA, M1, M2 and NS2. The DNA mixtures contained, in addition, four plasmids encoding the three subunits of the polymerase and NP from either A/Victoria/3/75 (A) or B/Panama /45/90 (B) influenza viruses (as indicated under Pol NP). The amounts of type B plasmids included in the transfection mixture were: 1 µg of pgb-pb1-89.1, 0 5 µg of pgb-pb2-2, 0 5 µg of pgb-pa-4 and 3 µg of pgb-np-7. The cells were transfected again 5 h later with an NS-CAT RNA containing the conserved ends of either type A or B virus (indicated in RNA). At 72 h post-infection, cell supernatants were harvested and the COS-1 cell extracts were assayed for CAT (left panel). Supernatants from COS-1 samples 1 4 were divided into two aliquots and added to MDCK cultures that were then infected with either influenza virus A/Puerto Rico/8/34 (A) or B/Panama /45/90 (B) as indicated (Inf. virus). Cell extracts were prepared 17 h later and assayed for CAT expression (right panel). electron microscopy (Fig. 5, samples NS1). A few filamentous particles were observed in the proximity of 5 30% of the cells examined (routinely more than 100) and similar structures were not observed in cultures that did not receive plasmid pgem-m1. These particles resembled authentic influenza virions in size and morphology and could be labelled specifically with an anti-ha antibody. It was therefore concluded that the particles detected corresponded to the VLPs that transmitted the CAT RNA. We have demonstrated that the supernatant from COS-1 cells expressing all structural proteins except NA was unable to transmit the CAT RNA to MDCK cells (Fig. 4 A). One explanation of this result would be that VLPs were not formed in the absence of the NA protein. Alternatively, on the basis of the results obtained with the NA-deficient viruses (see Introduction), it is possible that budded particles were formed but that they remained aggregated and attached to the surface of the COS-1 cells. To distinguish between these two possibilities, COS-1 cultures not expressing NA were examined by electron microscopy (Fig. 5, samples NA). Particles that had apparently completed budding were observed in 5 30% of the cells examined. Virtually all particles observed were found in aggregates or associated with the cell surface singly or in small groups. The aggregated particles could be decorated with an anti-ha MAb and structures of this kind were not observed in cultures expressing all structural proteins except NA and M1. On the basis of these findings, we concluded that the filamentous, membrane-associated particles were in fact VLPs lacking NA. Interactions between RNPs and other viral proteins are essential for formation of functional VLPs It was then decided to test whether the rescue system could provide functional evidence of the importance of interactions between RNPs and other viral proteins for the formation of mature virions. To this end, transfection mixtures containing plasmids encoding either the influenza A or B virus core proteins and, in addition, plasmids expressing the influenza A virus HA, NA, M1, M2 and NS2 proteins were prepared. These mixtures were transfected into COS-1 cells that were then further transfected with either an influenza A or B virustype CAT RNA. The supernatants from these cultures were then assayed for CAT RNA transmission to MDCK cells by the standard protocol, infecting the cultures with either influenza A or B virus. The results obtained in a representative experiment are shown in Fig. 6. As we have described previously (Jambrina et al., 1997), both the influenza A and B core proteins were capable of replicating amplifying the heterotypic RNAs. However, CAT expression in MDCK cells was only observed in the samples in which all recombinant proteins and the helper virus were from the type A virus, independent of the transfected CAT RNA. These results suggest that virus type-specific contacts between the RNPs and other viral polypeptides are required for formation of functional VLPs. Discussion We have described here an optimized system that allows efficient rescue of synthetic CAT RNA into functional VLPs. Several interesting observations were made during the optimization experiments. Overexpression of the NS2 protein had an inhibitory effect on CAT expression in COS-1 cells, suggesting that this protein reduced the level of transcription and or replication of the input CAT RNA. It is suggested that the observed effect may involve interactions of NS2 with virus factors other than the core proteins, since other authors did not observe such an effect in cells expressing exclusively NS2 and the four core proteins (Huang et al., 1990; Enami et al., 1994). It remains to be tested whether the NS2 effect may have something to do with the poorly characterized role of the protein in replication of viral RNAs (Odagiri & Tobita, 1990) BGEC
9 Influenza VLPs and viral protein expression and or with the role of NS2 in transport of RNPs from the nucleus to the cytoplasm (O Neill et al., 1998). Overexpression of M2 completely blocked CAT RNA transmission to MDCK cultures. M2 functions as an ion channel protein (Sugrue et al., 1990; Pinto et al., 1992) and it has been shown that, when this protein is overexpressed, the intracellular transport of co-expressed HA is inhibited and the accumulation of HA at the plasma membrane is reduced by 75 80% (Sakaguchi et al., 1996; Henkel & Weisz, 1998). It is suggested that a similar effect occurs in the rescue system, so that, by overexpressing M2, the accumulation of virus membrane proteins at the plasma membrane is reduced and hence there is a drastic reduction in the number of functional VLPs produced. The situation with NS1 is peculiar, in that the protein was not needed for formation of functional VLPs but functional VLPs were not detected when the protein was overexpressed. The NS1 protein is an RNA-binding protein that has been found associated with RNPs (Mario n et al., 1997) and that inhibits nucleocytoplasmic transport of RNAs, stimulates translation of viral mrnas and modulates splicing of influenza virus mrnas (Nemeroff et al., 1998; reviewed in Lamb & Krug, 1996; Ortı n, 1998). We hypothesize that, when NS1 protein is overexpressed, the CAT RNPs are sequestered into the cell nucleus and therefore cannot be packaged into VLPs. It should be mentioned that the three proteins (NS1, NS2 and M2) that most dramatically affected CAT expression are translated in infected cells from spliced mrnas (M2 and NS2) or from an mrna that is a substrate for splicing (NS1) (Lamb & Lai, 1980; Lamb et al., 1981). This is consistent with previous evidence indicating that splicing of viral mrnas is tightly regulated in infected cells (Alonso-Caplen & Krug, 1991; Valca rcel et al., 1991; Shih et al., 1995). We have shown here, using three different plasmid preparations, that all virus structural proteins were required for formation of functional VLPs. It should be mentioned that in our previous report (Mena et al., 1996), similar results were obtained with two plasmid preparations but in the case of a third, it was observed that expression of M2 was not required for formation of functional VLPs. We found here that transfecting very small amounts of the M2 plasmid was sufficient for a positive CAT signal to be detected in the MDCK cells. It is therefore suggested that the results obtained previously with one of the plasmid preparations were probably due to minor contamination with the M2 plasmid. It was demonstrated by electron microscopic analysis that particles similar to wild-type virions were formed in the absence of NA expression. As indicated above, the NAdeficient mutants described previously, which were selected in the presence of bacterial NA, retained the capacity to encode an N-terminal NA peptide (Yang et al., 1997). Our data indicate that expression of this NA fragment is not essential for virus particle formation, although we cannot rule out that its expression could confer a selective advantage to the NAdeficient virus. It has been described that viruses lacking the cytoplasmic tail of NA have a higher tendency to be filamentous (Mitnaul et al., 1996; Jin et al., 1997). This observation has been made with the WSN strain, which produces primarily spherical particles. It has not been possible to determine whether a similar effect occurs with the A Victoria 3 75 VLPs formed in the absence of NA, since both the wild-type virions and the VLPs containing NA appear mainly as filamentous viruses. Functional VLPs were not detected when the core proteins were derived from an influenza B virus and the rest of the structural proteins from a type A virus. This result suggests strongly that critical virus type-specific interactions between the RNP components and other viral proteins take place during virion assembly. These interactions probably involve protein protein rather than RNA protein contacts, since an influenza B virus RNA could be transmitted to MDCK cells by the type A proteins. In the RNP complexes, the polymerase complex holds the RNA ends together, whereas NP interacts with the rest of the RNA, exposing the RNA bases to the solvent (Klumpp et al., 1997). We propose that the interactions that affect RNP packaging involve the NP polypeptide, since the polymerase subunits are minor components of the RNPs. An interesting observation made during the experiments involving heterotypic virus components was that the type A CAT RNPs were expressed in MDCK cells infected with an influenza A virus but not when the cells were infected with an influenza B virus (samples 1 and 2, Fig. 6). We had hypothesized (Mena et al., 1996) that the CAT RNPs delivered into MDCK cells are capable, in the absence of helper virus infection, of primary transcription, but that detection of CAT activity depends on the replication (amplification) of the CAT RNPs, mediated by the proteins provided in trans by the helper virus. The lack of expression of the type A CAT RNPs in influenza B virusinfected cells indicates that the influenza B virus proteins cannot switch the type A CAT RNPs from primary transcription to replication. In summary, we have developed an efficient encapsidation packaging system and we have presented evidence that shows its usefulness in the characterization of the roles of viral proteins during the influenza virus life-cycle. This work was supported by the Fondo de Investigaciones Sanitarias (Grant ). P. Go mez-puertas and I. Mena were supported by fellowships from the Instituto de Salud Carlos III and Comunidad Auto noma de Madrid, respectively. We thank J. Ortı n for critically reading the manuscript and A. del Pozo for the artwork. We thank W. Barclay, A. Hay, B. Moss and P. Palese for the reagents provided. References Alonso-Caplen, F. V. & Krug, R. M. (1991). Regulation of the extent of splicing of influenza virus NS1 mrna: role of the rates of splicing and of the nucleocytoplasmic transport of NS1 mrna. Molecular and Cellular Biology 11, BGED
10 P. Go mez-puertas and others Arrese, M. & Portela, A. (1996). Serine 3 is critical for phosphorylation at the N-terminal end of the nucleoprotein of influenza virus A Victoria Journal of Virology 70, Barclay, W. S. & Palese, P. (1995). Influenza B viruses with site-specific mutations introduced into the HA gene. Journal of Virology 69, Bridgen, A. & Elliott, R. M. (1996). Rescue of a segmented negativestrand RNA virus entirely from cloned complementary DNAs. Proceedings of the National Academy of Sciences, USA 93, Compans, R. W. & Dimmock, N. J. (1969). An electron microscopic study of single-cycle infection of chick embryo fibroblasts by influenza virus. Virology 39, Conzelmann, K. K. & Schnell, M. (1994). Rescue of synthetic genomic RNA analogs of rabies virus by plasmid-encoded proteins. Journal of Virology 68, Dunn, E. F., Pritlove, D. C., Jin, H. & Elliott, R. M. (1995). Transcription of a recombinant bunyavirus RNA template by transiently expressed bunyavirus proteins. Virology 211, Enami, K., Sato, T. A., Nakada, S. & Enami, M. (1994). Influenza virus NS1 protein stimulates translation of the M1 protein. Journal of Virology 68, Fuerst, T. R., Niles, E. G., Studier, F. W. & Moss, B. (1986). Eukaryotic transient-expression system based on recombinant vaccinia virus that synthesizes bacteriophage T7 RNA polymerase. Proceedings of the National Academy of Sciences, USA 83, Garcı a-sastre, A. & Palese, P. (1995). The cytoplasmic tail of the neuraminidase protein of influenza A virus does not play an important role in the packaging of this protein into viral envelopes. Virus Research 37, Henkel, J. R. & Weisz, O. A. (1998). Influenza virus M2 protein slows traffic along the secretory pathway. ph perturbation of acidified compartments affects early Golgi transport steps. Journal of Biological Chemistry 273, Herz, C., Stavnezer, E., Krug, R. M. & Gurney, T., Jr (1981). Influenza virus, an RNA virus, synthesizes its messenger RNA in the nucleus of infected cells. Cell 26, Huang, T.-S., Palese, P. & Krystal, M. (1990). Determination of influenza virus proteins required for genome replication. Journal of Virology 64, Jambrina, E., Ba rcena, J., Uez, O. & Portela, A. (1997). The three subunits of the polymerase and the nucleoprotein of influenza B virus are the minimum set of viral proteins required for expression of a model RNA template. Virology 235, Jin, H., Leser, G. P. & Lamb, R. A. (1994). The influenza virus hemagglutinin cytoplasmic tail is not essential for virus assembly or infectivity. EMBO Journal 13, Jin, H., Leser, G. P., Zhang, J. & Lamb, R. A. (1997). Influenza virus hemagglutinin and neuraminidase cytoplasmic tails control particle shape. EMBO Journal 16, Klumpp, K., Ruigrok, R. W. H. & Baudin, F. (1997). Roles of the influenza virus polymerase and nucleoprotein in forming a functional RNP structure. EMBO Journal 16, Lamb, R. A. (1989). Genes and proteins of the influenza viruses. In The Influenza Viruses, pp Edited by R. M. Krug. New York: Plenum. Lamb, R. A. & Krug, R. M. (1996). Orthomyxoviridae: the viruses and their replication. In Fields Virology, 3rd edn, pp Edited by B. N. Fields, D. M. Knipe & P. M. Howley. Philadelphia: Lippincott-Raven. Lamb, R. A. & Lai, C.-J. (1980). Sequence of interrupted and uninterrupted mrnas and cloned DNA coding for the two overlapping nonstructural proteins of influenza virus. Cell 21, Lamb, R. A., Lai, C.-J. & Choppin, P. W. (1981). Sequences of mrnas derived from genome RNA segment 7 of influenza virus: colinear and interrupted mrnas code for overlapping proteins. Proceedings of the National Academy of Sciences, USA 78, Liu, C., Eichelberger, M. C., Compans, R. W. & Air, G. M. (1995). Influenza type A virus neuraminidase does not play a role in viral entry, replication, assembly, or budding. Journal of Virology 69, Lo pez, J. A., Guillen, M., Sa nchez-fauquier, A. & Melero, J. A. (1986). An antigen-binding assay to determine the specificity of monoclonal antibodies against influenza virus and mapping of epitopes. Journal of Virological Methods 13, Mario n, R. M., Zu rcher, T., de la Luna, S. & Ortı n, J. (1997). Influenza virus NS1 protein interacts with viral transcription replication complexes in vivo. Journal of General Virology 78, Martin, K. & Helenius, A. (1991). Nuclear transport of influenza virus ribonucleoproteins: the viral matrix protein (M1) promotes export and inhibits import. Cell 67, Mena, I., de la Luna, S., Albo, C., Martı n, J., Nieto, A., Ortı n, J. & Portela, A. (1994). Synthesis of biologically active influenza virus core proteins using a vaccinia virus T7 RNA polymerase expression system. Journal of General Virology 75, Mena, I., Vivo, A., Pe rez, E. & Portela, A. (1996). Rescue of a synthetic chloramphenicol acetyltransferase RNA into influenza virus-like particles obtained from recombinant plasmids. Journal of Virology 70, Mitnaul, L. J., Castrucci, M. R., Murti, K. G. & Kawaoka, Y. (1996). The cytoplasmic tail of influenza A virus neuraminidase (NA) affects NA incorporation into virions, virion morphology, and virulence in mice but is not essential for virus replication. Journal of Virology 70, Murti, K. G., Brown, P. S., Bean, W. J., Jr & Webster, R. G. (1992). Composition of the helical internal components of influenza virus as revealed by immunogold labeling electron microscopy. Virology 186, Nemeroff, M. E., Barabino, S. M. L., Li, Y., Keller, W. & Krug, R. M. (1998). Influenza virus NS1 protein interacts with the cellular 30 kda subunit of CPSF and inhibits 3 end formation of cellular pre-mrnas. Molecular Cell 1, Odagiri, T. & Tobita, K. (1990). Mutation in NS2, a nonstructural protein of influenza A virus, extragenically causes aberrant replication and expression of the PA gene and leads to generation of defective interfering particles. Proceedings of the National Academy of Sciences, USA 87, O Neill, R. E., Talon, J. & Palese, P. (1998). The influenza virus NEP (NS2 protein) mediates the nuclear export of viral ribonucleoproteins. EMBO Journal 17, Ortı n, J. (1998). Multiple levels of posttranscriptional regulation of influenza virus gene expression. Seminars in Virology 8, Palese, P., Tobita, K., Ueda, M. & Compans, R. W. (1974). Characterization of temperature sensitive influenza virus mutants defective in neuraminidase. Virology 61, Pattnaik, A. K. & Wertz, G. W. (1990). Replication and amplification of defective interfering particle RNAs of vesicular stomatitis virus in cells expressing viral proteins from vectors containing cloned cdnas. Journal of Virology 64, Pattnaik, A. K. & Wertz, G. W. (1991). Cells that express all five proteins of vesicular stomatitis virus from cloned cdnas support replication, assembly, and budding of defective interfering particles. Proceedings of the National Academy of Sciences, USA 88, BGEE
11 Influenza VLPs and viral protein expression Piccone, M. E., Ferna ndez-sesma, A. & Palese, P. (1993). Mutational analysis of the influenza virus vrna promoter. Virus Research 28, Pinto, L. H., Holsinger, L. J. & Lamb, R. A. (1992). Influenza virus M2 protein has ion channel activity. Cell 69, Sakaguchi, T., Leser, G. P. & Lamb, R. A. (1996). The ion channel activity of the influenza virus M2 protein affects transport through the Golgi apparatus. Journal of Cell Biology 133, Sa nchez-fauquier, A., Villanueva, N. & Melero, J. A. (1987). Isolation of cross-reactive, subtype-specific monoclonal antibodies against influenza virus HA1 and HA2 hemagglutinin subunits. Archives of Virology 97, Sanz-Ezquerro, J. J., de la Luna, S., Ortı n, J. & Nieto, A. (1995). Individual expression of influenza virus PA protein induces degradation of coexpressed proteins. Journal of Virology 69, Shih, S. R., Nemeroff, M. E. & Krug, R. M. (1995). The choice of alternative 5 splice sites in influenza virus M1 mrna is regulated by the viral polymerase complex. Proceedings of the National Academy of Sciences, USA 92, Sugrue, R. J., Bahadur, G., Zambon, M. C., Hall-Smith, M., Douglas, A. R. & Hay, A. J. (1990). Specific structural alteration of the influenza haemagglutinin by amantadine. EMBO Journal 9, Teng, M. N. & Collins, P. L. (1998). Identification of the respiratory syncytial virus proteins required for formation and passage of helperdependent infectious particles. Journal of Virology 72, Valca rcel, J., Portela, A. & Ortı n, J. (1991). Regulated M1 mrna splicing in influenza virus-infected cells. Journal of General Virology 72, Whittaker, G., Bui, M. & Helenius, A. (1996). The role of nuclear import and export in influenza virus infection. Trends in Cell Biology 6, Yang, P., Bansal, A., Liu, C. & Air, G. M. (1997). Hemagglutinin specificity and neuraminidase coding capacity of neuraminidase-deficient influenza viruses. Virology 229, Yasuda, J., Nakada, S., Kato, A., Toyoda, T. & Ishihama, A. (1993). Molecular assembly of influenza virus: association of the NS2 protein with virion matrix. Virology 196, Zhirnov, O. P. (1992). Isolation of matrix protein M1 from influenza viruses by acid-dependent extraction with nonionic detergent. Virology 186, Zvonarjev, A. Y. & Ghendon, Y. Z. (1980). Influence of membrane (M) protein on influenza A virus virion transcriptase activity in vitro and its susceptibility to rimantadine. Journal of Virology 33, Received 6 January 1999; Accepted 16 March 1999 BGEF
Plasmid-Driven Formation of Influenza Virus-Like Particles
 JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza Virus Matrix Protein Is the Major Driving Force in Virus Budding
 JOURNAL OF VIROLOGY, Dec. 2000, p. 11538 11547 Vol. 74, No. 24 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Influenza Virus Matrix Protein Is the Major
JOURNAL OF VIROLOGY, Dec. 2000, p. 11538 11547 Vol. 74, No. 24 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Influenza Virus Matrix Protein Is the Major
Reverse Genetics of RNA Viruses
 Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus from eight plasmids
 Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
The accumulation of influenza A virus segment 7 spliced mrnas is regulated by the NS1 protein
 Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
 http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Influenza or flu is a
 Clinical and Research Area Infectious Diseases Influenza Virus Types A and B Influenza or flu is a respiratory illness that is caused by influenza viruses. Influenza viruses type A and type B cause seasonal
Clinical and Research Area Infectious Diseases Influenza Virus Types A and B Influenza or flu is a respiratory illness that is caused by influenza viruses. Influenza viruses type A and type B cause seasonal
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Viral reproductive cycle
 Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Size nm m m
 1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
Temperature-Sensitive Mutants Isolated from Hamster and
 JOURNAL OF VIROLOGY, Nov. 1975, p. 1332-1336 Copyright i 1975 American Society for Microbiology Vol. 16, No. 5 Printed in U.S.A. Temperature-Sensitive Mutants Isolated from Hamster and Canine Cell Lines
JOURNAL OF VIROLOGY, Nov. 1975, p. 1332-1336 Copyright i 1975 American Society for Microbiology Vol. 16, No. 5 Printed in U.S.A. Temperature-Sensitive Mutants Isolated from Hamster and Canine Cell Lines
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
Ralf Wagner Paul-Ehrlich-Institut
 www.pei.de Other Assays for the Detection of Neuraminidase (NA)-Specific Antibodies Ralf Wagner Paul-Ehrlich-Institut Overview to presented assays Assay principle based on: Chemical substrates: Protein
www.pei.de Other Assays for the Detection of Neuraminidase (NA)-Specific Antibodies Ralf Wagner Paul-Ehrlich-Institut Overview to presented assays Assay principle based on: Chemical substrates: Protein
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP
 Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome
 Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
~Lentivirus production~
 ~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
Virology Introduction. Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment.
 DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
Influenza A H1N1 HA ELISA Pair Set
 Influenza A H1N1 HA ELISA Pair Set for H1N1 ( A/Puerto Rico/8/1934 ) HA Catalog Number : SEK11684 To achieve the best assay results, this manual must be read carefully before using this product and the
Influenza A H1N1 HA ELISA Pair Set for H1N1 ( A/Puerto Rico/8/1934 ) HA Catalog Number : SEK11684 To achieve the best assay results, this manual must be read carefully before using this product and the
MINIREVIEW. Recovery of Negative-Strand RNA Viruses from Plasmid DNAs: A Positive Approach Revitalizes a Negative Field
 VIROLOGY 247, 1 6 (1998) ARTICLE NO. VY989250 MINIREVIEW Recovery of Negative-Strand RNA Viruses from Plasmid DNAs: A Positive Approach Revitalizes a Negative Field Anjeanette Roberts and John K. Rose
VIROLOGY 247, 1 6 (1998) ARTICLE NO. VY989250 MINIREVIEW Recovery of Negative-Strand RNA Viruses from Plasmid DNAs: A Positive Approach Revitalizes a Negative Field Anjeanette Roberts and John K. Rose
Nature Medicine: doi: /nm.4322
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
Patricia Fitzgerald-Bocarsly
 FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
Rama Nada. - Malik
 - 2 - Rama Nada - - Malik 1 P a g e We talked about HAV in the previous lecture, now we ll continue the remaining types.. Hepatitis E It s similar to virus that infect swine, so its most likely infect
- 2 - Rama Nada - - Malik 1 P a g e We talked about HAV in the previous lecture, now we ll continue the remaining types.. Hepatitis E It s similar to virus that infect swine, so its most likely infect
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
HIV-1 p24 ELISA Pair Set Cat#: orb54951 (ELISA Manual)
 HIV-1 p24 ELISA Pair Set Cat#: orb54951 (ELISA Manual) BACKGROUND Human Immunodeficiency Virus ( HIV ) can be divided into two major types, HIV type 1 (HIV-1) and HIV type 2 (HIV-2). HIV-1 is related to
HIV-1 p24 ELISA Pair Set Cat#: orb54951 (ELISA Manual) BACKGROUND Human Immunodeficiency Virus ( HIV ) can be divided into two major types, HIV type 1 (HIV-1) and HIV type 2 (HIV-2). HIV-1 is related to
Interaction of the Influenza Virus Nucleoprotein with the Cellular CRM1-Mediated Nuclear Export Pathway
 JOURNAL OF VIROLOGY, Jan. 2001, p. 408 419 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.408 419.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Interaction of
JOURNAL OF VIROLOGY, Jan. 2001, p. 408 419 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.408 419.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Interaction of
Reoviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Virus and Prokaryotic Gene Regulation - 1
 Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Overview of virus life cycle
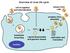 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Chapter 13B: Animal Viruses
 Chapter 13B: Animal Viruses 1. Overview of Animal Viruses 2. DNA Viruses 3. RNA Viruses 4. Prions 1. Overview of Animal Viruses Life Cycle of Animal Viruses The basic life cycle stages of animal viruses
Chapter 13B: Animal Viruses 1. Overview of Animal Viruses 2. DNA Viruses 3. RNA Viruses 4. Prions 1. Overview of Animal Viruses Life Cycle of Animal Viruses The basic life cycle stages of animal viruses
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
Unexpected complexity in the interference activity of a cloned influenza defective interfering RNA
 Meng et al. Virology Journal (2017) 14:138 DOI 10.1186/s12985-017-0805-6 RESEARCH Unexpected complexity in the interference activity of a cloned influenza defective interfering RNA Open Access Bo Meng
Meng et al. Virology Journal (2017) 14:138 DOI 10.1186/s12985-017-0805-6 RESEARCH Unexpected complexity in the interference activity of a cloned influenza defective interfering RNA Open Access Bo Meng
11/15/2011. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Viral Genomes. Structural Features and Characteristics
 Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide gel electrophoresis/genetics)
 Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Influenza A H7N9 (A/Anhui/1/2013) Hemagglutinin / HA ELISA Pair Set
 Influenza A H7N9 (A/Anhui/1/2013) Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK40103 To achieve the best assay results, this manual must be read carefully before using this product and the assay
Influenza A H7N9 (A/Anhui/1/2013) Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK40103 To achieve the best assay results, this manual must be read carefully before using this product and the assay
Viral Genetics. BIT 220 Chapter 16
 Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
HS-LS4-4 Construct an explanation based on evidence for how natural selection leads to adaptation of populations.
 Unit 2, Lesson 2: Teacher s Edition 1 Unit 2: Lesson 2 Influenza and HIV Lesson Questions: o What steps are involved in viral infection and replication? o Why are some kinds of influenza virus more deadly
Unit 2, Lesson 2: Teacher s Edition 1 Unit 2: Lesson 2 Influenza and HIV Lesson Questions: o What steps are involved in viral infection and replication? o Why are some kinds of influenza virus more deadly
Instructions. Fuse-It-mRNA easy. Shipping and Storage. Overview. Kit Contents. Specifications. Note: Important Guidelines
 Membrane fusion is a highly efficient method for transfecting various molecules and particles into mammalian cells, even into sensitive and primary cells. The Fuse-It reagents are cargo-specific liposomal
Membrane fusion is a highly efficient method for transfecting various molecules and particles into mammalian cells, even into sensitive and primary cells. The Fuse-It reagents are cargo-specific liposomal
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS
 Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Supplementary Figure 1. Prevalence of U539C and G540A nucleotide and E172K amino acid substitutions among H9N2 viruses. Full-length H9N2 NS nucleotide sequences (a, b) or amino acid sequences (c) from
Medical Virology. Herpesviruses, Orthomyxoviruses, and Retro virus. - Herpesviruses Structure & Composition: Herpesviruses
 Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recommended laboratory tests to identify influenza A/H5 virus in specimens from patients with an influenza-like illness
 World Health Organization Recommended laboratory tests to identify influenza A/H5 virus in specimens from patients with an influenza-like illness General information Highly pathogenic avian influenza (HPAI)
World Health Organization Recommended laboratory tests to identify influenza A/H5 virus in specimens from patients with an influenza-like illness General information Highly pathogenic avian influenza (HPAI)
Human Immunodeficiency Virus type 1 (HIV-1) p24 / Capsid Protein p24 ELISA Pair Set
 Human Immunodeficiency Virus type 1 (HIV-1) p24 / Capsid Protein p24 ELISA Pair Set Catalog Number : SEK11695 To achieve the best assay results, this manual must be read carefully before using this product
Human Immunodeficiency Virus type 1 (HIV-1) p24 / Capsid Protein p24 ELISA Pair Set Catalog Number : SEK11695 To achieve the best assay results, this manual must be read carefully before using this product
19 Viruses BIOLOGY. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Structural Features and Characteristics
 9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
The M2 Ectodomain Is Important for Its Incorporation into Influenza A Virions
 JOURNAL OF VIROLOGY, Mar. 1998, p. 2449 2455 Vol. 72, No. 3 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology The M2 Ectodomain Is Important for Its Incorporation into Influenza A
JOURNAL OF VIROLOGY, Mar. 1998, p. 2449 2455 Vol. 72, No. 3 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology The M2 Ectodomain Is Important for Its Incorporation into Influenza A
Influenza B Hemagglutinin / HA ELISA Pair Set
 Influenza B Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK11053 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
Influenza B Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK11053 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
Cover Page. The handle holds various files of this Leiden University dissertation
 Cover Page The handle http://hdl.handle.net/1887/35908 holds various files of this Leiden University dissertation Author: Soema, Peter Title: Formulation of influenza T cell peptides : in search of a universal
Cover Page The handle http://hdl.handle.net/1887/35908 holds various files of this Leiden University dissertation Author: Soema, Peter Title: Formulation of influenza T cell peptides : in search of a universal
Segment-specific and common nucleotide sequences in the
 Proc. Nati. Acad. Sci. USA Vol. 84, pp. 2703-2707, May 1987 Biochemistry Segment-specific and common nucleotide sequences in the noncoding regions of influenza B virus genome RNAs (viral transcription/viral
Proc. Nati. Acad. Sci. USA Vol. 84, pp. 2703-2707, May 1987 Biochemistry Segment-specific and common nucleotide sequences in the noncoding regions of influenza B virus genome RNAs (viral transcription/viral
Morphogenesis of Influenza Virus Particle
 !!! Morphogenesis of Influenza Virus Particle ( )!!!! CONTENTS! PREFACE 3 Chapter I The Native Morphology of Influenza Virions I.1 ABSTRACT 9 I.2 INTRODUCTION 11 I.3 MATERIALS AND METHODS 13 I.4 RESULTS
!!! Morphogenesis of Influenza Virus Particle ( )!!!! CONTENTS! PREFACE 3 Chapter I The Native Morphology of Influenza Virions I.1 ABSTRACT 9 I.2 INTRODUCTION 11 I.3 MATERIALS AND METHODS 13 I.4 RESULTS
Dr. Ahmed K. Ali Attachment and entry of viruses into cells
 Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Laboratory of Infectious Diseases, National Institute of Allergy and Infectious Diseases, 7 Center Drive MSC 0720, Bethesda, MD
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 11259 11264, September 1999 Biochemistry The M2 2 protein of human respiratory syncytial virus is a regulatory factor involved in the balance between RNA replication
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 11259 11264, September 1999 Biochemistry The M2 2 protein of human respiratory syncytial virus is a regulatory factor involved in the balance between RNA replication
7.012 Problem Set 6 Solutions
 Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
The Influenza A Virus M 2 Cytoplasmic Tail Is Required for Infectious Virus Production and Efficient Genome Packaging
 JOURNAL OF VIROLOGY, Mar. 2005, p. 3595 3605 Vol. 79, No. 6 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.6.3595 3605.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. The Influenza
JOURNAL OF VIROLOGY, Mar. 2005, p. 3595 3605 Vol. 79, No. 6 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.6.3595 3605.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. The Influenza
Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics
 Hepadnaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Hepatitis viruses A group of unrelated pathogens termed hepatitis viruses cause the vast majority
Hepadnaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Hepatitis viruses A group of unrelated pathogens termed hepatitis viruses cause the vast majority
ISG15 sirna # Ctrl sirna+ifn+wt. Virus titer (Pfu/ml) hours post infection. d USP18 sirna #2 IFN
 a ISG15 sirna Ctrl #1 #2 ISG15 conjugates IB: ISG15 Virus titer (Pfu/ml) b ISG15 sirna #2 10 7 Ctrl sirna+ifn+wt Ctrl sirna+ifn+67 10 6 ISG15 sirna+ifn+wt ISG15 sirna+ifn+67 10 5 10 4 10 3 Free ISG15 10
a ISG15 sirna Ctrl #1 #2 ISG15 conjugates IB: ISG15 Virus titer (Pfu/ml) b ISG15 sirna #2 10 7 Ctrl sirna+ifn+wt Ctrl sirna+ifn+67 10 6 ISG15 sirna+ifn+wt ISG15 sirna+ifn+67 10 5 10 4 10 3 Free ISG15 10
Virus Basics. General Characteristics of Viruses. Chapter 13 & 14. Non-living entities. Can infect organisms of every domain
 Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-forms Commonly referred to by organism they infect
Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-forms Commonly referred to by organism they infect
Relative activity (%) SC35M
 a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
The RNA Polymerase of Influenza Virus, Bound to the 5 End of Virion RNA, Acts in cis To Polyadenylate mrna
 JOURNAL OF VIROLOGY, Oct. 1998, p. 8214 8219 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. The RNA Polymerase of Influenza Virus, Bound to
JOURNAL OF VIROLOGY, Oct. 1998, p. 8214 8219 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. The RNA Polymerase of Influenza Virus, Bound to
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Influenza A virus hemagglutinin and neuraminidase act as novel motile machinery. Tatsuya Sakai, Shin I. Nishimura, Tadasuke Naito, and Mineki Saito
 Supplementary Information for: Influenza A virus hemagglutinin and neuraminidase act as novel motile machinery Tatsuya Sakai, Shin I. Nishimura, Tadasuke Naito, and Mineki Saito Supplementary Methods Supplementary
Supplementary Information for: Influenza A virus hemagglutinin and neuraminidase act as novel motile machinery Tatsuya Sakai, Shin I. Nishimura, Tadasuke Naito, and Mineki Saito Supplementary Methods Supplementary
 This article was published in an Elsevier journal. The attached copy is furnished to the author for non-commercial research and education use, including for instruction at the author s institution, sharing
This article was published in an Elsevier journal. The attached copy is furnished to the author for non-commercial research and education use, including for instruction at the author s institution, sharing
Virus Basics. General Characteristics of Viruses 5/9/2011. General Characteristics of Viruses. Chapter 13 & 14. Non-living entities
 Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-formsf Commonly referred to by organism they infect
Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-formsf Commonly referred to by organism they infect
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions
 Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication
 Journal of General Virology (2002), 83, 723 734. Printed in Great Britain... The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication Agustı n Portela 1 and
Journal of General Virology (2002), 83, 723 734. Printed in Great Britain... The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication Agustı n Portela 1 and
Identification of New Influenza B Virus Variants by Multiplex Reverse Transcription-PCR and the Heteroduplex Mobility Assay
 JOURNAL OF CLINICAL MICROBIOLOGY, June 1998, p. 1544 1548 Vol. 26, No. 6 0095-1137/98/$04.00 0 Copyright 1998, American Society for Microbiology Identification of New Influenza B Virus Variants by Multiplex
JOURNAL OF CLINICAL MICROBIOLOGY, June 1998, p. 1544 1548 Vol. 26, No. 6 0095-1137/98/$04.00 0 Copyright 1998, American Society for Microbiology Identification of New Influenza B Virus Variants by Multiplex
Chapter 13 Viruses, Viroids, and Prions. Biology 1009 Microbiology Johnson-Summer 2003
 Chapter 13 Viruses, Viroids, and Prions Biology 1009 Microbiology Johnson-Summer 2003 Viruses Virology-study of viruses Characteristics: acellular obligate intracellular parasites no ribosomes or means
Chapter 13 Viruses, Viroids, and Prions Biology 1009 Microbiology Johnson-Summer 2003 Viruses Virology-study of viruses Characteristics: acellular obligate intracellular parasites no ribosomes or means
Severe Acute Respiratory Syndrome (SARS) Coronavirus
 Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Regulation of a nuclear export signal by an adjacent inhibitory sequence: The effector domain of the influenza virus NS1 protein
 Proc. Natl. Acad. Sci. USA Vol. 95, pp. 4864 4869, April 1998 Biochemistry Regulation of a nuclear export signal by an adjacent inhibitory sequence: The effector domain of the influenza virus NS1 protein
Proc. Natl. Acad. Sci. USA Vol. 95, pp. 4864 4869, April 1998 Biochemistry Regulation of a nuclear export signal by an adjacent inhibitory sequence: The effector domain of the influenza virus NS1 protein
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Influenza Virus NS1 Protein Stimulates Translation of the Ml Protein
 JOURNAL OF VROLOGY, Mar. 1994, p. 1432-1437 Vol. 68, No. 3 0022-538X/94/$04.00+0 Copyright 1994, American Society for Microbiology nfluenza Virus Protein Stimulates Translation of the Ml Protein KAZUE
JOURNAL OF VROLOGY, Mar. 1994, p. 1432-1437 Vol. 68, No. 3 0022-538X/94/$04.00+0 Copyright 1994, American Society for Microbiology nfluenza Virus Protein Stimulates Translation of the Ml Protein KAZUE
Introductory Virology. Ibrahim Jamfaru School of Medicine UHAS
 Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Rabies virus-like particles expressed in HEK293 cells
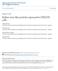 Engineering Conferences International ECI Digital Archives Vaccine Technology IV Proceedings Spring 5-21-2012 Rabies virus-like particles expressed in HEK293 cells Diego Fontana Cell Culture Laboratory
Engineering Conferences International ECI Digital Archives Vaccine Technology IV Proceedings Spring 5-21-2012 Rabies virus-like particles expressed in HEK293 cells Diego Fontana Cell Culture Laboratory
Overview: Chapter 19 Viruses: A Borrowed Life
 Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Influenza A H1N1 (Swine Flu 2009) Hemagglutinin / HA ELISA Pair Set
 Influenza A H1N1 (Swine Flu 2009) Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK001 To achieve the best assay results, this manual must be read carefully before using this product and the assay
Influenza A H1N1 (Swine Flu 2009) Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK001 To achieve the best assay results, this manual must be read carefully before using this product and the assay
Variation in the HindlII Restriction Fragments of DNA from the Chinese Tian Tan Strain of Vaccinia Virus
 J. gen. irol. (1985), 66, 1819-1823. Printed in Great Britain 1819 Key words: vaccinia virus~vaccine~restriction Jragrnent variation ariation in the Hindl Restriction Fragments of DNA from the Chinese
J. gen. irol. (1985), 66, 1819-1823. Printed in Great Britain 1819 Key words: vaccinia virus~vaccine~restriction Jragrnent variation ariation in the Hindl Restriction Fragments of DNA from the Chinese
Persistent Infection of MDCK Cells by Influenza C Virus: Initiation and Characterization
 J. gen. Virol. (199), 70, 341-345. Printed in Great Britain 341 Key words: influenza C virus/interferon/persistent infection Persistent Infection of MDCK Cells by Influenza C Virus: Initiation and Characterization
J. gen. Virol. (199), 70, 341-345. Printed in Great Britain 341 Key words: influenza C virus/interferon/persistent infection Persistent Infection of MDCK Cells by Influenza C Virus: Initiation and Characterization
Chapter 19: The Genetics of Viruses and Bacteria
 Chapter 19: The Genetics of Viruses and Bacteria What is Microbiology? Microbiology is the science that studies microorganisms = living things that are too small to be seen with the naked eye Microorganisms
Chapter 19: The Genetics of Viruses and Bacteria What is Microbiology? Microbiology is the science that studies microorganisms = living things that are too small to be seen with the naked eye Microorganisms
Fine Mapping of a cis-acting Sequence Element in Yellow Fever Virus RNA That Is Required for RNA Replication and Cyclization
 JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
