Whole Genome Sequencing of Influenza C Virus. Karin Pachler and Reinhard Vlasak Department of Molecular Biology University Salzburg, Austria
|
|
|
- Clifton McCarthy
- 5 years ago
- Views:
Transcription
1 Whole Genome Sequencing of Influenza C Virus Karin Pachler and Reinhard Vlasak Department of Molecular Biology University Salzburg, Austria
2 1 Introduction Influenza C virus belongs to the Orthomyxoviridae, a virus family divided into five different genera. These genera are influenzaviruses A, B, and C, Thogotovirus, and Isavirus (see Table 1). Orthomyxoviruses are characterised by a segmented single-stranded RNA genome of negative polarity. The genome of influenza A and B viruses is distributed among eight segments, while influenza C viruses harbour only seven RNA segments. Thogotovirus and Isavirus bear six and eight genomic segments, respectively. The seven influenza C virus RNA segments have a total size of approximately 12.9 kb. Genus Members Hosts Influenza A virus Human influenza virus Avian influenza virus Equine influenza virus Porcine influenza virus Human, bird, horse, pig, seal Number of Surface Protein(s) Segments 8 HA, NA Influenza B Influenza B virus Human, seal 8 HA, NA virus Influenza C Influenza C virus Human, pig, dog 7 HEF virus Thogotovirus Thogoto virus, Dhori virus Tick, cow, sheep, 6 Glycoprotein (G) goat, rodents Isavirus Infectious salmon anaemia virus salmon 8 HE Table 1: Overview of the virus family Orthomyxoviridae Influenza A, B, and C viruses not only differ in the number of genomic segments, but also in viral antigens and host range. While influenza B viruses mainly circulate in humans, influenza A viruses infect humans, domestic animals like pigs and horses, and wild birds. Influenza C viruses were isolated from humans and pigs, and the presence of specific antibodies was demonstrated in the serum of dogs as well (Brown et al., 1995, Manuguerra & Hannoun 1992, Manuguerra et al., 1993, Ohwada et al., 1987, Yamaoka et al., 1991). In contrast to influenza A and B viruses, which encode two major surface glycoproteins, influenza C viruses display only one major glycoprotein on their surfaces. Influenza C viruses, which are distributed worldwide, are divided into six antigenic and genetic groups (Matsuzaki et al., 2003, Muraki et al., 1996), and reassortment between cocirculating viruses frequently occurs (Matsuzaki et al., 1994, 2003; Peng et al., 1994). Currently, more than 160 different influenza C virus isolates are known 1. Like all influenza viruses, the influenza C virus is transmitted via aerosol droplets, and it infects the epithelial cells of the upper respiratory tract (Taylor 1951). Infection normally results in mild upper respiratory disease and sometimes in lower respiratory illness in young children (Katagiri et al., 1983, Katagiri et al., 1987, Matsuzaki et al., 2006). Influenza C viruses can provoke seasonal epidemics 1
3 (Matsuzaki et al., 2004). Influenza A viruses are best known to cause influenza virus epidemics and pandemics. Influenza B viruses account for seasonal epidemics of respiratory infections. 2 Structure of Influenza Virus Particle According to electron microscopic analyses, influenza viruses are enveloped viruses occurring in spherical or filamentous forms with an average diameter of 80 to 120 nm (Noda et al., 2006). Their lipid membrane, which is derived from the host cell, carries virus-encoded surface proteins (spikes). These are HA and NA for influenza A and B viruses, and HEF (haemagglutinin-esterase-fusion protein) for influenza C virus. In fig. 1, an influenza C virus particle is schematically illustrated. The HEF protein, which is present as homotrimers, combines the functions of influenza A and B virus HA and NA proteins. In addition, few copies of the glycosylated CM2 protein are inserted into the lipid membrane. The matrix protein (M1) lies beneath the envelope giving rigidity to the virion. M1 is the most abundant protein in the virion. The core of the influenza C virus particle is comprised of viral ribonucleoprotein (RNP) complexes consisting of the viral RNA segments, the three polymerase proteins PB2, PB1, and P3, and the nucleoprotein (NP). The seven negative-sense single-stranded RNA segments, which probably form panhandle/corkscrew-like structures, are fully associated with NP and with the polymerase proteins at their base-paired ends. The RNPs are associated with M1 via electrostatic attraction (Zhirnov & Grigoriev, 1994). Some copies of NEP/NS2 (nuclear export protein/nonstructural protein 2) are incorporated into the virion as well. On the surface of infected cells, influenza C virus was found to form long (500 µm) cord-like structures, and M1 was demonstrated as determinant of these formations (Muraki et al., 2007, Muraki et al., 2004, Nishimura et al., 1990, Nishimura et al., 1994). Figure 1: Schematic representation of an influenza C virus particle. The influenza C virus is made of seven single-stranded RNA segments of negative polarity assembled to vrnp complexes, a lipid membrane derived from the host cell displaying HEF spikes and CM2. M1 lies beneath the envelope, and a small amount of NEP/NS2 proteins is present in the virion.
4 3 Genome Structure and Encoded Proteins of Influenza C Virus Influenza C viruses have a genome of seven single-stranded negative-sense RNA segments. Negativesense means that this RNA is complementary to mrna ( positive sense ). These seven RNA segments code for nine viral proteins. In fig. 2, an overview of the influenza C virus genome is depicted. The three largest influenza C virus segments code for the polymerase proteins PB2, PB1, and P3. The fourth RNA encodes the HEF glycoprotein. The nucleoprotein NP is encoded by the fifth RNA. The RNA segments six and seven are bicistronic genes. The spliced mrna of segment six encodes the matrix protein M1. The unspliced mrna of the segment encodes protein P42, which is cleaved by a signal peptidase to give M1 and CM2 (Hongo et al. 1999, Pekosz & Lamb, 1998). CM2 is a surface glycoprotein, possibly with ion channel activity (Betakova & Hay, 2007, Hongo et al., 2004). RNA seven codes for NS1 (nonstructural protein 1) and, via a spliced mrna, for NEP/NS2 protein. NS1 is an interferon antagonist (Pachler & Vlasak, 2011). NEP/NS2 possesses nuclear export activity (Paragas et al. 2001). It is thought to be engaged in the export of newly synthesised vrnps. The coding regions of the RNAs from all influenza viruses are flanked by 3 and 5 noncoding (nc) ends. The ultimate nucleotides of the nc ends are conserved among all segments of the respective influenza genus, and these conserved regions are followed by segment-specific nc regions. Figure 2: Genome structure of influenza C/JJ/50 virus. The seven RNA segments are shown in positive-sense orientation. Numbers indicate the nucleotide positions along the segments. The lines at the 3 and 5 ends represent the noncoding regions. The V-shaped lines in segments six and seven indicate introns. M1 is encoded by a spliced transcript of segment six, into which stop codon TGA is introduced as a result of splicing. P42 is encoded by a collinear transcript of segment six. It is cleaved by a signal peptidase at an internal cleavage site (black arrow) to generate M1 and CM2. RNA segment seven codes for NS1 and, from a spliced mrna, for NEP/NS2. In the C-terminal region of NEP/NS2 (grey area) the ORF is +1.
5 4 The Different RNA Species in Influenza Viruses and the Role of the Noncoding RNA Ends The influenza virions are packaged with segments of negative-sense vrna. Each of these genomic vrnas contains noncoding sequences at its 3 and 5 ends that flank the coding region. The sequence and length of the nc ends vary between the vrna segments. For influenza C virus strain C/JJ/50, the 5 nc region consists of 19 nucleotides (segment PB2) to 102 nucleotides (segment NP), and the 3 nc region has between 17 nucleotides (segment PB1) and 29 nucleotides (segment NP). But the ultimate nucleotides of the nc ends are conserved between all segments. Table 2 shows these conserved ends for an influenza C virus strain. In the course of the influenza virus life cycle, the vrnas are transcribed into mrnas. The mrnas are incomplete complementary copies of the vrnas and are capped and polyadenylated. For replication of the viral genome, full-length positive-sense copies of the vrnas are generated as intermediate products. These RNA species are termed complementary RNAs (crnas) and are complete complementary copies of the vrnas. The crnas in turn serve as templates to manifold the vrnas (reviewed in Neumann et al., 2004). Segment 3 nc end (ultimate 20 nucleotides) a 5 nc end (first 20 nucleotides) a PB2 b UCCAAUCCUCUGCUUCUGCU AGCAGUAGCAAGAGGAUUUU PB1 b CAUAAUCCUCUGCUUCUGCU AGCAGUAGCAAGAGGAUUUU P3 UCGGAUCCCCUGCUUCUGCU AGCAGUAGCAAGGGGAUUUU HEF AUUAAACCCCUGCUUCUGCU AGCAGUAGCAAGGGGAUUUU NP b CAAAAUCUCCUGCUUCUGCU AGCAGUAGCAAGGAGAUUUU M AGAAAUCCCCUGCUUCUGCU AGCAGUAGCAAGGGGAUUUU NS b AAAGUACCCCUGCUUCUGCU AGCAGGAGCAAGGGGUUUUU Table 2: Comparison of the conserved 3 and 5 noncoding ends of the vrna segments from influenza C/JJ/50 virus (from Pachler et al., 2010). a Underlined: Conserved sequences. b In bold: Variations at positions 12, 13, 6, 13 and 14 (5 nucleotides are marked with a prime). Note that sequences are shown in 5-3 orientation and numbering of nucleotides starts from both ends. For proper transcription and replication of the vrnas, the nc regions play a crucial role. Most studies to elucidate nature and function of the nc ends were carried out with influenza A viruses. The conserved 3 and 5 ends were found to be partially complementary to each other, which allows a basepaired double-stranded region of four to eight nucleotides between the 3 and 5 ends for influenza A virus strains. This partially double-stranded region was first described as a panhandle/fork structure (Fodor et al., 1994, Hsu et al., 1987). Later, the corkscrew model was introduced (Flick & Hobom 1999, Flick et al., 1996) suggesting that the base-paired region is flanked by local secondary structures at the ultimate nucleotides (see figure 3A). The conserved noncoding ends were found to be required for activity of vrna and crna promoters (Fodor et al., 1995, Fodor et al., 19945, Pritlove et al., 1995, Tiley et al., 1994), endonuclease activity of the viral polymerase complex (Cianci et al., 1995, Hagen et al., 1994), and mrna polyadenylation (Li & Palese, 1994, Luo et al., 1991, Pritlove et al., 1998).
6 The importance of the nc regions was also proved for influenza C viruses. Crescenzo-Chaigne et al. (Crescenco-Chaigne et al., 1999, Crescenco-Chaigne & van der Werf 2001) found that a model RNA flanked by the nc regions of segment NS from influenza C/JHB/1/66 virus was transcribed in COS-1 cells under the presence of the viral polymerase proteins (PB2, PB1, P3) and the nucleoprotein (NP). They could thus demonstrate that the nc regions of influenza C virus are involved in replication and transcription of the viral genome. In a study to generate recombinant influenza C viruses (Pachler et al. 2010), we found that a single mutation within the proposed double-stranded region of the nc ends of one segment prevented the rescue of recombinant influenza C viruses. The single mutation was at position 13 of the 5 -end of segment PB1 and abolished base-pairing of nucleotides 13 and 12 (see figure 3B, note that 5 nucleotides are marked with a prime). When this single mutation was complemented by a mutation at position 12 of the 3 -end of PB1, which restored base-pairing between nucleotides 13 and 12, a recombinant virus was gained. But this virus was impaired in growth. Therefore, not only the ability to form base-paired parts, but also the exact nucleotide sequence at the panhandle/corkscrew region of the nc ends is of major importance for a proper viral life cycle. Figure 3: Representation of the noncoding terminal sequences of a viral segment (segment PB1) of influenza C virus in the corkscrew model in A (Crescenzo-Chaigne et al., 2001, Flick et al., 1996). In B, single and base-pair mutations at positions 13 and 12 within the proposed six base pairs long double-stranded region of the nc end of PB1 from influenza C virus are depicted. Picture taken from Pachler et al., Strategy for Whole Genome Sequencing of Influenza C Virus As outlined in the previous chapter, the nucleotide sequences of the nc viral ends are extremely important for viral life cycle. Therefore, a thorough approach to determine the entire genome of an influenza virus must include the nc ends as well. In contrast to DNA, which can be sequenced directly and bears two complementary strands both applicable for sequencing, the influenza virus RNA must first be reverse-
7 transcribed and PCR-amplified. As a consequence, sequences beyond both ends are needed for primer annealing to produce PCR fragments, which may then be sequenced. And to obtain such sequences beyond the ends is the major challenge in sequencing an RNA virus. To identify the viral 5 - and 3 -ends of all seven RNA segments from influenza virus strain C/JJ/50, we applied the SMART-RACE technique. 5.1 Overview of SMART-RACE Method The SMART technology was developed by Clontech, Inc. SMART stands for switching mechanism at RNA termini and is a method for synthesis of first strand cdna from a 3 polyadenylated RNA template (see figure 4). The conversion of RNA to cdna is catalysed by M-MuLV-RT (Moloney Murine Leukemia Virus Reverse Transcriptase) mutated in the RNase H domain. M-MuLV-RT possesses not only reverse transcription activity, but also terminal deoxynucleotidyl transferase (TdT) activity. It has therefore the very useful feature to add a few nontemplate deoxycytosines (d(c)) to the 3 -end of a newly synthesised cdna strand. Reverse transcription of the RNA starts from an oligo-d(t) primer that anneals to the poly(a) tail of the RNA (primer TRsa in fig. 4). The oligo-d(t) primer additionally contains a defined sequence at the 5 -end, which serves as target site for the subsequent PCR amplification of the viral 3 -end. Figure 4: Schematic outline of cdna synthesis using the SMART method with oligo-(d)t primer TRsa and template-switching primer TS according to Matz (Matz 2003). A second primer is added to the RT reaction, the so-called template-switch (TS) primer. This primer harbours an oligo(rg) sequence at its 3 -end. Upon reaching the 5 -end of the RNA template, the M- MuLV-RT adds a few nontemplate d(c) residues to the 3 -end of the cdna due to its TdT activity. The oligo(rg) of the TS primer consequently anneals to the un-templated d(c) residues, and the M-MuLV-RT switches the templates and continues reverse transcription with the TS primer as template. The TS primer also contains a defined sequence at its 5 -end, which is then the target site for primer annealing in the subsequent PCR amplification of the viral 5 -end. With this technique, first strand cdna is generated
8 with defined 5 - and 3 -end sequences flanking the sequence of the starting RNA. Subsequently, the unknown 5 - and 3 -ends of the RNA can be amplified by RACE-PCR (rapid amplification of cdna ends- PCR). For RACE-PCR (see fig. 5), the 5 -end of the cdna (corresponding to the 3 -end of the vrna) is PCR-amplified with a primer annealing to the cdna internally together with the oligo-d(t) primer (TRsa). The 3 -end of the cdna (corresponding to the 5 -end of the vrna) is manifolded with a primer containing the defined sequence of the TS oligo (but lacking the oligo(rg), termed TS-PCR) along with an internal primer specific for the cdna. The PCR fragments can then be sequenced. Figure 5: Schematic outline of 5 - and 3 -RACE-PCR for amplification of viral noncoding regions (ncr) starting from cdna. 5.2 Protocol for Sequencing the Influenza C/JJ/50 Virus Genome We performed the SMART-RACE technique with modifications according to Matz (Matz 2003). As the starting material required is polyadenylated RNA, we first attached a poly(a) tail to the 3 end of isolated vrna with a commercially available poly(a) kit. For SMART, we added the oligos TRsa (5 -CGCAGTCGGTACTTTTTTTTTTTTTTTTTTCA- 3 ) and TS (5 -AAGCAGTGGTATCAACGCAGAGTACGCrGrGrG-3 ). At the end of the first strand cdna synthesis reaction, MnCl 2 was added to the reaction mix, as it increases the efficiency of nontemplate C addition to the cdna and thus results in higher yield following cdna amplification (Matz 2003). The exact protocol for cdna synthesis was the following: First strand cdna synthesis µl poly(a)-vrna, containing approximately 5 µg vrna - 1 µl primer TRsa (10 µm) - 1 µl primer TS (10 µm) incubated at 65 C for 5 minutes and at 4 C for 5 minutes, then addition of: - 4 µl 5x M-MuLV RT buffer - 2 µl dntps (10 mm) µl RNasin Plus RNase inhibitor (20 U), Promega incubated at 37 C for 5 minutes, then addition of: - 1 µl M-MuLV RT (200 U, RevertAid TM H Minus RT, Fermentas)
9 incubated at 42 C for 1 hour, then addition of: - 2 µl MnCl 2 (25 mm) incubated at 42 C for 15 minutes and at 70 C for 10 minutes, kept at 4 C This cdna was then used as template for amplification of the nc ends from all seven viral segments. For each influenza C virus segment, the 5 nc end was manifolded with an internal primer specific for the respective viral segment s coding region and primer TS-PCR (5 - AAGCAGTGGTATCAACGCAGAGT-3 ), while the 3 end was manifolded with primer TRsa together with a primer specific for the segment s coding region. (see fig. 5). Sequences for segment-specific internal forward and reverse primers were taken from published sequences of other influenza C viruses. In the course of determining the genomic sequences of the seven entire segments from strain C/JJ/50, the regions serving as internal primer binding sites for nc ends amplification were exactly sequenced as well. The exact protocol for RACE-PCR was the following: RACE-PCR reaction mix (25 µl): µl cdna µl primer TRsa or primer TS-PCR (both 10 µm) µl segment-specific reverse primer or segment-specific forward primer (both 10 µm) µl dntps (2 mm) - 5 µl 5x buffer µl DNA polymerase (Herculase II Fusion DNA polymerase, Stratagene) µl H 2 O PCR programme: Denaturation 95 C 20 sec. Denaturation 95 C 20 sec. Annealing 60 C 1 min. 30 cycles Elongation 72 C 1 min. Final elongation 72 C 4 min. Final hold 4 C After amplification of the ends from all seven segments, the ends could be sequenced by standard methods, employing internal segment-specific primers. Once the ends of the seven viral segments had been determined (see table 2 for sequences of the conserved viral ends), the full-length segments were PCR-amplified with primers complementary to the last 12 nucleotides of the 5 and 3 nc ends. The entire segments were then sequenced with overlapping internal primers. Again, the sequences of these internal primers were taken from published sequences of other influenza C virus strains, and the regions, where the internal sequencing primers had annealed to, were covered by sequencing of overlapping regions. The resulting partial sequences can then be compiled and aligned with the help alignment software. Crescenzo-Chaigne (Crescenco-Chaigne & van der Werf 2007) applied a different method to identify the nc ends of influenza C/JHB/1/66 virus. For sequencing of the 3 -ends, they first also polyadenylated the vrnas. cdna fragments complementary to the 3 -viral ends were then prepared with AMV-RT (Avian Myeloblastosis Virus Reverse Transcriptase) and an oligo-d(t) as primer. Amplification of the
10 fragments was performed with an anchored oligo-d(t) together with an internal primer specific for the coding sequence of each of the seven vrnas. For sequencing of the 5 -ends, they first reversetranscribed each of the seven vrnas with primers specific for the respective coding sequences. The cdnas were then elongated with a TdT in the presence of datp. Afterwards, the cdnas containing poly(a) were amplified in two steps with an anchored oligo-d(t) primer together with oligos specific for the coding region of each segment. This protocol is a bit more laborious than the SMART-RACE-PCR protocol we applied. With the SMART-RACE-PCR method, only a single reverse transcription reaction is required to obtain cdnas of all seven influenza C virus segments covering both viral ends at once. In contrast, with the protocol used by Crescenzo-Chaigne (Crescenco-Chaigne & van der Werf 2007), one reverse transcription reaction has to be performed to get cdnas of the seven viral 3 -ends, and seven separate reverse transcription reactions are necessary to obtain cdnas of every segment s 5 -end. Moreover, sequence analysis of the 5 -ends requires an additional polyadenylation step of the seven cdnas as well as two rounds of PCR amplification. In summary, this protocol is more time-consuming and also more vrna template-consuming. On the other hand, weak template switching during cdna synthesis according to the SMART method can lead to low PCR product yields for the viral 5 -ends. Yields may be higher with the more complicated protocol described by Crescenzo-Chaigne (Crescenco-Chaigne & van der Werf 2007), where cdnas of the seven 5 -ends are polyadenylated prior to PCR amplification. For larger vrnas, the SMART method may also require to synthesise cdnas corresponding to the viral 5 - and 3 -ends separately. However, the SMART-RACE-PCR is a simple and fast method for sequence analysis. With the sequence information obtained by the SMART-RACE-PCR method for influenza C virus strain C/JJ/50, we were able to rescue recombinant influenza C viruses and to study the effect of mutations within the nc ends (Pachler et al., 2010). SMART-PCR can be applied for sequence determination of other RNA viruses as well. Any RNA can be manifolded with this method, and gene hunters often use this technique to reverse-transcribe and amplify mrna pools from diverse tissues. 6 Comparison of the Genomic Sequences of Different Influenza C Virus Strains In the 1980s, the complete cdna sequences including the nc regions of segments PB2, PB1, P3, and M from influenza C/JJ/50 virus were published (Yamashita et al., 1989, Yamashita et al., 1988), as well as partial sequences from other influenza C virus strains (Clern-van Haaster, 1984, Desselberger et al, 1980). Applying the SMART-RACE technique, we could easily sequence the nc regions of all seven segments from influenza C/JJ/50 virus and afterwards elucidate the entire genomic sequence of this strain (see table 3). We identified some differences in comparison to the published sequences from the 1980s (Pachler et al. 2010). Only segment P3 from C/JJ/50 does not show any difference compared to the one sequenced by Yamashita et al. (Yamashita et al., 1989). Two other groups have also sequenced the entire genomes of other influenza C virus strains, namely influenza C/JHB/1/66 virus (Crescenzo-Chaigne & van der Werf 2007) and influenza C/Ann Arbor/1/50 virus (Muraki et al., 2007). With the aid of alignment programmes (like Multalin, or ClustalW2, the nucleotide sequences of different influenza
11 virus strains can be compared. In such an approach to compare the nucleotide sequences of the three fully sequenced influenza C virus strains, we found high sequence identities (97 to 99.15%) between the respective segments of these strains (see table 4). Notable differences were within segments NP and NS. The NP segments of strains C/JJ/50 and C/JHB/1/66 have both a size of 1802 nt, while the respective segment of strain C/Ann Arbor/1/50 is 5 nt longer. This difference lies within the open reading frame (ORF) of NP and leads to a later stop codon for strain C/Ann Arbor/1/50. Therefore, the NP of C/Ann Arbor/1/50 is 9 amino acids longer, while the 5 nc end is 22 nt shorter. vrna Segment Number of nucleotides Total 3 ncr Coding region 5 ncr Accession no. a PB FR PB FR P M28062 b HEF FR NP FR M FR NS FR Table 3: The whole genome of influenza C/JJ/50 virus. a GenBank/EMBL/DDJ accession numbers, b Sequence M28062 was deposited at the GenBank Database by Yamashita et al The NS segment of strain C/JJ/50 is 21 nt shorter than the NS segments of C/JHB/1/66 and C/Ann Arbor/1/50. This deletion is located within the ORF of NS1, which is therefore 7 amino acids shorter than the NS1 proteins of the two other strains. vrna Number of Sequence Difference in Difference in segment nucleotides identity 3 ncr 5 ncr Difference in cr a PB % nt PB % nt P % - 1 nt 41 nt HEF % 1 nt 7 nt 55 nt NP 1802/1807 b 98.4% 2 nt 12 nt 15 nt c M % 3 nt - 17 nt NS 914/935 d 98.9% 1 nt - 9 nt e Table 4: Comparison of nucleotide sequences of all viral segments from the influenza virus strains C/JJ/50, C/JHB/1/66, and C/Ann Arbor/1/50. a coding region, b NP from strain C/Ann Arbor/1/50 has a size of 1807 nt, c Besides this difference, NP from C/Ann Arbor/1/50 has additional 5 nt in the ORF leading to later stop codon, d NS from strain C/JJ/50 has a size of 914 nt, e In addition to this difference, NS from C/JJ/50 has 21 nt less within the cr, leading to a shorter NS1 protein. The differences in the nc regions between these three influenza C virus strains, as shown in table 4, do not affect the ultimate parts of the nc ends. The ultimate parts of the nc ends the regions conserved among the different segments are also conserved among the three virus strains (see Table 2). Regarding the 3 -ends, the first eleven nucleotides and nucleotide 14 are identical in all seven segments,
12 whereas nucleotides 12 and 13 show a slight variability. In the 5 -ends, the first twelve nucleotides as well as nucleotide 15 are conserved among the segments, with exception of segment NS, where nucleotide 6 differs from the others. In all segments from these three virus strains, nucleotides (3 -viral ends) are complementary to nucleotides (5 -viral ends), which would allow formation of a double-stranded structure of five base pairs at this region. For segments PB2, PB1, P3, NP, and M, even six base pairs would be possible. Besides the alignment of several sequences via Multalin or ClustalW2, it is also possible to search for sequence similarities between a sequence of interest and sequences deposited at the GenBank Database, for example with the Basic Local Alignment Search Tool (BLAST) from the National Center for Biotechnology Information, ncbi ( 7 Glossary AMV-RT BLAST cdna CM2 HEF M1 M-MuLV RT mrna nc ncbi ncr nt NEP (NS2) NP NS1 NS2 ORF P3 PB1 PB2 RACE-PCR RNP RT SMART TdT TS vrna Avian Myeloblastosis Virus Reverse Transcriptase Basic Local Alignment Search Tool complementary DNA surface glycoprotein from influenza C virus haemagglutinin-esterase-fusion protein from influenza C virus matrix protein from influenza C virus Moloney Murine Leukemia Virus Reverse Transcriptase messenger RNA noncoding National Center for Biotechnology Information noncoding region nucleotide nuclear export protein (nonstructural protein) from influenza virus nucleoprotein from influenza virus nonstructural protein 1 from influenza virus nonstructural protein 2 from influenza virus open reading frame polymerase protein from influenza C virus polymerase protein from influenza virus polymerase protein from influenza virus rapid amplification of cdna ends polymerase chain reaction ribonucleoprotein reverse transcription/ reverse transcriptase switching mechanism at RNA termini terminal deoxynucleotidyl transferase template switch viral RNA
13 References Betakova, T., and A. J. Hay Evidence that the CM2 protein of influenza C virus can modify the ph of the exocytic pathway of transfected cells. J Gen Virol 88: Brown, I. H., P. A. Harris, and D. J. Alexander Serological studies of influenza viruses in pigs in Great Britain Epidemiol Infect 114: Cianci, C., L. Tiley, and M. Krystal Differential activation of the influenza virus polymerase via template RNA binding. J Virol 69: Clern-van Haaster, C. M., and H. Meier-Ewert '-Terminal sequences of influenza C virion RNA. Brief report. Arch Virol 80: Crescenzo-Chaigne, B., N. Naffakh, and S. van der Werf Comparative analysis of the ability of the polymerase complexes of influenza viruses type A, B and C to assemble into functional RNPs that allow expression and replication of heterotypic model RNA templates in vivo. Virology 265: Crescenzo-Chaigne, B., and S. van der Werf Nucleotides at the extremities of the viral RNA of influenza C virus are involved in type-specific interactions with the polymerase complex. J Gen Virol 82: Crescenzo-Chaigne, B., and S. van der Werf Rescue of influenza C virus from recombinant DNA. J Virol 81: Desselberger, U., V. R. Racaniello, J. J. Zazra, and P. Palese The 3' and 5'-terminal sequences of influenza A, B and C virus RNA segments are highly conserved and show partial inverted complementarity. Gene 8: Flick, R., and G. Hobom Interaction of influenza virus polymerase with viral RNA in the 'corkscrew' conformation. J Gen Virol 80 ( Pt 10): Flick, R., G. Neumann, E. Hoffmann, E. Neumeier, and G. Hobom Promoter elements in the influenza vrna terminal structure. Rna 2: Fodor, E., D. C. Pritlove, and G. G. Brownlee Characterization of the RNA-fork model of virion RNA in the initiation of transcription in influenza A virus. J Virol 69: Fodor, E., D. C. Pritlove, and G. G. Brownlee The influenza virus panhandle is involved in the initiation of transcription. J Virol 68: Hagen, M., T. D. Chung, J. A. Butcher, and M. Krystal Recombinant influenza virus polymerase: requirement of both 5' and 3' viral ends for endonuclease activity. J Virol 68: Hongo, S., K. Ishii, K. Mori, E. Takashita, Y. Muraki, Y. Matsuzaki, and K. Sugawara Detection of ion channel activity in Xenopus laevis oocytes expressing Influenza C virus CM2 protein. Arch Virol 149: Hongo, S., K. Sugawara, Y. Muraki, Y. Matsuzaki, E. Takashita, F. Kitame, and K. Nakamura Influenza C virus CM2 protein is produced from a 374-amino-acid protein (P42) by signal peptidase cleavage. J Virol 73: Hsu, M. T., J. D. Parvin, S. Gupta, M. Krystal, and P. Palese Genomic RNAs of influenza viruses are held in a circular conformation in virions and in infected cells by a terminal panhandle. Proc Natl Acad Sci U S A 84: Katagiri, S., A. Ohizumi, and M. Homma An outbreak of type C influenza in children's home. J Infect Dis. 148: Katagiri, S., A. Ohizumi, S. Ohyama, and M. Homma Follow-up study of type C influenza outbreak in children's home. Microbiol Immunol. 31: Li, X., and P. Palese Characterization of the polyadenylation signal of influenza virus RNA. J Virol 68:
14 Luo, G. X., W. Luytjes, M. Enami, and P. Palese The polyadenylation signal of influenza virus RNA involves a stretch of uridines followed by the RNA duplex of the panhandle structure. J Virol 65: Manuguerra, J. C., and C. Hannoun Natural infection of dogs by influenza C virus. Res Virol 143: Manuguerra, J. C., C. Hannoun, F. Simon, E. Villar, and J. A. Cabezas Natural infection of dogs by influenza C virus: a serological survey in Spain. New Microbiol 16: Matsuzaki, Y., N. Katsushima, Y. Nagai, M. Shoji, T. Itagaki, M. Sakamoto, S. Kitaoka, K. Mizuta, and H. Nishimura Clinical Features of Influenza C Virus Infection in Children. J Infect Dis. 193: Matsuzaki, Y., K. Mizuta, K. Sugawara, E. Tsuchiya, Y. Muraki, S. Hongo, H. Suzuki, and H. Nishimura Frequent reassortment among influenza C viruses. J Virol 77: Matsuzaki, Y., Y. Muraki, K. Sugawara, S. Hongo, H. Nishimura, F. Kitame, N. Katsushima, Y. Numazaki, and K. Nakamura Cocirculation of two distinct groups of influenza C virus in Yamagata City, Japan. Virology 202: Matsuzaki, Y., S. Takao, S. Shimada, K. Mizuta, K. Sugawara, E. Takashita, Y. Muraki, S. Hongo, and H. Nishimura Characterization of antigenically and genetically similar influenza C viruses isolated in Japan during the season. Epidemiol Infect 132: Matz, M. V Amplification of representative cdna pools from microscopic amounts of animal tissue. Methods Mol Biol 221: Muraki, Y., S. Hongo, K. Sugawara, F. Kitame, and K. Nakamura Evolution of the haemagglutinin-esterase gene of influenza C virus. J Gen Virol 77 ( Pt 4): Muraki, Y., T. Murata, E. Takashita, Y. Matsuzaki, K. Sugawara, and S. Hongo A mutation on influenza C virus M1 protein affects virion morphology by altering the membrane affinity of the protein. J Virol 81: Muraki, Y., H. Washioka, K. Sugawara, Y. Matsuzaki, E. Takashita, and S. Hongo Identification of an amino acid residue on influenza C virus M1 protein responsible for formation of the cord-like structures of the virus. J Gen Virol 85: Neumann, G., G. G. Brownlee, E. Fodor, and Y. Kawaoka Orthomyxovirus replication, transcription, and polyadenylation. Curr Top Microbiol Immunol 283: Nishimura, H., M. Hara, K. Sugawara, F. Kitame, K. Takiguchi, Y. Umetsu, A. Tonosaki, and K. Nakamura Characterization of the cord-like structures emerging from the surface of influenza C virus-infected cells. Virology 179: Nishimura, H., S. Hongo, K. Sugawara, Y. Muraki, F. Kitame, H. Washioka, A. Tonosaki, and K. Nakamura The ability of influenza C virus to generate cord-like structures is influenced by the gene coding for M protein. Virology 200: Noda, T., H. Sagara, A. Yen, A. Takada, H. Kida, R. H. Cheng, and Y. Kawaoka Architecture of ribonucleoprotein complexes in influenza A virus particles. Nature 439: Ohwada, K., F. Kitame, K. Sugawara, H. Nishimura, M. Homma, and K. Nakamura Distribution of the antibody to influenza C virus in dogs and pigs in Yamagata Prefecture, Japan. Microbiol Immunol 31: Pachler, K., J. Mayr, and R. Vlasak A seven plasmid-based system for the rescue of influenza C virus. J Mol Genet Med 4: Pachler, K., and R. Vlasak Influenza C virus NS1 protein counteracts RIG-I-mediated IFN signalling. Virology Journal 8:48. Paragas, J., J. Talon, R. E. O'Neill, D. K. Anderson, A. Garcia-Sastre, and P. Palese Influenza B and C Virus NEP (NS2) Proteins Possess Nuclear Export Activities. J. Virol. 75:
15 Pekosz, A., and R. A. Lamb Influenza C virus CM2 integral membrane glycoprotein is produced from a polypeptide precursor by cleavage of an internal signal sequence. Proc Natl Acad Sci U S A 95: Peng, G., S. Hongo, Y. Muraki, K. Sugawara, H. Nishimura, F. Kitame, and K. Nakamura Genetic reassortment of influenza C viruses in man. J Gen Virol 75 ( Pt 12): Pritlove, D. C., E. Fodor, B. L. Seong, and G. G. Brownlee In vitro transcription and polymerase binding studies of the termini of influenza A virus crna: evidence for a crna panhandle. J Gen Virol 76 ( Pt 9): Pritlove, D. C., L. L. Poon, E. Fodor, J. Sharps, and G. G. Brownlee Polyadenylation of influenza virus mrna transcribed in vitro from model virion RNA templates: requirement for 5' conserved sequences. J Virol 72: Taylor, R. M A further note on 1233 influenza C virus. Arch Gesamte Virusforsch 4: Tiley, L. S., M. Hagen, J. T. Matthews, and M. Krystal Sequence-specific binding of the influenza virus RNA polymerase to sequences located at the 5' ends of the viral RNAs. J Virol 68: Yamaoka, M., H. Hotta, M. Itoh, and M. Homma Prevalence of antibody to influenza C virus among pigs in Hyogo Prefecture, Japan. J Gen Virol 72 ( Pt 3): Yamashita, M., M. Krystal, and P. Palese Comparison of the three large polymerase proteins of influenza A, B, and C viruses. Virology 171: Yamashita, M., M. Krystal, and P. Palese Evidence that the matrix protein of influenza C virus is coded for by a spliced mrna. J Virol 62: Zhirnov, O. P., and V. B. Grigoriev Disassembly of influenza C viruses, distinct from that of influenza A and B viruses requires neutral-alkaline ph. Virology 200:
A seven plasmid-based system for the rescue of influenza C virus
 239 RESEARCH ARTICLE A seven plasmid-based system for the rescue of influenza C virus Karin Pachler, Juliane Mayr and Reinhard Vlasak* Department of Molecular Biology, University of Salzburg, Billrothstrasse
239 RESEARCH ARTICLE A seven plasmid-based system for the rescue of influenza C virus Karin Pachler, Juliane Mayr and Reinhard Vlasak* Department of Molecular Biology, University of Salzburg, Billrothstrasse
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
into 293T cells together with four (PB2, PB1, PA, and NP) or nine (PB2, PB1, PA, HA, NP, NA, M1, M2, and NS2) viral protein expressing plasmid DNAs. D
 Jpn. J. Infect. Dis., 63, 157 165, 2010 Review The Molecular Virology and Reverse Genetics of Influenza C Virus Yasushi Muraki 1,2 * and Seiji Hongo 2 1 Department of Microbiology, Kanazawa Medical University
Jpn. J. Infect. Dis., 63, 157 165, 2010 Review The Molecular Virology and Reverse Genetics of Influenza C Virus Yasushi Muraki 1,2 * and Seiji Hongo 2 1 Department of Microbiology, Kanazawa Medical University
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
The RNA Polymerase of Influenza Virus, Bound to the 5 End of Virion RNA, Acts in cis To Polyadenylate mrna
 JOURNAL OF VIROLOGY, Oct. 1998, p. 8214 8219 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. The RNA Polymerase of Influenza Virus, Bound to
JOURNAL OF VIROLOGY, Oct. 1998, p. 8214 8219 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. The RNA Polymerase of Influenza Virus, Bound to
Reverse Genetics of RNA Viruses
 Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Patricia Fitzgerald-Bocarsly
 FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus from eight plasmids
 Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Mutational Analysis of the Influenza Virus crna Promoter and Identification of Nucleotides Critical for Replication
 JOURNAL OF VIROLOGY, June 2004, p. 6263 6270 Vol. 78, No. 12 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.12.6263 6270.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Mutational
JOURNAL OF VIROLOGY, June 2004, p. 6263 6270 Vol. 78, No. 12 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.12.6263 6270.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Mutational
 http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Plasmid-Driven Formation of Influenza Virus-Like Particles
 JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
Rama Nada. - Malik
 - 2 - Rama Nada - - Malik 1 P a g e We talked about HAV in the previous lecture, now we ll continue the remaining types.. Hepatitis E It s similar to virus that infect swine, so its most likely infect
- 2 - Rama Nada - - Malik 1 P a g e We talked about HAV in the previous lecture, now we ll continue the remaining types.. Hepatitis E It s similar to virus that infect swine, so its most likely infect
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
The accumulation of influenza A virus segment 7 spliced mrnas is regulated by the NS1 protein
 Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
Segment-specific and common nucleotide sequences in the
 Proc. Nati. Acad. Sci. USA Vol. 84, pp. 2703-2707, May 1987 Biochemistry Segment-specific and common nucleotide sequences in the noncoding regions of influenza B virus genome RNAs (viral transcription/viral
Proc. Nati. Acad. Sci. USA Vol. 84, pp. 2703-2707, May 1987 Biochemistry Segment-specific and common nucleotide sequences in the noncoding regions of influenza B virus genome RNAs (viral transcription/viral
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
CDC website:
 Hepatitis B virus CDC website: http://www.cdc.gov/ncidod/diseases/hepatitis/slideset/hep_b/slide_1.htm Relevance Key Features es of Hepatitis t B Virus 250 million people infected worldwide. In areas of
Hepatitis B virus CDC website: http://www.cdc.gov/ncidod/diseases/hepatitis/slideset/hep_b/slide_1.htm Relevance Key Features es of Hepatitis t B Virus 250 million people infected worldwide. In areas of
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Bernadette Crescenzo-Chaigne, Nadia Naffakh, and Sylvie van der Werf 1
 Virology 265, 342 353 (1999) Article ID viro.1999.0059, available online at http://www.idealibrary.com on Comparative Analysis of the Ability of the Polymerase Complexes of Influenza Viruses Type A, B
Virology 265, 342 353 (1999) Article ID viro.1999.0059, available online at http://www.idealibrary.com on Comparative Analysis of the Ability of the Polymerase Complexes of Influenza Viruses Type A, B
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP
 Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Genetics. Instructor: Dr. Jihad Abdallah Transcription of DNA
 Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Overview: Chapter 19 Viruses: A Borrowed Life
 Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
Interaction of influenza virus polymerase with viral RNA in the corkscrew conformation
 Journal of General Virology (1999), 80, 2565 2572. Printed in Great Britain... Interaction of influenza virus polymerase with viral RNA in the corkscrew conformation Ramon Flick and Gerd Hobom Institut
Journal of General Virology (1999), 80, 2565 2572. Printed in Great Britain... Interaction of influenza virus polymerase with viral RNA in the corkscrew conformation Ramon Flick and Gerd Hobom Institut
Mutagenic Analysis of the 5 Arm of the Influenza A Virus Virion RNA Promoter Defines the Sequence Requirements for Endonuclease Activity
 JOURNAL OF VIROLOGY, Jan. 2001, p. 134 142 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.134 142.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Mutagenic Analysis
JOURNAL OF VIROLOGY, Jan. 2001, p. 134 142 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.134 142.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Mutagenic Analysis
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
Medical Virology. Herpesviruses, Orthomyxoviruses, and Retro virus. - Herpesviruses Structure & Composition: Herpesviruses
 Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Role of the CM2 Protein in the Influenza C Virus Replication Cycle
 JOURNAL OF VIROLOGY, Feb. 2011, p. 1322 1329 Vol. 85, No. 3 0022-538X/11/$12.00 doi:10.1128/jvi.01367-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Role of the CM2 Protein
JOURNAL OF VIROLOGY, Feb. 2011, p. 1322 1329 Vol. 85, No. 3 0022-538X/11/$12.00 doi:10.1128/jvi.01367-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Role of the CM2 Protein
Coronaviruses cause acute, mild upper respiratory infection (common cold).
 Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Influenza Genome Sequencing Project Proposal
 Date of Proposal (MM/DD/YY): 06/13/12 TITLE: Role of the Untranslated Regions of the Influenza A Virus Replication and Vaccines Purpose/Objective-please provide a brief one to two paragraph description
Date of Proposal (MM/DD/YY): 06/13/12 TITLE: Role of the Untranslated Regions of the Influenza A Virus Replication and Vaccines Purpose/Objective-please provide a brief one to two paragraph description
Phylogenetic analysis of influenza C virus nonstructural (NS) protein genes and identification of the NS2 protein
 Journal of General Virology (2000), 81, 1933 1940. Printed in Great Britain... Phylogenetic analysis of influenza C virus nonstructural (NS) protein genes and identification of the NS2 protein A. S. M.
Journal of General Virology (2000), 81, 1933 1940. Printed in Great Britain... Phylogenetic analysis of influenza C virus nonstructural (NS) protein genes and identification of the NS2 protein A. S. M.
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Reoviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Hepadnaviruses: Variations on the Retrovirus Theme
 WBV21 6/27/03 11:34 PM Page 377 Hepadnaviruses: Variations on the Retrovirus Theme 21 CHAPTER The virion and the viral genome The viral replication cycle The pathogenesis of hepatitis B virus A plant hepadnavirus
WBV21 6/27/03 11:34 PM Page 377 Hepadnaviruses: Variations on the Retrovirus Theme 21 CHAPTER The virion and the viral genome The viral replication cycle The pathogenesis of hepatitis B virus A plant hepadnavirus
Viral reproductive cycle
 Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Hierarchy among Viral RNA (vrna) Segments in Their Role in vrna Incorporation into Influenza A Virions
 JOURNAL OF VIROLOGY, Mar. 2006, p. 2318 2325 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2318 2325.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Hierarchy among
JOURNAL OF VIROLOGY, Mar. 2006, p. 2318 2325 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2318 2325.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Hierarchy among
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
11/15/2011. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Viral Genomes. Structural Features and Characteristics
 Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics
 Hepadnaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Hepatitis viruses A group of unrelated pathogens termed hepatitis viruses cause the vast majority
Hepadnaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Hepatitis viruses A group of unrelated pathogens termed hepatitis viruses cause the vast majority
Tempo and Mode in the Molecular Evolution of Influenza C
 Tempo and Mode in the Molecular Evolution of Influenza C December 7, 2010 1 Derek Gatherer 1 MRC-University of Glasgow Centre for Virus Research, 8 Church Street, Glasgow G11 5JR, UK. Gatherer D. Tempo
Tempo and Mode in the Molecular Evolution of Influenza C December 7, 2010 1 Derek Gatherer 1 MRC-University of Glasgow Centre for Virus Research, 8 Church Street, Glasgow G11 5JR, UK. Gatherer D. Tempo
Unité de Génétique Moléculaire des Virus Respiratoires, URA 1966 CNRS, Institut Pasteur, 25 rue du Dr. Roux, PARIS Cedex 15, France
 Virology 345 (2006) 73 87 www.elsevier.com/locate/yviro Recombinant influenza A viruses harboring optimized dicistronic NA segment with an extended native 5V terminal sequence: Induction of heterospecific
Virology 345 (2006) 73 87 www.elsevier.com/locate/yviro Recombinant influenza A viruses harboring optimized dicistronic NA segment with an extended native 5V terminal sequence: Induction of heterospecific
Reassortment of influenza A virus genes linked to PB1 polymerase gene
 International Congress Series 1263 (2004) 714 718 Reassortment of influenza A virus genes linked to PB1 polymerase gene Jean C. Downie* www.ics-elsevier.com Centre for Infectious Diseases and Microbiology,
International Congress Series 1263 (2004) 714 718 Reassortment of influenza A virus genes linked to PB1 polymerase gene Jean C. Downie* www.ics-elsevier.com Centre for Infectious Diseases and Microbiology,
Transcription of the German Cockroach Densovirus BgDNV Genome: Alternative Processing of Viral RNAs
 ISSN 1607-6729, Doklady Biochemistry and Biophysics, 2008, Vol. 421, pp. 176 180. Pleiades Publishing, Ltd., 2008. Original Russian Text T.V. Kapelinskaya, E.U. Martynova, A.L. Korolev, C. Schal, D.V.
ISSN 1607-6729, Doklady Biochemistry and Biophysics, 2008, Vol. 421, pp. 176 180. Pleiades Publishing, Ltd., 2008. Original Russian Text T.V. Kapelinskaya, E.U. Martynova, A.L. Korolev, C. Schal, D.V.
Influenza Viruses A Review
 Influenza Viruses A Review AVIAN INFLUENZA: INTERSECTORAL COLLABORATION Larnaca, Cyprus 20 22 July 2009 Kate Glynn Scientific and Technical Department, OIE Influenza Viruses C. Goldsmith,1981 Influenza
Influenza Viruses A Review AVIAN INFLUENZA: INTERSECTORAL COLLABORATION Larnaca, Cyprus 20 22 July 2009 Kate Glynn Scientific and Technical Department, OIE Influenza Viruses C. Goldsmith,1981 Influenza
NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome
 Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Fine Mapping of a cis-acting Sequence Element in Yellow Fever Virus RNA That Is Required for RNA Replication and Cyclization
 JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
Human Genome Complexity, Viruses & Genetic Variability
 Human Genome Complexity, Viruses & Genetic Variability (Learning Objectives) Learn the types of DNA sequences present in the Human Genome other than genes coding for functional proteins. Review what you
Human Genome Complexity, Viruses & Genetic Variability (Learning Objectives) Learn the types of DNA sequences present in the Human Genome other than genes coding for functional proteins. Review what you
New Aspects of Influenza Viruses
 CLINICAL MICROBIOLOGY REVIEWS, Jan. 1992, p. 74-92 Vol. 5, No. 1 0893-8512192/010074-19$02.00/0 Copyright 1992, American Society for Microbiology New Aspects of Influenza Viruses MICHAEL W. SHAW,* NANCY
CLINICAL MICROBIOLOGY REVIEWS, Jan. 1992, p. 74-92 Vol. 5, No. 1 0893-8512192/010074-19$02.00/0 Copyright 1992, American Society for Microbiology New Aspects of Influenza Viruses MICHAEL W. SHAW,* NANCY
Existence of reassortant A (H1N2) swine influenza viruses in Saitama Prefecture, Japan
 International Congress Series 1263 (2004) 749 753 Existence of reassortant A (H1N2) swine influenza viruses in Saitama Prefecture, Japan Shin ichi Shimada a, *, Takayasu Ohtsuka b, Masayuki Tanaka b, Munehito
International Congress Series 1263 (2004) 749 753 Existence of reassortant A (H1N2) swine influenza viruses in Saitama Prefecture, Japan Shin ichi Shimada a, *, Takayasu Ohtsuka b, Masayuki Tanaka b, Munehito
AP Biology. Viral diseases Polio. Chapter 18. Smallpox. Influenza: 1918 epidemic. Emerging viruses. A sense of size
 Hepatitis Viral diseases Polio Chapter 18. Measles Viral Genetics Influenza: 1918 epidemic 30-40 million deaths world-wide Chicken pox Smallpox Eradicated in 1976 vaccinations ceased in 1980 at risk population?
Hepatitis Viral diseases Polio Chapter 18. Measles Viral Genetics Influenza: 1918 epidemic 30-40 million deaths world-wide Chicken pox Smallpox Eradicated in 1976 vaccinations ceased in 1980 at risk population?
HS-LS4-4 Construct an explanation based on evidence for how natural selection leads to adaptation of populations.
 Unit 2, Lesson 2: Teacher s Edition 1 Unit 2: Lesson 2 Influenza and HIV Lesson Questions: o What steps are involved in viral infection and replication? o Why are some kinds of influenza virus more deadly
Unit 2, Lesson 2: Teacher s Edition 1 Unit 2: Lesson 2 Influenza and HIV Lesson Questions: o What steps are involved in viral infection and replication? o Why are some kinds of influenza virus more deadly
Picornaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
LESSON 4.6 WORKBOOK. Designing an antiviral drug The challenge of HIV
 LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
Technology Overview. Summary
 Live Attenuated Influenza Vaccines with Altered NS1 Technology Overview Summary Transformative Technology: Live attenuated influenza vaccines (LAIVs) with precise, genetically stable truncations of the
Live Attenuated Influenza Vaccines with Altered NS1 Technology Overview Summary Transformative Technology: Live attenuated influenza vaccines (LAIVs) with precise, genetically stable truncations of the
Influenzavirus. Influenzavirus 10/27/15. Lord Randolphs report from Queen Marys trip to Scotland in 1562
 A disease under the influence of the stars or the moon. Lord Randolphs report from Queen Marys trip to Scotland in 1562 She fell acquaintance with a new disease which passed through her whole court neither
A disease under the influence of the stars or the moon. Lord Randolphs report from Queen Marys trip to Scotland in 1562 She fell acquaintance with a new disease which passed through her whole court neither
Original Article Development and Sequence Analysis of a Cold-Adapted Strain of Influenza A/New Caledonia/20/1999(H1N1) Virus
 Iranian Journal of Virology 2011;5(4): 6-10 2011, Iranian Society for Virology Original Article Development and Sequence Analysis of a Cold-Adapted Strain of Influenza A/New Caledonia/20/1999(H1N1) Virus
Iranian Journal of Virology 2011;5(4): 6-10 2011, Iranian Society for Virology Original Article Development and Sequence Analysis of a Cold-Adapted Strain of Influenza A/New Caledonia/20/1999(H1N1) Virus
Cover Page. The handle holds various files of this Leiden University dissertation
 Cover Page The handle http://hdl.handle.net/1887/35908 holds various files of this Leiden University dissertation Author: Soema, Peter Title: Formulation of influenza T cell peptides : in search of a universal
Cover Page The handle http://hdl.handle.net/1887/35908 holds various files of this Leiden University dissertation Author: Soema, Peter Title: Formulation of influenza T cell peptides : in search of a universal
Viral Genetics. BIT 220 Chapter 16
 Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
VIROLOGY. Engineering Viral Genomes: Retrovirus Vectors
 VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
Reverse genetic platform for inactivated and live-attenuated influenza vaccine
 EXPERIMENTAL and MOLECULAR MEDICINE, Vol. 42, No. 2, 116-121, February 2010 Reverse genetic platform for inactivated and live-attenuated influenza vaccine Eun-Ju Jung, Kwang-Hee Lee and Baik Lin Seong
EXPERIMENTAL and MOLECULAR MEDICINE, Vol. 42, No. 2, 116-121, February 2010 Reverse genetic platform for inactivated and live-attenuated influenza vaccine Eun-Ju Jung, Kwang-Hee Lee and Baik Lin Seong
Protein Synthesis
 Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
ORTHOMYXOVIRUSES INFLUENZA VIRUSES. (A,B and C)
 ORTHOMYXOVIRUSES INFLUENZA VIRUSES (A,B and C) Orthomyxoviridae Influenza Viruses Epidemiology: Influenza A virus is so subjected to major antigenic changes that cause occasional world wide pandemics when
ORTHOMYXOVIRUSES INFLUENZA VIRUSES (A,B and C) Orthomyxoviridae Influenza Viruses Epidemiology: Influenza A virus is so subjected to major antigenic changes that cause occasional world wide pandemics when
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Severe Acute Respiratory Syndrome (SARS) Coronavirus
 Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Influenza viruses are classified as members of the family
 Rewiring the RNAs of influenza virus to prevent reassortment Qinshan Gao a and Peter Palese a,b,1 Departments of a Microbiology and b Medicine, Mount Sinai School of Medicine, New York, NY 10029 Contributed
Rewiring the RNAs of influenza virus to prevent reassortment Qinshan Gao a and Peter Palese a,b,1 Departments of a Microbiology and b Medicine, Mount Sinai School of Medicine, New York, NY 10029 Contributed
Characterization of the 1918 influenza virus polymerase genes
 Vol 437 6 October 2005 doi:10.1038/nature04230 Characterization of the 1918 influenza virus polymerase genes Jeffery K. Taubenberger 1, Ann H. Reid 1, Raina M. Lourens 1, Ruixue Wang 1, Guozhong Jin 1
Vol 437 6 October 2005 doi:10.1038/nature04230 Characterization of the 1918 influenza virus polymerase genes Jeffery K. Taubenberger 1, Ann H. Reid 1, Raina M. Lourens 1, Ruixue Wang 1, Guozhong Jin 1
Cristina Cassetti, Ph.D.
 NIAID Extramural Research Update: Recombinant Influenza Viruses and Biosafety Cristina Cassetti, Ph.D. Influenza Program Officer Division of Microbiology and Infectious Diseases NIAID Influenza virus DMID
NIAID Extramural Research Update: Recombinant Influenza Viruses and Biosafety Cristina Cassetti, Ph.D. Influenza Program Officer Division of Microbiology and Infectious Diseases NIAID Influenza virus DMID
VIRUSES. 1. Describe the structure of a virus by completing the following chart.
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
19 Viruses BIOLOGY. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Structural Features and Characteristics
 9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Unit 2: Lesson 2 Case Studies: Influenza and HIV LESSON QUESTIONS
 1 Unit 2: Lesson 2 Case Studies: Influenza and HIV LESSON QUESTIONS What steps are involved in viral infection and replication? Why are some kinds of influenza virus more deadly than others? How do flu
1 Unit 2: Lesson 2 Case Studies: Influenza and HIV LESSON QUESTIONS What steps are involved in viral infection and replication? Why are some kinds of influenza virus more deadly than others? How do flu
Mutations at Alternative 5 Splice Sites of M1 mrna Negatively Affect Influenza A Virus Viability and Growth Rate
 JOURNAL OF VIROLOGY, Nov. 2008, p. 10873 10886 Vol. 82, No. 21 0022-538X/08/$08.00 0 doi:10.1128/jvi.00506-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Mutations at Alternative
JOURNAL OF VIROLOGY, Nov. 2008, p. 10873 10886 Vol. 82, No. 21 0022-538X/08/$08.00 0 doi:10.1128/jvi.00506-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Mutations at Alternative
Biol115 The Thread of Life"
 Biol115 The Thread of Life" Lecture 9" Gene expression and the Central Dogma"... once (sequential) information has passed into protein it cannot get out again. " ~Francis Crick, 1958! Principles of Biology
Biol115 The Thread of Life" Lecture 9" Gene expression and the Central Dogma"... once (sequential) information has passed into protein it cannot get out again. " ~Francis Crick, 1958! Principles of Biology
Unexpected complexity in the interference activity of a cloned influenza defective interfering RNA
 Meng et al. Virology Journal (2017) 14:138 DOI 10.1186/s12985-017-0805-6 RESEARCH Unexpected complexity in the interference activity of a cloned influenza defective interfering RNA Open Access Bo Meng
Meng et al. Virology Journal (2017) 14:138 DOI 10.1186/s12985-017-0805-6 RESEARCH Unexpected complexity in the interference activity of a cloned influenza defective interfering RNA Open Access Bo Meng
Influenza vaccines: present and future
 PERSPECTIVE SERIES The future of vaccine design Peter Palese and Adolfo García-Sastre, Series Editors Influenza vaccines: present and future Peter Palese and Adolfo García-Sastre Department of Microbiology,
PERSPECTIVE SERIES The future of vaccine design Peter Palese and Adolfo García-Sastre, Series Editors Influenza vaccines: present and future Peter Palese and Adolfo García-Sastre Department of Microbiology,
Overview of virus life cycle
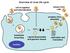 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Virus Basics. General Characteristics of Viruses. Chapter 13 & 14. Non-living entities. Can infect organisms of every domain
 Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-forms Commonly referred to by organism they infect
Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-forms Commonly referred to by organism they infect
Virus and Prokaryotic Gene Regulation - 1
 Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Islamic University Faculty of Medicine
 Islamic University Faculty of Medicine 2012 2013 2 RNA is a modular structure built from a combination of secondary and tertiary structural motifs. RNA chains fold into unique 3 D structures, which act
Islamic University Faculty of Medicine 2012 2013 2 RNA is a modular structure built from a combination of secondary and tertiary structural motifs. RNA chains fold into unique 3 D structures, which act
7.014 Problem Set 7 Solutions
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
MedChem 401~ Retroviridae. Retroviridae
 MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
INFLUENZA VIRUS. INFLUENZA VIRUS CDC WEBSITE
 INFLUENZA VIRUS INFLUENZA VIRUS CDC WEBSITE http://www.cdc.gov/ncidod/diseases/flu/fluinfo.htm 1 THE IMPACT OF INFLUENZA Deaths: PANDEMICS 1918-19 S p a n is h flu 5 0 0,0 0 0 U S 2 0,0 0 0,0 0 0 w o rld
INFLUENZA VIRUS INFLUENZA VIRUS CDC WEBSITE http://www.cdc.gov/ncidod/diseases/flu/fluinfo.htm 1 THE IMPACT OF INFLUENZA Deaths: PANDEMICS 1918-19 S p a n is h flu 5 0 0,0 0 0 U S 2 0,0 0 0,0 0 0 w o rld
Virus Basics. General Characteristics of Viruses 5/9/2011. General Characteristics of Viruses. Chapter 13 & 14. Non-living entities
 Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-formsf Commonly referred to by organism they infect
Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-formsf Commonly referred to by organism they infect
MECHANISMS FOR THE ENHANCEMENT OF INFLUENZA B VIRUS VACCINE ANTIGEN YIELD
 MECHANISMS FOR THE ENHANCEMENT OF INFLUENZA B VIRUS VACCINE ANTIGEN YIELD HYUNSUH KIM DOCTOR OF PHILOSOPHY 2012 RMIT MECHANISMS FOR THE ENHANCEMENT OF INFLUENZA B VIRUS VACCINE ANTIGEN YIELD Hyunsuh Kim
MECHANISMS FOR THE ENHANCEMENT OF INFLUENZA B VIRUS VACCINE ANTIGEN YIELD HYUNSUH KIM DOCTOR OF PHILOSOPHY 2012 RMIT MECHANISMS FOR THE ENHANCEMENT OF INFLUENZA B VIRUS VACCINE ANTIGEN YIELD Hyunsuh Kim
Rescue of Recombinant Thogoto Virus from Cloned cdna
 JOURNAL OF VIROLOGY, Oct. 2001, p. 9282 9286 Vol. 75, No. 19 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.19.9282 9286.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Rescue of
JOURNAL OF VIROLOGY, Oct. 2001, p. 9282 9286 Vol. 75, No. 19 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.19.9282 9286.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Rescue of
Lecture 19 Evolution and human health
 Lecture 19 Evolution and human health The evolution of flu viruses The evolution of flu viruses Google Flu Trends data US data Check out: http://www.google.org/flutrends/ The evolution of flu viruses the
Lecture 19 Evolution and human health The evolution of flu viruses The evolution of flu viruses Google Flu Trends data US data Check out: http://www.google.org/flutrends/ The evolution of flu viruses the
Evolutionary distances between proteins of the Influenza A Virus
 Evolutionary distances between proteins of the Influenza A Virus Hatem Nassrat B00393388 Bioinformatics: Winter 06/07 Dr. Christian Blouin Table of Contents Evolutionary distances between proteins of the
Evolutionary distances between proteins of the Influenza A Virus Hatem Nassrat B00393388 Bioinformatics: Winter 06/07 Dr. Christian Blouin Table of Contents Evolutionary distances between proteins of the
