* * p=0.04 p=0.007 p=0.013 p=0.04 p= p= U/L. Myc
|
|
|
- Laura Mathews
- 6 years ago
- Views:
Transcription
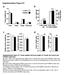
1 Supplemental Figure 1 Supplemental Figure 1 A H&E CV WT Alb-Myc CV PV PV * * Total bilirubin AST Alk phos Triglycerides p=.3 p=.4 p=.7 p=.13 p=.4 p=.14 U/L mg/dl U/L 2 * * WT Myc βδex3 βδex3: Myc 25 2 * 5 5 WT Myc βδex3 βδex3: Myc p=.1 mg/dl B β catδex3 β catδex3 CV WT Myc βδex3 βδex3: Myc WT Myc βδex3 βδex3: Myc C IHC: SOX9 IHC:β-catenin WT β-catδex3 PV PV PV CV Supplemental Figure 1. Pan-hepatic expression of mutant β-catenin induces severe liver damage and dysfunction. (A) Photomicrographs of H&E stained liver sections from 28 day old WT (top left), Alb-c-Myc (top right) and β-catδex3 (bottom left and right) mice. Note that while expression of the Alb-Myc transgene induces mild hepatocyte cell and nuclear enlargement and hyperchromatic nuclei (inset, top right), liver architecture is otherwise normal. Note atrophic hepatocytes indicative of possible impending necrosis (inset, bottom left), areas of overt focal necrosis (!) and inflammatory infiltrates (arrowheads) in livers expressing mutant β-catenin. PV, portal vein; CV, central vein. (B) Graphs depicting biochemical analysis of liver function in WT, Alb-c-Myc, βcatδex3 and β-catδex3:myc mice showing that mutant β-catenin results in significant liver dysfunction characterized by elevations in aspartate aminotransferase (AST), alkaline phosphatase (Alk phos) and triglycerides. Note that while liver function is unaffected by expression of the Alb-c-Myc transgene at 28 days, co-
2 expression of the Alb-c-Myc transgene with mutant β-catenin partially ameliorates β-catenin-induced liver injury in β-catδex3:myc mice as evidenced by lower alkaline phosphatase and triglyceride levels relative to β-catδex3 mice. n=3 mice/group except for AST and bilirubin where n=4 for the WT group. P values generated by a twotailed unpaired t-test. Bars represent mean +/- SEM. (C) Photomicrographs of IHC for CTNNB1 (β-catenin) (top panels) and SOX9 (bottom panels) in WT (left top and bottom panels) and β-catδex3 (right top and bottom panels) livers. β-catenin is localized to the outer membranes of hepatocytes and biliary cells (black arrowhead) in WT liver (top left panel), but is overexpressed and mislocalized to the cytoplasm of hepatocytes in β-catδex3 mice (upper right panel) (β-catenin, red (AEC chromagen); hematoxylin counterstain, blue). In WT livers, SOX9, is exclusively expressed in the nuclei of biliary cells immediately adjacent to the portal vein (PV), but not in hepatocytes (bottom left panel). In β-catδex3 livers, SOX9-expressing cells are no longer restricted to the periportal area, but are visible throughout the lobule and a subset of hepatocytes now express SOX9 (insets) (bottom right panel), indicating possible induction of a regenerative response in an attempt to counteract hepatocyte loss and improve function. A higher magnification of the SOX9-stained bile ducts around the portal vein (PV) in the same WT liver (bottom left panel in C) is also shown in Figure 4B. (AEC chromagen, red/brown; hematoxylin counterstain, blue). Scale bars represent 5µm.
3 Supplemental Figure 2 Age Alb-c-Myc Northern Blot Total Liver RNA P28-42 Endogenous c-myc (E) c-myc Transgene (T) Albumin Supplemental Figure 2. Hepatic c-myc mrna expression driven by the Albumin-c-Myc transgene in adult liver. Northern blot showing levels of endogenous c-myc (E) expressed in the livers of WT and Albumin-c-Myc transgenic mice (lanes 1, 2, 3 and 4) and expression of the Albumin-c-Myc transgene (T) in the livers of adult transgenic mice (lanes 3 and 4). Although transgene-derived c-myc mrna first becomes detectable by Northern blot during the first week after birth (Figure 2), maximal expression is not achieved until the liver has fully matured at ~ 3-4 weeks of age. Despite this high level of c-myc, Alb-c-Myc mice do not develop liver tumors (hepatocellular carcinomas) until they reach ~1-12 months of age. Lanes 1-4 represent RNA samples that were run on the same gel, but were non-contiguous between lanes 2 and 3 (see separating black line).
4 Supplemental Figure 3 Liver: H & E A B C β-catδex3:myc mice Tet-O-Myc mice D E F G H I J K L M N O HBs Other tumors HBs P Q R Supplemental Figure 3. Histopathology of HBs and other liver tumors that develop in β-catδex3 and Tet-O-Myc mice. Photomicrographs of H&E stained sections showing spectrum of HBs (A-I) and other liver tumors (J-L) in 3-6 week old β-catδex3 mice and week old Tet-O-Myc mice (M-R). (A) Low-power image of fetal HBs showing adjacent tumors with characteristic clear and dark cell morphologies. (B) Higher power image of a fetal HB with clear cell morphology (left) juxtaposed to one with dark cell morphology (right). (C) Leading edge of a fetal HB depicting border with normal liver. (D) High power image of fetal HB with clear cell morphology. Extramedullary hematopoiesis (EMH) is prominent in many HBs (arrowheads), particularly those in A-D. (E) Mixed fetal-embryonal HB. (F) Mixed fetal-embryonal HB with a microtrabecular growth pattern. (G) Low power image of solitary fetal-embryonal HB. Contrast the well-demarcated border of the tumor in (G) with the less defined border of the fetal HB in C. (H and I) Embyonal HBs with macrotabecular (H) or solid (I) growth patterns. Cells have a high nuclear/cytoplasmic ratio, angulated nuclei and demonstrate reduced cohesion between cells. EMH is also largely absent. (J) Hepatocellular malignant neoplasm; not otherwise specified (HMN NOS). (K) Anaplastic HCC. (L) Trabecular HCC. (M-R) Tet-O-Myc-mice develop poorly differentiated HBs with primitive features. HBs contain small angulated cells that display reduced cohesion with one another (M, N, O) with tumors developing within the context of non-injured, relatively normal liver (M, P). Tet-O-Mycderived HBs also contain tubulo-glandular structures (P, Q), and display trabecular (O) or solid (R) growth patterns, have clearly demarcated boundaries and generally lack EMH. Most β-catδex3:myc-derived HBs resemble cells present during the latter stages of embryonic liver development (E15 onwards), while Tet-O-Mycderived HBs are more similar to embryonic liver cells present at E Scale bars represent 5µm except for panels A and G where they represent 1µm.
5 Supplemental Figure 4 c-myc IHC β-catenin WT A B Mouse Liver Tet-O-Myc β-cat ΔEx3 :Myc C E D F G H Supplemental Figure 4. Activated wnt/β-catenin and c-myc cooperate to drive tumorigenesis in β- catδex3:myc and Tet-O-Myc mice. Photomicrographs if immunohistochemistry with c-myc- and β-cateninspecific antibodies showing expression and localization of c-myc (A, C, E and G) and β-catenin (B, D, F and H) in livers of WT (A, B) β-catδex3:myc (C-F) and Tet-O-Myc (G, H) mice. c-myc is undetectable in WT liver (A), but is highly expressed in the nuclei of tumor cells in β-catδex3:myc (C, E) and Tet-O-Myc (G) mice. β-catenin is localized to the outer membranes of hepatocytes and bile ducts (arrowhead) in WT livers (B), but is predominantly cytoplasmic in HBs from β-catδex3:myc (D, F) and Tet-O-Myc (H) mice, although some membranous staining is also evident in (H). In each case β-catenin and c-myc are more abundant in tumors relative to surrounding liver. Note the marked absence or reduction in membranous β-catenin in non-tumorous liver of β-catδex3:myc mice in (D). c-myc and β-catenin (AEC chromagen, red/brown), hematoxylin counterstain (blue). Scale bars represent 5µm.
6 Supplemental Figure 5 A Total Liver Protein ΔEx3 ΔEx3 WT Myc β β :Myc 9kD 68kD Wild Type β-cat Mutant β-cat 37kD GAPDH B 25 Wild type β-cat Relative Expression WT Myc β ΔEx3 β ΔEx3: Myc 25 Mutant β-cat Relative Expression WT Myc β ΔEx3 β ΔEx3: Myc Supplemental Figure 5. Hepatic expression of mutant β-catenin results in reduced expression of endogenous β-catenin. (A) Western blotting of liver extracts prepared from 2 individual WT, Alb-c-Myc (Myc), β- catδex3 (β ΔEx3 ) and β-catδex3:myc (β ΔEx3 :Myc) mice using β-catenin- and GAPDH-specific antibodies. Note that while 9kD wild-type (endogenous) β-catenin is abundantly expressed in WT and Alb-c-Myc livers, there is a 65-8% reduction in wild-type β-catenin in livers of β-catδex3 (lanes 5 & 6) and β-catδex3:myc (lanes 7 & 8) mice expressing mutant β-catenin. (B) Graphs showing relative hepatic expression of wild-type (upper graph) and mutant (lower graph) β-catenin. Graphs were generated by importing scanned images of each gel into Image J software and quantifying signal intensity. Bars represent the mean +/- SEM.
7 Supplemental Figure 6 Early hepatoblasts (E1-14) Late hepatoblasts (E15-E19) Normal adult hepatocytes Normal adult biliary cells Well-differentiated (fetal) high (hi) Mouse HBs Increasing differentiation Moderately-differentiated (embryonal) HNF4α SOX9 Well-differentiated HBs AFP DLK1 GLUL* CITED1 EpCAM LIN28A LIN28B MYC Magnitude of expression Moderately differentiated HBs intermediate (int) low negative/ (neg) negligible HNF4α (hi) AFP (hi) (low) DLK1 (hi) GLUL (hi) CITED1 EpCAM LIN28A (hi) LIN28B (int) MYC cyto-β-cat (neg) SOX9 (neg) (neg) (hi) HNF4α(hi) AFP (low-int) (hi) DLK1 GLUL(low) (hi) CITED1 EpCAM LIN28A (hi) LIN28B (hi) MYC cyto-β-cat SOX9(neg) (low-int) (low-int) (hi) * Perivenous hepatocytes only cyto-β-cat; cytoplasmic localized β-catenin Supplemental Figure 6. Scheme summarizing the expression of oncofetal genes and lineage markers as a function of tumor cell differentiation in HBs from β-catδex3:myc and Tet-O-Myc mice. The magnitude of expression of lineage-restricted and developmentally regulated genes is represented as a function of tumor cell differentiation in tumors (long vertical bars) from both strains of mice. Small squares depict the magnitude of expression of each gene in normal hepatoblasts present during the early and latter stages of embryonic liver development, adult hepatocytes and biliary cells. Both types of HB express abundant HNF4α, a marker of hepatocytic commitment, but little to no SOX9, a marker of biliary cell commitment. IHC shows that well-differentiated fetal HBs can be discriminated from embryonal HBs on the basis of high AFP and GLUL expression and low or absent expression of DLK1, EpCAM and LIN28A. Fetal HBs completely lack expresssion of LIN28A and EpCAM suggesting that these may be the most useful markers for discriminating between well and poorly-differentiated HBs. LIN28B was highly expressed in both HB subtypes and provided no discrimination. Embryonal HBs also generally expressed higher levels of c- Myc than fetal HBs. Black, gray and white color indicates high, intermediate and low/negligible expression, respectively.
8 Supplemental Figure 7 A mrna WT Myc βδex3 β ΔEx3: Myc Dlk1 Igf2 S1a9 Peg3 Phgdh Afp Nope Wnt4 Cited1 Ppia B Dlk1 Igf2 S1a9 Peg3 Phgdh Relative expression (mrna) W M β βm W M β βm W M β βm W M β βm W M β βm Relative expression (mrna) Afp Nope Wnt4 Cited1 Ppia W M β βm W M β βm W M β βm W M β βm W M β βm W:WT M:Myc β:β-catδex3 βm:β-catδex3:myc Supplemental Figure 7. Post-array verification of tumor-enriched and c-myc-associated DEGs. (A) Ethidium bromide stained gels of RT-PCR-amplified products for selected tumor-enriched DEGs in Table S1 as identified by array. Total RNA isolated from livers of 2 individual WT, Alb-c-Myc (Myc), β-catδex3 (βδex3) and β-catδex3:myc (βδex3:myc) mice was converted to cdna and amplified using gene-specific primers listed in Supplemental Table 11. (B) Graphs showing quantitation of gels in (A) depicting relative hepatic mrna expression of each gene. Graphs were generated by importing scanned images of each gel into Image J software and quantifying signal intensity relative to Ppia (cyclophilin A). W, WT; M, Myc; β, βδex3 and βm, βδex3:myc mice. Horizontal bars for each genotype represents the mean of two individual samples. Error bars represent +/- SEM.
9 Supplemental Figure 8 (A) Putative tumor-enriched DEGs; genes differentially expressed between β-cat:myc & β-cat livers. (corresponding to yellow box, Fig. 5) I. Enriched Oncogenic Signature Gene Sets Bcat vs Bcat+M gene list NOM FDR GS DETAILS SIZE ES NES p-val q-val KRAS.PROSTATE_UP.V1_DN P53_DN.V2_UP P53_DN.V2_DN CYCLIN_D1_P.V1_DN KRAS.AMP.LUNG_UP.V1_UP BMI1_DN_MEL18_DN.V1_UP CSR_LATE_UP.V1_DN MEL18_DN.V1_UP KRAS.DF.V1_DN BMI1_DN.V1_UP II. Enriched Gene Ontology Biologocal Processes Terms Bcat+Myc vs Bcat gene list NOM FDR GS DETAILS SIZE ES NES p-val q-val MITOTIC_CELL_CYCLE CELL_CYCLE_PHASE GLYCOPROTEIN_BIOSYNTHETIC_PROCESS CELL_CYCLE_PROCESS REGULATION_OF_CYCLIN_DEPENDENT_ PROTEIN_KINASE_ACTIVITY CELL_CELL_SIGNALING CELLULAR_MORPHOGENESIS_ DURING_DIFFERENTIATION AXONOGENESIS NEURITE_DEVELOPMENT NEURON_DEVELOPMENT Supplemental Figure 8A. GSEA (Enriched oncogenic signature gene sets and gene ontology biological processes terms) of the putative tumor-enriched DEGs in Supplemental Table 1 (β-cat:myc vs. β-cat comparison) (Fig. 5, yellow box).
10 Supplemental Figure 8 (B) DEGs co-regulated by β-cat & Myc; genes differentially expressed between β-cat:myc and WT livers. (corresponding to green box, Fig. 5) I. Enriched Oncogenic Signature Gene Sets Bcat+Myc vs Normal gene list NOM FDR GS DETAILS SIZE ES NES p-val q-val NFE2L2.V ESC_J1_UP_LATE.V1_UP LEF1_UP.V1_DN ATM_DN.V1_DN ESC_V6.5_UP_LATE.V1_UP KRAS.LUNG_UP.V1_UP KRAS.DF.V1_UP KRAS.BREAST_UP.V1_UP KRAS.6.LUNG.BREAST_UP.V1_UP RPS14.DN.V1_UP II. Enriched Gene Ontology Biological Processes Terms Bcat:Myc vs WT gene list NOM FDR GS DETAILS SIZE ES NES p-val q-val PROTEIN_OLIGOMERIZATION CHROMATIN_MODIFICATION MRNA_PROCESSING_GO_ M_PHASE_OF_MITOTIC_CELL_CYCLE MITOCHONDRION_ORGANIZATION _AND_BIOGENESIS G_PROTEIN_COUPLED_RECEPTOR_ PROTEIN_SIGNALING_PATHWAY LOCOMOTORY_BEHAVIOR CELL_SURFACE_RECEPTOR_LINKED_ SIGNAL_TRANSDUCTION_GO_7166 IMMUNE_RESPONSE IMMUNE_SYSTEM_PROCESS Supplemental Figure 8B. GSEA (Enriched oncogenic signature gene sets and gene ontology biological processes terms) of the DEGs co-regulated by β-catenin and Myc (β-cat:myc vs. WT comparison) (Fig. 5, green box). (Related to Online GEO submission GSE 7984).
11 Supplemental Figure 8. (C) β-catenin-regulated DEGs; genes differentially expressed between β-cat and WT livers. (corresponding to red box, Fig. 5) I. Enriched oncogenic Oncogenic signature Signature Gene gene Sets sets Bcat vs WT gene list NOM FDR GS DETAILS SIZE ES NES p-val q-val NFE2L2.V LEF1_UP.V1_DN MEK_UP.V1_UP ESC_J1_UP_LATE.V1_UP ESC_V6.5_UP_LATE.V1_DN P53_DN.V2_UP KRAS.6.LUNG.BREAST_UP.V1_UP KRAS.BREAST_UP.V1_UP JAK2_DN.V1_UP KRAS.PROSTATE_UP.V1_UP Bcat:Myc vs WT gene list II. Enriched Gene Ontology Biological Processes Terms Enriched Gene Ontology Biologial Processes Terms Bcat vs WT gene list NOM FDR GS DETAILS SIZE ES NES p-val q-val PPROTEIN_OLIGOMERIZATION REPRODUCTIVE_PROCESS AMINE_TRANSPORT NUCLEOTIDE_METABOLIC_PROCESS RESPONSE_TO_ABIOTIC_STIMULUS IMMUNE_RESPONSE IMMUNE_SYSTEM_PROCESS LOCOMOTORY_BEHAVIOR DEFENSE_RESPONSE CELLULAR_DEFENSE_RESPONSE INFLAMMATORY_RESPONSE G_PROTEIN_COUPLED_RECEPTOR_ PROTEIN_SIGNALING_PATHWAY CELL_SURFACE_RECEPTOR_LINKED_ SIGNAL_TRANSDUCTION_GO_7166 BEHAVIOR RESPONSE_TO_EXTERNAL_STIMULUS Supplemental Figure 8C. GSEA (Enriched oncogenic signature gene sets and enriched gene ontology biological processes terms) of the DEGs regulated by β-catenin alone (β-cat vs. WT) (Fig. 5, red box). (Related to Figure 6 and Online GEO submission GSE 7984).
12 Supplemental Figure 9 A IHC: c-myc B IHC: NQO1 Ton HB1 HB1 Ton HB1 HB1 HB2 HB2 HB3 HB3 HB4 HB4 HB2 HB2 HB3 HB3 HB4 NL1 HB6 HB9 HB9 HB12 HB12 HB14 HB14 NL4 NL4 HB5 HB5 NL2 NL2 HB8 HB8 HB11 HB11 HB13 HB13 HB16 HB16 No Hematoxylin HB18 With Hematoxylin NL1 HB9 HB12 HB14 NL4 HB5 NL2 NL2 HB5 HB6 HB8 HB8 HB9 HB11 HB11 HB12 HB13 HB13 HB14 HB16 HB16 NL4 With Hematoxylin HB18 NL2 HB1 HB3 NL4 HB8 HB9 Emb Fe HB5 HB16 HB18 HB3 HB14 HB16 Supplemental Figure 9. Expression of c-myc and the canonical Nrf2 target, NQO1, in a panel of human HBs. (Tissue microarray (TMA) format). IHC analysis was available for 14 cases after discounting loss of cores due to sectioning and IHC processing. Higher magnification images of a representative example of staining in normal liver (NL) and the range of staining in 5 of the HBs on the TMA is shown below. (A) 7/14 (5%) HBs expressed high levels of c-myc in a significant proportion of cells. In c-myc-positive HBs, c-myc appears more intense in the embryonal (Emb) component than in the fetal (Fe) component (see HB1). (B) 8/14 (57%) HBs expressed NQO1 in a moderate-high number of cells. NQO1 is localized to the cytoplasm of tumor cells and is heterogeneous within tumors (see HB14 and HB16). In normal liver (NL4), NQO1 is undetectable in hepatocytes, yet readily detectable in normal biliary cells (inset, arrowheads). Scoring of c-myc and NQO1 expression for each sample is shown in Supplemental Table 7. Slides were scanned on a NanoZoomer slide scanner (Hamamatsu Photonics, Hamamatsu, Japan). Low and high power images were captured using NDP View Software (Hamamatsu Photonics, Japan). AEC chromagen, red; hematoxylin, blue. HB, hepatoblastoma; NL, normal (noncancerous) liver; Ton, tonsil. Scale bars represent 5µm.
13 Supplemental Table 2: IPA Summary of Putative Tumor Enriched DEGs (β-catδex3:myc vs β-cat Comparison). Molecular and Cellular Functions Name p-value No. of Molecules Cell-To-Cell Signaling and Interaction 2.7E E-3 86 Cell Growth and Proliferation 2.66E E Cellular Movement 3.73E E-3 91 Cellular Development 7.42E E-3 13 Cell Death and Survival 1.67E E-3 18 Top Canonical Pathways Name p-value Ratio Wnt/β-catenin Signaling 3.31E-4 1/169(.59) Agranulocyte Adhesion and Diapedesis 2.95E-3 9/189 (.48) Colorectal Cancer and Metastasis 4.7E-3 1/236(.42) Granulocyte Adhesion and Diapedesis 6.73E-3 8/177 (.45) VDR/RXR Activation 7.46E-3 5/78 (.64) Top Tox Lists Name p-value Ratio Hepatic Fibrosis 1.29E-4 8/96 (.83) Increased Renal Proliferation 9.54E-4 8/129 (.62) Hepatic Stellate Cell Activation 2.8E-3 4/35(.114) Renal Glomerulus Panel (Human) 2.17E-3 3/17 (.176) LPS/IL1-Mediated Inhibition of RXR Function 6.23E-3 1/51 (.4) Hepatotoxicty Name p-value No. of Molecules Liver Fibrosis 8.2E E-1 8 Liver Inflammation/Hepatitis 9.41E-4-5.E-1 13 Liver Damage 4.65E-3-1.4E-1 1 Liver Hyperplasia/Hyperproliferation 1.13E E-1 32 Liver Steatosis 1.44E E-1 1
14 Supplemental Table 3: IPA Summary of DEGs co-regulated by β-catenin and Myc (β-catδex3:myc Livers vs WT Comparison). Molecular and Cellular Functions Name p-value No. of Molecules Cellular Movement 4.61E E Cellular Function and Maintenance 6.8E E Cell-To-Cell Signaling and Interaction 8.2E E Cellular Development 1.94E E Protein Synthesis 1.23E E Top Canonical Pathways Name p-value Ratio Complement System 2.73E-9 13/33(.394) B-Cell Development 4.E-8 12/34 (.353) LPS/IL-1-Mediated Inhibition of RXR Function 5.98E-7 3/219(.137) icos-icosl Signaling in T-Helper Cells 1.68E-6 19/18 (.176) T-Helper Cell Differentiation 1.94E-6 15/71 (.211) Top Tox Lists Name p-value Ratio LPS/IL-1-Mediated Inhibition of RXR Function 3.42E-13 44/251 (.175) NRF2-Mediated Oxidative Stress Response 8.25E-8 33/234 (.141) Xenobiotic Metabolism Signaling 1.5E-6 39/336 (.116) Persistent Renal Ischemia-Reperfusion Injury (Mouse) 1.9E-5 9/3 (.3) Cytochrome P45 Panel-Substrate is a Xenobiotic (Mouse) 2.E-5 8/25 (.32) Hepatotoxicty Name p-value No. of Molecules Liver Steatosis 1.1E E-1 35 Liver Inflammation/Hepatitis 1.34E E-1 36 Hepatocellular Carcinoma 4.41E-6-1.6E-1 49 Liver Hyperplasia/Hyperproliferation 4.41E E-1 9 Liver Cirrhosis 2.22E-5-5.3E-2 23
15 Supplemental Table 4: IPA Summary of β-cat regulated DEGs (β-cat vs WT Comparison) Molecular and Cellular Functions Name p-value No. of Molecules Cellular Function and Movement 1.66E E Cell Morphology 1.81E E Protein Synthesis 6.78E E-3 97 Cell-To-Cell Signaling and Interaction 1.23E E Cell Death and Survival 9.3E-9-1.3E Top Canonical Pathways Name p-value Ratio LPS/IL-1-Mediated Inhibition of RXR Function 1.18E-1 33/219 (.151) B-Cell Development 6.7E-8 11/34 (.324) Nicotine Degradation 5.13E-6 12/6 (.2) Xenobiotic Metabolism Signaling 8.33E-6 28/271 (.13) Glutathione-Mediated Detoxification 1.22E-5 8/28 (.286) Top Tox Lists Name p-value Ratio LPS/IL-1-Mediated Inhibition of RXR Function 3.26E-16 44/251 (.175) NRF2-Mediated Oxidative Stress Response 3.97E-11 35/234 (.15) Xenobiotic Metabolism Signaling 5.62E-1 41/336 (.122) Cytochrome P45 Panel-Substrate is a Xenobiotic (Mouse) 2.55E-1 1/25 (.4) Oxidative Stress 5.56E-8 14/57 (.246) Hepatotoxicty Name p-value No. of Molecules Liver Cholestasis 1.14E E-1 16 Glutathione Depletion 1.15E E-1 8 Liver Inflammation/Hepatitis 3.35E E-1 26 Liver Damage 8.46E-4-6.5E-1 22 Liver Necrosis/Cell Death 9.15E E-1 23
16 Supplemental Table 5: Stress and toxicity-associated gene expression in livers of β-cat:myc, β-cat and Alb-c-Myc mice Gene Symbol Gene Name Stress Pathway Fold change vs. WT Liver β-catδex3 β-catδex3:c-myc c-myc Nqo1 NAD(P)H dehydrogenase, quinone 1 Ox Slc2a1 Solute carrier family 2 (facilitated glucose transporter), member 1 Hyp Gsr Glutathione reductase Ox Hmox1 Heme oxygenase (decycling) 1 Ox Gclc Glutamate-cysteine ligase, catalytic subunit Ox Gstp1 Glutathione S-transferase Pi 1 Ox Edn1 Endothelin 1 Os Serpine1 Serpin peptidase inhibitor, clade E (plasminogen activator inhibitor type 1), member 1 Hyp Txnrd1 Thioredoxin reductase 1 Ox Tnfrsf1b Tumor necrosis factor receptor superfamily, member 1b CD/Nec Atg7 Autophagy related 7 CD/Aut Fth1 Ferritin, heavy polypeptide 1 Ox Atr Ataxia telangiectasia and Rad3 related DDR Txn1 Thioredoxin Ox Sqstm1 Sequestosome 1 Ox Bid BH3 Interacting domain death agonist HSP/UPR/CD/Apop Mre11a MRE11 Meiotic recombination 11 homolog A (S. Cerevisiae) DDR Pvr Poliovirus receptor CD/Nec Prdx1 Peroxiredoxin 1 Ox Gapdh Glyceraldehyde-3-phosphate dehydrogenase Chek1 Checkpoint kinase 1 DDR Atg12 Autophagy related 12 CD/Aut Cftr Cystic fibrosis transmembrane conductance regulator Os Gclm Glutamate-cysteine ligase, modifier subunit Ox Cdkn1a Cyclin-dependent kinase inhibitor 1a DDR Adm Adrenomedullin Hyp Rad51 RAD51 recombinase DDR Chek2 Checkpoint kinase 2 DDR Parp1 Poly (ADP-Ribose) polymerase 1 CD/Nec Arnt Aryl hydrocarbon receptor nuclear translocator Hyp Ddb2 Damage-specific DNA binding protein 2 DDR Bnip3l BCL2/Adenovirus E1B 19kDa interacting protein 3-like Hyp Atf6 Activating transcription factor 6 HSP/UPR Page 1
17 Supplemental Table 5: Stress and toxicity-associated gene expression in livers of β-cat:myc, β-cat and Alb-c-Myc mice Gene Symbol Gene Name Stress Pathway Fold change vs. WT Liver β-catδex3 β-catδex3:c-myc c-myc Xpc Xeroderma pigmentosum, complementation group C DDR Atf6b Activating transcription factor 6b HSP/UPR Tnfrsf1a Tumor necrosis factor receptor superfamily, member 1A CD/Nec Ccl12 Chemokine (C-C motif) ligand 12 IR Hus1 HUS1 checkpoint homolog (S. Pombe) DDR Hsp9ab1 Heat shock protein 9 alpha (cytosolic), class B member Becn1 Beclin 1 CD/Aut Txnl4b Thioredoxin-like 4B CD/Nec Grb2 Growth factor receptor-bound protein 2 CD/Nec Actb Actin, beta Hspa4 Heat shock 7kDa protein 4 HSP/UPR Ripk1 Receptor (TNFRSF)-interacting serine-threonine kinase 1 CD/Nec Rad9 RAD9 homolog A (S. Pombe) DDR Rad17 RAD17 homolog (S. Pombe) DDR Trp53 Tumor protein P53 DDR Hsp9aa1 Heat shock protein 9kDa alpha (cytosolic), class A member 1 HSP/UPR Atg5 Autophagy related 5 CD/Aut Nbn Nibrin DDR Atm Ataxia telangiectasia mutated DDR Akr1b3 Aldo-keto reductase family 1, member B1 (aldose reductase) Os Atf4 Activating transcription factor 4 HSP/UPR Mcl1 Myeloid cell leukemia sequence 1 (BCL2-Related) CD/Apop Hspa4l Heat shock 7kDa protein 4-Like HSP/UPR Vegfa Vascular endothelial growth factor A Hyp B2m Beta-2 microglobulin Nfat5 Nuclear factor of activated T-cells 5, tonicity-responsive Os Ddit3 DNA-damage-inducible transcript 3 DDR Dnajc3 DnaJ (Hsp4) homolog, subfamily C, member 3 HSP/UPR Hspa5 Heat shock 7kDa protein 5 (glucose-regulated protein, 78kDa) HSP/UPR Calr Calreticulin HSP/UPR Hsp9b1 Heat shock protein 9kDa beta (Grp94), member 1 HSP/UPR Gusb Glucuronidase, beta Epo Erthyropoietin Hyp Page 2
18 Supplemental Table 5: Stress and toxicity-associated gene expression in livers of β-cat:myc, β-cat and Alb-c-Myc mice Gene Symbol Gene Name Stress Pathway Fold change vs. WT Liver β-catδex3 β-catδex3:c-myc c-myc Gadd45a Growth arrest and DNA-Damage-inducible, alpha DDR Il1a Interleukin 1, alpha IR Gadd45g Growth arrest and DNA-damage-inducible, gamma DDR Xbp1 X-Box binding protein 1 HSP/UPR Car9 Carbonic anhydrase IX Hyp Ldha Lactate dehydrogenase A Hyp Aqp2 Aquaporin 2 Os Crp C-reactive protein IR Fas Fas cell surface death receptor/cd95 CD/Apop Tnf Tumor necrosis factor IR Tlr4 Toll-like receptor 4 IR Il6 Interleukin 6 (interferon, beta 2) IR Ripk3 Receptor-interacting serine-threonine kinase 3 CD/Nec Aqp4 Aquaporin 4 Os Aqp1 Aquaporin 1 Os Slc5a3 Solute carrier family 5 (sodium/myo-inositol cotransporter), member 3 Os Il1b Interleukin 1, beta IR Ulk1 Unc-51 like autophagy activating kinase 1 CD/Aut Casp1 Caspase 1 CD/Apop Mmp9 Matrix metallopeptidase 9 (gelatinase B, 92kDa type IV collagenase) Hyp Slc9a3 Solute carrier family 9, subfamily A, member 3 Os Cd4lg CD4 ligand IR Ifng Interferon, gamma IR Housekeeping genes Stress Pathway Ox: Oxidative Stress Os: Osmotic Stress Hyp: Hypoxia IR: Inflammatory Response CD: Cell Death: Apop:Apoptosis, Aut:Autophagy, Nec: Necrosis HSP/UPR: Heat Shock Proteins/Unfolded Protein Response Page 3
19 Supplemental Table 6: Oxidative stress-associated gene expression in livers of β-cat:myc, β-cat and Alb-c-Myc mice Gene Symbol Gene Name Fold-change vs. WT Liver β-catδex3 β-catδex3:c-myc c-myc nrf2 target Gpx2 Glutathione peroxidase Yes Srxn1 Sulfiredoxin 1 homolog (S. cerevisiae) Yes Nqo1 NAD(P)H dehydrogenase, quinone Yes Gpx3 Glutathione peroxidase Yes Gsr Glutathione reductase Yes Aox1 Aldehyde oxidase Yes Gclc Glutamate-cysteine ligase, catalytic subunit Yes Gstp1 Glutathione S-transferase, pi Yes Hmox1 Heme oxygenase (decycling) Yes Fmo2 Flavin containing monooxygenase Yes Nox1 NADPH oxidase Yes Gpx6 Glutathione peroxidase Yes Prdx3 Peroxiredoxin Yes Txnrd1 Thioredoxin reductase Yes Noxo1 NADPH oxidase organizer Yes Atr Ataxia telangiectasia and rad3 related Sqstm1 Sequestosome Yes Fth1 Ferritin heavy chain Yes Gpx4 Glutathione peroxidase Yes Prdx6 Peroxiredoxin Yes Prdx5 Peroxiredoxin Yes Gss Glutathione synthetase Yes Gapdh Glyceraldehyde-3-phosphate dehydrogenase Prdx2 Peroxiredoxin Yes Park7 Parkinson disease (autosomal recessive, early onset) Psmb5 Proteasome (prosome, macropain) subunit, beta type Recql4 RecQ protein-like Sod2 Superoxide dismutase 2, mitochondrial Gclm Glutamate-cysteine ligase, modifier subunit Yes Ift172 Intraflagellar transport 172 homolog (Chlamydomonas) Txn1 Thioredoxin Yes Duox1 Dual oxidase Gpx7 Glutathione peroxidase Page 1
20 Supplemental Table 6: Oxidative stress-associated gene expression in livers of β-cat:myc, β-cat and Alb-c-Myc mice Gene Symbol Gene Name Fold-change vs. WT Liver β-catδex3 β-catδex3:c-myc c-myc nrf2 target Prdx1 Peroxiredoxin Hsp9ab1 Heat shock protein 9 alpha (cytosolic), class B member Xpa Xeroderma pigmentosum, complementation group A Epx Eosinophil peroxidase Gstk1 Glutathione S-transferase kappa Actb Actin, beta Ccs Copper chaperone for superoxide dismutase Dnm2 Dynamin Prnp Prion protein Il19 Interleukin Il22 Interleukin Lpo Lactoperoxidase Mb Myoglobin Sod1 Superoxide dismutase 1, soluble Ercc6 Excision repair cross-complementing rodent repair deficiency, complementation group Cygb Cytoglobin Ptgs2 Prostaglandin-endoperoxide synthase Apc Adenomatosis polyposis coli Cat Catalase Gpx1 Glutathione peroxidase Apoe Apolipoprotein E Ercc2 Excision repair cross-complementing rodent repair deficiency, complementation group Rag2 Recombination activating gene Vim Vimentin Ctsb Cathepsin B Txnrd3 Thioredoxin reductase Txnip Thioredoxin interacting protein Gusb Glucuronidase, beta Serpinb1b Serine (or cysteine) peptidase inhibitor, clade B, member 1b Nos2 Nitric oxide synthase 2, inducible Ucp3 Uncoupling protein 3 (mitochondrial, proton carrier) Fancc Fanconi anemia, complementation group C Txnrd2 Thioredoxin reductase Page 2
21 Gene Symbol Gene Name Supplemental Table 6: Oxidative stress-associated gene expression in livers of β-cat:myc, β-cat and Alb-c-Myc mice Fold-change vs. WT Liver β-catδex3 β-catδex3:c-myc c-myc nrf2 target Gpx5 Glutathione peroxidase B2m Beta-2 microglobulin Ehd2 EH-domain containing Idh1 Isocitrate dehydrogenase 1 (NADP+), soluble Ccl5 Chemokine (C-C motif) ligand Krt1 Keratin Als2 Amyotrophic lateral sclerosis 2 (juvenile) homolog (human) Cyba Cytochrome b-245, alpha polypeptide Ucp2 Uncoupling protein 2 (mitochondrial, proton carrier) Noxa1 NADPH oxidase activator Alb Albumin Ncf2 Neutrophil cytosolic factor Ncf1 Neutrophil cytosolic factor Ngb Neuroglobin Ptgs1 Prostaglandin-endoperoxide synthase Slc38a1 Solute carrier family 38, member Sod3 Superoxide dismutase 3, extracellular Nox4 NADPH oxidase Prdx4 Peroxiredoxin Mpo Myeloperoxidase Hspa1a Heat shock protein 1A Tpo Thyroid peroxidase Scd1 Stearoyl-Coenzyme A desaturase Housekeeping genes Page 3
22 Supplemental Table 7. Scoring of c-myc and NQO1 Protein Expression in Human HBs c-myc NQO1 Tissue a Sample Intensity Score b c-myc c Tissue d Intensity Score b Core 1 Core 2 Avg Status Sample Core 1 Core 2 Avg Tonsil 3. NA 3. + Tonsil NA - NL1.5 NA.5 - NL1 NA - NL2 - NL2 - NL4 - NL4* - HB HB HB HB HB HB HB HB4 NA - HB HB HB6.5 NA.5 - HB6.5 NA.5 - HB8* HB HB9* HB9 - HB HB11 - HB NA HB12 NA - HB HB HB HB HB HB HB NA HB NA a NL- normal human liver, HB- hepatoblastoma. b NA- duplicate sample: not available or insufficient tissue c The sample was designated c-myc-positive if the average intensity score of 2 cores was > 1. Using this criterium, 5% of HBs were c-myc-positive and 5% were c-myc-negative. HBs 1, 2, 5, 12, 13, 14 and 18 were strongly positive for c-myc. * HB8 & HB9 demonstrated high background staining. NQO1 e Status d NL- normal human liver, HB- hepatoblastoma. * NL4 had no staining in hepatocyes but had robust staining in bile ducts. b NA- duplicate sample: not available or insufficient tissue e The sample was designated NQO1-positive if the average intensity score of 2 cores was equal to or greater than 1.5. Using this criterium, 57% of HBs were NQO1-positive and 43% were NQO1-negative. HBs 3, 5, 8 & 18 were strongly positive for NQO1.
23 Supplemental Table 8 Cell Line Organ Derivation- Tumor Type NFE2L2 (reads/million) NFE2L2 (compared to median) UHR Universal Daoy Brain Hep293TT Liver-HB Hep3b Liver-Pediatric HCC HepG2 Liver-Pediatric HCC MOLT4 ALL (Relapse) RDES Bone-Ewings SKES1 Ewings SKNBE2 Adrenal-NB SuPB15 ALL/Ph WERI-RB-1 Eye-Retinoblastoma Y79 Eye-Retinoblastoma WiT49 Kidney-Wilms Tumor Supplemental Table 8. Table of RNA-seq data showing relative abundance of NFE2L2 in Hep293TT cells relative to other human pediatric tumor-derived cell lines. Hep293TT cells were derived from an aggressive human HB and express NFE2L2 at a level that is 2-fold above the median. UHR; Universal Human Reference comprising 1 different human cell lines, HB; hepatoblastoma, HCC; hepatocellular carcinoma, ALL; Acute lymphoblastic leukemia, PH+; Philadelphia Chromosome-positive.
24 Supplemental Table 9: Primer sequences and PCR conditions for genotyping of mice and analysis of recombination of the β-cateninδex3 allele Genotyping Allele Forward Primer (5'-3') Reverse Primer (5'-3') Amplicon Size(s) (bp) Albumin-c-Myc ATC AGC AAC AAC CGC AAG TGC CTC CAT ACC ACC CCC CTC ~3 WT & β-cat lox(ex3) AGG GTA CCT GAA GCT CAG CG CAG TGG CTG ACA GCA GCT TT 412 (wt β-cat); 645 (β-cat lox(ex3) ) Alb-Cre GCT GCC ACG ACC AAG TGA CAG CAA TG GTA GTT ATT CGG ATC ATC AGC CAC AC 42 Allele Albumin-c-Myc WT & β-cat lox(ex3) Alb-Cre PCR Conditions 95 o C 3 min: 54 o C 1 min (1 cycle): 95 o C 1min, 54 o C 2 min, 72 o C 3 min (25 cycles): 95 o C 1 min, 54 o C 2 min, 72 o C 1 min (1 cycle): 4 o C hold 95 o C 3 min: 56 o C 1 min (1 cycle): 95 o C 1min, 56 o C 2 min, 72 o C 3 min (35 cycles): 95 o C 1 min, 56 o C 2 min, 72 o C 1 min (1 cycle): 4 o C hold 95 o C 2 min: 58 o C 2 min, 72 o C 3 min (1 cycle): 95 o C 1 min, 58 o C 3 sec, 72 o C 2 min (25 cycles): 72 o C 1 min (1 cycle): 4 o C hold Recombination Allele Forward Primer (5'-3') Reverse Primer (5'-3') Amplicon Size(s) (bp) β-catδex3 GGT AGG TGA AGC TCA GCG CAG AGC GGC CAG TAC TAG TGA ACC TCT TCG 343 PCR Conditions 95 o C 1 min (1 cycle): 95 o C 3 secs, 6 o C 1 min, 72 o C 1 min (35 cycles): 72 o C 1 min (1 cycle): 4 o C hold
25 Supplemental Table 1: Antibodies and IHC Reagents/Conditions IHC Primary Antibody Species Supplier Cat. No. Dilution/Conc Antigen Retrieval Buffer Time (mins) Afp Rabbit DAKO A8 1 in 2 Not Required β-catenin Mouse BD Biosciences in 5 Tris/EDTA/Tween ph 8 5 Cited1 Rabbit Sigma AV in 3 Citrate ph 6 12 Dlk1 Rabbit Santa Cruz sc in 1 Citrate ph 6 15 Epcam Rabbit Sino Biological 5591-RP2 1 µg/ml Citrate ph6 15 Glul Mouse BD Biosciences in 1 Citrate ph 6 2 Lin28a Rabbit Cell Signaling in 5 Citrate ph 6 12 Lin28b Rabbit Proteintech AP 1 in 5 Not Required Myc Rabbit Santa Cruz sc in 15 Citrate ph 6 5 Nqo1 Rabbit Sigma HPA738 1 in 5 Citrate ph6 3 PCNA Mouse Novocastra NCL-L-PCNA 1 in 75 Citrate ph 6 1 Sox9 Rabbit Millipore AB in 1 Citrate ph 6 12 Secondary Antibody Species Supplier Cat. No. Dilution Biotinylated anti-rabbit Goat Jackson Immunoresearch in 5 Biotinylated anti-mouse Rabbit Jackson Immunoresearch in 175 Other Reagents Supplier Cat. No. Dilution Streptavidin-HRP Hematoxylin QS AquaPolymount AEC Chromagen Invitrogen in 75 Vector Labs H-344 As supplied Polysciences Inc As supplied Invitrogen -27 As supplied Western Blotting Primary Antibody Species Supplier Cat. No. Dilution β-catenin Mouse BD Biosciences in 2 GAPDH Rabbit Cell Signaling in 2 Nqo1 Mouse Santa Cruz SC in 1 p62/sqstm1 Rabbit Cell Signaling in 1
26 Supplemental Table 11: Primers for Semi-quantitative (Sq) RT-PCR Analysis of Gene Expression (Mice) mrna Forward Primer (5' - 3') Reverse Primer (5' - 3') Amplicon Size (bp) Source Afp CTT CCC TCA TCC TCC TGC TAC ACA AAC TGG GTA AAG GTG ATG G 145 Primer Bank Bex1 AGT GAG TCC AAA GAT CAA GGC G CTG GCT CCC TTC TGA TGG TA 16 Primer Bank Cbr3 GTA ACT GGG GCT AAC AAA GGC TTG ACC AGC ACG TTA AGT CCC 239 Primer Bank Cited1 AAC CTT GGA GTG AAG GAT CGC GTA GGA GAG CCT ATT GGA GAT GT 128 Primer Bank Cyclophilin A (Ppia) GAG CTG TTT GCA GAC AAA GTT C CCC TGG CAC ATG AAT CCT GG 125 Primer Bank Dlk1 AGT GCG AAA CCT GGG TGT C GCC TCC TTG TTG AAA GTG GTC A 148 Primer Bank Ggt1 TTT GTC ATC ATC GGC CTC TGT CCC GTC CAA TCT CTG AGC AG 115 Primer Bank Gpr49 AAT CGC GGT AGT GGA CAT TC GAT TCG GAA GCA AAA ATG GA 155 Giera et al. (21) Gpx2 GCC TCA AGT ATG TCC GAC CTG GGA GAA CGG GTC ATC ATA AGG G 143 Primer Bank Gsta1 AAG CCC GTG CTT CAC TAC TTC GGG CAC TTG GTC AAA CAT CAA A 159 Primer Bank Gsta4 TGA TTG CCG TGG CTC CAT TTA CAA CGA GAA AAG CCT CTC CGT 135 Primer Bank Gstm2 TGG AAC CCA AAG TAG GAT TAC AAA TGA GGA CCA AGG CAG CAC AC 313 Giera et al. (21) Igf2 GTG CTG CAT CGC TGC TTA C ACG TCC CTC TCG GAC TTG G 222 Primer Bank Nope CTC TGC AAG AGA CAT TCT CAG AC GTG GGA GCT TTG TCG TAG GC 125 Primer Bank Nqo1 TAT CCT TCC GAG TCA TCT CTA GCA TCT GCA GCT TCC AGC TCC TTG 85 Qu et al. (214) Peg3 GAG ACC ATC ATG CCT AGC CC GTT CAC GCT CAT TAT CTC TGT CA 172 Primer Bank Phgdh ATG GCC TTC GCA AAT CTG C AGT TCA GCT ATC AGC TCC TCC 134 Primer Bank Rasl1b GAC AGC TTT GAG TAC GTC AAG A TAG CCG CAC TTC CAG GTC T 173 Primer Bank S1a9 ATA CTC TAG GAA GGA AGG ACA CC TCC ATG ATG TCA TTT ATG AGG GC 129 Primer Bank Wisp2 CGC TGT GAT GAC GGT GGT TT CCT GGC ACC TGT ATT CTC CTG 11 Primer Bank Wnt4 AGA CGT GCG AGA AAC TCA AAG GGA ACT GGT ATT GGC ACT CCT 126 Primer Bank References: Geira et al. (21) Toxicol. Sciences. 115: Qu et al. (214) Toxicology. 324:35-42
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53
 1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at
 Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Materials
 Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
CD31 5'-AGA GAC GGT CTT GTC GCA GT-3' 5 ' -TAC TGG GCT TCG AGA GCA GT-3'
 Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Supplemental Data. Shin et al. Plant Cell. (2012) /tpc YFP N
 MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
Abbreviations: P- paraffin-embedded section; C, cryosection; Bio-SA, biotin-streptavidin-conjugated fluorescein amplification.
 Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most
 Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1 a
 Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Figure S1. Analysis of genomic and cdna sequences of the targeted regions in WT-KI and
 Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
Figure S1. Analysis of genomic and sequences of the targeted regions in and indicated mutant KI cells, with WT and corresponding mutant sequences underlined. (A) cells; (B) K21E-KI cells; (C) D33A-KI cells;
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards
 Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. Lats1/2 deleted ihbs and ihps showed decreased transcripts of hepatocyte related genes (a and b) Western blots (a) and recombination PCR (b) of control and
Supplementary Figure 1 Supplementary Figure 1. Lats1/2 deleted ihbs and ihps showed decreased transcripts of hepatocyte related genes (a and b) Western blots (a) and recombination PCR (b) of control and
SUPPLEMENTARY DATA. Supplementary Table 1. Primer sequences for qrt-pcr
 Supplementary Table 1. Primer sequences for qrt-pcr Gene PRDM16 UCP1 PGC1α Dio2 Elovl3 Cidea Cox8b PPARγ AP2 mttfam CyCs Nampt NRF1 16s-rRNA Hexokinase 2, intron 9 β-actin Primer Sequences 5'-CCA CCA GCG
Supplementary Table 1. Primer sequences for qrt-pcr Gene PRDM16 UCP1 PGC1α Dio2 Elovl3 Cidea Cox8b PPARγ AP2 mttfam CyCs Nampt NRF1 16s-rRNA Hexokinase 2, intron 9 β-actin Primer Sequences 5'-CCA CCA GCG
Supplementary Table 2. Conserved regulatory elements in the promoters of CD36.
 Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
Journal of Cell Science Supplementary information. Arl8b +/- Arl8b -/- Inset B. electron density. genotype
 J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
Citation for published version (APA): Oosterveer, M. H. (2009). Control of metabolic flux by nutrient sensors Groningen: s.n.
 University of Groningen Control of metabolic flux by nutrient sensors Oosterveer, Maaike IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
University of Groningen Control of metabolic flux by nutrient sensors Oosterveer, Maaike IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature05883 SUPPLEMENTARY INFORMATION Supplemental Figure 1 Prostaglandin agonists and antagonists alter runx1/cmyb expression. a-e, Embryos were exposed to (b) PGE2 and (c) PGI2 (20μM) and
doi: 10.1038/nature05883 SUPPLEMENTARY INFORMATION Supplemental Figure 1 Prostaglandin agonists and antagonists alter runx1/cmyb expression. a-e, Embryos were exposed to (b) PGE2 and (c) PGI2 (20μM) and
Table S1. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9. Assay (s) Target Name Sequence (5 3 ) Comments
 SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
Supplementary Figure 1
 Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Nature Immunology: doi: /ni.3836
 Supplementary Figure 1 Recombinant LIGHT-VTP induces pericyte contractility and endothelial cell activation. (a) Western blot showing purification steps for full length murine LIGHT-VTP (CGKRK) protein:
Supplementary Figure 1 Recombinant LIGHT-VTP induces pericyte contractility and endothelial cell activation. (a) Western blot showing purification steps for full length murine LIGHT-VTP (CGKRK) protein:
Astaxanthin prevents and reverses diet-induced insulin resistance and. steatohepatitis in mice: A comparison with vitamin E
 Supplementary Information Astaxanthin prevents and reverses diet-induced insulin resistance and steatohepatitis in mice: A comparison with vitamin E Yinhua Ni, 1,2 Mayumi Nagashimada, 1 Fen Zhuge, 1 Lili
Supplementary Information Astaxanthin prevents and reverses diet-induced insulin resistance and steatohepatitis in mice: A comparison with vitamin E Yinhua Ni, 1,2 Mayumi Nagashimada, 1 Fen Zhuge, 1 Lili
BIOLOGY 621 Identification of the Snorks
 Name: Date: Block: BIOLOGY 621 Identification of the Snorks INTRODUCTION: In this simulation activity, you will examine the DNA sequence of a fictitious organism - the Snork. Snorks were discovered on
Name: Date: Block: BIOLOGY 621 Identification of the Snorks INTRODUCTION: In this simulation activity, you will examine the DNA sequence of a fictitious organism - the Snork. Snorks were discovered on
Supplementary Figure 1a
 Supplementary Figure 1a Hours: E-cadherin TGF-β On TGF-β Off 0 12 24 36 48 24 48 72 Vimentin βactin Fig. S1a. Treatment of AML12 cells with TGF-β induces EMT. Treatment of AML12 cells with TGF-β results
Supplementary Figure 1a Hours: E-cadherin TGF-β On TGF-β Off 0 12 24 36 48 24 48 72 Vimentin βactin Fig. S1a. Treatment of AML12 cells with TGF-β induces EMT. Treatment of AML12 cells with TGF-β results
Description of Supplementary Files. File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables
 Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
Supplemental Information. Cancer-Associated Fibroblasts Neutralize. the Anti-tumor Effect of CSF1 Receptor Blockade
 Cancer Cell, Volume 32 Supplemental Information Cancer-Associated Fibroblasts Neutralize the Anti-tumor Effect of CSF1 Receptor Blockade by Inducing PMN-MDSC Infiltration of Tumors Vinit Kumar, Laxminarasimha
Cancer Cell, Volume 32 Supplemental Information Cancer-Associated Fibroblasts Neutralize the Anti-tumor Effect of CSF1 Receptor Blockade by Inducing PMN-MDSC Infiltration of Tumors Vinit Kumar, Laxminarasimha
Supplementary information
 Supplementary information Unique polypharmacology nuclear receptor modulator blocks inflammatory signaling pathways Mi Ra Chang 1, Anthony Ciesla 1, Timothy S. Strutzenberg 1, Scott J. Novick 1, Yuanjun
Supplementary information Unique polypharmacology nuclear receptor modulator blocks inflammatory signaling pathways Mi Ra Chang 1, Anthony Ciesla 1, Timothy S. Strutzenberg 1, Scott J. Novick 1, Yuanjun
Supplemental Information. Th17 Lymphocytes Induce Neuronal. Cell Death in a Human ipsc-based. Model of Parkinson's Disease
 Cell Stem Cell, Volume 23 Supplemental Information Th17 Lymphocytes Induce Neuronal Cell Death in a Human ipsc-based Model of Parkinson's Disease Annika Sommer, Franz Maxreiter, Florian Krach, Tanja Fadler,
Cell Stem Cell, Volume 23 Supplemental Information Th17 Lymphocytes Induce Neuronal Cell Death in a Human ipsc-based Model of Parkinson's Disease Annika Sommer, Franz Maxreiter, Florian Krach, Tanja Fadler,
Plasmids Western blot analysis and immunostaining Flow Cytometry Cell surface biotinylation RNA isolation and cdna synthesis
 Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Cross-talk between mineralocorticoid and angiotensin II signaling for cardiac
 ONLINE SUPPLEMENT TO Crosstalk between mineralocorticoid and angiotensin II signaling for cardiac remodeling An Di ZHANG,,3, Aurelie NGUYEN DINH CAT*,,3, Christelle SOUKASEUM *,,3, Brigitte ESCOUBET, 4,
ONLINE SUPPLEMENT TO Crosstalk between mineralocorticoid and angiotensin II signaling for cardiac remodeling An Di ZHANG,,3, Aurelie NGUYEN DINH CAT*,,3, Christelle SOUKASEUM *,,3, Brigitte ESCOUBET, 4,
Culture Density (OD600) 0.1. Culture Density (OD600) Culture Density (OD600) Culture Density (OD600) Culture Density (OD600)
 A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
SUPPORTING INFORMATION
 SUPPORTING INFORMATION Biology is different in small volumes: endogenous signals shape phenotype of primary hepatocytes cultured in microfluidic channels Amranul Haque, Pantea Gheibi, Yandong Gao, Elena
SUPPORTING INFORMATION Biology is different in small volumes: endogenous signals shape phenotype of primary hepatocytes cultured in microfluidic channels Amranul Haque, Pantea Gheibi, Yandong Gao, Elena
A smart acid nanosystem for ultrasensitive. live cell mrna imaging by the target-triggered intracellular self-assembly
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
 www.lessonplansinc.com Topic: Protein Synthesis - Sentence Activity Summary: Students will simulate transcription and translation by building a sentence/polypeptide from words/amino acids. Goals & Objectives:
www.lessonplansinc.com Topic: Protein Synthesis - Sentence Activity Summary: Students will simulate transcription and translation by building a sentence/polypeptide from words/amino acids. Goals & Objectives:
Supplementary Materials and Methods
 DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because
 Supplementary Figure S1 Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because importin-α6 was shown to be testis-specific. Human and chicken importin protein
Supplementary Figure S1 Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because importin-α6 was shown to be testis-specific. Human and chicken importin protein
Supplementary Information
 Supplementary Information Remodeling of heterochromatin structure slows neuropathological progression and prolongs survival in an animal model of Huntington s disease Junghee Lee, Yu Jin Hwang, Yunha Kim,
Supplementary Information Remodeling of heterochromatin structure slows neuropathological progression and prolongs survival in an animal model of Huntington s disease Junghee Lee, Yu Jin Hwang, Yunha Kim,
Lezione 10. Sommario. Bioinformatica. Lezione 10: Sintesi proteica Synthesis of proteins Central dogma: DNA makes RNA makes proteins Genetic code
 Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Lezione 10: Sintesi proteica Synthesis of proteins Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza
Lezione 10 Bioinformatica Mauro Ceccanti e Alberto Paoluzzi Lezione 10: Sintesi proteica Synthesis of proteins Dip. Informatica e Automazione Università Roma Tre Dip. Medicina Clinica Università La Sapienza
A basic helix loop helix transcription factor controls cell growth
 A basic helix loop helix transcription factor controls cell growth and size in root hairs Keke Yi 1,2, Benoît Menand 1,3, Elizabeth Bell 1, Liam Dolan 1,4 Supplementary note Low soil phosphate availability
A basic helix loop helix transcription factor controls cell growth and size in root hairs Keke Yi 1,2, Benoît Menand 1,3, Elizabeth Bell 1, Liam Dolan 1,4 Supplementary note Low soil phosphate availability
Hepatoblastoma modeling in mice places Nrf2 within a cancer field established by mutant b- catenin
 Hepatoblastoma modeling in mice places Nrf2 within a cancer field established by mutant b- catenin Sarah A. Comerford,, Gail E. Tomlinson, Robert E. Hammer JCI Insight. 2016;1(16):e88549. https://doi.org/10.1172/jci.insight.88549.
Hepatoblastoma modeling in mice places Nrf2 within a cancer field established by mutant b- catenin Sarah A. Comerford,, Gail E. Tomlinson, Robert E. Hammer JCI Insight. 2016;1(16):e88549. https://doi.org/10.1172/jci.insight.88549.
Formylpeptide receptor2 contributes to colon epithelial homeostasis, inflammation, and tumorigenesis
 Supplementary Data Formylpeptide receptor2 contributes to colon epithelial homeostasis, inflammation, and tumorigenesis Keqiang Chen, Mingyong Liu, Ying Liu, Teizo Yoshimura, Wei Shen, Yingying Le, Scott
Supplementary Data Formylpeptide receptor2 contributes to colon epithelial homeostasis, inflammation, and tumorigenesis Keqiang Chen, Mingyong Liu, Ying Liu, Teizo Yoshimura, Wei Shen, Yingying Le, Scott
ice-cold 70% ethanol with gentle vortexing, incubated at -20 C for 4 hours, and washed with PBS.
 Cell cycle analysis For cell cycle analysis, single cell suspensions of E12.5 fetal liver cells were suspended in 4 ml ice-cold 7% ethanol with gentle vortexing, incubated at -2 C for 4 hours, and washed
Cell cycle analysis For cell cycle analysis, single cell suspensions of E12.5 fetal liver cells were suspended in 4 ml ice-cold 7% ethanol with gentle vortexing, incubated at -2 C for 4 hours, and washed
Single-Molecule Analysis of Gene Expression Using Two-Color RNA- Labeling in Live Yeast
 Supplemental Figures, Tables and Results Single-Molecule Analysis of Gene Expression Using Two-Color RNA- Labeling in Live Yeast Sami Hocine 1, Pascal Raymond 2, Daniel Zenklusen 2, Jeffrey A. Chao 1 &
Supplemental Figures, Tables and Results Single-Molecule Analysis of Gene Expression Using Two-Color RNA- Labeling in Live Yeast Sami Hocine 1, Pascal Raymond 2, Daniel Zenklusen 2, Jeffrey A. Chao 1 &
Supplementary Figure 1
 Metastatic melanoma Primary melanoma Healthy human skin Supplementary Figure 1 CD22 IgG4 Supplementary Figure 1: Immunohisochemical analysis of CD22+ (left) and IgG4 (right), cells (shown in red and indicated
Metastatic melanoma Primary melanoma Healthy human skin Supplementary Figure 1 CD22 IgG4 Supplementary Figure 1: Immunohisochemical analysis of CD22+ (left) and IgG4 (right), cells (shown in red and indicated
Supplementary Information. Bamboo shoot fiber prevents obesity in mice by. modulating the gut microbiota
 Supplementary Information Bamboo shoot fiber prevents obesity in mice by modulating the gut microbiota Xiufen Li 1,2, Juan Guo 1, Kailong Ji 1,2, and Ping Zhang 1,* 1 Key Laboratory of Tropical Plant Resources
Supplementary Information Bamboo shoot fiber prevents obesity in mice by modulating the gut microbiota Xiufen Li 1,2, Juan Guo 1, Kailong Ji 1,2, and Ping Zhang 1,* 1 Key Laboratory of Tropical Plant Resources
BHP 2-7 and Nthy-ori 3-1 cells were grown in RPMI1640 medium (Hyclone) supplemented with 10% fetal bovine serum (Gibco), 2mM L-glutamine, and 100 U/mL
 1 2 3 4 Materials and Methods Cell culture BHP 2-7 and Nthy-ori 3-1 cells were grown in RPMI1640 medium (Hyclone) 5 supplemented with 10% fetal bovine serum (Gibco), 2mM L-glutamine, and 100 U/mL 6 penicillin-streptomycin.
1 2 3 4 Materials and Methods Cell culture BHP 2-7 and Nthy-ori 3-1 cells were grown in RPMI1640 medium (Hyclone) 5 supplemented with 10% fetal bovine serum (Gibco), 2mM L-glutamine, and 100 U/mL 6 penicillin-streptomycin.
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
SUPPLEMENTAL FIGURE 1
 SUPPLEMENTL FIGURE 1 C Supplemental Figure 1. pproach for removal of snorns from Rpl13a gene. () Wild type Rpl13a exonintron structure is shown, with exo in black and intronic snorns in red rectangles.
SUPPLEMENTL FIGURE 1 C Supplemental Figure 1. pproach for removal of snorns from Rpl13a gene. () Wild type Rpl13a exonintron structure is shown, with exo in black and intronic snorns in red rectangles.
TetR repressor-based bioreporters for the detection of doxycycline using Escherichia
 Supplementary materials TetR repressor-based bioreporters for the detection of doxycycline using Escherichia coli and Acinetobacter oleivorans Hyerim Hong and Woojun Park * Department of Environmental
Supplementary materials TetR repressor-based bioreporters for the detection of doxycycline using Escherichia coli and Acinetobacter oleivorans Hyerim Hong and Woojun Park * Department of Environmental
Loyer, et al. microrna-21 contributes to NASH Suppl 1/15
 Loyer, et al. microrna-21 contributes to NASH Suppl 1/15 SUPPLEMENTARY MATERIAL: Liver MicroRNA-21 is Overexpressed in Non Alcoholic Steatohepatitis and Contributes to the Disease in Experimental Models
Loyer, et al. microrna-21 contributes to NASH Suppl 1/15 SUPPLEMENTARY MATERIAL: Liver MicroRNA-21 is Overexpressed in Non Alcoholic Steatohepatitis and Contributes to the Disease in Experimental Models
PATIENTS AND METHODS. Subjects
 PATIENTS AND METHODS Subjects Twenty-nine morbidly obese subjects involved in a gastric surgery program were enrolled in the study between October 25 and March 21. Bariatric surgery was performed in patients
PATIENTS AND METHODS Subjects Twenty-nine morbidly obese subjects involved in a gastric surgery program were enrolled in the study between October 25 and March 21. Bariatric surgery was performed in patients
McAlpine PERK-GSK3 regulates foam cell formation. Supplemental Material. Supplementary Table I. Sequences of real time PCR primers.
 Mclpine PERK-GSK3 regulates foam cell formation Supplemental Material Supplementary Table I. Sequences of real time PCR primers. Primer Name Primer Sequences (5-3 ) Product Size (bp) GRP78 (human) Fwd:
Mclpine PERK-GSK3 regulates foam cell formation Supplemental Material Supplementary Table I. Sequences of real time PCR primers. Primer Name Primer Sequences (5-3 ) Product Size (bp) GRP78 (human) Fwd:
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Supplemental Figures: Supplemental Figure 1
 Supplemental Figures: Supplemental Figure 1 Suppl. Figure 1. BM-DC infection with H. pylori does not induce cytotoxicity and treatment of BM-DCs with H. pylori sonicate, but not heat-inactivated bacteria,
Supplemental Figures: Supplemental Figure 1 Suppl. Figure 1. BM-DC infection with H. pylori does not induce cytotoxicity and treatment of BM-DCs with H. pylori sonicate, but not heat-inactivated bacteria,
Isoprene-Derived Secondary Organic Aerosol Induces the Expression of. Oxidative Stress Response Genes in Human Lung Cells
 1 2 3 Supporting Information Isoprene-Derived Secondary Organic Aerosol Induces the Expression of Oxidative Stress Response Genes in Human Lung Cells 4 5 6 7 8 9 10 11 12 13 14 15 Ying-Hsuan Lin 1,a, Maiko
1 2 3 Supporting Information Isoprene-Derived Secondary Organic Aerosol Induces the Expression of Oxidative Stress Response Genes in Human Lung Cells 4 5 6 7 8 9 10 11 12 13 14 15 Ying-Hsuan Lin 1,a, Maiko
Supporting Information
 Supporting Information Malapeira et al. 10.1073/pnas.1217022110 SI Materials and Methods Plant Material and Growth Conditions. A. thaliana seedlings were stratified at 4 C in the dark for 3 d on Murashige
Supporting Information Malapeira et al. 10.1073/pnas.1217022110 SI Materials and Methods Plant Material and Growth Conditions. A. thaliana seedlings were stratified at 4 C in the dark for 3 d on Murashige
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
CIRCRESAHA/2004/098145/R1 - ONLINE 1. Validation by Semi-quantitative Real-Time Reverse Transcription PCR
 CIRCRESAHA/2004/098145/R1 - ONLINE 1 Expanded Materials and Methods Validation by Semi-quantitative Real-Time Reverse Transcription PCR Expression patterns of 13 genes (Online Table 2), selected with respect
CIRCRESAHA/2004/098145/R1 - ONLINE 1 Expanded Materials and Methods Validation by Semi-quantitative Real-Time Reverse Transcription PCR Expression patterns of 13 genes (Online Table 2), selected with respect
SUPPLEMENTARY INFORMATION
 BASELINE ISCHAEMIA a b Phd2 +/- c d Collateral growth and maintenance SMC recruitment SMC proliferation Phd2 +/- NF- B off NF- B on NF- B on NF- B on Endothelial cell Smooth muscle cell Pro-arteriogenic
BASELINE ISCHAEMIA a b Phd2 +/- c d Collateral growth and maintenance SMC recruitment SMC proliferation Phd2 +/- NF- B off NF- B on NF- B on NF- B on Endothelial cell Smooth muscle cell Pro-arteriogenic
Viral hepatitis, which affects half a billion people
 GASTROENTEROLOGY 2006;130:435 452 BASIC LIVER, PANCREAS, AND BILIARY TRACT Natural Killer Cells Ameliorate Liver Fibrosis by Killing Activated Stellate Cells in NKG2D-Dependent and Tumor Necrosis Factor
GASTROENTEROLOGY 2006;130:435 452 BASIC LIVER, PANCREAS, AND BILIARY TRACT Natural Killer Cells Ameliorate Liver Fibrosis by Killing Activated Stellate Cells in NKG2D-Dependent and Tumor Necrosis Factor
SUPPLEMENTARY RESULTS
 SUPPLEMENTARY RESULTS Supplementary Table 1. hfpr1- Flpln-CHO hfpr2-flpln-cho pec 50 E max (%) Log( /K A) Log( /K A) N pec 50 E max (%) Log( /K A) Log( /K A) n ERK1/2 phosphorylation fmlp 9.0±0.6 80±7
SUPPLEMENTARY RESULTS Supplementary Table 1. hfpr1- Flpln-CHO hfpr2-flpln-cho pec 50 E max (%) Log( /K A) Log( /K A) N pec 50 E max (%) Log( /K A) Log( /K A) n ERK1/2 phosphorylation fmlp 9.0±0.6 80±7
Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients
 Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients Shuichi OTABE, Karine CLEMENT, Séverine DUBOIS, Frederic LEPRETRE, Veronique PELLOUX,
Mutation Screening and Association Studies of the Human UCP 3 Gene in Normoglycemic and NIDDM Morbidly Obese Patients Shuichi OTABE, Karine CLEMENT, Séverine DUBOIS, Frederic LEPRETRE, Veronique PELLOUX,
Integration Solutions
 Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
Supporting Information. Mutational analysis of a phenazine biosynthetic gene cluster in
 Supporting Information for Mutational analysis of a phenazine biosynthetic gene cluster in Streptomyces anulatus 9663 Orwah Saleh 1, Katrin Flinspach 1, Lucia Westrich 1, Andreas Kulik 2, Bertolt Gust
Supporting Information for Mutational analysis of a phenazine biosynthetic gene cluster in Streptomyces anulatus 9663 Orwah Saleh 1, Katrin Flinspach 1, Lucia Westrich 1, Andreas Kulik 2, Bertolt Gust
Advanced Subsidiary Unit 1: Lifestyle, Transport, Genes and Health
 Write your name here Surname Other names Edexcel GCE Centre Number Candidate Number Biology Advanced Subsidiary Unit 1: Lifestyle, Transport, Genes and Health Thursday 8 January 2009 Morning Time: 1 hour
Write your name here Surname Other names Edexcel GCE Centre Number Candidate Number Biology Advanced Subsidiary Unit 1: Lifestyle, Transport, Genes and Health Thursday 8 January 2009 Morning Time: 1 hour
without LOI phenotype by breeding female wild-type C57BL/6J and male H19 +/.
 Sakatani et al. 1 Supporting Online Material Materials and methods Mice and genotyping: H19 mutant mice with C57BL/6J background carrying a deletion in the structural H19 gene (3 kb) and 10 kb of 5 flanking
Sakatani et al. 1 Supporting Online Material Materials and methods Mice and genotyping: H19 mutant mice with C57BL/6J background carrying a deletion in the structural H19 gene (3 kb) and 10 kb of 5 flanking
Malignant Amelanotic Melanoma of the Pleura without Primary Skin Lesion: An Autopsy Case Report. a a*
 2009 63 6 379 384 Malignant Amelanotic Melanoma of the Pleura without Primary Skin Lesion: An Autopsy Case Report a b a a a* a b 380 63 6 Chest x ray and computed tomography (CT). A, Chest x ray on admission
2009 63 6 379 384 Malignant Amelanotic Melanoma of the Pleura without Primary Skin Lesion: An Autopsy Case Report a b a a a* a b 380 63 6 Chest x ray and computed tomography (CT). A, Chest x ray on admission
Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and Gerard D. Wright.
 Supplementary Data for TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and
Supplementary Data for TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and
Baseline clinical characteristics for the 81 CMML patients Routine diagnostic testing and statistical analyses... 3
 Next-Generation Sequencing Technology Reveals a Characteristic Pattern of Molecular Mutations in 72.8% of Chronic Myelomonocytic Leukemia (CMML) by Detecting Frequent Alterations in TET2, CBL, RAS, and
Next-Generation Sequencing Technology Reveals a Characteristic Pattern of Molecular Mutations in 72.8% of Chronic Myelomonocytic Leukemia (CMML) by Detecting Frequent Alterations in TET2, CBL, RAS, and
Nucleotide Sequence of the Australian Bluetongue Virus Serotype 1 RNA Segment 10
 J. gen. Virol. (1988), 69, 945-949. Printed in Great Britain 945 Key words: BTV/genome segment lo/nucleotide sequence Nucleotide Sequence of the Australian Bluetongue Virus Serotype 1 RNA Segment 10 By
J. gen. Virol. (1988), 69, 945-949. Printed in Great Britain 945 Key words: BTV/genome segment lo/nucleotide sequence Nucleotide Sequence of the Australian Bluetongue Virus Serotype 1 RNA Segment 10 By
Relationship of the APOA5/A4/C3/A1 gene cluster and APOB gene polymorphisms with dyslipidemia
 elationship of the APOA5/A4/C3/A1 gene cluster and APOB gene polymorphisms with dyslipidemia H.J. Ou 1, G. Huang 2, W. Liu 3, X.L. Ma 2, Y. Wei 4, T. Zhou 5 and Z.M. Pan 3 1 Department of Neurology, The
elationship of the APOA5/A4/C3/A1 gene cluster and APOB gene polymorphisms with dyslipidemia H.J. Ou 1, G. Huang 2, W. Liu 3, X.L. Ma 2, Y. Wei 4, T. Zhou 5 and Z.M. Pan 3 1 Department of Neurology, The
SUPPLEMENTAL METHODS Cell culture RNA extraction and analysis Immunohistochemical analysis and laser capture microdissection (LCM)
 SUPPLEMENTAL METHODS Cell culture Human peripheral blood mononuclear cells were isolated from healthy donors by Ficoll density gradient centrifugation. Monocyte differentiation to resting macrophages ()
SUPPLEMENTAL METHODS Cell culture Human peripheral blood mononuclear cells were isolated from healthy donors by Ficoll density gradient centrifugation. Monocyte differentiation to resting macrophages ()
An epithelial circadian clock controls pulmonary inflammation and glucocorticoid action
 An epithelial circadian clock controls pulmonary inflammation and glucocorticoid action Supplementary Figure : Expression levels of toll-like receptor 4 (Tlr4) in muse lung does not change throughout the
An epithelial circadian clock controls pulmonary inflammation and glucocorticoid action Supplementary Figure : Expression levels of toll-like receptor 4 (Tlr4) in muse lung does not change throughout the
Expression of Selected Inflammatory Cytokine Genes in Bladder Biopsies
 Borneo Journal of Resource Science and Technology (2013) 3(2): 15-20 Expression of Selected Inflammatory Cytokine Genes in Bladder Biopsies EDMUND UI-HANG SIM *1, NUR DIANA ANUAR 2, TENG-AIK ONG 3, GUAN-
Borneo Journal of Resource Science and Technology (2013) 3(2): 15-20 Expression of Selected Inflammatory Cytokine Genes in Bladder Biopsies EDMUND UI-HANG SIM *1, NUR DIANA ANUAR 2, TENG-AIK ONG 3, GUAN-
*To whom correspondence should be addressed. This PDF file includes:
 www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
Supplementary Figure S1
 Supplementry Figure S Tissue weights (g).... Liver Hert Brin Pncres Len mss (g) 8 6 -% +% 8 6 Len mss Len mss (g) (% ody weight) Len mss (% ody weight) c Tiilis nterior weight (g).6...... Qudriceps weight
Supplementry Figure S Tissue weights (g).... Liver Hert Brin Pncres Len mss (g) 8 6 -% +% 8 6 Len mss Len mss (g) (% ody weight) Len mss (% ody weight) c Tiilis nterior weight (g).6...... Qudriceps weight
Figure S1. IRF5 mrna expression is not expressed modulated by steatosis grade in
 9-INS-RG-TR- SUPPLEMENTRY MTERILS B IRF mrn expression 1 Control Fatty liver NSH HCV αsm IRF 1 ERG IRF < -33 33- Steatosis (%) < Figure S1. IRF mrn expression is not expressed modulated by steatosis grade
9-INS-RG-TR- SUPPLEMENTRY MTERILS B IRF mrn expression 1 Control Fatty liver NSH HCV αsm IRF 1 ERG IRF < -33 33- Steatosis (%) < Figure S1. IRF mrn expression is not expressed modulated by steatosis grade
Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany
 HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
HLA ISSN 2059-2302 BRIEF COMMUNICATION Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany C. J. Hernández-Frederick 1, N. Cereb 2,A.S.Giani 1, J.
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/473/eaai7696/dc1 Supplementary Materials for Astrocyte-shed extracellular vesicles regulate the peripheral leukocyte response to inflammatory brain lesions
www.sciencesignaling.org/cgi/content/full/10/473/eaai7696/dc1 Supplementary Materials for Astrocyte-shed extracellular vesicles regulate the peripheral leukocyte response to inflammatory brain lesions
Isolate Sexual Idiomorph Species
 SUPLEMENTARY TABLE 1. Isolate identification, sexual idiomorph and species of each isolate used for MAT locus distribution in Paracoccidioides species. Isolate Sexual Idiomorph Species Pb01 MAT1-1 P. lutzii
SUPLEMENTARY TABLE 1. Isolate identification, sexual idiomorph and species of each isolate used for MAT locus distribution in Paracoccidioides species. Isolate Sexual Idiomorph Species Pb01 MAT1-1 P. lutzii
University of Groningen. Vasoregression in incipient diabetic retinopathy Pfister, Frederick
 University of Groningen Vasoregression in incipient diabetic retinopathy Pfister, Frederick IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from
University of Groningen Vasoregression in incipient diabetic retinopathy Pfister, Frederick IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from
Mutation analysis of a Chinese family with oculocutaneous albinism
 /, 2016, Vol. 7, (No. 51), pp: 84981-84988 Mutation analysis of a Chinese family with oculocutaneous albinism Xiong Wang 1, Yaowu Zhu 1, Na Shen 1, Jing Peng 1, Chunyu Wang 1, Haiyi Liu 2, Yanjun Lu 1
/, 2016, Vol. 7, (No. 51), pp: 84981-84988 Mutation analysis of a Chinese family with oculocutaneous albinism Xiong Wang 1, Yaowu Zhu 1, Na Shen 1, Jing Peng 1, Chunyu Wang 1, Haiyi Liu 2, Yanjun Lu 1
Cancer Genetics 204 (2011) 45e52
 Cancer Genetics 204 (2011) 45e52 Exon scanning by reverse transcriptaseepolymerase chain reaction for detection of known and novel EML4eALK fusion variants in nonesmall cell lung cancer Heather R. Sanders
Cancer Genetics 204 (2011) 45e52 Exon scanning by reverse transcriptaseepolymerase chain reaction for detection of known and novel EML4eALK fusion variants in nonesmall cell lung cancer Heather R. Sanders
3) Table_S1: Clinical Characteristics of Breast Cancer Patients. 5) Table_S3: Primer sequences used for qt-pcr of ChIP samples
 Supplemental Section: 1) Eight supplemental figures and legends 2) Supplemental Materials and Methods 3) Table_S1: Clinical Characteristics of Breast Cancer Patients 4) Table_S2: Oligonucleotide sequences
Supplemental Section: 1) Eight supplemental figures and legends 2) Supplemental Materials and Methods 3) Table_S1: Clinical Characteristics of Breast Cancer Patients 4) Table_S2: Oligonucleotide sequences
What do you think of when you here the word genome?
 What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
Mechanistic and functional insights into fatty acid activation in Mycobacterium tuberculosis SUPPLEMENTARY INFORMATION
 Mechanistic and functional insights into fatty acid activation in Mycobacterium tuberculosis Pooja Arora 1, Aneesh Goyal 2, Vivek T atarajan 1, Eerappa Rajakumara 2, Priyanka Verma 1, Radhika Gupta 3,
Mechanistic and functional insights into fatty acid activation in Mycobacterium tuberculosis Pooja Arora 1, Aneesh Goyal 2, Vivek T atarajan 1, Eerappa Rajakumara 2, Priyanka Verma 1, Radhika Gupta 3,
Development of RT-qPCR-based molecular diagnostic assays for therapeutic target selection of breast cancer patients
 Development of RT-qPCR-based molecular diagnostic assays for therapeutic target selection of breast cancer patients Sangjung Park The Graduate School Yonsei University Department of Biomedical Laboratory
Development of RT-qPCR-based molecular diagnostic assays for therapeutic target selection of breast cancer patients Sangjung Park The Graduate School Yonsei University Department of Biomedical Laboratory
Figure 1. Effects of FGF21 on adipose tissue. (A) Representative histological. findings of epididymal adipose tissue (B) mrna expression of
 SUPPLEMENTAL MATERIAL EN-12-2276 Figure 1. Effects of FGF21 on adipose tissue. (A) Representative histological findings of epididymal adipose tissue (B) mrna expression of adipocytokines in adipose tissue.
SUPPLEMENTAL MATERIAL EN-12-2276 Figure 1. Effects of FGF21 on adipose tissue. (A) Representative histological findings of epididymal adipose tissue (B) mrna expression of adipocytokines in adipose tissue.
Supplementary Data. Clinical Setup. References. Program for Embryo Donation
 Supplementary Data Clinical Setup Notwithstanding their value in stem cell research, the source of supernumerary human blastocysts for the derivation of new stem cell lines and the modalities of their
Supplementary Data Clinical Setup Notwithstanding their value in stem cell research, the source of supernumerary human blastocysts for the derivation of new stem cell lines and the modalities of their
SUPPLEMENTARY FIG. S2. b-galactosidase staining of
 SUPPLEMENTARY FIG. S1. b-galactosidase staining of senescent cells in 500 mg=dl glucose (10 magnification). SUPPLEMENTARY FIG. S3. Toluidine blue staining of chondrogenic-differentiated adipose-tissue-derived
SUPPLEMENTARY FIG. S1. b-galactosidase staining of senescent cells in 500 mg=dl glucose (10 magnification). SUPPLEMENTARY FIG. S3. Toluidine blue staining of chondrogenic-differentiated adipose-tissue-derived
Characterizing intra-host influenza virus populations to predict emergence
 Characterizing intra-host influenza virus populations to predict emergence June 12, 2012 Forum on Microbial Threats Washington, DC Elodie Ghedin Center for Vaccine Research Dept. Computational & Systems
Characterizing intra-host influenza virus populations to predict emergence June 12, 2012 Forum on Microbial Threats Washington, DC Elodie Ghedin Center for Vaccine Research Dept. Computational & Systems
Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction. Marker Sequence (5 3 ) Accession No.
 Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
RESEARCH COMMUNICATION. Genotype Frequencies of Cyclooxygenease 2 (COX2) Rare Polymorphisms for Japanese with and without Colorectal Cancer
 COX2 Rare Polymorphisms and Colorectal Cancer RESEARCH COMMUNICATION Genotype Frequencies of Cyclooxygenease 2 (COX2) Rare Polymorphisms for Japanese with and without Colorectal Cancer Nobuyuki Hamajima
COX2 Rare Polymorphisms and Colorectal Cancer RESEARCH COMMUNICATION Genotype Frequencies of Cyclooxygenease 2 (COX2) Rare Polymorphisms for Japanese with and without Colorectal Cancer Nobuyuki Hamajima
Grade-dependent effects on cell cycle progression and apoptosis in response to doxorubicin in human bladder cancer cell lines
 INTERNATIONAL JOURNAL OF ONCOLOGY 34: 137-160, 2009 137 Grade-dependent effects on cell cycle progression and apoptosis in response to doxorubicin in human bladder cancer cell lines DIMITRIOS J. STRAVOPODIS
INTERNATIONAL JOURNAL OF ONCOLOGY 34: 137-160, 2009 137 Grade-dependent effects on cell cycle progression and apoptosis in response to doxorubicin in human bladder cancer cell lines DIMITRIOS J. STRAVOPODIS
Supplementary Figure 1: GPCR profiling and G q signaling in murine brown adipocytes (BA). a, Number of GPCRs with 2-fold lower expression in mature
 Supplementary Figure 1: GPCR profiling and G q signaling in murine brown adipocytes (BA). a, Number of GPCRs with 2-fold lower expression in mature BA vs. preadipocytes. b, Number of GPCRs with 2-fold
Supplementary Figure 1: GPCR profiling and G q signaling in murine brown adipocytes (BA). a, Number of GPCRs with 2-fold lower expression in mature BA vs. preadipocytes. b, Number of GPCRs with 2-fold
