The oral microbiome in human immunodeficiency virus (HIV)-positive individuals
|
|
|
- Charles Adams
- 6 years ago
- Views:
Transcription
1 Journal of Medical Microbiology (215), 64, DOI 1.199/jmm..128 The oral microbiome in human immunodeficiency virus (HIV)-positive individuals James O. Kistler, 1 Pratanporn Arirachakaran, 2 Yong Poovorawan, 3 Gunnar Dahlén 4 and William G. Wade 1 Correspondence William G. Wade w.wade@qmul.ac.uk 1 Centre for Immunobiology, Blizard Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, London, UK 2 Infectious Diseases Clinic, Dental Hospital, Faculty of Dentistry, Chulalongkorn University, Bangkok, Thailand 3 Center of Excellence in Clinical Virology, Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand 4 Oral Microbiology and Immunology, Institute of Odontology, Sahlgrenska Academy, University of Gothenburg, Sweden Received 25 March 215 Accepted 7 July 215 Human immunodeficiency virus (HIV) infection is associated with a range of oral conditions, and increased numbers of disease-associated microbial species have previously been found in HIV-positive subjects. The aim of this study was to use next-generation sequencing to compare the composition of the oral microbiome in HIV-positive and -negative individuals. Plaque and saliva were collected from 37 HIV-positive individuals and 37 HIV-negative individuals, and their bacterial composition determined by pyrosequencing of partial 16S rrna genes. A total of sequences were analysed. The number of species-level operational taxonomic units (OTUs) detected was significantly lower in the saliva of HIV-positive individuals (mean533.3) than in that of HIV-negative individuals (mean5365.5) (P,.3). Principal coordinates analysis (PCoA) based on community membership (Jaccard index) and structure (Yue and Clayton measure of dissimilarity) showed significant separation of plaque and saliva samples [analysis of molecular variance (AMOVA), P,.1]. PCoA plots did not show any clear separation based on HIV status. However, AMOVA indicated that there was a significant difference in the community membership of saliva between HIV-positive and -negative groups (P5.1). Linear discriminant analysis effect size revealed an OTU identified as Haemophilus parainfluenzae to be significantly associated with HIV-positive individuals, whilst Streptococcus mitis/hot473 was most significantly associated with HIV-negative individuals. In conclusion, this study has confirmed that the microbial composition of saliva and plaque is different. The oral microbiomes of HIV-positive and -negative individuals were found to be similar overall, although there were minor but significant differences in the composition of the salivary microbiota of the two groups. INTRODUCTION There were an estimated 35 million people infected with the human immunodeficiency virus (HIV) at the end of Abbreviations: AMOVA, analysis of molecular variance; HAART, highly active antiretroviral therapy; HIV, human immunodeficiency virus; LDA, linear discriminant analysis; LEfSe, LDA effect size; OTU, operational taxonomic units; PCoA, principal coordinates analysis The NCBI SRA database accession number for the raw sequence data generated in this study is SRP Four supplementary tables, one supplementary figure and a supplementary dataset are available with the online Supplementary Material. 213 and AIDS-related illnesses were responsible for 1.5 million deaths in a single year (Joint United Nations Programme on HIV/AIDS, 214). Wider and improved access to highly active antiretroviral therapy (HAART) in recent years has led to an increase in life expectancy of HIV-infected individuals (World Health Organization, 213). HIV infection is associated with a number of oral conditions linked to lowered CD4 + T-cell counts. These include opportunistic infections by Candida spp. (candidiasis) and by viruses such as herpes simplex virus, varicella-zoster virus, Epstein Barr virus and human papilloma viruses (Reznik, 25). Oral candidiasis is a G 215 The Authors Printed in Great Britain IP: On: Sat, 3 Dec 217 2:43:55
2 HIV infection and the oral microbiome particularly common problem in HIV-infected individuals, although the introduction of HAART has led to a decline in its incidence (Hodgson et al., 26). Infection with HIV is also associated with the periodontal diseases: linear gingival erythema, nectrotizing ulcerative gingivitis (NUG) and acute necrotizing periodontitis (ANP) (Mataftsi et al., 211). NUG and ANP typically have a sudden onset and can cause severe pain. ANP is characterized by a rapid destruction of periodontal tissue, which may result in tooth loss. Whilst a number of studies have investigated the bacteria associated with these forms of periodontal disease (Aas et al., 27; Murray et al., 1989; Paster et al., 22; Ramos et al., 212; Rams et al., 1991), relatively few studies have compared the normal oral microbiome in HIV-infected and uninfected individuals. One study used the Human Oral Microbe Identification Microarray (HOMIM) to compare the proportions of approximately 3 of the most prevalent oral species on the dorsal surfaces of the tongues of antiretroviral-treated and untreated HIVinfected individuals, and an uninfected control group (Dang et al., 212). The authors detected significantly increased proportions of pathogenic/disease-associated species, including Campylobacter concisus, Campylobacter rectus, Prevotella pallens and Megasphaera micronuciformis, in the untreated HIV-infected group relative to the uninfected control group (Dang et al., 212). Another study using HOMIM, denaturing gradient gel electrophoresis and culture compared the salivary microbiota in HIV-positive and HIV-negative individuals, and monitored the changes in microbial diversity after HAART in the HIVpositive group (Li et al., 214). The authors found that the microbial diversity of saliva was lower in the HIV-positive group than in the HIV-negative group, and that diversity was reduced following HAART treatment. A recent study used an open-ended deep sequencing approach to characterize the composition of both the oral mycobiome and bacteriome in 12 HIV-infected and 12 uninfected individuals (Mukherjee et al., 214). The authors reported no significant differences in the bacterial community composition of oral rinse samples from these groups. However, they revealed differences in the fungal communities (mycobiome). In particular, Pichia species were associated with uninfected individuals and had a negative correlation to the presence of Candida species. The authors also showed that Pichia spp. could inhibit Candida growth in vitro. Whilst the authors reported that the bacteriomes of HIV-infected and uninfected individuals were similar, it is possible that differences in specific habitats such as tooth surfaces were overlooked, as a single oral rinse sample was used for each individual. In addition, the number of participants in each group was relatively low. The aim of this study was to use next-generation sequencing to compare the composition of oral bacterial communities present in plaque and saliva of HIV-positive and -negative individuals. METHODS Study groups. Ethical approval for this study was granted by the Ethics Committee of the Faculty of Dentistry, Chulalongkorn University, Bangkok, Thailand, and written informed consent was obtained for all of the participants. A group of HIV-positive patients was recruited for cross-sectional comparison to a group of HIVnegative subjects (Tables 1 and S1, available in the online Supplementary Material). The groups were matched for age and gender and all of the subjects were non-smokers who had not used antibiotics during the 6 months prior to sampling. All of the individuals were Thai who lived in Bangkok and visited Chulalongkorn University dental clinics for dental care. The subjects did not have any recent history of oral lesions (6 months prior to sampling), oral mucositis, active caries involvement or dental implants. HIV-positive subjects had been infected for at least 1 years prior to sampling and had been undergoing antiretroviral therapy for at least 5 years. The HIV viral load of the HIV-seropositive patients was determined using quantitative real-time PCR (Roche Molecular Systems) and expressed as the number of viral copies ml 21. The number of CD4 + cells of the HIVpositive subjects was determined by flow cytometry with a Becton Dickinson instrument and expressed as cells mm 23. Sample collection. Dental plaque and saliva were collected from both groups. Probing for pocket depth and bleeding was performed after saliva collection, but before plaque collection, during the same visit. Plaque samples were taken using a sterile curette from the buccal surfaces of three teeth (one molar, one premolar and one incisor) in each quadrant by scraping along the tooth in the gingival sulcus and pooled into.5 ml of sterile Tris/EDTA buffer. No sites with deep pockets (w3 mm) were included in the sampling. If a deep pocket was located at a sampling tooth/site, the adjacent tooth was used. There was no evidence of bleeding on probing at the sites used for sampling plaque. Two millilitres of saliva was collected by expectoration into 1 ml sterile tubes. Plaque and saliva samples were stored at 27 uc and processed within 24 h of collection. DNA extraction. DNA was extracted from saliva and plaque using the GenElute Bacterial DNA Extraction kit (Sigma-Aldrich), with an additional lysis step to increase the recovery of Gram-positive bacterial DNA: the samples were incubated with a 45 mg lysozyme ml 21 solution at 37 uc for 3 min. 16S rrna gene PCR and 454 pyrosequencing. The protocol used was as described previously (Kistler et al., 213) with some minor modifications. PCR amplification of a fragment of the 16S rrna gene, approximately 5 bp in length covering the V1 V3 hypervariable regions, was performed using composite fusion primers. The fusion primers comprised the broad-range 16S rrna gene primers 27 Table 1. Summary of the study groups No. of subjects HIV-positive HIV-negative Mean ( SD) age (years) 44 ( 1) 46 ( 1) Male : female 18 : : 2 Mean CD4 + T-cell count ( SD) (cells mm 23 ) 51 ( 232) NA NA, Not applicable. IP: On: Sat, 3 Dec 217 2:43:55
3 J. O. Kistler and others FYM (Frank et al., 28) and 519 R (Lane et al., 1985) along with Roche GS-FLX Titanium Series adaptor sequences (A and B) for 454 pyrosequencing using the Lib-L emulsion-pcr method. Previously described 12-base error-correcting Golay barcode sequences (Fierer et al., 28) were incorporated into the forward primers to enable multiplexing of samples in the same sequencing run. The appropriate barcoded forward primer and reverse primer were used in PCRs along with Extensor High-Fidelity PCR Mastermix (Thermo-Scientific). The PCR conditions were as follows: 5 min initial denaturation step at 95 uc, followed by 25 cycles of 95 uc for 45 s, 53 uc for 45 s and 72 uc for 45 s and a final extension of 72 uc for 5 min. PCR amplicons were then purified using the QIAquick PCR purification kit (Qiagen) according to the manufacturer s instructions. The size and purity of the amplicons were checked using the Agilent DNA 1 kit and the Agilent 21 Bioanalyser. Quantification of the amplicons was performed by means of a fluorometric assay using the Quant-iT Picogreen fluorescent nucleic acid stain (Invitrogen). The amplicons were then pooled together at equimolar concentrations (1 1 9 molecules ml 21 ). Emulsion-PCR and unidirectional sequencing of the samples were performed using the Lib-L kit and the Roche 454 GS- FLX+Titanium series sequencer by the Department of Biochemistry, Cambridge University, Cambridge, UK. Sequence analysis. Sequence analysis was performed using the mothur software suite version 1.33 (Schloss et al., 29), following the 454 standard operating procedure (Schloss & Westcott, 211) on The sequences were denoised using the AmpliconNoise algorithm, as implemented by mothur. Sequences that were less than 44 bases in length and/or had one of the following: w2 mismatches to the primer, w1 mismatch to the barcode regions, and homopolymers of w8 bases in length, were discarded. The remaining sequences were trimmed to remove primers and barcodes and aligned to the SILVA 16S rrna gene reference alignment. The UChime algorithm was used to identify chimeric sequences, which were then removed from the dataset. Sequences were clustered into operational taxonomic units (OTUs) at a genetic distance of.15 using the average neighbour algorithm and identified using a naive Bayesian classifier with the Human Oral Microbiome Database (HOMD) reference set (version 13). A distance of.15 was chosen because many named oral bacterial species have high sequence identity in their 16S rrna genes, particularly among the streptococci, Actinomyces and Neisseria. Therefore, a number of different species are likely to be combined into the same OTU when applying the more commonly used distance of.3. In addition, species-level phylotypes in the HOMD have been defined based on a 98.5 % 16S rrna gene sequence identity cut-off (Dewhirst et al., 21). For those OTUs that could not be identified using the Bayesian classifier, representative sequences were obtained in mothur (the sequence with the smallest distance to all other sequences in that OTU) and identified using BLAST against the HOMD reference set. The possible alternatives for the species identification were then provided. In order to perform OTU-based comparisons of bacterial diversity, the number of sequences in each sample was normalized by subsampling down to the number of sequences equal to that of the sample with the lowest number. The diversity of the communities was estimated and compared in mothur using Simpson s inverse diversity index (Simpson, 1949). Good s non-parametric coverage estimator was used to determine how well sampled the communities were (Good, 1953). The beta-diversity of the samples was analysed using distance matrices generated using the Jaccard index for community membership and the Yue and Clayton measure of dissimilarity (thetayc calculator) for community structure (Yue & Clayton, 25). The distance matrices were visualized using dendrograms and principal coordinates analysis (PCoA) plots. Three-dimensional PCoA plots were generated in R ( using the rgl package. Statistical analysis. Analysis of molecular variance (AMOVA) (Excoffier et al., 1992), as implemented by mothur, was used to determine if there were any statistically significant differences between groups and sample types in the distance matrices based on the Jaccard and thetayc indices. Linear discriminant analysis (LDA) effect size (LEfSe) (Segata et al., 211) was used to identify OTUs that were significantly differentially abundant between saliva and plaque and between HIV-positive and -negative groups. The alpha values for the factorial Kruskal Wallis and pairwise Wilcoxon tests were set to.5 and the LDA score threshold for discriminative features was set to 3.5. Two sample t-tests were performed in R to determine if there were any significant differences in OTU richness and diversity between HIV-positive and -negative groups. Correlations between the relative abundances of OTUs and metadata for the HIV-positive group, including CD4 + cell counts and subject age, were determined using Pearson s rank correlation coefficient as implemented by mothur in the otu.association command; OTUs that were detected in less than 5 % of the saliva or plaque samples were not included in the analysis. RESULTS Pyrosequencing A total of sequences were included in the final analysis after filtering and removal of chimeric sequences. This resulted in a mean of 598 per sample (range: ). Five samples were not included in the pyrosequencing run due to poor PCR amplification. This resulted in a final total of 36 plaque and 36 saliva samples in the HIV-positive group and 35 plaque and 36 saliva samples in the HIV-negative group for comparison. The mean numbers of quality-filtered sequences per sample were 6355 (range: ) and 5599 (range: ) in the HIV-positive and -negative groups respectively. Composition of plaque and saliva samples The sequences were assigned to a total of 12 phyla. The predominant phyla in both plaque and saliva were Actinobacteria, Bacteroidetes, Firmicutes, Fusobacteria and Proteobacteria. Other phyla detected, but not in every sample, included Chloroflexi, GN2, SR1, Spirochaetes, Synergistetes, TM7 and Tenericutes. The relative abundances of the predominant phyla in plaque and saliva are shown in Fig. S1. A total of 168 different genera were detected among all of the samples. The predominant genera in plaque (constituting w5 % mean relative abundance) were Streptococcus, Corynebacterium, Prevotella, Capnocytophaga, Actinomyces, Veillonella and Selenomonas, whilst in saliva the predominant genera were Streptococcus, Prevotella, Veillonella and Rothia. The bacterial composition of all the samples (classified to species level where possible) is shown in Dataset S1. Comparisons of OTU richness and diversity Table S2 shows the OTU richness and diversity (Simpson s inverse diversity index) of each of the plaque and saliva samples included in the analysis (when subsampled to 3583 sequences per sample). The mean number of OTUs 196 IP: Journal of Medical Microbiology 64 On: Sat, 3 Dec 217 2:43:55
4 HIV infection and the oral microbiome detected in the saliva of the HIV-positive individuals was significantly lower than in the HIV-negative individuals (two sample t-test, Pv.3) (Fig. 1a). Two saliva samples from HIV-positive individuals had a particularly reduced richness with 18 and 184 OTUs detected in each sample. Interestingly, the same individuals had the lowest richness in their plaque samples (141 and 177 OTUs). There was, however, no significant difference (P5.58) between the mean number of OTUs detected in the plaque samples of the HIV-positive and -negative groups (Fig. 1a). In addition, there were no significant differences in the Simpson s inverse diversity index of saliva or plaque samples of the two groups. The mean (a) Number of OTUs (b) Simpson's inverse diversity index Plaque_HIV_neg Plaque_HIV_pos Saliva_HIV_neg Saliva_HIV_pos Plaque_HIV_neg Plaque_HIV_pos Saliva_HIV_neg Saliva_HIV_pos Fig. 1. Bar charts comparing the mean species-level OTU richness (a) and Simpson s inverse diversity index (b) in plaque and saliva samples of the HIV-positive and -negative groups. The saliva of HIV-positive individuals had a significantly lower richness than the saliva of HIV-negative individuals (indicated by the asterisk) using a two-sample t-test. There was no significant difference in Simpson s diversity index between groups. The error bars indicate SEM. * diversity of plaque samples was higher than that of saliva samples (Fig. 1b). Comparisons of bacterial community membership and structure The community membership and structure of plaque and saliva samples from HIV-positive and -negative groups was compared using PCoA plots. Comparisons of samples from both HIV-positive and negative groups in PCoA plots revealed clear separation of plaque and saliva (Fig. 2). AMOVA tests confirmed that saliva and plaque were highly significantly different (Pv.1 for both membership and structure). The plots did not show a clear separation/difference between the HIV-positive and -negative groups for either the plaque or saliva samples (Fig. 3). AMOVA confirmed that there was no significant difference in community structure between the groups. However, AMOVA indicated that there was a significant difference between the community membership of saliva, but not plaque, of HIV-negative and -positive subjects (P5.1) (Fig. 4). Comparison using LEfSe revealed 121 OTUs that were significantly differentially abundant between saliva and plaque samples. The differentially abundant OTUs with LDA scores of i3.5 are shown in Fig. 5. The OTU most significantly associated with saliva was identified as Streptococcus salivarius/streptococcus vestibularis, whilst a Corynebacterium matruchotii OTU was most significantly associated with plaque. Comparison of the saliva of HIV-positive subjects to that of HIV-negative subjects revealed 116 OTUs that were significantly differentially abundant. However, only three OTUs had LDA scores of i3.5 and many of the associated OTUs were detected at low relative abundances. Of the OTUs with LDA scores of i3.5, a Haemophilus parainfluenzae OTU was associated with HIVpositive individuals, whilst a Streptococcus mitis/streptococcus pneumoniae/hot473 OTU and a Streptococcus mitis bv.2 OTU were associated with HIV-negative individuals. LEfSe comparison of plaque from HIV-positive and -negative individuals was not performed, as there was no significant difference in community membership or structure by AMOVA. Correlation of OTUs with CD4 1 cell counts and subject age The OTUs that showed a statistically significant correlation (Pearson s rank correlation coefficient) with CD4 + T-cell counts and with subject age in the HIV-positive group are shown in Tables S3 and S4. For both plaque and saliva samples, stronger correlations were obtained with age than with the CD4 + cell counts. The OTUs that showed the strongest negative correlations with CD4 + cell counts were OTU 3 Alloprevotella tannerae (2.42, Pv.9) and OTU 373 Eubacterium yurii (2.45, Pv.4) in saliva and plaque, respectively. The OTUs IP: On: Sat, 3 Dec 217 2:43:55
5 J. O. Kistler and others (a) (b) Fig. 2. PCoA plots comparing plaque and saliva from HIV-positive and -negative individuals based on their community membership (Jaccard index) (a) and structure (thetayc calculator) (b). Plaque samples are shown in red and saliva samples are in blue. Plaque and saliva had a highly significantly different community membership and structure by AMOVA (P,.1 for both). The proportion of variance shown for (a) and (b), respectively, was: 56.1 % and 18.4 %, 52.4 % and 7.3 %, 51.5 % and 5.5 %. (a) (b) Fig. 3. PCoA plots comparing plaque and saliva from HIV-positive and -negative individuals based on their community membership (Jaccard index) (a) and structure (thetayc calculator) (b). Red, saliva from HIV-positive subjects; blue, saliva from HIV-negative subjects; purple, plaque from HIV-positive subjects; green, plaque from HIV-negative subjects. There was a significant difference between the community membership, but not structure, of saliva between groups using AMOVA (P5.1). No other significant differences were detected. The proportion of variance shown for (a) and (b), respectively, was: 56.1 % and 18.4 %, 52.4 % and 7.3 %, 51.5 % and 5.5 %. 198 Journal of Medical Microbiology 64 IP: On: Sat, 3 Dec 217 2:43:55
6 HIV infection and the oral microbiome Fig. 4. PCoA plot comparing saliva from HIV-positive and -negative individuals based on their community membership (Jaccard index). Red, saliva from HIV-positive subjects; blue, saliva from HIV-negative subjects. There was a significant difference in community membership between the two groups by AMOVA (P5.1). The proportion of variance shown is: 56.1 %, 52.4 %, 51.5 %. most strongly positively correlated with CD4 + T-cell counts were OTU 623 H. parainfluenzae (.46, Pv.4) and OTU 269 Actinomyces sp. (.5, Pv.2) in saliva and plaque, respectively. DISCUSSION This study has comprehensively characterized the bacterial composition of dental plaque and saliva of HIV-positive individuals, at the species level, and compared it with that of HIV-negative individuals. The results confirmed the findings of previous studies, which showed that saliva and plaque have a different bacterial composition (Segata et al., 212; Simón-Soro et al., 213). In addition, the results showed that plaque had a higher bacterial diversity (Simpson s inverse diversity index) than saliva. Species that were most strongly associated with saliva included salivarius- and mitis-group streptococci, Rothia mucilaginosa and Prevotella melaninogenica whilst species most strongly associated with plaque included Corynebacterium matruchotii, Streptococcus sp. HOT58 and Capnocytophaga granulosa. The hard, non-shedding tooth surfaces on which plaque forms provide a distinct microbial habitat to the shedding mucosal surfaces of the mouth and is thus associated with a differing microbial community structure (Mager et al., 23; Segata et al., 212). Using chequerboard DNA DNA hybridization, Mager et al. (23) showed that the distribution of 4 selected bacterial species in saliva was most similar to that found on the tongue surfaces, suggesting that saliva predominantly comprises species that are shed from tongue biofilms. In that study, saliva and tongue communities were characterized by increased numbers of Prevotella melaninogenica and Veillonella parvula relative to other oral sites/samples. Nextgeneration sequencing data from the Human Microbiome Project also showed that saliva samples were highly similar in composition to samples from the tongue dorsum (Segata et al., 212). Supra- and subgingival plaque had a distinct composition from samples from other oral sites and was characterized by having significantly higher proportions of members of the orders Burkholderiales, Cardiobacteriales, Flavobacteriales, Fusobacteriales, Neisseriales and Pasteurellales. However, the sequences were not classified to the species level. Comparisons of the community composition of plaque and saliva of HIV-positive and HIV-negative individuals in PCoA plots showed that their bacterial communities were largely similar, supporting the findings of a recent deepsequencing study (Mukherjee et al., 214). The plaque samples in the present study were, however, only taken from periodontally healthy sites in both groups. It is therefore possible that there are significant differences in plaque composition between HIV-infected and uninfected individuals at inflamed/diseased sites. Differences in OTU community membership between groups were detected in saliva by AMOVA and a number of species-level OTUs were identified as being significantly differentially abundant. In addition, the saliva of the HIV-positive individuals had a significantly lower OTU richness than that of the HIV-negative individuals. This is in agreement with the findings of a previous study that compared the salivary microbiota of HIV-positive and -negative individuals using culture and other culture-independent methods (Li et al., 214). The authors reported a decreased microbial diversity (Shannon diversity index) in the saliva of HIVpositive subjects compared with that of the negative control group and also showed that the microbial diversity within the HIV-positive group was reduced following HAART. There is evidence that HAART can reduce salivary flow (Navazesh et al., 29) and this in turn might affect the richness and composition of the salivary microbiota. A previous study has shown that individuals with reduced salivary secretion have increased numbers of lactobacilli, which are caries-associated, in their saliva compared with a control group (Almståhl & Wikström, 1999). All of the HIV-positive patients in the present study were undergoing HAART. The importance of the OTUs that were associated with HIV-positive individuals is unclear as they were found to be relatively common oral commensal species. An H. parainfluenzae OTU was most significantly associated with the HIV-positive group. This commensal species is a member of the HACEK group of Gram-negative bacilli that have been implicated in various opportunistic IP: On: Sat, 3 Dec 217 2:43:55
7 J. O. Kistler and others OTU2_Streptococcus_salivarius_vestibularis OTU3_Streptococcus_mitis_pneumoniae_HOT423_HOT64 OTU7_Rothia_mucilaginosa OTU9_Streptococcus_infantis_HOT65 OTU4_Prevotella_melaninogenica OTU1_Haemophilus_parainfluenzae OTU16_Gemella_sanguinis OTU19_Streptococcus_mitis_HOT431 OTU17_Streptococcus_parasanguinis OTU13_Neisseria_flavescens_subflava OTU15_Granulicatella_adiacens OTU35_Prevotella_histicola OTU41_Prevotella_sp_HOT36 OTU22_Gemella_morbillorum_haemolysans OTU44_Actinomyces_odontolyticus_HOT18 OTU47_Streptococcus_mitis_bv2 OTU34_Prevotella_pallens OTU3_Alloprevotella_tannerae OTU52_Bacteroidales_G2_sp_HOT274 OTU42_Ottowia_sp_HOT894 OTU28_Cardiobacterium_hominis OTU36_Cardiobacterium_valvulum OTU31_Actinomyces_sp_HOT448 OTU27_Capnocytophaga_leadbetteri OTU26_Campylobacter_gracilis OTU24_Dialister_invisus OTU25_Tannerella_sp_HOT286 OTU12_Streptococcus_cristatus OTU21_Capnocytophaga_granulosa OTU11_Streptococcus_sp_HOT58 OTU6_Corynebacterium_matruchotii OTU5_Corynebacterium_matruchotii LDA score (log 1 ) Plaque Saliva Fig. 5. Significantly differentially abundant OTUs between saliva and plaque using LEfSe. A total of 121 OTUs were significantly differentially abundant between saliva and plaque. The 32 OTUs shown are those with effect sizes of 3.5. infections including endocardial infections, upper respiratory tract infections, brain abscesses and joint infections (Janda, 213). Whilst H. parainfluenzae was associated with the HIVpositive individuals, within the HIV-positive group it was positively correlated with CD4 + cell counts. This suggests that H. parainfluenzae does not increase in proportion as the disease progresses and CD4 + cell numbers decline. In conclusion, this study found that the oral microbiomes of HIV-positive and -negative individuals were similar overall, although minor but significant differences in the composition of the salivary microbiota were detected. The results also confirmed that plaque and saliva have a distinct microbial composition. ACKNOWLEDGEMENTS This study was supported by the Higher Education Research Promotion and National Research University Project of Thailand, Office of the Higher Education Commission (HR1155A). This research utilized Queen Mary s MidPlus computational facilities, supported by QMUL Research-IT and funded by EPSRC grant EP/K128/1. REFERENCES Aas, J. A., Barbuto, S. M., Alpagot, T., Olsen, I., Dewhirst, F. E. & Paster, B. J. (27). Subgingival plaque microbiota in HIV positive patients. J Clin Periodontol 34, IP: Journal of Medical Microbiology 64 On: Sat, 3 Dec 217 2:43:55
8 HIV infection and the oral microbiome Almståhl, A. & Wikström, M. (1999). Oral microflora in subjects with reduced salivary secretion. J Dent Res 78, Dang, A. T., Cotton, S., Sankaran-Walters, S., Li, C.-S., Lee, C.-Y. M., Dandekar, S., Paster, B. J. & George, M. D. (212). Evidence of an increased pathogenic footprint in the lingual microbiome of untreated HIV infected patients. BMC Microbiol 12, 153. Dewhirst, F. E., Chen, T., Izard, J., Paster, B. J., Tanner, A. C., Yu, W. H., Lakshmanan, A. & Wade, W. G. (21). The human oral microbiome. J Bacteriol 192, Excoffier, L., Smouse, P. E. & Quattro, J. M. (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131, Fierer, N., Hamady, M., Lauber, C. L. & Knight, R. (28). The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci U S A 15, Frank, J. A., Reich, C. I., Sharma, S., Weisbaum, J. S., Wilson, B. A. & Olsen, G. J. (28). Critical evaluation of two primers commonly used for amplification of bacterial 16S rrna genes. Appl Environ Microbiol 74, Good, I. J. (1953). The population frequencies of species and the estimation of population parameters. Biometrika 4, Hodgson, T. A., Greenspan, D. & Greenspan, J. S. (26). Oral lesions of HIV disease and HAART in industrialized countries. Adv Dent Res 19, Janda, W. M. (213). Update on the HACEK group of fastidious gramnegative bacilli, Part I. Clin Microbiol Newsl 35, Joint United Nations Programme on HIV/AIDS (UNAIDS) (214). The Gap Report. Geneva: UNAIDS. Kistler, J. O., Booth, V., Bradshaw, D. J. & Wade, W. G. (213). Bacterial community development in experimental gingivitis. PLoS One 8, e Lane, D. J., Pace, B., Olsen, G. J., Stahl, D. A., Sogin, M. L. & Pace, N. R. (1985). Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci U S A 82, Li, Y., Saxena, D., Chen, Z., Liu, G., Abrams, W. R., Phelan, J. A., Norman, R. G., Fisch, G. S., Corby, P. M. & other authors (214). HIV infection and microbial diversity in saliva. J Clin Microbiol 52, Mager, D. L., Ximenez-Fyvie, L. A., Haffajee, A. D. & Socransky, S. S. (23). Distribution of selected bacterial species on intraoral surfaces. J Clin Periodontol 3, Mataftsi, M., Skoura, L. & Sakellari, D. (211). HIV infection and periodontal diseases: an overview of the post-haart era. Oral Dis 17, Mukherjee, P. K., Chandra, J., Retuerto, M., Sikaroodi, M., Brown, R. E., Jurevic, R., Salata, R. A., Lederman, M. M., Gillevet, P. M. & Ghannoum, M. A. (214). Oral mycobiome analysis of HIV-infected patients: identification of Pichia as an antagonist of opportunistic fungi. PLoS Pathog 1, e Murray, P. A., Grassi, M. & Winkler, J. R. (1989). The microbiology of HIV-associated periodontal lesions. J Clin Periodontol 16, Navazesh, M., Mulligan, R., Karim, R., Mack, W. J., Ram, S., Seirawan, H., Greenspan, J., Greenspan, D., Phelan, J., Alves, M. & Oral Substudy of the Women s Interagency HIV Study Collaborative Study Group* (29). Effect of HAART on salivary gland function in the Women s Interagency HIV Study (WIHS). Oral Dis 15, Paster, B. J., Russell, M. K., Alpagot, T., Lee, A. M., Boches, S. K., Galvin, J. L. & Dewhirst, F. E. (22). Bacterial diversity in necrotizing ulcerative periodontitis in HIV-positive subjects. Ann Periodontol 7, Ramos, M. P., de A., Ferreira, S. M. S., Silva-Boghossian, C. M., Souto, R., Colombo, A. P., Noce, C. W. & Gonçalves, L. S. (212). Necrotizing periodontal diseases in HIV-infected Brazilian patients: a clinical and microbiologic descriptive study. Quintessence Int 43, Rams, T. E., Andriolo, M. Jr, Feik, D., Abel, S. N., McGivern, T. M. & Slots, J. (1991). Microbiological study of HIV-related periodontitis. J Periodontol 62, Reznik, D. A. (25). Oral manifestations of HIV disease. Top HIV Med 13, Schloss, P. D. & Westcott, S. L. (211). Assessing and improving methods used in operational taxonomic unit-based approaches for 16S rrna gene sequence analysis. Appl Environ Microbiol 77, Schloss, P. D., Westcott, S. L., Ryabin, T., Hall, J. R., Hartmann, M., Hollister, E. B., Lesniewski, R. A., Oakley, B. B., Parks, D. H. & other authors (29). Introducing mothur: open-source, platformindependent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75, Segata, N., Izard, J., Waldron, L., Gevers, D., Miropolsky, L., Garrett, W. S. & Huttenhower, C. (211). Metagenomic biomarker discovery and explanation. Genome Biol 12, R6. Segata, N., Haake, S. K., Mannon, P., Lemon, K. P., Waldron, L., Gevers, D., Huttenhower, C. & Izard, J. (212). Composition of the adult digestive tract bacterial microbiome based on seven mouth surfaces, tonsils, throat and stool samples. Genome Biol 13, R42. Simpson, E. H. (1949). Measurement of diversity. Nature 163, 688. Simón-Soro, A., Tomás, I., Cabrera-Rubio, R., Catalan, M. D., Nyvad, B. & Mira, A. (213). Microbial geography of the oral cavity. J Dent Res 92, World Health Organization (213). Global Update on HIV Treatment 213: Results, Impact and Opportunities. Geneva: World Health Organization. Yue, J. C. & Clayton, M. K. (25). A similarity measure based on species proportions. Commun Stat Theory Methods 34, IP: On: Sat, 3 Dec 217 2:43:55
Bacterial Community Development in Experimental Gingivitis
 Bacterial Community Development in Experimental Gingivitis James O. Kistler 1, Veronica Booth 2, David J. Bradshaw 3, William G. Wade 1 * 1 Microbiology Unit, King s College London Dental Institute, London,
Bacterial Community Development in Experimental Gingivitis James O. Kistler 1, Veronica Booth 2, David J. Bradshaw 3, William G. Wade 1 * 1 Microbiology Unit, King s College London Dental Institute, London,
Microbiota Transplantation Workshop: Oral Cavity
 Microbiota Transplantation Workshop: Oral Cavity December 3, 2015 Floyd E. Dewhirst, DDS, PhD Department of Microbiology The Forsyth Institute Outline Microbiota of the oral cavity Diseases of oral dysbiosis
Microbiota Transplantation Workshop: Oral Cavity December 3, 2015 Floyd E. Dewhirst, DDS, PhD Department of Microbiology The Forsyth Institute Outline Microbiota of the oral cavity Diseases of oral dysbiosis
The oral microbiome and human health
 201 Journal of Oral Science, Vol. 59, No. 2, 201-206, 2017 Review The oral microbiome and human health Yoshihisa Yamashita 1) and Toru Takeshita 1,2) 1) Section of Preventive and Public Health Dentistry,
201 Journal of Oral Science, Vol. 59, No. 2, 201-206, 2017 Review The oral microbiome and human health Yoshihisa Yamashita 1) and Toru Takeshita 1,2) 1) Section of Preventive and Public Health Dentistry,
Effect of maltitol-containing chewing gum use on the composition of dental plaque microbiota in subjects with active dental caries
 Journal of Oral Microbiology ISSN: (Print) 2000-2297 (Online) Journal homepage: http://www.tandfonline.com/loi/zjom20 Effect of maltitol-containing chewing gum use on the composition of dental plaque microbiota
Journal of Oral Microbiology ISSN: (Print) 2000-2297 (Online) Journal homepage: http://www.tandfonline.com/loi/zjom20 Effect of maltitol-containing chewing gum use on the composition of dental plaque microbiota
Index. Note: Page numbers of article titles are in boldface type.
 Note: Page numbers of article titles are in boldface type. A Actinobacteria, 307 308 Actinomycetes, 307 308 Actinomycosis, in oral cavity, 288 290 Aggregatibacter, 308 309 Amoxicillin, in odontogenic infections,
Note: Page numbers of article titles are in boldface type. A Actinobacteria, 307 308 Actinomycetes, 307 308 Actinomycosis, in oral cavity, 288 290 Aggregatibacter, 308 309 Amoxicillin, in odontogenic infections,
Materials and Methods: Literature review and Authors opinion.
 Haffajee AD, Bogren A, Hasturk H et al. Subgingival microbiota of chronic periodontitis subjects from different geographic locations. J Clin Periodontol 2004; 31:996-1002. Purpose: To compare the subgingival
Haffajee AD, Bogren A, Hasturk H et al. Subgingival microbiota of chronic periodontitis subjects from different geographic locations. J Clin Periodontol 2004; 31:996-1002. Purpose: To compare the subgingival
Imbalances in the Oral Microbiota associated with Oral Mucosal Diseases
 2nd Microbiome R&D and Business Collaboration Congress: Asia 216. 2. 29 Imbalances in the Oral Microbiota associated with Oral Mucosal Diseases Youngnim Choi, D.D.S., Ph.D. C Huttenhower et al. Nature
2nd Microbiome R&D and Business Collaboration Congress: Asia 216. 2. 29 Imbalances in the Oral Microbiota associated with Oral Mucosal Diseases Youngnim Choi, D.D.S., Ph.D. C Huttenhower et al. Nature
Supplementary Figures. Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies.
 Supplementary Figures Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies. Supplementary Figure 2. SIV replication and CD4 + T cell count. (A) Log SIVmac239 copies/ml
Supplementary Figures Supplementary Figure 1. Treatment schematic of SIV infection and ARV and PP therapies. Supplementary Figure 2. SIV replication and CD4 + T cell count. (A) Log SIVmac239 copies/ml
Oral Health Applications for Probiotics
 Oral Health Applications for Probiotics Andrew McBain Biofilm Research Group Manchester Pharmacy School University of Manchester Overview Oral microbiology introduction Dental probiotics and replacement
Oral Health Applications for Probiotics Andrew McBain Biofilm Research Group Manchester Pharmacy School University of Manchester Overview Oral microbiology introduction Dental probiotics and replacement
Oral microbiota in relation to HIV infection and oral infection in perinatally HIV-infected adolescents
 Oral microbiota in relation to HIV infection and oral infection in perinatally HIV-infected adolescents Bruce J. Paster Jacqueline R. Starr Kyu Ha Lee Lina Faller Barbara Moscicki Russell Van Dyke Mark
Oral microbiota in relation to HIV infection and oral infection in perinatally HIV-infected adolescents Bruce J. Paster Jacqueline R. Starr Kyu Ha Lee Lina Faller Barbara Moscicki Russell Van Dyke Mark
Cariogenic microflora and ph in superficial and deep layers of occlusal carious lesions - A metagenomic analysis
 Journal of Advanced Clinical & Research Insights (2017), 4, 124 128 ORIGINAL ARTICLE Cariogenic microflora and ph in superficial and deep layers of occlusal carious lesions - A metagenomic analysis S.
Journal of Advanced Clinical & Research Insights (2017), 4, 124 128 ORIGINAL ARTICLE Cariogenic microflora and ph in superficial and deep layers of occlusal carious lesions - A metagenomic analysis S.
Review Article Importance of Diversity in the Oral Microbiota including Candida Species Revealed by High-Throughput Technologies
 International Dentistry, Article ID 454391, 5 pages http://dx.doi.org/10.1155/2014/454391 Review Article Importance of Diversity in the Oral Microbiota including Candida Species Revealed by High-Throughput
International Dentistry, Article ID 454391, 5 pages http://dx.doi.org/10.1155/2014/454391 Review Article Importance of Diversity in the Oral Microbiota including Candida Species Revealed by High-Throughput
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12198 1. Supplementary results 1.1. Associations between gut microbiota, glucose control and medication Women with T2D who used metformin had increased levels of Enterobacteriaceae (i.e.
doi:10.1038/nature12198 1. Supplementary results 1.1. Associations between gut microbiota, glucose control and medication Women with T2D who used metformin had increased levels of Enterobacteriaceae (i.e.
Multi-view learning and deep learning for heterogeneous biological data to maintain oral health Imangaliyev, Sultan
 UvA-DARE (Digital Academic Repository) Multi-view learning and deep learning for heterogeneous biological data to maintain oral health Imangaliyev, Sultan Link to publication Citation for published version
UvA-DARE (Digital Academic Repository) Multi-view learning and deep learning for heterogeneous biological data to maintain oral health Imangaliyev, Sultan Link to publication Citation for published version
Egija Zaura Professor in Oral Microbial Ecology Department of Preventive Dentistry Academic Centre for Dentistry Amsterdam (ACTA)
 Egija Zaura Professor in Oral Microbial Ecology Department of Preventive Dentistry Academic Centre for Dentistry Amsterdam (ACTA) Women & their Microbes, June 2, 2017, Amsterdam Joined Faculty of Dentistry
Egija Zaura Professor in Oral Microbial Ecology Department of Preventive Dentistry Academic Centre for Dentistry Amsterdam (ACTA) Women & their Microbes, June 2, 2017, Amsterdam Joined Faculty of Dentistry
Cell counting (EtBr) Before cell-lysis. Cell-lysis by 3% SDS beads beating. After cell-lysis
 Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
The Gut Microbiota: Evidence For Gut Microbes as Contributors to Weight Gain
 The Gut Microbiota: Evidence For Gut Microbes as Contributors to Weight Gain Michael T. Bailey, Ph.D. Center for Microbial Pathogenesis The Research Institute, Nationwide Children s Hospital Department
The Gut Microbiota: Evidence For Gut Microbes as Contributors to Weight Gain Michael T. Bailey, Ph.D. Center for Microbial Pathogenesis The Research Institute, Nationwide Children s Hospital Department
Microbial markers of oral carcinogenesis in Fanconi Anemia and cancer patients Flavia Teles, DDS, MS, DMSc
 Microbial markers of oral carcinogenesis in Fanconi Anemia and cancer patients Flavia Teles, DDS, MS, DMSc The Forsyth Institute, Boston, MA Harvard School of Dental Medicine Introduction Infection as
Microbial markers of oral carcinogenesis in Fanconi Anemia and cancer patients Flavia Teles, DDS, MS, DMSc The Forsyth Institute, Boston, MA Harvard School of Dental Medicine Introduction Infection as
Bacterial composition in the supragingival plaques of children with and without dental caries
 182 Ro kiewicz D, et al. Advances in Medical Sciences Vol. 51 2006 Suppl. 1 Bacterial composition in the supragingival plaques of children with and without dental caries Ro kiewicz D 1, Daniluk T 2, Zaremba
182 Ro kiewicz D, et al. Advances in Medical Sciences Vol. 51 2006 Suppl. 1 Bacterial composition in the supragingival plaques of children with and without dental caries Ro kiewicz D 1, Daniluk T 2, Zaremba
Interactions between the host genome and environmental factors,
 ORIGINAL ARTICLE Alterations of the Subgingival Microbiota in Pediatric Crohn s Disease Studied Longitudinally in Discovery and Validation Cohorts Judith Kelsen, MD,* Kyle Bittinger, PhD, Helen Pauly-Hubbard,
ORIGINAL ARTICLE Alterations of the Subgingival Microbiota in Pediatric Crohn s Disease Studied Longitudinally in Discovery and Validation Cohorts Judith Kelsen, MD,* Kyle Bittinger, PhD, Helen Pauly-Hubbard,
Evidence of an increased pathogenic footprint in the lingual microbiome of untreated HIV infected patients
 Dang et al. BMC Microbiology 2012, 12:153 RESEARCH ARTICLE Open Access Evidence of an increased pathogenic footprint in the lingual microbiome of untreated HIV infected patients Angeline T Dang 1,4, Sean
Dang et al. BMC Microbiology 2012, 12:153 RESEARCH ARTICLE Open Access Evidence of an increased pathogenic footprint in the lingual microbiome of untreated HIV infected patients Angeline T Dang 1,4, Sean
Defining the Normal Bacterial Flora of the Oral Cavity
 JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2005, p. 5721 5732 Vol. 43, No. 11 0095-1137/05/$08.00 0 doi:10.1128/jcm.43.11.5721 5732.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2005, p. 5721 5732 Vol. 43, No. 11 0095-1137/05/$08.00 0 doi:10.1128/jcm.43.11.5721 5732.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
FACULTY OF HEALTH SCIENCES INSTITUTE OF CLINICAL DENTISTRY
 Professor Gottfried Schmalz Editor-in-Chief Clinical Oral Investigations Universität Regensburg Fakultät für Medizin Poliklinik für Zahnerhatungskunde und Parodontologie Frans-Josef-Strauss-Allee 11 93053
Professor Gottfried Schmalz Editor-in-Chief Clinical Oral Investigations Universität Regensburg Fakultät für Medizin Poliklinik für Zahnerhatungskunde und Parodontologie Frans-Josef-Strauss-Allee 11 93053
Periodontal Microbiology (Dental Plaque)
 Periodontal Microbiology (Dental Plaque) Dental plaque Dental plaque is soft deposits that form the biofilm adhering to the tooth surface of other hard surface in the oral cavity, including removable and
Periodontal Microbiology (Dental Plaque) Dental plaque Dental plaque is soft deposits that form the biofilm adhering to the tooth surface of other hard surface in the oral cavity, including removable and
Analysis of the effect of probiotics on shaping human gut microbiota
 Analysis of the effect of probiotics on shaping human gut microbiota Masahira HATTORI Center for Omics and Bioinformatics, The University of Tokyo http://www.cb.k.u-tokyo.ac.jp/hattorilab
Analysis of the effect of probiotics on shaping human gut microbiota Masahira HATTORI Center for Omics and Bioinformatics, The University of Tokyo http://www.cb.k.u-tokyo.ac.jp/hattorilab
Evidence Based Management of Dental Caries
 Evidence Based Management of Dental Caries Understanding the etiology of dental caries Dr. Wenyuan Shi Professor and Chair, Oral Biology, UCLA SOD Professor, Microbiology & Immunology, UCLA SOM Disruptive
Evidence Based Management of Dental Caries Understanding the etiology of dental caries Dr. Wenyuan Shi Professor and Chair, Oral Biology, UCLA SOD Professor, Microbiology & Immunology, UCLA SOM Disruptive
Research and Reviews: Journal of Dental Sciences
 Research and Reviews: Journal of Dental Sciences A Review on the Human Oral Microflora Sowmya Y * Department of Microbiology, Andhra University, Visakhapatnam, Andhra Pradesh, India Review Article Received:
Research and Reviews: Journal of Dental Sciences A Review on the Human Oral Microflora Sowmya Y * Department of Microbiology, Andhra University, Visakhapatnam, Andhra Pradesh, India Review Article Received:
Dental Research Journal
 Dental Research Journal Original Article Microflora and periodontal disease Luca Scapoli 1, Ambra Girardi 1, Annalisa Palmieri 2, Tiziano Testori 3, Francesco Zuffetti 3, Riccardo Monguzzi 4, Dorina Lauritano
Dental Research Journal Original Article Microflora and periodontal disease Luca Scapoli 1, Ambra Girardi 1, Annalisa Palmieri 2, Tiziano Testori 3, Francesco Zuffetti 3, Riccardo Monguzzi 4, Dorina Lauritano
actinomycetemcornitans and
 Periodontology 2000, Vol. 20, 1999, 341-362 Printed in Denmark. All rights reserved Copyright 8 Munksgaard 1999 PERIODONTOLOGY 2000 ISSN 0906-6713 Ecological considerations in the treatment of Actinobacillus
Periodontology 2000, Vol. 20, 1999, 341-362 Printed in Denmark. All rights reserved Copyright 8 Munksgaard 1999 PERIODONTOLOGY 2000 ISSN 0906-6713 Ecological considerations in the treatment of Actinobacillus
Detection of Herpes Simplex Virus in Chronic Generalized Periodontitis via Polymerase Chain Reaction: A Pilot Study
 Journal of Periodontology & Implant Dentistry Research Article Detection of Herpes Simplex Virus in Chronic Generalized Periodontitis via Polymerase Chain Reaction: A Pilot Study Thomas George Veliyaveetil
Journal of Periodontology & Implant Dentistry Research Article Detection of Herpes Simplex Virus in Chronic Generalized Periodontitis via Polymerase Chain Reaction: A Pilot Study Thomas George Veliyaveetil
Microbiology - Problem Drill 21: Microbial Diseases of the Digestive System
 Microbiology - Problem Drill 21: Microbial Diseases of the Digestive System No. 1 of 10 1. Which of the following organs is not part of the gastrointestinal tract (GI)? (A) Esophagus (B) Small intestine
Microbiology - Problem Drill 21: Microbial Diseases of the Digestive System No. 1 of 10 1. Which of the following organs is not part of the gastrointestinal tract (GI)? (A) Esophagus (B) Small intestine
collected for biochemical and molecular microbiological analyses at baseline, week 4 and
 SUPPLEMENTARY FIGURE LEGENDS Figure S1 Study design. Figure was adapted from Paramsothy et al. 5 Blood and stool samples were collected for biochemical and molecular microbiological analyses at baseline,
SUPPLEMENTARY FIGURE LEGENDS Figure S1 Study design. Figure was adapted from Paramsothy et al. 5 Blood and stool samples were collected for biochemical and molecular microbiological analyses at baseline,
Comparison of the Respiratory Microbiome in Healthy Nonsmokers and Smokers
 Comparison of the Respiratory Microbiome in Healthy Nonsmokers and Smokers Weiguang (Wayne) Mao Morris, A., Beck, J. M., Schloss, P. D., Campbell, T. B., Crothers, K., Curtis, J. L.,... & Weinstock, G.
Comparison of the Respiratory Microbiome in Healthy Nonsmokers and Smokers Weiguang (Wayne) Mao Morris, A., Beck, J. M., Schloss, P. D., Campbell, T. B., Crothers, K., Curtis, J. L.,... & Weinstock, G.
Peri-implantitis Review
 1 2 Aro n J. Saf f er D DS M S Di pl omate o f th e Ame ric an Bo ard of P eri od o n tol og y A quarterly review of the latest publications related to the study of Peri-implant inflammation and bone loss
1 2 Aro n J. Saf f er D DS M S Di pl omate o f th e Ame ric an Bo ard of P eri od o n tol og y A quarterly review of the latest publications related to the study of Peri-implant inflammation and bone loss
Role of microbial communities in the pathogenesis of periodontal diseases and caries
 J Clin Periodontol 2017; 44 (Suppl.18): S23 S38 doi: 10.1111/jcpe.12671 Role of microbial communities in the pathogenesis of periodontal diseases and caries Alex Mira 1, A. Simon-Soro 1 and M. A. Curtis
J Clin Periodontol 2017; 44 (Suppl.18): S23 S38 doi: 10.1111/jcpe.12671 Role of microbial communities in the pathogenesis of periodontal diseases and caries Alex Mira 1, A. Simon-Soro 1 and M. A. Curtis
Alterations of oral microbiota distinguish children with autism spectrum disorders from healthy controls
 www.nature.com/scientificreports Received: 23 February 2017 Accepted: 10 January 2018 Published: xx xx xxxx OPEN Alterations of oral microbiota distinguish children with autism spectrum disorders from
www.nature.com/scientificreports Received: 23 February 2017 Accepted: 10 January 2018 Published: xx xx xxxx OPEN Alterations of oral microbiota distinguish children with autism spectrum disorders from
PREVALENCE OF CYTOMEGALOVIRUS, HUMAN HERPESVIRUS-6, AND EPSTEIN-BARR VIRUS IN PERIODONTITIS PATIENTS AND HEALTHY SUBJECTS IN THE THAI POPULATION
 CMV, HHV-6, AND EBV-1 IN PERIODONTITIS AND HEALTHY PATIENTS PREVALENCE OF CYTOMEGALOVIRUS, HUMAN HERPESVIRUS-6, AND EPSTEIN-BARR VIRUS IN PERIODONTITIS PATIENTS AND HEALTHY SUBJECTS IN THE THAI POPULATION
CMV, HHV-6, AND EBV-1 IN PERIODONTITIS AND HEALTHY PATIENTS PREVALENCE OF CYTOMEGALOVIRUS, HUMAN HERPESVIRUS-6, AND EPSTEIN-BARR VIRUS IN PERIODONTITIS PATIENTS AND HEALTHY SUBJECTS IN THE THAI POPULATION
Microbiota and Oral Disease Prof. Dennis Cvitkovitch
 1 Professor Dennis Cvitkovitch Faculty of Dentistry Dental Research Institute University of Toronto The human microbiome We are a composite species: eukaryotic, bacterial, archeal Every human harbors over
1 Professor Dennis Cvitkovitch Faculty of Dentistry Dental Research Institute University of Toronto The human microbiome We are a composite species: eukaryotic, bacterial, archeal Every human harbors over
Analysis of Run-to-Run Variation of Bar-Coded Pyrosequencing for Evaluating Bacterial Community Shifts and Individual Taxa Dynamics
 Analysis of Run-to-Run Variation of Bar-Coded Pyrosequencing for Evaluating Bacterial Community Shifts and Individual Taxa Dynamics Yuan Ge 1,2,3, Joshua P. Schimel 3,4, Patricia A. Holden 1,2,3 * 1 Earth
Analysis of Run-to-Run Variation of Bar-Coded Pyrosequencing for Evaluating Bacterial Community Shifts and Individual Taxa Dynamics Yuan Ge 1,2,3, Joshua P. Schimel 3,4, Patricia A. Holden 1,2,3 * 1 Earth
Oral bacterial interactions in periodontal health and disease
 Vol. 6(5), pp. 51-57, August 2014 DOI: 10.5897/JDOH2014.0127 Article Number: 84910BB46956 ISSN 2006-9871 Copyright 2014 Author(s) retain the copyright of this article http://www.academicjournals.org/jdoh
Vol. 6(5), pp. 51-57, August 2014 DOI: 10.5897/JDOH2014.0127 Article Number: 84910BB46956 ISSN 2006-9871 Copyright 2014 Author(s) retain the copyright of this article http://www.academicjournals.org/jdoh
A MODIFIED DIP-SLIDE TEST FOR MICROBIOLOGICAL RISK IN CARIES ASSESSMENT
 A MODIFIED DIP-SLIDE TEST FOR MICROBIOLOGICAL RISK IN CARIES ASSESSMENT Boonyanit Thaweboon, Sroisiri Thaweboon, Cherdkul Sopavanit and Reda Kasetsuwan Department of Microbiology, Department of Community
A MODIFIED DIP-SLIDE TEST FOR MICROBIOLOGICAL RISK IN CARIES ASSESSMENT Boonyanit Thaweboon, Sroisiri Thaweboon, Cherdkul Sopavanit and Reda Kasetsuwan Department of Microbiology, Department of Community
Show biofilm the red card
 Show biofilm the red card BIOFILM-ERASER 1 A NEW APPLICATION Effective treatment for periodontal and peri-implant inflammation Periodontitis and peri-implantitis are bacterial inflammations with similar
Show biofilm the red card BIOFILM-ERASER 1 A NEW APPLICATION Effective treatment for periodontal and peri-implant inflammation Periodontitis and peri-implantitis are bacterial inflammations with similar
Manuscript Title: Responses in ileal and cecal bacteria to low and high amylose/amylopectin ratio diets in growing pigs
 Journal name: Applied Microbiology and Biotechnology Manuscript Title: Responses in ileal and cecal bacteria to low and high amylose/amylopectin ratio diets in growing pigs The names of the authors: Yu-heng
Journal name: Applied Microbiology and Biotechnology Manuscript Title: Responses in ileal and cecal bacteria to low and high amylose/amylopectin ratio diets in growing pigs The names of the authors: Yu-heng
Overview. Dental biofilm bacteria. Biofilm. Multi-species dental biofilms. Biofilm work group meeting Marts 15th Irene Dige
 Overview Multi-species dental biofilms Introduction to oral and dental biofilms Previous studies of dental biofilms Recent studies of multispecies dental biofilms Methodological considerations Irene Dige
Overview Multi-species dental biofilms Introduction to oral and dental biofilms Previous studies of dental biofilms Recent studies of multispecies dental biofilms Methodological considerations Irene Dige
Diversity and Morphology of Members of the Phylum Synergistetes in Periodontal Health and Disease
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, June 2009, p. 3777 3786 Vol. 75, No. 11 0099-2240/09/$08.000 doi:10.1128/aem.02763-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Diversity
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, June 2009, p. 3777 3786 Vol. 75, No. 11 0099-2240/09/$08.000 doi:10.1128/aem.02763-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Diversity
Biomarkers into saliva
 Biomarkers into saliva Saliva used as biomarker for risk for caries Cario-Analyse saliva testing strategies Saliva composition? Saliva comes from blood serum associated with oral microflora and viruses
Biomarkers into saliva Saliva used as biomarker for risk for caries Cario-Analyse saliva testing strategies Saliva composition? Saliva comes from blood serum associated with oral microflora and viruses
NIH Public Access Author Manuscript Quintessence Int. Author manuscript; available in PMC 2011 August 6.
 NIH Public Access Author Manuscript Published in final edited form as: Quintessence Int. 2007 April ; 38(4): e213 e217. Microbial acid production (Clinpro Cario L-Pop) and dental caries in infants and
NIH Public Access Author Manuscript Published in final edited form as: Quintessence Int. 2007 April ; 38(4): e213 e217. Microbial acid production (Clinpro Cario L-Pop) and dental caries in infants and
The Adult Cystic Fibrosis Airway Microbiota Is Stable over Time and Infection Type, and Highly Resilient to Antibiotic Treatment of Exacerbations
 The Adult Cystic Fibrosis Airway Microbiota Is Stable over Time and Infection Type, and Highly Resilient to Antibiotic Treatment of Exacerbations Anthony A. Fodor 1 *., Erich R. Klem 2, Deirdre F. Gilpin
The Adult Cystic Fibrosis Airway Microbiota Is Stable over Time and Infection Type, and Highly Resilient to Antibiotic Treatment of Exacerbations Anthony A. Fodor 1 *., Erich R. Klem 2, Deirdre F. Gilpin
ACCEPTED MANUSCRIPT. Community Acquired Rhinovirus Infection Is Associated With Changes in the Airway Microbiome
 Community Acquired Rhinovirus Infection Is Associated With Changes in the Airway Microbiome Kirsten M. Kloepfer, MD, MSa,, Vishal K. Sarsani, MSb, Valeriy Poroyko, PhDc, Wai Ming Lee, PhDd, Tressa E. Pappas,
Community Acquired Rhinovirus Infection Is Associated With Changes in the Airway Microbiome Kirsten M. Kloepfer, MD, MSa,, Vishal K. Sarsani, MSb, Valeriy Poroyko, PhDc, Wai Ming Lee, PhDd, Tressa E. Pappas,
LIFESTYLE-ASSOCIATED ALTERATIONS OF INTESTINAL
 LIFESTYLE-ASSOCIATED ALTERATIONS OF INTESTINAL MICROBIOTA AND THEIR POTENTIAL ROLE IN ETIOLOGY OF IBD: A METAGENOMIC ANALYSIS Stephan J. Ott, University Hospital (UKSH), Campus Kiel Institute for Clinical
LIFESTYLE-ASSOCIATED ALTERATIONS OF INTESTINAL MICROBIOTA AND THEIR POTENTIAL ROLE IN ETIOLOGY OF IBD: A METAGENOMIC ANALYSIS Stephan J. Ott, University Hospital (UKSH), Campus Kiel Institute for Clinical
Gut Lung Axis Implication of the Gut Microbiota beyond its niche
 Gut Lung Axis Implication of the Gut Microbiota beyond its niche Reema Subramanian PhD Candidate (4 th year) Supervisor: Prof. Margaret Ip Department of Microbiology, CUHK Joint Graduate Student Seminar
Gut Lung Axis Implication of the Gut Microbiota beyond its niche Reema Subramanian PhD Candidate (4 th year) Supervisor: Prof. Margaret Ip Department of Microbiology, CUHK Joint Graduate Student Seminar
Molecular Mapping to Species Level of the Tonsillar Crypt Microbiota Associated with Health and Recurrent Tonsillitis
 Molecular Mapping to Species Level of the Tonsillar Crypt Microbiota Associated with Health and Recurrent Tonsillitis Anders Jensen 1, Helena Fagö-Olsen 2, Christian Hjort Sørensen 2, Mogens Kilian 1 *
Molecular Mapping to Species Level of the Tonsillar Crypt Microbiota Associated with Health and Recurrent Tonsillitis Anders Jensen 1, Helena Fagö-Olsen 2, Christian Hjort Sørensen 2, Mogens Kilian 1 *
Microbial biofilms are
 Managing the complexity of a dynamic biofilm John G. Thomas, MS, PhD; Lindsay A. Nakaishi, BS Microbial biofilms are common in nature. Virtually any fluid environment in which microorganisms are subject
Managing the complexity of a dynamic biofilm John G. Thomas, MS, PhD; Lindsay A. Nakaishi, BS Microbial biofilms are common in nature. Virtually any fluid environment in which microorganisms are subject
Rate of cultivable subgingival periodontopathogenic bacteria in chronic periodontitis
 157 Journal of Oral Science, Vol. 46, No. 3, 157-161, 2004 Original Rate of cultivable subgingival periodontopathogenic bacteria in chronic periodontitis Mohammad Hossein Salari and Zainab Kadkhoda Department
157 Journal of Oral Science, Vol. 46, No. 3, 157-161, 2004 Original Rate of cultivable subgingival periodontopathogenic bacteria in chronic periodontitis Mohammad Hossein Salari and Zainab Kadkhoda Department
Gastric microbial community profiling reveals a dysbiotic cancer-associated microbiota
 Gastric microbial community profiling reveals a dysbiotic cancer-associated microbiota Rui M. Ferreira 1,2, Joana Pereira-Marques 1,2,3, Ines Pinto-Ribeiro 1,2,4, Jose L. Costa 1,2,4, Fatima Carneiro 1,2,4,5,
Gastric microbial community profiling reveals a dysbiotic cancer-associated microbiota Rui M. Ferreira 1,2, Joana Pereira-Marques 1,2,3, Ines Pinto-Ribeiro 1,2,4, Jose L. Costa 1,2,4, Fatima Carneiro 1,2,4,5,
Title: Cultivable Anaerobic Microbiota of Severe Early Childhood Caries.
 JCM Accepts, published online ahead of print on 2 February 2011 J. Clin. Microbiol. doi:10.1128/jcm.02427-10 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All
JCM Accepts, published online ahead of print on 2 February 2011 J. Clin. Microbiol. doi:10.1128/jcm.02427-10 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All
Supplementary Information
 Supplementary Information Methods Growth Characterisation Total Adenosine Triphosphate (ATP) was determined using a Molecular Probes ATP Determination Kit (Life Technologies, USA). 0.5 ml of sample was
Supplementary Information Methods Growth Characterisation Total Adenosine Triphosphate (ATP) was determined using a Molecular Probes ATP Determination Kit (Life Technologies, USA). 0.5 ml of sample was
Are microbes the missing piece of the gut comfort puzzle?
 Priority Research Programme Foods for improving gut function and comfort Are microbes the missing piece of the gut comfort puzzle? Wayne Young Senior Scientist, AgResearch Host institution Foods for gut
Priority Research Programme Foods for improving gut function and comfort Are microbes the missing piece of the gut comfort puzzle? Wayne Young Senior Scientist, AgResearch Host institution Foods for gut
Differences in Vaginal Bacterial Communities of Women in North America: Implications for disease diagnosis and prevention
 Differences in Vaginal Bacterial Communities of Women in North America: Implications for disease diagnosis and prevention Larry J. Forney, Pawel Gajer, Christopher J. Williams, Maria G. Schneider, Stacey
Differences in Vaginal Bacterial Communities of Women in North America: Implications for disease diagnosis and prevention Larry J. Forney, Pawel Gajer, Christopher J. Williams, Maria G. Schneider, Stacey
Going With Your Gut: The Microbiome and You
 Going With Your Gut: The Microbiome and You Robert T. Schooley, MD Professor of Medicine University of California San Diego San Diego, California Learning Objectives After attending this presentation,
Going With Your Gut: The Microbiome and You Robert T. Schooley, MD Professor of Medicine University of California San Diego San Diego, California Learning Objectives After attending this presentation,
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 NLRP12 is downregulated in biopsy samples from patients with active ulcerative colitis (UC). (a-g) NLRP12 expression in 7 UC mrna profiling studies deposited in NCBI GEO database.
Supplementary Figure 1 NLRP12 is downregulated in biopsy samples from patients with active ulcerative colitis (UC). (a-g) NLRP12 expression in 7 UC mrna profiling studies deposited in NCBI GEO database.
Bacterial Profiles of Root Caries in Elderly Patients
 JOURNAL OF CLINICAL MICROBIOLOGY, June 2008, p. 2015 2021 Vol. 46, No. 6 0095-1137/08/$08.00 0 doi:10.1128/jcm.02411-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Bacterial
JOURNAL OF CLINICAL MICROBIOLOGY, June 2008, p. 2015 2021 Vol. 46, No. 6 0095-1137/08/$08.00 0 doi:10.1128/jcm.02411-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Bacterial
Microbiome and Asthma
 제 12 차천식연구회 COPD 연구회공동심포지엄 Microbiome and Asthma 한양대학교병원호흡기알레르기내과 김상헌 Disclosure 내용 1 Lung Microbiome 2 Lung Microbiome and Asthma 3 Gut Microbiome and Asthma Microbiome and Microbiota human microbiome
제 12 차천식연구회 COPD 연구회공동심포지엄 Microbiome and Asthma 한양대학교병원호흡기알레르기내과 김상헌 Disclosure 내용 1 Lung Microbiome 2 Lung Microbiome and Asthma 3 Gut Microbiome and Asthma Microbiome and Microbiota human microbiome
Intra-oral microbial profiles of beagle dogs assessed by checkerboard DNA-DNA hybridization using human probes
 Intra-oral microbial profiles of beagle dogs assessed by checkerboard DNA-DNA hybridization using human probes M. Rober, M. Quirynen, A.D. Haffajee, E. Schepers, W. Teughels To cite this version: M. Rober,
Intra-oral microbial profiles of beagle dogs assessed by checkerboard DNA-DNA hybridization using human probes M. Rober, M. Quirynen, A.D. Haffajee, E. Schepers, W. Teughels To cite this version: M. Rober,
Probiotics In Gingivitis Management: A Randomized Clinical Trial. Jordi Espadaler, PhD. Director of Innovation
 Probiotics In Gingivitis Management: A Randomized Clinical Trial Jordi Espadaler, PhD. Director of Innovation espadaler@ab-biotics.com Improving oral care Mechanical plaque removal, brushing and rinsing
Probiotics In Gingivitis Management: A Randomized Clinical Trial Jordi Espadaler, PhD. Director of Innovation espadaler@ab-biotics.com Improving oral care Mechanical plaque removal, brushing and rinsing
Subgingival microbiota of chronic periodontitis subjects from different geographic locations
 J Clin Periodontol 2004; 31: 996 1002 doi: 10.1111/j.1600-051X.2004.00597.x Copyright r Blackwell Munksgaard 2004 Printed in Denmark. All rights reserved Subgingival microbiota of chronic periodontitis
J Clin Periodontol 2004; 31: 996 1002 doi: 10.1111/j.1600-051X.2004.00597.x Copyright r Blackwell Munksgaard 2004 Printed in Denmark. All rights reserved Subgingival microbiota of chronic periodontitis
Title:Pyrosequencing of supra- and subgingival biofilms from inflamed peri-implant and periodontal sites
 Author's response to reviews Title:Pyrosequencing of supra- and subgingival biofilms from inflamed peri-implant and periodontal sites Authors: Joerg Eberhard (eberhard.joerg@mh-hannover.de) Simone Schaumann
Author's response to reviews Title:Pyrosequencing of supra- and subgingival biofilms from inflamed peri-implant and periodontal sites Authors: Joerg Eberhard (eberhard.joerg@mh-hannover.de) Simone Schaumann
Citation for published version (APA): Volgenant, C. M. C. (2016). Red fluorescent dental plaque: An indicator of oral disease?
 UvA-DARE (Digital Academic Repository) Red fluorescent dental plaque: An indicator of oral disease? Volgenant, C.M.C. Link to publication Citation for published version (APA): Volgenant, C. M. C. (2016).
UvA-DARE (Digital Academic Repository) Red fluorescent dental plaque: An indicator of oral disease? Volgenant, C.M.C. Link to publication Citation for published version (APA): Volgenant, C. M. C. (2016).
Effect of temperature change of 0.2% chlorhexidine rinse on matured human plaque: an in vivo study.
 ISSN: 2278 0211 (Online) Effect of temperature change of 0.2% chlorhexidine rinse on matured human plaque: an in vivo study. Dr. Yashika Jain Senior Lecturer, Institution: SGT Dental College & Hospital,
ISSN: 2278 0211 (Online) Effect of temperature change of 0.2% chlorhexidine rinse on matured human plaque: an in vivo study. Dr. Yashika Jain Senior Lecturer, Institution: SGT Dental College & Hospital,
VIP: an integrated pipeline for metagenomics of virus
 VIP: an integrated pipeline for metagenomics of virus identification and discovery Yang Li 1, Hao Wang 2, Kai Nie 1, Chen Zhang 1, Yi Zhang 1, Ji Wang 1, Peihua Niu 1 and Xuejun Ma 1 * 1. Key Laboratory
VIP: an integrated pipeline for metagenomics of virus identification and discovery Yang Li 1, Hao Wang 2, Kai Nie 1, Chen Zhang 1, Yi Zhang 1, Ji Wang 1, Peihua Niu 1 and Xuejun Ma 1 * 1. Key Laboratory
Drinking alcohol is associated with variation in the human oral microbiome in a large study of American adults
 Fan et al. Microbiome (2018) 6:59 https://doi.org/10.1186/s40168-018-0448-x RESEARCH Drinking alcohol is associated with variation in the human oral microbiome in a large study of American adults Open
Fan et al. Microbiome (2018) 6:59 https://doi.org/10.1186/s40168-018-0448-x RESEARCH Drinking alcohol is associated with variation in the human oral microbiome in a large study of American adults Open
Identification of the microorganisms responsible for periodontopathy by Multiplex RT-PCR
 ORIGINAL ARTICLES Identification of the microorganisms responsible for periodontopathy by Multiplex RT-PCR R. SQUERI, V. LA FAUCI, G. CANNAVÒ *, G. LO GIUDICE **, L. SINDONI University of Messina, Department
ORIGINAL ARTICLES Identification of the microorganisms responsible for periodontopathy by Multiplex RT-PCR R. SQUERI, V. LA FAUCI, G. CANNAVÒ *, G. LO GIUDICE **, L. SINDONI University of Messina, Department
Rationale. A complete view of endometrial health
 Rationale The Human Microbiome Project (HMP) has highlighted the importance of different microorganisms and their genomes in human health and disease (Human Microbiome Project Consortium, 2012). This knowledge
Rationale The Human Microbiome Project (HMP) has highlighted the importance of different microorganisms and their genomes in human health and disease (Human Microbiome Project Consortium, 2012). This knowledge
Dental Caries Prevalence in Human Immunodeficiency Virus Infected Patients Receiving Highly Active Anti-Retroviral Therapy in Kermanshah, Iran
 Original Article Dental Caries Prevalence in Human Immunodeficiency Virus Infected Patients Receiving Highly Active Anti-Retroviral Therapy in Kermanshah, Iran Loghman Rezaei-Soufi, D.D.S., M.Sc. 1, Poorandokht
Original Article Dental Caries Prevalence in Human Immunodeficiency Virus Infected Patients Receiving Highly Active Anti-Retroviral Therapy in Kermanshah, Iran Loghman Rezaei-Soufi, D.D.S., M.Sc. 1, Poorandokht
The human body is made up of over cells of which
 Sci. and Cult. 83 (9 10) 318-322 (2017) Establishment of Oral Bacterial Flora An Overview Abstract : Until the time of birth the human infant is usually "germfree." The newborn then becomes suddenly exposed
Sci. and Cult. 83 (9 10) 318-322 (2017) Establishment of Oral Bacterial Flora An Overview Abstract : Until the time of birth the human infant is usually "germfree." The newborn then becomes suddenly exposed
Sequencing-Based Analysis for the Identification of Bacteria Associated with Severe Early Childhood Caries (SECC)
 Sequencing-Based Analysis for the Identification of Bacteria Associated with Severe Early Childhood Caries (SECC) Zarina Zainuddin 1, Abdul Qahhar Bin Paiman 1, Yunita Dewi Ardini 2 1 Department of Biotechnology,
Sequencing-Based Analysis for the Identification of Bacteria Associated with Severe Early Childhood Caries (SECC) Zarina Zainuddin 1, Abdul Qahhar Bin Paiman 1, Yunita Dewi Ardini 2 1 Department of Biotechnology,
Identification of Microbial Pathogens in Periodontal disease and Diabetic patients of South Indian Population
 www.bioinformation.net Research note Volume 10(4) Identification of Microbial Pathogens in Periodontal disease and Diabetic patients of South Indian Population Tikka Chiranjeevi 1, Osuru Hari Prasad 1,
www.bioinformation.net Research note Volume 10(4) Identification of Microbial Pathogens in Periodontal disease and Diabetic patients of South Indian Population Tikka Chiranjeevi 1, Osuru Hari Prasad 1,
JSS Medical and Dental College and Hospital, JSS University, Mysore, India *Author for Correspondence
 CLINICAL, MICROBIOLOGICAL AND MOLECULAR STUDY OF PORPHYROMONAS GINGIVALIS IN PATIENTS WITH CHRONIC PERIODONTITIS *Mayuri Vajawat 1, Vijay Kumar 2, K.G. Rajeshwari 3 and P.C. Deepika 4 1&4 Department of
CLINICAL, MICROBIOLOGICAL AND MOLECULAR STUDY OF PORPHYROMONAS GINGIVALIS IN PATIENTS WITH CHRONIC PERIODONTITIS *Mayuri Vajawat 1, Vijay Kumar 2, K.G. Rajeshwari 3 and P.C. Deepika 4 1&4 Department of
Normal Flora PROF. HANAN HABIB DEPARTMENT OF PATHOLOGY COLLEGE OF MEDICINE, KSU
 Normal Flora PROF. HANAN HABIB DEPARTMENT OF PATHOLOGY COLLEGE OF MEDICINE, KSU Objectives 1. Define the terms: Normal Flora, Resident flora, Transient flora and carrier state 2. Know the origin of normal
Normal Flora PROF. HANAN HABIB DEPARTMENT OF PATHOLOGY COLLEGE OF MEDICINE, KSU Objectives 1. Define the terms: Normal Flora, Resident flora, Transient flora and carrier state 2. Know the origin of normal
Development and Application of an Enteric Pathogens Microarray
 Development and Application of an Enteric Pathogens Microarray UC Berkeley School of Public Health Sona R. Saha, MPH Joseph Eisenberg, PhD Lee Riley, MD Alan Hubbard, PhD Jack Colford, MD PhD East Bay
Development and Application of an Enteric Pathogens Microarray UC Berkeley School of Public Health Sona R. Saha, MPH Joseph Eisenberg, PhD Lee Riley, MD Alan Hubbard, PhD Jack Colford, MD PhD East Bay
Persson GR, Salvi GE, Heitz-Mayfield LJA et al. Antimicrobial therapy using a local drug delivery system (Arestin) in the treatment of
 Persson GR, Salvi GE, Heitz-Mayfield LJA et al. Antimicrobial therapy using a local drug delivery system (Arestin) in the treatment of peri-implantitis I: microbiological outcomes. Clin Oral Imp Res 2006;
Persson GR, Salvi GE, Heitz-Mayfield LJA et al. Antimicrobial therapy using a local drug delivery system (Arestin) in the treatment of peri-implantitis I: microbiological outcomes. Clin Oral Imp Res 2006;
GSK Medicine: Study Number: Title: Rationale: Phase: Study Period Study Design: Group 1 (Test Group) Group 2 (Reference group):
 The study listed may include approved and non-approved uses, formulations or treatment regimens. The results reported in any single study may not reflect the overall results obtained on studies of a product.
The study listed may include approved and non-approved uses, formulations or treatment regimens. The results reported in any single study may not reflect the overall results obtained on studies of a product.
Normal Flora. CLS 212: Medical Microbiology
 Normal Flora CLS 212: Medical Microbiology Relationships between Organisms Symbiosis Permanent association between two different organisms. Neutralism Two organisms living together, and neither is affected
Normal Flora CLS 212: Medical Microbiology Relationships between Organisms Symbiosis Permanent association between two different organisms. Neutralism Two organisms living together, and neither is affected
SUPPLEMENTARY FIGURES & TABLES
 SUPPLEMENTARY FIGURES & TABLES Figures S1-S6 Tables S1-S5 A 6 Bacteroidetes/Firmicutes B 1.0 Prevotellaceae/Bacteroidetes Ratio 5 4 3 2 Ratio 0.8 0.6 0.4 1 0.2 0 1 2 3 4 5 6 BSS 0.0 1 2 3 4 5 6 BSS Supplementary
SUPPLEMENTARY FIGURES & TABLES Figures S1-S6 Tables S1-S5 A 6 Bacteroidetes/Firmicutes B 1.0 Prevotellaceae/Bacteroidetes Ratio 5 4 3 2 Ratio 0.8 0.6 0.4 1 0.2 0 1 2 3 4 5 6 BSS 0.0 1 2 3 4 5 6 BSS Supplementary
International Journal of Pharma and Bio Sciences PERIODONTAL PATHOGENS AMONG PRE PUBERTAL, PUBERTAL AND POST PUBERTAL GIRLS, IN CHENNAI, INDIA.
 Research Article Microbiology International Journal of Pharma and Bio Sciences ISSN 0975-6299 PERIODONTAL PATHOGENS AMONG PRE PUBERTAL, PUBERTAL AND POST PUBERTAL GIRLS, IN CHENNAI, INDIA. KANAKAM ELIZABETH
Research Article Microbiology International Journal of Pharma and Bio Sciences ISSN 0975-6299 PERIODONTAL PATHOGENS AMONG PRE PUBERTAL, PUBERTAL AND POST PUBERTAL GIRLS, IN CHENNAI, INDIA. KANAKAM ELIZABETH
Hundreds of bacterial species populate the body. The oral cavity provides a unique environment
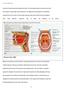 1.0 Introduction Hundreds of bacterial species populate the body. The oral cavity provides a unique environment that supports a wide range of bacterial species. The highly diverse flora grows in the different
1.0 Introduction Hundreds of bacterial species populate the body. The oral cavity provides a unique environment that supports a wide range of bacterial species. The highly diverse flora grows in the different
Bio Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease
 Bio 230 - Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease It has been calculated that the normal human houses about 10 12 bacteria on the skin, 10 10 in the mouth, and
Bio 230 - Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease It has been calculated that the normal human houses about 10 12 bacteria on the skin, 10 10 in the mouth, and
PERIODONTAL CASE PRESENTATION - 1
 PERIODONTAL CASE PRESENTATION - 1 Overview A 32 year-old patient presented with generalized aggressive periodontitis. Treatment included non-surgical therapy with adjunctive antibiotics and surgical treatment.
PERIODONTAL CASE PRESENTATION - 1 Overview A 32 year-old patient presented with generalized aggressive periodontitis. Treatment included non-surgical therapy with adjunctive antibiotics and surgical treatment.
Prevalent opportunistic infections associated with HIV-positive children 0-5 years in Benin city, Nigeria
 Malaysian Journal of Microbiology, Vol 4(2) 2008, pp. 11-14 http://dx.doi.org/10.21161/mjm.11508 Prevalent opportunistic infections associated with HIV-positive children 0-5 years in Benin city, Nigeria
Malaysian Journal of Microbiology, Vol 4(2) 2008, pp. 11-14 http://dx.doi.org/10.21161/mjm.11508 Prevalent opportunistic infections associated with HIV-positive children 0-5 years in Benin city, Nigeria
Subgingival microbiota of periodontally untreated Mexican subjects with generalized aggressive periodontitis
 J Clin Periodontol 2006; 33: 869 877 doi: 10.1111/j.1600-051X.2006.01006.x Subgingival microbiota of periodontally untreated Mexican subjects with generalized aggressive periodontitis Laurie Ann Ximenez-Fyvie,
J Clin Periodontol 2006; 33: 869 877 doi: 10.1111/j.1600-051X.2006.01006.x Subgingival microbiota of periodontally untreated Mexican subjects with generalized aggressive periodontitis Laurie Ann Ximenez-Fyvie,
Molecular Diagnosis Future Directions
 Molecular Diagnosis Future Directions Philip Cunningham NSW State Reference Laboratory for HIV/AIDS & Molecular Diagnostic Medicine Laboratory, SydPath St Vincent s Hospital Sydney Update on Molecular
Molecular Diagnosis Future Directions Philip Cunningham NSW State Reference Laboratory for HIV/AIDS & Molecular Diagnostic Medicine Laboratory, SydPath St Vincent s Hospital Sydney Update on Molecular
Microbial Risk Indicators of Early Childhood Caries
 JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2005, p. 5753 5759 Vol. 43, No. 11 0095-1137/05/$08.00 0 doi:10.1128/jcm.43.11.5753 5759.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2005, p. 5753 5759 Vol. 43, No. 11 0095-1137/05/$08.00 0 doi:10.1128/jcm.43.11.5753 5759.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
DHR International Journal Of Medical Sciences (DHR-IJMS) ISSN: X, Vol. 3(1), 2012 Available online
 DHR International Journal Of Medical Sciences (DHR-IJMS) ISSN: 2278-831X, Vol. 3(1), 2012 Available online http://www.doublehelixresearch.com/dhrijms Double Helix Research Oral probiotics: the beneficial
DHR International Journal Of Medical Sciences (DHR-IJMS) ISSN: 2278-831X, Vol. 3(1), 2012 Available online http://www.doublehelixresearch.com/dhrijms Double Helix Research Oral probiotics: the beneficial
Effect of Systemically Administered Azithromycin in Early Onset Aggressive Periodontitis
 CLINICAL AND RESEARCH REPORTS Effect of Systemically Administered Azithromycin in Early Onset Aggressive Periodontitis Takeo Fujii, Pao-Li Wang, Yoichiro Hosokawa, Shinichi Shirai, Atsumu Tamura, Kazuhiro
CLINICAL AND RESEARCH REPORTS Effect of Systemically Administered Azithromycin in Early Onset Aggressive Periodontitis Takeo Fujii, Pao-Li Wang, Yoichiro Hosokawa, Shinichi Shirai, Atsumu Tamura, Kazuhiro
DURATION OF THE STUDY: JUNE-OCTOBER 2008 COST OF STUDY; 9400KSH SOURCE OF FUNDS: SELF INVESTIGATOR: GIKUNDA MARY KATHURE
 J ORAL MANIFESTATIONS OF HIV INFECTION/AIDS AND THEIR INFLUENCE ON ORAL FUNCTIONS AND ORAL HYGIENE PRACTICES AMONG ADULT PATIENTS IN KENYATTA NATIONAL HOSPITAL COMPREHENSIVE CARE CENTRE. INVESTIGATOR:
J ORAL MANIFESTATIONS OF HIV INFECTION/AIDS AND THEIR INFLUENCE ON ORAL FUNCTIONS AND ORAL HYGIENE PRACTICES AMONG ADULT PATIENTS IN KENYATTA NATIONAL HOSPITAL COMPREHENSIVE CARE CENTRE. INVESTIGATOR:
Exploring a possible correlation between the human oral microbiome and body mass index
 University of Colorado, Boulder CU Scholar Undergraduate Honors Theses Honors Program Spring 2016 Exploring a possible correlation between the human oral microbiome and body mass index Ivy S. McDermott
University of Colorado, Boulder CU Scholar Undergraduate Honors Theses Honors Program Spring 2016 Exploring a possible correlation between the human oral microbiome and body mass index Ivy S. McDermott
THE ROLE OF MICROBIOME IN IBD
 Disclosures THE ROLE OF MICROBIOME IN IBD Janssen UCB No relevance to the talk Subra Kugathasan, MD Professor of Pediatrics & Human Genetics Marcus Professor of Pediatric Gastroenterology Emory University
Disclosures THE ROLE OF MICROBIOME IN IBD Janssen UCB No relevance to the talk Subra Kugathasan, MD Professor of Pediatrics & Human Genetics Marcus Professor of Pediatric Gastroenterology Emory University
The clinical, microbiological and systemic characteristics of periodontitis and their changes after periodontal therapy Bizzarro, S.
 UvA-DARE (Digital Academic Repository) The clinical, microbiological and systemic characteristics of periodontitis and their changes after periodontal therapy Bizzarro, S. Link to publication Citation
UvA-DARE (Digital Academic Repository) The clinical, microbiological and systemic characteristics of periodontitis and their changes after periodontal therapy Bizzarro, S. Link to publication Citation
Microbiome GI Disorders
 Microbiome GI Disorders Prof. Ram Dickman Neurogastroenterology Unit Rabin Medical Center Israel 1 Key Points Our gut microbiota Were to find them? Symbiosis or Why do we need them? Dysbiosis or when things
Microbiome GI Disorders Prof. Ram Dickman Neurogastroenterology Unit Rabin Medical Center Israel 1 Key Points Our gut microbiota Were to find them? Symbiosis or Why do we need them? Dysbiosis or when things
Breast tissue, oral and urinary microbiomes in breast cancer
 /, Advance Publications 2017 Breast tissue, oral and urinary microbiomes in breast cancer Hannah Wang 1,2, Jessica Altemus 1, Farshad Niazi 1, Holly Green 4, Benjamin C. Calhoun 5, Charles Sturgis 5, Stephen
/, Advance Publications 2017 Breast tissue, oral and urinary microbiomes in breast cancer Hannah Wang 1,2, Jessica Altemus 1, Farshad Niazi 1, Holly Green 4, Benjamin C. Calhoun 5, Charles Sturgis 5, Stephen
