Study of different types of ubiquitination
|
|
|
- Teresa Rich
- 6 years ago
- Views:
Transcription
1 Study of different types of ubiquitination Rudi Beyaert VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October 5, 2017 Provinciehuis, Leuven
2 Overview Introduction Ubiquitin Ubiquitination machinery Types of ubiquitination Simple biochemical methods Is protein X ubiquitinated? What type of ubiquitination is it? Troubleshooting
3 Ubiquitin 76 Amino Acids Compact, globular protein, 8.5 kda Highly conserved (3 aa difference from yeast to man) Expressed in every eukaryotic cell Post-translational modification on proteins Attaches on Lysine residues or N-terminal Methionine Multiple functions
4 Ubiquitin Ubiquitin Adapted from Komander et al., EMBO Rep., 2009
5 Ubiquitin The discovery of ubiquitin-mediated protein degradation Aaron Ciechanover Avram Hershko Irwin Rose Cells give a chemical "kiss of death" to proteins that need to be destroyed. A. Ciechanover et al., Biochem Biophys Res Comm 81 (1978)
6 The ubiquitination machinery Proteasomal degradation Signal transduction Endocytosis Vesicle and protein trafficking DNA repair Autophagy Cell cycle. Inflammation Cancer.
7 Types of ubiquitination G David Komander 7
8 Different ubiquitin chains have different architectures M1 8
9 The ubiquitin code Different types of ubiquitination are recognized by specific ubiquitin binding domains (UBD) in other proteins (Ub receptors) Simone Fulda et al. EMBO Mol Med. 2012;4:
10 Ubiquitination is key in NF-kB signaling
11 Phosphorylation versus Ubiquitination Sylvie Urbé 11
12 New complexity in the ubiquitin code K. Swatek and D. Komander, Cell Res,
13 Ubiquitination - questions Is my protein ubiquitinated? What type of ubiquitin chain? Which residue?
14 Is protein X ubiquitinated? Ubiquitination increases MW of protein X (immunoblotting) Ubiquitin = 8.5 kda; but mostly a high MW smear instead of defined bands differences in polyubiquitin chain length heterogeneously ubiquitinated at multiple sites different chain types lead to distinct motilities Can also reflect other types of PTMs treat with DUB before loading Emmerich C and Cohen P, BBRC, 2015
15 Is protein X ubiquitinated? In most cases a smear is not visible; reasons? protein ubiquitination is rapidly reversed by DUBs add DUB inhibitors (Nethylmaleimide/iodoacetamide + EDTA/EGTA) to the lysis buffer or work in cells in which DUB expression is silenced (M1- Ub) (K48)polyubiquitinated protein is rapidly degraded by the proteasome add proteasome inhibitors to the cells (e.g. MG132) only small fraction of protein is modified requires enrichment of ubiquitinated proteins Emmerich C and Cohen P, BBRC, 2015
16 Is protein X ubiquitinated? Enrichment of ubiquitinated proteins Immunoprecipitation (IP) with antiprotein X + immunoblotting for ubiquitin needs cell lysis in stringent (1% SDS) conditions to eliminate detection of other interacting proteins that are ubiquitinated; 10x dilution of cell lysate needed for IP (needs native conditions) sensitivity can be increased by using epitope tagged ubiquitin (caution!)
17 Overexpression of tagged Ub may render the system constitutive!
18 Commercial anti-ub antibodies do not recognize all ubiquitin linkages equally! Emmerich C and Cohen P, BBRC, 2015
19 Is protein X ubiquitinated? Enrichment of ubiquitinated proteins Affinity purification of His-tagged ubiquitin via Ni 2+ -affinity chromatography under denaturing conditions, followed by immunoblotting for protein X (caution: tagged Ub)
20 Is protein X ubiquitinated? Enrichment of ubiquitinated proteins Affinity purification with TUBEs TUBEs = ubiquitin traps = repeats of UBDs (with specificity for certain polyubiquitin-linkages) TUBEs show almost a 1000-fold increase in affinity for polyubiquitin moieties over a single UBD TUBEs protect against DUBs TUBEs can be immobilized in different ways: GST-TUBE, His-TUBE, Halo-TUBE,
21 TUBE affinity purification 21
22 TUBE affinity purification
23 Is protein X ubiquitinated? Global ubiquitin profiling using MS + enrichment for di-glylys containing peptides using digly-lys specific antibodies Determining the Lys acceptor site in the substrate (mutagenesis mostly does not work due to jumping sites)
24 D-GlyLys signature is not unique for ubiquitin! Need for ubiquitin enrichment by prior affinity purification of ubiquitinated proteins using e.g. anti-ubiquitin or Halo-TUBE
25 What type of ubiquitination do I have? Use of epitope tagged ubiquitin mutants (caution) Two complementary sets of ubiquitin mutants are available
26 What type of ubiquitination do I have? Use of ubiquitin linkage specific antibodies Emmerich C and Cohen P, BBRC, 2015
27 What type of ubiquitination do I have? Use of ubiquitin linkage specific TUBEs > 20 different UBDs
28 What type of ubiquitination do I have? Use of poly-ubiquitin linkage specific TUBEs
29 What type of ubiquitination do I have? Use of poly-ubiquitin linkage specific DUBs David Komander
30 What type of ubiquitination do I have? Ubiquitin chain restriction analysis (UbiCRest): linkage specific DUBs are used as restriction enzymes in ubiquitin sequencing M1 K63 K48 David Komander
31 What type of ubiquitination do I have? Ubiquitin chain restriction analysis (UbiCRest): linkage specific DUBs allow analysis of ubiquitin chain architecture (linkage-type + topology) K48 K6 K11 Mixed K6-K48 Ub5-6
32 What type of ubiquitination do I have? Ubiquitin chain restriction analysis (UbiCRest): linkage specific DUBs allow analysis of ubiquitin chain architecture FLAG-TNFa purification of TNF receptor complex Yogesh Kulathu
33 What type of ubiquitination do I have? Quantification of unique digly signature peptides for each polyubiquitin linkage after trypsin cleavage
34 Troubleshooting NEM is better at preserving K63- and M1-Ub chains than IAA (unstable, destroyed by light) IAA may interfere with the identification of Ub sites by MS (IAA reaction with Cys leads to covalent 2-acetamidoacetamide with a MW of 114Da = that of digly dipeptide (use NEM) MG132 can have cytotoxic effects; increased ubiquitination may be due to cellular stress Amount of ubiquitin in cells is very high (~85µM in HEK293) needs high amounts of anti-ubiquitin to IP all ubiquitinated proteins (first IP protein of interest and use anti-ub for immunoblotting) Antibody epitope may be blocked/masked by the presence of polyubiquitin chains (use polyclonal Ab) (e.g. IL-1 induced degradation of IRAK1 is false!)
35 Troubleshooting Ubiquitin = globular protein; epitopes may not be accessible to antibodies due to insufficient denaturation during SDS-PAGE signal strength can be enhanced by extra denaturation step (15 min boiling water, autoclaving, ) Overexpression of tagged Ub may lead to ubiquitination of proteins that are normally not ubiquitinated N- or C-terminally tagged ubiquitin can not be used in case of M1-Ub (use of internally Strep-tagged Ub) Ubiquitin mutants may not fold equally well, have different affinitiy for UBDs, affect deubiquitination, GGK-specific monoclonal antibodies do not recognize the characteristic GGMQIFVK signature peptides resulting from tryptic cleavage of M1-polyubiquitin
36 Troubleshooting UBD linkage specificity is not 100% Also use immobilized Ub-binding defective UBD mutant to establish that the interaction with the ubiquitinated protein is specific GST-TUBE forms dimers, which can alter linkage specificity
37 Plasmids for ubiquitin research available at BCCM- LMBP plasmid collection at UGent-IRC 37
38 38
The Nobel Prize in Chemistry 2004
 The Nobel Prize in Chemistry 2004 Ubiquitous Quality Control of Life C S Karigar and K R Siddalinga Murthy The Nobel Prize in Chemistry for 2004 is shared by Aaron Ciechanover, Avram Hershko and Irwin
The Nobel Prize in Chemistry 2004 Ubiquitous Quality Control of Life C S Karigar and K R Siddalinga Murthy The Nobel Prize in Chemistry for 2004 is shared by Aaron Ciechanover, Avram Hershko and Irwin
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
The ubiquitin-proteasome system. in bone biology. University of Nottingham. ECTS PhD Training July 5 th Dr Rob Layfield
 The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
Broad Spectrum Protease Inhibitor Cocktail
 Broad Spectrum Protease Inhibitor Cocktail Catalog number: AR1182 Boster s Broad Spectrum Protease Inhibitor Cocktail is a complex of various protease inhibitors, which has been tested for inhibiting proteases
Broad Spectrum Protease Inhibitor Cocktail Catalog number: AR1182 Boster s Broad Spectrum Protease Inhibitor Cocktail is a complex of various protease inhibitors, which has been tested for inhibiting proteases
The K48-K63 Branched Ubiquitin Chain Regulates NF-kB Signaling
 Article The K48-K63 Branched Ubiquitin Chain Regulates NF-kB Signaling Graphical Abstract Authors Fumiaki Ohtake, Yasushi Saeki, Satoshi Ishido, Jun Kanno, Keiji Tanaka Correspondence otake-fm@igakuken.or.jp
Article The K48-K63 Branched Ubiquitin Chain Regulates NF-kB Signaling Graphical Abstract Authors Fumiaki Ohtake, Yasushi Saeki, Satoshi Ishido, Jun Kanno, Keiji Tanaka Correspondence otake-fm@igakuken.or.jp
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Degradation needs no energy or does it?
 English Swedish The Nobel Prize in Chemistry 2004 Information for the Public 6 October 2004 A human cell contains some hundred thousand different proteins. These have numerous important functions: as accelerators
English Swedish The Nobel Prize in Chemistry 2004 Information for the Public 6 October 2004 A human cell contains some hundred thousand different proteins. These have numerous important functions: as accelerators
Nature Structural & Molecular Biology: doi: /nsmb.3218
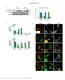 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1%
 FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
Why do cellular proteins linked to K63-polyubiquitin chains not associate with proteasomes?
 The EMBO Journal (2013) 32, 552 565 www.embojournal.org Why do cellular proteins linked to K63-polyubiquitin chains not associate with proteasomes? THE EMBO JOURNAL James A Nathan 1,2, Hyoung Tae Kim 1,
The EMBO Journal (2013) 32, 552 565 www.embojournal.org Why do cellular proteins linked to K63-polyubiquitin chains not associate with proteasomes? THE EMBO JOURNAL James A Nathan 1,2, Hyoung Tae Kim 1,
Complexity DNA. Genome RNA. Transcriptome. Protein. Proteome. Metabolites. Metabolome
 DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
E2Select Ubiquitin Conjugation Kit
 E2Select Ubiquitin Conjugation Kit Catalog Number K-982 This kit contains reagents sufficient to perform 2 Ubiquitination reactions for each E2 included in the kit. This package insert must be read in
E2Select Ubiquitin Conjugation Kit Catalog Number K-982 This kit contains reagents sufficient to perform 2 Ubiquitination reactions for each E2 included in the kit. This package insert must be read in
The Immunoassay Guide to Successful Mass Spectrometry. Orr Sharpe Robinson Lab SUMS User Meeting October 29, 2013
 The Immunoassay Guide to Successful Mass Spectrometry Orr Sharpe Robinson Lab SUMS User Meeting October 29, 2013 What is it? Hey! Look at that! Something is reacting in here! I just wish I knew what it
The Immunoassay Guide to Successful Mass Spectrometry Orr Sharpe Robinson Lab SUMS User Meeting October 29, 2013 What is it? Hey! Look at that! Something is reacting in here! I just wish I knew what it
Ub-CHOP-Reporter Deubiquitination Assay Kit Catalog Number PR1001
 - 1 - MANUAL Ub-CHOP-Reporter Deubiquitination Assay Kit Catalog Number PR1001 all products are for research use only not intended for human or animal diagnostic or therapeutic uses LifeSensors, Inc.,
- 1 - MANUAL Ub-CHOP-Reporter Deubiquitination Assay Kit Catalog Number PR1001 all products are for research use only not intended for human or animal diagnostic or therapeutic uses LifeSensors, Inc.,
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
crossmark Ca V subunits interact with the voltage-gated calcium channel
 crossmark THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 291, NO. 39, pp. 20402 20416, September 23, 2016 Author s Choice 2016 by The American Society for Biochemistry and Molecular Biology, Inc. Published in
crossmark THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 291, NO. 39, pp. 20402 20416, September 23, 2016 Author s Choice 2016 by The American Society for Biochemistry and Molecular Biology, Inc. Published in
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Cellular functions of protein degradation
 Protein Degradation Cellular functions of protein degradation 1. Elimination of misfolded and damaged proteins: Environmental toxins, translation errors and genetic mutations can damage proteins. Misfolded
Protein Degradation Cellular functions of protein degradation 1. Elimination of misfolded and damaged proteins: Environmental toxins, translation errors and genetic mutations can damage proteins. Misfolded
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Protein methylation CH 3
 Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Appendix. Table of Contents
 Appendix Table of Contents Appendix Figures Figure S1: Gp78 is not required for the degradation of mcherry-cl1 in Hela Cells. Figure S2: Indel formation in the MARCH6 sgrna targeted HeLa clones. Figure
Appendix Table of Contents Appendix Figures Figure S1: Gp78 is not required for the degradation of mcherry-cl1 in Hela Cells. Figure S2: Indel formation in the MARCH6 sgrna targeted HeLa clones. Figure
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
SUPPLEMENTARY INFORMATION
 Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
Figure S1. Silver staining and immunoblotting of the purified TAK1 kinase complex. The TAK1 kinase complex was purified through tandem affinity methods (Protein A and FLAG), and aliquots of the purified
Overview of the Expressway Cell-Free Expression Systems. Expressway Mini Cell-Free Expression System
 Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
20S Proteasome Activity Assay Kit
 20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
Protein Trafficking in the Secretory and Endocytic Pathways
 Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
supplementary information
 DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
DOI: 10.1038/ncb1875 Figure S1 (a) The 79 surgical specimens from NSCLC patients were analysed by immunohistochemistry with an anti-p53 antibody and control serum (data not shown). The normal bronchi served
Index. Index 371. Aggresome, cystic fibrosis transmembrane conductance regulator,
 Index 371 Index Aggresome, cystic fibrosis transmembrane conductance regulator, centrosome-associated inclusions, characterization of soluble and insoluble fractions, 319 formation induction, 318, 320,
Index 371 Index Aggresome, cystic fibrosis transmembrane conductance regulator, centrosome-associated inclusions, characterization of soluble and insoluble fractions, 319 formation induction, 318, 320,
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
SUPPLEMENTAL FIGURE LEGENDS
 SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
SUPPLEMENTAL FIGURE LEGENDS Supplemental Figure S1: Endogenous interaction between RNF2 and H2AX: Whole cell extracts from 293T were subjected to immunoprecipitation with anti-rnf2 or anti-γ-h2ax antibodies
Enhanced Purification of Ubiquitinated Proteins by Engineered Tandem Hybrid
 MCP Papers in Press. Published on February 3, 2016 as Manuscript O115.051839 Enhanced Purification of Ubiquitinated Proteins by Engineered Tandem Hybrid Ubiquitin-binding Domains (ThUBDs) Yuan Gao 1,ǂ,
MCP Papers in Press. Published on February 3, 2016 as Manuscript O115.051839 Enhanced Purification of Ubiquitinated Proteins by Engineered Tandem Hybrid Ubiquitin-binding Domains (ThUBDs) Yuan Gao 1,ǂ,
AccuMAP Low ph Protein Digestion Kits
 TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Ubiquitin modifications
 REVIEW Ubiquitin modifications Cell Research (2016) :1-24. www.nature.com/cr npg Kirby N Swatek 1, David Komander 1 1 Medical Research Council Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge,
REVIEW Ubiquitin modifications Cell Research (2016) :1-24. www.nature.com/cr npg Kirby N Swatek 1, David Komander 1 1 Medical Research Council Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge,
SHORT LINEAR MOTIFS. Holger Dinkel EMBO Practical Course Computational analysis of protein-protein interactions From sequences to networks
 Holger Dinkel EMBO Practical Course Computational analysis of protein-protein interactions From sequences to networks IMPORTANCE OF 2 / 19 IMPORTANCE OF 2 / 19 IMPORTANCE OF 2 / 19 IMPORTANCE OF 2 / 19
Holger Dinkel EMBO Practical Course Computational analysis of protein-protein interactions From sequences to networks IMPORTANCE OF 2 / 19 IMPORTANCE OF 2 / 19 IMPORTANCE OF 2 / 19 IMPORTANCE OF 2 / 19
Phosphoserine Detection Kit
 Kit 0701/PSER-KIT 02/080507 Background and Specificity extracellular signals to the nucleus. Phosphorylated epitopes may serve as docking sites for the assembley of protein complexes or may alter the 3-dimensional
Kit 0701/PSER-KIT 02/080507 Background and Specificity extracellular signals to the nucleus. Phosphorylated epitopes may serve as docking sites for the assembley of protein complexes or may alter the 3-dimensional
Supplementary Materials
 Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
Supplementary Materials Supplementary Figure S1 Regulation of Ubl4A stability by its assembly partner A, The translation rate of Ubl4A is not affected in the absence of Bag6. Control, Bag6 and Ubl4A CRISPR
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Signal-Transduction Cascades - 2. The Phosphoinositide Cascade
 Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
SMART Digest Kit Facilitating perfect digestion
 Questions Answers SMART Digest Kit Facilitating perfect digestion The modern biopharmaceutical and protein research laboratory is tasked with providing high quality analytical results, often in high-throughput,
Questions Answers SMART Digest Kit Facilitating perfect digestion The modern biopharmaceutical and protein research laboratory is tasked with providing high quality analytical results, often in high-throughput,
Tryptic digestion of ubiquitin standards reveals an improved strategy for identifying ubiquitinated proteins by mass spectrometry
 868 DOI 10.1002/pmic.200600410 Proteomics 2007, 7, 868 874 RESEARCH ARTICLE Tryptic digestion of ubiquitin standards reveals an improved strategy for identifying ubiquitinated proteins by mass spectrometry
868 DOI 10.1002/pmic.200600410 Proteomics 2007, 7, 868 874 RESEARCH ARTICLE Tryptic digestion of ubiquitin standards reveals an improved strategy for identifying ubiquitinated proteins by mass spectrometry
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
UBIQUITIN AT FOX CHASE
 UBIQUITIN AT FOX CHASE Nobel Lecture, December 8, 2004 by Irwin Rose University of California, Department of Physiology and Biophysics, College of Medicine, Irvine, CA 92697 4560, USA. My interest in protein
UBIQUITIN AT FOX CHASE Nobel Lecture, December 8, 2004 by Irwin Rose University of California, Department of Physiology and Biophysics, College of Medicine, Irvine, CA 92697 4560, USA. My interest in protein
supplementary information
 DOI: 10.1038/ncb2153 Figure S1 Ectopic expression of HAUSP up-regulates REST protein. (a) Immunoblotting showed that ectopic expression of HAUSP increased REST protein levels in ENStemA NPCs. (b) Immunofluorescent
DOI: 10.1038/ncb2153 Figure S1 Ectopic expression of HAUSP up-regulates REST protein. (a) Immunoblotting showed that ectopic expression of HAUSP increased REST protein levels in ENStemA NPCs. (b) Immunofluorescent
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Protocol. Signal-Seeker TM Ubiquitination Detection Kit. 30 Assays. Cat. # BK161. cytoskeleton.com. Cytoskeleton, Inc.
 The Protein Experts Protocol Cytoskeleton, Inc. V. 22.0 Signal-Seeker TM Ubiquitination Detection Kit 30 Assays Cat. # BK161 cytoskeleton.com Phone: (303) 322.2254 Fax: (303) 322.2257 Customer Service:
The Protein Experts Protocol Cytoskeleton, Inc. V. 22.0 Signal-Seeker TM Ubiquitination Detection Kit 30 Assays Cat. # BK161 cytoskeleton.com Phone: (303) 322.2254 Fax: (303) 322.2257 Customer Service:
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry
 Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
The Ubiquitin E3 Ligase RAUL Negatively Regulates Type I Interferon through Ubiquitination of the Transcription Factors IRF7 and IRF3
 Article The Ubiquitin E3 Ligase Negatively Regulates Type I Interferon through Ubiquitination of the Transcription Factors and Yanxing Yu 1,2, * and Gary S. Hayward 1, * 1 Viral Oncology Program, The Sidney
Article The Ubiquitin E3 Ligase Negatively Regulates Type I Interferon through Ubiquitination of the Transcription Factors and Yanxing Yu 1,2, * and Gary S. Hayward 1, * 1 Viral Oncology Program, The Sidney
Proteins: Proteomics & Protein-Protein Interactions Part I
 Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
S1a S1b S1c. S1d. S1f S1g S1h SUPPLEMENTARY FIGURE 1. - si sc Il17rd Il17ra bp. rig/s IL-17RD (ng) -100 IL-17RD
 SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
SUPPLEMENTARY FIGURE 1 0 20 50 80 100 IL-17RD (ng) S1a S1b S1c IL-17RD β-actin kda S1d - si sc Il17rd Il17ra rig/s15-574 - 458-361 bp S1f S1g S1h S1i S1j Supplementary Figure 1. Knockdown of IL-17RD enhances
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
SUPPLEMENTARY INFORMATION
 a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
AA s are the building blocks of proteins
 Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
Chapter 6. Antigen Presentation to T lymphocytes
 Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Stelter et al., http://www.jcb.org/cgi/content/full/jcb.201105042/dc1 S1 Figure S1. Dyn2 recruitment to edid-labeled FG domain Nups.
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Stelter et al., http://www.jcb.org/cgi/content/full/jcb.201105042/dc1 S1 Figure S1. Dyn2 recruitment to edid-labeled FG domain Nups.
MCB130 Midterm. GSI s Name:
 1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
USP10 inhibits genotoxic NF-κB activation by MCPIP1- facilitated deubiquitination of NEMO
 Manuscript EMBO-2013-84573 USP10 inhibits genotoxic NF-κB activation by MCPIP1- facilitated deubiquitination of NEMO Jixiao Niu, Yuling Shi, Jingyan Xue, Ruidong Miao, Shengping Huang, Tianyi Wang, Jiong
Manuscript EMBO-2013-84573 USP10 inhibits genotoxic NF-κB activation by MCPIP1- facilitated deubiquitination of NEMO Jixiao Niu, Yuling Shi, Jingyan Xue, Ruidong Miao, Shengping Huang, Tianyi Wang, Jiong
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Protein Modification Overview DEFINITION The modification of selected residues in a protein and not as a component of synthesis
 Lecture Four: Protein Modification & Cleavage [Based on Chapters 2, 9, 10 & 11 Berg, Tymoczko & Stryer] (Figures in red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Protein Modification
Lecture Four: Protein Modification & Cleavage [Based on Chapters 2, 9, 10 & 11 Berg, Tymoczko & Stryer] (Figures in red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Protein Modification
Post-translational modifications of proteins in gene regulation under hypoxic conditions
 203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
William C. Comb, Jessica E. Hutti, Patricia Cogswell, Lewis C. Cantley, and Albert S. Baldwin
 Molecular Cell, Volume 45 Supplemental Information p85 SH2 Domain Phosphorylation by IKK Promotes Feedback Inhibition of PI3K and Akt in Response to Cellular Starvation William C. Comb, Jessica E. Hutti,
Molecular Cell, Volume 45 Supplemental Information p85 SH2 Domain Phosphorylation by IKK Promotes Feedback Inhibition of PI3K and Akt in Response to Cellular Starvation William C. Comb, Jessica E. Hutti,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
Antigen Recognition by T cells
 Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
supplementary information
 DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
Supplementary Material
 Supplementary Material HLA-DM Captures Partially Empty HLA-DR Molecules for Catalyzed Peptide Removal Anne-Kathrin Anders, Melissa J. Call, Monika-Sarah E. D. Schulze, Kevin D. Fowler, David A. Schuert,
Supplementary Material HLA-DM Captures Partially Empty HLA-DR Molecules for Catalyzed Peptide Removal Anne-Kathrin Anders, Melissa J. Call, Monika-Sarah E. D. Schulze, Kevin D. Fowler, David A. Schuert,
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3076 Supplementary Figure 1 btrcp targets Cep68 for degradation during mitosis. a) Cep68 immunofluorescence in interphase and metaphase. U-2OS cells were transfected with control sirna
DOI: 10.1038/ncb3076 Supplementary Figure 1 btrcp targets Cep68 for degradation during mitosis. a) Cep68 immunofluorescence in interphase and metaphase. U-2OS cells were transfected with control sirna
AlphaScreen : A Straightforward and Powerful Alternative to ELISA. Martina Bielefeld-Sévigny Ph.D., R&D Director
 AlphaScreen : A Straightforward and Powerful Alternative to ELISA Martina Bielefeld-Sévigny Ph.D., R&D Director Overview AlphaScreen - an alternative to ELISA Why an alternative to ELISA? Assay principle
AlphaScreen : A Straightforward and Powerful Alternative to ELISA Martina Bielefeld-Sévigny Ph.D., R&D Director Overview AlphaScreen - an alternative to ELISA Why an alternative to ELISA? Assay principle
Introduction Session 1 and 2 (Ubiquitin, proteasome and human disease)
 Introduction Session 1 and 2 (Ubiquitin, proteasome and human disease) Courtesy of Sam Griffiths-Jones. Used with permission. Source: "Peptide models for protein beta-sheets." PhD thesis, University of
Introduction Session 1 and 2 (Ubiquitin, proteasome and human disease) Courtesy of Sam Griffiths-Jones. Used with permission. Source: "Peptide models for protein beta-sheets." PhD thesis, University of
Supplementary Figure 1
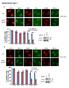 Supplementary Figure 1 a γ-h2ax MDC1 RNF8 FK2 BRCA1 U2OS Cells sgrna-1 ** 60 sgrna 40 20 0 % positive Cells (>5 foci per cell) b ** 80 sgrna sgrna γ-h2ax MDC1 γ-h2ax RNF8 FK2 MDC1 BRCA1 RNF8 FK2 BRCA1
Supplementary Figure 1 a γ-h2ax MDC1 RNF8 FK2 BRCA1 U2OS Cells sgrna-1 ** 60 sgrna 40 20 0 % positive Cells (>5 foci per cell) b ** 80 sgrna sgrna γ-h2ax MDC1 γ-h2ax RNF8 FK2 MDC1 BRCA1 RNF8 FK2 BRCA1
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
A. List of selected proteins with high SILAC (H/L) ratios identified in mass
 Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary material Figure S1. Interaction between UBL5 and FANCI A. List of selected proteins with high SILAC (H/L) ratios identified in mass spectrometry (MS)-based analysis of UBL5-interacting proteins,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
Biological Sciences 4087 Exam I 9/20/11
 Name: Biological Sciences 4087 Exam I 9/20/11 Total: 100 points Be sure to include units where appropriate. Show all calculations. There are 5 pages and 11 questions. 1.(20pts)A. If ph = 4.6, [H + ] =
Name: Biological Sciences 4087 Exam I 9/20/11 Total: 100 points Be sure to include units where appropriate. Show all calculations. There are 5 pages and 11 questions. 1.(20pts)A. If ph = 4.6, [H + ] =
Schwarz et al. Activity-Dependent Ubiquitination of GluA1 Mediates a Distinct AMPAR Endocytosis
 Schwarz et al Activity-Dependent Ubiquitination of GluA1 Mediates a Distinct AMPAR Endocytosis and Sorting Pathway Supplemental Data Supplemental Fie 1: AMPARs undergo activity-mediated ubiquitination
Schwarz et al Activity-Dependent Ubiquitination of GluA1 Mediates a Distinct AMPAR Endocytosis and Sorting Pathway Supplemental Data Supplemental Fie 1: AMPARs undergo activity-mediated ubiquitination
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature05732 SUPPLEMENTARY INFORMATION Supplemental Data Supplement Figure Legends Figure S1. RIG-I 2CARD undergo robust ubiquitination a, (top) At 48 h posttransfection with a GST, GST-RIG-I-2CARD
doi: 10.1038/nature05732 SUPPLEMENTARY INFORMATION Supplemental Data Supplement Figure Legends Figure S1. RIG-I 2CARD undergo robust ubiquitination a, (top) At 48 h posttransfection with a GST, GST-RIG-I-2CARD
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.10/nature10195 NCBI gene: Tagged Subunit(s: HA-Vpx; FLAG-Cul4 HA-DCAF1 FLAG-Cul4 HA-FLAG-Vpx Mock Vpx (SIVmac 100 (a ; 0.159 (b ; 0.05 DCAF1 DDB1 DDA1 Cul4A 1; 0.024591
SUPPLEMENTARY INFORMATION doi:10.10/nature10195 NCBI gene: Tagged Subunit(s: HA-Vpx; FLAG-Cul4 HA-DCAF1 FLAG-Cul4 HA-FLAG-Vpx Mock Vpx (SIVmac 100 (a ; 0.159 (b ; 0.05 DCAF1 DDB1 DDA1 Cul4A 1; 0.024591
