Eukaryotic transcription (III)
|
|
|
- Reynold Warner
- 5 years ago
- Views:
Transcription
1 Eukaryotic transcription (III)
2 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids or chromosome during cell division chromosomes
3 Euchromatin and heterochromatin Euchromatin: less condensed chromatin, light color under microscope, actively transcribed genomic regions Heterochromatin: highly condensed chromatin regions, such as centromeres and telomeres, which are not actively transcribed.
4 How chromatin structure affects transcription activity of genes? The degree of chromatin condensation (i.e. how tightly is chromatin packed) affects transcription. Usually the less condensed euchromatins are active because they are transcribed more readily than the highly condensed heterochromatins ( silent ).
5 Nucleosomes are the basic structural units of chromatins 10 nm kb per loop
6 Nucleosome and histones Nucleosomes are DNA-histone particles. Histones are basic proteins rich in Arg and Lys, they bind tight to negatively charged DNA. Nucleosome consists of 147 bp DNA wrapped around 8 core histone molecules. Core histones are H2A (~13 kd) H2B (~14 kd) H3 (~15 kd), H4 (~11 kd). Nucleosomes are linked by about 50bp linker DNA and linker histones H1 (~21kD). Histones are highly conserved.
7 Summary of chromatin structure Linker histone H1 (10 nm fiber) Nucleosomes composed of core histones 2x(H2A, H2B,H3, H4) 147bp DNA
8 2. Nucleosome and transcription 1). The position of nucleosomes is affected by the number of linker histone H1, histone modification, and chromosome remodeling proteins. 2). An active promoter is usually free of nucleosomes 3). Nucleosome localization is dynamic (nucleosome "moves"), the location and density of nucleosomes can be changed by the histone modification and chromatin remodeling process. 4). Histone modifying enzymes and chromatin remodeling proteins play critical roles in chromatin remodeling to affect transcription.
9 (1) Linker histone (H1) may inhibit transcription Nucleosomes are connected by histone H1, also called linker histone. H1 often represses transcription. H1 may repress transcription by simply occupying the position of DNA that is normally accessible to transcription activators. nucleosome H1
10 (2) Active promoter is free of nucleosomes SV40 minichromosome uncut BamHI or Bgl I uncut SV40 minichromosom e Experiment: 1. BglI cuts a circular SV40 viral minichromosome within the promoter and enhancer (P/E) region. 2. BamHI cuts at a site opposite to the P/E region. 3. The BamHI-cut SV40 DNA shows a nucleosome-free DNA (arrow) in the center, BglI-cut SV40 shows nucleosome-free DNA at the end. Conclusion: promoter/enhancer DNA (P/E) must be free of the nucleosome BamHI BglI
11 (3). Chromatin remodeling a). Chromatin remodeling (or chromosome remodeling) is an ATPdependent process that changes the chromatin architecture. Chromatin remodeling is accomplished by changes of the position and/or density of core nucleosomes. b). chromatin remodeling proteins or chromatin remodeler, such as SWI/SNF, SWR, ISWI, INO80, NuRD, etc consume ATP to change chromatin architecture. c). Chromatin remodeling can be accomplished by change of core histones or histone modifications. d). Chromatin remodeling may activate or repress transcription.
12 Mechanisms of chromatin remodeling Chromatin remodeling complexes (CRC) use hydrolyze ATP to change the nucleosomal position or density by at least 4 mechanisms. 1) Mobilization of nucleosome position (sliding), 2) Dissociation of DNA-histone contact (unwrapping), 3) Remove core histones (histone eviction). 4) Replacement of the common histone unit (exchange of histone variants), such as change from H2A to H2A.Z.
13 chromatin remodeling by replacing core histones (1) Nucleosomes are tightly packed by nucleosomes composed of the common H2A, H2B, H3, and H4. (2-3) A remodeler protein (pink) facilitates reposition the nucleosome to establish a nucleosome-free region (red). (4) Another remodeler protein (brown) facilitates replacement of the common core histone H2A (green) with the histone variant H2A.Z (dark pink) into nucleosomes flanking the nucleosome-free region. (5) The chromatin now possesses a stable nucleosome-free region that allows for productive transcription.
14 Histone modifications facilitates chromatin remodeling
15 2. Histone modifications (1) N-terminal of about amino acids of core histone (H2A, H2B, H4, H4) are called histone tails. Histone tails are not buried inside a core nucleosome, and often chemically modified. (2) The same residue of a histone may be modified by different chemical reactions under different conditions. (3) The same chemical modification in different residues of a histone may have different effect on transcription. (4) The combined pattern of histone modifications is referred to as histone codes. e.g.h3r2mk4mk9ack14ac This code means a histone H3 with Arg2 methylated, Lys4 methylated, Lys9 acetylated, Lys 14 acetylated, (5) Histone code is important to transcription
16 Four major histone modifications Acetyl- Methyl- Phosphoryl- ubiquitin Acetylation: addition of acetyl group to lysine residues. Acetylated histones have less positive charges and interact poorly with DNA. Histones in euchromatin are highly acetylated (hyperacetylation), histones of heterochromatin are rarely acetylated (hypoacetylation) Methylation: it often occurs to the same Lys residue as acetylation, so histone methylation competes with acetylation. It also occurs to Arg. Phosphorylation: to introduce negative charges Ubiquitination: Although poly-ubiquitination is usually a degradation signal of the ubiquitinated proteins, mono-ubiquitination (add only one ubq) is not a degradation signal. Mono-ubiquitination is part of histone code.
17 (1) Histone acetylation and deacetylation HAT (Acetyl-CoA) HDAC 1. Histones can be acetylated. HAT (histone acetyltransferase) transfers acetyl groups from the donor acetyl-coa to the lysine residues of histones. Only lysine (but not arginine) of histones are acetylated. Acetylated histones can be deacetylated by HDAC (histone deacetylase) that removes the acetyl group from histones. 2. Lysine acetylation neutralizes positive charges of histone. Histone acetylation tend to reduce the interactions between the neighboring nucleosomes, decreasing the overall chromatin condensation. Therefore, histone acetylation usually (not always) activates transcription.
18 activator to repressor conversion Transcription co-activators often recruit HAT to activate transcription. Transcription corepressors often recruit HDAC to repress transcription. When Max binds Myc, the Max-Myc dimer recruits HAT (CBP and GCN5) to acetylate histones, which activates transcription. When Max binds to Mad, the Max-Mad dimer recruits HDAC to deacetylate histone, resulting in suppression of transcription.
19 Experiment 1. Transform cells with a plasmid expressing FLAG-tagged HDAC2, or FLAG only (control), with or without the second plasmid expressing Mad1 or a mutant Mad1 (Mad1Pro). V= cells transformed with vector only. 2. Immunoprecipitate (IP) with anti-flag antibody, run IP in SDS-PAGE 3. Immunoblot with anti-sin3 (=msin3a) antibody first, strip the membrane and re-probe with the anti-mad1 antibody Result: 1. IP from the control (FLAG-only) cells pull down nothing. 2. IP from FLAG- HDAC2 cells pull down Sin3 regardless of Mad1. 3. IP of FLAG- HDAC2 cell also pull down Mad1, but not mutant Mad1 (Mad1Pro).
20 It was previously known that: - Sin3 can interact with HDAC2 but not DNA, so it is a co-repressor - Mad1 is a transcription repressor interacting with DNA and Sin3, - mutant Mad1Pro does not interact with Sin3 or inhibit transcription. Question: How does Mad1 repress transcription? Reasoning: It was found by the co-ip experiment that: 1. FLAG alone did not pull down Mad1 or Sin3; 2. FLAG-HDAC2 pulled down Sin3 in all cells (as previously known); 3. FLAG-HDAC2 pulled down Mad1; 4. FLAG-HDAC2 did not pull down mutant Mad1Pro. Conclusion: Mad1 must interact with the Sin3-HDAC2 complex to repress transcription. When Max binds to Mad1, the Max-Mad1 dimer recruits HDAC2, via the Sin3 bridge. HDAC2 deacetylates histones around the promoter region, resulting in inhibition of transcription. Sin3 HDAC2
21 Repressor to activator conversion Without thyroid hormone (TH), the thyroid receptor (TR) binds co-repressor NcoR to recruit Sin3, which binds HDAC to deacetylate histone and repress transcription With thyroid (TH), TR binds to the coactivator CBP-CAF-TAF250, which is a HAT complex that acetylates histone to activate transcription
22 Using ChIP (chromatin-imunoprecipitation) to identify chromatin DNA that is bound by a protein in vivo 1. Isolate total chromatins, crosslink chromatins to fix DNA-protein interaction 2. Cut chromatins to smaller fragments (e.g. about 500bp) by ultrasonication or mild nuclease digestion 3. Immunoprecipitation using antibodies against the transcription factor or modified histones (such as acetylated histone). The chromatin fragments bound to the protein are precipitated. 4. PCR amplification of DNA of the promoter of interest. The DNA of chromatin fragments bound by the protein (recognized by the antibody) should produce a PCR product.
23 Histone acetylation occur before transcription starts Experiment: 1. Infect cells with virus. 2. Isolate chromatin at different time after infection. 3. ChIP by antibodies against TBP or acetylated histone (e.g. α-ach4 K5, K8 is the antibody against histone H4 acetylated at the Lys5 or Lys8). 4. PCR the promoter DNA of the IFN- gene (encoding an interferon involved in immune responses) to examine how TBP or modified histones bind to the IFN- promoter DNA. Results: Acetylation of histones on the IFN- promoter occurred after viral infection but before TBP binding to the promoter and IFN- mrna expression.
24 Histone modification may also affect DNA-TBP interaction HDAC This model shows that the transcription repressor Mad/Max recruits HDAC to deacetylate histone and suppress transcription. In contrast, TAF250 (HAT) acetylate histone to facilitate TBP-DNA interaction between TATA box and TBP that is associated with TAF250. The ultimate level of transcription is dependent on the balance between HDAC and HAT.
25 (2) Histone methylation Histone can be methylated at both lysine and arginine residues. Methyltransferases (HMT) transfer (one to three) methyl groups from the donor SAM (s-adenosylmethionine) to lysine or arginine residues of histone tails. Methylation does not reduce the positive charges of lysine (or arginine), so it does not reduces nucleosome condensation. Instead, histone methylation often causes chromatin condensation and transcription inactivation.
26 Histone methylation may cause chromatin condensation to repress transcription If H3K9 is methylated, it binds to the chromosome remodeling protein HP1. HP1 recruits HMT, HMT methylates H3K9 in the next nucleosome, HP1 binds methylated histone to spread histone methylation and chromatin condensation to form heterochromatin. Heterochromatin cause gene silencing unless the gene is insulated from heterochromatin spreading by insulators.
27 (3) histone codes are dynamic and interactive Modification of one histone may affect other histones, different histone modifications have different effects on transcription. For example, H3S10 phosphorylation activates H3K14 acetylation, which inhibits H3K9 methylation and activates transcription. But if H3K9 is methylated first, it inhibits H3S10 phosphorylation and H3K14 acetylation, resulting in transcription repression.
28 A comparison Prokaryote RNAP II Eukaryote
29 Prokaryotic transcription: One RNA pol, holoenzyme of RNA pol =,,,ω + sigma factor Closed complex = holoenzyme + promoter DNA Promoters often contain TATA box at -10 position DNA regulatory elements (e.g. CAP-binding sites, operators, etc) are usually close to or within the promoter or the transcribed region Regulated by a few transcription factors, also regulated by changing of sigma factors, or even RNA polymerase (phage only) No histone, no DNA methylation, no complex chromatin structures Eukaryotic transcription: Three RNA polymerases, RNA pol II transcribes mrnas Holoenzyme of RNA pol II: more subunits, also need TFII s Preinitiation complex = holoenzyme + TFIID + promoter DNA Promoter often contains TATA box at -25 position DNA regulatory elements (e.g. enhancers and silencers) may be close to, within, or far away from the promoter or the transcribed region Regulated by many transcription factors, co-activator, and corepressors, also regulated by nucleosome position/density, histone codes, and other aspects of the chromatin structure.
Chromatin Structure & Gene activity part 2
 Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Lecture 10. Eukaryotic gene regulation: chromatin remodelling
 Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Transcription and chromatin. General Transcription Factors + Promoter-specific factors + Co-activators
 Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Gene Expression DNA RNA. Protein. Metabolites, stress, environment
 Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
On the way of revealing coactivator complexes cross talk during transcriptional activation
 DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
9/3/2009 DNA i DNA n euk euk yotes Organizatio Organ izatio n of o f gen ge e n tic Locati t on: In n ucleu e s material mater in e ial
 DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
Host cell activation
 Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
Control of nuclear receptor function by local chromatin structure
 MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
Organization of genetic material in eukaryotes
 Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS
 R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Epigenetics: The Future of Psychology & Neuroscience. Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1
 Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling
 Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Structure and function of active chromatin and DNase I hypersensitive sites
 MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
Where Splicing Joins Chromatin And Transcription. 9/11/2012 Dario Balestra
 Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Are you the way you are because of the
 EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
Chapter 19 Eukaryotic Genomes
 Chapter 19 Eukaryotic Genomes Lecture Outline Overview: How Eukaryotic Genomes Work and Evolve Two features of eukaryotic genomes present a major information-processing challenge. First, the typical multicellular
Chapter 19 Eukaryotic Genomes Lecture Outline Overview: How Eukaryotic Genomes Work and Evolve Two features of eukaryotic genomes present a major information-processing challenge. First, the typical multicellular
Protein methylation CH 3
 Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
How Cells Divide. Chapter 10
 How Cells Divide Chapter 10 Bacterial Cell Division Bacteria divide by binary fission. -the single, circular bacterial chromosome is replicated -replication begins at the origin of replication and proceeds
How Cells Divide Chapter 10 Bacterial Cell Division Bacteria divide by binary fission. -the single, circular bacterial chromosome is replicated -replication begins at the origin of replication and proceeds
Repression of IP-10 by Interactions between Histone Deacetylation and Hypermethylation in Idiopathic Pulmonary Fibrosis
 MOLECULAR AND CELLULAR BIOLOGY, June 2010, p. 2874 2886 Vol. 30, No. 12 0270-7306/10/$12.00 doi:10.1128/mcb.01527-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Repression of
MOLECULAR AND CELLULAR BIOLOGY, June 2010, p. 2874 2886 Vol. 30, No. 12 0270-7306/10/$12.00 doi:10.1128/mcb.01527-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Repression of
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Stacie Bulfer Proteins Research Proposal April 27, Background
 Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Chapter 11 How Genes Are Controlled
 Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Chapter 11 How Genes Are Controlled PowerPoint Lectures for Biology: Concepts & Connections, Sixth Edition Campbell, Reece, Taylor, Simon, and Dickey Copyright 2009 Pearson Education, Inc. Lecture by Mary
Histone octamer assembly
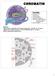 CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
Watson-Crick Model of B-DNA
 Reading: Ch8; 285-290 Ch24; 963-978 Problems: Ch8 (text); 9 Ch8 (study-guide: facts); 3 Ch24 (text); 5,7,9,10,14,16 Ch24 (study-guide: applying); 1 Ch24 (study-guide: facts); 1,2,4 NEXT Reading: Ch1; 29-34
Reading: Ch8; 285-290 Ch24; 963-978 Problems: Ch8 (text); 9 Ch8 (study-guide: facts); 3 Ch24 (text); 5,7,9,10,14,16 Ch24 (study-guide: applying); 1 Ch24 (study-guide: facts); 1,2,4 NEXT Reading: Ch1; 29-34
U3.2.3: Eukaryotic chromosomes are linear DNA molecules associated with histone proteins. (Oxford Biology Course Companion page 151).
 Cell Division Study Guide U3.2.3: Eukaryotic chromosomes are linear DNA molecules associated with histone proteins. (Oxford Biology Course Companion page 151). 1. Describe the structure of eukaryotic DNA
Cell Division Study Guide U3.2.3: Eukaryotic chromosomes are linear DNA molecules associated with histone proteins. (Oxford Biology Course Companion page 151). 1. Describe the structure of eukaryotic DNA
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
The Role of Protein Domains in Cell Signaling
 The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
 EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
The Process of Cell Division
 Lesson Overview 10.2 The Process of Cell Division THINK ABOUT IT What role does cell division play in your life? Does cell division stop when you are finished growing? Chromosomes What is the role of chromosomes
Lesson Overview 10.2 The Process of Cell Division THINK ABOUT IT What role does cell division play in your life? Does cell division stop when you are finished growing? Chromosomes What is the role of chromosomes
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Transcriptional and Epigenetic Mechanisms of Addiction
 Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome
 The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes
 Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Gene Regulation - 4. One view of the Lactose Operon
 Gene Regulation - 1 Regulating Genes We have been discussing the structure of DNA and that the information stored in DNA is used to direct protein synthesis. We've studied how RNA molecules are used to
Gene Regulation - 1 Regulating Genes We have been discussing the structure of DNA and that the information stored in DNA is used to direct protein synthesis. We've studied how RNA molecules are used to
The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation gene PHR1
 Published online 3 February 2011 Nucleic Acids Research, 2011, Vol. 39, No. 10 4151 4165 doi:10.1093/nar/gkr040 The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation
Published online 3 February 2011 Nucleic Acids Research, 2011, Vol. 39, No. 10 4151 4165 doi:10.1093/nar/gkr040 The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation
BIO 5099: Molecular Biology for Computer Scientists (et al)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential Expression of Genes Prokaryotes and eukaryotes precisely regulate gene expression in response to environmental conditions In multicellular eukaryotes,
Chapter 18 Regulation of Gene Expression Differential Expression of Genes Prokaryotes and eukaryotes precisely regulate gene expression in response to environmental conditions In multicellular eukaryotes,
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Transcription Through Chromatin - Dynamic Organization of Genes
 GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
Regulation of Gene Expression
 18 Regulation of Gene Expression KEY CONCEPTS Figure 18.1 What regulates the precise pattern of gene expression in the developing wing of a fly embryo? 18.1 Bacteria often respond to environmental change
18 Regulation of Gene Expression KEY CONCEPTS Figure 18.1 What regulates the precise pattern of gene expression in the developing wing of a fly embryo? 18.1 Bacteria often respond to environmental change
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Brd4 Coactivates Transcriptional Activation of NF- B via Specific Binding to Acetylated RelA
 MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
Campbell Biology 10. A Global Approach. Chapter 18 Control of Gene Expression
 Lecture on General Biology 2 Campbell Biology 10 A Global Approach th edition Chapter 18 Control of Gene Expression Chul-Su Yang, Ph.D., chulsuyang@hanyang.ac.kr Infection Biology Lab., Dept. of Molecular
Lecture on General Biology 2 Campbell Biology 10 A Global Approach th edition Chapter 18 Control of Gene Expression Chul-Su Yang, Ph.D., chulsuyang@hanyang.ac.kr Infection Biology Lab., Dept. of Molecular
RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh-
 1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
1 a b Supplementary Figure 1. Effects of GSK3b knockdown on poly I:C-induced cytokine production. RAW264.7 cells stably expressing control shrna (Con) or GSK3b-specific shrna (sh- GSK3b) were stimulated
Gene Regulation Part 2
 Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
'''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Five'Levels'of'Organiza-on' Molecular'
 '''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Anggraini'Barlian,' Iriawa-' Tjandra'Anggraeni' SITH4ITB' Five'Levels'of'Organiza-on' Molecular'
'''''''''''''''''Fundamental'Biology' BI'1101' ' an'interdisciplinary'approach'to'introductory'biology' Anggraini'Barlian,' Iriawa-' Tjandra'Anggraeni' SITH4ITB' Five'Levels'of'Organiza-on' Molecular'
Chromatin Remodeling: A Novel Mechanism. of Psychotropic Drug Action
 Molecular Pharmacology This article has Fast not been Forward. copyedited Published and formatted. The on final May version 25, 2006 may differ as doi:10.1124/mol.106.027078 from this version. Chromatin
Molecular Pharmacology This article has Fast not been Forward. copyedited Published and formatted. The on final May version 25, 2006 may differ as doi:10.1124/mol.106.027078 from this version. Chromatin
Post-translational modifications of proteins in gene regulation under hypoxic conditions
 203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being A Eukaryote. Eukaryotic Cells. Basic eukaryotes have:
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription
 9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription Friday, 20 th of November 2015, 13:00h Institute of Molecular Biology and Tumor Research (IMT) Philipps-University
9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription Friday, 20 th of November 2015, 13:00h Institute of Molecular Biology and Tumor Research (IMT) Philipps-University
Cellular Reproduction, Part 1: Mitosis Lecture 10 Fall 2008
 Cell Theory 1 Cellular Reproduction, Part 1: Mitosis Lecture 10 Fall 2008 Cell theory: All organisms are made of cells All cells arise from preexisting cells How do new cells arise? Cell division the reproduction
Cell Theory 1 Cellular Reproduction, Part 1: Mitosis Lecture 10 Fall 2008 Cell theory: All organisms are made of cells All cells arise from preexisting cells How do new cells arise? Cell division the reproduction
Nuclear receptors. Lecture 4. Never work alone: corepressors and coactivators
 Nuclear receptors Lecture 4 Never work alone: corepressors and coactivators General transcription factors Eucaryotic RNA polymerases cannot initiate transcription on their own; they require a set of proteins
Nuclear receptors Lecture 4 Never work alone: corepressors and coactivators General transcription factors Eucaryotic RNA polymerases cannot initiate transcription on their own; they require a set of proteins
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson
 Thesis for doctoral degree (Ph.D.) 2009 Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson Characterization of Gcn5 Histone
Thesis for doctoral degree (Ph.D.) 2009 Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson Characterization of Gcn5 Histone
Chapter 8 The Cell Cycle
 What molecule stores your genetic information or determines everything about you? DNA a nucleic acid How are DNA molecules arranged in the nucleus? As you can see DNA is: Chapter 8 The Cell Cycle 1. Arranged
What molecule stores your genetic information or determines everything about you? DNA a nucleic acid How are DNA molecules arranged in the nucleus? As you can see DNA is: Chapter 8 The Cell Cycle 1. Arranged
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Enzymes Part III: regulation II. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Epigenetics and Environmental Health A Step-by-Step Tutorial
 Powerful ideas for a healthier world Epigenetics and Environmental Health A Step-by-Step Tutorial Andrea Baccarelli, MD, PhD, MPH Laboratory of Environmental Epigenetics Objective of my presentation To
Powerful ideas for a healthier world Epigenetics and Environmental Health A Step-by-Step Tutorial Andrea Baccarelli, MD, PhD, MPH Laboratory of Environmental Epigenetics Objective of my presentation To
Chapter 10. 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation)
 Chapter 10 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation) 1 Fueling Processes Respiration 1 Most respiration involves use of an electron transport chain As electrons pass through the electron
Chapter 10 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation) 1 Fueling Processes Respiration 1 Most respiration involves use of an electron transport chain As electrons pass through the electron
Transcriptional repression is epigenetically marked by H3K9 methylation during SV40 replication
 Kallestad et al. Clinical Epigenetics 2014, 6:21 RESEARCH Open Access Transcriptional repression is epigenetically marked by H3K9 methylation during SV40 replication Les Kallestad, Kendra Christensen,
Kallestad et al. Clinical Epigenetics 2014, 6:21 RESEARCH Open Access Transcriptional repression is epigenetically marked by H3K9 methylation during SV40 replication Les Kallestad, Kendra Christensen,
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis
 Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis Prokaryotes Have a Simpler Cell Cycle Cell division in prokaryotes takes place in two stages, which together make up a simple cell cycle 1. Copy
Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis Prokaryotes Have a Simpler Cell Cycle Cell division in prokaryotes takes place in two stages, which together make up a simple cell cycle 1. Copy
5.2. Mitosis and Cytokinesis. Chromosomes condense at the start of mitosis. Connecting
 5.2 Mitosis and Cytokinesis KEY CONCEPT Cells divide during mitosis and cytokinesis. MAIN IDEAS Chromosomes condense at the start of mitosis. Mitosis and cytokinesis produce two genetically identical daughter
5.2 Mitosis and Cytokinesis KEY CONCEPT Cells divide during mitosis and cytokinesis. MAIN IDEAS Chromosomes condense at the start of mitosis. Mitosis and cytokinesis produce two genetically identical daughter
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis
 Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis Chromosomes Chromosomes were first observed by the German embryologist Walther Fleming in 1882. Chromosome number varies among organisms most
Breaking Up is Hard to Do (At Least in Eukaryotes) Mitosis Chromosomes Chromosomes were first observed by the German embryologist Walther Fleming in 1882. Chromosome number varies among organisms most
Phenomena first observed in petunia
 Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
Vectors for RNAi Phenomena first observed in petunia Attempted to overexpress chalone synthase (anthrocyanin pigment gene) in petunia. (trying to darken flower color) Caused the loss of pigment. Bill Douherty
