Histone octamer assembly
|
|
|
- Gary Jefferson
- 6 years ago
- Views:
Transcription
1 CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity References: -Special issue on Epigenetic and Chromatin Organization. Cell 2007 vol. 128 issue 4 -Special issue Focus on Chromatin. Nat Struct Mol Biol 2007 vol. 14 issue 11 Contact: jacques.cote@crhdq.ulaval.ca 1
2 Chromatin Domains 2
3 H1 Linker histone H2A H2B Core histones H3 H4 N helix variable conserved HISTONES are highly conserved, small, basic proteins Histone acetylation is a reversible modification of lysines in the N-termini of the core histones. Result: reduced binding to DNA destabilization of chromatin Histone octamer assembly H3-H4 tetramer Histone octamer H2A-H2B dimer 3
4 Where are the N-termini of the core histones? Luger, Mader, Richmond, Sargent & Richmond Nature 389, (1997) Question 1: What is the function of histone N-termini? Question 2: Are all N-termini functionally equivalent? 4
5 5
6 Factors regulating local chromatin/ accessibility to DNA -Histone modifying enzymes (HAT, HDAC, HMT, HDM, kinase) -ATP-dependent chromatin remodelers (SWI/SNF, BRG1/BRM, NuRD, NURF) -Histone variants (H2A.Z, H2A.X, CENP-A, macro-h2a) -Histone chaperones (ASF1, NAP1, FACT, CAF1) Acetylation of conserved lysines The N-termini of histones H4 and H3, and their acetylation patterns, are absolutely conserved. H4 N-terminus H3 N-terminus Ac Ac Ac Ac Ac or Me Ac-S-G-R-G-K-G-G-K-G-L-G-K-G-G-A-K-R-H-R-K-V-L-R-D Ac or Me Me Ac Ac Ac Ac A-R-T-K-Q-T-A-R-K-S-T-G-G-K-A-P-R-K-Q-L-A-T-K-A-A-R-K-S-A-P Acetyl-CoA Lysine HAT (Histone Acetyl-Transferase) CoA ε-n-acetyl-lysine O O N C C γ C C α β δ ε O O C C N+ - P P O O reversible reactions N C C C C C ε C N C O Histone Deacetylase C DNA backbone binding no DNA binding P
7 Chromatin fibers 30 nm chromatin fiber 11 nm (beads) + charged N termini (bind DNA and neigboring nucleosomes) highly acetylated core histones (especially H3 and H4) HIGH level of histone H1 NO gene transcription Reduced level of histone H1 Gene transcription possible Acetylation of Chromatin Domains Example: the chicken β-globin gene domain domain boundary DNA remaining kb β ρ β H β A β ε domain boundary α(ac-lys) antibody nucleosome ppt DNA loop domain β = β-globin genes: DNase I hyper-sensitive site General DNase I sensitivity Control: inactive gene chicken ovalbumin DNase I High levels of chromatin acetylation, across complete chromatin domains (DNA loops), induces chromatin changes detected as general DNase I sensitivity Within these chromatin domains, at functional genes or transcription factors, the chromatin structure is interrupted by small DNase I hypersensitive sites DNase I (U/ml) Hebbes, Clayton, Thorne, Crane-Robinson EMBO J. 13, 1823 (1994) 7
8 DOSAGE COMPENSATION MECHANISMS SPECIFIC ACETYLATION VS TRANSCRIPTION Drosophila Human Both X chromosomes are normaly expressed. The X chromosome is two-fold hypertranscribed K16 of histone H4 is hyperacetylated on the X chromosome. Only one X chromosome is transcribed The other one is hypoacetylated on all 4 sites of H4 The X chromosome is normaly expressed and acetylated. Acetylation at Promoters Transcriptional ACTIVATION TAF 250 II TBP HAT TATA ACGGTC ADA2 ADA3 GCN5 HAT pol.ii TH TH TACCCG p300 CPB HAT P/CAF + hormone HAT TR/RXR Thyroid Hormone Receptor example of DNA-binding Transcription Factor co-activator Histone Acetyl Transferases TAF 250 II TBP HAT Transcriptional REPRESSION Without Thyroid Hormone co-repressor TATA Histone Deacetylase TACCCG N-CoR Sin3 RPD3 Adapted from Wolffe Nature 387, 16 (1997) 8
9 S phase Gene Activation co-activator CPB HAT HAT TAF 250 II Rb co-repressor HDAC1 Histone Deacetylase Rb G2 phase Mitosis S phase G1 phase Cell cycle control: the G1-S transition (Restriction point) R cyclinecdk2 kinase Low activity HAT P + P Rb Weakly active Adapted from Ait-Si-Ali et al. Nature 396, 184 (1998) TBP TATA TAF 250 II TBP HAT S phase gene: OFF TATA Histone Acetyl Transferase CPB P HAT pol.ii Active HAT S phase gene: ON 9
10 Deacetylation of Chromatin Transcriptional Silencing X-chromosome Inactivation me 5 5..pCpGp..3 transcriptional repressor MeCP2 co-repressor Histone Deacetylase 5 me 3..pGpCp..5 º º Sin3 RPD3 l l l l 5meC CpG DNA modification is observed in repressed genes and inactivated X chromosomes 5meC CpG-methylation is maintained after DNA replication by Maintenance Methylase action on hemi-methylated DNA 5meC binds transcriptional repressor MeCP2 (MethylC-binding Protein-2) MeCP2 binds Sin3 with RPD3 histone deacetylase + + CpG methylation Hypo-acetylated repressed chromatin fiber Nan et al. Mol.Cell.Biol. 16, 414 (1996); Cell 88, 1 (1997); Jones et al. Nat.Genet. 19, 187 (1998) Sas2 Sir4 Sir4 Sir4 Sir4 Sir4 Sir4 Sir4 Sir4 Sir4 Sir2 Sir3 Sir3 Sir3 Sir3 Sir2 Sir2 Sir2 Sir2 Rap1 Heterochromatin Gene activity Euchromatin Sas2 Sir4 Sir4 Sir4 Rap1 Sir4 Sir4 Sir2 Sir2 Sir2 Sir2 Sir3 Sir3 Sir3 Sir3 Sir4 Sir4 Sir4 Sir4 Sir2 Sir2 Sir2 Sir4 Sir4 Sir4 Sir4 Sir3 Sir3 Sir3 Sir3 Sir2 Sir2 Gene repression Dot1 H3 MeK79 Heterochromatin Sir2 Sir4 Sir4 Sir4 Sir4 Sir4 Sir4 Sir3 Rap1 Gene activity Heterochromatin Euchromatin 10
11 11
12 ATP-dependent Chromatin Remodelers 12
13 13
14 14
15 G&D 13:2339(1999) 15
16 Histone Acetyltransferases 16
17 Recruitment of NuA4 in transcription activation 17
18 18
19 Phosphate rich (+Pi) -4-3 ClaI -2 UAS2-1 TATA UAS1 yng2 WT WT esa1 ts WT esa1 ts +Pi -Pi +Pi -Pi +Pi -Pi +Pi -Pi +Pi -Pi -Pi DNaseI Non Digested Digested RT 37 o C RT 37 o C RT NuA4 is required for chromatin remodeling at the PHO5 promoter p53 Element WT or Δyng2 p21-his3 + p53 WT p53 Δyng2 p53 Galactose WT Δyng2 + + p21-his3 GAL1 ACT1 25S rrna α p53 YNG2 is required for the transactivation function of p53 in yeast 19
20 WT+p21 WT+p21/p53 Δyng2+p21/p53 input αach4 αhyperach4 αach3 p53 targets NuA4 dependent histone H4 hyperacetylation to p21 promoter in yeast Cooperativity between Histone Acetylation and Chromatin Remodeling 20
21 21
22 HO URS 1 URS 2 Swi5 Swi5 URS 1 URS 2 HO Swi5 URS 1 URS 2 HO Swi/Snf Ash1 Swi5 URS 1 Swi/Snf URS 2 HO Swi5Ash1 URS 1 URS 2 HO SAGA HO URS 1 URS 2 Swi/Snf SAGA HO URS 1 URS 2 Swi6 SBF Swi4 Swi/Snf SAGA Swi6 Swi4 URS 1 URS 2 HO Transcription de HO 22
23 Bromodomain: an acetylated histone binding module 23
24 Bromodomains bind acetylated histone tails; mechanism of retention? Bromodomains bind acetylated histone tails; mechanism of retention? 24
25 A code formed by histone modifications 25
26 La modification des histones affecte la fonction de la chromatine Empreinte Épigénétique 26
27 Specific protein domains recognize different epigenetic marks (read a local code/signature?) 27
28 28
29 Mécanisme 29
30 Le code épigénétique des histones La théorie dit que différentes modifications à différents endroits et en différentes combinaisons sur les extrémités N terminales des histones formeraient des marques pour la formation d un état chromatinien transmissible particulier et pour des protéines qui les reconnaîtraient spécifiquement. 30
31 L héritage des histones parentales permet la transmission de l état chromatinien de génération en génération Tout comme pour la réplication de l ADN, la distribution des constituants des nucléosomes se fait selon un modèle SEMI-CONSERVATIF 31
32 32
33 Modulation of chromatin modifications during the cell cycle and in response to signaling pathways 33
34 Chromatin and the Transcription Cycle 34
35 Factors regulating local chromatin/ accessibility to DNA -Histone modifying enzymes (HAT, HDAC, HMT, HDM, kinase) -ATP-dependent chromatin remodelers (SWI/SNF, BRG1/BRM, NuRD, NURF) -Histone variants (H2A.Z, H2A.X, CENP-A, macro-h2a) -Histone chaperones (ASF1, NAP1, FACT, CAF1) Workman G&D
36 J. Mellor Mol. Cell 2005 Presetting of highly inducible genes by NuA4 The human NuA4/Tip60 complex is structurally equivalent to a physical merge of yeast NuA4 and SWR1 complexes Link between H4 acetylation and H2AZ-H2A exchange in chromatin? 36
37 NuA4-dependent chromatin acetylation influences H2AZ deposition on chromatin Chromatin domains along a transcription unit K4-trimethylated nucleosomes Lieb and Clarke, Cell
38 38
39 Figure 7. Model of the role of ubiquitylation and deubiquitylation at GAL1 Karl W. Henry et al. Genes Dev. 2003; 17:
40 Swr1-dependent K4-trimethylated nucleosomes (Lieb and Clarke, Cell 2005) H3 trimek36 (Set2) Rpd3 Sin3 H3 trimek36(set2) H4/H2A acetylation on promoter(nua4) H3 trimek4(set1) Histone deacetylation on coding region(rpd3) K4-trimethylated nucleosomes (Lieb and Clarke, Cell 2005) 2 HDAC complexes with opposite functions Repression at promoters Accurate transcription elongation 40
41 HBO1-JADE1 HAT complexes are enriched on the coding region of genes High resolution localization analysis (ChIP-chip) 41
42 Chromatin and Gene silencing 42
43 Modèle Modèle suggéré: La méthylation des histones induit la méthylation de l ADN. 1. Une séquence signal recrute un facteur x. 2. Facteur x recrute HDAC et HMT Induction d un état transcriptionnel inactif. 3. Facteurs y recruté sur les sites méthylés 4. DMTase recrutée par facteur y. Propagation de l état inactif 43
44 44
45 45
46 Chromatin and Genome stability 46
47 47
48 N H2A.X P C 48
49 Redon et al. JCB 1999 H2A.X is rapidly phosphorylated at sites of DNA damage A B, TIP60 (BAF53 + HAT) TIP60 (HAT + ruvb-like helicases) 49
50 Ordered recruitment of NuA4 and ATP-dependent remodelers at a DSB in vivo 50
51 Recruitment and role of the NuA4/TIP60 HAT complex at DNA double-strand breaks Chromatin and DNA Replication 51
52 52
53 Epigenetic inheritance 53
54 S-phase chromatin contains the bulk of acetylated H4 ING tumor suppressor family of proteins are critical regulators of chromatin acetylation in eukaryotes 54
55 ING5 HAT complexes interact with the MCM helicase and are essential for DNA replication Schematic representation of the assembly of the pre-replicative complex (Pre-RC) and its activation upon entry into S-phase to initiate replication fork progression. (from Takeda and Dutta, Oncogene 2005) 55
56 Essential role of ING complexes during DNA replication Larrieu and Pedeux Cell Cycle 2009 Modified from Ann E. Ehrenhofer-Murray EJB
57 Chromatin and Cancer Cancer Biology vs Chromatin Remodeling Translocations in Leukemia: -PML-RAR (functions trough HDAC) -MOZ/MORF-CBP/p300 (AML1-HATs) -MLL-CBP (HMT+HAT) -MLL-ENL/AF9 (HMT,SWI,HAT) -AML1-ETO (functions through HDAC) Tumor suppressors: -Rb, functions through SWI/SNF, HDAC, HMT and DNMT recruitment -p53, functions through HAT recruitment (ING) (SWI/SNF?) -BRCA1, is associated with SWI/SNF in vivo -Tip60, HAT subunit of NuA4 complex -ING1-5, subunits of HATs and HDACs Oncogenes: -Myc, functions through SWI/SNF and HAT recruitment -E2F, functions through HAT recruitment -E1A, disrupts HAT and HDAC complexes Mutations associated with leukemia and diverse tumors: -Ini1/Snf5, subunit of SWI/SNF -Brg1, subunit of SWI/SNF -HBO1, HAT subunit Mutations associated with metastasis/invasive growth -MTA1-3, subunits of NuRD -ING4, subunit of HBO1 HAT complex 57
58 58
59 The NuRD complex (MTA s) and Breast Cancer 59
60 60
61 Six hallmarks of cancer. Epigenetic silencing of tumor suppressor genes (examples in parentheses) plays a significant role in the development of each of these six tumor traits. Epigenetic therapy can resensitize cancer cells to conventional therapies. In a heterogenous tumor cell population, a small percentage of cells may respond directly to epigenetic drugs. The remaining tumor cells, however, could then be resensitized (e.g., by tumor suppressor gene re-expression) to conventional therapies. 61
62 Complex and Conserved roles of Chromatin in regulating Nuclear Functions 62
63 63
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Gene Expression DNA RNA. Protein. Metabolites, stress, environment
 Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Lecture 10. Eukaryotic gene regulation: chromatin remodelling
 Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Transcription and chromatin. General Transcription Factors + Promoter-specific factors + Co-activators
 Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
Host cell activation
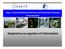 Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS
 R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
Chromatin Structure & Gene activity part 2
 Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Chromatin Structure & Gene activity part 2 Figure 13.30 Make sure that you understand it. It is good practice for identifying the purpose for various controls. Chromatin remodeling Acetylation helps to
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
The Role of Protein Domains in Cell Signaling
 The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics & cancer. Present by : Sanaz Zebardast Under supervision : Dr. Gheibi. 31 December 2016
 Epigenetics & cancer Present by : Sanaz Zebardast Under supervision : Dr. Gheibi 31 December 2016 1 contents Introduction Epigenetic & signaling pathways Epigenetic & integral protein Epigenetic & apoptosis
Epigenetics & cancer Present by : Sanaz Zebardast Under supervision : Dr. Gheibi 31 December 2016 1 contents Introduction Epigenetic & signaling pathways Epigenetic & integral protein Epigenetic & apoptosis
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Control of nuclear receptor function by local chromatin structure
 MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
Epigenetics: The Future of Psychology & Neuroscience. Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1
 Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Structure and function of active chromatin and DNase I hypersensitive sites
 MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
Stacie Bulfer Proteins Research Proposal April 27, Background
 Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
On the way of revealing coactivator complexes cross talk during transcriptional activation
 DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
5. Heterochromatin allows for silencing of large domains of the genome and utilizes a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
6. Heterochromatin allows for silencing of large domains of the genome and appears to utilize a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Supplemental Figure 1: Asymmetric chromatin maturation leads to epigenetic asymmetries on sister chromatids.
 Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
Epigenetics DNA methylation. Biosciences 741: Genomics Fall, 2013 Week 13. DNA Methylation
 Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Transcription Through Chromatin - Dynamic Organization of Genes
 GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
Are you the way you are because of the
 EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling
 Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Today. Genomic Imprinting & X-Inactivation
 Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Today 1. Quiz (~12 min) 2. Genomic imprinting in mammals 3. X-chromosome inactivation in mammals Note that readings on Dosage Compensation and Genomic Imprinting in Mammals are on our web site. Genomic
Pho23 Regulates Gene Expression through Histone Methylation and an Mck1-Controlled Pathway in Budding Yeast
 Pho23 Regulates Gene Expression through Histone Methylation and an Mck1-Controlled Pathway in Budding Yeast by Dennis Myers A thesis submitted in conformity with the requirements for the degree of Master
Pho23 Regulates Gene Expression through Histone Methylation and an Mck1-Controlled Pathway in Budding Yeast by Dennis Myers A thesis submitted in conformity with the requirements for the degree of Master
9/3/2009 DNA i DNA n euk euk yotes Organizatio Organ izatio n of o f gen ge e n tic Locati t on: In n ucleu e s material mater in e ial
 DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
DNA in eukaryotes Organization of genetic material in eukaryotes Location: In nucleus In mitochondria DNA in eukaryotes Nuclear DNA: Long, linear molecules; Chromatin chromosomes; 10% of DNA in genes,
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
The interplay of histone modifications writers that read
 Review Histones and Chromatin Review series The interplay of histone modifications writers that read Tianyi Zhang, Sarah Cooper *, & Neil Brockdorff Abstract Histones are subject to a vast array of posttranslational
Review Histones and Chromatin Review series The interplay of histone modifications writers that read Tianyi Zhang, Sarah Cooper *, & Neil Brockdorff Abstract Histones are subject to a vast array of posttranslational
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition
 Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson
 Thesis for doctoral degree (Ph.D.) 2009 Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson Characterization of Gcn5 Histone
Thesis for doctoral degree (Ph.D.) 2009 Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson Characterization of Gcn5 Histone
9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription
 9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription Friday, 20 th of November 2015, 13:00h Institute of Molecular Biology and Tumor Research (IMT) Philipps-University
9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription Friday, 20 th of November 2015, 13:00h Institute of Molecular Biology and Tumor Research (IMT) Philipps-University
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
DNA Methylation and Demethylation as Targets for Anticancer Therapy
 Biochemistry (Moscow), Vol. 70, No. 5, 2005, pp. 533-549. Translated from Biokhimiya, Vol. 70, No. 5, 2005, pp. 651-669. Original Russian Text Copyright 2005 by Szyf. DNA Methylation and Demethylation
Biochemistry (Moscow), Vol. 70, No. 5, 2005, pp. 533-549. Translated from Biokhimiya, Vol. 70, No. 5, 2005, pp. 651-669. Original Russian Text Copyright 2005 by Szyf. DNA Methylation and Demethylation
Epigenomics. Ivana de la Serna Block Health Science
 Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney
 BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Chromatin structure and epigenetic regulation of eukaryotic gene expression. Giacomo Cavalli
 Chromatin structure and epigenetic regulation of eukaryotic gene expression Giacomo Cavalli 12.09.2016 10,000 nm DNA compaction in a human nucleus 11 nm 30nm 1bp (0.3nm) compact size DNA length compaction
Chromatin structure and epigenetic regulation of eukaryotic gene expression Giacomo Cavalli 12.09.2016 10,000 nm DNA compaction in a human nucleus 11 nm 30nm 1bp (0.3nm) compact size DNA length compaction
THE HISTONE CODE AT DNA BREAKS: A GUIDE TO REPAIR? Haico van Attikum and Susan M. Gasser
 THE HISTONE CODE AT DNA BREAKS: A GUIDE TO REPAIR? Haico van Attikum and Susan M. Gasser Abstract Chromatin modifications are important for all cellular processes that involve DNA, including transcription,
THE HISTONE CODE AT DNA BREAKS: A GUIDE TO REPAIR? Haico van Attikum and Susan M. Gasser Abstract Chromatin modifications are important for all cellular processes that involve DNA, including transcription,
Structure and Function of Fusion Gene Products in. Childhood Acute Leukemia
 Structure and Function of Fusion Gene Products in Childhood Acute Leukemia Chromosomal Translocations Chr. 12 Chr. 21 der(12) der(21) A.T. Look, Science 278 (1997) Distribution Childhood ALL TEL-AML1 t(12;21)
Structure and Function of Fusion Gene Products in Childhood Acute Leukemia Chromosomal Translocations Chr. 12 Chr. 21 der(12) der(21) A.T. Look, Science 278 (1997) Distribution Childhood ALL TEL-AML1 t(12;21)
Protein methylation CH 3
 Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
The Pennsylvania State University. The Graduate School. Eberly College of Science THE MANY FACETS OF HISTONE TAILS IN REGULATING TRANSCRIPTION
 The Pennsylvania State University The Graduate School Eberly College of Science THE MANY FACETS OF HISTONE TAILS IN REGULATING TRANSCRIPTION A Dissertation in Biochemistry, Microbiology, and Molecular
The Pennsylvania State University The Graduate School Eberly College of Science THE MANY FACETS OF HISTONE TAILS IN REGULATING TRANSCRIPTION A Dissertation in Biochemistry, Microbiology, and Molecular
Chromatin Modifications by Methylation and Ubiquitination: Implications in the Regulation of Gene Expression
 Annu. Rev. Biochem. 2006. 75:243 69 First published online as a Review in Advance on February 22, 2006 The Annual Review of Biochemistry is online at biochem.annualreviews.org doi: 10.1146/ annurev.biochem.75.103004.142422
Annu. Rev. Biochem. 2006. 75:243 69 First published online as a Review in Advance on February 22, 2006 The Annual Review of Biochemistry is online at biochem.annualreviews.org doi: 10.1146/ annurev.biochem.75.103004.142422
The complex language of chromatin regulation during transcription Shelley L. Berger 1
 doi:10.1038/nature05915 The complex language of chromatin regulation during transcription Shelley L. Berger 1 An important development in understanding the influence of chromatin on gene regulation has
doi:10.1038/nature05915 The complex language of chromatin regulation during transcription Shelley L. Berger 1 An important development in understanding the influence of chromatin on gene regulation has
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Transcriptional and Epigenetic Mechanisms of Addiction
 Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE
 Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Organization of genetic material in eukaryotes
 Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
Organization of genetic material in eukaryotes biologiemoleculara.usmf.md pass.: bmgu e.usmf.md 1 DNA in eukaryotes Location: In nucleus In mitochondria biologiemoleculara.usmf.md e.usmf.md pass.: bmgu
HGD 5502 EPIGENETICS. Dr. Abhi Veerakumarasivam (2011)
 HGD 5502 EPIGENETICS Dr. Abhi Veerakumarasivam (2011) Outline History Definition Mechanisms of Epigenetics Epigenetic Processes Review History Conrad Hall Waddington (1905 1975) Developmental biologist,
HGD 5502 EPIGENETICS Dr. Abhi Veerakumarasivam (2011) Outline History Definition Mechanisms of Epigenetics Epigenetic Processes Review History Conrad Hall Waddington (1905 1975) Developmental biologist,
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
The role of epigenetics in breast. cancer progression
 The role of epigenetics in breast cancer progression Aslı Dilber Yıldırım January 2013 Master thesis, Cancer Genomics & Developmental Biology Under daily supervision of Rick van Nuland, MSc Examiners:
The role of epigenetics in breast cancer progression Aslı Dilber Yıldırım January 2013 Master thesis, Cancer Genomics & Developmental Biology Under daily supervision of Rick van Nuland, MSc Examiners:
Regulation of DNA replication-coupled histone gene expression
 /, 2017, Vol. 8, (No. 55), pp: 95005-95022 Regulation of DNA replication-coupled histone gene expression Qianyun Mei 1,*, Junhua Huang 1,*, Wanping Chen 1,2, Jie Tang 1, Chen Xu 1, Qi Yu 1, Ying Cheng
/, 2017, Vol. 8, (No. 55), pp: 95005-95022 Regulation of DNA replication-coupled histone gene expression Qianyun Mei 1,*, Junhua Huang 1,*, Wanping Chen 1,2, Jie Tang 1, Chen Xu 1, Qi Yu 1, Ying Cheng
DNMT1 SILENCING ELICITS DIFFERENT CELL CYCLE RESPONSES IN PRIMARY VERSUS TUMOR CELLS AND IS ASSOCIATED WITH ANEUPLOIDY GENERATION
 Ministero dell Istruzione, dell Università e della Ricerca Università degli Studi di Palermo Dottorato di Ricerca in Genomica e Proteomica nella ricerca Oncologica ed Endocrino-Metabolica Ciclo XXII: 2008-2010
Ministero dell Istruzione, dell Università e della Ricerca Università degli Studi di Palermo Dottorato di Ricerca in Genomica e Proteomica nella ricerca Oncologica ed Endocrino-Metabolica Ciclo XXII: 2008-2010
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Altered Histone Modifications in Gliomas
 REVIEW ARTICLE Brain Tumor Res Treat 2014;2(1):7-21 / pissn 2288-2405 / eissn 2288-2413 http://dx.doi.org/10.14791/btrt.2014.2.1.7 Altered Histone Modifications in Gliomas Young Zoon Kim Division of Neuro-Oncology,
REVIEW ARTICLE Brain Tumor Res Treat 2014;2(1):7-21 / pissn 2288-2405 / eissn 2288-2413 http://dx.doi.org/10.14791/btrt.2014.2.1.7 Altered Histone Modifications in Gliomas Young Zoon Kim Division of Neuro-Oncology,
Regulators of Cell Cycle Progression
 Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
Chapt 15: Molecular Genetics of Cell Cycle and Cancer
 Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Review II: Cell Biology
 Review II: Cell Biology Rajan Munshi BBSI @ Pitt 2006 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2006 Outline Cell Cycle Signal Transduction 1 Cell Cycle Four
Review II: Cell Biology Rajan Munshi BBSI @ Pitt 2006 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2006 Outline Cell Cycle Signal Transduction 1 Cell Cycle Four
CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS
 CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS Summary of the regulation of cyclin/cdk complexes during celll cycle Cell cycle phase Cyclin-cdk complex inhibitor activation Substrate(s) G1 Cyclin D/cdk 4,6
CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS Summary of the regulation of cyclin/cdk complexes during celll cycle Cell cycle phase Cyclin-cdk complex inhibitor activation Substrate(s) G1 Cyclin D/cdk 4,6
Epigenetics q&more 01.11
 Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Repressive Transcription
 Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Repressive Transcription The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Guenther, M. G., and R. A.
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Lecture 10. G1/S Regulation and Cell Cycle Checkpoints. G1/S regulation and growth control G2 repair checkpoint Spindle assembly or mitotic checkpoint
 Lecture 10 G1/S Regulation and Cell Cycle Checkpoints Outline: G1/S regulation and growth control G2 repair checkpoint Spindle assembly or mitotic checkpoint Paper: The roles of Fzy/Cdc20 and Fzr/Cdh1
Lecture 10 G1/S Regulation and Cell Cycle Checkpoints Outline: G1/S regulation and growth control G2 repair checkpoint Spindle assembly or mitotic checkpoint Paper: The roles of Fzy/Cdc20 and Fzr/Cdh1
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN
 Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Molecular Hematopathology Leukemias I. January 14, 2005
 Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
ubiquitinylation succinylation butyrylation hydroxylation crotonylation
 Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
Supplementary Information S1 (table) Overview of histone core-modifications histone, residue/modification H1 H2A methylation acetylation phosphorylation formylation oxidation crotonylation hydroxylation
H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the
 Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 825 H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the BY CLAES HOLMGREN ACTA UNIVERSITATIS
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 825 H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the BY CLAES HOLMGREN ACTA UNIVERSITATIS
Biochimica et Biophysica Acta
 BBAGRM-00286; No. of pages: 19; 4C: 8, 12 Biochimica et Biophysica Acta xxx (2010) xxx xxx Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbagrm
BBAGRM-00286; No. of pages: 19; 4C: 8, 12 Biochimica et Biophysica Acta xxx (2010) xxx xxx Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbagrm
The RSC Complex Localizes to Coding Sequences to Regulate Pol II and Histone Occupancy
 Article The RSC Complex Localizes to Coding Sequences to Regulate Pol II and Histone Occupancy Marla M. Spain, Suraiya A. Ansari, 2 Rakesh Pathak, Michael J. Palumbo, 2 Randall H. Morse, 2 and Chhabi K.
Article The RSC Complex Localizes to Coding Sequences to Regulate Pol II and Histone Occupancy Marla M. Spain, Suraiya A. Ansari, 2 Rakesh Pathak, Michael J. Palumbo, 2 Randall H. Morse, 2 and Chhabi K.
Epigenetic mechanisms behind cellular sensitivity to DNA damage
 www.cell-stress.com Review Epigenetic mechanisms behind cellular sensitivity to DNA damage Amanda K. Williamson 1, Zijing Zhu 1 and Zhi-Min Yuan 1, * 1 Department of Environmental Health, John B. Little
www.cell-stress.com Review Epigenetic mechanisms behind cellular sensitivity to DNA damage Amanda K. Williamson 1, Zijing Zhu 1 and Zhi-Min Yuan 1, * 1 Department of Environmental Health, John B. Little
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Epigenetic Inheritance
 (2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
(2) The role of Epigenetic Inheritance Lamarck Revisited Lamarck was incorrect in thinking that the inheritance of acquired characters is the main mechanism of evolution (Natural Selection more common)
Epigenetics ~from mechanism to therapy~ Literature seminar January 31, 2012 Soichi Ito (B4)
 Epigenetics ~from mechanism to therapy~ Literature seminar January 31, 2012 Soichi Ito (B4) 1 Contents Introduction Central dogma What is the Epigenetics? Topics 1. DNA methylation 2. Histone modification
Epigenetics ~from mechanism to therapy~ Literature seminar January 31, 2012 Soichi Ito (B4) 1 Contents Introduction Central dogma What is the Epigenetics? Topics 1. DNA methylation 2. Histone modification
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
I. Each cell of a multicellular eukaryote expresses only a small fraction of its genome
 CHAPTER 18 GENOME ORGANIZATION AND EXPRESSION IN EUKARYOTES OUTLINE I. Each cell of a multicellular eukaryote expresses only a small fraction of its genome II. III. IV. The structural organization of chromatin
CHAPTER 18 GENOME ORGANIZATION AND EXPRESSION IN EUKARYOTES OUTLINE I. Each cell of a multicellular eukaryote expresses only a small fraction of its genome II. III. IV. The structural organization of chromatin
The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation gene PHR1
 Published online 3 February 2011 Nucleic Acids Research, 2011, Vol. 39, No. 10 4151 4165 doi:10.1093/nar/gkr040 The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation
Published online 3 February 2011 Nucleic Acids Research, 2011, Vol. 39, No. 10 4151 4165 doi:10.1093/nar/gkr040 The histone H3K36 demethylase Rph1/KDM4 regulates the expression of the photoreactivation
Variants of core histones and their roles in cell fate decisions, development and cancer
 Variants of core histones and their roles in cell fate decisions, development and cancer Marcus Buschbeck 1,2 and Sandra B. Hake 3,4 Abstract Histone variants endow chromatin with unique properties and
Variants of core histones and their roles in cell fate decisions, development and cancer Marcus Buschbeck 1,2 and Sandra B. Hake 3,4 Abstract Histone variants endow chromatin with unique properties and
Controlling Epithelial to Mesenchymal Transition through Acetylation of Histone H2BK5
 Biological Chemistry Controlling Epithelial to Mesenchymal Transition through Acetylation of Histone H2BK5 Robert J. Mobley 1,* and Amy N. Abell 1,2 1 Department of Biological Sciences, 2 Department of
Biological Chemistry Controlling Epithelial to Mesenchymal Transition through Acetylation of Histone H2BK5 Robert J. Mobley 1,* and Amy N. Abell 1,2 1 Department of Biological Sciences, 2 Department of
