Title cooperative gene expression.
|
|
|
- Joella York
- 5 years ago
- Views:
Transcription
1 Title Rigor of cell fate decision cooperative gene expression by vari by p53 Author(s) Murakami, Yohei; Takada, Shoji Citation BIOPHYSICS (2012), 8: Issue Date 2012 URL (c) 2012 THE BIOPHYSICAL SOCIETY OF Right not the published version. Please c version. この論文は出版社版でありません 引用の際には出版社版をご確認ご利用ください Type Journal Article Textversion author Kyoto University
2 Rigor of cell fate decision by variable p53 pulses and roles of cooperative gene expression by p53 Yohei Murakami* and Shoji Takada Department of Biophysics, Division of Biology, Graduate School of Science, Kyoto University, Kyoto , Japan *Corresponding author: Department of Biophysics, Division of Biology, Graduate School of Science, Kyoto University, Kyoto , Japan. Phone: ; Fax: ABSTRACT Upon DNA damage, the cell fate decision between survival and apoptosis is largely regulated by p53-related networks. Recent experiments found a series of discrete p53 pulses in individual cells, which led to the hypothesis that the cell fate decision upon DNA damage is controlled by counting the number of p53 pulses. Under this hypothesis, Sun et al. (2009) modeled the Bax activation switch in the apoptosis signal transduction pathway that can rigorously count the number of uniform p53 pulses. Based on experimental evidence, here we use variable p53 pulses with Sun et al.'s model to investigate how the variability in p53 pulses affects the rigor of the cell fate decision by the pulse number. Our calculations showed that the experimentally anticipated variability in the pulse sizes reduces the rigor of the cell fate decision. In addition, we 1
3 tested the roles of the cooperativity in PUMA expression by p53, finding that lower cooperativity is plausible for more rigorous cell fate decision. This is because the variability in the p53 pulse height is more amplified in PUMA expressions with more cooperative cases. Key words: Apoptosis, cell death, computational biology, systems biology 2
4 Introduction DNA damage in cells leads to either the induction of apoptosis or activation of DNA repair and cell cycle arrest, depending on the level of DNA damage 1-3, which is termed the "cell fate decision". The precise decision is of central importance and disorder of apoptosis signal transduction often causes serious diseases such as cancer and neurodegenerative disease 4-6. In the cell fate decision process, p53, a well-known tumor-suppressor transcription factor, plays key roles by regulating the transcription of many kinds of genes responsible for cell cycle control, DNA repair, and apoptosis induction 2, 7. Recently, it was reported that, upon DNA damage by gamma ray radiation, a series of discrete pulse-like p53 concentration changes (called p53 pulses in this article) were observed in tumor cell lines in individual cells It was suggested that the level of DNA damage does not severely affect the mean size or shape of each pulse, but correlates with the number of p53 pulses 10. The p53 pulses may be generated when the level of DNA damage exceeds a particular threshold. A plausible hypothesis is that a cell counts the successive p53 pulses and decides the cell fate by the number of pulses 13. In other words, the difference in the p53 pulse number elicits a different cellular response, apoptosis or survival; however, the details of the p53 pulse counting mechanism of cells and the relationship between the p53 pulse number and the cell fate decision remain largely unclear. Currently, several studies using computational systems biology have elucidated the p53 pulse-counting mechanism and the relationship between the p53 pulse and the cell fate decision For example, Sun et al. (2009) modeled a Bax activation switch that 3
5 counts the number of p53 pulses and decides the cell fate. In their work, the dynamics was deterministic and the p53 pulse size and shape were fixed. Thus, when the pulse number was larger than the threshold, cells always underwent apoptosis, whereas when the pulse number was below the threshold, cells always survived. The cell fate decision is completely rigorous depending on the pulse number. The threshold of the p53 pulse number that divides apoptosis from survival depends on their model details and kinetic parameters. Experiments suggested, however, that the pulse size and shape are variable More precisely, the pulse heights (also called amplitude), pulse widths, and pulse intervals (also called period) are not constants, but are variable. For example, the pulse heights of different pulses in the same cell can vary by about 3-fold 10. This variability apparently complicates the rigorous cell fate decision. Zhang et al. (2009b) included variability in terms of the initial level of DNA damage and its repair dynamics, while the pulse size was still deterministic and fixed. The main purpose of this paper was to address the effects of variability in p53 pulses. How the variability affects the rigor of cell fate decision by counting the pulse number is investigated. Signal transductions from p53 to the induction of apoptosis have been intensively studied and many components identified experimentally. The p53-based apoptosis signals are transmitted to mitochondria, especially to Bcl-2 family proteins that mutually interact at the mitochondrial outer membrane. Bcl-2 family proteins induce cytochrome c release from mitochondria to cytoplasm, and released cytochrome c enhances the activation of caspase in cytoplasm. Then, caspase activation turns on cascade reactions, leading to various apoptotic morphological changes 5, 22. Most of the members of Bcl-2 family proteins are known to be located at the mitochondrial outer membrane and interact with each other. Bcl-2 family proteins can be classified into 3 groups, (1) 4
6 pro-apoptotic Bcl-2 family proteins (Bax etc), (2) anti-apoptotic Bcl-2 family proteins (Bcl-2, Mcl-1 etc), (3) BH3-only proteins (PUMA, NOXA, Bid etc). BH3-only proteins can be further divided into 2 subgroups, called activator BH3-only proteins (Bid etc) and sensitizer (also called enabler) BH3-only proteins (PUMA, NOXA etc). Pro-apoptotic Bcl-2 family proteins and BH3-only proteins facilitate apoptosis, whereas anti-apoptotic Bcl-2 family proteins counteract them and facilitate survival. Interactions and balances among Bcl-2 family proteins are thought to be crucial for the precise cell death decision For p53 to enhance apoptosis, p53 activates the expression of several kinds of Bcl-2 family proteins 22, 25. In particular, PUMA, one of the sensitizer BH3-only proteins, is a main transcriptional target of p53 in various tissues In computational systems biology studies, Sun et al. (2009) suggested that the Bax activation switch can count p53 pulses through PUMA accumulation and decide the cell fate. Our modeling is largely based on the signal transduction model of Sun et al. (2009). We note that Sun et al.'s model has not been verified experimentally. To experimentally verify their suggestion, we need to observe and examine the relationship between the number of p53 pulses which are directly related to the expression of PUMA and subsequent apoptosis. In the gene expression by p53, p53 binds to the target DNA in the tetramer form in a highly cooperative manner This highly cooperative binding of p53 to DNA is a source of its nonlinear nature. In the latter part of this paper, we investigate how cooperativity in PUMA expression by p53 affects the rigor of cell fate decision. In this study, based on the modeling of Sun et al. (2009), we investigated how the variable p53 pulse size affects the rigor of cell fate decision. We explored the factors 5
7 which influence the rigor of cell death decision by the apoptosis signal transduction pathway. In particular, we focused on the cooperativity of PUMA expression by p53 because a highly cooperative process is thought to be the source of its nonlinear nature and strongly influences the cell fate decision. Methods Model of the apoptosis signal transduction pathway In this study, we adopted a subset of the model of the apoptosis signal transduction pathway developed by Sun et al. (2009). In particular, to focus on the probabilistic nature of the apoptosis signal transduction pathway, we selected only the core of the bifurcation module, Bax activation switch module, from the model of Sun et al. (2009) (Figure 1). We briefly describe the model in Figure 1. The input stimulus for apoptosis induction in this model is p53 pulses, which is supposed to be produced by gamma ray radiation-based DNA damage. Increased p53 concentration enhances the transcription of PUMA, one of the sensitizer BH3-only proteins. The output signal for apoptosis in this model is pore formation on the mitochondrial outer membrane. The pore transports cytochrome c from the mitochondria to cytoplasm, and the cytoplasmic cytochrome c activates caspases, which then induce apoptosis. Since downstream from the pore formation is relatively straightforward, we do not include it here. The pore is an oligomer (here assumed as a dimer following Sun et al. (2009)) of activated Bax, pro-apoptotic Bcl-2 family proteins. A monomeric Bax takes two forms, inactive Bax (denoted as Bax) and activated Bax (denoted as AcBax). Binding of two AcBaxs as well as binding of 6
8 AcBax with Bax leads to pore formation. Activator BH3-only proteins (denoted as Act) activate Bax to form activated Bax (AcBax). Bcl-2, one of the anti-apoptotic Bcl-2 family proteins, is a key player, which binds to and sequesters activated Bax, and also inhibits Act to activate Bax. PUMA binds to Bcl-2, which reduces Bcl-2 binding to its targets, Act and AcBax. We note that PUMA expressed by p53 stimulus activates Bax not directly, but indirectly, leading to apoptosis enhancement In the model, all the protein interactions (all dashed lines in Figure 1) except the arrow from p53 to PUMA are protein interactions, and the arrow from p53 to PUMA represents transcription regulation. Based on static bifurcation analysis (see Steady state bifurcation analysis in Results), we can unambiguously define the onset of apoptosis when the pore concentration is over (nm) after a sufficient time (3000min in this work). Model representation by ordinary differential equations, kinetic parameters and initial conditions All protein interactions in this model are represented by ordinary differential equations following mass-action kinetics (Table 1). Only the expression process of PUMA by p53 is represented by the Hill function, as in Sun et al. (2009). By default, we used the same kinetic parameters as in Sun et al. (2009), but changed several kinetic parameters of the expression process of PUMA by p53 based on experimental data 28, 30 (Table 2). As for the maximum rate of PUMA synthesis by p53 ( kssp53 in Table 2), it was set 10 times higher than the PUMA basal production rate based on Yu et al. (2001). We set the Hill coefficient of PUMA expression by p53 as 2.0, by default, ( n in Table 2) because an in vitro experiment showed that p53 binding to DNA has a Hill coefficient of
9 In this study, for simplification, we assumed that p53 concentration is zero without DNA damage. The initial conditions of all proteins were set at the steady state concentrations when the p53 concentration was zero. p53 pulse representation Experiments suggested that p53 pulse heights, widths and intervals do not significantly depend on the level of DNA damage, and have their own mean values as well as their own variability The measured mean values and standard deviations (SD) of p53 pulse widths and intervals are ± (min) and ± (min), respectively (mean value ± SD) 10. As for the p53 pulse heights, although the coefficient of variation (CV = SD/mean value) was experimentally estimated as 0.7 9, the mean height itself was not reliably measured. Therefore, we chose the mean height for each simulation by a certain criterion (see Minimum pulse height for apoptosis in Results for the details); without the variability in the pulse, apoptosis occurs after a certain number of pulses. Instead of generating p53 pulses by the p53-mdm2 interaction network module, here we gave the p53 pulses as Gaussian functions. Namely, the concentration of the n-th p53 pulse is represented by. (1) Here [p53] is the p53 concentration (nm) at time t (min), H represents the pulse height (nm), μ is the peak time of the first pulse set as "pulse width/2"(min), and σ represents the width of the pulse set as p53 pulse width / 6 (min). Factor 6 comes from the difference in the definition of the "width". The experimentally measured "p53 pulse 8
10 width" is the time difference between the onset and the end of the pulse, while σ is the "width" in Gaussian function. D represents the pulse interval (min). To represent variable p53 pulse heights, widths, and intervals, we used random numbers. For σ, and D, random numbers were generated from the normal distribution with the above mentioned means and SDs. Pulse height was generated for the normal distribution with a given mean and SD = 0.7 x mean. Figure 2 illustrates the generated p53 pulses. Calculation of apoptosis probability We performed 1000 independent calculations with variable p53 pulses. The apoptosis probability is estimated as the ratio of the cases in which apoptosis occurred by the end of calculations (3000min). Numerical calculation Steady state concentrations of all proteins were calculated by solving the simultaneous equations obtained by setting all ordinary differential equations equal to zero with the standard Newton-Raphson method. Local stabilities of all steady states were determined by evaluating eigenvalues of Jacobian matrices, which were obtained by linearization of ordinary differential equations. In the dynamics calculations, the ordinary differential equations were numerically solved by the fourth-order Runge-Kutta method with a time step of 0.01 minute. Results 9
11 Steady state bifurcation analysis We started with the standard steady state bifurcation analysis of the model (Figure 3). In the figure, red and blue curves indicate stable and unstable steady states, respectively. As described above, steady states with a high pore concentration correspond to the apoptosis state, while those with low (basal) pore concentration represent the survival (non-apoptosis) state. Indeed, Figure 3A shows two discrete stable states, one apoptotic and the other survival. In this bistable system, increasing the p53 concentration, the survival state suddenly disappears and abrupt transition from the survival state to the apoptosis state occurs at a threshold p53 concentration (solid arrow in Figure 3A). Once a high level of pore is accumulated (i.e., apoptosis state), it is sustained even when the p53 concentration returns to the level below the threshold (x-marked dashed arrow in Figure 3A). Thus, the system shows irreversibility. It has been suggested experimentally as well as computationally that the interaction network of Bcl-2 family proteins at mitochondria exhibits bistable Bax activation and pore formation 16, In particular, two positive feedbacks, Bax auto-activation and Bcl-2 binding to activated Bax, were thought to be the origin of the bistable nature 32. Apparently, bistability and irreversibility are important for the apoptosis signal transduction pathway to avoid improper apoptosis induced by noise, or the appearance of malignant cells by improper survival In this study, we address the cooperativity of PUMA expression regulation by p53, which is represented by the Hill coefficient, n in Stimulus term in Eq. (7) in Table 1. The default is n = 2.0 (Figure 3A), but we will also examine cases with n = 1.0 and 4.0. Thus, we show bifurcation diagrams for n = 1.0 and n = 4.0 (Figure 3B and C). As the Hill coefficient increases, the threshold p53 concentration increases, whereas the pore 10
12 concentrations in the apoptosis and survival states do not alter very much. In all cases, Figures 3 robustly shows the bistability and irreversibility of pore formation by p53 concentration changes. These are essentially the same as the results in Sun et al. (2009), which dealt with more comprehensive signal transduction networks. Based on these bifurcation analyses, we set the criterion for the pore concentration in the apoptosis state as >100.0 (nm). Minimum pulse height for apoptosis We then performed dynamic analysis of the model. Before analyzing the roles of variability in the p53 pulses, a uniform p53 pulse was examined as a control. The minimum pulse height necessary for apoptosis induction was calculated for each given pulse number (Table 3). Here, the pulse width and interval were set as (min) and (min), respectively, based on the above argument (see p53 pulse representation in Methods). As the pulse number increases, the required minimal pulse height for apoptosis induction becomes smaller. In addition, a larger pulse height is necessary for apoptosis induction as the Hill coefficient becomes larger. The calculated data of the minimum pulse height was used in the following sections to calculate the probability of apoptosis onset. The rigor of cell fate decision and p53 pulse size variability Now, variability was introduced into the p53 pulse size to assess how it affected the rigor of cell fate decision. Experimentally, the number of p53 pulses to induce apoptosis is unclear. In the computational study of Sun et al. (2009), with their parameter choices, the cell underwent apoptosis when the pulse was 3. Here, here we simply followed 11
13 their model by setting the pulse height as (nm), which is between the minimum pulse height for 2-pulse apoptosis induction and that for 3-pulse apoptosis induction (Table 3). For the pulse size variety, SDs and the coefficient of variation described in p53 pulse representation in Methods were used as a reference. These values are based on experiments of tumor cell lines It was argued that normal cells may have less variability than tumor cells 36 ; therefore, based on the degree of variability in tumor cells (designed as "pulse_variability = 1"), we also tested reduced variability cases by a factor of 0.5 ("pulse_variability = 0.5 ), 0.25 ( pulse_variability = 0.25 ), and 0 ("pulse_variability = 0" corresponding to deterministic cases). For example, when pulse_variability = 0.5, the SD of pulse width was set to 80.0, the SD of pulse interval was set to 50.0, and the coefficient of variation of pulse height was set to Figure 4 shows apoptosis probability as a function of the number of variable p53 pulses. Without variability ( pulse_variability = 0 ), with our choice of pulse height, the apoptosis probability suddenly changed from 0 % for 2 pulses to 100 % for 3 pulses. Due to the deterministic nature, the cell fate decision is completely rigorous. With pulse_variability = 0.25, the cell fate decision curve softened, still showing sigmoidal behavior. As the pulse_variability increased, the sigmoidal nature was reduced, and the curves became hyperbolic. These results indicate that p53 pulse variability lowers the rigor of the cell fate decision. In normal cells, pulse height, width and interval are thought to fluctuate to some extent, as assumed in this study. With variable p53 pulse sizes, completely rigorous cell fate decision is difficult. Recently, Zhang et al. (2009b) also investigated the variability in the cell fate decision. They investigated the stochasticity of the initial DNA damage and its repair dynamics as the source of cell fate variability, whereas the pulse size was deterministic. It is probable 12
14 that variable initial DNA damage and its repair dynamics, together with intrinsic stochasticity, may induce variable p53 pulse sizes. Thus two means of variability are related, but not identical. In addition, all biochemical reactions should have intrinsic stochasticity, which further lowers the rigor of the cell fate decision. Considering these multiple sources of stochasticity will be a future study. Role of cooperativity in the rigor of cell fate decision by variable p53 pulses Finally, we investigated how the rigor of the cell fate decision by variable p53 pulse sizes is affected by the cooperativity in PUMA expression regulation by p53. Until this point we had used the default Hill coefficient of n = 2.0, but here we also used n = 1.0 and 4.0. The mean pulse heights were set to 77.5 (nm) for n = 1.0 and (nm) for n = 4.0, respectively, which were chosen in the same way as the default n = 2.0. Figure 5 shows apoptosis probability as a function of the pulse number. Quite surprisingly, it was seen that, for the same degree of pulse variability, a larger Hill coefficient exhibits less of a sigmoidal curve. Namely, with larger cooperativity in PUMA expression regulation by p53, the cell fate decision becomes less rigorous. Experimentally, the exact Hill coefficient for PUMA expression regulation by p53 in living cells is unclear. Weinberg et al. (2004) estimated it to be 1.8 by in vitro experiments using p53ct (residues ) and synthesized DNA. Some of the earlier computational studies used n = 3.0 or , 20, probably based on the fact that p53 is a tetramer and its binding to DNA is highly cooperative 29. Why did a larger Hill coefficient lead to a less pronounced sigmoid curve? First, it was investigated, among the three types of variability in the pulses, which variability had the 13
15 largest impact on the rigor of cell fate decision. We repeated the apoptosis probability calculations, but by setting one of three SDs as zero (Figure 6). In Figure 6, we can see that there was almost no difference among the three Hill coefficients when the pulse height was constant. In contrast, when the pulse width or pulse interval was constant, the rigor of cell fate decision seemed to depend on the Hill coefficient, similar to the Figure 5 result. These results suggest that the pulse height variability seems to be almost the sole factor influencing the rigor of cell fate decision. Next, we tried to elucidate why pulse height variability strongly influences the rigor of cell fate decision. For example, in the case of pulse_variability = 0.5, the p53 pulse height was distributed across nM when the Hill coefficient equaled 2.0. These ranges were input into the Stimulus term (2) to obtain the distribution range in PUMA concentration. We show these ranges in Table 4 for different Hill coefficients ( ST range and its width RWS ). For better comparison, Table 4 also contains the relative ranges, the ranges divided by the mean ( NRWS ). The results in Table 4 clearly shows that, as the Hill coefficient increases, the variability in the p53 pulse height more strongly amplifies the variability in the PUMA concentration (variability in Stimulus), which results in the uncertainty of cell fate decision. In the current model, PUMA accumulates over p53 pulses. When the PUMA concentration exceeds a threshold, Bax activation is switched on; therefore, the cumulative amount of p53 within a certain time scale is a key factor. We calculated the cumulative amount of p53 pulses above the pore formation thresholds in Figure 3 while varying each of the 14
16 three pulse characteristics (height, width and interval). Calculated averages of integrals and coefficient of variations (CV) of integrals were, respectively, (nm*min) and when the height was varied, (nm*min) and when the width was varied, and (nm*min) and when the interval was varied. These values were calculated by 1000 simulations of 5 pulses with pulse variability = 0.5. Thus, the pulse interval is not very important. With an experimentally anticipated size of variability in pulse height and width, the height variability modulates the cumulative amount of p53 more than the pulse width. Because of the variability amplification via the Hill formula, the apoptosis probability curves in Figure 5 become more hyperbolic with a larger Hill coefficient. Qualitatively similar results were also obtained by setting the pulse height to different values, e.g. the minimum pulse height for 3-pulse apoptosis induction (data not shown). Thus, the roles of the cooperativity of PUMA expression by p53 in the rigor of the cell fate decision found here were quite robust. Discussion and Conclusion We investigated how a variable p53 pulse size affects the rigor of cell fate decision, and the roles of the cooperativity in PUMA expression regulation by p53. By variable pulse sizes, the rigorous cell fate decision was disrupted. As the variability in p53 pulse size increases, the probability of the apoptosis induction curve changes from sigmoidal to hyperbolic. As the cooperativity of PUMA expression by p53 increases, the rigor of the cell fate decision is more disrupted. Among the three types of pulse variability, pulse height variability was the most influential on perturbation because the cooperativity in 15
17 PUMA expression amplified the variability in stimulus. In the current study, we set the Hill coefficient of PUMA expression by p53 as 2.0 by default, following the in vitro experiment which showed that p53 binding to DNA has a Hill coefficient of ; however, equation (7) in Table 1 for the PUMA expression by p53 is a simple modeling that, in reality, includes many complex processes. Thus, not only cooperative binding of p53 to the promoter of the PUMA gene but also many other transcription and translation processes, such as chromatin remodeling or mrna transport from the nucleus to cytoplasm, are included in the Stimulus term. Any factors which influence transcription and translation processes may change the cooperativity of gene expression by p53. Thus, it is possible that cells have various mechanisms to regulate the rigor of cell fate decision by controlling the cooperativity of gene expression by p53. The suggestion of Sun et al. (2009) and the current work on the influence of cooperativity of the PUMA expression by p53 on the rigor of cell fate decision need to be verified experimentally. For this, firstly, a mutant and/or modified p53 that enhances the expression of PUMA with different levels of cooperativity will be required. Also, cells should be prepared which can stably express such types of p53 after DNA damage. Alternatively, a method to control the cooperativity of the gene expression by p53 could be used to verify our suggestion. Secondly, cells should be damaged by gamma ray radiation at different intensities to induce different numbers of p53 pulses. Finally, the relationship between the number of p53 pulses and the ratio of apoptotic cells should be examined. Using this type of experiment, experimental evidence will be obtained of the relationship between the rigor of the cell fate decision and the cooperativity of gene expression by p53. To accurately count the number of p53 pulses in individual cells, 16
18 single cell measurements as in time-lapse microscopy are thought to be suitable, as indicated in Lahav et al. (2004). Although we limited ourselves to the p53 pulse counting mechanism of the Bax activation switch, other mechanisms are also possible. For example, downstream of the Bax activation switch, there exists a caspase cascade 37. It is thought that because of the existence of a positive feedback loop in the caspase cascade, caspase-3 activation occurs in an all-or-none irreversible manner,, similarly to the Bax activation switch 38. Although PUMA is the main transcriptional target of p53 in various tissues 26-28, p53 is known to enhance the expression of Apaf-1 modestly 25. Apaf-1 is known to form a complex with cytochrome c and to activate the caspase cascade 37. Pulse-like concentration changes of p53 may influence Apaf-1 concentration and subsequent caspase cascade activation; therefore, the caspase cascade may also count the number of p53 pulses. Since the two counting mechanisms are not mutually exclusive, both the Bax activation switch and caspase cascade may count the number of p53 pulses and decide the cell fate; however, even if the caspase cascade can count the number of p53 pulses, the influence of the cooperativity of gene expression by p53 is still thought to be important for the rigor of the cell fate decision, as indicated in the current study. The current study sheds light on the probabilistic nature of the cell fate decision, which may make cells fragile; however, homeostasis or the health of our tissues and organs may be ensured by higher order systems such as the immune system. Acknowledgements 17
19 This work was partly supported by a Grant-in-Aid for Scientific Research on Innovative Areas Molecular Science of Fluctuations toward Biological Functions, by Research and Development of the Next-Generation Integrated Simulation of Living Matter of MEXT, and by the Global COE Program Formation of a strategic base for biodiversity and evolutionary research: from genome to ecosystem of MEXT. Conflict of interest There are no conflicts of interest. Contributors Y.M. designed and performed the research and analyzed data; and Y.M. and S.T. wrote the paper. 18
20 References 1. Aylon, Y., & M. Oren. Living with p53, dying of p53. Cell 130, (2007). 2. Batchelor, E., A. Loewer, & G. Lahav. The ups and downs of p53: understanding protein dynamics in single cells. Nat Rev Cancer 9, (2009). 3. Vousden, K. H., & D. P. Lane. p53 in health and disease. Nat Rev Mol Cell Biol 8, (2007). 4. Adams, J. M., & S. Cory. The Bcl-2 apoptotic switch in cancer development and therapy. Oncogene 26, (2007). 5. Cotter, T. G. Apoptosis and cancer: the genesis of a research field. Nat Rev Cancer 9, (2009). 6. Mattson, M. P.. Apoptosis in neurodegenerative disorders. Nat Rev Mol Cell Biol 1, (2000). 7. Vogelstein, B., D. Lane, & A. J. Levine. Surfing the p53 network. Nature 408, (2000). 8. Batchelor, E., C. S. Mock, I. Bhan, A. Loewer, & G. Lahav. Recurrent initiation: a mechanism for triggering p53 pulses in response to DNA damage. Mol Cell 30, (2008). 9. Geva-Zatorsky, N., N. Rosenfeld, S. Itzkovitz, R. Milo, A. Sigal, E. Dekel, T. Yarnitzky, Y. Liron, P. Polak, G. Lahav, & U. Alon. Oscillations and variability in the p53 system. Mol Syst Biol 2, 0033 (2006). 10. Lahav, G., N. Rosenfeld, A. Sigal, N. Geva-Zatorsky, A. J. Levine, M. B. Elowitz, & U. Alon. Dynamics of the p53-mdm2 feedback loop in individual cells. Nat 19
21 Genet 36, (2004). 11. Loewer, A., E. Batchelor, G. Gaglia, & G. Lahav. Basal dynamics of p53 reveal transcriptionally attenuated pulses in cycling cells. Cell 142, (2010). 12. Ramalingam, S., P. Honkanen, L. Young, T. Shimura, J. Austin, P. S. Steeg, & S. Nishizuka. Quantitative assessment of the p53-mdm2 feedback loop using protein lysate microarrays. Cancer Res 67, (2007). 13. Tyson, J. J. Monitoring p53's pulse. Nat Genet 36, (2004). 14. Hamada, H., Y. Tashima, Y. Kisaka, K. Iwamoto, T. Hanai, Y. Eguchi, & M. Okamoto. Sophisticated framework between cell cycle arrest and apoptosis induction based on p53 dynamics. PLoS One 4, e4795 (2009). 15. Iwamoto, K., H. Hamada, Y. Eguchi, & M. Okamoto. Mathematical modeling of cell cycle regulation in response to DNA damage: exploring mechanisms of cell-fate determination. Biosystems 103, (2011). 16. Sun, T., C. Chen, Y. Wu, S. Zhang, J. Cui, & P. Shen. Modeling the role of p53 pulses in DNA damage- induced cell death decision. BMC Bioinformatics 10, 190 (2009). 17. Wee, K. B., U. Surana, & B. D. Aguda. Oscillations of the p53-akt network: implications on cell survival and death. PLoS One 4, e4407 (2009). 18. Zhang, T., P. Brazhnik, & J. J. Tyson.. Exploring mechanisms of the DNA-damage response: p53 pulses and their possible relevance to apoptosis. Cell Cycle 6, (2007). 19. Zhang, T., P. Brazhnik, & J. J. Tyson. Computational analysis of dynamical responses to the intrinsic pathway of programmed cell death. Biophys J 97, (2009). 20
22 20. Zhang, X. P., F. Liu, Z. Cheng, & W. Wang. Cell fate decision mediated by p53 pulses. Proc Natl Acad Sci U S A 106, (2009). 21. Zhang, X. P., F. Liu, & W. Wang. Coordination between cell cycle progression and cell fate decision by the p53 and E2F1 pathways in response to DNA damage. J Biol Chem 285, (2010). 22. Haupt, S., M. Berger, Z. Goldberg, & Y. Haupt. Apoptosis - the p53 network. J Cell Sci 116, (2003). 23. Brenner, D., & T. W. Mak. Mitochondrial cell death effectors. Curr Opin Cell Biol 21, (2009). 24. Youle, R. J., & A. Strasser. The BCL-2 protein family: opposing activities that mediate cell death. Nat Rev Mol Cell Biol 9, (2008). 25. Yu, J., & L. Zhang. The transcriptional targets of p53 in apoptosis control. Biochem Biophys Res Commun 331, (2005). 26. Nakano, K., & K. H. Vousden. PUMA, a novel proapoptotic gene, is induced by p53. Mol Cell 7, (2001). 27. Villunger, A., E. M. Michalak, L. Coultas, F. Mullauer, G. Bock, M. J. Ausserlechner, J. M. Adams, & A. Strasser. p53- and drug-induced apoptotic responses mediated by BH3-only proteins puma and noxa. Science 302, (2003). 28. Yu, J., L. Zhang, P. M. Hwang, K. W. Kinzler, & B. Vogelstein. PUMA induces the rapid apoptosis of colorectal cancer cells. Mol Cell 7, (2001). 29. Balagurumoorthy, P., H. Sakamoto, M. S. Lewis, N. Zambrano, G. M. Clore, A. M. Gronenborn, E. Appella, & R. E. Harrington. Four p53 DNA-binding domain peptides bind natural p53-response elements and bend the DNA. Proc Natl 21
23 Acad Sci U S A 92, (1995). 30. Weinberg, R. L., D. B. Veprintsev, & A. R. Fersht. Cooperative binding of tetrameric p53 to DNA. J Mol Biol 341, (2004). 31. Chen, C., J. Cui, H. Lu, R. Wang, S. Zhang, & P. Shen. Modeling of the role of a Bax-activation switch in the mitochondrial apoptosis decision. Biophys J 92, (2007). 32. Cui, J., C. Chen, H. Lu, T. Sun, & P. Shen. Two independent positive feedbacks and bistability in the Bcl-2 apoptotic switch. PLoS One 3, e1469 (2008). 33. Sun, T., X. Lin, Y. Wei, Y. Xu, & P. Shen. Evaluating bistability of Bax activation switch. FEBS Lett 584, (2010). 34. Albeck, J. G., J. M. Burke, B. B. Aldridge, M. Zhang, D. A. Lauffenburger, & P. K. Sorger. Quantitative analysis of pathways controlling extrinsic apoptosis in single cells. Mol Cell 30, (2008). 35. Vaughan, A. T., C. J. Betti, & M. J. Villalobos. Surviving apoptosis. Apoptosis 7, (2002). 36. Tyson, J. J. Another turn for p53. Mol Syst Biol 2, 0032 (2006). 37. Slee, E. A., M. T. Harte, R. M. Kluck, B. B. Wolf, C. A. Casiano, D. D. Newmeyer, H. G. Wang, J. C. Reed, D. W. Nicholson, E. S. Alnemri, D. R. Green, and S. J. Martin. Ordering the cytochrome c-initiated caspase cascade: hierarchical activation of caspases-2, -3, -6, -7, -8, and -10 in a caspase-9-dependent manner. J Cell Biol 144: (1999). 38. Legewie, S., N. Bluthgen, and H. Herzel. Mathematical modeling identifies inhibitors of apoptosis as mediators of positive feedback and bistability. PLoS Comput Biol 2:e120 (2006). 22
24 Figure Legends Fig. 1. Schematic diagram of the model. Solid arrows represent conformational change, dimerization or dissociation of the same protein, Bax. Dotted arrow from p53 to PUMA represents enhancement of transcription. Other dotted arrows represent activation. Dotted lines with horizontal bar represent inhibition. Abbreviations: AcBax: activated Bax, Act: activator BH3-only protein. Fig. 2. Examples of p53 pulses. Mean pulse height was set to (nm) as an example. (A) An example in which the pulse heights, widths and intervals were set to constant mean values. (B) An example in which the pulse heights, widths, and interval varied. Fig. 3. Bifurcation diagrams of the model system. Red curves indicate stable steady states. Blue curves indicate unstable steady states. (A) Hill coefficient = 2.0 (default value). Threshold p53 concentration (position of solid arrow) equals 75.0 (nm) (B) Hill coefficient = 1.0. Threshold p53 concentration equals 11.0 (nm) (C) Hill coefficient = 4.0. Threshold p53 concentration equals (nm). Fig. 4. Apoptosis percentage (Hill coefficient = 2.0). Blue curve: pulse variability = 0.0, Red curve: pulse variability = 0.25, Yellow curve: pulse variability = 0.5, Green curve: pulse variability =
25 Fig. 5. Difference of apoptosis percentage among different Hill coefficient systems. (A) pulse variability = (B) pulse variability = 0.5. (C) pulse variability = 1.0. Blue curve: Hill coefficient = 1.0, Red curve: Hill coefficient = 2.0, Green curve: Hill coefficient = 4.0. Fig. 6. Apoptosis percentage with constant pulse height or pulse width or pulse interval. (A) (C) Pulse height was set to constant value. (A) pulse variability = (B) pulse variability = 0.5. (C) pulse variability = 1.0. (D) (F) Pulse width was set to constant value. (D) pulse variability = (E) pulse variability = 0.5. (F) pulse variability = 1.0. (G) (I) Pulse interval was set to constant value. (G) pulse variability = (H) pulse variability = 0.5. (I) pulse variability = 1.0. Blue curve: Hill coefficient = 1.0, Red curve: Hill coefficient = 2.0, Green curve: Hill coefficient =
26 Table 1. Ordinary differential equations of the model. = ksx kdx [Bax]-kxa [Bax][Act] + kax [AcBax] kaxx [AcBax][Bax] (1) = -kdax [AcBax] + kxa [Bax][Act]-kfaxb2 [AcBax][Bcl2] + kraxb2 [AcBax/Bcl2]-kfaxab2 [AcBax][Act/Bcl2] + kraxab2 [AcBax/Bcl2][Act] kax [AcBax] -kfaxsb2 [AcBax][PUMA/Bcl2] + kraxsb2 [AcBax/Bcl2][PUMA] kaxx [AcBax][Bax] 2 kfaxax [AcBax] kraxax [Pore] (2) = ksb2-kdb2 [Bcl2] -kfaxb2 [AcBax][Bcl2] + kraxb2 [AcBax/Bcl2]-kfab2 [Act][Bcl2] + krab2 [Act/Bcl2] -kfsb2 [PUMA][Bcl2] + krsb2 [PUMA/Bcl2] (3) = ksa kda [Act]-kfab2 [Act][Bcl2] + krab2 [Act/Bcl2] + kfaxab2 [AcBax][Act/Bcl2] -kraxab2 [AcBax/Bcl2][Act] kfasb2 [Act][PUMA/Bcl2] + krasb2 [Act/Bcl2][PUMA] (4) = -kdab2 [Act/Bcl2] + kfab2 [Act][Bcl2] 25
27 -krab2 [Act/Bcl2]-kfaxab2 [AcBax][Act/Bcl2] + kraxab2 [AcBax/Bcl2][Act] + kfasb2 [Act][PUMA/Bcl2] -krasb2 [Act/Bcl2][PUMA] (5) = -kdaxb2 [AcBax/Bcl2] + kfaxb2 [AcBax][Bcl2] -kraxb2 [AcBax/Bcl2] + kfaxab2 [AcBax][Act/Bcl2] -kraxab2 [AcBax/Bcl2][Act] + kfaxsb2 [AcBax][PUMA/Bcl2] -kraxsb2 [AcBax/Bcl2][PUMA] (6) = kss kds [PUMA]-kfsb2 [PUMA][Bcl2] + krsb2 [PUMA/Bcl2] + kfasb2 [Act][PUMA/Bcl2]-krasb2 [Act/Bcl2][PUMA] + kfaxsb2 [AcBax][PUMA/Bcl2] -kraxsb2 [AcBax/Bcl2][PUMA] + stimulus (7) = -kdsb2 [PUMA/Bcl2] + kfsb2 [PUMA][Bcl2] -krsb2 [PUMA/Bcl2]-kfasb2 [Act][PUMA/Bcl2] + krasb2 [Act/Bcl2][PUMA]-kfaxsb2 [AcBax][PUMA/Bcl2] + kraxsb2 [AcBax/Bcl2][PUMA] (8) = -kdpo [Pore] + kaxx [AcBax][Bax] + kfaxax [AcBax] 2 26
28 kraxax [Pore] (9) 27
29 Table 2. Kinetic parameters of the model. Variable Value unit Reference Kp53 5.0e + 2 nm (16) kssp53 2.0e + 1 nm min -1 (28) n 2.0e (30) ksx 2.0e + 1 nm min -1 (16) ksa 2.0e + 0 nm min -1 (16) ksb2 6.0e + 0 nm min -1 (16) kss 2.0e + 0 nm min -1 (16) kdx 3.0e-2 min -1 (16) kdax 2.0e-3 min -1 (16) kda 1.0e-2 min -1 (16) kdb2 2.0e-3 min -1 (16) kdab2 2.0e-3 min -1 (16) kdaxb2 1.0e-2 min -1 (16) kds 1.0e-3 min -1 (16) kdsb2 5.0e-3 min -1 (16) kdpo 1.0e-2 min -1 (16) kxa 1.0e-4 nm -1 min -1 (16) kfaxb2 1.0e-3 nm -1 min -1 (16) kraxb2 1.0e-3 min -1 (16) kfab2 1.0e-2 nm -1 min -1 (16) krab2 6.0e-2 min -1 (16) 28
30 kfaxab2 5.0e-4 nm -1 min -1 (16) kraxab2 1.0e-5 nm -1 min -1 (16) kax 1.0e-3 min -1 (16) kfsb2 1.0e-4 nm -1 min -1 (16) krsb2 1.0e-3 min -1 (16) kfasb2 5.0e-5 nm -1 min -1 (16) krasb2 5.0e-4 nm -1 min -1 (16) kfaxsb2 5.0e-4 nm -1 min -1 (16) kraxsb2 1.0e-2 nm -1 min -1 (16) kaxx 2.0e-4 nm -1 min -1 (16) kfaxax 2.0e-4 nm -1 min -1 (16) kraxax 1.0e-2 min -1 (16) 29
31 Table 3. Minimum pulse height needed for apoptosis. Table 3A. Hill coefficient = 1.0 p53 pulse number p53 pulse height (nm) Table 3B. Hill coefficient = 2.0 p53 pulse number p53 pulse height (nm) Table 3C. Hill coefficient = 4.0 p53 pulse number p53 pulse height (nm)
32 Table 4. Relationship between p53 concentration and the value of Stimulus term. Here shows pulse variability = 0.5 as an example. HC pulse height (nm) ST range (nm/min) RWS (nm/min) NRWS Abbreviations: HC: Hill coefficient. ST range: Stimulus term range RWS: Range width of Stimulus term. NRWS: Normalized range width of Stimulus term. 31
33 Fig.1 32
34 Fig.2.A Fig.2.B 33
35 Fig.3.A Fig.3.B Fig.3.C 34
36 Fig.4 35
37 Fig.5.A Fig.5.B Fig.5.C 36
38 37
Supplementary Information
 Supplementary Information Significance of p53 dynamics in regulating apoptosis in response to ionizing radiation, and polypharmacological strategies Bing Liu 1,, Divesh Bhatt 1,, Zoltan N. Oltvai 2, Joel
Supplementary Information Significance of p53 dynamics in regulating apoptosis in response to ionizing radiation, and polypharmacological strategies Bing Liu 1,, Divesh Bhatt 1,, Zoltan N. Oltvai 2, Joel
Elsevier B.V.; この論文は出版社版でありま Right 引用の際には出版社版をご確認ご利用ください This is
 Title Refractory cutaneous lichenoid sarc tranilast. Author(s) Nakahigashi, Kyoko; Kabashima, Kenj Utani, Atsushi; Miyachi, Yoshiki Citation Journal of the American Academy of 63(1): 171-172 Issue Date
Title Refractory cutaneous lichenoid sarc tranilast. Author(s) Nakahigashi, Kyoko; Kabashima, Kenj Utani, Atsushi; Miyachi, Yoshiki Citation Journal of the American Academy of 63(1): 171-172 Issue Date
microrna as a Potential Vector for the Propagation of Robustness in Protein Expression and Oscillatory Dynamics within a cerna Network
 microrna as a Potential Vector for the Propagation of Robustness in Protein Expression and Oscillatory Dynamics within a cerna Network Claude Gérard*,Béla Novák Oxford Centre for Integrative Systems Biology,
microrna as a Potential Vector for the Propagation of Robustness in Protein Expression and Oscillatory Dynamics within a cerna Network Claude Gérard*,Béla Novák Oxford Centre for Integrative Systems Biology,
Supporting information (SI) Cell fate decision mediated by p53 pulses Xiao-Peng Zhang, Feng Liu, Zhang Cheng, and Wei Wang
 Supporting information (SI) Cell fate decision mediated by p53 pulses Xiao-Peng Zhang, Feng Liu, Zhang Cheng, and Wei Wang In this work, we aim at the p53 signaling network in untransformed mammalian cells,
Supporting information (SI) Cell fate decision mediated by p53 pulses Xiao-Peng Zhang, Feng Liu, Zhang Cheng, and Wei Wang In this work, we aim at the p53 signaling network in untransformed mammalian cells,
PDCD5-Regulated Cell Fate Decision after Ultraviolet-Irradiation-Induced DNA Damage
 2582 Biophysical Journal Volume December 2 2582 259 PDCD5-Regulated Cell Fate Decision after Ultraviolet-Irradiation-Induced DNA Damage Changjing Zhuge, Ying Chang, Yanjun Li, Yingyu Chen, and Jinzhi Lei
2582 Biophysical Journal Volume December 2 2582 259 PDCD5-Regulated Cell Fate Decision after Ultraviolet-Irradiation-Induced DNA Damage Changjing Zhuge, Ying Chang, Yanjun Li, Yingyu Chen, and Jinzhi Lei
Molecular biology :- Cancer genetics lecture 11
 Molecular biology :- Cancer genetics lecture 11 -We have talked about 2 group of genes that is involved in cellular transformation : proto-oncogenes and tumour suppressor genes, and it isn t enough to
Molecular biology :- Cancer genetics lecture 11 -We have talked about 2 group of genes that is involved in cellular transformation : proto-oncogenes and tumour suppressor genes, and it isn t enough to
この論文は出版社版でありません 引用の際には出版社版をご確認ご利用ください This is not the publis version. Please cite only the publi
 Title A case of progressive digital ische gemcitabine and S-1 in a patient wi Zaima, C; Kanai, M; Ishikawa, S; Ka Author(s) Mori, Y; Nishimura, T; Matsumoto, S T; Mimori, T Citation Japanese journal of
Title A case of progressive digital ische gemcitabine and S-1 in a patient wi Zaima, C; Kanai, M; Ishikawa, S; Ka Author(s) Mori, Y; Nishimura, T; Matsumoto, S T; Mimori, T Citation Japanese journal of
Modeling of Cancer Signaling Pathways
 Modeling of Cancer Signaling Pathways by Remziye Karabekmez A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree of Master of Mathematics in Applied
Modeling of Cancer Signaling Pathways by Remziye Karabekmez A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree of Master of Mathematics in Applied
Stochastic Simulation of P53, MDM2 Interaction
 Human Journals Research Article May 2017 Vol.:6, Issue:3 All rights are reserved by B.Odgerel et al. Stochastic Simulation of P53, MDM2 Interaction Keywords: DNA damage, P53, MDM2 interactions, propensity,
Human Journals Research Article May 2017 Vol.:6, Issue:3 All rights are reserved by B.Odgerel et al. Stochastic Simulation of P53, MDM2 Interaction Keywords: DNA damage, P53, MDM2 interactions, propensity,
Introduction to pathology lecture 5/ Cell injury apoptosis. Dr H Awad 2017/18
 Introduction to pathology lecture 5/ Cell injury apoptosis Dr H Awad 2017/18 Apoptosis = programmed cell death = cell suicide= individual cell death Apoptosis cell death induced by a tightly regulated
Introduction to pathology lecture 5/ Cell injury apoptosis Dr H Awad 2017/18 Apoptosis = programmed cell death = cell suicide= individual cell death Apoptosis cell death induced by a tightly regulated
MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1. Lecture 27: Systems Biology and Bayesian Networks
![MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1. Lecture 27: Systems Biology and Bayesian Networks MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1. Lecture 27: Systems Biology and Bayesian Networks](/thumbs/80/82116384.jpg) MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1 Lecture 27: Systems Biology and Bayesian Networks Systems Biology and Regulatory Networks o Definitions o Network motifs o Examples
MBios 478: Systems Biology and Bayesian Networks, 27 [Dr. Wyrick] Slide #1 Lecture 27: Systems Biology and Bayesian Networks Systems Biology and Regulatory Networks o Definitions o Network motifs o Examples
Apoptosis Chapter 9. Neelu Yadav PhD
 Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org 1 Apoptosis: Lecture outline Apoptosis a programmed cell death pathway in normal homeostasis Core Apoptosis cascade is conserved Compare
Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org 1 Apoptosis: Lecture outline Apoptosis a programmed cell death pathway in normal homeostasis Core Apoptosis cascade is conserved Compare
Apoptosis Oncogenes. Srbová Martina
 Apoptosis Oncogenes Srbová Martina Cell Cycle Control point Cyclin B Cdk1 Cyclin D Cdk4 Cdk6 Cyclin A Cdk2 Cyclin E Cdk2 Cyclin-dependent kinase (Cdk) have to bind a cyclin to become active Regulation
Apoptosis Oncogenes Srbová Martina Cell Cycle Control point Cyclin B Cdk1 Cyclin D Cdk4 Cdk6 Cyclin A Cdk2 Cyclin E Cdk2 Cyclin-dependent kinase (Cdk) have to bind a cyclin to become active Regulation
Apoptotic Pathways in Mammals Dr. Douglas R. Green
 Apoptotic Pathways in Mammals Douglas R. Green 1 Apoptosis A form of cell death that is defined morphologically, and features a number of biochemical events Programmed cell death Cell death that occurs
Apoptotic Pathways in Mammals Douglas R. Green 1 Apoptosis A form of cell death that is defined morphologically, and features a number of biochemical events Programmed cell death Cell death that occurs
Research Communication
 IUBMB Life, 58(11): 659 663, November 2006 Research Communication Sequestration Shapes the Response of Signal Transduction Cascades Nils Blu thgen Molecular Neurobiology, Free University Berlin, and Institute
IUBMB Life, 58(11): 659 663, November 2006 Research Communication Sequestration Shapes the Response of Signal Transduction Cascades Nils Blu thgen Molecular Neurobiology, Free University Berlin, and Institute
MicroRNA-mediated incoherent feedforward motifs are robust
 The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
The Second International Symposium on Optimization and Systems Biology (OSB 8) Lijiang, China, October 31 November 3, 8 Copyright 8 ORSC & APORC, pp. 62 67 MicroRNA-mediated incoherent feedforward motifs
Biochemistry 673 Your Name: Regulation of Metabolism Assoc. Prof. Jason Kahn Final Exam (150 points total) May 17, 2005
 Biochemistry 673 Your Name: Regulation of Metabolism Assoc. Prof. Jason Kahn Final Exam (150 points total) May 17, 2005 You have 120 minutes for this exam. Explanations should be concise and clear. Generous
Biochemistry 673 Your Name: Regulation of Metabolism Assoc. Prof. Jason Kahn Final Exam (150 points total) May 17, 2005 You have 120 minutes for this exam. Explanations should be concise and clear. Generous
Pancreatic Cancer Research and HMGB1 Signaling Pathway
 Pancreatic Cancer Research and HMGB1 Signaling Pathway Haijun Gong*, Paolo Zuliani*, Anvesh Komuravelli*, James R. Faeder #, Edmund M. Clarke* * # The Hallmarks of Cancer D. Hanahan and R. A. Weinberg
Pancreatic Cancer Research and HMGB1 Signaling Pathway Haijun Gong*, Paolo Zuliani*, Anvesh Komuravelli*, James R. Faeder #, Edmund M. Clarke* * # The Hallmarks of Cancer D. Hanahan and R. A. Weinberg
p53 and Apoptosis: Master Guardian and Executioner Part 2
 p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
Supplementary Material
 Supplementary Material Supplementary Text Text S1. Time distributions of the high FRET efficiency at different concentrations of EF-G.GTP From Fig. 1, the model of ribosomal translocation at non-saturating
Supplementary Material Supplementary Text Text S1. Time distributions of the high FRET efficiency at different concentrations of EF-G.GTP From Fig. 1, the model of ribosomal translocation at non-saturating
Stochastic simulations
 Stochastic simulations Application to circadian clocks Didier Gonze Circadian rhythms Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Circadian rhythms
Stochastic simulations Application to circadian clocks Didier Gonze Circadian rhythms Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Circadian rhythms
Citation Applied Surface Science (2009), Right 論文は出版社版でありません 引用の際には出版社版をご確認ご利用ください
 Title Emission spectroscopy of laser abla analysis of a target in water Author(s) Sakka, Tetsuo; Yamagata, Hajimu; Og Kazuhiro; Ogata, Yukio H. Citation Applied Surface Science (2009), 255 Issue Date 2009-09
Title Emission spectroscopy of laser abla analysis of a target in water Author(s) Sakka, Tetsuo; Yamagata, Hajimu; Og Kazuhiro; Ogata, Yukio H. Citation Applied Surface Science (2009), 255 Issue Date 2009-09
Cancer: Brief Introduction. First stage: Mutations in genes progressively accumulate so that there is unrestrained cell proliferation
 Cancer: Brief Introduction First stage: Mutations in genes progressively accumulate so that there is unrestrained cell proliferation There has to be mutational amplification Over lifetime about 10 16 divisions
Cancer: Brief Introduction First stage: Mutations in genes progressively accumulate so that there is unrestrained cell proliferation There has to be mutational amplification Over lifetime about 10 16 divisions
Citation Congenital Anomalies (2016), 56(2): version. この論文は出版社版でありません 引用の際には出版社版をご確認ご利用ください
 Title Correlation of external ear human embryos auricle Author(s) Ozeki-Sato, Maimi; Yamada, Shigehit Ishizu, Koichi; Takakuwa, Tetsuya Citation Congenital Anomalies (2016), 56(2): Issue Date 2016-03 URL
Title Correlation of external ear human embryos auricle Author(s) Ozeki-Sato, Maimi; Yamada, Shigehit Ishizu, Koichi; Takakuwa, Tetsuya Citation Congenital Anomalies (2016), 56(2): Issue Date 2016-03 URL
A Two-Step Mechanism for Cell Fate Decision by Coordination of Nuclear and Mitochondrial p53 Activities
 by Coordination of Nuclear and Mitochondrial p53 Activities Xiao-Jun Tian, Feng Liu*, Xiao-Peng Zhang, Jun Li, Wei Wang* National Laboratory of Solid State Microstructures and Department of Physics, Nanjing
by Coordination of Nuclear and Mitochondrial p53 Activities Xiao-Jun Tian, Feng Liu*, Xiao-Peng Zhang, Jun Li, Wei Wang* National Laboratory of Solid State Microstructures and Department of Physics, Nanjing
#19 Apoptosis Chapter 9. Neelu Yadav PhD
 #19 Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org Why cells decide to die? - Stress, harmful, not needed - Completed its life span Death stimulation or Stress Cell Survival Death Functions
#19 Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org Why cells decide to die? - Stress, harmful, not needed - Completed its life span Death stimulation or Stress Cell Survival Death Functions
Citation Journal of Cardiology Cases (2013), Ltd.; この論文は出版社版でありません 引用の際には出版社版をご確認ご利用ください This is not the publ
 Title Pathological analyses of very longimplantation in human coronary arte Author(s) Imai, Masao; Kimura, Takeshi; Tazak Erika; Inoue, Katsumi Citation Journal of Cardiology Cases (01), Issue Date 01-
Title Pathological analyses of very longimplantation in human coronary arte Author(s) Imai, Masao; Kimura, Takeshi; Tazak Erika; Inoue, Katsumi Citation Journal of Cardiology Cases (01), Issue Date 01-
Identification of influential proteins in the classical retinoic acid signaling pathway
 Ghaffari and Petzold Theoretical Biology and Medical Modelling (2018) 15:16 https://doi.org/10.1186/s12976-018-0088-7 RESEARCH Open Access Identification of influential proteins in the classical retinoic
Ghaffari and Petzold Theoretical Biology and Medical Modelling (2018) 15:16 https://doi.org/10.1186/s12976-018-0088-7 RESEARCH Open Access Identification of influential proteins in the classical retinoic
Cell cycle and Apoptosis. Chalermchai Mitrpant
 Cell cycle and Apoptosis 2556 Chalermchai Mitrpant Overview of the cell cycle Outline Regulatory mechanisms controlling cell cycle Progression of the cell cycle Checkpoint of the cell cycle Phases of the
Cell cycle and Apoptosis 2556 Chalermchai Mitrpant Overview of the cell cycle Outline Regulatory mechanisms controlling cell cycle Progression of the cell cycle Checkpoint of the cell cycle Phases of the
Stochastic simulations
 Circadian rhythms Stochastic simulations Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Application to circadian clocks Didier Gonze Circadian rhythms
Circadian rhythms Stochastic simulations Circadian rhythms allow living organisms to live in phase with the alternance of day and night... Application to circadian clocks Didier Gonze Circadian rhythms
Supplementary Figures
 Supplementary Figures Figure S1. Validation of kinase regulators of ONC201 sensitivity. Validation and screen results for changes in cell viability associated with the combination of ONC201 treatment (1
Supplementary Figures Figure S1. Validation of kinase regulators of ONC201 sensitivity. Validation and screen results for changes in cell viability associated with the combination of ONC201 treatment (1
Early cell death (FGF) B No RunX transcription factor produced Yes No differentiation
 Solution Key - Practice Questions Question 1 a) A recent publication has shown that the fat stem cells (FSC) can act as bone stem cells to repair cavities in the skull, when transplanted into immuno-compromised
Solution Key - Practice Questions Question 1 a) A recent publication has shown that the fat stem cells (FSC) can act as bone stem cells to repair cavities in the skull, when transplanted into immuno-compromised
Signaling Apoptosis. Scott André Oakes, M.D. Dept. of Pathology Univ. of Calif-San Francisco. Cyt c Release BAX/BAK. Apoptosome Formation
 Signaling Apoptosis Cyt c Release BAX/BAK Apoptosome Formation Caspase Activation Scott André Oakes, M.D. Dept. of Pathology Univ. of Calif-San Francisco Why do we need cell death? Sculpt Organs Control
Signaling Apoptosis Cyt c Release BAX/BAK Apoptosome Formation Caspase Activation Scott André Oakes, M.D. Dept. of Pathology Univ. of Calif-San Francisco Why do we need cell death? Sculpt Organs Control
Author(s) Umemura, Kenji; Sugihara, Osamu; Ka. Citation Journal of Wood Science (2013), 59(
 Title Investigation of a new natural adhe and sucrose for particleboard Author(s) Umemura, Kenji; Sugihara, Osamu; Ka Citation Journal of Wood Science (2013), 59( Issue Date 2013-06 URL http://hdl.handle.net/2433/189837
Title Investigation of a new natural adhe and sucrose for particleboard Author(s) Umemura, Kenji; Sugihara, Osamu; Ka Citation Journal of Wood Science (2013), 59( Issue Date 2013-06 URL http://hdl.handle.net/2433/189837
MATHEMATICAL MODELING OF PERIFUSION CELL CULTURE EXPERIMENTS ON GNRH SIGNALING
 MATHEMATICAL MODELING OF PERIFUSION CELL CULTURE EXPERIMENTS ON GNRH SIGNALING N Ezgi Temamogullari a,, H Frederik Nijhout b, Michael C Reed a arxiv:1606.00463v1 [q-bio.cb] 9 Dec 2015 Abstract a Department
MATHEMATICAL MODELING OF PERIFUSION CELL CULTURE EXPERIMENTS ON GNRH SIGNALING N Ezgi Temamogullari a,, H Frederik Nijhout b, Michael C Reed a arxiv:1606.00463v1 [q-bio.cb] 9 Dec 2015 Abstract a Department
Lecture 14 - The cell cycle and cell death
 02.17.10 Lecture 14 - The cell cycle and cell death The cell cycle: cells duplicate their contents and divide The cell cycle may be divided into 4 phases The cell cycle triggers essential processes (DNA
02.17.10 Lecture 14 - The cell cycle and cell death The cell cycle: cells duplicate their contents and divide The cell cycle may be divided into 4 phases The cell cycle triggers essential processes (DNA
Consecutive Positive Feedback Loops Create a Bistable Switch that Controls Preadipocyte-to-Adipocyte Conversion
 Cell Reports Article Consecutive Positive Feedback Loops Create a Bistable Switch that Controls Preadipocyte-to-Adipocyte Conversion Byung Ouk Park, 1 Robert Ahrends, 1 and Mary N. Teruel 1, * 1 Department
Cell Reports Article Consecutive Positive Feedback Loops Create a Bistable Switch that Controls Preadipocyte-to-Adipocyte Conversion Byung Ouk Park, 1 Robert Ahrends, 1 and Mary N. Teruel 1, * 1 Department
A simple theoretical framework for understanding heterogeneous differentiation of CD4 + T cells
 Hong et al. BMC Systems Biology 2012, 6:66 RESEARCH ARTICLE Open Access A simple theoretical framework for understanding heterogeneous differentiation of CD4 + T cells Tian Hong 1, Jianhua Xing 2, Liwu
Hong et al. BMC Systems Biology 2012, 6:66 RESEARCH ARTICLE Open Access A simple theoretical framework for understanding heterogeneous differentiation of CD4 + T cells Tian Hong 1, Jianhua Xing 2, Liwu
Title processing in milliseconds in rats. Citation Behavioural brain research (2014),
 Title Novel behavioral tasks to explore c processing in milliseconds in rats. Author(s) Yamaguchi, Kenji; Sakurai, Yoshio Citation Behavioural brain research (2014), Issue Date 2014-04-15 URL http://hdl.handle.net/2433/182931
Title Novel behavioral tasks to explore c processing in milliseconds in rats. Author(s) Yamaguchi, Kenji; Sakurai, Yoshio Citation Behavioural brain research (2014), Issue Date 2014-04-15 URL http://hdl.handle.net/2433/182931
Representational Switching by Dynamical Reorganization of Attractor Structure in a Network Model of the Prefrontal Cortex
 Representational Switching by Dynamical Reorganization of Attractor Structure in a Network Model of the Prefrontal Cortex Yuichi Katori 1,2 *, Kazuhiro Sakamoto 3, Naohiro Saito 4, Jun Tanji 4, Hajime
Representational Switching by Dynamical Reorganization of Attractor Structure in a Network Model of the Prefrontal Cortex Yuichi Katori 1,2 *, Kazuhiro Sakamoto 3, Naohiro Saito 4, Jun Tanji 4, Hajime
Supplementary Figures
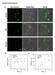 Supplementary Figures Supplementary Figure 1. Single cell gene expression analysis during collective cell migration. (a) Florescence and bright-field images of MCF7 cells transfected with dslna probes
Supplementary Figures Supplementary Figure 1. Single cell gene expression analysis during collective cell migration. (a) Florescence and bright-field images of MCF7 cells transfected with dslna probes
A Self-Propelled Biological Process Plk1-Dependent Product- Activated, Feed-Forward Mechanism
 A Self-Propelled Biological Process Plk1-Dependent Product- Activated, Feed-Forward Mechanism The Harvard community has made this article openly available. Please share how this access benefits you. Your
A Self-Propelled Biological Process Plk1-Dependent Product- Activated, Feed-Forward Mechanism The Harvard community has made this article openly available. Please share how this access benefits you. Your
Chapter 1. Introduction
 Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
Hypothesis Testing with Evolutionary and Systems Biology Models. Seongho Ryu
 Hypothesis Testing with Evolutionary and Systems Biology Models by Seongho Ryu A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Biology
Hypothesis Testing with Evolutionary and Systems Biology Models by Seongho Ryu A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Biology
BioNSi: A Discrete Biological Network Simulator Tool
 BioNSi: A Discrete Biological Network Simulator Tool Amir Rubinstein 1, Noga Bracha 1, Liat Rudner 1, Noga Zucker 1, Hadas E. Sloin 2, and Benny Chor 1 1 Blavatnik School of Computer Science, Tel Aviv
BioNSi: A Discrete Biological Network Simulator Tool Amir Rubinstein 1, Noga Bracha 1, Liat Rudner 1, Noga Zucker 1, Hadas E. Sloin 2, and Benny Chor 1 1 Blavatnik School of Computer Science, Tel Aviv
Information-theoretic stimulus design for neurophysiology & psychophysics
 Information-theoretic stimulus design for neurophysiology & psychophysics Christopher DiMattina, PhD Assistant Professor of Psychology Florida Gulf Coast University 2 Optimal experimental design Part 1
Information-theoretic stimulus design for neurophysiology & psychophysics Christopher DiMattina, PhD Assistant Professor of Psychology Florida Gulf Coast University 2 Optimal experimental design Part 1
Programmed Cell Death (apoptosis)
 Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Chapter 1: Exploring Data
 Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Annals of RSCB Vol. XVI, Issue 1
 ENDOPLASMIC RETICULUM INVOLVEMENT IN APOPTOSIS OF NORMAL AND TREATED GINGIVAL FIBROBLASTS Ancuţa Goriuc, Raluca Jipu, Roxana Irina Iancu, M. Costuleanu GR. T. POPA UNIVERSITY OF MEDICINE AND PHARMACY,
ENDOPLASMIC RETICULUM INVOLVEMENT IN APOPTOSIS OF NORMAL AND TREATED GINGIVAL FIBROBLASTS Ancuţa Goriuc, Raluca Jipu, Roxana Irina Iancu, M. Costuleanu GR. T. POPA UNIVERSITY OF MEDICINE AND PHARMACY,
Visualizing Cancer Heterogeneity with Dynamic Flow
 Visualizing Cancer Heterogeneity with Dynamic Flow Teppei Nakano and Kazuki Ikeda Keio University School of Medicine, Tokyo 160-8582, Japan keiohigh2nd@gmail.com Department of Physics, Osaka University,
Visualizing Cancer Heterogeneity with Dynamic Flow Teppei Nakano and Kazuki Ikeda Keio University School of Medicine, Tokyo 160-8582, Japan keiohigh2nd@gmail.com Department of Physics, Osaka University,
Ch.20 Dynamic Cue Combination in Distributional Population Code Networks. Ka Yeon Kim Biopsychology
 Ch.20 Dynamic Cue Combination in Distributional Population Code Networks Ka Yeon Kim Biopsychology Applying the coding scheme to dynamic cue combination (Experiment, Kording&Wolpert,2004) Dynamic sensorymotor
Ch.20 Dynamic Cue Combination in Distributional Population Code Networks Ka Yeon Kim Biopsychology Applying the coding scheme to dynamic cue combination (Experiment, Kording&Wolpert,2004) Dynamic sensorymotor
Mathematical and Computational Analysis of Adaptation via Feedback Inhibition in Signal Transduction Pathways
 806 Biophysical Journal Volume 93 August 2007 806 821 Mathematical and Computational Analysis of Adaptation via Feedback Inhibition in Signal Transduction Pathways Marcelo Behar,* Nan Hao, y Henrik G.
806 Biophysical Journal Volume 93 August 2007 806 821 Mathematical and Computational Analysis of Adaptation via Feedback Inhibition in Signal Transduction Pathways Marcelo Behar,* Nan Hao, y Henrik G.
The cell cycle plays a prominent role in development,
 Dynamics of the mammalian cell cycle in physiological and pathological conditions Claude Gérard 1 and Albert Goldbeter 1,2 * A network of cyclin-dependent kinases (Cdks) controls progression along the
Dynamics of the mammalian cell cycle in physiological and pathological conditions Claude Gérard 1 and Albert Goldbeter 1,2 * A network of cyclin-dependent kinases (Cdks) controls progression along the
Silibinin i activates p53-caspase-2 pathway and causes caspase-mediated cleavage of Cip1/p21 in apoptosis
 Silibinin i activates p53-caspase-2 pathway and causes caspase-mediated cleavage of Cip1/p21 in apoptosis induction in bladder transitional-cell papilloma RT4 cells: Evidence for a regulatory loop between
Silibinin i activates p53-caspase-2 pathway and causes caspase-mediated cleavage of Cip1/p21 in apoptosis induction in bladder transitional-cell papilloma RT4 cells: Evidence for a regulatory loop between
Author(s) Takemoto, Mitsuru; Nakamura, Takash. Citation Skeletal radiology (2011), 40(12):
 Title Paraarticular osteochondroma of a c presenting as myelopathy. Author(s) Okamoto, Takeshi; Neo, Masashi; Fuj Takemoto, Mitsuru; Nakamura, Takash Citation Skeletal radiology (0), 0(): Issue Date 0-
Title Paraarticular osteochondroma of a c presenting as myelopathy. Author(s) Okamoto, Takeshi; Neo, Masashi; Fuj Takemoto, Mitsuru; Nakamura, Takash Citation Skeletal radiology (0), 0(): Issue Date 0-
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cancer develops after somatic mutations overcome the multiple
 Genetic variation in cancer predisposition: Mutational decay of a robust genetic control network Steven A. Frank* Department of Ecology and Evolutionary Biology, University of California, Irvine, CA 92697-2525
Genetic variation in cancer predisposition: Mutational decay of a robust genetic control network Steven A. Frank* Department of Ecology and Evolutionary Biology, University of California, Irvine, CA 92697-2525
PRC2 crystal clear. Matthieu Schapira
 PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
NF -κb oscillations and cell-to-cell variability
 NF -κb oscillations and cell-to-cell variability arxiv:q-bio/0510025v1 [q-bio.qm] 13 Oct 2005 F. Hayot and C. Jayaprakash Department of Physics Ohio State University Columbus, OH 43210-1106, USA February
NF -κb oscillations and cell-to-cell variability arxiv:q-bio/0510025v1 [q-bio.qm] 13 Oct 2005 F. Hayot and C. Jayaprakash Department of Physics Ohio State University Columbus, OH 43210-1106, USA February
Title: LATS2 is De-methylated and Overexpressed in Nasopharyngeal Carcinoma and Predicts Poor Prognosis of the Patients
 Author's response to reviews Title: LATS2 is De-methylated and Overexpressed in Nasopharyngeal Carcinoma and Predicts Poor Prognosis of the Patients Authors: Yan Zhang (zhangy2@sysucc.org.cn) Chun-Fang
Author's response to reviews Title: LATS2 is De-methylated and Overexpressed in Nasopharyngeal Carcinoma and Predicts Poor Prognosis of the Patients Authors: Yan Zhang (zhangy2@sysucc.org.cn) Chun-Fang
Cells under stresses such as DNA damage, hypoxia, and
 A plausible model for the digital response of p53 to DNA damage Lan Ma, John Wagner, John Jeremy Rice, Wenwei Hu, Arnold J. Levine, and Gustavo A. Stolovitzky IBM Thomas J. Watson Research Center, Yorktown
A plausible model for the digital response of p53 to DNA damage Lan Ma, John Wagner, John Jeremy Rice, Wenwei Hu, Arnold J. Levine, and Gustavo A. Stolovitzky IBM Thomas J. Watson Research Center, Yorktown
AperTO - Archivio Istituzionale Open Access dell'università di Torino
 AperTO - Archivio Istituzionale Open Access dell'università di Torino From the nucleus to the mitochondria and backthe odyssey of a multitask STAT3 This is the author's manuscript Original Citation: From
AperTO - Archivio Istituzionale Open Access dell'università di Torino From the nucleus to the mitochondria and backthe odyssey of a multitask STAT3 This is the author's manuscript Original Citation: From
Modeling of epidemic spreading with white Gaussian noise
 Article Statistical Physics and Mathematics for Complex Systems December 20 Vol.56 No.34: 3683 3688 doi: 0.007/s434-0-4753-z SPECIAL TOPICS: Modeling of epidemic spreading with white Gaussian noise GU
Article Statistical Physics and Mathematics for Complex Systems December 20 Vol.56 No.34: 3683 3688 doi: 0.007/s434-0-4753-z SPECIAL TOPICS: Modeling of epidemic spreading with white Gaussian noise GU
Mechanisms of Cell Death
 Mechanisms of Cell Death CELL DEATH AND FORMATION OF THE SEMICIRCULAR CANALS Carol M. Troy August 25, 2008 FROM: Fekete et al., Development 124: 2451 (1997) PHENOMENOLOGY OF CELL DEATH I. DEVELOPMENT A.
Mechanisms of Cell Death CELL DEATH AND FORMATION OF THE SEMICIRCULAR CANALS Carol M. Troy August 25, 2008 FROM: Fekete et al., Development 124: 2451 (1997) PHENOMENOLOGY OF CELL DEATH I. DEVELOPMENT A.
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
2012 Course: The Statistician Brain: the Bayesian Revolution in Cognitive Sciences
 2012 Course: The Statistician Brain: the Bayesian Revolution in Cognitive Sciences Stanislas Dehaene Chair of Experimental Cognitive Psychology Lecture n 5 Bayesian Decision-Making Lecture material translated
2012 Course: The Statistician Brain: the Bayesian Revolution in Cognitive Sciences Stanislas Dehaene Chair of Experimental Cognitive Psychology Lecture n 5 Bayesian Decision-Making Lecture material translated
Application of Bayesian Probabilistic Approach on Ground Motion Attenuation Relations. Rong-Rong Xu. Master of Science in Civil Engineering
 Application of Bayesian Probabilistic Approach on Ground Motion Attenuation Relations by Rong-Rong Xu Master of Science in Civil Engineering August 2013 Faculty of Science and Technology University of
Application of Bayesian Probabilistic Approach on Ground Motion Attenuation Relations by Rong-Rong Xu Master of Science in Civil Engineering August 2013 Faculty of Science and Technology University of
#19 Apoptosis Chapter 9. Neelu Yadav PhD
 #19 Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org Why cells decide to die? - Stress, harmful, not needed - Completed its life span Death stimulation or Stress Cell Survival Death Functions
#19 Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org Why cells decide to die? - Stress, harmful, not needed - Completed its life span Death stimulation or Stress Cell Survival Death Functions
Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning
 Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning Adriana Birlutiu (1),(2), Paul Bulzu (1), Irina Iereminciuc (1), and Alexandru Floares (1) (1) SAIA & OncoPredict, Cluj-Napoca,
Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning Adriana Birlutiu (1),(2), Paul Bulzu (1), Irina Iereminciuc (1), and Alexandru Floares (1) (1) SAIA & OncoPredict, Cluj-Napoca,
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Title matching process in object recognit. Right でありません 引用の際には出版社版をご確認ご利用ください
 Title Effect of active exploration of 3-D matching process in object recognit Author(s) Sasaoka, Takafumi; Asakura, Nobuhik Citation Perception (2010), 39(3): 289-308 Issue Date 2010-03 URL http://hdl.handle.net/2433/139452
Title Effect of active exploration of 3-D matching process in object recognit Author(s) Sasaoka, Takafumi; Asakura, Nobuhik Citation Perception (2010), 39(3): 289-308 Issue Date 2010-03 URL http://hdl.handle.net/2433/139452
REGULATION OF ENZYME ACTIVITY. Medical Biochemistry, Lecture 25
 REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 Name: Key 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function
BIO360 Fall 2013 Quiz 1 Name: Key 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function
Effects of Light Stimulus Frequency on Phase Characteristics of Brain Waves
 SICE Annual Conference 27 Sept. 17-2, 27, Kagawa University, Japan Effects of Light Stimulus Frequency on Phase Characteristics of Brain Waves Seiji Nishifuji 1, Kentaro Fujisaki 1 and Shogo Tanaka 1 1
SICE Annual Conference 27 Sept. 17-2, 27, Kagawa University, Japan Effects of Light Stimulus Frequency on Phase Characteristics of Brain Waves Seiji Nishifuji 1, Kentaro Fujisaki 1 and Shogo Tanaka 1 1
ZOLTAN SZALLASI Department of Pharmacology, Uniformed Services University of the Health Sciences, Bethesda, MD,
 MODELING THE NORMAL AND NEOPLASTIC CELL CYCLE WITH REALISTIC BOOLEAN GENETIC NETWORKS : THEIR APPLICATION FOR UNDERSTANDING CARCINOGENESIS AND ASSESSING THERAPEUTIC STRATEGIES. ZOLTAN SZALLASI Department
MODELING THE NORMAL AND NEOPLASTIC CELL CYCLE WITH REALISTIC BOOLEAN GENETIC NETWORKS : THEIR APPLICATION FOR UNDERSTANDING CARCINOGENESIS AND ASSESSING THERAPEUTIC STRATEGIES. ZOLTAN SZALLASI Department
Mitochondria in apoptosis. Jean-Claude Martinou, MD, Ph.D Department of cell biology University of Geneva Geneva, Switzerland
 Mitochondria in apoptosis Jean-Claude Martinou, MD, Ph.D Department of cell biology University of Geneva Geneva, Switzerland The dual role of mitochondria in life and death QuickTime and a TIFF (Uncompressed)
Mitochondria in apoptosis Jean-Claude Martinou, MD, Ph.D Department of cell biology University of Geneva Geneva, Switzerland The dual role of mitochondria in life and death QuickTime and a TIFF (Uncompressed)
Title: Molecular Constraints on Synaptic Tagging and Maintenance of Long-Term Potentiation: A Predictive Model
 Title: Molecular Constraints on Synaptic Tagging and Maintenance of Long-Term Potentiation: A Predictive Model Authors: Paul Smolen, Douglas A. Baxter, and John H. Byrne Laboratory of Origin: Department
Title: Molecular Constraints on Synaptic Tagging and Maintenance of Long-Term Potentiation: A Predictive Model Authors: Paul Smolen, Douglas A. Baxter, and John H. Byrne Laboratory of Origin: Department
GMS 6644: Apoptosis. Introduction
 GMS 6644: Apoptosis Introduction (Feb. 15, 2006) Lei Xiao, Ph.D. Department of Anatomy & Cell Biology UF Shands Cancer Center ARB Rm R4-250, 846-1199, lxiao@ufl.edu Outline of the Lecture Different types
GMS 6644: Apoptosis Introduction (Feb. 15, 2006) Lei Xiao, Ph.D. Department of Anatomy & Cell Biology UF Shands Cancer Center ARB Rm R4-250, 846-1199, lxiao@ufl.edu Outline of the Lecture Different types
Supplemental Material
 Supplemental Material Recording technique Multi-unit activity (MUA) was recorded from electrodes that were chronically implanted (Teflon-coated platinum-iridium wires) in the primary visual cortex representing
Supplemental Material Recording technique Multi-unit activity (MUA) was recorded from electrodes that were chronically implanted (Teflon-coated platinum-iridium wires) in the primary visual cortex representing
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Citation Psychological science (2016), 27(2) 論文は出版社版でありません 引用の際には出版社版をご確認ご利用ください
 Title Location-Unbound Color-Shape Bindin Visual Working Memory. Author(s) Saiki, Jun Citation Psychological science (2016), 27(2) Issue Date 2016-02-09 URL http://hdl.handle.net/2433/202916 The final,
Title Location-Unbound Color-Shape Bindin Visual Working Memory. Author(s) Saiki, Jun Citation Psychological science (2016), 27(2) Issue Date 2016-02-09 URL http://hdl.handle.net/2433/202916 The final,
Problem-solving Test: The Mechanism of Protein Synthesis
 Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Balancing speed and accuracy of polyclonal T cell activation: A role for extracellular feedback
 Balancing speed and accuracy of polyclonal T cell activation: A role for extracellular feedback The Harvard community has made this article openly available. Please share how this access benefits you.
Balancing speed and accuracy of polyclonal T cell activation: A role for extracellular feedback The Harvard community has made this article openly available. Please share how this access benefits you.
Molecular Constraints on Synaptic Tagging and Maintenance of Long-Term Potentiation: A Predictive Model
 Molecular Constraints on Synaptic Tagging and Maintenance of Long-Term Potentiation: A Predictive Model Paul Smolen, Douglas A. Baxter, and John H. Byrne Department of Neurobiology and Anatomy W. M. Keck
Molecular Constraints on Synaptic Tagging and Maintenance of Long-Term Potentiation: A Predictive Model Paul Smolen, Douglas A. Baxter, and John H. Byrne Department of Neurobiology and Anatomy W. M. Keck
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Title water intake in dry and lactating c. Author(s) Kume, S.; Nonaka, K.; Oshita, T.; K. Citation Livestock Science (2010), 128(1-3):
 Title Evaluation of drinking water intake water intake in dry and lactating c Author(s) Kume, S.; Nonaka, K.; Oshita, T.; K Citation Livestock Science (10), 128(1-3): Issue Date 10-03 URL http://hdl.handle.net/2433/102278
Title Evaluation of drinking water intake water intake in dry and lactating c Author(s) Kume, S.; Nonaka, K.; Oshita, T.; K Citation Livestock Science (10), 128(1-3): Issue Date 10-03 URL http://hdl.handle.net/2433/102278
Part I Molecular Cell Biology
 1 Part I Molecular Cell Biology RNA Regulation: Advances in Molecular Biology and Medicine, First Edition. Edited by Robert A. Meyers. 2014 Wiley-VCH Verlag GmbH & Co. KGaA. Published 2014 by Wiley-VCH
1 Part I Molecular Cell Biology RNA Regulation: Advances in Molecular Biology and Medicine, First Edition. Edited by Robert A. Meyers. 2014 Wiley-VCH Verlag GmbH & Co. KGaA. Published 2014 by Wiley-VCH
Chapter 7 Conclusions
 VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
Definition 1: A fixed point iteration scheme to approximate the fixed point, p, of a function g, = for all n 1 given a starting approximation, p.
 Supplemental Material: A. Proof of Convergence In this Appendix, we provide a computational proof that the circadian adjustment method (CAM) belongs to the class of fixed-point iteration schemes (FPIS)
Supplemental Material: A. Proof of Convergence In this Appendix, we provide a computational proof that the circadian adjustment method (CAM) belongs to the class of fixed-point iteration schemes (FPIS)
Empirical Formula for Creating Error Bars for the Method of Paired Comparison
 Empirical Formula for Creating Error Bars for the Method of Paired Comparison Ethan D. Montag Rochester Institute of Technology Munsell Color Science Laboratory Chester F. Carlson Center for Imaging Science
Empirical Formula for Creating Error Bars for the Method of Paired Comparison Ethan D. Montag Rochester Institute of Technology Munsell Color Science Laboratory Chester F. Carlson Center for Imaging Science
Introduction to Computational Neuroscience
 Introduction to Computational Neuroscience Lecture 5: Data analysis II Lesson Title 1 Introduction 2 Structure and Function of the NS 3 Windows to the Brain 4 Data analysis 5 Data analysis II 6 Single
Introduction to Computational Neuroscience Lecture 5: Data analysis II Lesson Title 1 Introduction 2 Structure and Function of the NS 3 Windows to the Brain 4 Data analysis 5 Data analysis II 6 Single
An Ideal Observer Model of Visual Short-Term Memory Predicts Human Capacity Precision Tradeoffs
 An Ideal Observer Model of Visual Short-Term Memory Predicts Human Capacity Precision Tradeoffs Chris R. Sims Robert A. Jacobs David C. Knill (csims@cvs.rochester.edu) (robbie@bcs.rochester.edu) (knill@cvs.rochester.edu)
An Ideal Observer Model of Visual Short-Term Memory Predicts Human Capacity Precision Tradeoffs Chris R. Sims Robert A. Jacobs David C. Knill (csims@cvs.rochester.edu) (robbie@bcs.rochester.edu) (knill@cvs.rochester.edu)
Supplementary data for:
 Supplementary data for: Prediction of treatment efficacy for prostate cancer using a mathematical model Huiming Peng 1, Weiling Zhao 1, Hua Tan 1, Zhiwei Ji 1, Jingsong Li 2, King Li 1, Xiaobo Zhou 1,*
Supplementary data for: Prediction of treatment efficacy for prostate cancer using a mathematical model Huiming Peng 1, Weiling Zhao 1, Hua Tan 1, Zhiwei Ji 1, Jingsong Li 2, King Li 1, Xiaobo Zhou 1,*
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Lack of Association between Endoplasmic Reticulum Stress Response Genes and Suicidal Victims
 Kobe J. Med. Sci., Vol. 53, No. 4, pp. 151-155, 2007 Lack of Association between Endoplasmic Reticulum Stress Response Genes and Suicidal Victims KAORU SAKURAI 1, NAOKI NISHIGUCHI 2, OSAMU SHIRAKAWA 2,
Kobe J. Med. Sci., Vol. 53, No. 4, pp. 151-155, 2007 Lack of Association between Endoplasmic Reticulum Stress Response Genes and Suicidal Victims KAORU SAKURAI 1, NAOKI NISHIGUCHI 2, OSAMU SHIRAKAWA 2,
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
