Enrichment and Separation of Mono- and Multiply Phosphorylated Peptides Using Sequential Elution from IMAC Prior to Mass Spectrometric Analysis
|
|
|
- Merilyn Poole
- 5 years ago
- Views:
Transcription
1 Chapter 6 Enrichment and Separation of Mono- and Multiply Phosphorylated Peptides Using Sequential Elution from IMAC Prior to Mass Spectrometric Analysis Tine E. Thingholm, Ole N. Jensen, and Martin R. Larsen Summary Phospho-proteomics relies on methods for efficient purification and sequencing of phosphopeptides from highly complex biological systems using low amounts of starting material. Current methods for phosphopeptide enrichment, e.g., immobilized metal affinity chromatography and titanium dioxide chromatography, provide varying degrees of selectivity and specificity for phosphopeptide enrichment. Furthermore, the number of multiply phosphorylated peptides that are identified in most published studies is rather low. Here the protocol for a new strategy that separates mono-phosphorylated peptides from multiply phosphorylated peptides using sequential elution from immobilized metal affinity chromatography is described. The two separate phosphopeptide fractions are subsequently analyzed by mass spectrometric methods optimized for mono-phosphorylated and multiply phosphorylated peptides, respectively, resulting in improved identification of especially multiply phosphorylated peptides from a minimum amount of starting material. The new method increases the coverage of the phosphoproteome significantly. Key words: Phosphopeptide enrichment, Multiply phosphorylated peptides, Immobilized metal affinity chromatography (IMAC), Sequential elution, SIMAC, Titanium dioxide (TiO 2 ) chromatography, Mass spectrometry. 1. Introduction Several techniques exist for phosphopeptide enrichment prior to mass spectrometric analysis. Today the most commonly used methods are immobilized metal affinity chromatography (IMAC) and titanium dioxide (TiO 2 ) chromatography (see Chapters Marjo de Graauw (ed.), Phospho-Proteomics, Methods and Protocols, vol Humana Press, a part of Springer Science + Business Media, New York, NY Book DOI: / _6 67
2 68 Thingholm, Jensen, and Larsen Enrichment and Characterization of Phosphopeptides by Immobilized Metal Affinity Chromatography (IMAC) and Mass Spectrometry and The Use of Titanium Dioxide Micro-Columns to Selectively Isolate Phosphopeptides from Proteolytic Digests ). Recent studies comparing three different phosphopeptide enrichment methods, including phosphoramidate chemistry (PAC) (1), IMAC, and TiO 2 chromatography, showed that each method isolated distinct, partially overlapping segments of a phosphoproteome, whereas none of the tested methods were able to provide a whole phosphoproteome (2). One of the challenges in large-scale phospho-proteomics is the analysis of multiply phosphorylated peptides. Multiply phosphorylated peptides are in general suppressed in the ionization process in the mass spectrometric (MS) analysis in the presence of mono- or non-phosphorylated peptides and thereby the chance to detect and identify them decreases. In addition, most mass spectrometers are only able to perform a limited number of tandem MS experiments (MS/MS) in a given time period resulting in the negligence of the less abundant multiply phosphorylated peptides. Furthermore, collision-induced dissociation (CID) fragmentation of phosphopeptides usually results in the loss of phosphoric acid with poor peptide backbone fragmentation and consequently little sequence information is obtained. Phosphorylation-directed multistage tandem MS (pdms 3 ), where an ion originating from a neutral loss signal detected in the MS 2 is selected for a second round of CID (MS 3 ), provides better sequence information for mono-phosphorylated peptide ions with relatively high abundance. However, multiply phosphorylated peptides will result in additional losses of phosphoric acid. Optimized pdms 3 (3), multistage activation (MSA) (4), or electron capture/transfer dissociation (ECD/ETD) (5, 6) will most likely give better results for multiply phosphorylated peptides. Though, multiply phosphorylated peptides need to be separated from mono-phosphorylated peptides in order to analyze them using different mass spectrometric setups. Recently, we developed a new method for the separation of mono-phosphorylated peptides from multiply phosphorylated peptides where we are using sequential elution from IMAC (SIMAC) (3). In this strategy the peptide mixture is incubated with IMAC beads, which have a stronger selectivity for multiply phosphorylated peptides than for mono-phosphorylated peptides (7). After incubation, the sample is split in three elution fractions (see Fig. 1), an IMAC flow-through fraction, an acidic (1% TFA) fraction, and a basic (ph 11.3) fraction. The IMAC flow-through and acidic fractions that contain mainly mono-phosphorylated peptides in addition to a significant number of nonphosphorylated peptides are further enriched by
3 Enrichment and Separation of Mono- and Multiply Phosphorylated Peptides 69 Fig. 1. The setup used for the enrichment and separation of mono- from multiply phosphorylate peptides using the SIMAC strategy. The peptide sample is mixed with the IMAC beads and incubated for 30 min in a Thermomixer at room temperature. After incubation the beads are packed into a GELoader tip forming an IMAC micro-column. The IMAC flow-through is collected and further enriched using TiO 2 chromatography. The monophosphorylated peptides are eluted from the IMAC micro-column using acidic elution conditions (1% TFA, ph 1.0) and for complex samples this eluate is also further enriched using TiO 2 chromatography. The multiply phosphorylated peptides are subsequently eluted from the IMAC micro-column using basic elution conditions (ammonia water, ph 11.3). All three fractions are then ready for mass spectrometric analysis. TiO 2 chromatography to achieve pure mono-phosphorylated peptide fractions prior to tandem MS analysis. The basic fraction is analyzed directly by tandem MS analysis without further TiO 2 purification, as multiply phosphorylated peptides binds very hard to TiO 2 material and may be lost at this step (8). Furthermore, this fraction in general has a relatively low level of nonphosphorylated peptides, and thus no further enrichment is necessary. SIMAC greatly improves the number of phosphorylation sites identified even from very low amounts of starting material and offers a way to identify and characterize multiply phosphorylated peptides at large-scale levels (3).
4 70 Thingholm, Jensen, and Larsen 2. Materials 2.1. Model Proteins 2.2. Reduction, Alkylation, and Digestion of Model Proteins 2.3. Cell Culture and Lysis of Human Mesenchymal Stem Cells (hmscs) 2.4. Extraction, Reduction, Alkylation, and Digestion of hmsc Proteins 1. Transferrin (human). 2. Serum albumin (bovine). 3. β-lactoglobulin (bovine). 4. Carbonic anhydrase (bovine). 5. β-casein (bovine). 6. α-casein (bovine). 7. Ovalbumin (chicken). 8. Ribonuclease B (bovine pancreas). 9. Alcohol dehydrogenase (Baker s yeast). 10. Myoglobin (whale skeletal muscle). 11. Lysozyme (chicken). 12. α-amylase (Bacillus species). All proteins were from Sigma (St. Louis. MO). 1. Ammonium bicarbonate. 2. Dithiotreitol (DTT). 3. Iodoacetamide. 4. Modified trypsin. (see Note 1) 1. MEM (EARLES) media w/o phenol red, with glutamax-i (GibcoTM) containing 1% penicillin/streptomycin (GibcoTM) and 10% fetal bovine serum (FBS) (GibcoTM). 2. Phosphate saline buffer (PBS). 3. Phosphatase inhibitor cocktail 1 and 2 (P2850 and P5726, Sigma ). 4. Cell dissociation buffer (CDB) (GibcoTM). 1. Lysis buffer: 7 M urea, 2 M thiourea, 1% N-octyl glycoside, 40 mm Tris-base, 300 U benzonase. 2. Dithiotreitol (DTT). 3. Iodoacetamide. 4. Acetone. 5. Endoproteinase Lys-C (lysyl endopeptidase, WAKO). 6. Ammonium bicarbonate. 7. Modified trypsin (see Note 1).
5 Enrichment and Separation of Mono- and Multiply Phosphorylated Peptides Immobilized Metal Affinity Chromatography (IMAC) 2.6. Titanium Dioxide (TiO 2 ) Chromatography 2.7. Reversed Phase (RP) Chromatography 1. Ironcoated PHOS-select metal chelate beads (Sigma ). 2. IMAC Loading buffer: 0.1% trifluoroacetic acid (TFA), 50% acetonitrile, HPLC Grade. 3. GELoader tips from Alpha Laboratories (Hampshire, S050 4NU, UK). 4. IMAC Elution buffer 1: 1.0% TFA, 20% acetonitrile. 5. IMAC Elution buffer 2: 20 μl ammonia solution (25%), 980 μl UHQ water (ph 11.3). 6. Formic acid. 7. Water was obtained from a Milli-Q system (Millipore, Bedford, MA) (UHQ water). (see Notes 1 and 2). 1. Titanium dioxide (TiO 2 ) beads (Titansphere, 5 μm, GL sciences Inc., 2. GELoader tips (Eppendorf, Hamburg, Germany) or p10 pipette tips depending on the size of the column needed (see Note 3). 3. 3M Empore C8 disk (3M, Bioanalytical Technologies, St. Paul, MN). 4. Acetonitrile. 5. Syringe for HPLC loading (P/N , N25/500-LC PKT 5, SGE, Ringwood, Victoria, Australia) to create a small plug of C 8 membrane material. 6. TiO 2 loading buffer: 1 M glycolic acid in 5% TFA, 80% acetonitrile. 7. TiO 2 washing buffer: 1% TFA, 80% acetonitrile. 8. TiO 2 elution buffer 1: 20 μl ammonia solution (25%), 980 μl UHQ water. 9. TiO 2 elution buffer 2: 30% acetonitrile. 10. Formic acid. 1. POROS Oligo R3 reversed phase material (Applied Biosystems, Foster City, CA). 2. GELoader tips (Eppendorf, Hamburg, Germany) or p10 pipette tips depending on the size of the column needed (see Note 4). 3. 3M Empore C18 disk (3M, Bioanalytical Technologies, St. Paul, MN). 4. Syringe for HPLC loading (P/N , N25/500-LC PKT 5, SGE, Ringwood, Victoria, Australia) to create a small plug of C 18 membrane material.
6 72 Thingholm, Jensen, and Larsen 5. RP Loading buffer: 0.1% TFA. 6. RP Elution buffer (for LC-ESI MSMS analysis): 70% acetonitrile, 0.1% TFA. 7. DHB Elution buffer (for MALDI MS analysis): 20 mg/ml DHB in 50% acetonitrile, 1% ortho-phosphoric acid (Bie & Berntsen A/S, UK). 3. Methods The principle of the SIMAC method is illustrated in this chapter using a peptide mixture originating from tryptic digestions of 12 standard proteins (Model proteins) (see Notes 5 and 6). The protocol is then applied to enrich for phosphorylated peptides from whole cell lysates from 120 μg protein from human mesenchymal stem cells (hmscs). The SIMAC purification method is a simple and very straightforward method that can be adapted to already existing methods. It is fast and efficient for enrichment of phosphopeptides even from highly complex samples (3, 8). The experimental setup of the method is illustrated in Fig. 1. For illustrating the anticipated results, a peptide mixture originating from tryptic digestions of 12 standard proteins was prepared Model Proteins and Peptide Mixture 3.2. Human Mesenchymal Stem Cell (hmsc) Proteins See details in Subheading 3.1, Chapter Enrichment and Characterization of Phosphopeptides by Immobilized Metal Affinity Chromatography (IMAC) and Mass Spectrometry. The large-scale experiments shown in this article were performed using a peptide mixture originating from tryptic digestion of 120-μg total protein from cell extract from hmscs. 1. Grow human mesenchymal stem cells (hmsc-tert20) in T75 flasks in MEM media without phenol red, with glutamax- I containing penicillin/streptomycin and fetal bovine serum at 37 C until they reached 90% confluence. Wash the confluent cells once with PBS buffer (37 C) and add 5-mL media (37 C) to cover the cells. Add phosphatase inhibitor cocktails (50 μl of each) to the media. Incubate the cells with the phosphatase inhibitors for 30 min at 37 C. Wash the cells with ice-cold PBS buffer harvest them using cell dissociation buffer. 2. After harvesting the cells, resuspend the cell pellet in 1.5-mL lysis buffer. Sonicate the cells at 40% output with intervals three times 15 s on ice to disrupt the cells and incubate at 80 C for 30 min. Add dithiotreitol (DTT, 20 mm) and incubate the sample at room temperature for 35 min. After incubation, add
7 Enrichment and Separation of Mono- and Multiply Phosphorylated Peptides mm iodoacetamide and incubate for 35 min at room temperature in the dark. 3. Add 14-mL ice-cold acetone to the solution and incubate at 20 C for 20 min to precipitate the proteins. Pellet the proteins by centrifugation at 6,000 g for 10 min at 4 C. Lyophilize the pellet and store it at 20 C until further use. 4. Redissolve a total of 1 mg of the precipitated and lyophilized protein from hmsc (weight out) in 6 M urea, 2 M thiourea to a concentration of 50 μg protein/μl. Incubate the proteins with endoproteinase Lys-C (1 2% w/w) at room temperature for 3 h. After incubation, dilute the endoproteinase Lys-C digested sample five times with 50 mm NH 4 HCO 3 and add 1 2% (w/w) chemically modified trypsin. Incubate the sample at room temperature for 18 h Batch Incubation with IMAC Beads 3.4. Packing the IMAC Micro-Column 1. Always adjust the amount of IMAC beads to the amount of sample in order to reduce the level of nonspecific binding from nonphosphorylated peptides. For 1 pmol tryptic digest use 7-μL IMAC beads (see Chapter Enrichment and Characterization of Phosphopeptides by Immobilized Metal Affinity Chromatography (IMAC) and Mass Spectrometry ). For more complex samples, where more material is available, use more IMAC beads. The following is describing the protocol when using 120-μg tryptic digest. 2. Transfer 40 μl of IMAC beads to a fresh 1-mL Eppedorf tube. 3. Wash the IMAC beads twice using 200 μl of IMAC loading buffer. 4. Resuspend the beads in 150 μl of IMAC loading buffer and add the sample (here 12 μl). 5. Incubate the sample with IMAC beads in a Thermomixer (Eppendorf) for 30 min at 20 C. 1. Squeeze the end of a 200-μL GELoader tip to prevent the IMAC beads from leaking. 2. After incubation pack the beads in the constricted end of the GELoader tip by application of air pressure forming an IMAC micro-column essentially as described in Chapter Enrichment and Characterization of Phosphopeptides by Immobilized Metal Affinity Chromatography (IMAC) and Mass Spectrometry (9). 3. It is critical to collect the IMAC flow-through in a fresh 1.5- ml Eppendorf tube for further analysis by TiO 2 chromatography. 4. Wash the IMAC column using 50-μL IMAC loading buffer and pool this with the IMAC flow-through.
8 74 Thingholm, Jensen, and Larsen 5. Lyophilize the IMAC flow-through and wash and enrich for mono-phosphorylated peptides using TiO 2 chromatography (see Subheading ) Elution of Monophosphorylated Peptides from the IMAC Micro-Column 1. Elute the mono-phosphorylated peptides bound to the IMAC micro-column using 50 μl of IMAC elution buffer 1 (Fig. 1). 2. Dry the eluate by lyophilisation. 3. Redissolve the peptides in 5-μL 1% SDS and purify further by TiO 2 chromatography (see Subheadings ) Elution of Multiply phosphorylated Peptides from the IMAC Micro-Column 1. Elute the multiply phosphorylated peptides bound to the IMAC micro-column using 70 μl of IMAC elution buffer 2 (Fig. 1). 2. For LC-ESI MS n analysis this IMAC eluate should be acidified using 100% formic acid (ph should be 2 3) and desalted/ concentrated using reversed phase chromatography Packing the TiO 2 Micro-Column 1. Prepare six TiO 2 micro-columns of 4 mm by packing the TiO 2 beads on top of the C 8 membrane disks in P10 tips as described in Subheading 3.2, Chapter The Use of Titanium Dioxide Micro-Columns to Selectively Isolate Phosphopeptides from Proteolytic Digests Loading the Sample onto the TiO 2 Micro-Column 1. Divide the IMAC flow-through + wash between three different TiO 2 micro-columns. Do the same for the IMAC eluate. See details for loading onto the TiO 2 micro-columns in Subheading 3.3, Chapter SILAC for Global Phosphoproteomic Analysis Elution of Phosphorylated Peptides from the TiO 2 Micro-Column 1. Elute the phosphopeptides bound to each TiO 2 micro-columns using at least 35 μl of TiO 2 elution buffer 1 followed by an elution step using 1 μl of TiO 2 elution buffer 2 and pool the eluates as described in Subheading 3.4, Chapter The Use of Titanium Dioxide Micro-Columns to Selectively Isolate Phosphopeptides from Proteolytic Digests (see Notes 7 and 8) Packing a Reversed Phase (RP) Micro-Column for Desalting/Concentrating the Sample 1. Prepare a RP micro-column of ~5 6 mm by packing the R3 beads on top of the C 18 membrane disk in P10 tips as described for TiO 2 micro-columns in Subheading 3.2, Chapter The Use of Titanium Dioxide Micro-Columns to Selectively Isolate Phosphopeptides from Proteolytic Digests..
9 Enrichment and Separation of Mono- and Multiply Phosphorylated Peptides Loading the Enriched Phosphopeptides onto the RP Micro-Column Elution of Phosphorylated Peptides from the RP Micro- Column 1. Load the acidified sample slowly onto the RP micro-column (~1 drop/s). 2. Wash the RP micro-column using 30 µl of 0.1% TFA. 1. Elute the phosphopeptides from the RP micro-column using 70 μl RP elution buffer, followed by lyophilization of the phosphopeptides. (N.B. For simple samples, the peptides can be eluted off the RP micro-column directly onto the MALDI target using 1 μl DHB solution). 2. LC-ESI-MS n analysis: Redissolve the dried phosphopeptides in 0.5 μl 100% formic acid and immediately dilute to 10 μl with UHQ water. LC-ESI-MS n analysis of multiply phosphorylated peptides is improved by redissolving the phosphopeptides by sonication in an EDTA containing buffer prior to LC-ESI-MS n analysis (10). Fig. 2. Results obtained from 1 pmol peptide mixture using the SIMAC strategy. MALDI MS spectra of peptides identified from the IMAC flow-through after further enrichment using TiO 2 chromatography (a), MALDI MS spectra of peptides eluted from the IMAC micro-column using 1% TFA (b) and MALDI MS spectra of peptides eluted from the IMAC microcolumn using ammonia water (ph 11.30) (c). The number of phosphate groups on the individual phosphopeptides are indicated by #P. Asterisk indicates the loss of phosphoric acid. (see Note 9).
10 76 Thingholm, Jensen, and Larsen Figure 2 illustrates the results obtained by using the SIMAC strategy on 1 pmol standard peptide mixture. MALDI MS spectra of the IMAC flow-through after TiO 2 enrichment (a) and MALDI MS spectra of the phosphopeptides eluted from the IMAC material using acidic conditions (1% TFA, ph 1.0) (b) show that mainly mono-phosphorylated peptides are identified in these two peptide fractions. Mainly multiply phosphorylated peptides are identified when using basic elution conditions (ammonia water, ph 11.3) (c) (see Note 9). Figure 3 compares the number of mono-phosphorylated and multiply phosphorylated as well as nonphosphorylated peptides identified from 120-µg hmsc tryptic digests using either TiO 2 enrichment or the SIMAC strategy. The results show that mainly mono-phosphorylated peptides are identified by TiO 2 chromatography, whereas only 17% (54 of 325) of the identified peptides are multiply phosphorylated. In the SIMAC experiment, the IMAC flow-through as well as the acidic elution fraction consist mainly of mono-phosphorylated peptides and the level of multiply phosphorylated peptides identified is even lower than for TiO 2 chromatography (3% and 4%, respectively). However, the level of multiply phosphorylated peptides identified is significantly increased by basic elution (68%). In total SIMAC gives rise to more than three times as many multiply phosphorylated peptides identified from 120-µg hmsc tryptic digest when compared with TiO 2 chromatography. Fig. 3. Results obtained from the analysis of phosphorylated peptides from acetone precipitated proteins from human mesenchymal stem cells (hmscs) using TiO 2 chromatography or SIMAC. The number of mono-phosphorylated peptides, multiply phosphorylated peptides and nonphosphorylated peptides identified from 120-μg hmsc tryptic digests using TiO 2 enrichment and the number of mono-phosphorylated peptides, multiply phosphorylated peptides and nonphosphorylated peptides identified in the three peptide fractions obtained from SIMAC analysis of 120-μg hmsc tryptic digests.
11 Enrichment and Separation of Mono- and Multiply Phosphorylated Peptides Notes 1. It is important to obtain the highest purity of all chemicals used. 2. All solutions should be prepared in UHQ water. 3. The capacity of the TiO 2 beads have been tested; however, more experiments need to be performed. The capacity of the beads appears to be very high, but when working with high amounts of material it is recommended to use P10 pipette tips instead of GELoader tips and to split the peptide sample and load it onto several TiO 2 micro-columns. The resulting eluates can then be pooled prior to MS analysis. For the experiments using 120-μg hmsc protein, six TiO 2 micro-columns of ~4 mm were packed in P10 tips. 4. For the experiments using 120-μg hmsc protein, a R3 microcolumn of ~5 6 mm were packed in P10 tips. 5. Always start by testing the method using a model peptide mixture. It is important to freshly prepare the peptide mixture as peptides bind to the surface of the plastic tubes in which they are stored. In addition, avoid transferring the peptide sample between different tubes to minimize adsorptive losses of the sample. 6. The peptide mixture used for the experiment illustrated in this chapter contained peptides originating from tryptic digestions of 1 pmol of each of the 12 proteins. 7. Use at least 35 μl of TiO 2 elution buffer 1 due to the size of the TiO 2 micro-columns. It is important to follow the first elution step using 1 μl of TiO 2 elution buffer 2, to elute peptides bound to the C 8 membrane plug. 8. At this step, pool the eluates from the three TiO 2 micro-columns with IMAC flow-through + wash by eluting the phosphopeptides into the same Eppendorf tube. Do the same for the three TiO 2 micro-columns with IMAC eluates. 9. The results obtained using this protocol will differ according to the mass spectrometer used for the analysis of the phosphopeptides, not only between MALDI MS and ESI MS but also within different MALDI MS instruments, depending on laser optics, laser frequency, instrumental configuration, sensitivity etc. Acknowledgements This work was supported by the Danish Natural Science Research Council (grant no (M.R.L) ) and the Danish Strategic Research Council (O.N.J).
12 78 Thingholm, Jensen, and Larsen References 1. Zhou, H. L., Watts, J. D., and Aebersold, R. A. (2001) A systematic approach to the analysis of protein phosphorylation. Nat. Biotechnol. 19, Bodenmiller, B., Mueller, L. N., Mueller, M., Domon, B., and Aebersold, R. (2007) Reproducible isolation of distinct, overlapping segments of the phosphoproteome. Nat. Methods 4, Thingholm, T. E., Jensen, O. N., Robinson, P. J., and Larsen, M. R. (2007) SIMAC A phosphoproteomic strategy for the rapid separation of mono-phosphorylated from multiply phosphorylated peptides. Published online by MCP on November 27th 2007 (M MCP200). 4. Schroeder, M. J., Shabanowitz, J., Schwartz, J. C., Hunt, D. F., and Coon, J. J. (2004) A neutral loss activation method for improved phosphopeptide sequence analysis by quadrupole ion trap mass spectrometry. Analytical Chemistry 76, Chalmers, M. J., Hakansson, K., Johnson, R., Smith, R., Shen, J., Emmett, M. R., and Marshall, A. G. (2004) Protein kinase A phosphorylation characterized by tandem Fourier transform ion cyclotron resonance mass spectrometry. Proteomics 4, Schroeder, M. J., Webb, D. J., Shabanowitz, J., Horwitz, A. F., and Hunt, D. F. (2005) Methods for the detection of paxillin posttranslational modifications and interacting proteins by mass spectrometry. J Proteome Res 4, Ficarro, S. B., McCleland, M. L., Stukenberg, P. T., Burke, D. J., Ross, M. M., Shabanowitz, J., Hunt, D. F., and White, F. M. (2002) Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nat Biotechnol 20, Jensen, S. S., and Larsen, M. R. (2007) Evaluation of the impact of some experimental procedures on different phosphopeptide enrichment techniques. Rapid Commun. Mass Spectrom. 21, Gobom, J., Nordhoff, E., Mirgorodskaya, E., Ekman, R., and Roepstorff, P. (1999) Sample purification and preparation technique based on nano-scale reversed-phase columns for the sensitive analysis of complex peptide mixtures by matrix-assisted laser desorption/ionization mass spectrometry. J. Mass Spectrom. 34, Liu, S., Zhang, C., Campbell, J. L., Zhang, H., Yeung, K. K., Han, V. K., and Lajoie, G. A. (2005) Formation of phosphopeptidemetal ion complexes in liquid chromatography/electrospray mass spectrometry and their influence on phosphopeptide detection. Rapid Commun Mass Spectrom. 19,
Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins
 Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins Loreta Bllaci, Silje B. Torsetnes,, Celina Wierzbicka, Sudhirkumar Shinde, Börje Sellergren,
Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins Loreta Bllaci, Silje B. Torsetnes,, Celina Wierzbicka, Sudhirkumar Shinde, Börje Sellergren,
PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX
 PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX E. Claude 1, E. Emekwue 2, M. Snel 1, T. McKenna 1 and J. Langridge 1 1: Waters Corporation, Manchester, UK 2:
PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX E. Claude 1, E. Emekwue 2, M. Snel 1, T. McKenna 1 and J. Langridge 1 1: Waters Corporation, Manchester, UK 2:
In-Solution Digestion for proteomics
 In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Supporting information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Analytical strategies for phosphoproteomics
 Proteomics 2009, 9, 1451 1468 DOI 10.1002/pmic.200800454 1451 REVIEW Analytical strategies for phosphoproteomics Tine E. Thingholm 1, 2, Ole N. Jensen 2 and Martin R. Larsen 2 1 KMEB, Department of Endocrinology,
Proteomics 2009, 9, 1451 1468 DOI 10.1002/pmic.200800454 1451 REVIEW Analytical strategies for phosphoproteomics Tine E. Thingholm 1, 2, Ole N. Jensen 2 and Martin R. Larsen 2 1 KMEB, Department of Endocrinology,
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Electronic Supplementary Information
 Electronic Supplementary Information Microwave-assisted Kochetkov amination followed by permanent charge derivatization: A facile strategy for glycomics Xin Liu a,b, Guisen Zhang* a, Kenneth Chan b and
Electronic Supplementary Information Microwave-assisted Kochetkov amination followed by permanent charge derivatization: A facile strategy for glycomics Xin Liu a,b, Guisen Zhang* a, Kenneth Chan b and
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2011
 Supporting Information Experimental details 1. Materials Ferric chloride (FeCl 3 6H 2 O), Ethylene glycol (EG) and Ammonium hydroxide (NH 3 H 2 O) were obtained from Beijing Chemical Regent Co. Ltd. (Beijing,
Supporting Information Experimental details 1. Materials Ferric chloride (FeCl 3 6H 2 O), Ethylene glycol (EG) and Ammonium hydroxide (NH 3 H 2 O) were obtained from Beijing Chemical Regent Co. Ltd. (Beijing,
Supplementary Figure 1. Method development.
 Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
Application Note. Authors. Abstract. Automated Peptide Sample Preparation for LC/MS
 Agilent AssayMAP Bravo Technology Enables Reproducible Automated Phosphopeptide Enrichment from Complex Mixtures Using High Capacity Fe(III) NTA Cartridges Application Note Automated Peptide Sample Preparation
Agilent AssayMAP Bravo Technology Enables Reproducible Automated Phosphopeptide Enrichment from Complex Mixtures Using High Capacity Fe(III) NTA Cartridges Application Note Automated Peptide Sample Preparation
TiO 2 Mag Sepharose. SepharoseTM. GE Healthcare. Simple handling
 GE Healthcare Data file 28-9539-44 AA Protein sample preparation TiO 2 Mag Sepharose TiO 2 Mag Sepharose magnetic beads use titanium dioxide (TiO 2 )-based chromatography to simplify capture and enrichment
GE Healthcare Data file 28-9539-44 AA Protein sample preparation TiO 2 Mag Sepharose TiO 2 Mag Sepharose magnetic beads use titanium dioxide (TiO 2 )-based chromatography to simplify capture and enrichment
Improved Detection of Hydrophilic Phosphopeptides Using Graphite Powder Microcolumns and Mass Spectrometry
 Research Improved Detection of Hydrophilic Phosphopeptides Using Graphite Powder Microcolumns and Mass Spectrometry EVIDENCE FOR IN VIVO DOUBLY PHOSPHORYLATED DYNAMIN I AND DYNAMIN III* Martin R. Larsen,
Research Improved Detection of Hydrophilic Phosphopeptides Using Graphite Powder Microcolumns and Mass Spectrometry EVIDENCE FOR IN VIVO DOUBLY PHOSPHORYLATED DYNAMIN I AND DYNAMIN III* Martin R. Larsen,
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g.
 Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
This document is the unedited Author s version of a Submitted Work that was subsequently accepted
 This document is the unedited Author s version of a Submitted Work that was subsequently accepted for publication in Analytical Chemistry, copyright American Chemical Society after peer review. To access
This document is the unedited Author s version of a Submitted Work that was subsequently accepted for publication in Analytical Chemistry, copyright American Chemical Society after peer review. To access
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Protein sequence mapping is commonly used to
 Reproducible Microwave-Assisted Acid Hydrolysis of Proteins Using a Household Microwave Oven and Its Combination with LC-ESI MS/MS for Mapping Protein Sequences and Modifications Nan Wang and Liang Li
Reproducible Microwave-Assisted Acid Hydrolysis of Proteins Using a Household Microwave Oven and Its Combination with LC-ESI MS/MS for Mapping Protein Sequences and Modifications Nan Wang and Liang Li
(III) MALDI instrumentation
 Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
MALDI-TOF. Introduction. Schematic and Theory of MALDI
 MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
VaTx1 VaTx2 VaTx3. VaTx min Retention Time (min) Retention Time (min)
 a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
Supporting Information
 Supporting Information Experimental Materials Ammonium dihydrogen phosphate (NH 4 H 2 PO 4 ) and Ammonium hydroxide (NH 3 H 2 O) were obtained from Beijing Chemical Regent Co., Ltd. (Beijing, China). Lanthanum
Supporting Information Experimental Materials Ammonium dihydrogen phosphate (NH 4 H 2 PO 4 ) and Ammonium hydroxide (NH 3 H 2 O) were obtained from Beijing Chemical Regent Co., Ltd. (Beijing, China). Lanthanum
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Protocol for purification of recombinant protein from 300 ml yeast culture
 Protocol for purification of recombinant protein from 300 ml yeast culture Equipment and reagents needed: Zirconia beads (0.5 mm diameter from BSP, Germany) Paint Shaker (at 4 C) Tube rotator for 15 ml
Protocol for purification of recombinant protein from 300 ml yeast culture Equipment and reagents needed: Zirconia beads (0.5 mm diameter from BSP, Germany) Paint Shaker (at 4 C) Tube rotator for 15 ml
Proteomics Grade. Protocol. Catalog # Agilent Technologies. Research Use Only. Not for use in Diagnostic Procedures. Version A, January 2010
 Proteomics Grade Trypsin Catalog #204310 Protocol Version A, January 2010 Research Use Only. Not for use in Diagnostic Procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010 No part of
Proteomics Grade Trypsin Catalog #204310 Protocol Version A, January 2010 Research Use Only. Not for use in Diagnostic Procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010 No part of
ProteaseMAX Surfactant, Trypsin Enhancer
 Technical Bulletin ProteaseMAX Surfactant, Trypsin Enhancer INSTRUCTIONS FOR USE OF PRODUCTS V2071 AND V2072. PRINTED IN USA. Revised 1/10 ProteaseMAX Surfactant, Trypsin Enhancer All technical literature
Technical Bulletin ProteaseMAX Surfactant, Trypsin Enhancer INSTRUCTIONS FOR USE OF PRODUCTS V2071 AND V2072. PRINTED IN USA. Revised 1/10 ProteaseMAX Surfactant, Trypsin Enhancer All technical literature
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Dienes Derivatization MaxSpec Kit
 Dienes Derivatization MaxSpec Kit Item No. 601510 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
Dienes Derivatization MaxSpec Kit Item No. 601510 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
Quantitative Analysis of Protein Phosphorylation in Mouse Brain by Hypothesis-Driven Multistage Mass Spectrometry
 Anal. Chem. 2005, 77, 7845-7851 Accelerated Articles Quantitative Analysis of Protein Phosphorylation in Mouse Brain by Hypothesis-Driven Multistage Mass Spectrometry Mi Jin, Helen Bateup, Júlio C. Padovan,
Anal. Chem. 2005, 77, 7845-7851 Accelerated Articles Quantitative Analysis of Protein Phosphorylation in Mouse Brain by Hypothesis-Driven Multistage Mass Spectrometry Mi Jin, Helen Bateup, Júlio C. Padovan,
Europium Labeling Kit
 Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
rev. 04/26/16 MXRXXS*/T* 9% RXXS* 45% LXRXXS*/T* 32% RXXT* 14%
 Store at 20 C #5564 PTMScan Phospho-AMPK Substrate Motif (LXRXXS*/T*) Kit 3 n 1 Kit (10 assays) rev. 04/26/16 Orders n 877-616-CELL (2355) orders@cellsignal.com Support n 877-678-TECH (8324) info@cellsignal.com
Store at 20 C #5564 PTMScan Phospho-AMPK Substrate Motif (LXRXXS*/T*) Kit 3 n 1 Kit (10 assays) rev. 04/26/16 Orders n 877-616-CELL (2355) orders@cellsignal.com Support n 877-678-TECH (8324) info@cellsignal.com
Ion Source. Mass Analyzer. Detector. intensity. mass/charge
 Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
DetergentOUT Tween. DetergentOUT GBS10. OrgoSol DetergentOUT
 252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2008
 Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for
 Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Trypsin Digestion Mix
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
Babu Antharavally, Ryan Bomgarden, and John Rogers Thermo Fisher Scientific, Rockford, IL
 A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
In-Gel Tryptic Digestion Kit
 INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
Bench Protocol to Perform TAILS v4
 Bench Protocol to Perform TAILS v4 OVERALL Lab Protocols May 2016 We have been performing TAILS for 12 years and this is now a very streamlined and highly optimized approach. The one request we have is
Bench Protocol to Perform TAILS v4 OVERALL Lab Protocols May 2016 We have been performing TAILS for 12 years and this is now a very streamlined and highly optimized approach. The one request we have is
Nat Rev Mol Cell Biol (2006) 7, Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312,
 Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Determination of β2-agonists in Pork Using Agilent SampliQ SCX Solid-Phase Extraction Cartridges and Liquid Chromatography-Tandem Mass Spectrometry
 Determination of β2-agonists in Pork Using Agilent SampliQ SCX Solid-Phase Extraction Cartridges and Liquid Chromatography-Tandem Mass Spectrometry Application Note Food Safety Authors Chenhao Zhai Agilent
Determination of β2-agonists in Pork Using Agilent SampliQ SCX Solid-Phase Extraction Cartridges and Liquid Chromatography-Tandem Mass Spectrometry Application Note Food Safety Authors Chenhao Zhai Agilent
REDOX PROTEOMICS. Roman Zubarev.
 REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
Highly efficient Phosphopeptide Enrichment by Calcium Phosphate. Precipitation Combined with Subsequent Serial Immobilized Metal Ion
 MCP Papers in Press. Published on August 4, 2007 as Manuscript M700278-MCP200 Highly efficient Phosphopeptide Enrichment by Calcium Phosphate Precipitation Combined with Subsequent Serial Immobilized Metal
MCP Papers in Press. Published on August 4, 2007 as Manuscript M700278-MCP200 Highly efficient Phosphopeptide Enrichment by Calcium Phosphate Precipitation Combined with Subsequent Serial Immobilized Metal
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
Neosolaniol. [Methods listed in the Feed Analysis Standards]
![Neosolaniol. [Methods listed in the Feed Analysis Standards] Neosolaniol. [Methods listed in the Feed Analysis Standards]](/thumbs/86/94003482.jpg) Neosolaniol [Methods listed in the Feed Analysis Standards] 1 Simultaneous analysis of mycotoxins by liquid chromatography/ tandem mass spectrometry [Feed Analysis Standards, Chapter 5, Section 1 9.1 ]
Neosolaniol [Methods listed in the Feed Analysis Standards] 1 Simultaneous analysis of mycotoxins by liquid chromatography/ tandem mass spectrometry [Feed Analysis Standards, Chapter 5, Section 1 9.1 ]
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
MALDI Sepsityper Kit
 Instructions for Use MALDI Sepsityper Kit Kit for identification of microorganisms from positive blood cultures using the MALDI Biotyper system CARE products are designed to support our worldwide customers
Instructions for Use MALDI Sepsityper Kit Kit for identification of microorganisms from positive blood cultures using the MALDI Biotyper system CARE products are designed to support our worldwide customers
Advances in Hybrid Mass Spectrometry
 The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
Minute TM Plasma Membrane Protein Isolation and Cell Fractionation Kit User Manual (v5)
 Minute TM Plasma Membrane Protein Isolation and Cell Fractionation Kit Catalog number: SM-005 Description Minute TM plasma membrane (PM) protein isolation kit is a novel and patented native PM protein
Minute TM Plasma Membrane Protein Isolation and Cell Fractionation Kit Catalog number: SM-005 Description Minute TM plasma membrane (PM) protein isolation kit is a novel and patented native PM protein
Proteins: Proteomics & Protein-Protein Interactions Part I
 Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
The Detection of Allergens in Food Products with LC-MS
 The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
AccuMAP Low ph Protein Digestion Kits
 TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
Faster Mass Spec: Same-Day Sample Prep Now a Reality. Michael M. Rosenblatt, Ph.D.
 Faster Mass Spec: Same-Day Sample Prep Now a Reality Michael M. Rosenblatt, Ph.D. Applications of Mass Spec in Biology Biomarker Discovery Protein Interactions Protein Expression Drug Discovery Subcellular
Faster Mass Spec: Same-Day Sample Prep Now a Reality Michael M. Rosenblatt, Ph.D. Applications of Mass Spec in Biology Biomarker Discovery Protein Interactions Protein Expression Drug Discovery Subcellular
Title: Column Chromatography of Green Fluorescent Protein
 Title: Column Chromatography of Green Fluorescent Protein Approvals: Preparer Date_07Oct06 Reviewer: Mary Jane Kurtz Date 09Jul13 Part I Crude Isolation of GFP from Lysed Cells q Page 1 of 6 1. Purpose:
Title: Column Chromatography of Green Fluorescent Protein Approvals: Preparer Date_07Oct06 Reviewer: Mary Jane Kurtz Date 09Jul13 Part I Crude Isolation of GFP from Lysed Cells q Page 1 of 6 1. Purpose:
Solving practical problems. Maria Kuhtinskaja
 Solving practical problems Maria Kuhtinskaja What does a mass spectrometer do? It measures mass better than any other technique. It can give information about chemical structures. What are mass measurements
Solving practical problems Maria Kuhtinskaja What does a mass spectrometer do? It measures mass better than any other technique. It can give information about chemical structures. What are mass measurements
Automated Sample Preparation/Concentration of Biological Samples Prior to Analysis via MALDI-TOF Mass Spectroscopy Application Note 222
 Automated Sample Preparation/Concentration of Biological Samples Prior to Analysis via MALDI-TOF Mass Spectroscopy Application Note 222 Joan Stevens, Ph.D.; Luke Roenneburg; Tim Hegeman; Kevin Fawcett
Automated Sample Preparation/Concentration of Biological Samples Prior to Analysis via MALDI-TOF Mass Spectroscopy Application Note 222 Joan Stevens, Ph.D.; Luke Roenneburg; Tim Hegeman; Kevin Fawcett
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Glycosylation analysis of blood plasma proteins
 Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
High Resolution Glycopeptide Mapping of EPO Using an Agilent AdvanceBio Peptide Mapping Column
 High Resolution Glycopeptide Mapping of EPO Using an Agilent AdvanceBio Peptide Mapping Column Application Note BioPharma Authors James Martosella, Phu Duong, and Alex Zhu Agilent Technologies, Inc. Abstract
High Resolution Glycopeptide Mapping of EPO Using an Agilent AdvanceBio Peptide Mapping Column Application Note BioPharma Authors James Martosella, Phu Duong, and Alex Zhu Agilent Technologies, Inc. Abstract
Western Immunoblotting Preparation of Samples:
 Western Immunoblotting Preparation of Samples: Total Protein Extraction from Culture Cells: Take off the medium Wash culture with 1 x PBS 1 ml hot Cell-lysis Solution into T75 flask Scrap out the cells
Western Immunoblotting Preparation of Samples: Total Protein Extraction from Culture Cells: Take off the medium Wash culture with 1 x PBS 1 ml hot Cell-lysis Solution into T75 flask Scrap out the cells
Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
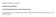 Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
N-Glycosidase F Deglycosylation Kit
 For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
for the Identification of Phosphorylated Peptides
 Application of a Data Dependent Neutral-Loss Experiment on the Finnigan LTQ for the Identification of Phosphorylated Peptides Gargi Choudhary Diane Cho Thermo Electron, San Jose, CA Abstracted from posters
Application of a Data Dependent Neutral-Loss Experiment on the Finnigan LTQ for the Identification of Phosphorylated Peptides Gargi Choudhary Diane Cho Thermo Electron, San Jose, CA Abstracted from posters
Fractionation of phosphopeptides on strong anion-exchange capillary trap column for large-scale phosphoproteome analysis of microgram samples
 J. Sep. Sci. 2010, 33, 1879 1887 1879 Fangjun Wang Guanghui Han Zhiyuan Yu Xinning Jiang Shutao Sun Rui Chen Mingliang Ye Hanfa Zou CAS Key Lab of Separation Sciences for Analytical Chemistry, National
J. Sep. Sci. 2010, 33, 1879 1887 1879 Fangjun Wang Guanghui Han Zhiyuan Yu Xinning Jiang Shutao Sun Rui Chen Mingliang Ye Hanfa Zou CAS Key Lab of Separation Sciences for Analytical Chemistry, National
Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics
 Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
Supporting Information for:
 Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
DetergentOUT Detergent Removal Systems
 252PR-04 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents from Peptide
252PR-04 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents from Peptide
Isomeric Separation of Permethylated Glycans by Porous Graphitic Carbon (PGC)-LC-MS/MS at High- Temperatures
 Supplementary Information Isomeric Separation of Permethylated Glycans by Porous Graphitic Carbon (PGC)-LC-MS/MS at High- Temperatures Shiyue Zhou 1, Yifan Huang 1, Xue Dong 1, Wenjing Peng 1, Lucas Veillon
Supplementary Information Isomeric Separation of Permethylated Glycans by Porous Graphitic Carbon (PGC)-LC-MS/MS at High- Temperatures Shiyue Zhou 1, Yifan Huang 1, Xue Dong 1, Wenjing Peng 1, Lucas Veillon
An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27
 An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27 Top-down HRAM Bioanalysis of Native Proteins/Molecules Relative Abundance 100
An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27 Top-down HRAM Bioanalysis of Native Proteins/Molecules Relative Abundance 100
Chromatin IP (Isw2) Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles.
 Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
LABORATÓRIUMI GYAKORLAT SILLABUSZ SYLLABUS OF A PRACTICAL DEMONSTRATION. financed by the program
 TÁMOP-4.1.1.C-13/1/KONV-2014-0001 projekt Az élettudományi-klinikai felsőoktatás gyakorlatorientált és hallgatóbarát korszerűsítése a vidéki képzőhelyek nemzetközi versenyképességének erősítésére program
TÁMOP-4.1.1.C-13/1/KONV-2014-0001 projekt Az élettudományi-klinikai felsőoktatás gyakorlatorientált és hallgatóbarát korszerűsítése a vidéki képzőhelyek nemzetközi versenyképességének erősítésére program
COMPONENT NAME COMPONENT # QUANTITY STORAGE SHELF LIFE FORMAT. Store at 2-8 C. Do not freeze. Store at 2-8 C. Do not freeze.
 This document is available at www.stemcell.com/pis EasySep Mouse Monocyte Isolation Kit Catalog #19861 For processing 1 x 10^9 cells Description Isolate untouched and highly purified monocytes from mouse
This document is available at www.stemcell.com/pis EasySep Mouse Monocyte Isolation Kit Catalog #19861 For processing 1 x 10^9 cells Description Isolate untouched and highly purified monocytes from mouse
BabyBio IMAC columns DATA SHEET DS
 BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
Mass Spectrometry and Proteomics. Professor Xudong Yao Bioanalytical Chemistry Spring 2007
 Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein
![2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein](/thumbs/86/94743397.jpg) Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
Molecular Cell, Volume 46. Supplemental Information
 Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Mammalian Membrane Protein Extraction Kit
 Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Development of a Cell-penetrating Peptide that Exhibits Responsive. Changes in its Secondary Structure in the Cellular Environment
 Development of a Cell-penetrating Peptide that Exhibits Responsive Changes in its Secondary Structure in the Cellular Environment iroko Yamashita, 1 Takuma Kato, 2 Makoto ba, 2 Takashi Misawa, 1 Takayuki
Development of a Cell-penetrating Peptide that Exhibits Responsive Changes in its Secondary Structure in the Cellular Environment iroko Yamashita, 1 Takuma Kato, 2 Makoto ba, 2 Takashi Misawa, 1 Takayuki
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
1. Sample Introduction to MS Systems:
 MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
Protein Identification and Phosphorylation Site Determination by de novo sequencing using PepFrag TM MALDI-Sequencing kit
 Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Biological Mass spectrometry in Protein Chemistry
 Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
L6 GLUT4myc Cell Growth Protocol
 L6 GLUT4myc Cell Growth Protocol Background: Parental L6 cells selected for high fusion (2, 3) were stably transfected with a rat GLUT4 cdna carrying a myc epitope (recognized by the commercially available
L6 GLUT4myc Cell Growth Protocol Background: Parental L6 cells selected for high fusion (2, 3) were stably transfected with a rat GLUT4 cdna carrying a myc epitope (recognized by the commercially available
Rapid antigen-specific T cell enrichment (Rapid ARTE)
 Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Protein and peptide separations,
 A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
Mitochondrial DNA Isolation Kit
 Mitochondrial DNA Isolation Kit Catalog Number KA0895 50 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
Mitochondrial DNA Isolation Kit Catalog Number KA0895 50 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
Posttranslational modification of proteins Stephen Barnes, PhD
 Posttranslational modification of proteins Stephen Barnes, PhD sbarnes@uab.edu General classes of modification Biochemical event involving peptide processing Biochemical event stimulated by enzymes Chemical
Posttranslational modification of proteins Stephen Barnes, PhD sbarnes@uab.edu General classes of modification Biochemical event involving peptide processing Biochemical event stimulated by enzymes Chemical
Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection. EPL-BAS Method No.
 Page 1 of 10 Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection EPL-BAS Method No. 205G881B Method Summary: Residues of 6-CPA are
Page 1 of 10 Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection EPL-BAS Method No. 205G881B Method Summary: Residues of 6-CPA are
RayBio KinaseSTAR TM Akt Activity Assay Kit
 Activity Assay Kit User Manual Version 1.0 March 13, 2015 RayBio KinaseSTAR TM Akt Activity Kit Protocol (Cat#: 68AT-Akt-S40) RayBiotech, Inc. We Provide You With Excellent Support And Service Tel:(Toll
Activity Assay Kit User Manual Version 1.0 March 13, 2015 RayBio KinaseSTAR TM Akt Activity Kit Protocol (Cat#: 68AT-Akt-S40) RayBiotech, Inc. We Provide You With Excellent Support And Service Tel:(Toll
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
LC-Based Lipidomics Analysis on QTRAP Instruments
 LC-Based Lipidomics Analysis on QTRAP Instruments Junhua Wang and Paul RS Baker SCIEX LC-Based Lipidomics Analysis Topics Covered Lipid extraction techniques Hydrophilic Interaction Chromatography (HILIC)
LC-Based Lipidomics Analysis on QTRAP Instruments Junhua Wang and Paul RS Baker SCIEX LC-Based Lipidomics Analysis Topics Covered Lipid extraction techniques Hydrophilic Interaction Chromatography (HILIC)
