Supplementary Table S1. Histopathologic findings for multi-region analysis.
|
|
|
- Sheila Floyd
- 5 years ago
- Views:
Transcription
1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY TABLES Supplementary Table S1. Histopathologic findings for multi-region analysis. Patient Histology R1: clear cell RCC, Fuhrman 2-3, acinar pattern 1 R2: clear cell RCC, Fuhrman 3-4, large alveolar pattern R3: clear cell RCC, Fuhrman 3-4, large alveolar pattern R1: clear cell RCC, Fuhrman 3, large alveolar pattern R2: clear cell RCC, Fuhrman 3-4, large alveolar pattern 2 R3: clear cell RCC, Fuhrman 3-4, large alveolar pattern M1: clear cell RCC, Fuhrman 3-4, large alveolar pattern R1: clear cell RCC, Fuhrman 2-3, acinar and tubulopapillary patterns R2: clear cell RCC, Fuhrman 3-4, large alveolar and solid patterns 3 R3: clear cell RCC, Fuhrman 3-4, solid pattern R4: clear cell RCC, Fuhrman 3-4, solid pattern R1: clear cell RCC, Fuhrman 2, acinar pattern 4 M1: clear cell RCC, Fuhrman 2, acinar pattern 5 R1: Unclassified RCC, intermixed clear and eosinophilic cells, Fuhrman 2-3, papillary architecture 1
2 Supplementary Table S2. IMPACT gene list and their respective chromosomal positions. Gene Symbol RefSeq ID Chromosome ABL1 NM_ q34.1 ABL2 NM_ q25.2 AKT1 NM_ q32.32-q32.33 AKT2 NM_ q13.1-q13.2 AKT3 NM_ q44 ALK NM_ p23 ALOX12B NM_ p13.1 APC NM_ q21-q22 AR NM_ Xq12 ARAF NM_ Xp11.3-p11.23 ARHGAP26 NM_ q31 ARID1A NM_ p36.1-p35 ASXL1 NM_ q11 ATM NM_ q22-q23 ATRX NM_ Xq21.1 AURKA NM_ q13 BAP1 NM_ p21.31-p21.2 BCL2L1 NM_ q11.21 BCL6 NM_ q27 BIRC2 NM_ q22 BRAF NM_ q34 BRCA1 NM_ q21-q24 BRCA2 NM_ q12-q13 CARD11 NM_ p22 CBL NM_ q23.3-qter CBLB NM_ q CBLC NM_ q13.2 CCND1 NM_ q13 CCNE1 NM_ q12 CD79B NM_ q23 CDC42EP2 NM_ q13 CDC73 NM_ q25 CDH1 NM_ q22.1 CDK4 NM_ q13 CDK6 NM_ q21-q22 CDK8 NM_ q12 CDKN2A NM_ p21 CDKN2B NM_ p21 CDKN2C NM_ p32.3 CEBPA NM_ q13.1 CHEK1 NM_ q24.2 CHEK2 NM_ q12.1 2
3 CREBBP NM_ p13.3 CRKL NM_ q11.21 CRLF2 NM_ Xp22.3 and Yp11.3 CSF1R NM_ q32 CTNNB1 NM_ p21 CYLD NM_ q12-q13 DAXX NM_ p21.3 DDR2 NM_ q12-q23 DICER1 NM_ q32.2 DIS3 NM_ q21.32 DNMT1 NM_ p13.2 DNMT3A NM_ p23 DNMT3B NM_ q11.2 EGFR NM_ p12 EIF4EBP1 NM_ p12 EP300 NM_ q13.2 EPHA3 NM_ p11.2 EPHA5 NM_ q13.1 EPHA6 NM_ q12.1 EPHA7 NM_ q16.3 EPHA8 NM_ p36.12 EPHB1 NM_ q21-q23 EPHB4 NM_ q22 EPHB6 NM_ q33-q35 ERBB2 NM_ q11.2-q12 ERBB3 NM_ q13 ERBB4 NM_ q33.3-q34 ERG NM_ q22.3 ESR1 NM_ q24-q27 ETV1 NM_ p22 ETV6 NM_ p13 EZH2 NM_ q35-q36 FAM123B NM_ Xq11.1 FAM46C NM_ p12 FAS NM_ q24.1 FBXW7 NM_ q31.23 FGFR1 NM_ p12 FGFR2 NM_ q25.3-q26 FGFR3 NM_ p16.3 FGFR4 NM_ q33-qter FH NM_ q42.1 FLCN NM_ p11.2 FLT1 NM_ q12 FLT3 NM_ q12 FOXL2 NM_ q23 GATA1 NM_ Xp11.23 GATA2 NM_ q21 GATA3 NM_ p15 3
4 GNA11 NM_ p13.3 GNAQ NM_ q21 GNAS NM_ q13.2-q13.3 GOLPH3 NM_ p13.2 GRIN2A NM_ p13.2 GSK3B NM_ q13.3 HDAC2 NM_ q21 HIF1A NM_ q23.2 HMGA2 NM_ q15 HNF1A NM_ q24.31 HRAS NM_ p15.5 HSP90AA1 NM_ q32.33 IDH1 NM_ q32-qter IDH2 NM_ q21-qter IGF1R NM_ q26.3 IGFBP7 NM_ q12 IKBKE NM_ q31 IKZF1 NM_ pter-7qter INSR NM_ p13.3-p13.2 IRS1 NM_ q36 IRS2 NM_ q34 JAK1 NM_ p32.3-p31.3 JAK2 NM_ p24 JAK3 NM_ p13-p12 JUN NM_ p32-p31 KDM5C NM_ Xp11.22-p11.21 KDM6A NM_ Xp11.2 KDR NM_ q11-q12 KEAP1 NM_ p13.2 KIT NM_ q11-q12 KLF6 NM_ p15 KRAS NM_ p12.1 LDHA NM_ p15.1 LGR6 NM_ q32.1 MAGI2 NM_ q21 MAP2K1 NM_ q22.1-q22.33 MAP2K2 NM_ p13.3 MAP2K4 NM_ p11.2 MAP3K8 NM_ p11.2 MCL1 NM_ q21 MDM2 NM_ q13-q14 MDM4 NM_ q32 MEN1 NM_ q13 MET NM_ q31 MITF NM_ p14.1-p12.3 MLH1 NM_ p22.3 MLL NM_ q23 MLL2 NM_ q12-q13 4
5 MLL3 NM_ q36 MLST8 NM_ p13.3 MPL NM_ p34 MSH2 NM_ p21 MSH6 NM_ p16 mtor NM_ p36 MYB NM_ q22-q23 MYC NM_ q24 MYCL1 NM_ p34.3 MYCN NM_ p24.3 NCOA2 NM_ q13 NF1 NM_ q11.2 NF2 NM_ q12.2 NFE2L2 NM_ q31 NFKB1 NM_ q24 NFKB2 NM_ q24 NKX2-1 NM_ q13.3 NOTCH1 NM_ q34.3 NOTCH2 NM_ p13-p11 NOTCH3 NM_ p13.2-p13.1 NOTCH4 NM_ p21.3 NPM1 NM_ q35.1 NRAS NM_ p13.2 NTRK1 NM_ q21-q22 NTRK2 NM_ q22.1 NTRK3 NM_ q24-q25 PAK7 NM_ p12 PARK2 NM_ q25.2-q27 PARP1 NM_ q41-q42 PAX5 NM_ p13.2 PBRM1 NM_ p21 PDGFRA NM_ q12 PDGFRB NM_ q33.1 PHOX2B NM_ p13 PIK3C2G NM_ p12 PIK3CA NM_ q26.3 PIK3CB NM_ q21-qter PIK3CD NM_ p36.2 PIK3CG NM_ q22 PIK3R1 NM_ q13.1 PIK3R2 NM_ q13.2-q13.4 PIK3R3 NM_ p34.1 PKM2 NM_ q22-qter PLK2 NM_ q12.1-q13.2 PNRC1 NM_ q16.1 PREX2 NM_ q13.1 PRKAR1A NM_ q23-q24 PRKCI NM_ q26.3 5
6 PTCH1 NM_ q22.1-q31 PTEN NM_ q23 PTPN11 NM_ q24.1 PTPRD NM_ p24.1-p23 PTPRS NM_ p13.3 RAF1 NM_ p25 RARA NM_ q21.1 RB1 NM_ q14.2 REL NM_ p13-p12 RET NM_ q11.2 RICTOR NM_ p13.1 RPTOR NM_ q25.3 RUNX1 NM_ q22.3 SDHB NM_ p36.1-p35 SETD2 NM_ p21.31 SHQ1 NM_ p13 SMAD4 NM_ q21.1 SMARCA4 NM_ p13.3 SMARCB1 NM_ q11.23 SMO NM_ q32.1 SOCS1 NM_ p13.13 SOX2 NM_ q26.3-q27 SPOP NM_ q21.33 SRC NM_ q12-q13 STK11 NM_ p13.3 SUFU NM_ q24.32 TBK1 NM_ q14.2 TEK NM_ p21 TERT NM_ p15.33 TET1 NM_ q21 TET2 NM_ q24 TGFBR2 NM_ p22 TMPRSS2 NM_ q22.3 TNFAIP3 NM_ q23-q25 TOP1 NM_ q12-q13.1 TP53 NM_ p13.1 TP63 NM_ q27-q29 TSC1 NM_ q34 TSC2 NM_ p13.3 TSHR NM_ q24-q31 VHL NM_ p25.3 WT1 NM_ p13 YAP1 NM_ q13 YES1 NM_ p11.31-p
7 Supplementary Table S3. Impact assay run statistics on all samples. Please separate excel spreadsheet, table too large for inclusion here. Supplementary Table S4. Impact assay exon coverage on all samples. Please separate excel spreadsheet, table too large for inclusion here. 7
8 Supplementary Table S5. WEC run statistics on patients #4 and #5. PF UQ BASES ALIGNED PCT SELECTED BASES MEAN TARGET COVERAGE PCT USABLE BASES ON TARGET PCT TARGET BASES 2X PCT TARGET BASES 10X PCT TARGET BASES 20X PCT TARGET BASES 30X TOTAL SAMPLE READS Pt 4 (N) 70,182,339 5,123,974, % % 96.54% 91.96% 86.08% 79.47% Pt 4 (T) 54,314,779 3,920,678, % % 95.86% 89.77% 81.73% 72.13% Pt 5 (N) 77,253,432 5,659,907, % % 96.39% 91.70% 86.10% 80.08% Pt 5 (T) 93,018,700 6,812,455, % % 96.67% 92.90% 88.33% 83.49% 8
9 Supplementary Table S6. List of all non-variant mutations detected by IMPACT assays in individual patient samples. Gene Sample Chr Genomic Coordinates (GRCh37) REF ALT AA Change Effect Transcript ID Allele Freq % VHL Pt G T E94* Nonsense NM_ PBRM1 Pt T A E991D Missense NM_ PHOX2B Pt C A G20V Missense NM_ NFKB1 Pt T C L611P Missense NM_ NFKB1 Pt C T A623V Missense NM_ TSC1 Pt G - P311fs Frameshift NM_ VHL Pt C A H115N Missense NM_ TP53 Pt C T R273H Missense NM_ JAK1 Pt C - R110fs Frameshift NM_ IGF1R Pt C S847fs Frameshift NM_ BAP1 Pt C A Splice e9-1 Splice Site NM_ VHL Pt GA - G212 Frameshift NM_ MTOR Pt G T Q2223K Missense NM_ MLL3 Pt A G V2060A Missense NM_ VHL Pt T C L118P Missense NM_ PBRM1 Pt TCACTGCTGAA - E1360fs Frameshift NM_ ATM Pt T - N1044fs Frameshift NM_ DAXX Pt G A T684M Missense NM_ KEAP1 Pt G A S102L Missense NM_
10 Supplementary Table S7. Non-variant mutations identified by WEC in patients #4 and #5. Genomic Coordinates (GRCh37) REF ALT AA Gene Pt ID Chr Change Effect Transcript.ID AKR7A3 Pt A T M/K Missense NM_ SLC35A3 Pt C T L/F Missense NM_ TROVE2 Pt GA G. Frameshift NR_ CAD Pt C G S/R Missense NM_ OXER1 Pt T A H/L Missense NM_ RANBP2 Pt A T N/Y Missense NM_ ZNF717 Pt TC T. Frameshift NM_ ATP6V1G2- DDX39B Pt T C K/E Missense NR_ ABCF1 Pt T C F/S Missense NM_ ALDH8A1 Pt T C S/G Missense NM_ JARID2 Pt GA G. Frameshift NM_ NEUROD6 Pt G A P/S Missense NM_ TOPORS Pt T C N/S Missense NM_ HABP4 Pt G A A/T Missense NM_ PBLD Pt A T I/N Missense NM_ FAM171A1 Pt TG T. Frameshift NM_ ATM Pt AT A. Frameshift NM_ KLF5 Pt G T W/L Missense NM_ ANKRD20A9P Pt C CA. Frameshift NR_ MIR1197 Pt GA G. Frameshift NR_ PLA2G15 Pt T G L/W Missense NM_ ITGA3 Pt C A P/Q Missense NM_ TMX4 Pt G A R/W Missense NM_ C20orf118 Pt T A F/I Missense NM_ TSHZ2 Pt A T K/* Nonsense NM_ KIF17 Pt C A R/M Missense NM_ AGL Pt A G K/E Missense NM_ IGSF8 Pt G T A/E Missense NM_ PRG4 Pt A T R/* Nonsense NM_ FBXO2 Pt C CGCG A/AP Frameshift NM_ WDR54 Pt A G S/G Missense NM_ STAMBP Pt CT C. Frameshift NM_ PVRL3 Pt G A W/* Nonsense NM_ ISY1 Pt C G. Splice Site NM_ ISY1-RAB43 Pt G T L/I Missense NM_ C3orf25 Pt T C K/E Missense NM_ SI Pt A C V/G Missense NM_ COL7A1 Pt CT C. Frameshift NM_ DCP1A Pt TA T. Frameshift NM_ PARP14 Pt T TAC. Frameshift NM_ Allele Freq % 10
11 PPEF2 Pt G T P/H Missense NM_ DAB2 Pt T C K/E Missense NM_ SSBP2 Pt T C R/G Missense NM_ NMUR2 Pt C T C/Y Missense NM_ TAP1 Pt G A P/S Missense NM_ DAXX Pt G A R/* Nonsense NR_ FTSJD2 Pt G A E/K Missense NM_ STL Pt A T Y/N Missense NR_ GTPBP10 Pt A C E/A Missense NM_ SSPO Pt GC G. Frameshift NM_ VCPIP1 Pt T A N/I Missense NM_ TJP2 Pt A T T/S Missense NM_ ODF2 Pt A G E/G Missense NM_ NOXA1 Pt G C G/R Missense NM_ GAD2 Pt G A G/S Missense NM_ ZNF33A Pt A G I/V Missense NM_ MCU Pt T C Y/H Missense NM_ P4HA1 Pt T C N/S Missense NM_ KIAA0913 Pt T C V/A Missense NM_ ECHS1 Pt T G K/T Missense NM_ AGAP4 Pt CA C. Frameshift NM_ IFIT5 Pt G GT. Frameshift NM_ NAP1L4 Pt T G E/D Missense NM_ SPON1 Pt C G P/R Missense NM_ SLC22A24 Pt A G F/S Missense NM_ MALAT1 Pt A T K/N Missense NR_ MALAT1 Pt A T I/F Missense NR_ FAM138D Pt GT G. Frameshift NR_ ATP8A2 Pt A G K/R Missense NM_ ANKRD20A9P Pt C CA. Frameshift NR_ MIS18BP1 Pt C G D/H Missense NM_ NEMF Pt G T S/* Nonsense NM_ TDP1 Pt G A R/Q Missense NM_ SAV1 Pt GA G. Frameshift NM_ SPATA5L1 Pt A G T/A Missense NM_ SMYD4 Pt T A R/* Nonsense NM_ FLJ90757 Pt C A R/M Missense NR_ C19orf28 Pt A G L/P Missense NM_ KEAP1 Pt G A S/L Missense NM_ KLK9 Pt G A L/F Missense NM_ NLRP12 Pt G C T/R Missense NM_ SLC9A8 Pt C A P/T Missense NM_ SON Pt G A R/Q Missense NM_ PI4KA Pt G A R/* Nonsense NM_ POM121L8P Pt AC A. Frameshift NR_ TLR8 Pt 5 X T G F/V Missense NM_ CYBB Pt 5 X G T. Splice Site NM_
12 SUPPLEMENTARY FIGURES Supplementary Figure 1. Flow chart depicts the IMPACT assay mutation identification and filtering algorithm. Images Intensities Base Calls and Qualities Fastq Conversion, Demultiplexing Alignment Re-Alignment (Local MSA) Remove Duplicates Recalibrate Base Qualities Performance Metrics Alignment Rate Coverage Distribution On-Target Specificity Library Complexity Sequence Variants SNVs Structural Variants Cancer Variants Somatic Mutations Indels Copy Alterations Rearrangements 12
13 Supplementary Figure 2. Flow chart depicts the WEC assay mutation identification and filtering algorithm. Remove Adapters Align BWA (hg19) Resolve Overlaps Mark Duplicates (Picard) Remove Multi-Mapper GATK Realignment Annotations dbsnp SNP eff Intersection of Mu Tect /GATK Germline Low Qual <3x in Tumor GATK BaseQ (Recalibration) Mu Tect/GATK Unified Genotyper VCF (Total Mutations) VQSR (Recalibrate VCF) Annotated VCF MAF Mark Potential Errors Near Indels SNP Cluster Low Qual 13
14 Supplementary Figure 3. Sanger validations of mutations in the targeted pathway indentified by IMPACT assay for R1 of patients #1,#2, and #3. 14
15 Supplementary Figure 4. Sanger validations of mutations in the targeted pathway indentified by IMPACT assay for additional regions analyzed for patients #1, and #3. Nucleotide changes are circled in red. 15
16 Supplementary Figure 5. The Q2223K mtor mutation provokes hyperactivation of mtorc1 in serum free growth conditions. Immunoblot showing that a higher level of phosphorylation of S6K (T389) induced by Q2223K compared to wild-type mtor can be observed for hours following initiation of serum deprivation. HEK293T cells were co-transfected like in Figure 3d, and 24 hours later, cells were washed and subsequently maintained in serum free medium for the indicated lengths of time (1 to 20 hours) before harvest. 16
17 Supplementary Figure 6. Hyperactivation of mtorc1 induced by Q2223K is sensitive to rapalog-based mtor inhibition. Immunoblot showing that phosphorylation of S6K (T389), promoted by wild-type and Q2223K mtor, is markedly reduced by everolimus, temsirolimus, and rapamycin. Like in Figure 3e, HEK293T cells were co-transfected with Myc-tagged S6K and an expression plasmid for wild-type or Q2223K mtor. 24 hours later, cells were washed with serum free medium and exposed to 10% serum for 1 hour in the absence (-) or presence of the indicated concentrations of rapalog. 17
18 Supplementary Figure 7. Immunoblots detecting phosphorylated mtor at serine residues 2448 (S2448) and 2481 (S2481). Compared to wild-type mtor, the Q2223K mutant shows elevated phosphorylation at mtor (S2448), but not at (S2481). HEK293T cells were co-transfected and treated like in Figure 3d. 18
19 Supplementary Figure 8. Copy number plots for multiple tumor regions in patient #3 showing the loss of chromosome 9 only in tumor regions carrying the TSC1 nonsense mutation (R3, R4). R1 R2 R3 9- R chr 19
20 Supplementary Figure 9. Copy number plots for patient with tuberous sclerosis complex and metastatic unclassified RCC with extended benefit from everolimus therapy (44+ months). Plots show inherited hemizygous of TSC2 in the germline DNA and the current somatic loss (homozygous deletion) in the tumor 1 (T1), while tumor 2 (T2) had the inherited TSC2 loss with a concomitant nonsense mutation in the other allele (Q794*) shown in Sanger validation. N (vs unmatched N) - Germline T1 (vs unmatched N) 16p (-/+) 16p (-/-) 20
21 Supplementary Figure 10. PI3K pathway alterations as reported across 418 cases of clear cell RCC in TCGA. Alterations are detected in 42% of cases. Queried through the cbio cancer genomics portal (Cerami et al., Cancer Discov May;2(5): doi: / CD ). 21
22 Supplementary Figure 11: Hematoxylin and eosin (H&E) stains and immunohistochemistry for 4EBP1 phosphorylation (Thr 37/46), an mtorc1 downstream effector, in primary kidney tumors and adjacent normal kidney tissue for patients
Assessing the Economics of Genomic Medicine. Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center
 Assessing the Economics of Genomic Medicine Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center DISCLOSURES No conflicts Patents unenforced Consultancies unpaid
Assessing the Economics of Genomic Medicine Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center DISCLOSURES No conflicts Patents unenforced Consultancies unpaid
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study. February 2018
 Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Jocelyn Chapman, MD Division of Gynecologic Oncology. Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic
 Jocelyn Chapman, MD Division of Gynecologic Oncology Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic Genetics underlies all cancers Somatic or tumor genetics Germline or inherited genetics
Jocelyn Chapman, MD Division of Gynecologic Oncology Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic Genetics underlies all cancers Somatic or tumor genetics Germline or inherited genetics
Jennifer Hauenstein Oncology Cytogenetics Emory University Hospital Atlanta, GA
 Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Frampton et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing Supplementary Figure 1A Base substitution detection
SUPPLEMENTARY INFORMATION Frampton et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing Supplementary Figure 1A Base substitution detection
Supplementary Materials
 Supplementary Materials Recurrent SMARCA4 Mutations in Small Cell Carcinoma of the Ovary Petar Jelinic, Jennifer J. Mueller, Narciso Olvera, Fanny Dao, Sasinya N. Scott, Ronak Shah, JianJiong Gao, Nikolaus
Supplementary Materials Recurrent SMARCA4 Mutations in Small Cell Carcinoma of the Ovary Petar Jelinic, Jennifer J. Mueller, Narciso Olvera, Fanny Dao, Sasinya N. Scott, Ronak Shah, JianJiong Gao, Nikolaus
Illumina s Cancer Research Portfolio and Dedicated Workflows
 Illumina s Cancer Research Portfolio and Dedicated Workflows Michael Sohn Clinical Sales Specialist Spain&Italy 2017 2017 Illumina, Inc. All rights reserved. Illumina, 24sure, BaseSpace, BeadArray, BlueFish,
Illumina s Cancer Research Portfolio and Dedicated Workflows Michael Sohn Clinical Sales Specialist Spain&Italy 2017 2017 Illumina, Inc. All rights reserved. Illumina, 24sure, BaseSpace, BeadArray, BlueFish,
Sample Metrics. Allele Frequency (%) Read Depth Ploidy. Gene CDS Effect Protein Effect. LN Metastasis Tumor Purity Computational Pathology 80% 60%
 Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
SureSelect Cancer All-In-One Custom and Catalog NGS Assays
 SureSelect Cancer All-In-One Custom and Catalog NGS Assays Detect all cancer-relevant variants in a single SureSelect assay SNV Indel TL SNV Indel TL Single DNA input Single AIO assay Single data analysis
SureSelect Cancer All-In-One Custom and Catalog NGS Assays Detect all cancer-relevant variants in a single SureSelect assay SNV Indel TL SNV Indel TL Single DNA input Single AIO assay Single data analysis
Clinically Useful Next Generation Sequencing and Molecular Testing in Gliomas MacLean P. Nasrallah, MD PhD
 Clinically Useful Next Generation Sequencing and Molecular Testing in Gliomas MacLean P. Nasrallah, MD PhD Neuropathology Fellow Division of Neuropathology Center for Personalized Diagnosis (CPD) Glial
Clinically Useful Next Generation Sequencing and Molecular Testing in Gliomas MacLean P. Nasrallah, MD PhD Neuropathology Fellow Division of Neuropathology Center for Personalized Diagnosis (CPD) Glial
Precision Oncology: Experience at UW
 Precision Oncology: Experience at UW Colin Pritchard MD, PhD University of Washington, Department of Lab Medicine WSMOS Meeting November 1, 2013 Conflict of Interest Disclosures I declare the following,
Precision Oncology: Experience at UW Colin Pritchard MD, PhD University of Washington, Department of Lab Medicine WSMOS Meeting November 1, 2013 Conflict of Interest Disclosures I declare the following,
Table S1. Demographics of patients and tumor characteristics.
 Supplemental Tables for: A Clinical Actionability of Comprehensive Genomic Profiling for Management of Rare or Refractory Cancers Kim Hirshfield et al. Table S1. Demographics of patients and tumor characteristics.
Supplemental Tables for: A Clinical Actionability of Comprehensive Genomic Profiling for Management of Rare or Refractory Cancers Kim Hirshfield et al. Table S1. Demographics of patients and tumor characteristics.
What is the status of the technologies of "precision medicine?
 Session 2: What is the status of the technologies of "precision medicine? Gideon Blumenthal, MD, Clinical Team Leader, Thoracic and Head/Neck Oncology, Center for Drug Evaluation and Research (CDER), U.S.
Session 2: What is the status of the technologies of "precision medicine? Gideon Blumenthal, MD, Clinical Team Leader, Thoracic and Head/Neck Oncology, Center for Drug Evaluation and Research (CDER), U.S.
# 1 India s Best Selling Cancer Genomics Test Precision treatment for every Cancer Patient Patient Guide
 # 1 India s Best Selling Cancer Genomics Test Precision treatment for every Cancer Patient Patient Guide PositiveSelect is India s widely used Cancer Genomics test with every 8 out of 10 patients opting
# 1 India s Best Selling Cancer Genomics Test Precision treatment for every Cancer Patient Patient Guide PositiveSelect is India s widely used Cancer Genomics test with every 8 out of 10 patients opting
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
Changing the Culture of Cancer Care II. Eric Holland Fred Hutchinson Cancer Research Center University of Washington Seattle
 Changing the Culture of Cancer Care II Eric Holland Fred Hutchinson Cancer Research Center University of Washington Seattle Transforming Cancer Therapy Eric Holland Fred Hutchinson Cancer Research Center
Changing the Culture of Cancer Care II Eric Holland Fred Hutchinson Cancer Research Center University of Washington Seattle Transforming Cancer Therapy Eric Holland Fred Hutchinson Cancer Research Center
Accel-Amplicon Panels
 Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Clinical Grade Genomic Profiling: The Time Has Come
 Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Exons from 137 genes were targeted for hybrid selection and massively parallel sequencing.
 SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
Fluxion Biosciences and Swift Biosciences Somatic variant detection from liquid biopsy samples using targeted NGS
 APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST PROSPECTIVE
 AMP COMPANION MEETING SYMPOSIUM AT USCAP 2015 NEXT-GENERATION OF PATHOLOGY: ROLE OF PATHOLOGIST IN NGS-BASED PERSONALIZED MEDICINE APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST
AMP COMPANION MEETING SYMPOSIUM AT USCAP 2015 NEXT-GENERATION OF PATHOLOGY: ROLE OF PATHOLOGIST IN NGS-BASED PERSONALIZED MEDICINE APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each
 Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
The Center for PERSONALIZED DIAGNOSTICS
 The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope
 Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope National Medical Center Disclosures I have no disclosures
Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope National Medical Center Disclosures I have no disclosures
Next generation histopathological diagnosis for precision medicine in solid cancers
 Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Diagnostica Molecolare!
 Diagnostica Molecolare! Aldo Scarpa Unità Diagnostica Molecolare Azienda Ospedaliera Universitaria Integrata di Verona e ARC-NET Centro di Ricerca Applicata sul Cancro PDTA CARCINOMA POLMONARE - IL PAZIENTE
Diagnostica Molecolare! Aldo Scarpa Unità Diagnostica Molecolare Azienda Ospedaliera Universitaria Integrata di Verona e ARC-NET Centro di Ricerca Applicata sul Cancro PDTA CARCINOMA POLMONARE - IL PAZIENTE
Provide your cancer patients personalized treatment options with ClariFind
 ONCOLOGY ClariFind employs next-generation sequencing (NGS) for comprehensive genomic profiling of a patient s tumor sample to identify potential clinically actionable targets and associated therapies.
ONCOLOGY ClariFind employs next-generation sequencing (NGS) for comprehensive genomic profiling of a patient s tumor sample to identify potential clinically actionable targets and associated therapies.
Oncology Update: The 10 Most Talked About Breast Cancer Topics of The Top 10 10/24/2013. The Awareness Debate
 Oncology Update: The 10 Most Talked About Breast Cancer Topics of 2013 Michaela Tsai, MD Martha Bacon Stimpson Chair of Breast Oncology Virginia Piper Cancer Institute Minnesota Oncology The Top 10 1.
Oncology Update: The 10 Most Talked About Breast Cancer Topics of 2013 Michaela Tsai, MD Martha Bacon Stimpson Chair of Breast Oncology Virginia Piper Cancer Institute Minnesota Oncology The Top 10 1.
NeoTYPE Cancer Profiles
 NeoTYPE Cancer Profiles 30+ Multimethod Assays for Hematologic Diseases and Solid Tumors Molecular FISH Anatomic Pathology The next generation of diagnostic, prognostic, and therapeutic assessment What
NeoTYPE Cancer Profiles 30+ Multimethod Assays for Hematologic Diseases and Solid Tumors Molecular FISH Anatomic Pathology The next generation of diagnostic, prognostic, and therapeutic assessment What
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Illumina Trusight Myeloid Panel validation A R FHAN R A FIQ
 Illumina Trusight Myeloid Panel validation A R FHAN R A FIQ G E NETIC T E CHNOLOGIST MEDICAL G E NETICS, CARDIFF To Cover Background to the project Choice of panel Validation process Genes on panel, Protocol
Illumina Trusight Myeloid Panel validation A R FHAN R A FIQ G E NETIC T E CHNOLOGIST MEDICAL G E NETICS, CARDIFF To Cover Background to the project Choice of panel Validation process Genes on panel, Protocol
NeoTYPE Cancer Profiles
 NeoTYPE Cancer Profiles Multimethod Analysis of 25+ Hematologic Diseases and Solid Tumors Anatomic Pathology FISH Molecular The next generation of diagnostic, prognostic, and therapeutic assessment NeoTYPE
NeoTYPE Cancer Profiles Multimethod Analysis of 25+ Hematologic Diseases and Solid Tumors Anatomic Pathology FISH Molecular The next generation of diagnostic, prognostic, and therapeutic assessment NeoTYPE
Identification and clinical detection of genetic alterations of pre-neoplastic lesions Time for the PML ome? David Sidransky MD Johns Hopkins
 Identification and clinical detection of genetic alterations of pre-neoplastic lesions Time for the PML ome? David Sidransky MD Johns Hopkins February 3-5, 2016 Lansdowne Resort, Leesburg, VA Molecular
Identification and clinical detection of genetic alterations of pre-neoplastic lesions Time for the PML ome? David Sidransky MD Johns Hopkins February 3-5, 2016 Lansdowne Resort, Leesburg, VA Molecular
Clinical implication of MEN1 mutation in surgically resected thymic carcinoid patients
 Case Report Clinical implication of MEN1 in surgically resected thymic carcinoid patients Xiongfei Li 1 *, Mingbiao Li 1 *, Tao Shi 2 *, Renwang Liu 1, Dian Ren 1, Fan Yang 1, Sen Wei 1, Gang Chen 1, Jun
Case Report Clinical implication of MEN1 in surgically resected thymic carcinoid patients Xiongfei Li 1 *, Mingbiao Li 1 *, Tao Shi 2 *, Renwang Liu 1, Dian Ren 1, Fan Yang 1, Sen Wei 1, Gang Chen 1, Jun
Nature Genetics: doi: /ng Supplementary Figure 1. Depths and coverages in whole-exome and targeted deep sequencing data.
 Supplementary Figure 1 Depths and coverages in whole-exome and targeted deep sequencing data. Depth (top) and coverage (bottom) of whole-exome sequencing for 38 independent JPN cases (mean depth = 130)
Supplementary Figure 1 Depths and coverages in whole-exome and targeted deep sequencing data. Depth (top) and coverage (bottom) of whole-exome sequencing for 38 independent JPN cases (mean depth = 130)
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Systematic investigation of cancer-associated somatic point mutations in SNP databases HyunChul Jung 1,2, Thomas Bleazard 3, Jongkeun Lee 1 and Dongwan Hong 1 1. Cancer Genomics
SUPPLEMENTARY INFORMATION Systematic investigation of cancer-associated somatic point mutations in SNP databases HyunChul Jung 1,2, Thomas Bleazard 3, Jongkeun Lee 1 and Dongwan Hong 1 1. Cancer Genomics
Secuenciación masiva: papel en la toma de decisiones
 Secuenciación masiva: papel en la toma de decisiones Cancer is a Genetic Disease Development of cancer is driven by the acquisition of somatic genetic alterations: Nonsynonymous point mutations: missense.
Secuenciación masiva: papel en la toma de decisiones Cancer is a Genetic Disease Development of cancer is driven by the acquisition of somatic genetic alterations: Nonsynonymous point mutations: missense.
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester
 Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
Click to edit Master /tle style
 Click to edit Master /tle style Tel: (314) 747-7337 Toll Free: (866) 450-7697 Fax: (314) 747-7336 Email: gps@wustl.edu Website: gps.wustl.edu GENETIC TESTING IN CANCER Ka/nka Vigh-Conrad, PhD Genomics
Click to edit Master /tle style Tel: (314) 747-7337 Toll Free: (866) 450-7697 Fax: (314) 747-7336 Email: gps@wustl.edu Website: gps.wustl.edu GENETIC TESTING IN CANCER Ka/nka Vigh-Conrad, PhD Genomics
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making
 Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
IntelliGENSM. Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community.
 IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
Supplementary Figure 1. Copy Number Alterations TP53 Mutation Type. C-class TP53 WT. TP53 mut. Nature Genetics: doi: /ng.
 Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
The International Association for the Study of Lung Cancer (IASLC) Lung Cancer Staging Project, Data Elements
 Page 1 Contents 1.1. Registration... 2 1.2. Patient Characteristics... 3 1.3. Laboratory Values at Diagnosis... 5 1.4. Lung Cancers with Multiple Lesions... 8 1.5. Primary Tumour Description... 10 1.6.
Page 1 Contents 1.1. Registration... 2 1.2. Patient Characteristics... 3 1.3. Laboratory Values at Diagnosis... 5 1.4. Lung Cancers with Multiple Lesions... 8 1.5. Primary Tumour Description... 10 1.6.
Out-Patient Billing CPT Codes
 Out-Patient Billing CPT Codes Updated Date: August 3, 08 Client Billed Molecular Tests HPV DNA Tissue Testing 8764 No Medicare Billed - Molecular Tests NeoARRAY NeoARRAY SNP/Cytogenetic No 89 NeoLAB NeoLAB
Out-Patient Billing CPT Codes Updated Date: August 3, 08 Client Billed Molecular Tests HPV DNA Tissue Testing 8764 No Medicare Billed - Molecular Tests NeoARRAY NeoARRAY SNP/Cytogenetic No 89 NeoLAB NeoLAB
I have no conflicts of interest
 Deploying lung cancer molecular pathology guidelines in real life Neal Lindeman, MD Director, Molecular Pathology Brigham & Women s Hospital USCAP Annual Meeting AMP Companion Meeting San Antonio, T Sunday,
Deploying lung cancer molecular pathology guidelines in real life Neal Lindeman, MD Director, Molecular Pathology Brigham & Women s Hospital USCAP Annual Meeting AMP Companion Meeting San Antonio, T Sunday,
Diagnostic application of SNParrays to brain cancers
 Diagnostic application of SNParrays to brain cancers Adriana Olar 4/17/2018 No disclosures 55 yo M, focal motor seizure T2 T1-post C DIAGNOSIS BRAIN, LEFT FRONTAL LOBE, BIOPSY: - DIFFUSE GLIOMA, OLIGODENDROGLIAL
Diagnostic application of SNParrays to brain cancers Adriana Olar 4/17/2018 No disclosures 55 yo M, focal motor seizure T2 T1-post C DIAGNOSIS BRAIN, LEFT FRONTAL LOBE, BIOPSY: - DIFFUSE GLIOMA, OLIGODENDROGLIAL
EXAMPLE. - Potentially responsive to PI3K/mTOR and MEK combination therapy or mtor/mek and PKC combination therapy. ratio (%)
 Dr Kate Goodhealth Goodhealth Medical Clinic 123 Address Road SUBURBTOWN NSW 2000 Melanie Citizen Referring Doctor Your ref Address Dr John Medico 123 Main Street, SUBURBTOWN NSW 2000 Phone 02 9999 9999
Dr Kate Goodhealth Goodhealth Medical Clinic 123 Address Road SUBURBTOWN NSW 2000 Melanie Citizen Referring Doctor Your ref Address Dr John Medico 123 Main Street, SUBURBTOWN NSW 2000 Phone 02 9999 9999
Hematology Fusion/Expression Profile
 Hematology Fusion/Expression Profile Patient Name: Ordered By Date of Birth: Ordering Physician: Gender (M/F): Physician ID: Client: Accession #: Case #: Specimen Type: Body Site: Specimen ID: Ethnicity:
Hematology Fusion/Expression Profile Patient Name: Ordered By Date of Birth: Ordering Physician: Gender (M/F): Physician ID: Client: Accession #: Case #: Specimen Type: Body Site: Specimen ID: Ethnicity:
IMPLEMENTING NEXT GENERATION SEQUENCING IN A PATHOLOGY LABORATORY
 Madrid, Spain IMPLEMENTING NEXT GENERATION SEQUENCING IN A PATHOLOGY LABORATORY Dr. JL Rodríguez Peralto NGS Ion Torrent Oncomine Focus Assay - Implementation experience for EGFR mutation detection
Madrid, Spain IMPLEMENTING NEXT GENERATION SEQUENCING IN A PATHOLOGY LABORATORY Dr. JL Rodríguez Peralto NGS Ion Torrent Oncomine Focus Assay - Implementation experience for EGFR mutation detection
Thyroid Cancer Genomics. James A. Fagin MD Memorial Sloan Kettering Cancer Center
 Thyroid Cancer Genomics James A. Fagin MD Memorial Sloan Kettering Cancer Center Disclosure Information James Fagin I have the following financial relationships to disclose: Consultant for: LOXO Oncology
Thyroid Cancer Genomics James A. Fagin MD Memorial Sloan Kettering Cancer Center Disclosure Information James Fagin I have the following financial relationships to disclose: Consultant for: LOXO Oncology
What is Precision Medicine?
 Precision Medicine and the Treatment of Cancer Carlos L. Arteaga, MD Center for Cancer Targeted Therapies VICC Breast Cancer Program Vanderbilt-Ingram Cancer Center (VICC) Departments of Medicine and Cancer
Precision Medicine and the Treatment of Cancer Carlos L. Arteaga, MD Center for Cancer Targeted Therapies VICC Breast Cancer Program Vanderbilt-Ingram Cancer Center (VICC) Departments of Medicine and Cancer
Supplementary Figure 1. The overlap between statistically-significant and highconfidence
 Supplementary figures Supplementary Figure 1. The overlap between statistically-significant and highconfidence PPIs with previously reported PPIs in public databases. Venn diagrams showing the overlap
Supplementary figures Supplementary Figure 1. The overlap between statistically-significant and highconfidence PPIs with previously reported PPIs in public databases. Venn diagrams showing the overlap
Personalised cancer care Information for Medical Specialists. A new way to unlock treatment options for your patients
 Personalised cancer care Information for Medical Specialists A new way to unlock treatment options for your patients Contents Optimised for clinical benefit 4 Development history 4 Full FIND IT panel vs
Personalised cancer care Information for Medical Specialists A new way to unlock treatment options for your patients Contents Optimised for clinical benefit 4 Development history 4 Full FIND IT panel vs
Predictive biomarker profiling of > 1,900 sarcomas: Identification of potential novel treatment modalities
 Predictive biomarker profiling of > 1,900 sarcomas: Identification of potential novel treatment modalities Sujana Movva 1, Wenhsiang Wen 2, Wangjuh Chen 2, Sherri Z. Millis 2, Margaret von Mehren 1, Zoran
Predictive biomarker profiling of > 1,900 sarcomas: Identification of potential novel treatment modalities Sujana Movva 1, Wenhsiang Wen 2, Wangjuh Chen 2, Sherri Z. Millis 2, Margaret von Mehren 1, Zoran
August 17, Dear Valued Client:
 August 7, 08 Re: CMS Announces 6-Month Period of Enforcement Discretion for Laboratory Date of Service Exception Policy Under the Medicare Clinical Laboratory Fee Schedule (the 4 Day Rule ) Dear Valued
August 7, 08 Re: CMS Announces 6-Month Period of Enforcement Discretion for Laboratory Date of Service Exception Policy Under the Medicare Clinical Laboratory Fee Schedule (the 4 Day Rule ) Dear Valued
Webinar Series. Characterizing cancers from liquid biopsies and FFPE samples: The rise of targeted sequencing panels. Participating experts
 Characterizing cancers from liquid biopsies and FFPE samples: The rise of targeted sequencing panels April 13, 2016 Webinar Series Brought to you by the Science/ AAAS Custom Publishing Office Participating
Characterizing cancers from liquid biopsies and FFPE samples: The rise of targeted sequencing panels April 13, 2016 Webinar Series Brought to you by the Science/ AAAS Custom Publishing Office Participating
Targeted Molecular Diagnostics for Targeted Therapies in Hematological Disorders
 Targeted Molecular Diagnostics for Targeted Therapies in Hematological Disorders Richard D. Press, MD, PhD Dept of Pathology Knight Cancer Institute Knight Diagnostic Labs Oregon Health & Science University
Targeted Molecular Diagnostics for Targeted Therapies in Hematological Disorders Richard D. Press, MD, PhD Dept of Pathology Knight Cancer Institute Knight Diagnostic Labs Oregon Health & Science University
Oncomine Focus assay panel and Oncomine Knowledgebase Reporter.
 Oncomine Focus assay panel and Oncomine Knowledgebase Reporter. How it can help to identify relevant alteration and early phase trials. Dr Isabelle SOUBEYRAN Dr Emmanuel KHALIFA Molecular Pathology Unit
Oncomine Focus assay panel and Oncomine Knowledgebase Reporter. How it can help to identify relevant alteration and early phase trials. Dr Isabelle SOUBEYRAN Dr Emmanuel KHALIFA Molecular Pathology Unit
The Next Generation in Cancer Diagnostics.
 The Next Generation in Cancer Diagnostics. OncoTarget was created specifically for cancer patients. Every patient s cancer is unique, which is why discovering what makes it unique can be essential for
The Next Generation in Cancer Diagnostics. OncoTarget was created specifically for cancer patients. Every patient s cancer is unique, which is why discovering what makes it unique can be essential for
Blastic Plasmacytoid Dendritic Cell Neoplasm with DNMT3A and TET2 mutations (SH )
 Blastic Plasmacytoid Dendritic Cell Neoplasm with DNMT3A and TET2 mutations (SH2017-0314) Habibe Kurt, Joseph D. Khoury, Carlos E. Bueso-Ramos, Jeffrey L. Jorgensen, Guilin Tang, L. Jeffrey Medeiros, and
Blastic Plasmacytoid Dendritic Cell Neoplasm with DNMT3A and TET2 mutations (SH2017-0314) Habibe Kurt, Joseph D. Khoury, Carlos E. Bueso-Ramos, Jeffrey L. Jorgensen, Guilin Tang, L. Jeffrey Medeiros, and
Why Pathway/Network Analysis?
 6/8/16 More Depth on Pathway and Network Analysis Lincoln Stein Why Pathway/Network Analysis? Drama@c data size reduc@on: 1000 s of genes => dozens of pathways. Increase sta@s@cal power by reducing mul@ple
6/8/16 More Depth on Pathway and Network Analysis Lincoln Stein Why Pathway/Network Analysis? Drama@c data size reduc@on: 1000 s of genes => dozens of pathways. Increase sta@s@cal power by reducing mul@ple
Supporting Information
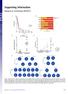 Supporting Information Rampal et al. 10.1073/pnas.1407792111 Fig. S1. Genetic events in leukemic transformation of chronic-phase MPNs. (A) Survival of post-mpn AML patients according to mutational status
Supporting Information Rampal et al. 10.1073/pnas.1407792111 Fig. S1. Genetic events in leukemic transformation of chronic-phase MPNs. (A) Survival of post-mpn AML patients according to mutational status
Mass spectrometric genotyping (OncoMap) Sensitivity of genotyping Laser capture microdissection Quantifying Langerhans cells in LCH samples
 Mass spectrometric genotyping (OncoMap) DNA was quantified using picogreen analysis, then subjected to whole genome amplification and used for further analysis only if DNA fingerprinting demonstrated non-biased
Mass spectrometric genotyping (OncoMap) DNA was quantified using picogreen analysis, then subjected to whole genome amplification and used for further analysis only if DNA fingerprinting demonstrated non-biased
Detecting Oncogenic Mutations in Whole Blood
 WHITE PAPER Detecting Oncogenic Mutations in Whole Blood Analytical validation of Cynvenio Biosystems LiquidBiopsy circulating tumor cell (CTC) capture and next-generation sequencing (NGS) September 2013
WHITE PAPER Detecting Oncogenic Mutations in Whole Blood Analytical validation of Cynvenio Biosystems LiquidBiopsy circulating tumor cell (CTC) capture and next-generation sequencing (NGS) September 2013
Vertical Magnetic Separation of Circulating Tumor Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients
 Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
Next Generation Sequencing in Haematological Malignancy: A European Perspective. Wolfgang Kern, Munich Leukemia Laboratory
 Next Generation Sequencing in Haematological Malignancy: A European Perspective Wolfgang Kern, Munich Leukemia Laboratory Diagnostic Methods Cytomorphology Cytogenetics Immunophenotype Histology FISH Molecular
Next Generation Sequencing in Haematological Malignancy: A European Perspective Wolfgang Kern, Munich Leukemia Laboratory Diagnostic Methods Cytomorphology Cytogenetics Immunophenotype Histology FISH Molecular
ACTIVITY 2: EXAMINING CANCER PATIENT DATA
 OVERVIEW Refer to the Overview of Cancer Discovery Activities for Key Concepts and Learning Objectives, Curriculum Connections, and Prior Knowledge, as well as background information, references, and additional
OVERVIEW Refer to the Overview of Cancer Discovery Activities for Key Concepts and Learning Objectives, Curriculum Connections, and Prior Knowledge, as well as background information, references, and additional
The lung cancer program of Alliance Against Cancer. Ruggero De Maria, President
 The lung cancer program of Alliance Against Cancer Ruggero De Maria, President Alliance Against Cancer (ACC) was established in 2002 by the Italian Ministry of Health with the task of promoting an active
The lung cancer program of Alliance Against Cancer Ruggero De Maria, President Alliance Against Cancer (ACC) was established in 2002 by the Italian Ministry of Health with the task of promoting an active
Clinical Grade Biomarkers in the Genomic Era Observations & Challenges
 Clinical Grade Biomarkers in the Genomic Era Observations & Challenges IOM Committee on Policy Issues in the Clinical Development & Use of Biomarkers for Molecularly Targeted Therapies March 31-April 1,
Clinical Grade Biomarkers in the Genomic Era Observations & Challenges IOM Committee on Policy Issues in the Clinical Development & Use of Biomarkers for Molecularly Targeted Therapies March 31-April 1,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
Figure S4. 15 Mets Whole Exome. 5 Primary Tumors Cancer Panel and WES. Next Generation Sequencing
 Figure S4 Next Generation Sequencing 15 Mets Whole Exome 5 Primary Tumors Cancer Panel and WES Get coverage of all variant loci for all three Mets Variant Filtering Sequence Alignments Index and align
Figure S4 Next Generation Sequencing 15 Mets Whole Exome 5 Primary Tumors Cancer Panel and WES Get coverage of all variant loci for all three Mets Variant Filtering Sequence Alignments Index and align
Best of ASCO 2014 Sarcoma
 Best of ASCO 2014 Sarcoma Robin L Jones Seattle Cancer Care Alliance University of Washington Fred Hutchinson Cancer Research Center Presentation Outline Overview progress made in sarcoma Highlight 2 trials
Best of ASCO 2014 Sarcoma Robin L Jones Seattle Cancer Care Alliance University of Washington Fred Hutchinson Cancer Research Center Presentation Outline Overview progress made in sarcoma Highlight 2 trials
Karl Kashofer, Phd Institut für Pathologie Medizinische Universität Graz
 Expanding on WHO guideline compliant molecular testing of central nervous system tumors by low density whole genome sequencing. Karl Kashofer, Phd Institut für Pathologie Medizinische Universität Graz
Expanding on WHO guideline compliant molecular testing of central nervous system tumors by low density whole genome sequencing. Karl Kashofer, Phd Institut für Pathologie Medizinische Universität Graz
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
A brain tumor and NGS/multiplexing
 A brain tumor and NGS/multiplexing Santiago Ramón y Cajal Jefe de Servicio. Hospital Vall d Hebron Catedrático de Anatomía Patológica U.A.B. Académico de Número de la Real Academia Nacional de Medicina
A brain tumor and NGS/multiplexing Santiago Ramón y Cajal Jefe de Servicio. Hospital Vall d Hebron Catedrático de Anatomía Patológica U.A.B. Académico de Número de la Real Academia Nacional de Medicina
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies
 Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Molecular. Oncology & Pathology. Diagnostic, Prognostic, Therapeutic, and Predisposition Tests in Precision Medicine. Liquid Biopsy.
 Molecular Oncology & Pathology Hereditary Cancer Somatic Cancer Liquid Biopsy Next-Gen Sequencing qpcr Sanger Sequencing Diagnostic, Prognostic, Therapeutic, and Predisposition Tests in Precision Medicine
Molecular Oncology & Pathology Hereditary Cancer Somatic Cancer Liquid Biopsy Next-Gen Sequencing qpcr Sanger Sequencing Diagnostic, Prognostic, Therapeutic, and Predisposition Tests in Precision Medicine
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Evolución Clonal de los Tumores y Biopsia Líquida en Cáncer de Pulmón
 Evolución Clonal de los Tumores y Biopsia Líquida en Cáncer de Pulmón Ignacio I. Wistuba, M.D. Professor and Chair Department of Translational Molecular Pathology The University of Texas M. D. Anderson
Evolución Clonal de los Tumores y Biopsia Líquida en Cáncer de Pulmón Ignacio I. Wistuba, M.D. Professor and Chair Department of Translational Molecular Pathology The University of Texas M. D. Anderson
Case Study. Chikako Shimizu, MD Breast and Medical Oncology Division National Cancer Center Hospital
 Case Study Chikako Shimizu, MD Breast and Medical Oncology Division National Cancer Center Hospital Triplenegative paradox Cortazar P et al. Lancet 2014 Chief Complaints A lump in the right breast Fertility
Case Study Chikako Shimizu, MD Breast and Medical Oncology Division National Cancer Center Hospital Triplenegative paradox Cortazar P et al. Lancet 2014 Chief Complaints A lump in the right breast Fertility
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Genomic alterations detectable in cfdna of EGFR-mutant p.t790m-positive and p.t790m-negative patients. (a-b) Lolliplots of gene level alterations in EGFR-mutant p.t790m mutant positive
Supplementary Figure 1 Genomic alterations detectable in cfdna of EGFR-mutant p.t790m-positive and p.t790m-negative patients. (a-b) Lolliplots of gene level alterations in EGFR-mutant p.t790m mutant positive
Schedule of Accreditation issued by United Kingdom Accreditation Service 2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK
 2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK Haematopathology and Oncology Diagnostic Services (HODS) Addenbrookes Hospital Hills Road Cambridge CB2 0QQ Contact: Brian Warner Tel: +44
2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK Haematopathology and Oncology Diagnostic Services (HODS) Addenbrookes Hospital Hills Road Cambridge CB2 0QQ Contact: Brian Warner Tel: +44
SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B and B
 SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B1-0916 and B1-0713. Copy number changes of the human chromosome 8 are common in many types of tumours. In most cases, losses of 8p sequences and gains of 8q
SALSA MLPA Probemix P014-B1 Chromosome 8 Lot B1-0916 and B1-0713. Copy number changes of the human chromosome 8 are common in many types of tumours. In most cases, losses of 8p sequences and gains of 8q
DNA Genetic Cancer Risk Test. Test Report
 Test Report CONTENTS SECTION 1 1-1. Customer Information 1-2. Test Report Summary 1-3. Test Report Details 1-4. List of Cancers / Tumors Tested SECTION 2 2-1. About the Test 2-2. References 1 CL0040-1_B
Test Report CONTENTS SECTION 1 1-1. Customer Information 1-2. Test Report Summary 1-3. Test Report Details 1-4. List of Cancers / Tumors Tested SECTION 2 2-1. About the Test 2-2. References 1 CL0040-1_B
Germline Multigene Panel Testing in Oncology: Genetic Counseling Perspective
 Germline Multigene Panel Testing in Oncology: Genetic Counseling Perspective Sarah L. Campian, MS CGC Certified Genetic Counselor Nancy & James Grosfeld Cancer Genetics Center Objectives Identify patients/families
Germline Multigene Panel Testing in Oncology: Genetic Counseling Perspective Sarah L. Campian, MS CGC Certified Genetic Counselor Nancy & James Grosfeld Cancer Genetics Center Objectives Identify patients/families
Next generation sequencing on FFPE tumor tissues for lung and colorectal carcinoma diagnostics and research Sakari Knuutila
 Next generation sequencing on FFPE tumor tissues for lung and colorectal carcinoma diagnostics and research Sakari Knuutila Faculty of Medicine / Haartman Institute / Department of Pathology / Sakari Knuutila
Next generation sequencing on FFPE tumor tissues for lung and colorectal carcinoma diagnostics and research Sakari Knuutila Faculty of Medicine / Haartman Institute / Department of Pathology / Sakari Knuutila
Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics
 Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics Director of Cytogenomics and Molecular Pathology Evidence-based
Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics Director of Cytogenomics and Molecular Pathology Evidence-based
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Individualisierte Therapie des CUP-Syndroms - Fakt oder Fiktion -
 Individualisierte Therapie des CUP-Syndroms - Fakt oder Fiktion - Alwin Krämer Klinische Kooperationseinheit Molekulare Hämatologie/Onkologie Medizinische Klinik V, Universität Heidelberg und Deutsches
Individualisierte Therapie des CUP-Syndroms - Fakt oder Fiktion - Alwin Krämer Klinische Kooperationseinheit Molekulare Hämatologie/Onkologie Medizinische Klinik V, Universität Heidelberg und Deutsches
The role of liquid biopsies in the treatment of advanced colon cancer
 The role of liquid biopsies in the treatment of advanced colon cancer Clara Montagut Hospital del Mar, Barcelona ESMO CRC Preceptorship Valencia, May 12, 2017 1 ESMO consensus guidelines for the management
The role of liquid biopsies in the treatment of advanced colon cancer Clara Montagut Hospital del Mar, Barcelona ESMO CRC Preceptorship Valencia, May 12, 2017 1 ESMO consensus guidelines for the management
Application of Whole Genome Microarrays in Cancer: You should be doing this test!!
 Application of Whole Genome Microarrays in Cancer: You should be doing this test!! Daynna Wolff, Ph.D. Director, Cytogenetics and Genomics Disclosures Clinical Laboratory Director and Employee, Medical
Application of Whole Genome Microarrays in Cancer: You should be doing this test!! Daynna Wolff, Ph.D. Director, Cytogenetics and Genomics Disclosures Clinical Laboratory Director and Employee, Medical
patient guide CancerNext-Expanded genetic testing for hereditary cancer Because knowing your risk can mean early detection and prevention
 patient guide CancerNext-Expanded genetic testing for hereditary cancer Because knowing your risk can mean early detection and prevention Know the Basics Cancer occurs in about 1 in 3 adults in their lifetime
patient guide CancerNext-Expanded genetic testing for hereditary cancer Because knowing your risk can mean early detection and prevention Know the Basics Cancer occurs in about 1 in 3 adults in their lifetime
COSMIC - Catalogue of Somatic Mutations in Cancer
 COSMIC - Catalogue of Somatic Mutations in Cancer http://cancer.sanger.ac.uk/cosmic https://academic.oup.com/nar/articl e-lookup/doi/10.1093/nar/gkw1121 Data In Large-scale systematic screens Detailed
COSMIC - Catalogue of Somatic Mutations in Cancer http://cancer.sanger.ac.uk/cosmic https://academic.oup.com/nar/articl e-lookup/doi/10.1093/nar/gkw1121 Data In Large-scale systematic screens Detailed
Paradigm Cancer Diagnostic (PCDx)
 Paradigm Cancer Diagnostic (PCDx) Date of Birth: PCDx Case#: Physician: Facility: PCDx-18-0XXXX Case/Specimen ID: Collection Site: Collection Date: Received for testing: Turnaround: 4 business days Tumor
Paradigm Cancer Diagnostic (PCDx) Date of Birth: PCDx Case#: Physician: Facility: PCDx-18-0XXXX Case/Specimen ID: Collection Site: Collection Date: Received for testing: Turnaround: 4 business days Tumor
CentoCancer STRIVE FOR THE MOST COMPLETE INFORMATION
 CentoCancer STRIVE FOR THE MOST COMPLETE INFORMATION CentoCancer our most comprehensive oncogenetics panel for hereditary mutations Hereditary pathogenic variants confer an increased risk of developing
CentoCancer STRIVE FOR THE MOST COMPLETE INFORMATION CentoCancer our most comprehensive oncogenetics panel for hereditary mutations Hereditary pathogenic variants confer an increased risk of developing
