Direct Identification of Gram-Positive Cocci from Routine Blood Cultures by Using AccuProbe Tests
|
|
|
- Brooke Sharp
- 5 years ago
- Views:
Transcription
1 JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 2004, p Vol. 42, No /04/$ DOI: /JCM Copyright 2004, American Society for Microbiology. All Rights Reserved. Direct Identification of Gram-Positive Cocci from Routine Blood Cultures by Using AccuProbe Tests Laura Lindholm* and Hannu Sarkkinen Department of Clinical Microbiology, Päijät-Häme Central Hospital, Lahti, Finland 1 Received 23 January 2004/Returned for modification 20 April 2004/Accepted 3 August 2004 Rapid and reliable identification of bacteria directly from blood cultures is important in clinical practice to guide appropriate antibiotic therapy. In this study, the performance of the AccuProbe (Gen-Probe, Inc., San Diego, Calif.) in direct identification of Staphylococcus aureus, Streptococcus pneumoniae, enterococci, and group A and B streptococci from positive blood culture bottles was evaluated by using 6-year routine clinical laboratory blood culture material from Päijät-Häme Central Hospital, Lahti, Finland. With the enterococcal and group A and B streptococcal probes, the diagnostic performance of the test was excellent at a cutoff value of 50,000 relative light units (RLU) as recommended by the manufacturer. However, with the S. aureus probe, although the specificity was very high (99.8%), the sensitivity was low (72.4%). To improve the clinical usability of the direct AccuProbe identification, optimal cutoff values for the individual AccuProbe tests were defined by using receiver-operating characteristic analysis. Consequently, cutoff values for S. aureus and S. pneumoniae tests were adjusted to 30,000 RLU and for enterococci and to 55,000 RLU for group A and B streptococci. With these adjustments, the performance of the AccuProbe tests, especially that for S. aureus, was significantly improved. Rapid and reliable identification of bacteria from blood culture is indisputably one of the most important functions of a clinical microbiology laboratory. Bloodstream infection is a potentially life-threatening condition that requires rapid, appropriate therapy. The increasing rate of nosocomial infections during the past decade (21), together with the threat from multiresistant infectious agents, further highlight the importance of early identification and tailored antimicrobial therapy. A rapid and accurate method for direct identification of pathogens from positive blood culture broth, combined with automated, continuously monitored blood culture systems, may greatly reduce the overall time from specimen collection to final identification. This may result in a reduction in the use of broad-spectrum antibiotics, since the diagnostic information will be available to the clinician earlier. However, relatively few methods for direct identification of bacteria from blood culture broth have been described. Standard assays comprising the immunological, tube coagulase, and stable endonuclease methods routinely used for identification of Staphylococcus aureus after subculture have been applied directly to positive blood culture bottles, but variable sensitivities and specificities have been reported (3, 12, 20). The use of PCR for the direct identification of Streptococcus pneumoniae from blood has also been described (7, 17, 22), but overall its use for this purpose has been rather limited owing to the amplification inhibitors present in blood (6). The extreme sensitivity of the method may also be a problem (14). However, rapid identification of S. aureus from blood culture by using the LightCycler system has already been reported (18), and the use of real-time fluorescence PCR for rapid identification of bacteria from blood culture may increase in the future. Fluorescent in situ hybridization (FISH) (8, 15, 16), as well as the * Corresponding author. Mailing address: Päijät-Häme Central Hospital, Department of Clin. Microbiology, Keskussairaalankatu 7, FIN , Lahti, Finland. Phone: Fax: laura.lindholm@phks.fi. hybridization protection assay (2), have also been shown to be quite reasonable methods for direct identification of microorganisms from blood. Direct identification of the most common gram-positive bacteria in blood cultures by using the commercially available DNA probe kit AccuProbe (Gen-Probe, Inc., San Diego, Calif.), which utilizes hybridization protection assay technology, has already been demonstrated (2; H. Sarkkinen, Abstr. 10th Int. Cong. Infect. Dis., p. 161, 2002). However, the test has been validated by the manufacturer only with the use of fresh growth from solid medium or from broth cultures and not with direct clinical specimens. We report here our 6-year routine clinical laboratory experience with the use of AccuProbe for direct identification of S. aureus, S. pneumoniae, enterococci, and group A and B streptococci from positive blood culture bottles. The aim of the present study was also to define the optimal conditions (cutoff values) of the individual Accu- Probe tests for direct identification. MATERIALS AND METHODS The present study involved all blood cultures obtained between June 1997 and July 2003 in Päijät-Häme Central Hospital, Lahti, Finland. The blood cultures were carried out at the request of clinicians as part of the routine care of patients suspected of bacteremia and were processed as part of our normal clinical microbiology workflow. Blood culture bottles (BacT/Alert FAN aerobic and standard anaerobic bottles) were incubated in a BacT/Alert continuous monitoring system (Organon Technica Corp., Durham, N.C.; biomérieux, Marcy l Etoile, France [since July 2001]) in accordance with the manufacturer s recommendations. Organon Tecnica replaced its vented BacT/Alert aerobic blood culture bottle with a redesigned nonvent blood culture bottle in 2000, so the BacT/Alert FAN nonvent aerobic blood culture bottle was adopted in our blood culture routine in February These two types of bottles (nonvent and vented) have, however, been shown to be equivalent (10, 19). When blood culture bottles were signaled as positive, broth from the positive bottle was Gram stained, and bottles containing gram-positive cocci were screened for S. aureus enterococcal or streptococcal rrna. Probe selection was guided by the Gram stain pattern: when the stain revealed gram-positive cocci in clusters, the S. aureus probe was selected and, if the gram-positive cocci were clearly in pairs or chains, then Str. pneumoniae, enterococcal, and group A and B streptococcal probes were tested. All five tests were performed when the Gram 5609
2 5610 LINDHOLM AND SARKKINEN J. CLIN. MICROBIOL. Downloaded from FIG. 1. AccuProbe test results with adjusted cutoff values (30,000 RLU with S. aureus and S. pneumoniae probes and 55,000 RLU with enterococcus, group A and group B streptococcus probes) versus culture. (a to e) Box and whisker plot representations show the distribution of true-positive and -negative and false-positive and -negative test results. The central box represents the distance between the first and third quartiles with the median RLU value between them marked with a horizontal line. The minimum RLU value is the origin of the lower whisker, and the maximum is the limit of the upper whisker. A /, AccuProbe positive/negative; C /, culture positive/negative. on September 7, 2017 by guest stain showed gram-positive cocci without a predominance of clusters or chains. Subcultures on both chocolate and fastidious anaerobe agar plates were performed to provide isolates for conventional identification methods and susceptibility testing. AccuProbe technology (Gen-Probe) utilizes a chemiluminescent acridinium ester labeled DNA probe directed toward the rrna of the target organisms. The manufacturer s instructions for specimen preparation were modified to use a pellet of bacteria made directly from positive blood culture broth as previously described (2). Briefly, 1.5 ml of broth from the positive blood culture bottle was centrifuged for 6sinamicrocentrifuge at 10,000 g to sediment the red blood cells. The supernatant was then centrifuged at 10,000 g for 1 min to concentrate the bacteria. A 1- l loopful of pelleted bacteria was then tested with the appropriate AccuProbe in accordance with the manufacturer s instructions by using reagents supplied by the manufacturer. The chemiluminescence of the hybridized probe was measured with a Gen-Probe Leader 50i luminometer. A positive result registered in the laboratory records was a luminometer reading above the cutoff value of 50,000 relative light units (RLU). This cutoff value has been determined by the manufacturer on the basis of readings obtained from in-house testing (sample prepared by using a colony from an agar plate). We reviewed all consecutive signal positive blood cultures in which direct Gram stain had revealed gram-positive cocci from laboratory records. The recorded luminometer readings were tabulated with the culture results and analyzed: the culture result being considered the gold standard. If both bottles in the same blood culture set were positive, only the first AccuProbe result per set was included in the analysis. During the period from June 1997 to July 2003, the total number of sets of blood cultures in our hospital was 33,695 (with one aerobic and one anaerobic bottle per set). During the same period there were 2,228 patients whose blood
3 VOL. 42, 2004 DIRECT IDENTIFICATION OF BACTERIA FROM BLOOD CULTURE 5611 FIG. 1 Continued. culture was positive. We tested 454 consecutive signal positive bottles (one per patient) directly with the AccuProbe when gram-positive cocci were seen in direct Gram stain. The number of different AccuProbe tests performed was as follows: 732 S. aureus tests (203 were culture positive and 529 were culture negative), 423 S. pneumoniae tests (142 were culture positive and 281 were culture negative), 377 enterococcus tests (55 were culture positive and 322 were culture negative), 372 group A streptococcus tests (18 were culture positive and 354 were culture negative), and 372 group B streptococcus tests (36 were culture positive and 336 were culture negative). Receiver-operating characteristic (ROC) analysis is a method for assessing the value of a quantitative diagnostic test and deciding where to set the threshold when the test is used. In the present study, ROC analysis (StatsDirect statistical software; StatsDirect, Ltd.) was used to adjust the cutoff values (measured in RLU) for the individual AccuProbe tests. RESULTS There were 203 culture positive and 529 culture negative S. aureus blood culture samples in the material. A total of 147 samples were found to be positive and 528 were negative by both methods. A total of 56 samples were found to be negative with the AccuProbe test but positive in the culture. In addition, there was one culture-negative but AccuProbe-positive case, with the very high RLU value of 562,252, which was clearly negative when repeated. However, since only the first test result of each patient was included, it was considered as an AccuProbe-positive but culture-negative case in all analyses. The sensitivity and specificity values of the S. aureus Accu- Probe test were 72.4% (95% confidence interval [CI] to 78.44%) and 99.8% (95% CI 98.95% to 100%), respectively. The predictive value of the positive test was 99.3% (95% CI to 99.98%), but the predictive value of the negative test was only 90.4% (95% CI to 92.68%). Owing to the low sensitivity of the test, 56 of 203 (27.6%) culture-positive cases were missed by the AccuProbe. On the other hand, the specificity of the test was very high, producing only one falsepositive case. To improve the clinical usability of the S. aureus AccuProbe test, ROC analysis was used to find a cutoff value that would increase the sensitivity of the test without excessively compromising the specificity of the test. When the cutoff value was set to 16,000 RLU, an equal number of false-positive and falsenegative cases were found. Setting the cutoff value higher than 16,000 RLU therefore increased the likelihood of a positive test result being a true-positive case. When the cutoff value was elevated to 30,000 RLU (Fig. 1a), 17 more culture-positive cases were detected by the AccuProbe at the expense of six more false-positive results. The median RLU value for these false-positive S. aureus samples was 31,472 RLU (Table 1), i.e., very close to the cutoff value. Similarly, with S. pneumoniae there were four false-negative AccuProbe results with 50,000 RLU as the cutoff value, and three false-negative results with 30,000 as the cutoff value (Fig. 1b and Table 1). For enterococcus, five false-positive Accu- Probe results were found with a cutoff value of 50,000 RLU and two fewer when 55,000 was used as the cutoff value. In addition, one false-negative result was also found (Fig. 1c and Table 1). With group A streptococcus, there was one falsepositive finding (50,750 RLU) with the cutoff value of 50,000 RLU but no false-negative or false-positive samples with a cutoff value of 55,000 RLU (Fig. 1d and Table 1). With group B (Fig. 1e) streptococcus there was only one false-positive sample, which turned out to be Streptococcus equi. The manufacturer reports cross-reaction with some isolates of S. equi in the package insert of the Group B Streptococcus Culture Identification Test. The diagnostic characteristics of all AccuProbe tests with the selected ROC analysis based cutoff values are given in Table 2. DISCUSSION Our study showed that for streptococcal and enterococcal probes, the sensitivity, specificity, and positive and negative TABLE 1. Contradictory results between AccuProbe tests with adjusted cutoff values versus culture AccuProbe test (organism and result) No. of samples RLU a Min Max Med S. aureus False positive 7 30, ,252 31,472 False negative 39 2,271 28,350 12,055 S. pneumoniae False positive 0 False negative 3 10,140 24,253 23,141 Enterococcus False positive 3 55,386 79,709 56,407 False negative 1 4,729 4,729 4,729 Group A streptococcus False positive 0 False negative 0 Group B streptococcus False positive 1 309, , ,773 False negative 0 a Abbreviations: Min, minimum RLU value for giving a false-positive or falsenegative test result; Max, maximum RLU value for giving a false-positive or false-negative test result; Med, median RLU value for giving a false-positive or false-negative test result.
4 5612 LINDHOLM AND SARKKINEN J. CLIN. MICROBIOL. Cutoff RLU and organism TABLE 2. Performance characteristics of AccuProbe tests with adjusted cutoff values % (95% CI) Sensitivity Specificity Positive predictive value Negative predictive value 30,000 S. aureus 80.8 ( ) 98.7 ( ) 95.9 ( ) 93.1 ( ) S. pneumoniae 97.9 ( ) 100 ( ) 100 ( ) 98.4 ( ) 55,000 Enterococcus 98.2 ( ) 99.1 ( ) 94.7 ( ) 99.7 ( ) Group A streptococcus 100 ( ) 100 ( ) 100 ( ) 100 ( ) Group B streptococcus 100 ( ) 99.7 ( ) 97.3 ( ) 100 ( ) predictive values of the AccuProbe tests were already excellent when the cutoff values recommended by the manufacturer are used. However, with the S. aureus probe, although the specificity was very high (99.8%), the sensitivity of the test (72.4%) was low. It was clear that the sensitivity of the test was unacceptable. The blood culture procedure in the present study included incubation in a continuous monitoring system, Gram staining, and use of the AccuProbe test when the stain demonstrated gram-positive cocci. According to this procedure, the sensitivity of the S. pneumoniae AccuProbe test for the direct identification was excellent. Pneumococci, however, are known to be subject in liquid cultures to autolysis, which first renders the cells gram negative and later brings about lysis of the culture. Consequently, it is possible that when autolysis has occurred, the Gram stain may not demonstrate gram-positive cocci, which may diminish the sensitivity of the blood culture procedure as a whole. However, we have never found a sample giving a positive signal in a continuous monitoring system that would have resulted in a negative Gram stain and then later positive subcultures on agar plates or positive acridine orange stains. Defining cutoff levels for diagnostic tests is a difficult task that combines ethical and practical considerations with statistical evidence. The main difficulty is not statistical but rather the need to decide how good the test should be in order for it to be clinically relevant. Therefore, the best clinical cutoff value is not necessarily the same as the best statistical cutoff value. Considering the potentially life-threatening nature of the sepsis, we were willing to accept a moderate number of false-positive results if the sensitivity of the test could be improved. To study the effects of different cutoff values on the diagnostic performance of the AccuProbe tests, ROC curve analysis was used (1, 23). On the basis of our analysis, the cutoff values for individual AccuProbe tests were adjusted as indicated in Table 2. With these adjustments the performance of the AccuProbe tests, especially that for S. aureus, was significantly improved. In our hospital, and in Finland generally, the most common blood culture pathogens are Escherichia coli, coagulase-negative staphylococci, S. aureus, S. pneumoniae, and Enterococcus spp. Along with enterococci, S. aureus and coagulase-negative staphylococci are among the causes of blood culture-positive nosocomial infections, the rate of which has increased during the past decade (4, 9). Our ROC analysis-based cutoff value adjustment improved the clinical usability of the S. aureus AccuProbe test, but the test still does not perform as well as the other AccuProbe tests. It may be that the sensitivity of the S. aureus AccuProbe test could be further improved with optimization of sample preparation protocol. A probe capable of identifying E. coli, as well as other gram-negative rods, at the species level would also be highly desirable. The development of such a probe has been hampered by the fact that the target sites are often detected not only in E. coli but also in Shigella spp. and in other enteric bacteria. However, an evaluation of a DNA probe matrix that contains a probe for E. coli has recently been described (11). That particular study examined the use of a Gen-Probe research prototype assay for direct identification of microorganisms from blood. Commercial application of this assay is not yet available. Overall, as noted in the introduction, relatively few methods for direct identification of bacteria from blood culture broth have been described, and their performance in large-scale routine use has not been reported. Very recently, M. Sogaard et al. (Abstr. 14th Eur. Cong. Clin. Microbiol. Infect. Dis., abstr. O57, 2004) reported the use in a routine setting of a PNA FISH assay, which is now available as a kit. In the future, a study comparing the performances of the AccuProbe and the PNA FISH kit in routine clinical laboratory use would be of great interest. Munson et al. recently reported that, as far as antimicrobial management is concerned, the most important information provided by the clinical microbiology laboratory is the first telephone call reporting positive blood culture and Gram stain results (13). Their data also demonstrated that release of antimicrobial susceptibility test results had little impact on antimicrobial usage. Since we have optimized the cutoff values for direct identification, on the basis of the high predictive values, we can reliably report these results, i.e., the species of the clinically important gram-positive cocci, at almost the same time (within 1 h) as the Gram stain results. In addition, a statistical antibiotic resistance pattern based on local data can be added to the report for guidance. Such a preliminary resistance pattern is indeed relevant, since the antibiotic resistance of the clinically most important bacteria, such as S. aureus, is still relatively stable in Finland (5, 9). However, the final decision about antimicrobial therapy is always made in conjunction with the use of other laboratory and clinical data. For a method to be successfully implemented in a routine microbiology laboratory, it should be fast, reliable, cost-effective, and simple to perform. The modified microcentrifuge method takes only 30 min to perform and was easy to adopt in our normal daily routines with only a few positive blood cultures per day. The sample preparation needs no special expertise, and the results are objective since they are measured
5 VOL. 42, 2004 DIRECT IDENTIFICATION OF BACTERIA FROM BLOOD CULTURE 5613 with a luminometer, which automatically calculates and prints out the test result. The AccuProbe culture identification test is clearly more expensive than identification by conventional methods. The additional cost per sample consists of reagent costs (ca. 6$ per probe) and labor costs of ca. 30 min per sample (one or more probes). However, considering the clinical significance of blood culture and the timely identification provided by the direct AccuProbe method, the additional cost is justified. In conclusion, on the basis of the present study we can now offer timely and accurate information to clinicians to assist them in the application of antimicrobial therapy in the early stages of infection. REFERENCES 1. Altman, D. G., and J. M. Bland Statistics notes: diagnostic tests. 3. Deceiver operating characteristic plots. BMJ 309: Davis, T. E., and D. D. Fuller Direct identification of bacterial isolates in blood cultures by using a DNA probe. J. Clin. Microbiol. 29: Davis, T. E., D. D. Fuller, and E. C. Aeschleman Rapid, direct identification of Staphylococcus aureus and Streptococcus pneumoniae from blood cultures using commercial immunologic kits and modified conventional tests. Diagn. Microbiol. Infect. Dis. 15: Edmond, M. B., S. E. Wallace, D. K. McClish, M. A. Pfaller, R. N. Jones, and R. P. Wenzel Nosocomial bloodstream infections in United States hospitals: a three-year analysis. Clin. Infect. Dis. 29: European Antibiotic Resistance Surveillance System EARSS annual report [Online.] 6. Fredricks, D. N., and D. A. Relman Improved amplification of microbial DNA from blood cultures by removal of the PCR inhibitor sodium polyanetholesulfonate. J. Clin. Microbiol. 36: Hassan-King, M., I. Baldeh, O. Secka, A. Falade, and B. Greenwood Detection of Streptococcus pneumoniae DNA in blood cultures by PCR. J. Clin. Microbiol. 32: Kempf, V. A. J., K. Trebesius, and I. B. Autenrieth Fluorescent in situ hybridization allows rapid identification of microorganisms in blood cultures. J. Clin. Microbiol. 38: Lyytikäinen, O., J. Lumio, H. Sarkkinen, E. Kolho, A. Kostiala, P. Ruutu, et al Nosocomial bloodstream infections in Finnish hospitals during Clin. Infect. Dis. 35: Marlowe, E. M., L. Gibson, J. Hogan, S. Kaplan, and D. A. Bruckner Conventional and molecular methods for verification of results obtained with BacT/Alert nonvent blood culture bottles. J. Clin. Microbiol. 41: Marlowe, E. M., J. J. Hogan, J. F. Hindler, I. Andruszkiewicz, P. Gordon, and D. A. Bruckner Application of an rrna probe matrix for rapid identification of bacteria and fungi from routine blood cultures. J. Clin. Microbiol. 41: McDonald, C. L., and K. Chapin Rapid identification of Staphylococcus aureus from blood culture bottles by a classic 2-hour tube coagulase test. J. Clin. Microbiol. 33: Munson, E. L., D. J. Diekema, S. E. Beekmann, K. C. Chapin, and G. V. Doern Detection and treatment of bloodstream infection: laboratory reporting and antimicrobial management. J. Clin. Microbiol. 41: Nikkari, S., I. J. McLaughlin, W. Bi, D. E. Dodge, and D. A. Relman Does blood of healthy subjects contain bacterial ribosomal DNA? J. Clin. Microbiol. 39: Oliveira, K., S. M. Brecher, A. Durbin, D. S. Shapiro, D. R. Schwartz, P. C. De Girolami, J. Dakos, G. W. Procop, D. Wilson, C. S. Hanna, G. Haase, H. Peltroche-Llacsahuanga, K. C. Chapin, M. C. Musgnug, M. H. Levi, C. Shoemaker, and H. Stender Direct identification of Staphylococcus aureus from positive blood culture bottles. J. Clin. Microbiol. 41: Oliveira, K., G. W. Procop, D. Wilson, J. Coull, and H. Stender Rapid identification of Staphylococcus aureus directly from blood cultures by fluorescence in situ hybridization with peptide nucleic acid probes. J. Clin. Microbiol. 40: Rudolph, K. M., A. J. Parkinson, C. M. Black, and L. W. Mayer Evaluation of polymerase chain reaction for diagnosis of pneumococcal pneumonia. J. Clin. Microbiol. 31: Shrestha, N. K., M. J. Tuohy, G. S. Hall, C. M. Isada, and G. W. Procop Rapid identification of Staphylococcus aureus and the meca gene from BacT/ALERT blood culture bottles by using the LightCycler system. J. Clin. Microbiol. 40: Snyder, J. W., K. S. Benzing, G. K. Munier, G. D. Bostic, P. S. Bozigar, R. Hanna, and M. J. Zervos Clinical comparison of nonvented aerobic BacT/Alert blood culture bottle and standard aerobic bottle for detection of microorganism in blood. J. Clin. Microbiol. 38: Speers, D. J., T. R. Olma, and G. L. Gilbert Evaluation of four methods for rapid identification of Staphylococcus aureus from blood cultures. J. Clin. Microbiol. 36: Weinstein, M. P., M. L. Towns, S. M. Quartey, S. Mirrett, L. G. Reimer, G. Parmigiani, and L. B. Reller The clinical significance of positive blood cultures in the 1990s: a prospective comprehensive evaluation of the microbiology, epidemiology, and outcome of bacteremia and fungemia in adults. Clin. Infect. Dis. 24: Zhang, Y., D. J. Isaacman, R. M. Wadowsky, J. Rydquist-White, J. C. Post, and G. D. Ehrlich Detection of Streptococcus pneumoniae in whole blood by PCR. J. Clin. Microbiol. 33: Zweig, M. H., and G. Campbell Receiver-operating characteristic (ROC) plots: a fundamental evaluation tool in clinical medicine. Clin. Chem. 39:
ESCMID Online Lecture Library. by author
 Microbiological evaluation: how to report the results Alvaro Pascual MD, PhD Infectious Diseases and Clinical Microbiology Unit. University Hospital Virgen Macarena University of Sevilla BSI management
Microbiological evaluation: how to report the results Alvaro Pascual MD, PhD Infectious Diseases and Clinical Microbiology Unit. University Hospital Virgen Macarena University of Sevilla BSI management
Blood culture 壢新醫院 病理檢驗科 陳啟清技術主任
 Blood culture 壢新醫院 病理檢驗科 陳啟清技術主任 A Positive Blood Culture Clinically Important Organism Failure of host defenses to contain an infection at its primary focus Failure of the physician to effectively eradicate,
Blood culture 壢新醫院 病理檢驗科 陳啟清技術主任 A Positive Blood Culture Clinically Important Organism Failure of host defenses to contain an infection at its primary focus Failure of the physician to effectively eradicate,
Epidemiology and Outcome of Nosocomial and Community-Onset Bloodstream Infection
 JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 2003, p. 3655 3660 Vol. 41, No. 8 0095-1137/03/$08.00 0 DOI: 10.1128/JCM.41.8.3655 3660.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 2003, p. 3655 3660 Vol. 41, No. 8 0095-1137/03/$08.00 0 DOI: 10.1128/JCM.41.8.3655 3660.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
on April 30, 2018 by guest
 JCM Accepted Manuscript Posted Online 21 October 2015 J. Clin. Microbiol. doi:10.1128/jcm.02024-15 Copyright 2015, American Society for Microbiology. All Rights Reserved. Cost effectiveness of 30-ml blood
JCM Accepted Manuscript Posted Online 21 October 2015 J. Clin. Microbiol. doi:10.1128/jcm.02024-15 Copyright 2015, American Society for Microbiology. All Rights Reserved. Cost effectiveness of 30-ml blood
The microbial diagnosis of infective endocarditis (IE)
 The microbial diagnosis of infective endocarditis (IE) Pierrette Melin Medical Microbiology pm-chulg sbimc 10.05.2007 1 Introduction for diagnosis Review of microbiological investigation of IE and perspectives
The microbial diagnosis of infective endocarditis (IE) Pierrette Melin Medical Microbiology pm-chulg sbimc 10.05.2007 1 Introduction for diagnosis Review of microbiological investigation of IE and perspectives
Impact of rapid in situ hybridization testing on coagulase-negative staphylococci positive blood cultures
 Journal of Antimicrobial Chemotherapy (2006) 58, 154 158 doi:10.1093/jac/dkl146 Advance Access publication 24 April 2006 Impact of rapid in situ hybridization testing on coagulase-negative staphylococci
Journal of Antimicrobial Chemotherapy (2006) 58, 154 158 doi:10.1093/jac/dkl146 Advance Access publication 24 April 2006 Impact of rapid in situ hybridization testing on coagulase-negative staphylococci
Reducing Blood Culture Contamination in Hospitalized Pediatric Patients
 ISSN: 2319-7706 Volume 4 Number 12 (2015) pp. 200-208 http://www.ijcmas.com Original Research Article Reducing Blood Culture Contamination in Hospitalized Pediatric Patients S. H. Khater Enas 1 * and Taha
ISSN: 2319-7706 Volume 4 Number 12 (2015) pp. 200-208 http://www.ijcmas.com Original Research Article Reducing Blood Culture Contamination in Hospitalized Pediatric Patients S. H. Khater Enas 1 * and Taha
Timing of Specimen Collection for Blood Cultures from Febrile Patients with Bacteremia
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2008, p. 1381 1385 Vol. 46, No. 4 0095-1137/08/$08.00 0 doi:10.1128/jcm.02033-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Timing of
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2008, p. 1381 1385 Vol. 46, No. 4 0095-1137/08/$08.00 0 doi:10.1128/jcm.02033-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Timing of
The Clinical Significance of Blood Cultures. Presented BY; Cindy Winfrey, MSN, RN, CIC, DON- LTC TM, VA- BC TM
 The Clinical Significance of Blood Cultures Presented BY; Cindy Winfrey, MSN, RN, CIC, DON- LTC TM, VA- BC TM OVERVIEW Blood cultures are considered an important laboratory tool used to diagnose serious
The Clinical Significance of Blood Cultures Presented BY; Cindy Winfrey, MSN, RN, CIC, DON- LTC TM, VA- BC TM OVERVIEW Blood cultures are considered an important laboratory tool used to diagnose serious
Clinical significance of potential contaminants in blood cultures among patients in a medical center
 Potential J Microbiol contaminants Immunol Infect. in blood cultures 2007;40:438-444 Original Article Clinical significance of potential contaminants in blood cultures among patients in a medical center
Potential J Microbiol contaminants Immunol Infect. in blood cultures 2007;40:438-444 Original Article Clinical significance of potential contaminants in blood cultures among patients in a medical center
MALDI-TOF MS: Translating Microbiology Laboratory Alphabet Soup to Optimize Antibiotic Therapy
 MALDI-TOF MS: Translating Microbiology Laboratory Alphabet Soup to Optimize Antibiotic Therapy September 8, 2017 Amy Carr, PharmD PGY-2 Infectious Diseases Pharmacy Resident Seton Healthcare Family amy.carr@ascension.org
MALDI-TOF MS: Translating Microbiology Laboratory Alphabet Soup to Optimize Antibiotic Therapy September 8, 2017 Amy Carr, PharmD PGY-2 Infectious Diseases Pharmacy Resident Seton Healthcare Family amy.carr@ascension.org
ARTICLE. Resource Utilization and Contaminated Blood Cultures in Children at Risk for Occult Bacteremia
 Resource Utilization and Contaminated Blood Cultures in Children at Risk for Occult Bacteremia Gershon S. Segal, MD; James M. Chamberlain, MD ARTICLE Objective: To measure the increases in resource utilization
Resource Utilization and Contaminated Blood Cultures in Children at Risk for Occult Bacteremia Gershon S. Segal, MD; James M. Chamberlain, MD ARTICLE Objective: To measure the increases in resource utilization
Tandem MS in Microbiology. Chris Doern, PhD D(ABMM) UT Southwestern Medical Center Dallas, TX
 Tandem MS in Microbiology Chris Doern, PhD D(ABMM) UT Southwestern Medical Center Dallas, TX Learning Objectives After the presentation you should be able to: 1. Understand the mass spectrometry methodology
Tandem MS in Microbiology Chris Doern, PhD D(ABMM) UT Southwestern Medical Center Dallas, TX Learning Objectives After the presentation you should be able to: 1. Understand the mass spectrometry methodology
Staphylococci. Gram stain: gram positive cocci arranged in clusters.
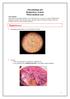 Microbiology lab Respiratory system Third medical year Lab contents: Gram positive bacteria (Staphylococcus and Streptococcus spp), two types of filamentous fungi (Aspergillus and Penicillium spp), and
Microbiology lab Respiratory system Third medical year Lab contents: Gram positive bacteria (Staphylococcus and Streptococcus spp), two types of filamentous fungi (Aspergillus and Penicillium spp), and
CONSIDERATIONS IN UTI DETECTION AND POTENTIAL IMPACT ON ANTIBIOTIC STEWARDSHIP
 CONSIDERATIONS IN UTI DETECTION AND POTENTIAL IMPACT ON ANTIBIOTIC STEWARDSHIP ERIN H. GRAF, PHD, D(ABMM) Director, Infectious Disease Diagnostics Laboratory Assistant Professor, Clinical Pathology and
CONSIDERATIONS IN UTI DETECTION AND POTENTIAL IMPACT ON ANTIBIOTIC STEWARDSHIP ERIN H. GRAF, PHD, D(ABMM) Director, Infectious Disease Diagnostics Laboratory Assistant Professor, Clinical Pathology and
Microbiological diagnosis of infective endocarditis; what is new?
 Microbiological diagnosis of infective endocarditis; what is new? Dr Amani El Kholy, MD Professor of Clinical Pathology (Microbiology), Faculty of Medicine, Cairo University ESC 2017 1 Objectives Lab Diagnostic
Microbiological diagnosis of infective endocarditis; what is new? Dr Amani El Kholy, MD Professor of Clinical Pathology (Microbiology), Faculty of Medicine, Cairo University ESC 2017 1 Objectives Lab Diagnostic
EDUCATIONAL COMMENTARY THROAT CULTURES LEARNING OUTCOMES. Upon completion of this exercise, the participant should be able to:
 EDUCATIONAL COMMENTARY THROAT CULTURES LEARNING OUTCOMES Upon completion of this exercise, the participant should be able to: distinguish three types of hemolysis produced by bacterial colonies. discuss
EDUCATIONAL COMMENTARY THROAT CULTURES LEARNING OUTCOMES Upon completion of this exercise, the participant should be able to: distinguish three types of hemolysis produced by bacterial colonies. discuss
Received 21 April 1997/Returned for modification 30 June 1997/Accepted 28 August 1997
 JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 1997, p. 3258 3263 Vol. 35, No. 12 0095-1137/97/$04.00 0 Copyright 1997, American Society for Microbiology Comparison of Agar Dilution, Broth Microdilution, E-Test,
JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 1997, p. 3258 3263 Vol. 35, No. 12 0095-1137/97/$04.00 0 Copyright 1997, American Society for Microbiology Comparison of Agar Dilution, Broth Microdilution, E-Test,
Lab 4. Blood Culture (Media) MIC AMAL-NORA-ALJAWHARA 1
 Lab 4. Blood Culture (Media) 2018 320 MIC AMAL-NORA-ALJAWHARA 1 Blood Culture 2018 320 MIC AMAL-NORA-ALJAWHARA 2 What is a blood culture? A blood culture is a laboratory test in which blood is injected
Lab 4. Blood Culture (Media) 2018 320 MIC AMAL-NORA-ALJAWHARA 1 Blood Culture 2018 320 MIC AMAL-NORA-ALJAWHARA 2 What is a blood culture? A blood culture is a laboratory test in which blood is injected
320 MBIO Microbial Diagnosis. Aljawharah F. Alabbad Noorah A. Alkubaisi 2017
 320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Blood Culture What is a blood culture? A blood culture is a laboratory test in which blood is injected into bottles with culture
320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Blood Culture What is a blood culture? A blood culture is a laboratory test in which blood is injected into bottles with culture
Effects of Blood Volume Monitoring on the Rate of Positive Blood Cultures from the Emergency Room
 Ann Clin Microbiol Vol. 19, No. 3, September, 2016 http://dx.doi.org/10.5145/acm.2016.19.3.70 pissn 2288-0585 eissn 2288-6850 Effects of Blood Volume Monitoring on the Rate of Positive Blood Cultures from
Ann Clin Microbiol Vol. 19, No. 3, September, 2016 http://dx.doi.org/10.5145/acm.2016.19.3.70 pissn 2288-0585 eissn 2288-6850 Effects of Blood Volume Monitoring on the Rate of Positive Blood Cultures from
Bacteremia in Children: Etiologic Agents, Focal Sites, and Risk Factors
 Bacteremia in Children: Etiologic Agents, Focal Sites, and Risk Factors by L. F. Nimri, a M. Rawashdeh, b and M. M. Meqdam a a Department of Applied Biology, and b Department of Pediatrics, Jordan University
Bacteremia in Children: Etiologic Agents, Focal Sites, and Risk Factors by L. F. Nimri, a M. Rawashdeh, b and M. M. Meqdam a a Department of Applied Biology, and b Department of Pediatrics, Jordan University
Minimizing the Workup of Blood Culture Contaminants: Implementation and Evaluation of a Laboratory-Based Algorithm
 JOURNAL OF CLINICAL MICROBIOLOGY, July 2002, p. 2437 2444 Vol. 40, No. 7 0095-1137/02/$04.00 0 DOI: 10.1128/JCM.40.7.2437 2444.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, July 2002, p. 2437 2444 Vol. 40, No. 7 0095-1137/02/$04.00 0 DOI: 10.1128/JCM.40.7.2437 2444.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
II- Streptococci. Practical 3. Objective: Required materials: Classification of Streptococci: Streptococci can be classified according to:
 Practical 3 II- Streptococci Objective: 1. Use of blood agar to differentiate between,, and hemolytic streptococci. 2. To know Gram reaction, shape and arrangement of streptococci. 3. To differentiate
Practical 3 II- Streptococci Objective: 1. Use of blood agar to differentiate between,, and hemolytic streptococci. 2. To know Gram reaction, shape and arrangement of streptococci. 3. To differentiate
MALDI-TOF mass spectrometry for direct bacterial identification
 JCM Accepts, published online ahead of print on 17 February 2010 J. Clin. Microbiol. doi:10.1128/jcm.01780-09 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All
JCM Accepts, published online ahead of print on 17 February 2010 J. Clin. Microbiol. doi:10.1128/jcm.01780-09 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All
MALDI-TOF Mass Spectrometry: A New Rapid ID Method in Clinical Microbiology
 MALDI-TOF Mass Spectrometry: A New Rapid ID Method in Clinical Microbiology Patrick R. Murray, PhD WW Director, Scientific Affairs BD Diagnostic Systems Outline MALDI-TOF is the most important innovation
MALDI-TOF Mass Spectrometry: A New Rapid ID Method in Clinical Microbiology Patrick R. Murray, PhD WW Director, Scientific Affairs BD Diagnostic Systems Outline MALDI-TOF is the most important innovation
Screening for Urinary Tract Infection with the Sysmex UF-1000i Urine Flow Cytometer
 JOURNAL OF CLINICAL MICROBIOLOGY, Mar. 2011, p. 1025 1029 Vol. 49, No. 3 0095-1137/11/$12.00 doi:10.1128/jcm.01669-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Screening for
JOURNAL OF CLINICAL MICROBIOLOGY, Mar. 2011, p. 1025 1029 Vol. 49, No. 3 0095-1137/11/$12.00 doi:10.1128/jcm.01669-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Screening for
Retrospective and Prospective Verification of the Cepheid Xpert Flu Assay
 JCM Accepts, published online ahead of print on 20 July 2011 J. Clin. Microbiol. doi:10.1128/jcm.01162-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
JCM Accepts, published online ahead of print on 20 July 2011 J. Clin. Microbiol. doi:10.1128/jcm.01162-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Infective endocarditis (IE) By Assis. Prof. Nader Alaridah MD, PhD
 Infective endocarditis (IE) By Assis. Prof. Nader Alaridah MD, PhD Infective endocarditis (IE) is an inflammation of the endocardium.. inner of the heart muscle & the epithelial lining of heart valves.
Infective endocarditis (IE) By Assis. Prof. Nader Alaridah MD, PhD Infective endocarditis (IE) is an inflammation of the endocardium.. inner of the heart muscle & the epithelial lining of heart valves.
Veerle Compernolle, Gerda Verschraegen, and Geert Claeys* Department of Microbiology, Ghent University Hospital, Ghent, Belgium
 JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 2007, p. 154 158 Vol. 45, No. 1 0095-1137/07/$08.00 0 doi:10.1128/jcm.01115-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Combined Use
JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 2007, p. 154 158 Vol. 45, No. 1 0095-1137/07/$08.00 0 doi:10.1128/jcm.01115-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Combined Use
Evaluation of the feasibility of the VACUETTE Urine CCM tube for microbial testing of urine samples
 Evaluation of the feasibility of the VACUETTE Urine CCM tube for microbial testing of urine samples Background The VACUETTE Urine CCM tube is for the collection, transport and storage of urine samples
Evaluation of the feasibility of the VACUETTE Urine CCM tube for microbial testing of urine samples Background The VACUETTE Urine CCM tube is for the collection, transport and storage of urine samples
OUTLINE Laboratory Detection and Reporting of Streptococcus agalactiae
 OUTLINE Laboratory Detection and Reporting of Streptococcus agalactiae I. Importance of prenatal screening strategies II. Past approaches Erik Munson Clinical Microbiology Wheaton Franciscan Laboratory
OUTLINE Laboratory Detection and Reporting of Streptococcus agalactiae I. Importance of prenatal screening strategies II. Past approaches Erik Munson Clinical Microbiology Wheaton Franciscan Laboratory
Detection of microorganisms in blood specimens using matrix-assisted laser desorption ionization time-of-flight mass spectrometry: a review
 REVIEW 10.1111/j.1469-0691.2010.03290.x Detection of microorganisms in blood specimens using matrix-assisted laser desorption ionization time-of-flight mass spectrometry: a review M. Drancourt Unité de
REVIEW 10.1111/j.1469-0691.2010.03290.x Detection of microorganisms in blood specimens using matrix-assisted laser desorption ionization time-of-flight mass spectrometry: a review M. Drancourt Unité de
Simultaneous and Rapid Detection of Causative Pathogens in Community-acquired Pneumonia by Real-time PCR (1167)
 From the Japanese Association of Medical Sciences The Japanese Association for Infectious Diseases Simultaneous and Rapid Detection of Causative Pathogens in Community-acquired Pneumonia by Real-time PCR
From the Japanese Association of Medical Sciences The Japanese Association for Infectious Diseases Simultaneous and Rapid Detection of Causative Pathogens in Community-acquired Pneumonia by Real-time PCR
Laboratory Detection and Reporting of Streptococcus agalactiae
 Laboratory Detection and Reporting of Streptococcus agalactiae Erik Munson Clinical Microbiology Wheaton Franciscan Laboratory Milwaukee, Wisconsin The presenter states no conflict of interest and has
Laboratory Detection and Reporting of Streptococcus agalactiae Erik Munson Clinical Microbiology Wheaton Franciscan Laboratory Milwaukee, Wisconsin The presenter states no conflict of interest and has
Labquality External Quality Assessment Programmes General Bacteriology 1 2/2013
 Labquality External Quality Assessment Programmes General Bacteriology 1 2/2013 Photos and text: Markku Koskela, M.D., Ph.D. Clinical microbiology specialist Oulu, Finland Sample 11/2013 Pus sample from
Labquality External Quality Assessment Programmes General Bacteriology 1 2/2013 Photos and text: Markku Koskela, M.D., Ph.D. Clinical microbiology specialist Oulu, Finland Sample 11/2013 Pus sample from
Getting the Point of Injection Safety
 Getting the Point of Injection Safety Barbara Montana, MD, MPH, FACP Medical Director Communicable Disease Service Outbreak of Enterococcus faecalis endocarditis associated with an oral surgery practice
Getting the Point of Injection Safety Barbara Montana, MD, MPH, FACP Medical Director Communicable Disease Service Outbreak of Enterococcus faecalis endocarditis associated with an oral surgery practice
Enrichment culture of CSF is of limited value in the diagnosis of neonatal meningitis
 Enrichment culture of CSF is of limited value in the diagnosis of neonatal S. H. Chaudhry, D. Wagstaff, Anupam Gupta, I. C. Bowler, D. P. Webster To cite this version: S. H. Chaudhry, D. Wagstaff, Anupam
Enrichment culture of CSF is of limited value in the diagnosis of neonatal S. H. Chaudhry, D. Wagstaff, Anupam Gupta, I. C. Bowler, D. P. Webster To cite this version: S. H. Chaudhry, D. Wagstaff, Anupam
Chain of Infection Agent Mode of transmission Contact (direct, indirect, droplet spread) Airborne Common-vehicle spread Host
 Goals Microbiology of Healthcare-associated Infections William A. Rutala, Ph.D., M.P.H. Director, Statewide Program for Infection Control and Epidemiology and Research Professor of Medicine, University
Goals Microbiology of Healthcare-associated Infections William A. Rutala, Ph.D., M.P.H. Director, Statewide Program for Infection Control and Epidemiology and Research Professor of Medicine, University
Cell counting (EtBr) Before cell-lysis. Cell-lysis by 3% SDS beads beating. After cell-lysis
 Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
Is the Volume of Blood Cultured Still a Significant Factor in the Diagnosis of Bloodstream Infections?
 JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 2007, p. 2765 2769 Vol. 45, No. 9 0095-1137/07/$08.00 0 doi:10.1128/jcm.00140-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Is the Volume
JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 2007, p. 2765 2769 Vol. 45, No. 9 0095-1137/07/$08.00 0 doi:10.1128/jcm.00140-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Is the Volume
Evaluation of the Clinical Performance of an Automated Procalcitonin Assay for the Quantitative Detection of Bloodstream Infection
 Korean J Lab Med 2010;30:153-9 DOI 10.3343/kjlm.2010.30.2.153 Original Article Clinical Microbiology Evaluation of the Clinical Performance of an Automated Procalcitonin Assay for the Quantitative Detection
Korean J Lab Med 2010;30:153-9 DOI 10.3343/kjlm.2010.30.2.153 Original Article Clinical Microbiology Evaluation of the Clinical Performance of an Automated Procalcitonin Assay for the Quantitative Detection
Bloodstream infection has been a primary
 Bacteremia in children at the University Hospital in Riyadh, Saudi Arabia Fahad Abdullah Al-Zamil Riyadh, Saudi Arabia 118 Background: Bacteremia is a major pediatric health care problem despite the availability
Bacteremia in children at the University Hospital in Riyadh, Saudi Arabia Fahad Abdullah Al-Zamil Riyadh, Saudi Arabia 118 Background: Bacteremia is a major pediatric health care problem despite the availability
Labquality External Quality Assessment Programmes General Bacteriology 1 3/2014
 Labquality External Quality Assessment Programmes General Bacteriology 1 3/2014 Photos and text: Markku Koskela, M.D., Ph.D. Clinical microbiology specialist Nordlab Oulu, Finland Specimen 31/2014 Abscess
Labquality External Quality Assessment Programmes General Bacteriology 1 3/2014 Photos and text: Markku Koskela, M.D., Ph.D. Clinical microbiology specialist Nordlab Oulu, Finland Specimen 31/2014 Abscess
IMMULEX S. PNEUMONIAE OMNI
 IMMULEX S. PNEUMONIAE OMNI ImmuLex S. Pneumoniae OMNI For in vitro diagnostic use Application The ImmuLex S. pneumoniae Omni is a ready-touse latex test for detection of all 92 Streptococcus pneumoniae
IMMULEX S. PNEUMONIAE OMNI ImmuLex S. Pneumoniae OMNI For in vitro diagnostic use Application The ImmuLex S. pneumoniae Omni is a ready-touse latex test for detection of all 92 Streptococcus pneumoniae
MALDI Sepsityper Kit
 Instructions for Use MALDI Sepsityper Kit Kit for identification of microorganisms from positive blood cultures using the MALDI Biotyper system CARE products are designed to support our worldwide customers
Instructions for Use MALDI Sepsityper Kit Kit for identification of microorganisms from positive blood cultures using the MALDI Biotyper system CARE products are designed to support our worldwide customers
A new selective blood agar medium for Streptococcus pyogenes and other haemolytic streptococci
 J. clin. Path. (1964), 17, 231 A new selective blood agar medium for Streptococcus pyogenes and other haemolytic streptococci E. J. L. LOWBURY, A. KIDSON, AND H. A. LILLY From the Medical Research Council
J. clin. Path. (1964), 17, 231 A new selective blood agar medium for Streptococcus pyogenes and other haemolytic streptococci E. J. L. LOWBURY, A. KIDSON, AND H. A. LILLY From the Medical Research Council
Rapid Detection and Identification of Candidemia by Direct Blood Culturing on Solid
 JCM Accepted Manuscript Posted Online 19 October 2016 J. Clin. Microbiol. doi:10.1128/jcm.01787-16 Copyright 2016 American Society for Microbiology. 1 2 Rapid Detection and Identification of Candidemia
JCM Accepted Manuscript Posted Online 19 October 2016 J. Clin. Microbiol. doi:10.1128/jcm.01787-16 Copyright 2016 American Society for Microbiology. 1 2 Rapid Detection and Identification of Candidemia
Affinity of Doripenem and Comparators to Penicillin-Binding Proteins in Escherichia coli and ACCEPTED
 AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Multi-clonal origin of macrolide-resistant Mycoplasma pneumoniae isolates. determined by multiple-locus variable-number tandem-repeat analysis
 JCM Accepts, published online ahead of print on 30 May 2012 J. Clin. Microbiol. doi:10.1128/jcm.00678-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Multi-clonal origin
JCM Accepts, published online ahead of print on 30 May 2012 J. Clin. Microbiol. doi:10.1128/jcm.00678-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Multi-clonal origin
Frozen Master Mix Modification of Commercial Reverse-transcriptase PCR. for Detection of Influenza and Respiratory Syncytial Viruses
 JCM Accepted Manuscript Posted Online 28 January 2015 J. Clin. Microbiol. doi:10.1128/jcm.03457-14 Copyright 2015, American Society for Microbiology. All Rights Reserved. 1 2 Frozen Master Mix Modification
JCM Accepted Manuscript Posted Online 28 January 2015 J. Clin. Microbiol. doi:10.1128/jcm.03457-14 Copyright 2015, American Society for Microbiology. All Rights Reserved. 1 2 Frozen Master Mix Modification
Bacterial Screening of Platelet Concentrates
 Bacterial Screening of Platelet Concentrates Enda Cadden Platelet Donations Very Short shelf-life
Bacterial Screening of Platelet Concentrates Enda Cadden Platelet Donations Very Short shelf-life
PRESENTER: DENNIS NYACHAE MOSE KENYATTA UNIVERSITY
 18/8/2016 SOURCES OF MICROBIAL CONTAMINANTS IN BIOSAFETY LABORATORIES IN KENYA PRESENTER: DENNIS NYACHAE MOSE KENYATTA UNIVERSITY 1 INTRODUCTION Contamination occurs through avoidable procedural errors
18/8/2016 SOURCES OF MICROBIAL CONTAMINANTS IN BIOSAFETY LABORATORIES IN KENYA PRESENTER: DENNIS NYACHAE MOSE KENYATTA UNIVERSITY 1 INTRODUCTION Contamination occurs through avoidable procedural errors
Updated Review of Blood Culture Contamination
 CLINICAL MICROBIOLOGY REVIEWS, Oct. 2006, p. 788 802 Vol. 19, No. 4 0893-8512/06/$08.00 0 doi:10.1128/cmr.00062-05 Copyright 2006, American Society for Microbiology. All Rights Reserved. Updated Review
CLINICAL MICROBIOLOGY REVIEWS, Oct. 2006, p. 788 802 Vol. 19, No. 4 0893-8512/06/$08.00 0 doi:10.1128/cmr.00062-05 Copyright 2006, American Society for Microbiology. All Rights Reserved. Updated Review
Clinical Utility of the Time-to-Positivity/ Procalcitonin Ratio to Predict Bloodstream Infection Due to Coagulase-Negative Staphylococci
 Clinical Utility of the Time-to-Positivity/ Procalcitonin Ratio to Predict Bloodstream Infection Due to Coagulase-Negative Staphylococci Binghuai Lu, MS, 1* Lina Shi, BS, 1 Fengxia Zhu, BS, 1 Hua Zhao,
Clinical Utility of the Time-to-Positivity/ Procalcitonin Ratio to Predict Bloodstream Infection Due to Coagulase-Negative Staphylococci Binghuai Lu, MS, 1* Lina Shi, BS, 1 Fengxia Zhu, BS, 1 Hua Zhao,
Overview of Mycobacterial Culture, Identification, and Drug Susceptibility Testing
 Overview of Mycobacterial Culture, Identification, and Drug Susceptibility Testing 1. Essentials for the Mycobacteriology Laboratory: Promoting Quality Practices 1.1 Overview: Mycobacterial Culture, Identification,
Overview of Mycobacterial Culture, Identification, and Drug Susceptibility Testing 1. Essentials for the Mycobacteriology Laboratory: Promoting Quality Practices 1.1 Overview: Mycobacterial Culture, Identification,
Mixed gram positive organisms uti
 Mixed gram positive organisms uti The Borg System is 100 % Mixed gram positive organisms uti Complicated UTIs are caused by a broader spectrum of bacteria, including Gram- positive in addition to Gramnegative
Mixed gram positive organisms uti The Borg System is 100 % Mixed gram positive organisms uti Complicated UTIs are caused by a broader spectrum of bacteria, including Gram- positive in addition to Gramnegative
Significance of coagulase negative Staphylococcus from blood cultures: persisting problems and partial progress in resource constrained settings
 Volume 8 Number 6 (December 2016) 366-371 ORIGINAL ARTICLE Significance of coagulase negative Staphylococcus from blood cultures: persisting problems and partial progress in resource constrained settings
Volume 8 Number 6 (December 2016) 366-371 ORIGINAL ARTICLE Significance of coagulase negative Staphylococcus from blood cultures: persisting problems and partial progress in resource constrained settings
Objectives 12/4/2013. Disclosure. Culture of Orthopaedic Infections. Microbiology Testing in the Diagnosis of Prosthetic Joint Infections
 Culture of Orthopaedic Infections Microbiology Testing in the Diagnosis of Prosthetic Joint Infections December 9, 2013 Raymond P. Podzorski, Ph.D., D(ABMM) Clinical Microbiologist ProHealth Care Laboratories
Culture of Orthopaedic Infections Microbiology Testing in the Diagnosis of Prosthetic Joint Infections December 9, 2013 Raymond P. Podzorski, Ph.D., D(ABMM) Clinical Microbiologist ProHealth Care Laboratories
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES
 OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
Diagnosis of Pneumococcal Disease
 Diagnosis of Pneumococcal Disease Limitations of Surveillance for Invasive Disease David Murdoch University of Otago, Christchurch New Zealand Key Points We are still reliant on culture-based methods for
Diagnosis of Pneumococcal Disease Limitations of Surveillance for Invasive Disease David Murdoch University of Otago, Christchurch New Zealand Key Points We are still reliant on culture-based methods for
Detection of Bacteriuria and Pyuria by URISCREEN, a Rapid Enzymatic Screening Test
 JOURNAL OF CLINICAL MICROBIOLOGY, Mar. 1992, p. 680-684 0095-1137/92/03680-05$02.00/0 Copyright 1992, American Society for Microbiology Vol. 30, No. 3 Detection of Bacteriuria and Pyuria by URISCREEN,
JOURNAL OF CLINICAL MICROBIOLOGY, Mar. 1992, p. 680-684 0095-1137/92/03680-05$02.00/0 Copyright 1992, American Society for Microbiology Vol. 30, No. 3 Detection of Bacteriuria and Pyuria by URISCREEN,
A Two-Year Analysis of Risk Factors and Outcome in Patients with Bloodstream Infection
 Jpn. J. Infect. Dis., 56, 1-7, 2003 Original Article A Two-Year Analysis of Risk Factors and Outcome in Patients with Bloodstream Infection Andrea Endimiani, Antonio Tamborini, Francesco Luzzaro, Gianluigi
Jpn. J. Infect. Dis., 56, 1-7, 2003 Original Article A Two-Year Analysis of Risk Factors and Outcome in Patients with Bloodstream Infection Andrea Endimiani, Antonio Tamborini, Francesco Luzzaro, Gianluigi
Virginia Commonwealth Medical Center, Medical College of Virginia Campus
 JCM Accepts, published online ahead of print on 5 June 2013 J. Clin. Microbiol. doi:10.1128/jcm.01354-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11
JCM Accepts, published online ahead of print on 5 June 2013 J. Clin. Microbiol. doi:10.1128/jcm.01354-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11
Standard Operating Procedure
 Standard Operating Procedure 1. Definitions Section: Laboratory Version: FINAL Initials: AD Title: 2.0 Testing Algorithm Revision Date: 20 Oct 2011 1.1 SOP: Standard Operating Procedure 1.2 CRF: Case Report
Standard Operating Procedure 1. Definitions Section: Laboratory Version: FINAL Initials: AD Title: 2.0 Testing Algorithm Revision Date: 20 Oct 2011 1.1 SOP: Standard Operating Procedure 1.2 CRF: Case Report
The MALDI Biotyper An In Vitro Diagnostic System (IVD) for Identification of Bacteria and Yeasts with a Global Reach
 The MALDI Biotyper An In Vitro Diagnostic (IVD) for Identification of Bacteria and Yeasts with a Global Reach The MALDI Biotyper identifies microorganisms using MALDI-TOF (Matrix Assisted Laser Desorption
The MALDI Biotyper An In Vitro Diagnostic (IVD) for Identification of Bacteria and Yeasts with a Global Reach The MALDI Biotyper identifies microorganisms using MALDI-TOF (Matrix Assisted Laser Desorption
Aerobic bacteria isolated from diabetic septic wounds
 Aerobic bacteria isolated from diabetic septic wounds Eithar Mohammed Mahgoub*, Mohammed Elfatih A. Omer Faculty of Pharmacy, Omdurman Islamic University Department of Pharmaceutical Microbiology, Omdurman
Aerobic bacteria isolated from diabetic septic wounds Eithar Mohammed Mahgoub*, Mohammed Elfatih A. Omer Faculty of Pharmacy, Omdurman Islamic University Department of Pharmaceutical Microbiology, Omdurman
Evaluation of the Oxoid Xpect Legionella test kit for Detection of Legionella
 JCM Accepts, published online ahead of print on 6 May 2009 J. Clin. Microbiol. doi:10.1128/jcm.00397-09 Copyright 2009, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
JCM Accepts, published online ahead of print on 6 May 2009 J. Clin. Microbiol. doi:10.1128/jcm.00397-09 Copyright 2009, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
INVESTIGATING: WOUND INFECTION
 INVESTIGATING: WOUND INFECTION Diagnosing infection in surgical and other wounds involves nurses being able to observe the clinical signs in a wound rather than simply obtaining positive microbiology results
INVESTIGATING: WOUND INFECTION Diagnosing infection in surgical and other wounds involves nurses being able to observe the clinical signs in a wound rather than simply obtaining positive microbiology results
Received 30 March 2005; returned 16 June 2005; revised 8 September 2005; accepted 12 September 2005
 Journal of Antimicrobial Chemotherapy (2005) 56, 1047 1052 doi:10.1093/jac/dki362 Advance Access publication 20 October 2005 Evaluation of PPI-0903M (T91825), a novel cephalosporin: bactericidal activity,
Journal of Antimicrobial Chemotherapy (2005) 56, 1047 1052 doi:10.1093/jac/dki362 Advance Access publication 20 October 2005 Evaluation of PPI-0903M (T91825), a novel cephalosporin: bactericidal activity,
Labquality External Quality Assesment Programmes General Bacteriology 1 1/2010
 Labquality External Quality Assesment Programmes General Bacteriology 1 1/2010 Photos and text: Markku Koskela, M.D., Ph.D. Clinical microbiology specialist Oulu, Finland Sample 1/2010 Pus from an infected
Labquality External Quality Assesment Programmes General Bacteriology 1 1/2010 Photos and text: Markku Koskela, M.D., Ph.D. Clinical microbiology specialist Oulu, Finland Sample 1/2010 Pus from an infected
Direct identification of bacteria from positive anaerobic BacT/Alert blood cultures by. Contents Category: Diagnostics, typing and identification
 1 2 3 Direct identification of bacteria from positive anaerobic BacT/Alert blood cultures by MALDI-TOF MS: MALDI Sepsityper kit (Bruker) versus in-house saponin method for bacterial extraction 4 5 Running
1 2 3 Direct identification of bacteria from positive anaerobic BacT/Alert blood cultures by MALDI-TOF MS: MALDI Sepsityper kit (Bruker) versus in-house saponin method for bacterial extraction 4 5 Running
320 MBIO Microbial Diagnosis. Aljawharah F. Alabbad Noorah A. Alkubaisi 2017
 320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Pathogens of the Urinary tract The urinary system is composed of organs that regulate the chemical composition and volume of
320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Pathogens of the Urinary tract The urinary system is composed of organs that regulate the chemical composition and volume of
Bacterial Identification
 JOURNAL OF CLINICAL MICROBIOLOGY, July 1994, p. 1757-1762 0095-1137/94/$04.00+0 Copyright C 1994, American Society for Microbiology Vol. 32, No. 7 Clinical Impact of Rapid In Vitro Susceptibility Testing
JOURNAL OF CLINICAL MICROBIOLOGY, July 1994, p. 1757-1762 0095-1137/94/$04.00+0 Copyright C 1994, American Society for Microbiology Vol. 32, No. 7 Clinical Impact of Rapid In Vitro Susceptibility Testing
SURVEILLANCE BLOODSTREAM INFECTIONS IN BELGIAN HOPITALS ( SEP ) RESULTS ANNUAL REPORT data
 SURVEILLANCE BLOODSTREAM INFECTIONS IN BELGIAN HOPITALS ( SEP ) RESULTS ANNUAL REPORT data 2000-2014 SEP Workgroup Meeting 24 June 2015 Dr. Naïma Hammami Dr. Marie-Laurence Lambert naima.hammami@wiv-isp.be
SURVEILLANCE BLOODSTREAM INFECTIONS IN BELGIAN HOPITALS ( SEP ) RESULTS ANNUAL REPORT data 2000-2014 SEP Workgroup Meeting 24 June 2015 Dr. Naïma Hammami Dr. Marie-Laurence Lambert naima.hammami@wiv-isp.be
Clinical Trial Data Supporting FDA Clearance of the T2Bacteria Panel Versus Blood Culture for the Diagnosis of Bacteremia
 Clinical Trial Data Supporting FDA Clearance of the T2Bacteria Panel Versus Blood Culture for the Diagnosis of Bacteremia New diagnostic test overcomes time, sensitivity, and antimicrobial interference
Clinical Trial Data Supporting FDA Clearance of the T2Bacteria Panel Versus Blood Culture for the Diagnosis of Bacteremia New diagnostic test overcomes time, sensitivity, and antimicrobial interference
Mt. San Antonio College Microbiology 22 Lab Schedule for Fall 2017 Tues/Thurs. Split Lab Sections ONLY
 Mt. San Antonio College Microbiology 22 Lab Schedule for Fall 2017 Tues/ Split Lab Sections ONLY Wk 1 Aug. 29 Orientation with Introductions & Safety Rules/Regulations Aug. 31 Orientation with Pathogen
Mt. San Antonio College Microbiology 22 Lab Schedule for Fall 2017 Tues/ Split Lab Sections ONLY Wk 1 Aug. 29 Orientation with Introductions & Safety Rules/Regulations Aug. 31 Orientation with Pathogen
Routine endotracheal cultures for the prediction of sepsis in ventilated babies
 Archives of Disease in Childhood, 1989, 64, 34-38 Routine endotracheal cultures for the prediction of sepsis in ventilated babies T A SLAGLE, E M BIFANO, J W WOLF, AND S J GROSS Department of Pediatrics,
Archives of Disease in Childhood, 1989, 64, 34-38 Routine endotracheal cultures for the prediction of sepsis in ventilated babies T A SLAGLE, E M BIFANO, J W WOLF, AND S J GROSS Department of Pediatrics,
Unit II Problem 2 Microbiology Lab: Pneumonia
 Unit II Problem 2 Microbiology Lab: Pneumonia - What are the steps needed to obtain a proper sputum specimen? You need the following: A wide-mouth labeled container. Gloves. Water. Mouth wash + tissues.
Unit II Problem 2 Microbiology Lab: Pneumonia - What are the steps needed to obtain a proper sputum specimen? You need the following: A wide-mouth labeled container. Gloves. Water. Mouth wash + tissues.
Normal Human Flora. (Human Microbiome) Dr.Sarmad M.H. Zeiny Baghdad College of Medicine
 Normal Human Flora (Human Microbiome) Dr.Sarmad M.H. Zeiny Baghdad College of Medicine 2014-2015 Objectives Describe important human normal flora. Demonstrate the epidemiology of human normal flora. Determine
Normal Human Flora (Human Microbiome) Dr.Sarmad M.H. Zeiny Baghdad College of Medicine 2014-2015 Objectives Describe important human normal flora. Demonstrate the epidemiology of human normal flora. Determine
Blood cultures in ED. Dr Sebastian Chang MBBS FACEM
 Blood cultures in ED Dr Sebastian Chang MBBS FACEM Why do we care about blood cultures? blood cultures are the most direct method for detecting bacteraemia in patients a positive blood culture: 1. can
Blood cultures in ED Dr Sebastian Chang MBBS FACEM Why do we care about blood cultures? blood cultures are the most direct method for detecting bacteraemia in patients a positive blood culture: 1. can
Steven D. Brown* and Maria M. Traczewski. The Clinical Microbiology Institute, 9725 SW Commerce Circle, Wilsonville, Oregon 97070
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 2010, p. 2063 2069 Vol. 54, No. 5 0066-4804/10/$12.00 doi:10.1128/aac.01569-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Comparative
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 2010, p. 2063 2069 Vol. 54, No. 5 0066-4804/10/$12.00 doi:10.1128/aac.01569-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Comparative
Enzyme Immunoassay versus Latex Agglutination Cryptococcal Antigen Assays in Adults With non-hiv-related Cryptococcosis
 JCM Accepts, published online ahead of print on 24 September 2014 J. Clin. Microbiol. doi:10.1128/jcm.02017-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9
JCM Accepts, published online ahead of print on 24 September 2014 J. Clin. Microbiol. doi:10.1128/jcm.02017-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9
Mongolia September 2012
 MVZ DORTMUND - Dr.Eberhard u. Partner - MICROBIOLOGY MALDI-TOF (MS) evaluation for routine diagnostics MIKROBIOLOGY www.labmed.de / mikro@labmed.de Mongolia September 2012 accreditation since april 2003
MVZ DORTMUND - Dr.Eberhard u. Partner - MICROBIOLOGY MALDI-TOF (MS) evaluation for routine diagnostics MIKROBIOLOGY www.labmed.de / mikro@labmed.de Mongolia September 2012 accreditation since april 2003
A Multicentre Study about Pattern and Organisms Isolated in Follow-up Blood Cultures
 Ann Clin Microbiol Vol., No., March, 0 http://dx.doi.org/0./acm.0... ISSN -0 A Multicentre Study about Pattern and Organisms Isolated in Follow-up Blood Cultures Jeong Hwan Shin, Eui Chong Kim, Sunjoo
Ann Clin Microbiol Vol., No., March, 0 http://dx.doi.org/0./acm.0... ISSN -0 A Multicentre Study about Pattern and Organisms Isolated in Follow-up Blood Cultures Jeong Hwan Shin, Eui Chong Kim, Sunjoo
Evaluation of a Two-Minute Test for Urine Screening
 JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1983. p. 697-701 0095-1137/83/090697-05$02.00/0 Copyright 1983, American Society for Microbiology Vol. 18, No. 3 Evaluation of a Two-Minute Test for Urine Screening
JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1983. p. 697-701 0095-1137/83/090697-05$02.00/0 Copyright 1983, American Society for Microbiology Vol. 18, No. 3 Evaluation of a Two-Minute Test for Urine Screening
Impact of Antimicrobial Stewardship Intervention on Coagulase-Negative Staphylococcus Blood. Cultures in Conjunction with Rapid Diagnostic Testing
 JCM Accepts, published online ahead of print on 28 May 2014 J. Clin. Microbiol. doi:10.1128/jcm.00682-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 Impact of Antimicrobial
JCM Accepts, published online ahead of print on 28 May 2014 J. Clin. Microbiol. doi:10.1128/jcm.00682-14 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 Impact of Antimicrobial
Corporate Medical Policy
 Corporate Medical Policy Identification of Microorganisms Using Nucleic Acid Probes File Name: Origination: Last CAP Review: Next CAP Review: Last Review: identification_of_microorganisms_using_nucleic_acid_probes
Corporate Medical Policy Identification of Microorganisms Using Nucleic Acid Probes File Name: Origination: Last CAP Review: Next CAP Review: Last Review: identification_of_microorganisms_using_nucleic_acid_probes
Poor Predictive Ability of Urinalysis and Microscopic Examination to Detect Urinary Tract Infection
 Microbiology and Infectious Disease / POOR PREDICTIVE ABILITY OF URINALYSIS Poor Predictive Ability of Urinalysis and Microscopic Examination to Detect Urinary Tract Infection Joy D. Van Nostrand, MS,
Microbiology and Infectious Disease / POOR PREDICTIVE ABILITY OF URINALYSIS Poor Predictive Ability of Urinalysis and Microscopic Examination to Detect Urinary Tract Infection Joy D. Van Nostrand, MS,
CLINICAL MICROBIOLOGY AND INFECTIOUS DISEASE
 CLINICAL MICROBIOLOGY AND INFECTIOUS DISEASE Original Article S o n i c a t e d V a s c u l a r C a t h e t e r T i p C u l t u r e s Quantitative Association With Catheter-related Sepsis and the Non-Utility
CLINICAL MICROBIOLOGY AND INFECTIOUS DISEASE Original Article S o n i c a t e d V a s c u l a r C a t h e t e r T i p C u l t u r e s Quantitative Association With Catheter-related Sepsis and the Non-Utility
ENG MYCO WELL D- ONE REV. 1.UN 29/09/2016 REF. MS01283 REF. MS01321 (COMPLETE KIT)
 ENG MYCO WELL D- ONE MYCO WELL D-ONE System for the presumptive identification and antimicrobial susceptibility test of urogenital mycoplasmas, Gardnerella vaginalis, Trichomonas vaginalis, Candida albicans
ENG MYCO WELL D- ONE MYCO WELL D-ONE System for the presumptive identification and antimicrobial susceptibility test of urogenital mycoplasmas, Gardnerella vaginalis, Trichomonas vaginalis, Candida albicans
Antimicrobial prophylaxis in liver transplant A multicenter survey endorsed by the European Liver and Intestine Transplant Association
 Antimicrobial prophylaxis in liver transplant A multicenter survey endorsed by the European Liver and Intestine Transplant Association Els Vandecasteele, Jan De Waele, Dominique Vandijck, Stijn Blot, Dirk
Antimicrobial prophylaxis in liver transplant A multicenter survey endorsed by the European Liver and Intestine Transplant Association Els Vandecasteele, Jan De Waele, Dominique Vandijck, Stijn Blot, Dirk
Mt. San Antonio College Microbiology 22 Lab Schedule for Spring 2018 Tues/Thurs. Split Lab Sections ONLY
 Mt. San Antonio College Microbiology 22 Lab Schedule for Spring 2018 Tues/ Split Lab Sections ONLY Wk 1 Feb. 27 Orientation with Introductions & Safety Rules/Regulations March 1 Orientation with Pathogen
Mt. San Antonio College Microbiology 22 Lab Schedule for Spring 2018 Tues/ Split Lab Sections ONLY Wk 1 Feb. 27 Orientation with Introductions & Safety Rules/Regulations March 1 Orientation with Pathogen
Mt. San Antonio College Microbiology 22 Lab Schedule for Spring 2018 Mon/Weds. Split Lab Sections ONLY
 Mt. San Antonio College Microbiology 22 Lab Schedule for Spring 2018 Mon/ Split Lab Sections ONLY Wk 1 Feb. 26 Orientation with Introductions & Safety Rules/Regulations Feb. 28 Orientation with Pathogen
Mt. San Antonio College Microbiology 22 Lab Schedule for Spring 2018 Mon/ Split Lab Sections ONLY Wk 1 Feb. 26 Orientation with Introductions & Safety Rules/Regulations Feb. 28 Orientation with Pathogen
Vimta Labs Ltd., Life Sciences Facility, Plot No. 5, Alexandria Knowledge Park, Genome Valley, Shameerpet, Hyderabad, Telangana
 Last Amended on - Page 1 of 14 I. DRUGS & PHARMACEUTICALS 1. Biological Assays Antibiotics And Other Drugs Bulk Drugs & Their Formulations: Erythromycin, Gentamicin, Nystatin IP Appendix 9.1 2.2.10 BP
Last Amended on - Page 1 of 14 I. DRUGS & PHARMACEUTICALS 1. Biological Assays Antibiotics And Other Drugs Bulk Drugs & Their Formulations: Erythromycin, Gentamicin, Nystatin IP Appendix 9.1 2.2.10 BP
IP Lab Webinar 8/23/2012
 2 What Infection Preventionists need to know about the Laboratory Anne Maher, MS, M(ASCP), CIC Richard VanEnk PhD, CIC 1 Objectives Describe what the laboratory can do for you; common laboratory tests
2 What Infection Preventionists need to know about the Laboratory Anne Maher, MS, M(ASCP), CIC Richard VanEnk PhD, CIC 1 Objectives Describe what the laboratory can do for you; common laboratory tests
Molecular approaches in the diagnosis of sepsis in neutropenic patients with haematological malignances
 J prev med hyg 2012; 53: 104-108 Short article Molecular approaches in the diagnosis of sepsis in neutropenic patients with haematological malignances M. Guido 1, M. Quattrocchi 1, A. Zizza 2, G. Pasanisi
J prev med hyg 2012; 53: 104-108 Short article Molecular approaches in the diagnosis of sepsis in neutropenic patients with haematological malignances M. Guido 1, M. Quattrocchi 1, A. Zizza 2, G. Pasanisi
SYMPOSIUM 10TH MAY 2007 BELGIUM. Blood culture-negative endocarditis
 SYMPOSIUM 10TH MAY 2007 BELGIUM Blood culture-negative endocarditis Didier RAOULT Didier.raoult@gmail.com Modified Duke criteria for diagnosis of infective endocarditis (IE) Li JS, et al. Proposed modifications
SYMPOSIUM 10TH MAY 2007 BELGIUM Blood culture-negative endocarditis Didier RAOULT Didier.raoult@gmail.com Modified Duke criteria for diagnosis of infective endocarditis (IE) Li JS, et al. Proposed modifications
