Short communication Natural polymorphism of the HIV-1 integrase gene and mutations associated with integrase inhibitor resistance
|
|
|
- Francine Lambert
- 5 years ago
- Views:
Transcription
1 Short communication Natural polymorphism of the HIV-1 integrase gene and mutations associated with integrase inhibitor resistance Max Lataillade 1 *, Jennifer Chiarella 1 and Michael J Kozal 1,2 1 Yale University School of Medicine, New Haven, CT, USA 2 VA Connecticut Healthcare System, New Haven, CT, USA Antiviral Therapy 12: *Corresponding author: Tel: ; Fax ; Max.Lataillade@yale.edu Background: Two inhibitors of the HIV-1 integrase enzyme (INIs) are in late stage clinical development. To date, approximately 42 mutations within the HIV-1 integrase (IN) gene have been associated with INI drug resistance. Naturally occurring IN gene polymorphisms may have important implications for INI development. In this study, we evaluated clinical HIV-1 strains from INI-naive patients to determine the prevalence of IN gene polymorphisms, and the frequency of naturally occurring amino acid (aa) substitutions at positions associated with INI resistance and at sites crucial for LEDGF/p75 binding and HIV-1 integration. Methods: The IN gene from 67 INI-naive, HIV-1 clade B-infected patients were sequenced using standard population-based DNA sequencing methods. In addition, 176 unique full-length HIV-1 clade B IN gene sequences from INI-naive patients obtained from the HIV Los Alamos database were analysed. Results: Analysis of 243 IN genes from HIV-1 clade B, INI-naive clinical strains revealed that 64% of the aa positions were polymorphic. Of the 42 aa substitutions currently associated with INI resistance, 21 occurred as natural polymorphisms: V72I, L74I, T97A, T112I, A128T, E138K, Q148H, V151I, S153Y/A, M154I, N155H, K156N, E157Q, G163R, V165I, V201I, I203M, T206S, S230N and R263K. IN aa positions crucial to LEDGF/P75 binding and HIV-1 integration were well conserved. Conclusion: Major INI mutations within the catalytic domain and extended active sites associated with high level resistance to the compounds in late stage development, especially strand transfer inhibitors (STIs), were infrequent in our study, which may help explain the excellent virological responses demonstrated in clinical trials. Introduction Inhibitors of the HIV-1 integrase enzyme (INIs) block the integration of viral double-stranded DNA into the host cell s chromosomal DNA [1]. The integrase (IN) enzyme is essential and unique to HIV-1 without any similar human analogue, a characteristic that has so far led to low side effects and toxicity in INI clinical trials [2]. HIV-1 IN is a 288 amino acid (aa) long polypeptide chain that folds into three distinct functional domains: the N-terminal domain (NTD), the central or catalytic core domain (CCD) and the C-terminal domain (CTD). The NTD (aa 1 50) is believed to be involved in protein multimerization and contains a histidine histidine cystine cystine (HHCC) motif that coordinates zinc binding [1,3]. The CCD (aa ) contains the catalytic trio of acidic amino acids, aspartic acid (Asp-D) 64, Asp (D) 116 and glutamic acid (E) 152, the DDE motif [1,3]. The DDE motif forms an active site that coordinates a divalent metal ion, either magnesium or manganese, which is required for catalytic activity [3 5]. The CCD of IN binds to human lens epithelium-derived growth factor (LEDGF/p75), which is an essential HIV integration cofactor linking IN to chromatin [6 7]. The CTD of IN (aa ) has non-specific but strong DNA binding activity, similar to that of the full-length IN, and is thought to play a role in binding to viral and host DNA [1,3]. HIV-1 retroviral integration can be described as a multistep process: (1) assembly of a stable complex between IN and specific viral DNA sequences at the end of the HIV-1 long terminal repeats, (2) 3 processing, (3) preintegration complex translocation, (4) strand transfer, and (5) DNA gap repair and ligation [2,4]. All 2007 International Medical Press
2 M Lataillade et al. Table 1. Class and status of integrase inhibitors Integrase inhibitor Integrase inhibitor class Status Reference L-708,906 Diketo-Acid STI Preclinical [1,2,17,18,32,33,35] L-731,988 Diketo-Acid STI Preclinical [1,2,17,18,32,33,35] S-1360 Diketo-acid STI Previous Phase II: [14,33] withdrawn L870,810 Naphthyridine carboxamide Previous Phase II: [1,219,20,34,35] STI withdrawn MK-0518 (Raltegravir) Pyrimidone STI Phase III [2,8,9,21,34,35] MK-2048 Second Generation STI Preclinical [35] GS-9137 (Elvitegravir) Dihydroquinoline carboxylic Phase II [2,10,20,34,35] Acid STI Enrolling in Phase III L-CA L-Chicoric Acid STI Early compound [26] 5-CITEP DKA derivative STI Early compound [27] V-165 PyranoDipyriminidine Preclinical [1,2,15,16,32] integrase DNA Binding inhibitor FZ41 Styrylquinolines Preclinical [1,2,22,23] 3 Processing Inhibitor integration steps can potentially be inhibited and each step can be considered a possible drug target. Multiple INIs are currently in different phases of development and can be divided into five classes: (1) integrase DNA binding inhibitors, (2) 3 processing inhibitors, (3) nuclear translocation/ import inhibitors, (4) strand transfer inhibitors (STIs) and (5) gap repair inhibitors [1 2]. To date, the STIs have been the most successful class of INIs and presently are the only class of INIs in advanced human clinical trials (see Table 1) [8 10]. As with other antiretroviral classes, drug resistance has been shown to occur with the INIs. Each class of INIs has a distinct mechanism of action and carry their respective resistance profile and mutations. A summary of IN mutations associated with specific compounds, the decrease susceptibility of the IN mutants and INI cross resistance patterns can be found in Table 2 [11] Knowing the prevalence rate of mutations in viral strains from INI treatment-naive and -experienced patients can help investigators identify if clinically relevant viral mutations or natural polymorphisms are occurring under drug selection pressure. In addition, other IN aa positions have been identified as critical for LEDGF/p75/DNA binding and integration, yet little is known about the natural polymorphism of these IN sites [12 13]. In this study we determined the natural polymorphism of the HIV-1 clade B IN gene. Further, we identified the natural occurrence of IN aa substitutions at positions associated with INI resistance, IN-DNA binding sites and at IN-LEDGF/P75 binding sites that are crucial for integration. Methods The HIV-1 clade B IN genes from 67 INI naive US patients were sequenced. HIV-1-infected plasma samples underwent HIV RNA extraction using QIAamp RNA mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer s instructions. An IN gene amplification and sequencing protocol was kindly provided to us by Michael Miller and Daria Hazuda of Merck Research Labs. HIV RNA underwent RT-PCR using a Superscript one-step kit (Invitrogen, Carlsbad, CA, USA); the integrase gene was amplified by nested RT-PCR with the following primers for the first round: 1) INFORI (5 -GGAATCATTCAAGCACAACCAGA- 3 ; nucleotide (NT) positions relative to HXB2 4,050 4,081) and INREV-I (5 -TCTCCTGTATGCA- GACCCCAATAT-3 ; NT: 5,244 5,267). PCR cycle parameters are: one cycle of 48 C for 30 min and 94 C for 5 min; 35 cycles of 94 C for 30 sec, 60 C for 1 min and 72 C for 2 min; and one cycle of: 72 C for 7 min. For the second round two additional primers were used: HIV+4141 (5 -TCTACCTGGCATGGGTA CCA- 3 ; NT: 4,141 4,160) and INREVII (5 -CCTAGTGGG ATGTGTACTTCTGA-3 ; NT: 5,197 5,219). PCR cycle parameters are: one cycle of 94 C for 5 min; 35 cycles of 94 C for 30 sec, 60 C for 1 min and 72 C for 2 min; and one cycle of 72 C for 7 min. Population based sequencing (ABI Technology TM, Foster City, CA, USA) was performed on the samples. All sequences (forward and reverse strands) were aligned and manually read using BioEdit software (Ibis Therapeutics, Carlsbad, CA, USA; GenBank accession EF EF472787) International Medical Press
3 Integrase inhibitors and natural polymorphisms Table 2. Integrase mutations associated with integrase inhibitor (INI) resistance and cross resistance (CR) patterns between different classes of INIs Mutations, single Decrease susceptibility of INIs versus multiple HIV-1 mutants to INIs Evidence of CR between INIs L-708,906* T66I, S153Y, M154I M154I: 2 5-fold CR to S-1360: fold L74M, S230R, N155S S153Y: 2 3-fold T66I/L74M/S230R: 26-fold T66I/M154I: 20-fold CR to GS-9137: fold T66I/L74M/S230R: 10 fold S153Y+T66I: 259-fold; N155S: 116-fold CR to MK-0518 (N155S):~12-fold CR to L870,810: (N155S): ~76-fold L-731,988 T66I, S153Y, M154I M154I: 2 5-fold; S153Y: no effect CR to L 708,906; CR to S-1360 T66I / M154I: 20-fold CR to L870,810: 22-fold; T66I / S153Y: 22-fold S-1360 T66I, L74M, Q146K T66I: 4-fold CR to L-708,906; Moderate to high S153A, K160D, V165I T66I/Q146K/V201I: 8-fold V201I, Q148K, A128T, T66I/Q146K/V201I/L74M/A128T/S153A/ E138K, N155S K160D/V165I >62-fold L870,810 F121Y, T125K, V72I, F121Y/T125K (6 months): 16-fold CR to GS-9137: 1,000-fold V151I, T97A, K156N V72I/F121Y/T125K (9 months): 20-fold F121Y/T125K/V151I/V72I: 1,000-fold V72I/F121Y/T125K/V151I :100-fold CR from GS-9137: fold E92Q + S147G: 101-fold CR to MK-0518: (F121Y): 3-fold MK-0518 (Raltegravir) E138K/A, G140S/A, Q148K/G140A/E138A: 388-fold # CR from GS-9137: fold Q148H/K/R, N155H, E92Q/N155H: 64-fold # E92Q: 3.5-fold E92Q, V151I, T97A, G140S/Q148H: 399-fold # CR from L : 3-fold G163R, L74I/M/A, G140S/Q148R: 239-fold # F121Y: 3-fold Y143R/C, I203M, S230R Q148R: 26-fold; Q148K: 40-fold # CR to GS-9137: ~200-2,000-fold N155H +(E92Q,V151I,T97A, Q148K/G140A/E138A: 395-fold G163R,L74M): IP G140S/Q148H: 2,194-fold Y143R/C +(L74I/A,E92Q,T97A, G140S/Q148R: 268-fold I203M, S230R): IP Q148K: 169-fold E92Q/N155H: 207-fold MK-2048 # T112I; resistance Resistance testing in progress CR testing in progress testing in progress GS-9137 (Elvitegravir)** H51Y, E92Q, S147G, E92Q / S147G: 356-fold CR to L-870,810 ( fold) E157Q, T66I, R263K H51Y/ E92Q/ S147G: 699-fold E92Q + S147G: 101 fold H51Y/E92Q/S147G/E157Q: 1,000-fold CR to MK-0518 (E92Q): fold T66I/R263K: 98-fold CR from L870,810 (1,000-fold) T66I (15.1 fold), R263K (5.1-fold) F121Y/T125K/V72I/V151I: 1,000-fold E157 (6.3 fold), E92Q (36-fold) CR from MK-0518: ~200-2,000-fold H51Y/E92Q/S147G/E157Q: 182-fold Q148K/G140A/E138A: 395-fold Q148H/G140S: 2,194-fold L-CA G140S G140S: 100-fold CR to DKAs 5-CITEP T66I/M154I Most likely similar effect as DKAs CR to DKAs V-165 ## V165I, T206S, S230N V165I: 15-fold No CR to DKAs; No CR from DKAs FZ41*** C280Y, V165I/V249I C280Y: 2 5 fold; V165I/V249I: 9-fold No CR to DKAs; No CR from DKAs In red: natural polymorphisms associated with INI resistance. CR, cross resistance; IP, testing in progress; STI, strand transfer inhibitor. *See [1,2,17,18,32,33,35]; See [1,2,17,18,32,33,35]; See [14,33]; See [1,219,20,34,35]; See [2,8,9,21,34,35]; # See [35]; **See [2,10,20,34,35]; see [20]; see [34]; See [26]; See [27]; ## See [1,2,15,16,32]; ***See [1,2,22,23]. In addition, 176 worldwide HIV-1 clade B (includes B recombinants) treatment-naive unique sequences were obtained from the Los Alamos HIV Database ( accessed 4 April 2006). The combined set of sequences (67 USA Clade B and 176 worldwide clade B samples; n=243) were compared and analysed in BioEdit. A new IN nucleotide and aa consensus was determined using the 243 samples and compared with the 2004 HIV-1 clade B subtype consensus. Antiviral Therapy 12:4 565
4 M Lataillade et al. Results A total of 243 HIV-1 clade B IN gene sequences from unique patient strains were analysed to determine the natural polymorphism of the clade B IN gene. These IN gene sequences came from patients from the USA (n=116), followed by samples from Australia, Argentina, China, South East Asia, Western Europe, Eastern Europe and Latin America. Analysis of the 243 IN clade B samples compared with the 2004 Los Alamos HIV-1 clade B consensus revealed that 64% (184/288) of the aa positions of IN were naturally polymorphic (Figure 1). IN aa residues within the DDE triad and the HHCC motif were relatively well conserved (Figure 1) [11]. Of the 42 aa substitutions currently associated with INI resistance (Table 2), 21 occurred as natural polymorphisms: V72I, L74I, T97A, T112I, A128T, E138K, Q148H, Figure 1. Natural polymorphism associated with the IN enzyme and mutations associated with INI resistance HHCC motif DEE motif aa position associated with INI resistance NP Natural polymorphism associated with INI resistance The consensus for the integrase IN gene for the 243 samples is represented on the top line. Natural polymorphisms are listed directly under the consensus amino acid (aa). Analysis of the 243 IN clade B samples compared to the 2004 Los Alamos HIV-1 clade B consensus revealed that 64% (184/288) of the aa positions of IN were naturally polymorphic. Codon 289 is a stop codon for IN and is not shown. Natural polymorphism associated with the N-terminal domain (NTD) of IN is represented by aa HHCC motif in purple is relatively well conserved with only two natural polymorphisms associated with the motif. Natural polymorphisms associated with IN inhibitor (INI) resistance were not found in the N-terminal domain of IN. Natural polymorphism associated with the catalytic core domain (CCD) of IN is represented by aa DDE motif in bold yellow is relatively well conserved. Of the 37 mutations associated with INI resistance in this domain (green), 19 occurred as natural polymorphisms (I72V, L74I, T97A, T112I, A128T, E138K, Q148H, V151I, S153Y, S153A, M154I, N155H, K156N, E157Q, G163R, V165I, V201I, I203M and T206S). The unique samples containing N155H and Q148H came from Australia (ref. AF042104; B. AU.96.MBCC 98) and Spain (ref. AF450096; BG.ES.00.X605) in the Los Alamos Database. Major aa substitutions located within the catalytic domain and the extended active sites, known to contribute to high level INI resistance (positions H51Y, T66I, L74M/A, E92Q, F121Y, T125K, E138A, G140S/A, Y143R/C, Q146K, S147G, Q148K/R, N155S and K160D) were not found to occur as natural polymorphisms. K156N associated with increased susceptibility to L870,810 and Raltegravir was present as natural polymorphism in ~9% (21/243) of the samples. I72V and V201I occurred concurrently in 20% (50/243) of sequences in our study. I72 was the consensus aa in this large sample set (51%) instead of previously described V72. Natural polymorphism associated with the C-terminal domain (CTD) of Integrase is represented by aa Of the five mutations associated with INI resistance in this domain (S230N, S230R, V249I, R263K, and C280Y in green), two mutations (S230N and R263K) occurred as natural polymorphism (in red). Approximately 42 mutations so far have been described to be associated with resistance to various INIs in preclinical and clinical phases of development. Twenty-one of these mutations were found as natural polymorphisms in our study (red). Other IN positions identified as critical for DNA binding (Q148), HIV-1 integration and replication (Q62, H67, N120, N144, Q148 and N155) were relatively well conserved within the IN CCD. Several point mutations within IN (C130G, W131A, I161A, V165A, R166A, Q168A/L/P, E170G, L172A, K173A, Q214L and Q216L) reported to be defective for interaction with LEDGF/p75, impairing chromosome tethering and HIV-1 replication were not found as natural polymorphisms in our study. In fact, the majority of these IN positions were highly conserved, lending support to the finding of the importance of these sites for LEDGF/p75 binding International Medical Press
5 Integrase inhibitors and natural polymorphisms V151I, S153Y, S153A, M154I, N155H, K156N, E157Q, G163R, V165I, V201I, I203M, T206S, S230N and R263K (Table 2; Figure 1). Table 3 describes the IN natural polymorphisms associated with resistance to the different INIs. I72 was the consensus aa in this large sample set (51%) instead of previously described V72 [11]. IN polymorphisms at I72V (n=113), V201I (n=107), T206S (n=30), S230N (n=24) and K156N (n=21) occurred at high frequency. I72V and V201I occurred concurrently in 20% (50/243) of sequences in our study. V165I was found to occur in only 1 of 114 USA samples. The majority of samples containing a V165I were from Latin America (58%; 7/12). The unique samples containing N155H and Q148H came from Australia (ref. AF042104; B. AU.96.MBCC 98) and Spain (ref. AF450096; BG.ES.00.X605) in the Los Alamos Database. However, a number of major aa substitutions located within the catalytic domain and the extended active sites, known to contribute to high level INI resistance (positions H51Y, T66I, L74M/A, E92Q, F121Y, T125K, E138A, G140S/A, Y143R/C, Q146K, S147G, Q148K/R, N155S, K160D, S230R, V249I and C280Y), were not found to occur as natural polymorphisms (Table 2) [8,9,14 27,32 35]. Discussion INI resistance arises from the selection of viral variants with genetic mutations that change the target enzyme, such as, creating conformational changes in the IN catalytic site or affecting metal ion coordination which may result in reduced affinity for a drug [28]. Early in the development of protease inhibitors (PIs), investigators reported the HIV-1 clade B protease to be extremely polymorphic with 47.5% of the 99 aa positions being variable [29]. Some naturally occurring protease polymorphisms were found to contribute to PI resistance [30,31] Similar to PI resistance, the number of IN mutations present can carry an additive effect on INI resistance [8, 32 35]. In our study many of the mutations associated with resistance to INIs were found to occur naturally in clinical viral strains (Table 2) [8,34 35]. The identification of naturally occurring IN polymorphisms associated with INI resistance suggests that resistance to some INIs may occur faster in patients that harbour these polymorphic variants, since they may augment other mutation effects [8,9,20,32 35]. Recently, researchers reported resistance pathways to STIs, MK-0518 (Raltegravir) and GS-9137 (Elvitegravir; Table 2) [8,9,34]. Raltegravir failure was Table 3. Natural polymorphisms associated with integrase inhibitor resistance Class Compound Natural polymorphism Reference Strand transfer inhibitors L-708,906 S153Y/1, M154I/11 [1,2,5,8 10,14,17 21, 25 27,32 35] L-731,988 S153Y/1, M154I/11 S-1360 S153A/1, V165I/12 V201I/107, A128T/1 E138K/1 L870,810 I72V/113, V151I/12 T97A/1, K156N/21 MK-0518 N155H/1, Q148H/1 (L-900,612) L74I/1 T97A/1, E138K/1 K156N/21 G163R/5, I203M/5 GS-9137 E157Q/5, R263K/1 (JTK-303) MA/DA DKAs S153Y/1 5-CITEP M154I/11 MK-2048 T112I/14 In DNA binding inhibitors PDP: V165 V165I/12, T206S/30, [1,2,15,16,32] S230N/24 3 Processing inhibitors SQ L: FZ41 V165I/12 [1,2,22,23] NTI/NII SQ L: FZ41 V165I/12 [23] Bifunctional DKAs Profile under investigation [1,2,24] Natural polymorphisms described as: mutation/n (number of times seen within 243 samples). NTI/NII, nuclear translocation/import inhibitors. Antiviral Therapy 12:4 567
6 M Lataillade et al. generally associated with one of two genetic pathways corresponding with one of two major mutations N155H or Q148K/H/R: N155H+(E92Q, V151I, T97A, G163R or L74M) or Q148K/R/H+(G140S/A or E138K/A); another pathway Y143R/C+(L74A/I, E92Q, T97A, I203M or S230R) may also occur [8,9]. L74I, T97A, E138K, G163R, I203M, N155H and Q148H were all identified as natural polymorphisms in our study. Furthermore, Merck Research Laboratories described MK-2048 as a potent IN second generation STI [35]. MK-2048 has the potential to inhibit HIV-1-resistant variants generated with first generation compounds (high genetic barrier) and showed good pharmacokinetic profiles in preclinical species [35]. Primary mutation T112I observed with this compound was identified as a natural polymorphism in our study (Table 2, Table 3) [35]. In vitro studies have identified two primary resistance patterns for GS-9137 involving T66I or E92Q with secondary IN mutations H51Y, F121Y, S147G, S153Y, E157Q or R263K [20,34]. E157Q and R263K were identified as natural polymorphisms in our study. In an important study by Hackett et al., the HIV-1 IN gene diversity in group M, N and O viruses was evaluated [36]. The natural polymorphism rate for the subset of 81 clade B samples was ~39% [36]. This rate for clade B reported by Hackett et al. is lower than that found in our sample (64%); however, this disparity may be due to the sample size differences of the two studies. Recently, Zioni et al. reported an IN polymorphism rate (56%) similar to our study [37]. Furthermore, both Hackett et al. and Zioni et al. also reported that several IN aa substitutions associated with INI resistance occurred as natural polymorphisms: V72I, L74M, T97A, V151I, E157Q, G163R, V165I, V201I, I203M, T206S and S230N, and I141V, S153Y/A, M154I, N155H and K156N, respectively [36,37]. L74M reported by Hackett et al. associated with resistance to early compounds L-708,906 and S-1360 was not found as a natural polymorphism in our 243 clade B samples; however, the other mutations were confirmed as natural polymorphisms (Table 2, Figure 1) [36]. Other IN positions have been identified as critical for DNA binding (Q148), and HIV-1 integration and replication (Q62, H67, N120, N144, Q148 and N155) [38]. Our data demonstrate that these important positions are relatively well conserved within the IN CCD (Figure 1) [38]. Furthermore, LEDGF/p75 has been described as an essential HIV integration cofactor linking IN to chromatin [6,7,39]. The LEDGF/p75 IN binding domain is reported to interact with the IN CCD between aa and [6,7,12,13,40]. Several point mutations within IN, A128T, C130G, W131A, I161A, V165A, R166A, Q168A/L/P, E170G, L172A, K173A, Q214L and Q216L are defective for interaction with LEDGF/p75, impairing chromosome tethering and HIV-1 replication [6,7,12,13,40]. Besides A128T, none of these mutations were found as natural polymorphisms in our study. In fact, the majority of these IN positions were highly conserved, lending support to the finding of the importance of these sites for LEDGF/p75 binding. Our study is limited in that the sample size was composed of only 243 sequences from unique INInaive patients. Furthermore, as INI development has focused on clade B strains, we limited our analysis to only HIV-1 clade B samples which prevents us from extrapolating our results to broader more diverse populations. In addition, some of the mutations were identified only in a single sequence from a public database. These single mutation occurrences may be the result of sequencing errors and this could inflate the natural polymorphism rate of IN. However, the IN diversity is similar to that of the protease gene and suggests that the underlying diversity of this portion of HIV-1 pol is similar to other components. Populationbased sequencing was used in our study, and thus only the dominant variant sequence is likely represented. Future studies using ultrasensitive quantitative sequencing techniques would determine if additional minor viral-resistant variants are present. Despite these limitations, we were able to determine a consensus sequence for the clade B IN gene based on 243 samples, demonstrate the natural polymorphism of IN and identify the natural occurrence of many IN aa substitutions associated with decreased susceptibility to INIs. The identification of naturally occurring IN aa substitutions associated with resistance suggests that these changes may not adversely affect viral fitness. INIs that are affected by these substitutions could be at a disadvantage leading to rapid resistance development. The data also demonstrates the conserved nature of the IN-LEDGF/P75 binding sites and of aa positions crucial to integration. The launch of INIs represents a new era in HIV treatment as it provides clinicians with a potent new drug class to target HIV. Resistance to INIs likely will emerge in the clinic; ongoing studies will be needed to determine the effects of these naturally occurring IN polymorphisms on clinical responses in patients. Acknowledgements This data was presented in part at the XV International Workshop on HIV Drug Resistance, Sitges, Spain, abstract 23, June ML was supported by the NIH training grant: T32 AI MJK was supported by the VA Career Development Award International Medical Press
7 Integrase inhibitors and natural polymorphisms We would like to thank Daria Hazuda, PhD. and Michael Miller, PhD. of Merck Research Labs for sharing integrase gene amplification and sequencing protocols. Financial Disclosure MJK has received research grants form the VA, NIH and is the principal investigator on Merck integrase inhibitor trials at Yale University. ML is an investigator on Merck integrase inhibitor trials at Yale University. Role of funding source The sponsors had no role in study design, data collection, data analysis, data interpretation, writing, review or approval of manuscript. The corresponding authors had full access to all the data in the study and had final responsibility for the decision to submit for publication. References 1. Pommier Y, Johnson AA, Marchand C. Integrase inhibitors to treat HIV/AIDS. Nat Rev Drug Discov 2005; 4: Lataillade M, Kozal MJ. The hunt for HIV-1 integrase inhibitors. AIDS Patient Care STDS 2006; 7: Van Maele B, Debyser Z. HIV-1 integration: an interplay between HIV-1 integrase, cellular and viral Proteins. AIDS Rev 2005; 7: Grobler JA, Stillmock K, Hu B, et al. Diketo acid inhibitor mechanism and HIV-1 integrase: implications for metal binding in the active site of phosphotransferase enzymes. Proc Natl Acad Sci U S A 2002; 99: Hazuda DJ. Inhibitors of human immunodeficiency virus type I. Retrovirology 2006; 3 Suppl 1:S7. 6. Debyser Z, Hombrouck A, Witvrouw M, De Rijck J, Hendrix J. Virus evolution reveals LEDGF/p75 as the sole mediator of chromosomal tethering. 14th Conference on Retroviral and Opportunistic Infections 2007, Los Angeles CA, USA. Feb 25 28; Abs Hombrouck A, De Rijck J, Busschots K, Christ F Debyser Z. Structure-function analysis of the interface between HIV-1 integrase and the cellular co-factor LEDGF/p75. 14th Conference on Retroviral and Opportunistic Infections 2007, Los Angeles CA, USA. Feb 25 28; Abs Cooper D, Gatell J, Rockstroh J, Katlama C, Isaacs R, Nguyen B. Results of BENCHMRK-1, a Phase III study evaluating the efficacy and safety of MK-0518, a novel HIV-1 integrase inhibitor, in patients with triple-class resistant virus. 14th Conference on Retroviral and Opportunistic Infections February 2007, Los Angeles CA, USA. Abs. 105aLB. 9. Steigbigel R, Kumar P, Eron J, et al. Results of BENCHMRK-2, a Phase III study evaluating the efficacy and safety of MK-0518, a novel HIV-1 integrase inhibitor, in patients with triple-class resistant virus. 14th Conference on Retroviral and Opportunistic Infections February 2007, Los Angeles CA, USA. Abs. 105bLB. 10. Zolopa AR, Mullen M, Berger D, Ruane P, Hawkins T. The HIV integrase inhibitor GS-9137 demonstrates potent antiretroviral activity in treatment-experienced patients. 14th Conference on Retroviral and Opportunistic Infections February 2007, Los Angeles CA, USA. Abs. 143LB. 11. M Lataillade, J Chiarella, MJ Kozal. Natural polymorphism of the HIV-1 integrase gene and mutations associated with integrase inhibitor resistance. Antivir Ther 2006; 11:S Llano M, Vanegas M, Hutchins N, Thompson D, Delgado S, Poeschla EM. Identification and characterization of the chromatin-binding domains of the HIV-1 integrase interactor LEDGF/p75. J Mol Biol 2006; 360: Cherepanov P, Ambrosio AL, Rahman S, Ellenberger T, Engelman A. Structural basis for the recognition between HIV-1 integrase and transcriptional coactivator p75. Proc Natl Acad Sci U S A. 2005; 102: Billich A. S-1360 Shinogi-GlaxoSmithKline. Curr Opin Investig Drugs 2003; 4: Witvrouw M, Hantson A, Hombrouck A, et al. Multiple mutations in HIV-1 integrase confer resistance to the PyranoDipyrimidine V th HIV DRP Symposium on Antiretroviral Drug Resistance, November 2004 Chantilly, VA, USA. Abstract # Pannecouque C, Pluymers W, Van Maele B, et al. New class of HIV integrase inhibitors that block viral replication in cell culture. Curr Biol 2002; 12: Hazuda DJ, Felock P, Witmer M et al. Inhibitors of strand transfer that prevent integration and inhibit HIV-1 replication in cells. Science 2000; 287: Dayam R, Al-Mawsawi LQ, Neamati N. HIV-1 integrase inhibitors: an emerging clinical reality. Drugs R D 2007; 8: Hazuda DJ, Anthony NJ, Gomez RP, et. al. A naphthyridine carboxamide provides evidence for discordant resistance between mechanistically identical inhibitors of HIV-1 integrase. Proc Natl Acad Sci U S A 2004; 101: Kodama E, Shimura K, Sakagami Y, et al. In vitro antiviral activity and resistance profile of a novel HIV integrase inhibitor JTK-303/GS th Interscience Conference on Antimicrobial Agents and Chemotherapy September 27 30, 2006, San Francisco, CA, USA. Abstract H Grinsztejn B, Nguyen, BY, Katlama C, et al. Potent antiretroviral effect of MK-0518, a novel HIV-1 integrase inhibitor, in patients with triple-class resistant virus. 46th Interscience Conference on Antimicrobial Agents and Chemotherapy September , San Francisco CA, USA. Abstract H1670b. 22. Bonnenfant S, Thomas CM, Vita C, et al. Styrylquinolines, integrase inhibitors acting prior to integration: a new mechanism of action for anti-integrase agents. J Virol 2004; 78: Mousnier A, Leh H, Mouscadet JF, Dargemont C. Nuclear import of HIV-1 integrase is inhibited in vitro by styrylquinoline derivatives. Mol Pharmacol 2004; 66: Di Santo R, Costi R, Roux A, et al. Novel bifunctional quinolonyl diketo acid derivatives as HIV-1 integrase inhibitors: design, synthesis, biological activities, and mechanism of action. J Med Chem 2006; 49: Marchand C, Johnson AA, Karki RG, et al. Metal-dependent inhibition of HIV-1 integrase by beta-diketo acids and resistance of the soluble double-mutant (F185K/C280S). Mol Pharmacol 2003; 64: Lee DJ, Robinson WE Jr. Human immunodeficiency virus type 1 (HIV-1) integrase: resistance to DKA integrase inhibitors impairs HIV replication and integration and confers cross resistance to L-chicoric acid. J Virol, 2004; 78: Barreca ML, Lee KW, Chimirri A, Briggs JM. Molecular dynamics of the wild type and double mutant HIV-1 integrase complexed with 5-CITEP inhibitor: mechanism for inhibition and drug resistance. Biophys J 2003; 84: Johnson VA, Brun-Vezinet F, Clotet B, et al. Update of the drug resistance mutations in HIV-1: Top HIV Med 2005; 13: Kozal MJ, Shah N, Shen N, et al. Extensive polymorphisms observed in HIV-1 clade B protease gene using high-density oligonucleotide arrays. Nat Med 1996; 2: De Meyer S, Hill A, De Baere I, et al. Effect of baseline susceptibility and on-treatment mutations on TMC114 and control PI efficacy: preliminary analysis of data from PI-experienced patients from POWER 1 and POWER 2. 13th Conference on Retroviral and Opportunistic Infections 2006, Denver, CO, USA. Abstract # Kozal MJ, Huppler Hullsiek K, Leduc R, et al. Prevalence and impact of HIV-1 protease codon 33 mutations and polymorphisms in treatment-naïve and treatment- Antiviral Therapy 12:4 569
8 M Lataillade et al. experienced patients enrolling in clinical trials. Antivir Ther 2006; 11: Fikkert V, Van Maele B, Vercammen J, Hantson A, Van Remoortel B, Michiels M. Development of resistance against diketo derivatives of human immunodeficiency virus type 1 by progressive accumulation of integrase mutations. J Virol 2003; 77: Fikkert V, Hombrouk A, Van Remoortel B, et al. Multiple mutations in HIV-1 IN confer resistance to the clinical trial drug S AIDS 2004; 18: Jones G, Ledford R, Yu F, Miller M, Tsiang M, McColl D. Resistance Profile of HIV-1 Mutants in vitro Selected by the HIV-1 Integrase Inhibitor, GS-9137 (JTK-303). 14th Conference on Retroviral and Opportunistic Infections February 2007, Los Angeles CA, USA. Abstract Wai J, Fisher T, Embrey M, et al. Next generation of inhibitors of HIV-1 integrase strand transfer inhibitor: structural diversity and resistance profiles. 14th Conference on Retroviral and Opportunistic Infections February 2007, Los Angeles CA, USA. Abstract 87. Accepted for publication 16 March Hackett J, Jr, Swanson P, Harris B, Holzmayer V, Yamaguchi J, Bodelle P, Brennan C, Schochetman G, and Devare S. Comprehensive evaluation of HIV-1 integrase gene diversity in group M, N, and O viruses. 12th Conference on Retroviral and Opportunistic Infections, February 2005, Boston MA, USA. Abstract LB# Zioni, R, Rhee S, Liu T, Shafer R. Natural variation of HIV-1 group M integrase: implications for integrase inhibitor therapy. 14th Conference on Retroviral and Opportunistic Infections February 2007, Los Angeles CA, USA. Abstract Lu R, Limon A, Ghory HZ, Engelman A. Genetic analyses of DNA-binding mutants in the catalytic core domain of human immunodeficiency virus type 1 integrase. J Virol 2005; 79: Llano M, Saenz DT, Meehan A, et al. An essential role for LEDGF/p75 in HIV integration. Science 2006; 314: Emiliani S, Mousnier A, Busschots K, Maroun M, Van Maele B, Tempe D. Integrase mutants defective for interaction with LEDGF/p75 are impaired in chromosome tethering and HIV- 1 replication. J Biol Chem 2005; 280: International Medical Press
Resistance to inhibitors of the human immunodeficiency virus type 1 integration
 Resistance to inhibitors of the human immunodeficiency virus type 1 integration REVIEW ARTICLE ABSTRACT This review will summarize the role of integrase in HIV-1 infection, the mechanism of integrase inhibitors
Resistance to inhibitors of the human immunodeficiency virus type 1 integration REVIEW ARTICLE ABSTRACT This review will summarize the role of integrase in HIV-1 infection, the mechanism of integrase inhibitors
The HIV life cycle. integration. virus production. entry. transcription. reverse transcription. nuclear import
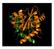 The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication
 Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Short communication Genetic barriers for integrase inhibitor drug resistance in HIV type-1 B and CRF02_AG subtypes
 Antiviral Therapy 14:123 129 Short communication Genetic barriers for integrase inhibitor drug resistance in HIV type-1 B and CRF2_AG subtypes Almoustapha-Issiaka Maïga 1, Isabelle Malet 1 *, Cathia Soulie
Antiviral Therapy 14:123 129 Short communication Genetic barriers for integrase inhibitor drug resistance in HIV type-1 B and CRF2_AG subtypes Almoustapha-Issiaka Maïga 1, Isabelle Malet 1 *, Cathia Soulie
Resistance Characteristics of Integrase Inhibitors
 Resistance Characteristics of Integrase Inhibitors Madrid, November 2016 Jonathan M Schapiro, MD National Hemophilia Center, Israel Stanford University School of Medicine, USA Disclaimer Presentation includes
Resistance Characteristics of Integrase Inhibitors Madrid, November 2016 Jonathan M Schapiro, MD National Hemophilia Center, Israel Stanford University School of Medicine, USA Disclaimer Presentation includes
Mutations to Integrase Inhibitors in real life
 Mutations to Integrase Inhibitors in real life González-Domenech CM, Viciana I, Sena Corrales G, Delgado M, De la Torre J, Torres Tortosa M, Téllez F, Jarilla F, Clavijo E, Santos J BACKGROUND INSTI Block
Mutations to Integrase Inhibitors in real life González-Domenech CM, Viciana I, Sena Corrales G, Delgado M, De la Torre J, Torres Tortosa M, Téllez F, Jarilla F, Clavijo E, Santos J BACKGROUND INSTI Block
Resistance to Integrase Strand Transfer Inhibitors
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
Analysis of Natural Sequence Variation and Covariation in Human Immunodeficiency Virus Type 1 Integrase
 JOURNAL OF VIROLOGY, Sept. 2008, p. 9228 9235 Vol. 82, No. 18 0022-538X/08/$08.00 0 doi:10.1128/jvi.01535-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Analysis of Natural
JOURNAL OF VIROLOGY, Sept. 2008, p. 9228 9235 Vol. 82, No. 18 0022-538X/08/$08.00 0 doi:10.1128/jvi.01535-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Analysis of Natural
Integrase variability and susceptibility to HIV integrase inhibitors: impact of subtypes, antiretroviral experience and duration of HIV infection
 Journal of Antimicrobial Chemotherapy Advance Access published December 9, 2009 J Antimicrob Chemother doi:10.1093/jac/dkp423 Integrase variability and susceptibility to HIV integrase inhibitors: impact
Journal of Antimicrobial Chemotherapy Advance Access published December 9, 2009 J Antimicrob Chemother doi:10.1093/jac/dkp423 Integrase variability and susceptibility to HIV integrase inhibitors: impact
Received 11 July 2007/Returned for modification 20 August 2007/Accepted 21 March 2008
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, June 2008, p. 2069 2078 Vol. 52, No. 6 0066-4804/08/$08.00 0 doi:10.1128/aac.00911-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Mutations
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, June 2008, p. 2069 2078 Vol. 52, No. 6 0066-4804/08/$08.00 0 doi:10.1128/aac.00911-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Mutations
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Drug Resistance to Integrase Strand Transfer Inhibitors in HIV-1 Chilean Patients. Frequency and evolution between the years 2013 and 2016.
 Drug Resistance to Integrase Strand Transfer Inhibitors in HIV-1 Chilean Patients. Frequency and evolution between the years 2013 and 2016. Sidgman F. 1, Valdés F. 1, Ferrer P. 2, Maureira V. 1, Afani
Drug Resistance to Integrase Strand Transfer Inhibitors in HIV-1 Chilean Patients. Frequency and evolution between the years 2013 and 2016. Sidgman F. 1, Valdés F. 1, Ferrer P. 2, Maureira V. 1, Afani
Transmission of integrase resistance HIV
 Transmission of integrase resistance HIV Charles Boucher, MD, PhD Clinical Virology, Dept. Viroscience, Erasmus Medical Center, Erasmus Universiy, The Netherlands Major resistance mutations (Stanford)
Transmission of integrase resistance HIV Charles Boucher, MD, PhD Clinical Virology, Dept. Viroscience, Erasmus Medical Center, Erasmus Universiy, The Netherlands Major resistance mutations (Stanford)
Introduction to the Impact of Resistance in Hepatitis C
 Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Acquired INSTI Resistance. Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Acquired INSTI Resistance Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosures The speaker has received consulting honoraria, speaker
Acquired INSTI Resistance Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosures The speaker has received consulting honoraria, speaker
Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
 Viruses 2010, 2, 2777-2781; doi:10.3390/v2122777 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
Viruses 2010, 2, 2777-2781; doi:10.3390/v2122777 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
History (August 2010) Therapy for Experienced Patients. History (September 2010) History (November 2010) 12/2/11
 (August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
(August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
HIV-1 Integrase Strand Transfer Inhibitors: Novel Insights into their Mechanism of Action
 CMMETARY SPECIAL ISSUE HIV-1 Integrase Strand Transfer Inhibitors: ovel Insights into their Mechanism of Action Krishan K. Pandey and Duane P. Grandgenett Institute for Molecular Virology, Saint Louis
CMMETARY SPECIAL ISSUE HIV-1 Integrase Strand Transfer Inhibitors: ovel Insights into their Mechanism of Action Krishan K. Pandey and Duane P. Grandgenett Institute for Molecular Virology, Saint Louis
Introduction to HIV Drug Resistance. Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School
 Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
ALLINIs modulate the interaction between HIV-1 Rev and integrase
 ALLINIs modulate the interaction between HIV-1 Rev and integrase Date: 13 June 2017 Author Shaakira Abrahams Salerwe Mosebi, Qasim Fish, Raymond Hewer, Maria Papathanasopoulos Outline Background Results
ALLINIs modulate the interaction between HIV-1 Rev and integrase Date: 13 June 2017 Author Shaakira Abrahams Salerwe Mosebi, Qasim Fish, Raymond Hewer, Maria Papathanasopoulos Outline Background Results
Pointer Elvitegravir: a new HIV integrase inhibitor
 Antiviral Chemistry & Chemotherapy 2009 20: 79 85 (doi: 10.3851/IMP1397) Pointer Elvitegravir: a new IV integrase inhibitor Kazuya Shimura 1 and Eiichi Kodama 1,2 * 1 Laboratory of Virus Control, Institute
Antiviral Chemistry & Chemotherapy 2009 20: 79 85 (doi: 10.3851/IMP1397) Pointer Elvitegravir: a new IV integrase inhibitor Kazuya Shimura 1 and Eiichi Kodama 1,2 * 1 Laboratory of Virus Control, Institute
It takes more than just a single target
 It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762)
 Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
HIV Drug Resistance: An Overview
 Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
HIV-1 Subtypes: An Overview. Anna Maria Geretti Royal Free Hospital
 HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
Update on HIV-1 Drug Resistance and Tropism Testing
 Update on HIV-1 Drug Resistance and Tropism Testing Daniel R. Kuritzkes, MD Section of Retroviral Therapeutics Brigham and Women s Hospital Harvard Medical School ACTHIV 2011: A State-of-the-Science Conference
Update on HIV-1 Drug Resistance and Tropism Testing Daniel R. Kuritzkes, MD Section of Retroviral Therapeutics Brigham and Women s Hospital Harvard Medical School ACTHIV 2011: A State-of-the-Science Conference
ORIGINAL ARTICLE /j x. Brescia, Italy
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E,
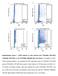 Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
HIV 101: Fundamentals of HIV Infection
 HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
Integrase strand-transfer inhibitors primary resistance in patients with acute/recent HIV infection
 Integrase strand-transfer inhibitors primary resistance in patients with acute/recent HIV infection Juan Ambrosioni, David Nicolás, Christian Manzardo, Fernando Agüero, José Luis Blanco, Maria del Mar
Integrase strand-transfer inhibitors primary resistance in patients with acute/recent HIV infection Juan Ambrosioni, David Nicolás, Christian Manzardo, Fernando Agüero, José Luis Blanco, Maria del Mar
Evaluation and Management of Virologic Failure
 National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
RESEARCH B/F/TAF in Treatment-Naïve HIV-1 and HIV-1 RNA Suppressed Switch Patients
 RESEARCH B/F/TAF in Treatment-Naïve HIV-1 and HIV-1 RNA Suppressed Switch Patients Kirsten White Gilead Sciences, Inc., Foster City, CA Background Bictegravir (BIC; B) is a novel, unboosted integrase strand
RESEARCH B/F/TAF in Treatment-Naïve HIV-1 and HIV-1 RNA Suppressed Switch Patients Kirsten White Gilead Sciences, Inc., Foster City, CA Background Bictegravir (BIC; B) is a novel, unboosted integrase strand
Update on HIV Drug Resistance. Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
The E138A substitution in HIV-1 reverse transcriptase decreases in vitro. susceptibility to emtricitabine as indicated by competitive fitness assays
 AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
modified dye uptake assay including formazan test EC 90 not tested plaque reduction assay
 Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
Understanding and. Pre-Integration. Post-Integration. By Andrea Low, md and Mark Muesing, phd. Nucleus. Cytoplasm
 Understanding and Inhibiting Integrase in the Treatment of HIV Disease Reprinted from The prn Notebook november 2006 Dr. James F. Braun, Editor-in-Chief Meri D. Pozo, PhD, Managing Editor Published in
Understanding and Inhibiting Integrase in the Treatment of HIV Disease Reprinted from The prn Notebook november 2006 Dr. James F. Braun, Editor-in-Chief Meri D. Pozo, PhD, Managing Editor Published in
Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosure The speaker serves as a consultant to, and has received
Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosure The speaker serves as a consultant to, and has received
Diagnostic Methods of HBV and HDV infections
 Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
Second-Line Therapy NORTHWEST AIDS EDUCATION AND TRAINING CENTER
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
Natural Polymorphisms of Human Immunodeficiency Virus Type 1 Integrase and Inherent Susceptibilities to a Panel of Integrase Inhibitors
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Oct. 2009, p. 4275 4282 Vol. 53, No. 10 0066-4804/09/$08.00 0 doi:10.1128/aac.00397-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Natural
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Oct. 2009, p. 4275 4282 Vol. 53, No. 10 0066-4804/09/$08.00 0 doi:10.1128/aac.00397-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Natural
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with
 Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
New options for new and old patients. Antonella Castagna
 New options for new and old patients Antonella Castagna Disclosures I have received funding for Advisory Boards, Speaker Panels and for preparation of educational materials from the following: Gilead Sciences
New options for new and old patients Antonella Castagna Disclosures I have received funding for Advisory Boards, Speaker Panels and for preparation of educational materials from the following: Gilead Sciences
Title. HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in
 Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation
 Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
Clinical utility of NGS for the detection of HIV and HCV resistance
 18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY. 16th International WAVLD symposium, 10th OIE Seminar
 NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY S. Van Borm, I. Monne, D. King and T. Rosseel 16th International WAVLD symposium, 10th OIE Seminar 07.06.2013 Viral livestock
NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY S. Van Borm, I. Monne, D. King and T. Rosseel 16th International WAVLD symposium, 10th OIE Seminar 07.06.2013 Viral livestock
Mutations Associated with Failure of Raltegravir Treatment Affect Integrase Sensitivity to the Inhibitor In Vitro
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 28, p. 1351 1358 Vol. 52, No. 4 66-484/8/$8. doi:1.1128/aac.1228-7 Copyright 28, American Society for Microbiology. All Rights Reserved. Mutations Associated
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Apr. 28, p. 1351 1358 Vol. 52, No. 4 66-484/8/$8. doi:1.1128/aac.1228-7 Copyright 28, American Society for Microbiology. All Rights Reserved. Mutations Associated
High Failure Rate of the ViroSeq HIV-1 Genotyping System for Drug Resistance Testing in Cameroon, a Country with Broad HIV-1 Genetic Diversity
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
Micropathology Ltd. University of Warwick Science Park, Venture Centre, Sir William Lyons Road, Coventry CV4 7EZ
 www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
Conference Reports for NATAP. S/GSK Integrase Inhibitor Resistance Profile
 Conference Reports for NATAP EACS - 12th European AIDS Conference November 11-14, 2009 Cologne, Germany Back S/GSK1349572 Integrase Inhibitor Resistance Profile Reported by Jules Levin EACS Oct 31-Nov
Conference Reports for NATAP EACS - 12th European AIDS Conference November 11-14, 2009 Cologne, Germany Back S/GSK1349572 Integrase Inhibitor Resistance Profile Reported by Jules Levin EACS Oct 31-Nov
Principles of HIV Drug Resistance: Resistance to New Drug Classes. Mark A Wainberg McGill University AIDS Centre Montreal, Quebec, Canada
 Principles of HIV Drug Resistance: Resistance to New Drug Classes Mark A Wainberg McGill University AIDS Centre Montreal, Quebec, Canada Why Is It Important to Understand HIV Drug Resistance? 1. Resistance
Principles of HIV Drug Resistance: Resistance to New Drug Classes Mark A Wainberg McGill University AIDS Centre Montreal, Quebec, Canada Why Is It Important to Understand HIV Drug Resistance? 1. Resistance
Transcriptional co-activator LEDGF/p75 modulates HIV-1 integrase ACCEPTED. mediated concerted integration
 JVI Accepts, published online ahead of print on 31 January 2007 J. Virol. doi:10.1128/jvi.02322-06 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 31 January 2007 J. Virol. doi:10.1128/jvi.02322-06 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Rajesh Kannangai Phone: ; Fax: ; *Corresponding author
 Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Antiviral Therapy 2011; 16: (doi: /IMP1851)
 Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
Relative activity (%) SC35M
 a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
2 nd Line Treatment and Resistance. Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012
 2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
Broad Antiretroviral Activity and Resistance Profile of the Novel Human Immunodeficiency Virus Integrase Inhibitor Elvitegravir (JTK-303/GS-9137)
 JOURNAL OF VIROLOGY, Jan. 2008, p. 764 774 Vol. 82, No. 2 0022-538X/08/$08.00 0 doi:10.1128/jvi.01534-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Broad Antiretroviral Activity
JOURNAL OF VIROLOGY, Jan. 2008, p. 764 774 Vol. 82, No. 2 0022-538X/08/$08.00 0 doi:10.1128/jvi.01534-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Broad Antiretroviral Activity
CL Vavro, 1 J Huang, 2 C Avatapally, 1 S Min, 1 and M Ait-Khaled 3. GlaxoSmithKline: 1 Research Triangle Park,NC, USA; 2 Toronto, ON, Canada; 3
 Durable Efficacy and Limited Integrase Resistance Evolution in Subjects Receiving Dolutegravir After Failing Prior Integrase Inhibitor (INI) Regimens: Week 48 Results from VIKING-3 CL Vavro, 1 J Huang,
Durable Efficacy and Limited Integrase Resistance Evolution in Subjects Receiving Dolutegravir After Failing Prior Integrase Inhibitor (INI) Regimens: Week 48 Results from VIKING-3 CL Vavro, 1 J Huang,
Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System
 Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System Joel O. Wertheim 1,2, Alexandra M. Oster 3, Jeffrey A. Johnson 3, William M. Switzer 3, Neeraja Saduvala
Transmission Fitness of Drug- Resistant HIV Revealed in the United States National Surveillance System Joel O. Wertheim 1,2, Alexandra M. Oster 3, Jeffrey A. Johnson 3, William M. Switzer 3, Neeraja Saduvala
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Identification of Novel Mutations Responsible for Resistance to MK-2048, a Second-Generation HIV-1 Integrase Inhibitor
 JOURNAL OF VIROLOGY, Sept. 2010, p. 9210 9216 Vol. 84, No. 18 0022-538X/10/$12.00 doi:10.1128/jvi.01164-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. Identification of Novel
JOURNAL OF VIROLOGY, Sept. 2010, p. 9210 9216 Vol. 84, No. 18 0022-538X/10/$12.00 doi:10.1128/jvi.01164-10 Copyright 2010, American Society for Microbiology. All Rights Reserved. Identification of Novel
Hepatitis C Resistance Associated Variants (RAVs)
 Hepatitis C Resistance Associated Variants (RAVs) Atif Zaman, MD MPH Oregon Health & Science University Professor of Medicine Division of Gastroenterology and Hepatology Nothing to disclose Disclosure
Hepatitis C Resistance Associated Variants (RAVs) Atif Zaman, MD MPH Oregon Health & Science University Professor of Medicine Division of Gastroenterology and Hepatology Nothing to disclose Disclosure
Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review
 pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
VIKING STUDIES Efficacy and safety of dolutegravir in treatment-experienced subjects
 VIKING STUDIES Efficacy and safety of dolutegravir in treatment-experienced subjects IL/DLG/0040/14 June 2014 GSK (Israel) Ltd. Basel 25, Petach Tikva. Tel-03-9297100 Medical information service: il.medinfo@gsk.com
VIKING STUDIES Efficacy and safety of dolutegravir in treatment-experienced subjects IL/DLG/0040/14 June 2014 GSK (Israel) Ltd. Basel 25, Petach Tikva. Tel-03-9297100 Medical information service: il.medinfo@gsk.com
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Effect of HIV-1 Subtype C integrase mutations implied using molecular modeling and docking data
 www.bioinformation.net Volume 12(3) Hypothesis Effect of HIV-1 Subtype C integrase mutations implied using molecular modeling and docking data Jaiprasath Sachithanandham 1, Karnati Konda Reddy 2, King
www.bioinformation.net Volume 12(3) Hypothesis Effect of HIV-1 Subtype C integrase mutations implied using molecular modeling and docking data Jaiprasath Sachithanandham 1, Karnati Konda Reddy 2, King
Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir.
 Abstract no. MOLBPEA13 Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir. G. Vansant,,A. Bruggemans, L. Vranckx, S. Saleh, I. Zurnic, F. Christ,
Abstract no. MOLBPEA13 Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir. G. Vansant,,A. Bruggemans, L. Vranckx, S. Saleh, I. Zurnic, F. Christ,
HIV Treatment: New and Veteran Drugs Classes
 HIV Treatment: New and Veteran Drugs Classes Jonathan M Schapiro, MD National Hemophilia Center Stanford University School of Medicine Rome, March 2013 Overview Many excellent antiretroviral agents are
HIV Treatment: New and Veteran Drugs Classes Jonathan M Schapiro, MD National Hemophilia Center Stanford University School of Medicine Rome, March 2013 Overview Many excellent antiretroviral agents are
Deep-Sequencing of HIV-1
 Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Inhibiting the HIV Integration Process: Past, Present, and the Future Roberto Di Santo*
 pubs.acs.org/jmc Terms of Use Inhibiting the HIV Integration Process: Past, Present, and the Future Roberto Di Santo* Dipartimento di Chimica e Tecnologie del Farmaco, Istituto Pasteur, Fondazione Cenci
pubs.acs.org/jmc Terms of Use Inhibiting the HIV Integration Process: Past, Present, and the Future Roberto Di Santo* Dipartimento di Chimica e Tecnologie del Farmaco, Istituto Pasteur, Fondazione Cenci
Disclosure. Companies/funders. Research grants (to the institute) Merck, Gilead, Janssen, ViiV. Speakers fee. Viiv, Abvie, Gilead.
 Disclosure Companies/funders Research grants (to the institute) Merck, Gilead, Janssen, ViiV Speakers fee Viiv, Abvie, Gilead Other Co-owner/Scientific director Virology Education Integration Insertion
Disclosure Companies/funders Research grants (to the institute) Merck, Gilead, Janssen, ViiV Speakers fee Viiv, Abvie, Gilead Other Co-owner/Scientific director Virology Education Integration Insertion
Disclosures. Introduction to ARV Drug Resistance New Clinicians Workshop 12/9/16. Introduction. ARS Question
 Disclosures Introduction to ARV Drug Resistance New Clinicians Workshop I have no disclosures Susa Coffey, MD Division of HIV, ID and Global Medicine ARS Question Which resistance test do you order for
Disclosures Introduction to ARV Drug Resistance New Clinicians Workshop I have no disclosures Susa Coffey, MD Division of HIV, ID and Global Medicine ARS Question Which resistance test do you order for
Received 13 October 2003/Accepted 27 January 2004
 JOURNAL OF VIROLOGY, June 2004, p. 5835 5847 Vol. 78, No. 11 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.11.5835 5847.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
JOURNAL OF VIROLOGY, June 2004, p. 5835 5847 Vol. 78, No. 11 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.11.5835 5847.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
Hepatitis C: Aplicaciones Clínicas de la Resistencia. Eva Poveda Division of Clinical Virology INIBIC-Complexo Hospitalario Universitario de A Coruña
 Hepatitis C: Aplicaciones Clínicas de la Resistencia Eva Poveda Division of Clinical Virology INIBIC-Complexo Hospitalario Universitario de A Coruña DAA agents approved or in more advanced stages of clinical
Hepatitis C: Aplicaciones Clínicas de la Resistencia Eva Poveda Division of Clinical Virology INIBIC-Complexo Hospitalario Universitario de A Coruña DAA agents approved or in more advanced stages of clinical
Scottish Medicines Consortium
 Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
Mapping evolutionary pathways of HIV-1 drug resistance using conditional selection pressure. Christopher Lee, UCLA
 Mapping evolutionary pathways of HIV-1 drug resistance using conditional selection pressure Christopher Lee, UCLA HIV-1 Protease and RT: anti-retroviral drug targets protease RT Protease: responsible for
Mapping evolutionary pathways of HIV-1 drug resistance using conditional selection pressure Christopher Lee, UCLA HIV-1 Protease and RT: anti-retroviral drug targets protease RT Protease: responsible for
Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies
 Research Article Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies Pawan Gupta 1,2 *, Prabha Garg 2 ABSTRACT Background and Aim: The
Research Article Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies Pawan Gupta 1,2 *, Prabha Garg 2 ABSTRACT Background and Aim: The
NIH Public Access Author Manuscript J Acquir Immune Defic Syndr. Author manuscript; available in PMC 2013 September 01.
 NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
From Mosquitos to Humans: Genetic evolution of Zika Virus
 Article: From Mosquitos to Humans: Genetic evolution of Zika Virus Renata Pellegrino, PhD Director, Sequencing lab Center for Applied Genomics The Children s Hospital of Philadelphia Journal Club Clinical
Article: From Mosquitos to Humans: Genetic evolution of Zika Virus Renata Pellegrino, PhD Director, Sequencing lab Center for Applied Genomics The Children s Hospital of Philadelphia Journal Club Clinical
Clinical Applications of Resistance Stuart C. Ray, MD
 Clinical Applications of Resistance Stuart C. Ray, MD Professor of Medicine and Oncology Director, Infectious Diseases Fellowship Training Program Johns Hopkins University School of Medicine Disclosures
Clinical Applications of Resistance Stuart C. Ray, MD Professor of Medicine and Oncology Director, Infectious Diseases Fellowship Training Program Johns Hopkins University School of Medicine Disclosures
Mapping Evolutionary Pathways of HIV-1 Drug Resistance. Christopher Lee, UCLA Dept. of Chemistry & Biochemistry
 Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
LESSON 4.6 WORKBOOK. Designing an antiviral drug The challenge of HIV
 LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
Case Study. Dr Sarah Sasson Immunopathology Registrar. HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital.
 Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
The Lentiviral Integrase Binding Protein LEDGF/p75 and HIV-1 Replication
 Review The Lentiviral Integrase Binding Protein LEDGF/p75 and HIV-1 Replication Alan Engelman 1 *, Peter Cherepanov 2 * 1 Department of Cancer Immunology and AIDS, Dana-Farber Cancer Institute, Division
Review The Lentiviral Integrase Binding Protein LEDGF/p75 and HIV-1 Replication Alan Engelman 1 *, Peter Cherepanov 2 * 1 Department of Cancer Immunology and AIDS, Dana-Farber Cancer Institute, Division
PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES
 PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES Mark A. Wainberg McGill University AIDS Centre Jewish General Hospital Montreal, Quebec,
PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES Mark A. Wainberg McGill University AIDS Centre Jewish General Hospital Montreal, Quebec,
Reassortment of influenza A virus genes linked to PB1 polymerase gene
 International Congress Series 1263 (2004) 714 718 Reassortment of influenza A virus genes linked to PB1 polymerase gene Jean C. Downie* www.ics-elsevier.com Centre for Infectious Diseases and Microbiology,
International Congress Series 1263 (2004) 714 718 Reassortment of influenza A virus genes linked to PB1 polymerase gene Jean C. Downie* www.ics-elsevier.com Centre for Infectious Diseases and Microbiology,
The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide polymorphisms at thymidine analogue mutation sites
 Journal of Antimicrobial Chemotherapy Advance Access published June 7, 2013 J Antimicrob Chemother doi:10.1093/jac/dkt204 The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide
Journal of Antimicrobial Chemotherapy Advance Access published June 7, 2013 J Antimicrob Chemother doi:10.1093/jac/dkt204 The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide
Nuclear import of HIV
 Center for Molecular Medicine Nuclear import of HIV Zeger Debyser MD PhD Molecular Virology and Gene Therapy, KULeuven Flanders, Europe HIV/AIDS: in search of novel drugs against new targets High genetic
Center for Molecular Medicine Nuclear import of HIV Zeger Debyser MD PhD Molecular Virology and Gene Therapy, KULeuven Flanders, Europe HIV/AIDS: in search of novel drugs against new targets High genetic
Emerging Resistance Profiles of Newly Approved Antiretroviral Drugs
 Perspective Emerging Resistance Profiles of Newly Approved Antiretroviral Drugs The antiretroviral treatment goal in highly treatment-experienced patients is now suppression of viral replication to undetectable
Perspective Emerging Resistance Profiles of Newly Approved Antiretroviral Drugs The antiretroviral treatment goal in highly treatment-experienced patients is now suppression of viral replication to undetectable
Whole genome deep sequencing of HIV reveals extensive multi-class drug resistance in Nigerian patients failing first-line antiretroviral therapy
 Whole genome deep sequencing of HIV reveals extensive multi-class drug resistance in Nigerian patients failing first-line antiretroviral therapy K El Bouzidi 1,, RP Datir 1, V Kwaghe 3, S Roy 1, D Frampton
Whole genome deep sequencing of HIV reveals extensive multi-class drug resistance in Nigerian patients failing first-line antiretroviral therapy K El Bouzidi 1,, RP Datir 1, V Kwaghe 3, S Roy 1, D Frampton
HIV-1 resistance testing from proviral DNA
 HIV-1 resistance testing from proviral DNA Alexander Thielen AREVIR 2018 Resistance testing from proviral DNA? amount of samples with low viral loads increasing desire to switch under successful therapy
HIV-1 resistance testing from proviral DNA Alexander Thielen AREVIR 2018 Resistance testing from proviral DNA? amount of samples with low viral loads increasing desire to switch under successful therapy
INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING
 INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING Dr. Danni Kirwan ID/Microbiology SpR St. George s Hospital, London ARV initiation in treatment-naïve patients BHIVA,
INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING Dr. Danni Kirwan ID/Microbiology SpR St. George s Hospital, London ARV initiation in treatment-naïve patients BHIVA,
Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal Introduction
 Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
Characterizing intra-host influenza virus populations to predict emergence
 Characterizing intra-host influenza virus populations to predict emergence June 12, 2012 Forum on Microbial Threats Washington, DC Elodie Ghedin Center for Vaccine Research Dept. Computational & Systems
Characterizing intra-host influenza virus populations to predict emergence June 12, 2012 Forum on Microbial Threats Washington, DC Elodie Ghedin Center for Vaccine Research Dept. Computational & Systems
Mutants and HBV vaccination. Dr. Ulus Salih Akarca Ege University, Izmir, Turkey
 Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Review HIV-1 integrase inhibitors: a review of their chemical development
 Antiviral Chemistry & Chemotherapy 2011; 22:95 105 (doi: 10.3851/IMP1740) Review IV-1 integrase inhibitors: a review of their chemical development Kundan B Ingale 1 *, Manish S Bhatia 1 1 Department of
Antiviral Chemistry & Chemotherapy 2011; 22:95 105 (doi: 10.3851/IMP1740) Review IV-1 integrase inhibitors: a review of their chemical development Kundan B Ingale 1 *, Manish S Bhatia 1 1 Department of
