Integrase variability and susceptibility to HIV integrase inhibitors: impact of subtypes, antiretroviral experience and duration of HIV infection
|
|
|
- Erika Cook
- 5 years ago
- Views:
Transcription
1 Journal of Antimicrobial Chemotherapy Advance Access published December 9, 2009 J Antimicrob Chemother doi: /jac/dkp423 Integrase variability and susceptibility to HIV integrase inhibitors: impact of subtypes, antiretroviral experience and duration of HIV infection Carolina Garrido 1, Anna Maria Geretti 2, Natalia Zahonero 1, Clare Booth 2, Angela Strang 2, Vincent Soriano 1 * and Carmen De Mendoza 1 1 Department of Infectious Diseases, Hospital Carlos III, Madrid, Spain; 2 Department of Virology, Royal Free Hampstead NHS Trust and University College London Medical School, London, UK *Corresponding author. Tel: þ ; Fax: þ ; vsoriano@dragonet.es Received 12 June 2009; returned 25 October 2009; revised 28 October 2009; accepted 29 October 2009 Background: Little is known about the extent and predictors of polymorphisms potentially influencing the susceptibility to HIV integrase inhibitors (INIs). Methods: Genetic sequences of HIV integrase were obtained from INI-naive patients at two European clinics. The 39 amino acid changes at 29 integrase positions so far associated with INI resistance were examined according to HIV clade, prior antiretroviral exposure and duration of HIV infection. Results: Integrase sequences were obtained from 418 patients, 294 (70.3%) infected with clade B and 124 (29.7%) infected with non-b variants (predominantly CRF02, A, C and D). Overall, 40% of patients were antiretroviral experienced and 32.8% were recent seroconverters. The most prevalent INI resistance-associated mutations were V72I (63.9%), V201I (54.8%), T206S (25.4%), I203M (9.8%) and K156N (7.4%). Major INI resistance mutations at positions 66, 92, 143, 148 and 155 were not detected. The mean number of polymorphic sites was greater in non-b than in B variants (2.17 versus 1.59; P, 0.001), and in antiretroviralexperienced than in drug-naive patients (1.89 versus 1.68; P¼0.034), whereas no significant differences were seen comparing recent seroconverters and chronically infected persons. Conclusions: Major INI resistance-associated mutations are very rare, if indeed ever present, in INI-naive patients. However, polymorphisms at positions which may influence the genetic barrier and/or drive the selection of specific INI resistance pathways are common, especially in HIV non-b subtypes. Keywords: polymorphisms, non-b subtypes, drug resistance, raltegravir, elvitegravir Introduction HIV replication requires three viral enzymes along with metabolites and machinery provided by the infected cell. The viral enzymes are encoded by the pol gene, and have been the main targets of antiretroviral drugs. 1 Integrase inhibitors (INIs) represent the latest approved drug family for the treatment of HIV infection. 2,3 The integrase enzyme is a 288 amino acid protein encoded by the 5 0 end of the pol gene. It consists of three different structural and functional domains: the N-terminal domain (amino acids 1 49), which contains the HHCC motif for zinc binding; the central catalytic domain (CCD; amino acids ), which contains the DDE motif; and the C-terminal domain (amino acids ), which displays DNA binding activity. 3,4 Integration is a complex process that comprises different steps: (i) assembly of the integrase enzyme at the end of the HIV long terminal repeat (LTR), forming the pre-integration complex; (ii) cleavage of the terminal GT dinucleotide from the 3 0 end of each LTR (known as 3 0 processing ); (iii) translocation of the pre-integration complex to the nucleus through the nucleopore; (iv) covalent linkage of the viral DNA into the cellular chromosome (known as strand transfer ); and (v) gap repair by cellular enzymes. 5 Although the integrase catalyses both 3 0 processing and strand transfer, the current INIs raltegravir and elvitegravir only inhibit its strand transfer activity. 3,6,7 Raltegravir (Isentress, Merck) is the first INI commercially available, and elvitegravir (GS-3197, Gilead) is currently in Phase III clinical development. 7 Other INIs are in earlier steps of clinical development, some of which (e.g. GSK or GSK ) may retain activity against viruses resistant to raltegravir or elvitegravir. Overall, all INIs display high potency against viruses resistant to other antiretroviral agents and constitute a valuable option for salvage therapy. 7 More recently # The Author Published by Oxford University Press on behalf of the British Society for Antimicrobial Chemotherapy. All rights reserved. For Permissions, please journals.permissions@oxfordjournals.org 1of7
2 Garrido et al. raltegravir has also received approval to be used as part of firstline antiretroviral therapy. However, resistance to INIs rapidly emerges as a result of selection of a few resistance-associated mutations (RAMs) within the integrase gene. Several changes influencing the susceptibility to INIs have been described both in vitro 8 10 and in vivo, including mutations that play a major role in resistance (major INI RAMs) and changes that play a contributing role (minor INI RAMs). The aim of this study was to assess the prevalence of naturally occurring polymorphisms within the HIV-1 integrase gene which might influence the susceptibility to INIs, including both major and minor INI RAMs. Particular attention was devoted to recognizing the influence of HIV-1 subtype, prior exposure to antiretroviral agents other than INIs, and duration of HIV infection on INI variability. Patients and methods Study population Plasma samples from HIV-1-infected individuals on regular follow-up at Hospital Carlos III (Madrid, Spain) and Royal Free Hampstead NHS Trust 3 p15 protease p66, p51 retrotranscriptase NTD aa 50 pol & UCL University College London Medical School (London, UK) were selected. All patients were naive to INIs. Integrase amplification and sequencing RNA was extracted from plasma using the automatic extractor EasyMag (Biomerieux, Madrid, Spain), which isolates nucleic acids based on the Boom methodology. 14 The integrase gene was amplified using an in-house PCR protocol designed on the basis of a well conserved integrase region in order to detect all HIV-1 variants. Briefly, an RT PCR was carried out using as primers Forward INtegrase EXternal (FINEX) (5 0 AGT GCT GGA ATC AGG AAA GTA C3 0 ) and Reverse INtegrase EXternal (RINEX) (5 0 TAA TCC TCA TCC TGT CTA CTT GC3 0 ). The reverse transcription step was carried out at 488C for 45 min, and PCR involved 35 repeated cycles (948C for 45 s, 578C for 45 s and 728C for 1 min) followed by incubation at 728C for 7 min. Then, a nested PCR was performed using as primers Forward INtegrase INternal (FININ) (5 0 TAG ATG GAA TAG ATA AGG CCC3 0 ) and the Reverse INtegrase INternal (RININ) (5 0 ATC ACC TGC CAT CTG TTT TCC3 0 ). PCR involved 35 repeated cycles (948C for 45 s, 578C for 45 s and 728C for 1 min) followed by incubation at 728C for 7 min. After checking the presence of amplicons with electrophoresis on an agarose gel, purification using Montage w PCR Centrifugal Filter Devices (Montáge, Madrid, Spain) was carried out. The integrase gene was then p15 RNase CCD P31 integrase aa 212 CTD 5 aa 288 H H C C D D E T L L E T F E G Y S Q V S N E G S D R I/A V/I M Q A Y K/A A/S/C R/H/C G K/R/H I Y H/S Q K/R R N K Figure 1. HIV-1 integrase gene and positions associated with resistance to integrase inhibitors. aa, amino acid; NTD, N-terminal domain; CTD, C-terminal domain; CCD, catalytic core domain. 2of7
3 Integrase variability in HIV non-b subtypes JAC sequenced using a cycle sequencing reaction with the Rhodamine terminator kit (Applied Biosystems, Foster City, CA, USA), and a set of four primers was used for complete coverage of both strands of the integrase region. Primers were as follows: two forward primers FININ (5 0 TAG ATG GAA TAG ATA AGG CCC3 0 ), FINMED (5 0 TAC AAT CCC CAA AGT C3 0 ), and two reverse primers RINEX (5 0 ATC ACC TGC CAT CTG TTT TCC3 0 ) and RINMED (5 0 ACT TTG GGG ATT GTA G3 0 ). Sequencing was carried out with the automated sequencer ABI Prism 3100 and integrase sequences were analysed using SeqScape v2.5 (Applied Biosystems). HIV-1 subtyping Phylogenetic and molecular evolutionary analyses were conducted using the MEGA v4 program. 15 All integrase sequences were first aligned together with reference subtype sequences available at the Los Alamos Database using ClustalX. A total of 56 reference sequences were considered, including 9 subtypes and 42 circulating recombinant forms (CRFs) of HIV-1 group M and one from HIV-1 group O, which was considered as an outgroup. INI resistance-associated mutations According to information derived from in vitro studies and clinical trials, to date 39 mutations at 29 positions within the integrase gene may influence INI susceptibility. 16,17 They are the following: H51Y, T66I, V72I, L74M/A/I, E92Q, T97A, F121Y, T125K, A128T, E138K/A, G140S/A, Y143R/C, Q146K/R, S147G, Q148K/R/H, V151I, S153A/Y, M154I, N155H/S, K156N, E157Q, K160D, G163R, V165I, V201I, I203M, T206S, S230M and V249I The major INI RAMs 16 are highlighted in bold in Figure 1. The prevalence of these mutations was examined in the whole study population. Additionally, the variability at the DDE and HHCC motifs, and the overall heterogeneity at the CCD was investigated. Results were stratified by HIV clade, prior antiretroviral exposure and duration of HIV infection. In addition, integrase sequence variability was examined longitudinally in plasma specimens collected from a subset of HIV-1-infected patients who began antiretroviral therapy and experienced virological failure on drugs other than INIs. Statistical analysis The main characteristics of the study population were reported as numbers, percentages or mean values. Fisher s exact test was used to compare the proportion of patients with distinct integrase amino acid changes. All statistical analyses were performed using SPSS v15.0 software (SPSS Inc., Chicago, IL, USA). Statistical significance was considered for two-tailed P values,0.05. Results Study population A total of 418 integrase sequences were obtained. Of these 294 (70.3%) were from patients infected with HIV-1 clade B and 124 (29.7%) from subjects infected with non-b variants. Seven different HIV-1 subtypes and 12 CRFs were represented in the latter group, the most prevalent being CRF02_AG, C, A, D and F. The main characteristics of the study population are recorded in Table 1. A total of 249 (60%) individuals were antiretroviral naive, including 137 who had been infected with HIV-1 within the last 12 months (recent seroconverters). Table 1. Main characteristics of the study population (n¼418) Prevalence of INI resistance-associated mutations The integrase sequences harboured between 0 and 5 INI RAMs, with a median number of 2 [interquartile range (IQR) 1 2]. A total of 33 (7.9%) integrase sequences did not show any resistance change, whereas 143 (34.2%) harboured one, 148 (35.4%) harboured two, 78 (18.7%) harboured three, 15 (3.6%) harboured four, and 1 (0.2%) harboured five INI RAMs. Of the 39 recognized INI RAMs, 22 did not appear in any sequence (Figure 2), comprising H51Y, E92Q, F121Y, T125K, A128T, E138K/A, G140S/A, Y143R/C, Q146R/K, S147G, Q148K/R/ H, S153A/Y, N155H/S and K160D, and including the major RAMs E92Q, Y143R, Q148K/H/R and N155H ,16 In contrast, the most frequent INI RAMs were V72I (63.9%), V201I (54.8%), T206S (25.4%), I203M (9.8%), K156N (7.4%) and L74M/A/I (6.9%), all of which are considered as minor (accessory or compensatory). Other mutations appeared less frequently, such as V165I (2.4%), V151I (2.2%), M154I (1%) or E157Q (1%). No. Percentage Male gender White ethnicity Risk group MSM IDU heterosexual others Antiretroviral naive Recent HIV seroconverters HIV-1 non-b subtypes A C D F G H J CRF01_AE CRF02_AG CRF CRF CRF CRF CRF12_BF CRF CRF14_BG CRF CRF CRF MSM, men who have sex with men; IDU, intravenous drug users. 3of7
4 Garrido et al. T66I V72I L74MAI T97A V151I M154I K156N E157Q G163R V165I V201I I203M T206S S230R V249I other subtypes Percentage of isolates with the change Figure 2. Prevalence of changes at the integrase gene associated with resistance to integrase inhibitors across HIV-1 subtypes. The following mutations were not found in any sample: H51Y, E92Q, F121Y, T125K, A128T, E138KA, G140SA, Y143RC, Q146KR, S147G, Q148KRH, S153AY, N155HS, K160D. CRF02 F D C A B Distribution of INI resistance-associated mutations Significant differences were found in the prevalence of INI RAMs when comparing clade B and non-b viruses, with a mean number of mutations of 1.59 and 2.17, respectively (P, 0.001). Similarly, antiretroviral-naive patients showed a significantly lower mean number of INI RAMs than antiretroviral-experienced patients (1.68 and 1.89; P¼0.034). Patients with recent infection showed fewer INI RAMs than chronically infected subjects (1.61 versus 1.86; P¼0.015). Nevertheless, the only clinically relevant difference was for HIV-1 clades (see Table 2). Several amino acid changes occurred significantly more commonly in non-b subtypes than in subtype B, including L74M/A/I (P¼0.001), T97A (P¼0.007), V165I (P, 0.001), V201I (P, 0.001) and T206S (P, 0.001). Conversely, K156N (P, 0.001) and I203M (P¼0.001) were more common in clade B than in non-b viruses. Comparing antiretroviral-naive with antiretroviral-experienced patients, the only significant differences were found for T97A (P¼0.025) and I203M (P¼0.012), which were more common in the latter. There were no significant differences in the prevalence of any INI RAMs when comparing recent seroconverters and chronically infected individuals. Variability at the integrase CCD A total of 163 amino acids (residues ) constitute the CCD of the HIV-1 integrase. A position was considered as polymorphic 4of7
5 Integrase variability in HIV non-b subtypes JAC Table 2. Rate of integrase inhibitor (INI) resistance-associated mutations in the study population, according to HIV subtype and antiretroviral experience Total B subtype Non-B subtypes Antiretroviral naive Antiretroviral experienced Patients (number) Number of INI resistance-associated mutations mean (SD) 1.76 (0.97) 1.59 (0.88) 2.17 (1.06) 1.68 (1.00) 1.89 (0.92) Number of polymorphic positions within the CCD mean (SD) 9.89 (5.05) 8.75 (4.58) (5.11) 9.82 (5.34) 9.9 (4.62) INI resistance-associated mutations T66I, G163R, S230R, V249I 1 (0.2%) V72I 267 (63.9%) 189 (64.3%) 78 (62.9%) 153 (61.4%) 114 (68.7%) L74A/M/I 29 (6.9%) 12 (4.1%) 17 (13.7%) 18 (7.2%) 10 (6%) T97A 4 (1%) 0 4 (3.2%) 0 4 (2.4%) V151I 9 (2.2%) 9 (3.1%) 0 6 (2.4%) 3 (1.8%) M154I 4 (1%) 3 (1%) 1 (0.8%) 2 (0.8%) 2 (1.2%) K156N 31 (7.4%) 31 (10.5%) 0 17 (6.8%) 13 (7.8%) E157Q 4 (1%) 3 (1%) 1 (0.8%) 4 (1.6%) 0 V165I 10 (2.4%) 1 (0.3%) 9 (7.3%) 6 (2.4%) 4 (2.4%) V201I 229 (54.8%) 130 (44.2%) 99 (78.8%) 138 (55.4%) 89 (53.6%) I203M 41 (9.8%) 36 (12.2%) 5 (4%) 17 (6.8%) 24 (14.5%) T206S 106 (25.4%) 54 (18.4%) 52 (41.9%) 57 (22.9%) 49 (29.5%) Bold text indicates significant differences between groups (P,0.05). when amino acid changes were observed in.1% of tested sequences. 17 Given that 418 sequences were analysed in our study, changes should be found in at least 5 samples. A total of 73 (44.7%) positions within the CCD were conserved, while the remaining 90 were polymorphic. The median number of polymorphic changes in the study population was 9 (IQR 7 12). DDE and HHCC motifs were relatively conserved, with polymorphic changes in 41 (9.8%) sequences, of which 31 (7.4%) occurred within the DDE triad and 10 (2.3%) within the HHCC motif. The number of integrase polymorphic sites was higher in non-b clades compared with subtype B. The mean number was 12.6 and 8.75, respectively (P, 0.001). No significant differences were found when comparing antiretroviral-experienced and drug-naive subjects (9.96 versus 9.82; P¼0.776), or recent seroconverters and chronically infected individuals (10.02 versus 9.82; P¼0.742). Moreover, the rate of polymorphisms was not influenced by plasma HIV RNA values (data not shown). Longitudinal analysis A total of 34 plasma samples were collected longitudinally from 17 patients before starting antiretroviral therapy and at subsequent virological failure. There was no evidence for selection of INI RAMs comparing paired samples in 13 out of 17 patients. However, four individuals selected one minor INI RAM upon failure under antiretroviral drugs other than INI. The selected change differed in each case (data not shown). Nine subjects showed variability within the CCD when comparing baseline and failing specimens. Differences ranged from 0.5 to 6 changes, considering 0.5 when only a mixture was seen either at baseline or at failure. The remaining eight patients did not show any change at the CCD motif. Discussion The introduction of INIs has represented a major breakthrough in the treatment of antiretroviral-experienced HIV-1 patients. In particular, raltegravir has shown high antiviral potency and good tolerability, allowing the virological rescue of many patients who had failed other antiretroviral drug families Nevertheless, as with other antiretroviral agents, selection of drug resistance represents an important drawback in patients treated with INIs ,21 Although a few key resistance mutations in the integrase have been well characterized as producing loss of susceptibility to INIs, 11,16 other multiple changes seem to modulate the degree of susceptibility to these compounds. 17 In this study, the variability within the integrase gene at positions associated with resistance to INIs was examined in a large group of HIV-1 patients, with the specific aim of analysing any relevant impact of HIV-1 subtypes, prior exposure to antiretroviral drugs other than INIs, and duration of HIV-1 infection. A total of 418 integrase sequences belonging to patients infected with 20 different HIV-1 clades were analysed. Interestingly, primary INI RAMs were not seen in INI-naive individuals, regardless of exposure to other antiretroviral agents, duration of HIV-1 disease or infection with diverse non-b subtypes. However, naturally occurring polymorphisms, which could influence INI susceptibility and the genetic barrier to resistance, were found to be relatively common, particularly in subjects infected with non-b subtypes. Similar results have recently been found by others. In contrast, the prevalence of these polymorphisms appeared to be only marginally influenced by prior antiretroviral exposure and/or the duration of HIV-1 infection. The absence of detectable major INI RAMs in our study is consistent with findings from others 22,23 and strongly argues in favour of a wide activity of INIs across HIV-1 subtypes, and 5of7
6 Garrido et al. regardless of exposure to other antiretroviral drugs or duration of HIV-1 infection. This observation is in contrast to sporadic reports of primary INI RAMs, including major (E92K) and minor (E157Q, G140S) changes, among INI-naive individuals Moreover, our findings indirectly support major INI RAMs significantly impairing viral fitness. 27,28 The recognition of a substantial variability within the integrase at secondary resistance positions which modulate the susceptibility to INIs, especially in non-b subtypes, may require further investigation. Prior studies examining in vitro the susceptibility to raltegravir and elvitegravir with large panels of clinical isolates, representing multiple HIV-1 subtypes with several polymorphisms, have concluded that fold changes in the IC 50 are almost always below the biological threshold Moreover, the more distant HIV-1 group O and HIV-2 viruses, which show 40% heterogeneity in the integrase gene compared with HIV-1 group M viruses, 32,33 have been shown to be susceptible to raltegravir inhibition. 24,34 37 It should be highlighted that the catalytic triad DDE and the HHCC and RKK motifs are fully conserved across all HIV variants. 17,38 Altogether, these data suggest that secondary INI RAMs per se do not compromise INI activity. Nevertheless, at this time it is unclear whether they may influence the genetic barrier to resistance or drive the selection of specific INI resistance pathways. In this regard, the presence of the V165I polymorphism might favour the selection of N155H upon raltegravir failure, while the presence of T97A could favour selection of Y143R. 39 Mutation E157Q was found in four sequences (1%) in our study, belonging to three subtype B viruses and one CRF06 virus. Although E157Q has been reported to cause resistance to raltegravir by itself, besides dramatically reducing the enzymatic activity of the integrase, 13 in vitro studies have demonstrated only a minimal impact of this change on phenotypic susceptibility to INIs (fold change of 1.14). 28 Finally, the results of our longitudinal study show that antiretroviral treatment with drugs other than INIs does not result in the selection of integrase changes which may significantly influence the susceptibility to INIs. Similar results have recently been found by others 27 and argue against previous reports suggesting that selection of resistance mutations at the protease and/or reverse transcriptase might condition changes at the integrase which subsequently might influence INI susceptibility. 38 In summary, major INI resistance-associated mutations for raltegravir and elvitegravir are not seen in HIV-infected persons naive for INIs, regardless of exposure to other antiretroviral agents. However, naturally occurring polymorphisms at integrase positions which may influence the genetic barrier and/or drive specific INI resistance pathways are common, especially in subjects infected with non-b subtypes. Funding This work was supported by grants from Fundación Investigación y Educación en SIDA (IES), Red de Investigación en SIDA (RIS ISCIII-RETIC RD06/006), Fondo de Investigación Sanitaria (FIS, projects: PI06/1826; CP06/0284; CP08/0214), NEAT and Agencia Laín Entralgo. Transparency declarations None to declare. References 1 Sarafianos S, Marchand B, Das K et al. Structure and function of HIV-1 reverse transcriptase: molecular mechanisms of polymerization and inhibition. J Mol Biol 2009; 385: Deeks S, Kar S, Gubernick S et al. Raltegravir. Nature Rev 2008; 7: Jegede O, Babu J, Di Santo R et al. HIV-1 integrase inhibitors: from basic research to clinical implications. AIDS Rev 2008; 10: Dyda F, Hickman A, Jenkins T et al. Crystal structure of the catalytic domain of HIV-1 integrase: similarity to other polynucleotidyl transferases. Science 1994; 266: Brown P. Integration of retroviral DNA. Curr Top Microbiol Immunol 1990; 157: Hazuda D, Felock P, Witmer M et al. Inhibitors of strand transfer that prevent integration and inhibit HIV-1 replication in cells. Science 2000; 287: Grant P, Zolopa A. Integrase inhibitors: a clinical review of raltegravir and elvitegravir. J HIV Ther 2008; 13: Marinello J, Marchand C, Mott B et al. Comparison of raltegravir and elvitegravir on HIV-1 integrase catalytic reactions and on a series of drug-resistant integrase mutants. Biochemistry 2008; 47: Goethals O, Clayton R, Van Ginderen M et al. Resistance mutations in HIV type 1 integrase selected with elvitegravir confer reduced susceptibility to a wide range of integrase inhibitors. J Virol 2008; 82: Shimura K, Kodama E, Sakagami Y et al. Broad antiretroviral activity and resistance profile of the novel HIV integrase inhibitor elvitegravir (JTK-303/GS-9137). J Virol 2008; 82: Cooper D, Steigbigel R, Gatell J et al. Subgroup and resistance analyses of raltegravir for resistant HIV-1 infection. N Engl J Med 2008; 359: Charpentier C, Karmochkine M, Laureillard D et al. Drug resistance profiles for the HIV integrase gene in patients failing raltegravir salvage therapy. HIV Med 2008; 9: Malet I, Delelis O, Valantin M et al. Mutations associated with failure of raltegravir treatment affect integrase sensitivity to the inhibitor in vitro. Antimicrob Agents Chemother 2008; 52: Boom R, Sol C, Salimans M et al. Rapid and simple method for purification of nucleic acids. J Clin Microbiol 1990; 28: Tamura K, Dudley J, Nei M et al. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 2007; 24: Johnson V, Brun-Vezinet F, Clotet B et al. Update of the drug resistance mutations in HIV-1. Top HIV Med 2008; 16: Ceccherini-Silberstein F, Malet I et al. Characterization and structural analysis of HIV-1 integrase conservation. AIDS Rev 2009; 11: De Jesus E, Berger D, Markowitz M et al. Antiviral activity, pharmacokinetics, and dose response of the HIV-1 integrase inhibitor GS-9137 (JTK-303) in treatment-naive and treatment-experienced patients. J Acquir Immune Defic Syndr 2006; 43: Grinsztejn B, Nguyen B, Katlama C et al. Safety and efficacy of the HIV-1 integrase inhibitor raltegravir (MK-0518) in treatment-experienced patients with multidrug-resistant virus: a phase II randomised controlled trial. Lancet 2007; 369: of7
7 Integrase variability in HIV non-b subtypes JAC 20 Steigbigel R, Cooper D, Kumar P et al. Raltegravir with optimized background therapy for resistant HIV-1 infection. N Engl J Med 2008; 359: Garrido C, Blanco F, Van Baelen K et al. Raltegravir clinical response and resistance in heavily experienced patients. Rev Antivir Ther 2008; 2: Lataillade M, Chiarella J, Kozal M. Natural polymorphisms of the HIV-1 integrase gene and mutations associated with integrase inhibitor resistance. Antivir Ther 2007; 12: Passaes C, Guimarães ML, Fernandez L et al. Lack of primary mutations associated with integrase inhibitors among HIV-1 subtypes B, C, and F circulating in Brazil. J Acquir Immune Defic Syndr 2009; 51: Eshleman S, Hudelson S, Smith P et al. Analysis of pol integrase sequences in diverse HIV type 1 strains using a prototype genotyping assay. AIDS Res Hum Retroviruses 2009; 25: Low A, Prada N, Topper M et al. Natural polymorphisms of HIV type 1 integrase and inherent susceptibilities to a panel of integrase inhibitors. Antimicrob Agents Chemother 2009; 53: Van Hal S, Herring B, Deris Z et al. HIV-1 integrase polymorphisms are associated with prior antiretroviral drug exposure. Retrovirology 2009; 6: Ferns R, Kirk S, Bennett J et al. The dynamics of appearance and disappearance of HIV-1 integrase mutations during and after withdrawal of raltegravir therapy. AIDS 2009; 23: Buzón M, Dalmau J, Puertas M et al. The HIV-1 integrase genotype strongly predicts raltegravir susceptibility but not viral fitness of primary virus isolates. AIDS, in press. 29 Van Baelen K, Van Eygen V, Rondelez E et al. Clade-specific HIV-1 integrase polymorphisms do not reduce raltegravir and elvitegravir phenotypic susceptibility. AIDS 2008; 22: Danovich R, Ke Y, Wan H et al. Raltegravir has similar in vitro antiviral potency, clinical efficacy, and resistance patterns in B subtype and non-b subtype HIV-1. In: Abstracts of the Seventeenth International AIDS Conference, Mexico City, Abstract TUAA0302 (www. aids2008-abstracts.org/aids2008_book_vol1_web.pdf). 31 Rondelez E, Van Baelen K, Ceccherini-Silverstein F et al. Preliminary biological cut-offs for GS-9137 and MK-0518 integrase inhibitors derived from clonal phenotypic analysis. Rev Antivir Ther 2008; 2: Leoz M, Depatureaux A, Vessière A et al. Integrase polymorphisms and HIV-1 group O diversity. AIDS 2008; 22: Xu L, Anderson J, Ferns B et al. Genetic diversity of integrase sequences in antiretroviral treatment-naïve and treatment-experienced HIV type 2 patients. AIDS Res Hum Retroviruses 2008; 24: Damond F, Lariven S, Roquebert B et al. Virological and immunological response to HAART regimen containing integrase inhibitors in HIV-2-infected patients. AIDS 2008; 22: Garrett N, Xu L, Smit E et al. Raltegravir treatment response in an HIV-2 infected patient: a case report. AIDS 2008; 22: Briz V, Garrido C, Poveda E et al. Raltegravir and etravirine are active against HIV type 1 group O. AIDS Res Hum Retroviruses 2009; 25: Salgado M, Toro C, Simón Aet al. Mutation N155H in HIV-2 integrase confers high phenotypic resistance to raltegravir and impairs replication capacity. J Clin Virol 2009; 46: Roquebert B, Damond F, Collin G et al. HIV-2 integrase gene polymorphism and phenotypic susceptibility of HIV-2 clinical isolates to the integrase inhibitors raltegravir and elvitegravir in vitro. J Antimicrob Chemother 2008; 62: Ceccherini-Silberstein F, Armenia D, D Arrigo R et al. Virological response and resistance in multi-experienced patients treated with raltegravir. Antivir Ther 2008; 13 Suppl 3: A20. 7of7
Resistance to inhibitors of the human immunodeficiency virus type 1 integration
 Resistance to inhibitors of the human immunodeficiency virus type 1 integration REVIEW ARTICLE ABSTRACT This review will summarize the role of integrase in HIV-1 infection, the mechanism of integrase inhibitors
Resistance to inhibitors of the human immunodeficiency virus type 1 integration REVIEW ARTICLE ABSTRACT This review will summarize the role of integrase in HIV-1 infection, the mechanism of integrase inhibitors
Short communication Genetic barriers for integrase inhibitor drug resistance in HIV type-1 B and CRF02_AG subtypes
 Antiviral Therapy 14:123 129 Short communication Genetic barriers for integrase inhibitor drug resistance in HIV type-1 B and CRF2_AG subtypes Almoustapha-Issiaka Maïga 1, Isabelle Malet 1 *, Cathia Soulie
Antiviral Therapy 14:123 129 Short communication Genetic barriers for integrase inhibitor drug resistance in HIV type-1 B and CRF2_AG subtypes Almoustapha-Issiaka Maïga 1, Isabelle Malet 1 *, Cathia Soulie
JCM (Revised version, June 23 th 2011)
 JCM Accepts, published online ahead of print on 6 July 2011 J. Clin. Microbiol. doi:10.1128/jcm.00908-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
JCM Accepts, published online ahead of print on 6 July 2011 J. Clin. Microbiol. doi:10.1128/jcm.00908-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Resistance to Integrase Strand Transfer Inhibitors
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
Integrase strand-transfer inhibitors primary resistance in patients with acute/recent HIV infection
 Integrase strand-transfer inhibitors primary resistance in patients with acute/recent HIV infection Juan Ambrosioni, David Nicolás, Christian Manzardo, Fernando Agüero, José Luis Blanco, Maria del Mar
Integrase strand-transfer inhibitors primary resistance in patients with acute/recent HIV infection Juan Ambrosioni, David Nicolás, Christian Manzardo, Fernando Agüero, José Luis Blanco, Maria del Mar
HIV-1 Subtypes: An Overview. Anna Maria Geretti Royal Free Hospital
 HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
HIV-1 Subtypes: An Overview Anna Maria Geretti Royal Free Hospital Group M Subtypes A (1, 2, 3) B C D F (1, 2) G H J K Mechanisms of HIV-1 genetic diversification Point mutations RT error rate: ~1 per
Supplemental Data. Shin et al. Plant Cell. (2012) /tpc YFP N
 MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
High Failure Rate of the ViroSeq HIV-1 Genotyping System for Drug Resistance Testing in Cameroon, a Country with Broad HIV-1 Genetic Diversity
 JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
JOURNAL OF CLINICAL MICROBIOLOGY, Apr. 2011, p. 1635 1641 Vol. 49, No. 4 0095-1137/11/$12.00 doi:10.1128/jcm.01478-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. High Failure
Short communication Natural polymorphism of the HIV-1 integrase gene and mutations associated with integrase inhibitor resistance
 Short communication Natural polymorphism of the HIV-1 integrase gene and mutations associated with integrase inhibitor resistance Max Lataillade 1 *, Jennifer Chiarella 1 and Michael J Kozal 1,2 1 Yale
Short communication Natural polymorphism of the HIV-1 integrase gene and mutations associated with integrase inhibitor resistance Max Lataillade 1 *, Jennifer Chiarella 1 and Michael J Kozal 1,2 1 Yale
Transmission of integrase resistance HIV
 Transmission of integrase resistance HIV Charles Boucher, MD, PhD Clinical Virology, Dept. Viroscience, Erasmus Medical Center, Erasmus Universiy, The Netherlands Major resistance mutations (Stanford)
Transmission of integrase resistance HIV Charles Boucher, MD, PhD Clinical Virology, Dept. Viroscience, Erasmus Medical Center, Erasmus Universiy, The Netherlands Major resistance mutations (Stanford)
It takes more than just a single target
 It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
The HIV life cycle. integration. virus production. entry. transcription. reverse transcription. nuclear import
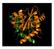 The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
Characterizing intra-host influenza virus populations to predict emergence
 Characterizing intra-host influenza virus populations to predict emergence June 12, 2012 Forum on Microbial Threats Washington, DC Elodie Ghedin Center for Vaccine Research Dept. Computational & Systems
Characterizing intra-host influenza virus populations to predict emergence June 12, 2012 Forum on Microbial Threats Washington, DC Elodie Ghedin Center for Vaccine Research Dept. Computational & Systems
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Second-Line Therapy NORTHWEST AIDS EDUCATION AND TRAINING CENTER
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Second-Line Therapy David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases University of Washington Presentation
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
Drug Resistance Mutations in HIV-Infected Patients Failing Tipranavir and Darunavir in the Spanish Drug Resistance Database
 AAC Accepts, published online ahead of print on 17 May 2010 Antimicrob. Agents Chemother. doi:10.1128/aac.00160-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions.
AAC Accepts, published online ahead of print on 17 May 2010 Antimicrob. Agents Chemother. doi:10.1128/aac.00160-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions.
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
ORIGINAL ARTICLE /j x. Brescia, Italy
 ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
ORIGINAL ARTICLE 10.1111/j.1469-0691.2004.00938.x Prevalence of drug resistance and newly recognised treatment-related substitutions in the HIV-1 reverse transcriptase and protease genes from HIV-positive
Mutations to Integrase Inhibitors in real life
 Mutations to Integrase Inhibitors in real life González-Domenech CM, Viciana I, Sena Corrales G, Delgado M, De la Torre J, Torres Tortosa M, Téllez F, Jarilla F, Clavijo E, Santos J BACKGROUND INSTI Block
Mutations to Integrase Inhibitors in real life González-Domenech CM, Viciana I, Sena Corrales G, Delgado M, De la Torre J, Torres Tortosa M, Téllez F, Jarilla F, Clavijo E, Santos J BACKGROUND INSTI Block
Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosure The speaker serves as a consultant to, and has received
Does Resistance Still Matter? Daniel R. Kuritzkes, M.D. Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Disclosure The speaker serves as a consultant to, and has received
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication
 Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Introduction to the Impact of Resistance in Hepatitis C
 Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
Introduction to the Impact of Resistance in Hepatitis C Sponsored by AbbVie 2/1/2017 Presented by Sammy Saab, MD, MPH, FACG, AGAF, FAASLD February 1 st, 2017 1 AbbVie disclosures This is an Abbvie sponsored
INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING
 INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING Dr. Danni Kirwan ID/Microbiology SpR St. George s Hospital, London ARV initiation in treatment-naïve patients BHIVA,
INTEGRASE INHIBITOR (INI) RESISTANCE IN HIV- POSITIVE PATIENTS UNDERGOING ROUTINE TESTING Dr. Danni Kirwan ID/Microbiology SpR St. George s Hospital, London ARV initiation in treatment-naïve patients BHIVA,
Introduction to HIV Drug Resistance. Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School
 Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
Introduction to HIV Drug Resistance Kevin L. Ard, MD, MPH Massachusetts General Hospital Harvard Medical School Objectives 1. Describe the epidemiology of HIV drug resistance in sub-saharan Africa. 2.
The E138A substitution in HIV-1 reverse transcriptase decreases in vitro. susceptibility to emtricitabine as indicated by competitive fitness assays
 AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
AAC Accepts, published online ahead of print on 13 January 2014 Antimicrob. Agents Chemother. doi:10.1128/aac.02114-13 Copyright 2014, American Society for Microbiology. All Rights Reserved. 1 2 The E138A
A smart acid nanosystem for ultrasensitive. live cell mrna imaging by the target-triggered intracellular self-assembly
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
History (August 2010) Therapy for Experienced Patients. History (September 2010) History (November 2010) 12/2/11
 (August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
(August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at
 Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
Supplementary Figure 1. ROS induces rapid Sod1 nuclear localization in a dosagedependent manner. WT yeast cells (SZy1051) were treated with 4NQO at different concentrations for 30 min and analyzed for
HIV Drug Resistance: An Overview
 Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
Human Journals Review Article October 2015 Vol.:1, Issue:1 All rights are reserved by Suraj Narayan Mali et al. HIV Drug Resistance: An Overview Keywords: HIV drug resistance mechanism, Antiretroviral
Update on HIV Drug Resistance. Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School
 Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
Update on HIV Drug Resistance Daniel R. Kuritzkes, MD Division of Infectious Diseases Brigham and Women s Hospital Harvard Medical School Learning Objectives Upon completion of this presentation, learners
Clinical utility of NGS for the detection of HIV and HCV resistance
 18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
18 th Annual Resistance and Antiviral Therapy Meeting v Professor Janke Schinkel Academic Medical Centre, Amsterdam, The Netherlands Thursday 18 September 2014, Royal College of Physicians, London Clinical
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762)
 Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
Resistance Post Week 48 in ART-Experienced, Integrase Inhibitor-Naïve Subjects with Dolutegravir (DTG) vs. Raltegravir (RAL) in SAILING (ING111762) MR Underwood 1, F DeAnda 1, D Dorey 2, K Hightower 1,
Supplementary Materials
 Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
Pointer Elvitegravir: a new HIV integrase inhibitor
 Antiviral Chemistry & Chemotherapy 2009 20: 79 85 (doi: 10.3851/IMP1397) Pointer Elvitegravir: a new IV integrase inhibitor Kazuya Shimura 1 and Eiichi Kodama 1,2 * 1 Laboratory of Virus Control, Institute
Antiviral Chemistry & Chemotherapy 2009 20: 79 85 (doi: 10.3851/IMP1397) Pointer Elvitegravir: a new IV integrase inhibitor Kazuya Shimura 1 and Eiichi Kodama 1,2 * 1 Laboratory of Virus Control, Institute
Conference Reports for NATAP. S/GSK Integrase Inhibitor Resistance Profile
 Conference Reports for NATAP EACS - 12th European AIDS Conference November 11-14, 2009 Cologne, Germany Back S/GSK1349572 Integrase Inhibitor Resistance Profile Reported by Jules Levin EACS Oct 31-Nov
Conference Reports for NATAP EACS - 12th European AIDS Conference November 11-14, 2009 Cologne, Germany Back S/GSK1349572 Integrase Inhibitor Resistance Profile Reported by Jules Levin EACS Oct 31-Nov
Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review
 pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
pissn 2349-2910 eissn 2395-0684 REVIEW Reverse transcriptase and protease inhibitor resistant mutations in art treatment naïve and treated hiv-1 infected children in India A Short Review Dinesh Bure, Department
Evaluation and Management of Virologic Failure
 National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
National HIV Curriculum PDF created November 3, 2018, 12:26 am Evaluation and Management of Virologic Failure This is a PDF version of the following document: Section 1: Antiretroviral Therapy Topic 5:
Micropathology Ltd. University of Warwick Science Park, Venture Centre, Sir William Lyons Road, Coventry CV4 7EZ
 www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53
 1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
1 2 3 4 5 6 7 8 9 10 Supplementary Figure 1. Induction of p53 LOH by MADM. a) Primary cultures derived from the pancreas of an 11-week-old Pdx1-Cre; K-MADM-p53 mouse revealed increased p53 KO/KO (green,
Resistance Characteristics of Integrase Inhibitors
 Resistance Characteristics of Integrase Inhibitors Madrid, November 2016 Jonathan M Schapiro, MD National Hemophilia Center, Israel Stanford University School of Medicine, USA Disclaimer Presentation includes
Resistance Characteristics of Integrase Inhibitors Madrid, November 2016 Jonathan M Schapiro, MD National Hemophilia Center, Israel Stanford University School of Medicine, USA Disclaimer Presentation includes
Supplementary Figure 1 a
 Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Supplementary Figure a Normalized expression/tbp (A.U.).6... Trip-br transcripts Trans Trans Trans b..5. Trip-br Ctrl LPS Normalized expression/tbp (A.U.) c Trip-br transcripts. adipocytes.... Trans Trans
Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal Introduction
 Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
Disclosure statement: Dr. Santoro reports personal fees from ViiV Healthcare, Gilead and JANSSEN Cilag Round table discussion Patients with multiresistant virus : A limited number, but a remarkable deal
HIV and drug resistance Simon Collins UK-CAB 1 May 2009
 HIV and drug resistance Simon Collins UK-CAB 1 May 2009 slides: thanks to Prof Clive Loveday, Intl. Clinical Virology Centre www.icvc.org.uk Tip of the iceberg = HIV result, CD4, VL Introduction: resistance
HIV and drug resistance Simon Collins UK-CAB 1 May 2009 slides: thanks to Prof Clive Loveday, Intl. Clinical Virology Centre www.icvc.org.uk Tip of the iceberg = HIV result, CD4, VL Introduction: resistance
Prevalence of Transmitted Drug Resistance Mutations among Naive HIV-infected patients ( ) in Northwest Spain
 Prevalence of Transmitted Drug Resistance Mutations among Naive HIV-infected patients (2014-2016) in Northwest Spain Berta Pernas 1, Andrés Tabernilla 1, Marta Grandal 1, Angelina Cañizares 2, Sofía Ortún
Prevalence of Transmitted Drug Resistance Mutations among Naive HIV-infected patients (2014-2016) in Northwest Spain Berta Pernas 1, Andrés Tabernilla 1, Marta Grandal 1, Angelina Cañizares 2, Sofía Ortún
Title. HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in
 Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
Title HIV-1 Protease and Reverse Transcriptase Mutations: Correlations with Antiretroviral Therapy in Subtype B Isolates and Implications for Drug-Resistance Surveillance October 13, 2004 Authors SY Rhee
Case Study. Dr Sarah Sasson Immunopathology Registrar. HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital.
 Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
Case Study Dr Sarah Sasson Immunopathology Registrar HIV, Immunology and Infectious Diseases Department and SydPath, St Vincent's Hospital Case 1: Case 1: 45F in Cameroon Cameroon HIV+ Presents with cutaneous
NNRTI Resistance NORTHWEST AIDS EDUCATION AND TRAINING CENTER
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER NNRTI Resistance David H. Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington Last Updated:
NORTHWEST AIDS EDUCATION AND TRAINING CENTER NNRTI Resistance David H. Spach, MD Principal Investigator, NW AETC Professor of Medicine, Division of Infectious Diseases University of Washington Last Updated:
Disclosures. Introduction to ARV Drug Resistance New Clinicians Workshop 12/9/16. Introduction. ARS Question
 Disclosures Introduction to ARV Drug Resistance New Clinicians Workshop I have no disclosures Susa Coffey, MD Division of HIV, ID and Global Medicine ARS Question Which resistance test do you order for
Disclosures Introduction to ARV Drug Resistance New Clinicians Workshop I have no disclosures Susa Coffey, MD Division of HIV, ID and Global Medicine ARS Question Which resistance test do you order for
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards
 Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
Toluidin-Staining of mast cells Ear tissue was fixed with Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) overnight at 4 C, afterwards incubated in 100 % ethanol overnight at 4 C and embedded in
2 nd Line Treatment and Resistance. Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012
 2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
2 nd Line Treatment and Resistance Dr Rohit Talwani & Dr Dave Riedel 12 th June 2012 Overview Basics of Resistance Treatment failure Strategies to manage treatment failure Mutation Definition: A change
Table S1. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9. Assay (s) Target Name Sequence (5 3 ) Comments
 SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide polymorphisms at thymidine analogue mutation sites
 Journal of Antimicrobial Chemotherapy Advance Access published June 7, 2013 J Antimicrob Chemother doi:10.1093/jac/dkt204 The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide
Journal of Antimicrobial Chemotherapy Advance Access published June 7, 2013 J Antimicrob Chemother doi:10.1093/jac/dkt204 The preferential selection of K65R in HIV-1 subtype C is attenuated by nucleotide
Diagnostic Methods of HBV and HDV infections
 Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
Low-frequency HIV-1 drug resistance mutations can be clinically significant but must be interpreted with caution
 J Antimicrob Chemother 2010; 65: 1322 1326 doi:10.1093/jac/dkq139 Advance Access publication 13 May 2010 Low-frequency HIV-1 drug resistance mutations can be clinically significant but must be interpreted
J Antimicrob Chemother 2010; 65: 1322 1326 doi:10.1093/jac/dkq139 Advance Access publication 13 May 2010 Low-frequency HIV-1 drug resistance mutations can be clinically significant but must be interpreted
CD31 5'-AGA GAC GGT CTT GTC GCA GT-3' 5 ' -TAC TGG GCT TCG AGA GCA GT-3'
 Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Table S1. The primer sets used for real-time RT-PCR analysis. Gene Forward Reverse VEGF PDGFB TGF-β MCP-1 5'-GTT GCA GCA TGA ATC TGA GG-3' 5'-GGA GAC TCT TCG AGG AGC ACT T-3' 5'-GAA TCA GGC ATC GAG AGA
Citation for published version (APA): Oosterveer, M. H. (2009). Control of metabolic flux by nutrient sensors Groningen: s.n.
 University of Groningen Control of metabolic flux by nutrient sensors Oosterveer, Maaike IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
University of Groningen Control of metabolic flux by nutrient sensors Oosterveer, Maaike IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
Tools to Monitor HIV Infection in 2013 and Beyond.
 Tools to Monitor HIV Infection in 2013 and Beyond. Federico García, fegarcia@ugr.es Servicio de Microbiología Univ. Hospital San Cecilio Granada, Spain Outline Address clinical questions: Ultra sensitive
Tools to Monitor HIV Infection in 2013 and Beyond. Federico García, fegarcia@ugr.es Servicio de Microbiología Univ. Hospital San Cecilio Granada, Spain Outline Address clinical questions: Ultra sensitive
Journal of Microbes and Infection,June 2007,Vol 2,No. 2. (HBsAg)2 , (PCR) 1762/ 1764
 68 2007 6 2 2 Journal of Microbes and Infection,June 2007,Vol 2,No. 2 2 S 1 1 1 2 2 3 1 (HBsAg)2 ( YIC) S 5 30g 60g YIC ( HBV) DNA > 2 log10 e (HBeAg), 6 DNA, 1 YIC 1, (PCR) (0 ) (44 ) HBV DNA S 2, S a
68 2007 6 2 2 Journal of Microbes and Infection,June 2007,Vol 2,No. 2 2 S 1 1 1 2 2 3 1 (HBsAg)2 ( YIC) S 5 30g 60g YIC ( HBV) DNA > 2 log10 e (HBeAg), 6 DNA, 1 YIC 1, (PCR) (0 ) (44 ) HBV DNA S 2, S a
Antiviral Therapy 2016; 21: (doi: /IMP2987)
 Antiviral Therapy 2016; 21:175 180 (doi: 10.3851/IMP2987) Case report Virological failure in two patients with HIV-1 RNA viral loads >1,000,000 copies/ml initiated on elvitegravir/cobicistat/emtricitabine/tenofovir
Antiviral Therapy 2016; 21:175 180 (doi: 10.3851/IMP2987) Case report Virological failure in two patients with HIV-1 RNA viral loads >1,000,000 copies/ml initiated on elvitegravir/cobicistat/emtricitabine/tenofovir
14 TH EUROPEAN HIV & HEPATITIS MEETING Abst#_O_06
 14 TH EUROPEAN HIV & HEPATITIS MEETING 2016 Abst#_O_06 Patients with pre-existent NRTI- and NNRTI-resistance have a higher risk to lose virological suppression under tenofovir/emtricitabine/rilpivirine
14 TH EUROPEAN HIV & HEPATITIS MEETING 2016 Abst#_O_06 Patients with pre-existent NRTI- and NNRTI-resistance have a higher risk to lose virological suppression under tenofovir/emtricitabine/rilpivirine
Supplementary Table 2. Conserved regulatory elements in the promoters of CD36.
 Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
Supplementary Table 1. RT-qPCR primers for CD3, PPARg and CEBP. Assay Forward Primer Reverse Primer 1A CAT TTG TGG CCT TGT GCT CTT TGA TGA GTC ACA GAA AGA ATC AAT TC 1B AGG AAA TGA ACT GAT GAG TCA CAG
VIKING STUDIES Efficacy and safety of dolutegravir in treatment-experienced subjects
 VIKING STUDIES Efficacy and safety of dolutegravir in treatment-experienced subjects IL/DLG/0040/14 June 2014 GSK (Israel) Ltd. Basel 25, Petach Tikva. Tel-03-9297100 Medical information service: il.medinfo@gsk.com
VIKING STUDIES Efficacy and safety of dolutegravir in treatment-experienced subjects IL/DLG/0040/14 June 2014 GSK (Israel) Ltd. Basel 25, Petach Tikva. Tel-03-9297100 Medical information service: il.medinfo@gsk.com
modified dye uptake assay including formazan test EC 90 not tested plaque reduction assay
 Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
HIV-1 Integrase Strand Transfer Inhibitors: Novel Insights into their Mechanism of Action
 CMMETARY SPECIAL ISSUE HIV-1 Integrase Strand Transfer Inhibitors: ovel Insights into their Mechanism of Action Krishan K. Pandey and Duane P. Grandgenett Institute for Molecular Virology, Saint Louis
CMMETARY SPECIAL ISSUE HIV-1 Integrase Strand Transfer Inhibitors: ovel Insights into their Mechanism of Action Krishan K. Pandey and Duane P. Grandgenett Institute for Molecular Virology, Saint Louis
*To whom correspondence should be addressed. This PDF file includes:
 www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
Monitoring for Drug Resistance by Genotyping. Urvi M Parikh, PhD MTN Virology Core Lab
 Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
BIOLOGY 621 Identification of the Snorks
 Name: Date: Block: BIOLOGY 621 Identification of the Snorks INTRODUCTION: In this simulation activity, you will examine the DNA sequence of a fictitious organism - the Snork. Snorks were discovered on
Name: Date: Block: BIOLOGY 621 Identification of the Snorks INTRODUCTION: In this simulation activity, you will examine the DNA sequence of a fictitious organism - the Snork. Snorks were discovered on
Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because
 Supplementary Figure S1 Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because importin-α6 was shown to be testis-specific. Human and chicken importin protein
Supplementary Figure S1 Phylogenetic analysis of human and chicken importins. Only five of six importins were studied because importin-α6 was shown to be testis-specific. Human and chicken importin protein
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most
 Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Antiviral Therapy 2011; 16: (doi: /IMP1851)
 Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
Antiviral Therapy 2011; 16:925 929 (doi: 10.3851/IMP1851) Short communication Prevalence of low-level HIV-1 variants with reverse transcriptase mutation K65R and the effect of antiretroviral drug exposure
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
Drug Resistance to Integrase Strand Transfer Inhibitors in HIV-1 Chilean Patients. Frequency and evolution between the years 2013 and 2016.
 Drug Resistance to Integrase Strand Transfer Inhibitors in HIV-1 Chilean Patients. Frequency and evolution between the years 2013 and 2016. Sidgman F. 1, Valdés F. 1, Ferrer P. 2, Maureira V. 1, Afani
Drug Resistance to Integrase Strand Transfer Inhibitors in HIV-1 Chilean Patients. Frequency and evolution between the years 2013 and 2016. Sidgman F. 1, Valdés F. 1, Ferrer P. 2, Maureira V. 1, Afani
COMPARISON OF HBV RIBONUCLEASE H DOMAIN IN NAÏVE AND DRUG RESISTANT PATIENTS
 HBV RIBONUCLEASE H DOMAIN IN PATIENTS WITH DRUG RESISTANT COMPARISON OF HBV RIBONUCLEASE H DOMAIN IN NAÏVE AND DRUG RESISTANT PATIENTS Surachai Amornsawadwattana, Pattaratida Sa-Nguanmoo, Preeyaporn Vichaiwattana,
HBV RIBONUCLEASE H DOMAIN IN PATIENTS WITH DRUG RESISTANT COMPARISON OF HBV RIBONUCLEASE H DOMAIN IN NAÏVE AND DRUG RESISTANT PATIENTS Surachai Amornsawadwattana, Pattaratida Sa-Nguanmoo, Preeyaporn Vichaiwattana,
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation
 Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
Perspective Resistance and Replication Capacity Assays: Clinical Utility and Interpretation Resistance testing has emerged as an important tool for antiretroviral management. Research continues to refine
Genotypic/phenotypic patterns of HIV-1 integrase resistance to raltegravir
 J Antimicrob Chemother ; : doi:.9/jac/dkp77 Advance publication 7 January Genotypic/phenotypic patterns of HIV- integrase resistance to raltegravir Filippo Canducci *, Maria Chiara Marinozzi, Michela Sampaolo,,
J Antimicrob Chemother ; : doi:.9/jac/dkp77 Advance publication 7 January Genotypic/phenotypic patterns of HIV- integrase resistance to raltegravir Filippo Canducci *, Maria Chiara Marinozzi, Michela Sampaolo,,
HIV-1 Dual Infection and Neurocognitive Impairment
 HIV-1 Dual Infection and Neurocognitive Impairment Gabriel Wagner, MD Assistant Professor of Medicine Infectious Diseases & Global Public Health UC San Diego HIV-Associated End Organ Damage Antiretroviral
HIV-1 Dual Infection and Neurocognitive Impairment Gabriel Wagner, MD Assistant Professor of Medicine Infectious Diseases & Global Public Health UC San Diego HIV-Associated End Organ Damage Antiretroviral
Scottish Medicines Consortium
 Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
Scottish Medicines Consortium raltegravir, 400mg film-coated tablet (Isentress) No. (461/08) Merck, Sharp and Dohme Limited 04 April 2008 The Scottish Medicines Consortium has completed its assessment
Mapping Evolutionary Pathways of HIV-1 Drug Resistance. Christopher Lee, UCLA Dept. of Chemistry & Biochemistry
 Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Resistance profile of the new nucleoside reverse transcriptase inhibitor apricitabine
 Journal of Antimicrobial Chemotherapy Advance Access published December 9, 2009 J Antimicrob Chemother doi:10.1093/jac/dkp422 esistance profile of the new nucleoside reverse transcriptase inhibitor apricitabine
Journal of Antimicrobial Chemotherapy Advance Access published December 9, 2009 J Antimicrob Chemother doi:10.1093/jac/dkp422 esistance profile of the new nucleoside reverse transcriptase inhibitor apricitabine
Original article Universal profiling of HIV-1 pol for genotypic study and resistance analysis across subtypes
 Antiviral Therapy 2011; 16:1267 1275 (doi: 10.3851/IMP1892) Original article Universal profiling of HIV-1 pol for genotypic study and resistance analysis across subtypes Ting Nie 1, Mervi Detorio 1, Raymond
Antiviral Therapy 2011; 16:1267 1275 (doi: 10.3851/IMP1892) Original article Universal profiling of HIV-1 pol for genotypic study and resistance analysis across subtypes Ting Nie 1, Mervi Detorio 1, Raymond
HIV . HIV-1 HIV. . NJplot. Nested RT-PCR . CRF35-AD
 (-) HIV-1 POL HIV.. HIV-1 HIV Nested RT- HIV-1. PTZ-57RT PCR.. ClustalW /. NJplot Nested RT-PCR.. CRF35-AD. CRF35-AD. HIV1 : // : // : : PhD : - PhD - HIV-1 POL. HIV.. HIV-1 PCR CRFs= ) (Criculating Recombinant
(-) HIV-1 POL HIV.. HIV-1 HIV Nested RT- HIV-1. PTZ-57RT PCR.. ClustalW /. NJplot Nested RT-PCR.. CRF35-AD. CRF35-AD. HIV1 : // : // : : PhD : - PhD - HIV-1 POL. HIV.. HIV-1 PCR CRFs= ) (Criculating Recombinant
Cross-clade simultaneous HIV drug resistance genotyping for reverse transcriptase, protease, and integrase inhibitor mutations by Illumina MiSeq
 Dudley et al. Retrovirology ( DOI 10.1186/s12977-014-0122-8 RESEARCH Open Access Cross-clade simultaneous HIV drug resistance genotyping for reverse transcriptase, protease, and integrase inhibitor mutations
Dudley et al. Retrovirology ( DOI 10.1186/s12977-014-0122-8 RESEARCH Open Access Cross-clade simultaneous HIV drug resistance genotyping for reverse transcriptase, protease, and integrase inhibitor mutations
Disclosures. Introduction to ARV Drug Resistance New Clinicians Workshop. Introduction. ARS Question 12/6/2017
 Disclosures Introduction to ARV Drug Resistance New Clinicians Workshop I have no disclosures Susa Coffey, MD Division of HIV, ID and Global Medicine ARS Question Which resistance test do you order for
Disclosures Introduction to ARV Drug Resistance New Clinicians Workshop I have no disclosures Susa Coffey, MD Division of HIV, ID and Global Medicine ARS Question Which resistance test do you order for
Viro-Immunological Response in HIV-1 Infected Patients with Multiple. Treatment Failures Receiving Raltegravir and Optimized Background Therapy,
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 Viro-Immunological Response in HIV-1 Infected Patients with Multiple Treatment Failures Receiving Raltegravir and Optimized Background
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 Viro-Immunological Response in HIV-1 Infected Patients with Multiple Treatment Failures Receiving Raltegravir and Optimized Background
Integrase Strand Transfer Inhibitors on the Horizon
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Integrase Strand Transfer Inhibitors on the Horizon David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, University of Washington Presentation
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Integrase Strand Transfer Inhibitors on the Horizon David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, University of Washington Presentation
Abbreviations: P- paraffin-embedded section; C, cryosection; Bio-SA, biotin-streptavidin-conjugated fluorescein amplification.
 Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
Supplementary Table 1. Sequence of primers for real time PCR. Gene Forward primer Reverse primer S25 5 -GTG GTC CAC ACT ACT CTC TGA GTT TC-3 5 - GAC TTT CCG GCA TCC TTC TTC-3 Mafa cds 5 -CTT CAG CAA GGA
HIV replication and selection of resistance: basic principles
 HIV replication and selection of resistance: basic principles 26th International HIV Drug Resistance and Treatment Strategies Workshop Douglas Richman 6 November 2017 CLINICAL DATA DURING SIXTEEN WEEKS
HIV replication and selection of resistance: basic principles 26th International HIV Drug Resistance and Treatment Strategies Workshop Douglas Richman 6 November 2017 CLINICAL DATA DURING SIXTEEN WEEKS
HIV, HBV, HCV Virology. Anna Maria Geretti Institute of Infection & Global Health University of Liverpool
 HIV, HBV, HCV Virology Anna Maria Geretti Institute of Infection & Global Health University of Liverpool HIV HBV HCV Many similarities Several fundamental differences High-level replication: HIV 10 10,
HIV, HBV, HCV Virology Anna Maria Geretti Institute of Infection & Global Health University of Liverpool HIV HBV HCV Many similarities Several fundamental differences High-level replication: HIV 10 10,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
Supplementary Figures Supplementary Figure 1. H3F3B expression in lung cancer. a. Comparison of H3F3B expression in relapsed and non-relapsed lung cancer patients. b. Prognosis of two groups of lung cancer
ALLINIs modulate the interaction between HIV-1 Rev and integrase
 ALLINIs modulate the interaction between HIV-1 Rev and integrase Date: 13 June 2017 Author Shaakira Abrahams Salerwe Mosebi, Qasim Fish, Raymond Hewer, Maria Papathanasopoulos Outline Background Results
ALLINIs modulate the interaction between HIV-1 Rev and integrase Date: 13 June 2017 Author Shaakira Abrahams Salerwe Mosebi, Qasim Fish, Raymond Hewer, Maria Papathanasopoulos Outline Background Results
Understanding and. Pre-Integration. Post-Integration. By Andrea Low, md and Mark Muesing, phd. Nucleus. Cytoplasm
 Understanding and Inhibiting Integrase in the Treatment of HIV Disease Reprinted from The prn Notebook november 2006 Dr. James F. Braun, Editor-in-Chief Meri D. Pozo, PhD, Managing Editor Published in
Understanding and Inhibiting Integrase in the Treatment of HIV Disease Reprinted from The prn Notebook november 2006 Dr. James F. Braun, Editor-in-Chief Meri D. Pozo, PhD, Managing Editor Published in
PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES
 PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES Mark A. Wainberg McGill University AIDS Centre Jewish General Hospital Montreal, Quebec,
PRINCIPLES and TRENDS in MANAGEMENT of HIV DISEASE: PROBLEMS OF DRUG RESISTANCE in VIRUSES of DIFFERENT SUBTYPES Mark A. Wainberg McGill University AIDS Centre Jewish General Hospital Montreal, Quebec,
Culture Density (OD600) 0.1. Culture Density (OD600) Culture Density (OD600) Culture Density (OD600) Culture Density (OD600)
 A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
